Introduction
Nasopharyngeal carcinoma (NPC) is an elusive
epithelial malignancy that arises in the nasopharynx and is
characterized by significant geographic variations in its
incidence. The analysis of epidemiological trends over the past
decade revealed a gradual decline in the incidence of NPC (1). Nevertheless, in Southeast Asia and
East Asia, particularly in the south of China, the incidence of NPC
remains significantly higher than those in other regions globally
(2), necessitating ongoing
vigilance from the Chinese medical community. Although various
etiological factors and key molecules implicated in the onset and
metastasis of NPC have been identified (3-6),
the current understanding of these factors and their associated
mechanisms is insufficient to completely elucidate the pathogenesis
of NPC. Consequently, conducting further mechanistic investigations
of NPC is an urgent requirement to deepen our understanding of the
disease and support its application in clinical interventions.
Since the identification of ferroptosis as an
emerging form of programmed cell death (PCD) in 2012 (7), it has attracted significant
attention from the scientific community. This form of cell death is
heavily dependent on iron, which causes excessive reactive oxygen
species (ROS) accumulation and, consequently, dangerously high
levels of lipid peroxides (8).
Since its discovery, ferroptosis has been closely associated with
cancer. It was first induced and characterized in human
fibrosarcoma cells expressing oncogenic Ras protein following
treatment with erastin and Ras-selective lethal small molecule 3
(7). Then, a particular type of
PCD was acknowledged to facilitate the elimination of cancer cells,
with evidence demonstrating that induced ferroptosis effectively
inhibits head and neck cancers (9,10), including NPC (11-13). A previous study identified that
ferroptosis is dependent on autophagy and is characterized by
excessive activation of autophagic mechanisms (14). Autophagy-dependent ferroptosis
also serves as a pivotal regulator in the inhibition of cancer
progression. In lung adenocarcinoma (15), pancreatic carcinoma (16) and triple-negative breast cancer
(17), key genes or crucial
bioactive compounds associated with autophagy-dependent ferroptosis
have been identified. Nevertheless, the mechanisms underlying
autophagy-dependent ferroptosis in NPC remain unclear. Therefore,
the present study aimed to identify the genes associated with
autophagy-dependent ferroptosis in NPC and explore the regulatory
mechanisms involved.
The regulatory mechanisms of ferroptosis are
generally diverse and intricate. To date, numerous signaling
pathways that regulate ferroptosis have been identified. Among
these, the nuclear factor-erythroid 2-related factor 2 (Nrf2)/heme
oxygenase (HO)-1/glutathione peroxidase 4 (GPX4) signaling pathway
has attracted considerable research interest. This pathway has been
recognized not only as a potential therapeutic target for malignant
tumors occurring in various body regions, including breast
(18), tongue (19), liver (20) and ovary (21) but also as being linked to various
diseases associated with ferroptosis (22). However, research on the potential
significance of this pathway in NPC remains limited.
The present study aimed to identify the key gene
that regulates NPC and ferroptosis through bioinformatics analysis
and an extensive review of the literature. Furthermore, in
vitro and in vivo methodologies were employed to
investigate the modulatory functions of this key gene in NPC and
mechanisms associated with autophagy-dependent ferroptosis, thereby
elucidating the potential of this gene as a therapeutic target for
NPC.
Materials and methods
Bioinformatics analyses
The GSE12452 dataset, containing 31 NPC samples and
10 controls, was sourced from Gene Expression Omnibus (GEO;
https://www.ncbi.nlm.nih.gov/geo/).
Ferroptosis-related genes (FRGs) were downloaded from the FerrDB
database (http://www.zhounan.org/ferrdb/current/).
Differentially expressed genes (DEGs) between NPC and healthy
controls were identified using the limma package in R with criteria
of |log2FC| >1.5 and adjusted P<0.05 (23). Key genes associated with NPC and
ferroptosis were then determined by intersecting DEGs with
FRGs.
To further verify whether the expression level of
lymphoid-specific helicase (HELLS) is higher in NPC, single-cell
RNA sequencing data was downloaded from the 10X Genomics platform
in the GSE120926 dataset, which includes 16 primary NPC tumors and
7 non-malignant rhinitis biopsies. The Read 10X function from the
Seurat package (version 4.30) was utilized to load the data. Cells
were filtered to retain those containing between 200-6,000 detected
genes and with a mitochondrial gene expression ratio of <10%,
resulting in a final dataset of 101,110 cells. Data normalization
was performed using Log Normalize, followed by identification of
highly variable genes with the Find Variable Features function (n
features=3,000), and the Scale Data function to standardize gene
expression levels within each cell to a mean of 0 and variance of
1, regressing out technical covariates [var. to. regress=c ('n
Count_ RNA', 'percent. Mito')]. Following dimensionality reduction
via principal component analysis, batch effects across different
samples were corrected using the Harmony package. A k-nearest
neighbor graph was constructed with the Find Neighbors function,
and cells were clustered using the Find Clusters function
(resolution=0.1). Finally, cell types were annotated based on
marker genes curated in the Cell Marker 2.0 database.
Transcriptome data of 12 pairs of NPC tissues with
high and low intratumoral bacterial loads were downloaded from the
GSE189642 dataset in the GEO database, with expression values
presented as FPKM. The expression levels of HELLS were extracted
from all samples, and the Wilcoxon rank-sum test was used to
calculate the differences in HELLS expression between the high and
low intratumoral bacterial load groups.
Patients
The present study included five patients diagnosed
with NPC who received treatment at the Department of
Otolaryngology, The Second Hospital of Jilin University, between
March 2023 and March 2024. Inclusion criteria for patient
enrollment were as follows: (i) Confirmed diagnosis of NPC by
pathological examination; (ii) aged 18 years or older; and (iii)
clear consciousness. Exclusion criteria were: (i) Comorbidity with
other major malignant tumors or severe diseases; (ii) presence of
mental or cognitive disorders; and (iii) inability to communicate
normally. Pathological specimens were obtained through endoscopic
nasopharyngeal biopsy for confirming the diagnosis of NPC and
endoscopic resection of lesions to remove localized tumor tissue.
Tumor and paracancerous normal tissue (the distance from the tumor
tissue was 2-3 cm) samples of suitable sizes were gathered
intraoperatively and subsequently preserved at −80°C. Waiver of
informed consent was approved by the Medical Ethics Committee of
The Second Hospital of Jilin University, which also granted ethical
approval for this study (approval no. 2023-301; Changchun, China).
Information of clinical patients (including sex and age) is
presented at Table I.
 | Table IClinicopathological characteristics
of the patients from which the samples were derived, including sex,
pathological type, clinical stage and clinical status. |
Table I
Clinicopathological characteristics
of the patients from which the samples were derived, including sex,
pathological type, clinical stage and clinical status.
| Patients with
NPC | Sex | Age, years | Pathological
type | Clinical stage | Clinical
status | Specimen collection
method |
|---|
| 1 | Male | 52 | Undifferentiated
non-keratinizing carcinoma | III | Initial
treatment | Endoscopic
biopsy |
| 2 | Female | 50 | Undifferentiated
non-keratinizing carcinoma | III | Initial
treatment | Endoscopic
biopsy |
| 3 | Male | 46 | Undifferentiated
non-keratinizing carcinoma | II | Initial
treatment | Endoscopic
biopsy |
| 4 | Male | 61 | Differentiated
non-keratinizing carcinoma | IVA | Recurrence | Endoscopic
biopsy |
| 5 | Female | 58 | Undifferentiated
non-keratinizing carcinoma | II | Initial
treatment | Endoscopic
biopsy |
Cell culture
Human NPC cell lines NPC/HK-1 (cat. no. IM-H533) and
C666-1 (cat. no. IM-H432) and nasopharyngeal epithelial cell line
NP69 (cat. no. IM-H435; all from Xiamen Immocell Biotechnology Co.,
Ltd.) were maintained in a humid and aseptic environment at 37°C
with 5% CO2. NPC/HK-1 and NP69 cell lines were cultured
in RPMI-1640 medium (Corning, Inc.), added with 10% fetal bovine
serum (FBS; Sijiqing; https://www.hzsjq.com/) and penicillin-streptomycin
solution (Biosharp Life Sciences). By contrast, C666-1 cells were
cultivated in DMEM with 15% FBS and a solution containing both
aforementioned antibiotics.
Cell transfection
Cells were plated in a 96-well plate at a uniform
concentration of 3×104 cells per well and cultivated for
24 h. The minimum puromycin concentration that can eliminate cells
was initially determined. Following this, cell transfection was
conducted using Lipofectamine 2000 (Thermo Fisher Scientific,
Inc.). The mixture comprising the transfection reagent and plasmid
DNA (0.3 μg per well, 1 μg/l) was prepared and
subsequently administered to cells. Gene expression was assessed
following a 48-h culture at 37°C with 5% CO2. The short
hairpin (sh) RNA sequences were as follows: sh-HELLS-1:
CAGGAAAGAATTCCTAGAATGTATG; sh-HELLS-2: CCAGGAAAGAATTCCTAGAATGTAT;
and sh-HELLS-3: GCTGCACCAGGAAAGAATTCCTAGA; sh-negative control
(NC): TTCTCCGAACGTGTCACGT.
Cell treatment and group setting
To modulate ferroptosis, cells transfected with
sh-NC and sh-HELLS sequences were treated for 24 h with either 10
μM of erastin (cat. no. HY-15763; MedChemExpress) or a
combination of 10 μM of Erastin and 2 μM
ferrostatin-1 (Fer-1; cat. no. HY-100579; MedChemExpress) (11,24). Furthermore, to investigate
apoptosis-dependent ferroptosis, cells were initially stimulated
with erastin and subsequently treated with either 500 nM rapamycin
(Rapa; cat. no. HY-10219; MedChemExpress) or 5 mM 3-methyladenine
(3-MA; cat. no. HY-19312; MedChemExpress) for 24 h (25,26). To investigate whether
autophagy-dependent ferroptosis was regulated through the
Nrf2/HO-1/GPX4 signaling pathway, cells were initially stimulated
with erastin and subsequently treated with 1 μM ML334 (cat.
no. HY-110258; MedChemExpress) for 24 h.
Cell proliferation assay
Cells were seeded into 96-well plates at a defined
cell density of 5×103 cells per well and cultured at
various time intervals. Upon achieving the defined treatment
period, the medium was removed, and 10 μl per well of
freshly prepared Cell Counting Kit-8 reagent (Kermey; http://www.jacksenbio.com/) was supplied to the cells
and incubated at 37°C for 1 h. The OD value at 450 nm was measured
promptly.
Cell apoptosis assay
Cells were suspended in 1X binding buffer, and
Annexin V-mCherry and GreenNuc™ Caspase-3 Substrate (all three
components of the Caspase-3 Activity and Apoptosis Detection Kit
for Live Cell; cat. no. C1077M; Beyotime Institute of
Biotechnology) were subsequently added. Following gentle mixing,
the mixture was incubated in the dark for 30 min to facilitate
staining. Subsequently, 1X binding buffer was added to halt the
staining process, and the thoroughly mixed cell suspension was
analyzed employing a BD FACSCaliburTM flow cytometer
(Becton, Dickinson and Company). Data analysis was performed with
FlowJo software (version 10.8.1; FlowJo LLC).
Transwell assay
Cell migration and invasion were assessed using the
Transwell assay with an 8-μm pore size. Specifically, in the
migration measurement, cells were inoculated into the upper chamber
at a density of 5×105 cells per well, whereas the lower
chamber was filled with the medium containing 15% FBS. In the
invasion assay, cells were placed in the same part, which was
pretreated with Matrigel (Corning, Inc.) at 37°C for 60 min, and
the lower chamber was filled with the same complete medium.
Following 24-h incubation at 37°C, the cells that stayed in the
upper part were removed. Conversely, the cells that migrated or
invaded through the filter pores were fixed with 100% methanol at
room temperature for 30 min, stained with 0.1% crystal violet (cat.
no. BL802A; Biosharp Life Sciences) at room temperature for 15 min,
and images were captured.
Glutathione assay
Glutathione (GSH) concentrations within the cells
were evaluated using the reduced GSH assay kit (cat. no. A006-2-1;
Nanjing Jiancheng Bioengineering Institute), following the
guidelines provided by the manufacturer.
Malondialdehyde (MDA) assay
The MDA levels were assessed using a lipid
peroxidation MDA assay kit (cat. no. S0131S; Beyotime Institute of
Biotechnology) based on the manufacturer's instructions.
Ferrous iron assay
Cell ferrous iron fluorometric assay kit (cat. no.
E-BC-F101; Elabscience Biotechnology, Inc.) was employed to
determine Fe2+ concentrations.
ROS assay
The immunofluorescence technique facilitates the
direct and visual detection of ROS levels. In this procedure, the
cells were washed twice with serum-free culture medium and
subsequently incubated in the dark at 37°C for 25 min with a 2',
7'-dichlorodihydrofluorescein diacetate working solution
(DCFH-DA/serum-free culture medium=1:1,000). Following incubation,
the cells were washed twice with serum-free culture medium to
remove any intracellular unbound DCFH-DA and subsequently imaged
using a fluorescence microscope.
Xenograft model
A total of 12 male Balb/c nude mice, obtained from
Charles River Laboratories, were housed in a specific pathogen-free
environment at 25°C with a 12/12-h light/dark cycle, and provided
with ad libitum access to food and water. The mice were
allocated randomly into two groups (sh-NC and sh-HELLS, n= 6). Each
animal experimental procedure gained approval from Institutional
Animal Care and Use Committee of Jilin University (approval no.
SY202406011; Changchun, China). A total of 5×106 cells
transfected with sh-HELLS or sh-NC plasmids were injected
subcutaneously into the dorsal region of each nude mouse,
respectively. Following cell implantation, tumor diameter was
measured every 3 days to construct a tumor growth curve. On day 19
post-implantation, the mice were euthanized by intravenous
injection of pentobarbital sodium at a dose of 150 mg/kg.
Subsequently, the tumors were surgically excised, weighed and
preserved for subsequent research.
Reverse transcription-quantitative
polymerase chain (RT-qPCR) reaction
The TRIzol (Invitrogen; Thermo Fisher Scientific,
Inc.) method was initially employed to extract the total RNA.
Subsequently, a reverse-transcription assay was conducted to
synthesize cDNA employing the TransScript II First-Strand cDNA
Synthesis SuperMix (TransGen Biotech Co., Ltd.) following the
manufacturer's instructions. ChamQ Universal SYBR qPCR master Mix
(Vazyme Biotech Co., Ltd.) was used for RT-qPCR assays to assess
gene expression, with thermocycling conditions as follows: Initial
denaturation at 95.0°C for 30 sec; followed by 40 cycles of
denaturation at 95.0°C for 10 sec, annealing and extension at
60.0°C for 30 sec. Relative expression levels were quantified using
the 2−ΔΔCq method, with GAPDH as the normalization
criterion (27). The primer
sequences are presented in Table
SI.
Western blot analysis
Radio immunoprecipitation assay buffer (Beyotime
Institute of Biotechnology), supplemented with
phenyl-methane-sulfonyl fluoride (PMSF) and protease inhibitor
cocktail (100X), was initially used to obtain the total protein.
Following the quantification of protein concentration utilizing a
bicinchoninic acid (BCA) assay kit, total proteins (20 μg
per lane) were separated via sodium dodecyl sulfate-polyacrylamide
gel electrophoresis (SDS-PAGE) on a 10% acrylamide gel and
subsequently transferred to polyvinylidene difluoride membranes
(MilliporeSigma). Following blocking with 5% non-fat milk (prepared
with 0.5 g milk powder in 10 ml TBS-T) at room temperature for 1 h,
the membranes were incubated with primary antibodies (1:1,000) at
4°C overnight, after which they were incubated with secondary
antibodies (1:5,000). An enhanced chemiluminescence (ECL) kit
(Beyotime Institute of Biotechnology) was employed to detect
protein blots using a gel imaging system (Chemidoc XRS, Bio-Rad
Laboratories, Inc.). Grayscale analysis of the protein bands was
performed using ImageJ software (Version 1.8.0; National Institutes
of Health). The antibodies included Lsh/HELLS (cat. no. A22289),
Beclin 1 (cat. no. A21191), MAP1LC3A (cat. no. A12319), COX2/PTGS2
(cat. no. A3560), ACSL4 (cat. no. A22901), HO-1/HMOX1 (cat. no.
A19062), GPX4 Rabbit mAb (cat. no. A11243; all from ABclonal
Biotech Co., Ltd.), anti-4 hydroxynonenal (cat. no. ab48506),
anti-Nrf2 antibody (cat. no. ab137550; both obtained from Abcam),
horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG (cat.
no. A0208) and HRP-conjugated goat anti-mouse IgG (cat. no. A0216;
both purchased from Beyotime Institute of Biotechnology).
Immunohistochemistry
The tissue, which had been fixed with 4% buffered
paraformaldehyde at room temperature for 6 h and dehydrated before
paraffin embedding, was sectioned into ~5-μm-thick slices.
The slides were then made transparent and dehydrated, and antigens
were retrieved through immersion in boiled sodium citrate buffer
(pH 6.0) for 10 min. Subsequently, a 0.5% Triton X-100 solution,
prepared in Dulbecco's phosphate-buffered saline, was incubated
with the slides for 20 min and then applied with a 3%
H2O2 solution to eliminate endogenous
peroxidases. Following a 30-min blocking period with a 5% bovine
serum albumin (Beyotime Institute of Biotechnology) solution, the
slides were incubated overnight with diluted primary antibody
against Lsh/HELLS (1:100; cat. no. A22289; ABclonal Biotech Co.,
Ltd.) at 4°C, then incubated for 30 min with HRP-conjugated goat
anti-mouse IgG secondary antibody (1:50; cat. no. A0216; both
purchased from Beyotime Institute of Biotechnology) and stained
using a DAB kit (Beyotime Institute of Biotechnology). The staining
results were then observed and imaged using a light microscope.
Co-immunoprecipitation (Co-IP)
CO-IP assays for HELLS and Nrf2 were conducted using
the Pierce® Co-IP Kit (cat. no. 26149; Thermo Fisher Scientific,
Inc.) following the manufacturer's protocol. Briefly, protein
lysates were prepared from HK-1 cells using PMSF buffer
supplemented with a protease inhibitor cocktail. Centrifuge columns
containing resin suspension were incubated with anti-HELLS antibody
(1:100; cat. no. 11955-1-AP; Proteintech Group, Inc.) or anti-Nrf2
antibody (1:100; cat. no. 2221102; Zen Bio; https://www.zen-bio.cn/); rabbit IgG (1:100; cat. no.
30000-0-AP; Proteintech Group, Inc.) served as the negative control
at 4°C for 2 h. Protein lysates were then incubated with the
antibody-loaded resin to capture target proteins at 4°C overnight,
which were eluted with elution buffer thereafter. The eluted
proteins were separated by SDS-PAGE and subsequently analyzed by
western blotting.
Statistical analysis
Statistical analyses were performed using GraphPad
Prism 8.0 software (Dotmatics). Before conducting t-tests and
one-way analysis of variance (ANOVA) on the experimental data, the
homogeneity of variance assumption of the data was verified using
the Homogeneity of Variance Test built into the software.
Specifically, unpaired Student's t test was used for comparisons
between two groups, while Welch's corrected t-test was applied if
variance homogeneity was not satisfied. For comparisons among
multiple groups, one-way ANOVA was used in combination with Tukey's
post hoc test. Results are expressed as the mean ± standard
deviation. P<0.05 was considered to indicate a statistically
significant difference. All experiments were conducted in
triplicate.
Results
HELLS exhibits elevated expression in
NPC
A total of 465 DEGs, comprising 125 upregulated and
331 downregulated genes, were identified from the GSE12452 dataset
(Fig. 1A). Additionally, a total
of 484 FRGs were obtained from FerrDB (Table SII), and 15 DEGs associated with
ferroptosis (FRDEGs) were obtained by the intersection between DEGs
and FRGs (Fig. 1B and Table SIII). In the present study,
HELLS was ultimately chosen from 15 FRDEGs for further
investigation based on a thorough literature review (28-32) and alignment with research
interests.
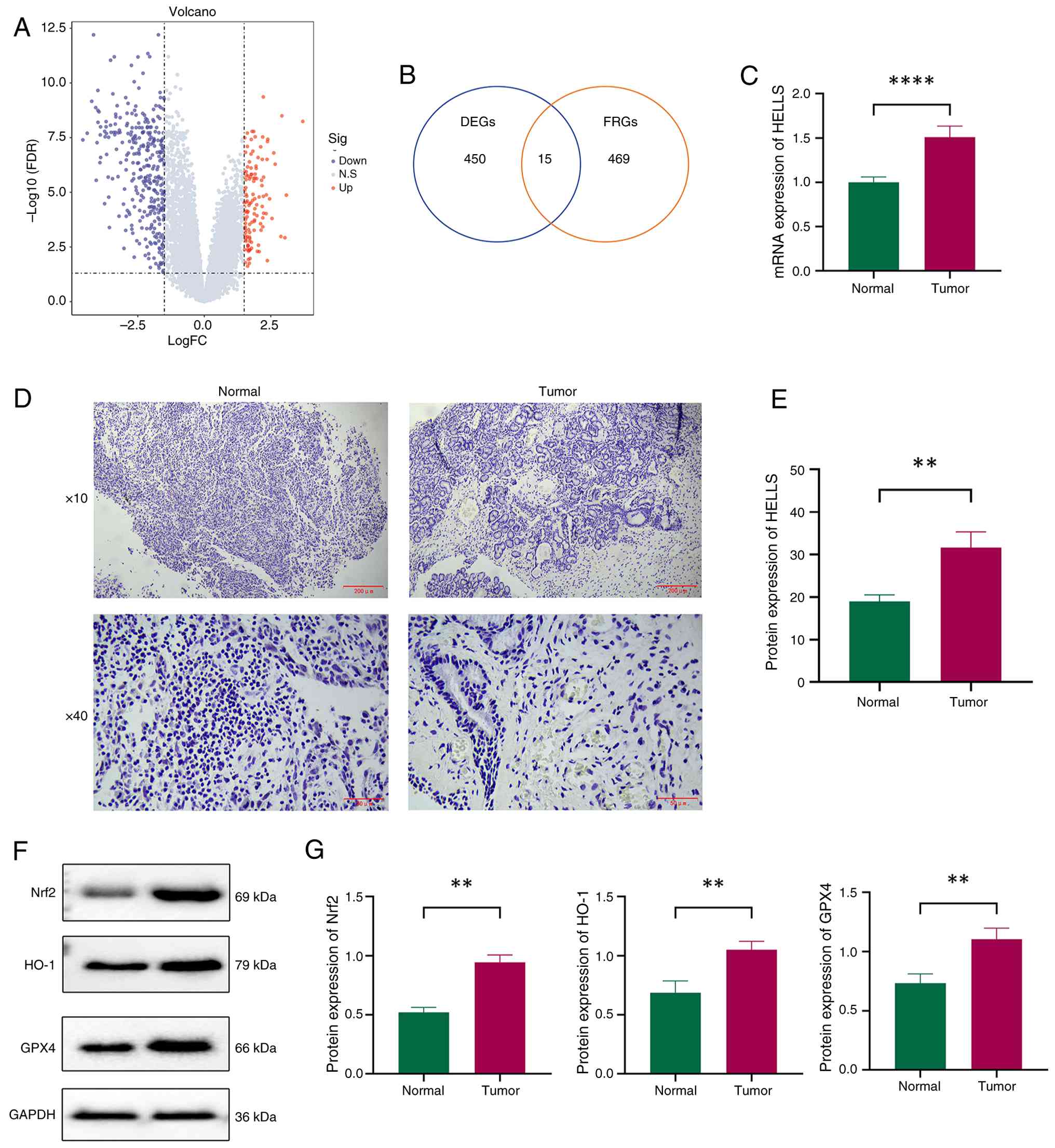 | Figure 1Identification and validation of the
expression of the key gene HELLS in NPC. (A) Volcano plot
illustrating DEGs identified through differential expression
analysis. (B) Venn diagram of the interaction between DEGs and
FRGs. (C) Relative mRNA expression levels of HELLS in NPC and
paracancerous tissue (n=5). (D) Representative images depicting the
immunohistochemical staining of HELLS in patients with NPC and
paracancerous tissue. (E) Relative protein expression levels of
HELLS in patients with NPC and paracancerous tissue. (F) The
expression of Nrf2, HO-1 and GPX4 protein were detected using WB in
NPC and adjacent. (G) Statistical analysis of Nrf2, HO-1 and GPX4
protein detected by WB. **P<0.01 and
****P<0.0001. HELLS, lymphoid-specific helicase; NPC,
nasopharyngeal carcinoma; DEGs, differentially expressed genes;
FRGs, ferroptosis-related genes; Nrf2, nuclear factor-erythroid
2-related factor 2; HO-1, heme oxygenase 1; GPX4, glutathione
peroxidase 4; WB, western blotting. |
Following the official Seurat workflow, after
performing dimensionality reduction and clustering on the cells, 9
major cell types were ultimately obtained, including: Plasma B
cells, B cells, plasmacytoid dendritic cells, myofibroblasts,
endothelial cells, epithelial cells, T cells, natural killer cells
and myeloid cells (Fig. S1A).
The marker gene corresponding to each cell type are shown in
Fig. S1B and C. Meanwhile, the
expression level of HELLS in total cell was significantly higher in
the primary tumor group (P<2.2×10−16, Fig. S1D), and the expression level of
HELLS in epithelial cells was also significantly higher in the
primary tumor group (P=3.82×10−8; Fig. S1E). To compare the expression
levels of HELLS among different NPC subtypes, the GSE189642 dataset
was obtained from the GEO database. The samples were grouped into
high and low intratumoral bacterial load groups based on the degree
of bacterial infiltration. The analysis results showed that the
expression level of HELLS in the high intratumoral bacterial load
group was significantly higher than that in the low load group
(P=0.02), suggesting a positive association between HELLS
expression and the degree of intratumoral bacterial infiltration
(Fig. S1F).
Subsequently, the mRNA and protein expression levels
of HELLS were validated by RT-qPCR and immunohistochemistry,
respectively. As illustrated in Fig.
1C-E, both mRNA and protein expression levels of HELLS were
significantly increased in the NPC tumor tissue in comparison with
the adjacent non-cancerous tissue (P<0.01), thereby providing
further evidence of the high expression of HELLS in NPC. Western
blotting showed that Nrf2, HO-1 and GPX4 protein were significantly
increased in NPC compared with adjacent tissue, suggesting that NPC
tumor cells may escape ferroptosis and promote survival by
activating this pathway (P<0.01, Fig. 1F and G).
HELLS promotes the progression of NPC in
vitro
The selection of an appropriate cell line for
research is of utmost importance. In alignment with the study's
objectives, cell models were selected based on their HELLS
expression. As demonstrated in Fig.
2A, HELLS expression was upregulated in NPC/HK1 and C666-1.
Meanwhile, the same expression trend was observed at the protein
level (Fig. 2B and C). Notably,
the expression level of HELLS in the NPC/HK1 cell line was
significantly higher than those in NP69 and C666-1 cell lines
(P<0.01). Consequently, the NPC/HK1 was selected for further
experimentation. Subsequently, HELLS was suppressed through the
application of cell transfection.
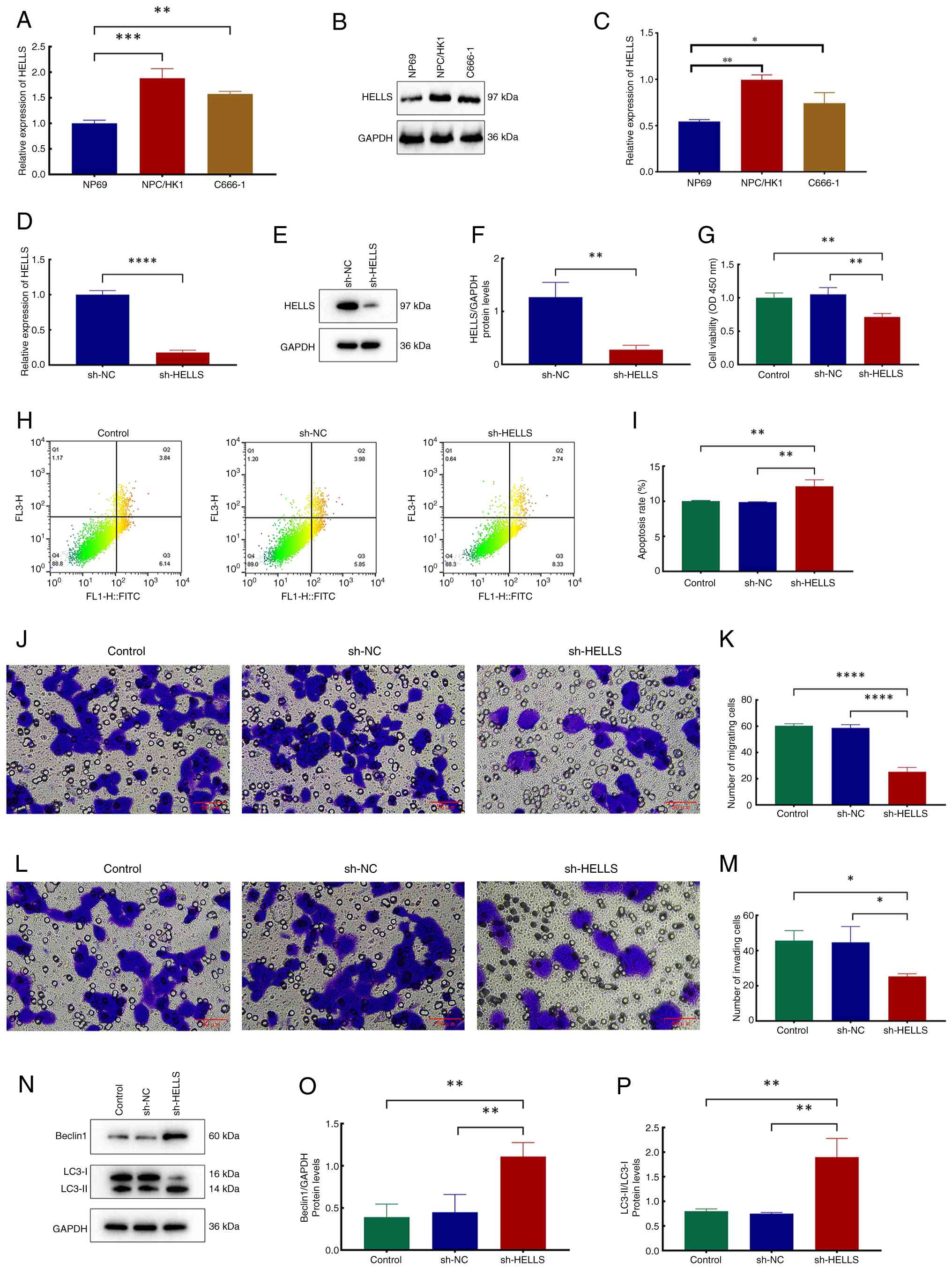 | Figure 2Exploration of the effects of HELLS
on NPC in vitro. (A) Relative mRNA expression levels of
HELLS in NPC cell lines NP69, NPC/HK1 and C666-1. (B)
Representative images of protein blots of HELLS. (C) Relative
protein expression levels of HELLS in the NPC cell lines NP69,
NPC/HK1 and C666-1. (D) Relative mRNA expression levels of HELLS in
the sh-NC and sh-HELLS groups. (E) Representative images of protein
blots of HELLS. (F) Relative protein expression levels of HELLS in
the sh-NC and sh-HELLS groups. (G) Relative viability of the
control, sh-NC and sh-HELLS groups. (H) Flow cytometry plots of the
control, sh-NC and sh-HELLS groups. (I) Analysis of cell apoptosis'
rates across the three groups. (J) Representative images depicting
crystal violet staining of migratory cells across the three groups.
(K) Quantitative analysis of migratory cells across the three
groups. (L) Representative images depicting crystal violet staining
of invasive cells across the three groups. (M) Quantitative
analysis of invading cells across the three groups. (N)
Representative images of protein blots of beclin1, LC3-I and
LC3-II. (O) Relative protein expression levels of beclin1 in the
three groups. (P) Relative ratios of LC3-II to LC3-I protein
expression levels in the three groups. *P<0.05,
**P<0.01, ***P<0.001 and
****P<0.0001. HELLS, lymphoid-specific helicase; NPC,
nasopharyngeal carcinoma; sh-, short hairpin; NC, negative
control. |
The effect of HELLS gene knockout on NPC cells was
subsequently investigated in detail. As revealed in Fig. 2D-F, the expression levels of both
the HELLS gene and protein significantly decreased in the sh-HELLS
group compared with those in the sh-NC group (P<0.01). The
evaluation of cell proliferation capacity revealed that the
sh-HELLS group demonstrated a significantly diminished
proliferative ability in comparison with both the sh-NC and control
groups (Fig. 2G; P<0.01).
Furthermore, cell apoptosis analysis revealed that low HELLS
expression significantly increased the apoptotic rate, which
reached 12.13±0.92% (Fig. 2H and
I; P<0.01). Subsequently, the migratory and invasive
properties of the cells were evaluated. As illustrated in Fig. 2J and K, when compared with the
control and sh-NC groups, the migratory abilities of sh-HELLS cells
were significantly decreased (P<0.0001). Moreover, the invasive
capabilities of sh-HELLS cells were significantly reduced (Fig. 2L and M; P<0.05). Furthermore,
the assessment of autophagy markers revealed that HELLS knockout
led to an increase in Beclin 1 expression levels and the
LC3-II/LC3-I ratio (Fig. 2N-P;
P<0.01), thereby providing evidence for an enhanced autophagic
process. Concurrent with these analyses, the identical experimental
procedures were replicated in the C666-1 cell line, which further
validated the robustness and reproducibility of the aforementioned
findings (Fig. S2).
HELLS suppresses ferroptosis
To evaluate the role of HELLS in ferroptosis, the
ferroptosis agonist erastin and the inhibitor ferrostatin-1 (Fer-1)
were administered to cells, and a comprehensive assessment of
ferroptosis-related biomarkers (GSH, MDA, Fe2+ and ROS)
was then conducted. In comparison with the sh-NC group, GSH levels
were significantly reduced in the sh-HELLS group (P<0.05).
Following treatment with erastin, GSH levels in the cells were
significantly diminished compared with the untreated group
(P<0.0001), whereas the subsequent administration of Fer-1 after
erastin administration effectively reversed this outcome (Fig. 3A; P<0.0001). Subsequent
analysis of MDA levels revealed a significant increase (P<0.05)
following HELLS knockdown. Furthermore, the erastin group exhibited
significantly high MDA levels, whereas treatment with Fer-1
resulted in a significant reduction of MDA levels in this context
(Fig. 3B; P<0.0001). The
results of Fe2+ concentration measurements closely align
with those obtained for MDA. Specifically, under identical
experimental conditions, the Fe2+ concentration in the
sh-HELLS group was remarkably high compared with that in the sh-NC
group (P<0.05). The findings also indicated that erastin
facilitates the enhancement of Fe2+ accumulation,
whereas Fer-1 appears to mitigate this phenomenon (Fig. 3C; P<0.0001). The evaluation of
the three aforementioned indicators presented substantial evidence
that HELLS significantly suppresses ferroptosis. Meanwhile, similar
results were likewise observed in C666-1 cells (Fig. S3).
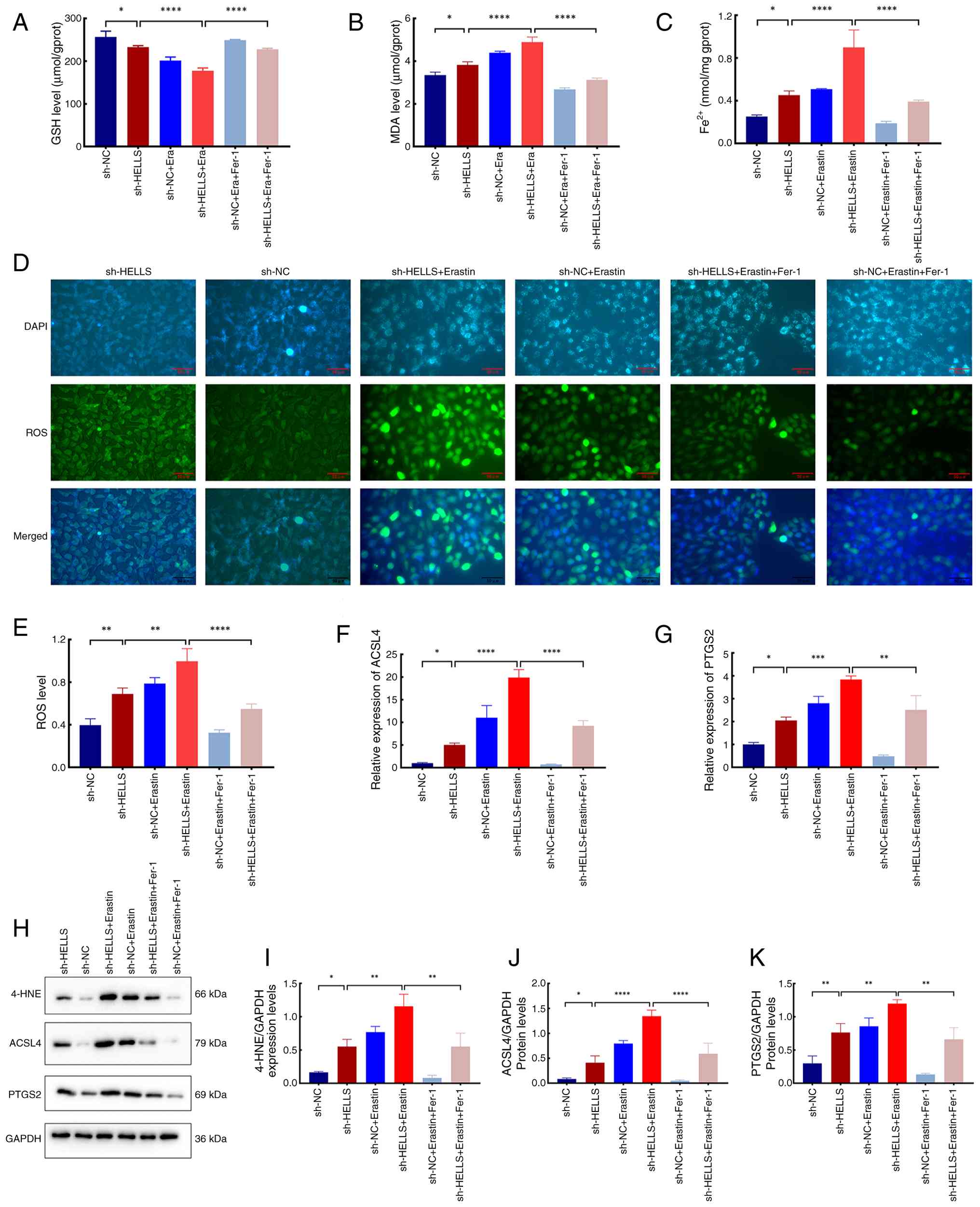 | Figure 3Effects of HELLS on ferroptosis.
Levels of (A) GSH, (B) MDA and (C) Fe2+ in the sh-HELLS,
sh-NC, sh-HELLS + Era, sh-NC + Era, sh-HELLS + Era + Fer-1 and
sh-NC + Era + Fer-1 groups. (D) Representative images illustrating
immunofluorescence staining of ROS probes in the six groups. (E)
ROS levels in the six groups. (F and G) Relative mRNA expression
levels of (F) ACSL4 and (G) PTGS2 in the six groups. (H)
Representative images of immunoblots of 4-HNE, ACSL4 and PTGS2. (I)
Relative expression levels of 4-HNE in the six groups. (J and K)
Relative protein expression levels of (J) ACSL4 and (K) PTGS2 in
the six groups. *P<0.05, **P<0.01,
***P<0.001 and ****P<0.0001. HELLS,
lymphoid-specific helicase; GSH, glutathione; MDA, malondialdehyde;
sh-, short hairpin; NC, negative control; Era, erastin; Fer-1,
ferrostatin-1; ROS, reactive oxygen species; 4-HNE,
4-hydroxynonenal; ACSL4, acyl-CoA synthetase long-chain family
member 4; PTGS2, prostaglandin-endoperoxide synthase 2. |
The ROS levels were further assessed, along with the
expression levels of ferroptosis-associated genes and proteins. ROS
levels significantly increased in the sh-HELLS group in comparison
with those in the sh-NC group (P<0.01). This trend persisted
following the administration of identical treatments with erastin
and erastin in conjunction with Fer-1 (Fig. 3D and E; P<0.05). The
expression levels of FRGs and proteins were consistent with the
results of the assessment of various ferroptosis markers. RT-qPCR
analyses revealed a significant upregulation of ACSL4 and
PTGS2 genes following HELLS suppression (P<0.05).
Additionally, erastin administration led to a significant increase
in the expression of these genes (P<0.05). Conversely, the
ferroptosis inhibitor Fer-1 significantly reversed this high
expression trend (Fig. 3F and G;
P<0.01). The protein expression patterns of these genes aligned
with their respective gene expressions (Fig. 3H, J and K).
4-Hydroxynonenal (4-HNE) functions as a significant
biomarker for ferroptosis-associated lipid peroxidation. Western
blot analysis showed that 4-HNE levels accumulated in patterns
consistent with the expression profiles of ACSL4 and PTGS2 across
all cell groups (Fig. 3H and I;
P<0.05), further supporting that HELLS significantly suppresses
ferroptosis.
HELLS inhibits autophagy-dependent
ferroptosis
The role of HELLS in autophagy-dependent ferroptosis
was subsequently investigated. Following erastin treatment, the
autophagy agonist Rapa and inhibitor 3-MA were administered to
sh-HELLS and sh-NC cells, and cell viability was assessed. The
sh-HELLS group exhibited significantly lower cell viability than
the sh-NC group (P<0.05; Fig.
4A). Furthermore, Rapa treatment significantly decreased cell
viability, whereas 3-MA treatment counteracted this effect. This
assessment preliminarily indicated that HELLS could inhibit
ferroptosis via an autophagy-dependent mechanism.
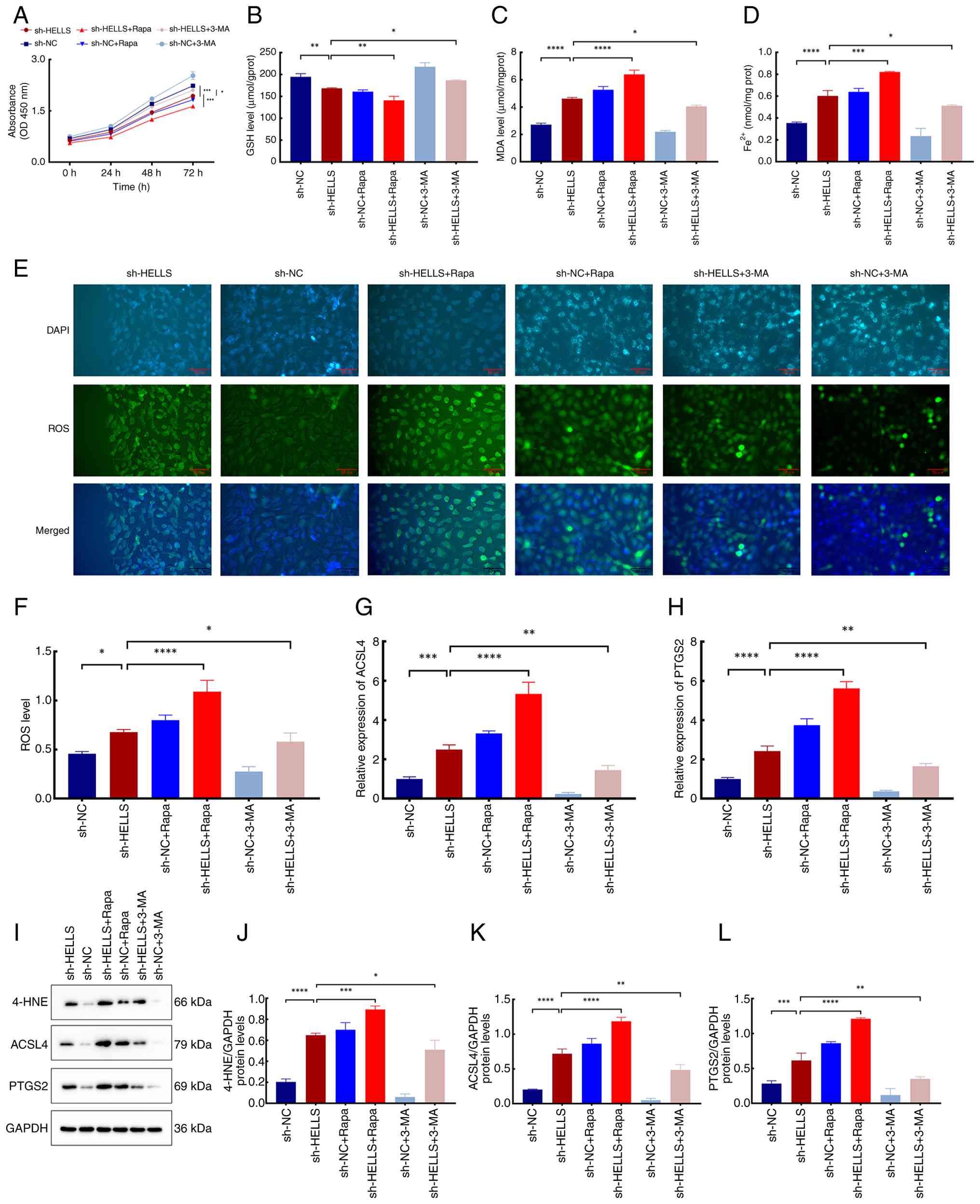 | Figure 4Effects of HELLS on
autophagy-dependent ferroptosis. (A) Relative viability of the
sh-HELLS, sh-NC, sh-HELLS + Rapa, sh-NC + Rapa, sh-HELLS + 3-MA,
and sh-NC + 3-MA groups at 0, 24, 48 and 72 h. Levels of (B) GSH,
(C) MDA and (D) Fe2+ in the six groups. (E)
Representative images illustrating immunofluorescence staining of
the ROS probes in the six groups. (F) ROS levels in the six groups.
(G and H) Relative mRNA expression levels of (G) ACSL4 and (H)
PTGS2 in the six groups. (I) Representative images of immunoblots
of 4-HNE, ACSL4, and PTGS2. (J) Relative expression levels of 4-HNE
in the six groups. Relative protein expression levels of (K) ACSL4
and (L) PTGS2 in the six groups. *P<0.05,
**P<0.01, ***P<0.001 and
****P<0.0001. HELLS, lymphoid-specific helicase; sh-,
short hairpin; NC, negative control; Rapa, rapamycin; 3-MA,
3-Methyladenine; GSH, glutathione; MDA, malondialdehyde; ROS,
reactive oxygen species; 4-HNE, 4-hydroxynonenal; ACSL4, acyl-CoA
synthetase long-chain family member 4; PTGS2,
prostaglandin-endoperoxide synthase 2. |
To validate the enhanced role of HELLS in
autophagy-dependent ferroptosis, GSH, MDA, Fe2+ and ROS
were re-analyzed after Rapa or 3-MA treatments. The experimental
results demonstrated that under consistent experimental conditions,
the levels of GSH in the sh-HELLS group were substantially reduced
compared with those in the sh-NC group (Fig. 4B; P<0.05). Additionally, Rapa
induced a significant reduction in GSH levels, whereas 3-MA
substantially increased them (P<0.05). Conversely, the levels of
MDA, Fe2+ and ROS in each cell group were inversely
associated with those of GSH (Fig.
4C-F; P<0.05).
Moreover, expression levels of ACSL4 and
PTGS2 were increased in the sh-HELLS group compared with
those in the sh-NC group (P<0.05). Similarly, Rapa or 3-MA
treatment resulted in the upregulation or downregulation of these
genes, respectively (Fig. 4G and
H; P<0.05). The expression patterns of the corresponding
proteins aligned with those of their respective genes, as well as
the consistent expression level of the biomarker 4-HNE (Fig. 4I-L; P<0.001 and P<0.0001).
Undoubtedly, these findings provide compelling evidence that HELLS
is crucial in inhibiting autophagy-dependent ferroptosis.
HELLS regulates autophagy-dependent
ferroptosis through the Nrf2/HO-1/GPX4 signaling pathway
To uncover the role of HELLS in modulating
ferroptosis via the Nrf2/HO-1/GPX4 signaling pathway, cells were
treated with the Nrf2 activator ML334 following erastin treatment,
and ferroptosis markers, as well as the pathway components, were
assessed.
The Co-IP results indicated that there was an
interaction between HELLs and Nrf2 (Fig. 5A). Cell viability assessment
revealed that the sh-HELLS group had significantly lower cell
viability than the sh-NC group (P<0.05). Nevertheless, following
ML334 treatment, cell viability significantly enhanced, indicating
that ML334 effectively mitigates the ferroptosis induced by HELLS
knockout (Fig. 5B; P<0.05).
Additionally, the levels of ferroptosis biomarkers (GSH, MDA and
Fe2+) were re-evaluated following ML334 treatment.
Consistent with previous investigations, GSH levels significantly
decreased after HELLS knockout, whereas MDA and Fe2+
levels comparatively increased (P<0.0001). Following ML334
treatment, GSH levels significantly increased, accompanied by a
simultaneous decrease in MDA and Fe2+ concentrations in
both cell groups, in comparison with pretreatment conditions
(Fig. 5C and D; P<0.05).
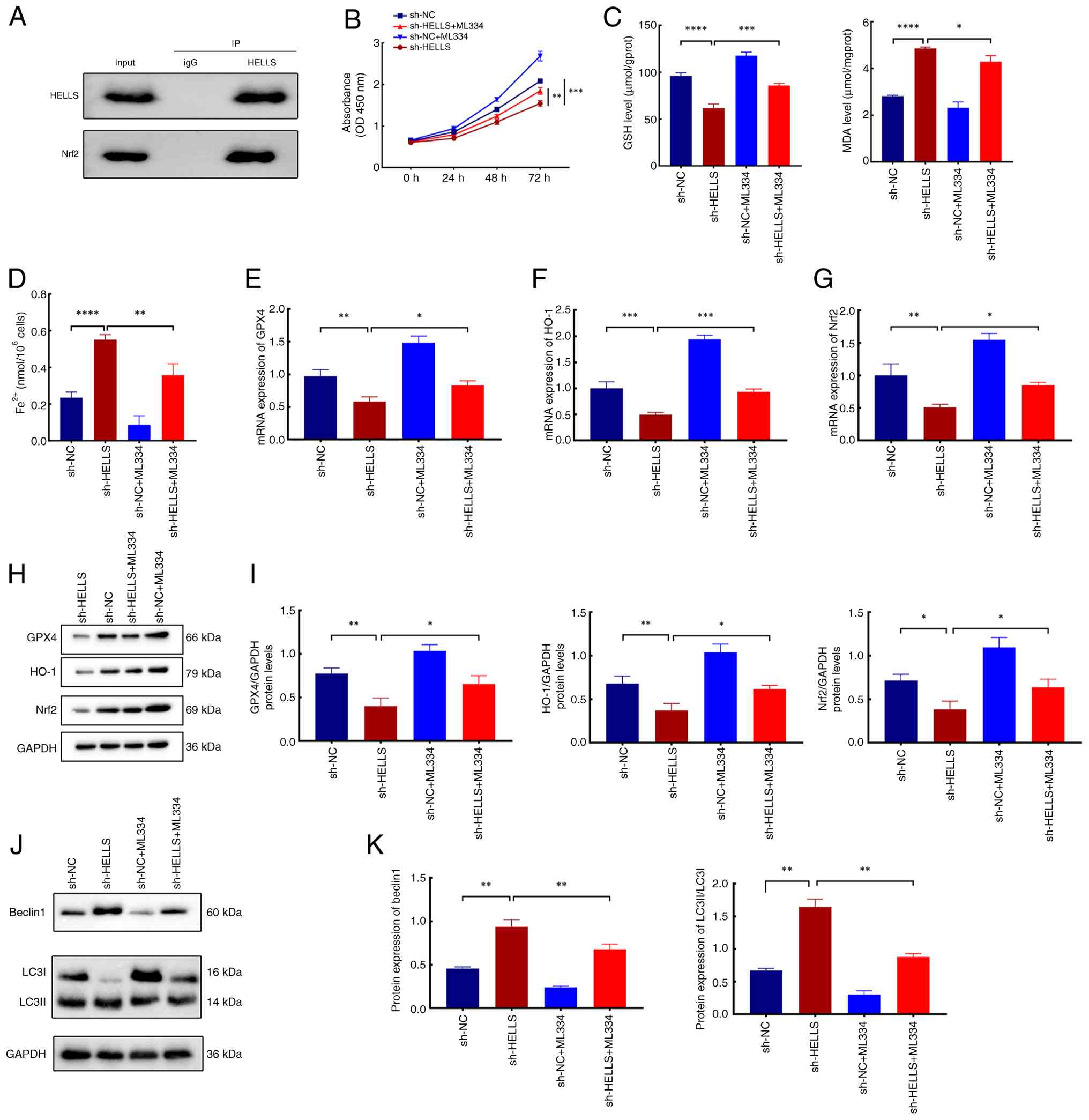 | Figure 5Mechanisms underlying the effects of
HELLS on autophagy-dependent ferroptosis. (A)
Co-immunoprecipitation was performed in HK-1 cells with either
HELLS antibody or normal rabbit IgG antibody, followed by
immunoblot analysis using the indicated antibodies. (B) Relative
viability of the sh-HELLS, sh-NC, sh-HELLS + ML334 and sh-NC +
ML334 groups at 0, 24, 48 and 72 h. Levels of (C) GSH, MDA and (D)
Fe2+ in the four groups. (E-G) Relative mRNA expression
levels of (E) GPX4, (F) HO-1 and (G) Nrf2 in the four groups. (H)
Representative images of protein blots of GPX4, HO-1 and Nrf2. (I)
Relative protein expression levels of GPX4 HO-1 and Nrf2 in the
four groups. (J) Representative images of protein blots of
Beclin1and LC3. (K) Relative protein expression levels of
Beclin1and LC3 in the four groups. *P<0.05,
**P<0.01, ***P<0.001 and
****P<0.0001. HELLS, lymphoid-specific helicase; sh-,
short hairpin; NC, negative control; GSH, glutathione; MDA,
malondialdehyde; GPX4, glutathione peroxidase 4; HO-1, heme
oxygenase-1; Nrf2, nuclear factor-erythroid 2-related factor 2. |
To further elucidate the involvement of HELLS in the
Nrf2/HO-1/GPX4 signaling pathway, the expression levels of
associated genes and proteins were evaluated. As demonstrated in
Fig. 5E-G, the expression levels
of Nrf2, HO-1 and GPX4 genes were
significantly downregulated in the sh-HELLS group (P<0.01).
Furthermore, ML334 administration led to a substantial increase in
the expression levels of these genes (P<0.05). Additionally, the
protein levels of Nrf2, HO-1 and GPX4 were evaluated, revealing
that their expression patterns in each cell group correspond with
their respective gene expression profiles (Fig. 5H and I; P<0.05 and P<0.01).
The aforementioned findings indicated that HELLS knockout
significantly inhibits the Nrf2/HO-1/GPX4 signaling pathway, which
acts as a critical regulator in the modulation of
autophagy-dependent ferroptosis. Meanwhile, after knockdown of
HELLS, the levels of Beclin1 and LC3 increased significantly
(P<0.05); subsequent addition of ML334 led to a significant
decrease in Beclin1 and LC3 (P<0.05). These findings indicated
that HELLS influences the progression of NPC by regulating
autophagy via Nrf2(Fig. 5J and
K).
HELLS facilitates NPC progression by
modulating the Nrf2/HO-1/GPX4 signaling pathway in vivo
In vitro studies have demonstrated that HELLS
inhibits ferroptosis by modulating the Nrf2/HO-1/GPX4 signaling
pathway, thereby facilitating NPC progression. To further validate
this, in vivo investigations were also conducted. Initially,
a tumorigenesis assay utilizing nude mice was conducted to
establish a xenograft model of NPC (Fig. 6A and B). The data presented
(Fig. 6C and D) clearly
indicated that the tumor volume and weight in the sh-HELLS group
were substantially reduced in comparison with those in the sh-NC
group (P<0.05), further indicating that HELLS acts as a critical
factor in promoting NPC development. Moreover, the expression level
of HELLS was significantly reduced in the tumor tissue in the
sh-HELLS group in comparison with that in the sh-NC group (Fig. 6E and F; P<0.01). Besides, mRNA
and protein expression levels of the autophagy markers LC3 and
Beclin 1 were significantly upregulated in tumors with suppressed
HELLS expression, indicating a suppressive effect of HELLS on
autophagy (Fig. 6G-K; P<0.05
and P<0.001).
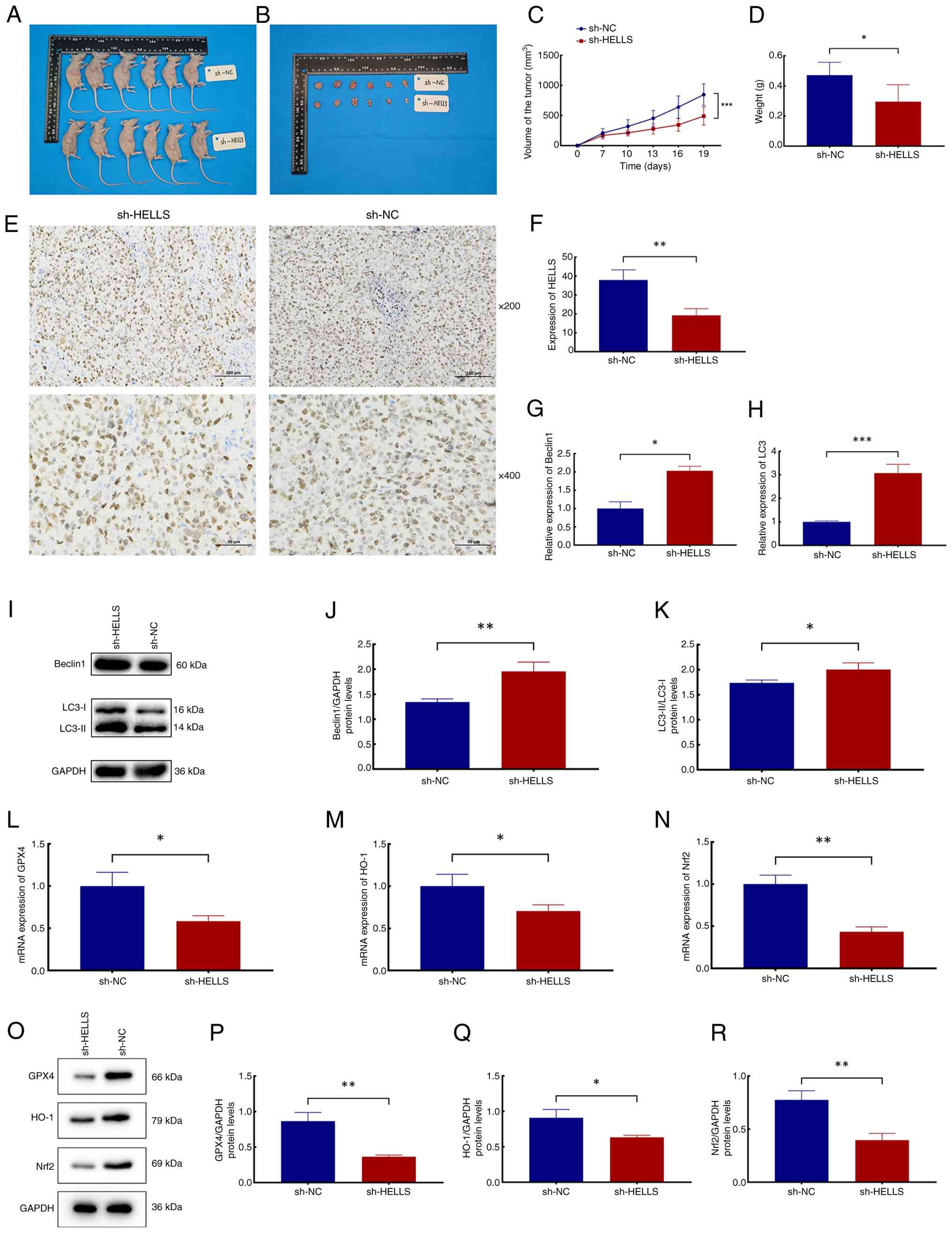 | Figure 6Effects of HELLS on
autophagy-dependent ferroptosis and its mechanism in vivo.
(A) Formation of subcutaneous nasopharyngeal carcinoma tumors in
nude mice. (B) Tumors. (C) Volume of tumors in the sh-NC and
sh-HELLS groups. (D) Weight of tumors in the two groups. (E)
Representative images illustrating immunohistochemical staining of
HELLS in tumors from the two groups (magnification, ×200 and ×400;
scale bars, 100 or 50 μm). (F) Relative protein expression
levels of HELLS in tumors from the two groups. (G and H) Relative
mRNA expression levels of (G) beclin1 and (H) LC3. (I)
Representative images of protein blots of beclin1, LC3-I and
LC3-II. (J) Relative protein expression levels of beclin1 in the
two groups. (K) Relative ratios of LC3-II to LC3-I protein
expression levels in the two groups. (L-N) Relative mRNA expression
levels of (L) GPX4, (M) HO-1 and (N) Nrf2 in tumors from the two
groups. (O) Representative images of protein blots of GPX4, HO-1
and Nrf2. (P-R) Relative protein expression levels of (P) GPX4, (Q)
HO-1 and (R) Nrf2 in tumors from the two groups.
*P<0.05, **P<0.01 and
***P<0.001. HELLS, lymphoid-specific helicase; sh-,
short hairpin; NC, negative control; GPX4, glutathione peroxidase
4; HO-1, heme oxygenase-1; Nrf2, nuclear factor-erythroid 2-related
factor 2. |
Subsequently, the molecular mechanisms by which
HELLS inhibits ferroptosis and promotes NPC progression were
validated in vivo. The mRNA and protein expression levels of
GPX4, HO-1 and Nrf2 genes were all
significantly downregulated in the tumors of the sh-HELLS group
compared with those of the sh-NC group (Fig. 6L-R; P<0.05 and P<0.01). The
consistent expression patterns of genes and proteins associated
with the Nrf2/HO-1/GPX4 signaling pathway observed both in
vivo and in vitro provide compelling evidence that HELLS
can inhibit ferroptosis and promote NPC progression through the
modulation of this pathway.
Discussion
In contrast to other regions globally, China,
particularly its southern provinces, has consistently been
demonstrated as a high-incidence area for NPC. Although the
incidence of this malignant carcinoma has declined in recent years,
the current epidemiological landscape remains critically concerning
and warrants substantial attention. Currently, the clinical
treatment regimens commonly used for NPC include radiotherapy,
chemotherapy, combined chemoradiotherapy and immunotherapy.
However, their application is often limited by adverse reactions,
drug resistance and other side effects (33-36). Therefore, investigating the
pathogenesis of NPC and exploring novel therapeutic strategies
remain the focus of current research. Despite research
advancements, considerable uncertainties persist regarding the
pathogenesis and progression of NPC, highlighting a substantial
knowledge gap in various aspects of the disease. A comprehensive
understanding of the pivotal genes and pathogenic mechanisms
associated with NPC may greatly enhance the prevention and
therapeutic strategies employed in clinical practice for this
challenging condition.
In the present study, a bioinformatics analysis and
a comprehensive literature review were conducted, which led to the
selection and subsequent determination of the HELLS gene for
in-depth investigation. This gene was hypothesized to play a
significant role in NPC development. Through experimental
validations, HELLS was found to function as an oncogene, playing a
critical regulatory role in NPC and substantially promoting disease
progression with high targeting specificity. This discovery can
address the treatment gap for patients with advanced NPC or
drug-resistant NPC who have high HELLS expression, providing a new
direction for targeted therapy of NPC.
Not only in NPC, HELLS has been formally recognized
to function as an oncogene in various cancers (29). The HELLS gene encodes a
lymphoid-specific helicase, which has been widely acknowledged to
function as a pivotal molecule in the maintenance of chromosome
stability and DNA repair. Broadly acknowledged, DNA damage is a
significant inducing factor in the etiology of cancers (37); therefore, DNA repair mechanisms
exhibit pivotality in cancer progression, with significant
contributions from the products encoded by the HELLS gene. A study
indicated that HELLS exhibited elevated expression across multiple
cancers and may serve as a potential pan-cancer diagnostic and
prognostic biomarker (28).
Moreover, HELLS has been identified as a promising biomarker for
pancreatic cancer (38,39), lung cancer (40) and hepatocellular carcinoma
(41). Furthermore, Liu et
al (30) indicated that
HELLS was significantly upregulated in colorectal cancer, and its
knockdown resulted in the arrest of the G2 and M cell cycle phases.
In addition, HELLS facilitates the proliferation and self-renewal
of glioma stem cells and interacts with E2F3 and MYC to regulate
gene expression patterns in glioblastoma (42). In light of these significant
findings regarding HELLS, HELLS is asserted to hold considerable
promise as a therapeutic target for various cancers, including
NPC.
Over more than a decade of extensive research, the
inducing factors, mechanisms of occurrence, and roles of
ferroptosis in various diseases have been redefined in light of its
original discovery. A previous study indicated that ferroptosis
requires the participation of autophagy (14). Subsequently, numerous studies
have elucidated that autophagy-dependent ferroptosis constitutes a
promising therapeutic target in cancer treatment (43), including glioblastoma (44) and colorectal cancer (45). In melanoma, Wang et al
(46) demonstrated that the
upregulation of the arachidonate 5-lipoxygenase (ALOX5) gene
not only promotes autophagy and ferroptosis but also inhibits tumor
progression. Given the substantial role of autophagy-dependent
ferroptosis in cancer, the present study not only investigated the
regulatory mechanisms through which HELLS influences ferroptosis
but also examined its role in autophagy-dependent ferroptosis. By
evaluating ferroptosis markers, in conjunction with the expression
levels of ACSL4 and PTGS2 following treatment with
ferroptosis agonist and inhibitor, as well as autophagy agonist and
inhibitor, the present study provided substantial evidence that the
HELLS gene significantly contributes to the inhibition of autophagy
and ferroptosis in NPC cells. These findings indicate that HELLS
may impede autophagy-dependent ferroptosis by downregulating
autophagy in NPC cells. These findings are consistent with existing
literature, highlighting the significance of autophagy-dependent
ferroptosis in cancer.
Nrf2 functions as a transcription factor and belongs
to a small family of basic leucine zipper proteins (47). This transcription factor is
crucial in regulating the expression of genes related to the
antioxidant stress response, including the HO-1 gene
(48), whose product is
instrumental in safeguarding against oxidative damage, modulating
cellular apoptosis, and mediating inflammatory responses (49). Researchers discovered that both
Nrf2 and HO-1 can regulate the expression level of GPX4 (50), which encodes a distinct isoform
of glutathione peroxidase, which has been recognized as a central
inhibitor of ferroptosis (47,51). Thus, the Nrf2/HO-1/GPX4 signaling
pathway is regarded as a crucial mechanism in ferroptosis
regulation. Yang et al (22) indicated that Maresin-1 is
protective against acute liver injury induced by ferroptosis
through the stimulation of the Nrf2/HO-1/GPX4 pathway. Furthermore,
Yang et al (20) revealed
that polyphyllin I can induce ferroptosis in hepatocellular
carcinoma cells by inhibiting the Nrf2/HO-1/GPX4 signaling pathway.
In the present study, Co-IP experiments confirmed a physical
interaction between HELLS and Nrf2. However, further verification
through chromatin immunoprecipitation, stability analysis, and
other assays is still needed to deepen our understanding of the
regulatory effect of HELLS on Nrf2. Additionally, the present study
further confirmed that the HELLS gene can inhibit
autophagy-dependent ferroptosis in NPC cells by regulating the
Nrf2/HO-1/GPX4 signaling pathway. Notably, based on existing
findings, including the overexpression of HELLS in NPC, its role in
promoting ferroptosis resistance, and its association with poor
prognosis, it is considered that regulating HELLS through small
molecule inhibitors or gene silencing technologies holds
significant therapeutic potential in the treatment of NPC. The
development of HELLS modulators is expected to become a potentially
effective therapeutic strategy for NPC.
Currently, studies on the functional role of HELLS
in NPC remain relatively limited. Immune escape and therapeutic
resistance are key issues restricting the clinical efficacy of NPC
treatment, yet their association with HELLS has not been explored.
He et al (52) found that
HELLS is highly expressed in NPC and can regulate cancer cell
metabolism to support the epithelial-mesenchymal transition
process, thereby promoting the malignant progression of NPC.
Another study explored the interaction between HELLS and
ferroptosis inducers, and the results showed that overexpression of
HELLS can significantly inhibit the death of cervical cancer cells
mediated by the ferroptosis inducer Erastin, suggesting that HELLS
may play a key role in regulating the ferroptosis sensitivity of
tumor cells (53). In addition,
immune escape and therapeutic resistance of NPC have become
research hotspots in the field. circBART2.2 promotes the
transcriptional activation of PD-L1 by binding to RIG-1, ultimately
leading to immune escape of tumor cells (54). Cai et al (55) found that in a hypoxic
microenvironment, Erastin and its target molecule BAP1 may serve as
effective intervention approaches to reduce the drug resistance of
NPC. However, at present, the specific mechanism by which HELLS
regulates immune escape and therapeutic resistance in NPC remains
unclear, and further in-depth exploration and verification are
required in subsequent studies.
There are certain limitations to the present study.
First, the representativeness of samples and cell models is
insufficient: The number of patient tumor samples is small, only
two NPC cell lines (NPC/HK1 and C666-1) were used, and no normal
nasopharyngeal epithelial cells were included as controls.
Meanwhile, large-sample clinical follow-up data are lacking, making
it impossible to clarify the association between HELLS and patient
prognosis. In the future, it is necessary to expand the sample
size, supplement external datasets, verify HELLS function in normal
epithelial cells, and combine follow-up analysis to explore its
prognostic value. Second, although it was confirmed that HELLS
regulates ferroptosis through the Nrf2/HO-1/GPX4 pathway, the
causal relationship between HELLS and this pathway has not been
clarified, the role of HELLS in the downstream of the pathway has
not been verified, and direct evidence for autophagic degradation
of ferroptosis-related substrates is lacking. In in vivo
experiments, neither ferroptosis markers were detected nor Nrf2
agonist rescue experiments were conducted. Subsequent studies need
to supplement experiments such as rescue experiments and
loss-of-function experiments to improve the mechanistic chain.
Third, the safety and specificity of targeted intervention have not
been evaluated. Although shRNA-mediated HELLS knockdown verified
target specificity, off-target effects have not been excluded, and
the toxicity of targeted intervention on normal tissues has not
been detected. In the future, it is necessary to use CRISPR-Cas9
and RNA-sequencing to verify specificity and conduct systematic
toxicity assessment. Fourth, the exploration of research extension
and clinical transformation value is insufficient. Other mechanisms
by which HELLS regulates ferroptosis have not been deeply explored,
and its role in the context of therapeutic resistance has not been
analyzed. Subsequent plans include combining ferroptosis inducers,
integrating tumor microenvironment analysis, and investigating the
association between HELLS and drug resistance to provide insights
for breaking through therapeutic bottlenecks.
In summary, the present study emphasizes the pivotal
role of the HELLS gene in the regulation of NPC and
autophagy-dependent ferroptosis. Findings of extensive research
will deepen the understanding of the pathogenesis of NPC, thereby
providing new insights for its precise clinical management. It is
proposed that the HELLS gene holds considerable promise as a
therapeutic target for NPC. Furthermore, the development of HELLS
modulators is expected to become a viable therapeutic strategy for
clinical application in NPC treatment, thereby contributing to the
advancement of precision clinical treatments for this highly
concerning disease.
In conclusion, HELLS can inhibit autophagy-dependent
ferroptosis in NPC cells through the activation of the
NRF2/HO-1/GPX4 signaling pathway, consequently facilitating NPC
progression. The HELLS gene may represent a significant therapeutic
target for NPC.
Supplementary Data
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
ZW and CJ designed the experiments. CJ and JL
performed experiments and collected data. CJ and SH analyzed the
data. CJ, CB and JY interpreted the data. All authors read and
approved the final version of the manuscript. ZW and CJ confirm the
authenticity of all the raw data.
Ethics approval and consent to
participate
Waiver of informed consent and ethical approval for
human studies (approval no. 2023-301) were granted by the Medical
Ethics Committee of The Second Hospital of Jilin University
(Changchun, China). Each animal experimental procedure gained
approval from Institutional Animal Care and Use Committee of Jilin
University (approval no. SY202406011; Changchun, China). The
experimental protocol was performed in accordance with the relevant
guidelines and regulations of the Basel Declaration. The study is
reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
The present study was supported by the Jilin Provincial Science
and Technology Development Plan Project (grant no.
20240401069YY).
References
|
1
|
Chen YP, Chan ATC, Le QT, Blanchard P, Sun
Y and Ma J: Nasopharyngeal carcinoma. Lancet. 394:64–80. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chang ET, Ye W, Zeng YX and Adami HO: The
evolving epidemiology of nasopharyngeal carcinoma. Cancer Epidemiol
Biomarkers Prev. 30:1035–1047. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Tang LL, Chen WQ, Xue WQ, He YQ, Zheng RS,
Zeng YX and Jia WH: Global trends in incidence and mortality of
nasopharyngeal carcinoma. Cancer Lett. 374:22–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ding S, Gao Y, Lv D, Tao Y, Liu S, Chen C,
Huang Z, Zheng S, Hu Y, Chow LK, et al: DNTTIP1 promotes
nasopharyngeal carcinoma metastasis via recruiting HDAC1 to DUSP2
promoter and activating ERK signaling pathway. EBioMedicine.
81:1041002022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhang B, Li J, Wang Y, Liu X, Yang X, Liao
Z, Deng S, Deng Y, Zhou Z, Tian Y, et al: Deubiquitinase USP7
stabilizes KDM5B and promotes tumor progression and cisplatin
resistance in nasopharyngeal carcinoma through the ZBTB16/TOP2A
axis. Cell Death Differ. 31:309–321. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Liu H, Tang L, Gong S, Xiao T, Yang H, Gu
W, Wang H and Chen P: USP7 inhibits the progression of
nasopharyngeal carcinoma via promoting SPLUNC1-mediated M1
macrophage polarization through TRIM24. Cell Death Dis. 14:8522023.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta
R, Zaitsev EM, Gleason CE, Patel DN, Bauer AJ, Cantley AM, Yang WS,
et al: Ferroptosis: An iron-dependent form of nonapoptotic cell
death. Cell. 149:1060–1072. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Tang D, Kang R, Berghe TV, Vandenabeele P
and Kroemer G: The molecular machinery of regulated cell death.
Cell Res. 29:347–364. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li M, Jin S, Zhang Z, Ma H and Yang X:
Interleukin-6 facilitates tumor progression by inducing ferroptosis
resistance in head and neck squamous cell carcinoma. Cancer Lett.
527:28–40. 2022. View Article : Google Scholar
|
|
10
|
Li S, Liu Y, Li J, Zhao X and Yu D:
Mechanisms of ferroptosis and application to head and neck squamous
cell carcinoma treatments. DNA Cell Biol. 40:720–732. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen P, Wang D, Xiao T, Gu W, Yang H, Yang
M and Wang H: ACSL4 promotes ferroptosis and M1 macrophage
polarization to regulate the tumorigenesis of nasopharyngeal
carcinoma. Int Immunopharmacol. 122:1106292023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen Y, Feng Y, Lin Y, Zhou X, Wang L,
Zhou Y, Lin K and Cai L: GSTM3 enhances radiosensitivity of
nasopharyngeal carcinoma by promoting radiation-induced ferroptosis
through USP14/FASN axis and GPX4. Br J Cancer. 130:755–768. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhou R, Qiu L, Zhou L, Geng R, Yang S and
Wu J: P4HA1 activates HMGCS1 to promote nasopharyngeal carcinoma
ferroptosis resistance and progression. Cell Signal.
105:1106092023. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhou B, Liu J, Kang R, Klionsky DJ,
Kroemer G and Tang D: Ferroptosis is a type of autophagy-dependent
cell death. Semin Cancer Biol. 66:89–100. 2020. View Article : Google Scholar
|
|
15
|
Miao H, Ren Q, Li H, Zeng M, Chen D, Xu C,
Chen Y and Wen Z: Comprehensive analysis of the autophagy-dependent
ferroptosis-related gene FANCD2 in lung adenocarcinoma. BMC Cancer.
22:2252022. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Liu J, Liu Y, Wang Y, Li C, Xie Y,
Klionsky DJ, Kang R and Tang D: TMEM164 is a new determinant of
autophagy-dependent ferroptosis. Autophagy. 19:945–956. 2023.
View Article : Google Scholar :
|
|
17
|
Chen YM, Xu W, Liu Y, Zhang JH, Yang YY,
Wang ZW, Sun DJ, Li H, Liu B and Chen LX: Anomanolide C suppresses
tumor progression and metastasis by ubiquitinating GPX4-driven
autophagy-dependent ferroptosis in triple negative breast cancer.
Int J Biol Sci. 19:2531–2550. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gong G, Ganesan K, Liu Y, Huang Y, Luo Y,
Wang X, Zhang Z and Zheng Y: Danggui Buxue Tang improves
therapeutic efficacy of doxorubicin in triple negative breast
cancer via ferroptosis. J Ethnopharmacol. 323:1176552024.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Du HF, Wu JW, Zhu YS, Hua ZH, Jin SZ, Ji
JC, Wang CS, Qian GY, Jin XD and Ding HM: Fucoxanthin induces
ferroptosis in cancer cells via downregulation of the
Nrf2/HO-1/GPX4 pathway. Molecules. 29:28322024. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yang R, Gao W, Wang Z, Jian H, Peng L, Yu
X, Xue P, Peng W, Li K and Zeng P: Polyphyllin I induced
ferroptosis to suppress the progression of hepatocellular carcinoma
through activation of the mitochondrial dysfunction via
Nrf2/HO-1/GPX4 axis. Phytomedicine. 122:1551352024. View Article : Google Scholar
|
|
21
|
Zhu X, Chen X, Qiu L, Zhu J and Wang J:
Norcantharidin induces ferroptosis via the suppression of NRF2/HO-1
signaling in ovarian cancer cells. Oncol Lett. 24:3592022.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yang W, Wang Y, Zhang C, Huang Y, Yu J,
Shi L, Zhang P, Yin Y, Li R and Tao K: Maresin1 protect against
ferroptosis-induced liver injury through ROS inhibition and
Nrf2/HO-1/GPX4 activation. Front Pharmacol. 13:8656892022.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wen JY, Chen G, Li JD, Luo JY, He J, Wang
RS and Qin LT: Downregulated miR-150-5p in the tissue of
nasopharyngeal carcinoma. Genet Res (Camb). 2022:24850552022.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Luo X, Gong Y, Jiang Q, Wang Q, Li S and
Liu L: Isoquercitrin promotes ferroptosis and oxidative stress in
nasopharyngeal carcinoma via the AMPK/NF-κB pathway. J Biochem Mol
Toxicol. 38:e235422024. View Article : Google Scholar
|
|
25
|
Yang T, Huang L, Qin H and Mai S: STRESS
granule-associated RNA-binding protein CAPRIN1 drives cancer
progression and regulates treatment response in nasopharyngeal
carcinoma. Med Oncol. 40:472022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang Y, Yin W and Zhu X: Blocked autophagy
enhances radiosensitivity of nasopharyngeal carcinoma cell line
CNE-2 in vitro. Acta Otolaryngol. 134:105–110. 2014. View Article : Google Scholar
|
|
27
|
Li M, Wei Y, Liu Y, Wei J, Zhou X, Duan Y,
Chen S, Xue C, Zhan Y, Zheng L, et al: BRD7 inhibits enhancer
activity and expression of BIRC2 to suppress tumor growth and
metastasis in nasopharyngeal carcinoma. Cell Death Dis. 14:1212023.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Liang X, Li L and Fan Y: Diagnostic,
prognostic, and immunological roles of HELLS in pan-cancer: A
bioinformatics analysis. Front Immunol. 13:8707262022. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Peixoto E, Khan A, Lewis ZA,
Contreras-Galindo R and Czaja W: The chromatin remodeler HELLS: A
new regulator in DNA repair, genome maintenance, and cancer. Int J
Mol Sci. 23:93132022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Liu X, Hou X, Zhou Y, Li Q, Kong F, Yan S,
Lei S, Xiong L and He J: Downregulation of the helicase
lymphoid-specific (HELLS) gene impairs cell proliferation and
induces cell cycle arrest in colorectal cancer cells. Onco Targets
Ther. 12:10153–10163. 2019. View Article : Google Scholar
|
|
31
|
Chen X, Li Y, Rubio K, Deng B, Li Y, Tang
Q, Mao C, Liu S, Xiao D, Barreto G and Tao Y: Lymphoid-specific
helicase in epigenetics, DNA repair and cancer. Br J Cancer.
126:165–173. 2022. View Article : Google Scholar :
|
|
32
|
Jia J, Shi Y, Chen L, Lai W, Yan B, Jiang
Y, Xiao D, Xi S, Cao Y, Liu S, et al: Decrease in lymphoid specific
helicase and 5-hydroxymethylcytosine is associated with metastasis
and genome instability. Theranostics. 7:3920–3932. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Guan S, Wei J, Huang L and Wu L:
Chemotherapy and chemo-resistance in nasopharyngeal carcinoma. Eur
J Med Chem. 207:1127582020. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang C, Chen J, Su L, Hua Y, Ye J, Song X,
Lv W, Zhang M, Huang F, Tian J and Hong J: The psychological status
in patients with nasopharyngeal carcinoma during radiotherapy. Eur
Arch Otorhinolaryngol. 279:1035–1042. 2022. View Article : Google Scholar
|
|
35
|
Huang H, Yao Y, Deng X, Huang Z, Chen Y,
Wang Z, Hong H, Huang H and Lin T: Immunotherapy for nasopharyngeal
carcinoma: Current status and prospects (Review). Int J Oncol.
63:972023. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Xiao L, Kang W, Liao J and Li Y: Efficacy
and tolerability of immunotherapy in advanced nasopharyngeal
carcinoma with or without chemotherapy: A meta-analysis. Braz J
Otorhinolaryngol. 88(Suppl 1): S70–S81. 2022. View Article : Google Scholar :
|
|
37
|
Basu AK: DNA damage, mutagenesis and
cancer. Int J Mol Sci. 19:9702018. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Hou X, Yang L, Wang K, Zhou Y, Li Q, Kong
F, Liu X and He J: HELLS, a chromatin remodeler is highly expressed
in pancreatic cancer and downregulation of it impairs tumor growth
and sensitizes to cisplatin by reexpressing the tumor suppressor
TGFBR3. Cancer Med. 10:350–364. 2021. View Article : Google Scholar :
|
|
39
|
Wang FJ, Jing YH, Cheng CS, Cao ZQ, Jiao
JY and Chen Z: HELLS serves as a poor prognostic biomarker and its
downregulation reserves the malignant phenotype in pancreatic
cancer. BMC Med Genomics. 14:1892021. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Zhu W, Li LL, Songyang Y, Shi Z and Li D:
Identification and validation of HELLS (Helicase,
Lymphoid-Specific) and ICAM1 (Intercellular adhesion molecule 1) as
potential diagnostic biomarkers of lung cancer. PeerJ. 8:e87312020.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Fang Y, Tang W, Zhao D, Zhang X, Li N,
Yang Y, Jin L, Li Z, Wei B, Miao Y, et al: Immunological function
and prognostic value of lymphoid-specific helicase in liver
hepatocellular carcinoma. Cancer Biomark. 38:225–239. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zhang G, Dong Z, Prager BC, Kim LJ, Wu Q,
Gimple RC, Wang X, Bao S, Hamerlik P and Rich JN: Chromatin
remodeler HELLS maintains glioma stem cells through E2F3 and MYC.
JCI Insight. 4:e1261402019. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Liu L, Li L, Li M and Luo Z:
Autophagy-dependent ferroptosis as a therapeutic target in cancer.
ChemMedChem. 16:2942–2950. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Xie Y, Hou T, Liu J, Zhang H, Liu X, Kang
R and Tang D: Autophagy-dependent ferroptosis as a potential
treatment for glioblastoma. Front Oncol. 13:10911182023. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Miao Q, Deng WQ, Lyu WY, Sun ZT, Fan SR,
Qi M, Qiu SH, Zhu YR, Lin JP, Chen MF, et al: Erianin inhibits the
growth and metastasis through autophagy-dependent ferroptosis in
KRASG13D colorectal cancer. Free Radic Biol Med.
204:301–312. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wang M, Zeng G, Xiong B, Zhu X, Guo J,
Chen D, Zhang S, Luo M, Guo L and Cai L: ALOX5 promotes
autophagy-dependent ferroptosis by activating the AMPK/mTOR pathway
in melanoma. Biochem Pharmacol. 212:1155542023. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Dang R, Wang M, Li X, Wang H, Liu L, Wu Q,
Zhao J, Ji P, Zhong L, Licinio J and Xie P: Edaravone ameliorates
depressive and anxiety-like behaviors via Sirt1/Nrf2/HO-1/Gpx4
pathway. J Neuroinflammation. 19:412022. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Somparn N, Prawan A, Senggunprai L,
Kukongviriyapan U, Jetsrisuparb A, Lee MH, Kim DH, Kukongviriyapan
V and Surh YJ: Cellular adaptation mediated through Nrf2-induced
glutamate cysteine ligase up-regulation against oxidative stress
caused by iron overload in β-thalassemia/HbE patients. Free Radic
Res. 53:791–799. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Ali T, Kim T, Rehman SU, Khan MS, Amin FU,
Khan M, Ikram M and Kim MO: Natural dietary supplementation of
anthocyanins via PI3K/Akt/Nrf2/HO-1 pathways mitigate oxidative
stress, neurodegeneration, and memory impairment in a mouse model
of Alzheimer's disease. Mol Neurobiol. 55:6076–6093. 2018.
View Article : Google Scholar
|
|
50
|
Hayes JD and Dinkova-Kostova AT: The Nrf2
regulatory network provides an interface between redox and
intermediary metabolism. Trends Biochem Sci. 39:199–218. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Brigelius-Flohé R and Flohé L: Regulatory
phenomena in the glutathione peroxidase superfamily. Antioxid Redox
Signal. 33:498–516. 2020. View Article : Google Scholar
|
|
52
|
He X, Yan B, Liu S, Jia J, Lai W, Xin X,
Tang CE, Luo D, Tan T, Jiang Y, et al: Chromatin remodeling factor
LSH drives cancer progression by suppressing the activity of
fumarate hydratase. Cancer Res. 76:5743–5755. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Tie W and Ge F: Lymphoid-specific helicase
inhibits cervical cancer cells ferroptosis by promoting Nrf2
expression. PeerJ. 11:e164512023. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Ge J, Wang J, Xiong F, Jiang X, Zhu K,
Wang Y, Mo Y, Gong Z, Zhang S, He Y, et al: Epstein-barr
virus-encoded circular RNA CircBART2.2 promotes immune escape of
nasopharyngeal carcinoma by regulating PD-L1. Cancer Res.
81:5074–5088. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Cai W, Wu S, Lin Z, Ming X, Yang X, Yang M
and Chen X: Hypoxia-induced BAP1 enhances erastin-induced
ferroptosis in nasopharyngeal carcinoma by stabilizing H2A. Cancer
Cell Int. 24:3072024. View Article : Google Scholar : PubMed/NCBI
|




















