Inflammatory bowel disease (IBD) comprises a set of
chronic, immune-driven disorders of the gastrointestinal tract,
defined by recurrent mucosal inflammation (1,2).
IBD, which comprises two major subtypes, Crohn's disease (CD) and
ulcerative colitis (UC), has become a global healthcare challenge
with steadily rising incidence rates (1), in Southeast Asia alone, the number
of cases increased from ~103,884 in 2017 to ~118,000 in 2020, and
is projected to reach ~199,000 by 2035 (3).
IBD arises from complex interactions among
environmental factors, gut microbiota and immune-mediated
mechanisms in genetically susceptible hosts (4). For decades, research on IBD
pathogenesis has focused on mucosal immunity, particularly T cell
responses. A previous study suggest that dysregulated innate and
adaptive immune mechanisms are central to the pathogenesis of
uncontrolled intestinal inflammation in IBD (5). Key components of intestinal
immunity include intestinal epithelial cells (IECs), macrophages,
dendritic cells (DCs), neutrophils, natural killer T (NKT) cells,
innate lymphoid cells (ILCs), T cells and B cells (6). As key mediators of adaptive
immunity, T cells, including CD4+ and CD8+
subsets, mature in the thymus, migrate to peripheral tissues and
orchestrate humoral and cellular immune responses (7).
Single-cell RNA sequencing (scRNA-seq) offers
distinct advantages over conventional bulk sequencing by enabling
high-dimensional analysis of individual cells with complete
transcriptome resolution. This technology characterizes cellular
composition and population dynamics and detects actively
transcribed genes at single-cell or cluster levels, reconstructs
cellular differentiation trajectories and subtype evolution,
identifies key regulatory genes governing these processes and
deciphers intercellular communication networks (13,14). ScRNA-seq, particularly, is
valuable for investigating CD8+ T cells, a functionally
heterogeneous population and the comprehensive nature of the
technology facilitates cross-tissue/cross-species cell type
identification and, through multimodal analytical approaches,
reveals previously unrecognized receptor-ligand interactions that
may inform therapeutic target discovery (15,16). The role of CD8+ T
cells in IBD has remained controversial for decades. While some
studies have identified their anti-colitic properties through
regulatory mechanisms, others have implicated specific subsets in
driving tissue inflammation and epithelial damage (10,17,18). These conflicting findings may
stem from their tissue-specific origins (for example, peripheral
blood mononuclear cells, intestinal epithelium or lamina propria)
and intrinsic heterogeneity among CD8+ T cell subsets,
which exhibit profound phenotypic and functional diversity
(19-21). Present investigations are
predominantly centered on conventional cytotoxic CD8+ T
cells and CD8+ regulatory T cells (CD8+
Tregs) (22,23), with additional attention given to
non-classical subsets such as type II cytokine-producing
CD8+ T cells, as well as CD8+ populations
secreting IL-9, IL-17 and IL-22 (21). The conventional approach of
RNA-seq, which is limited to analyzing specific cell types or
tissues under the assumption of population homogeneity, may
represent one key factor contributing to the current controversies
in CD8+ T cells research (20). By contrast, scRNA-seq provides
comprehensive transcriptome profiling at single-cell resolution,
enabling high-resolution examination of cellular heterogeneity
within precisely defined and well-characterized cell populations
(24,25). ScRNA-seq of the colonic tissues
of patients with IBD has identified highly heterogeneous
CD8+ T cell populations exhibiting distinct phenotypic
and functional characteristics (19,20).
In IBD research, scRNA-seq integrated with
complementary biological techniques has proven instrumental in
delineating cell type-specific transcriptional heterogeneity
(26,27). A multimodal study combining
scRNA-seq with subcellular spatial transcriptomics (ST) has
successfully tracked T cell populations associated with checkpoint
inhibitor (CPI) colitis in intestinal tissues and identified
distinctive features of mucosal IFN-γ-producing cytotoxic
CD8+ T cells, thereby elucidating pathogenic pathways
and informing novel preventive strategies (28). ScRNA-seq predicted that
chemokines that are important for wound healing are altered during
inflammation, in contrast to non-inflamed mucosa, with macrophages
and CD8+ T cells in inflamed mucosa identified as key
cellular targets (29).
Single-cell technologies have been utilized to
profile distinct immune cell subsets in mucosal tissues and
peripheral blood obtained from individuals with active or quiescent
CD and UC, as well as healthy donors. Furthermore, when
complemented with flow cytometry and RNA in situ
hybridization validation, these methodologies offer substantial
potential for developing precision therapies targeting distinct
cellular subsets in IBD pathogenesis (30). ScRNA-seq revealed marked immune
cell infiltration in 2,4,6-trinitrobenzene sulfonic acid
(TNBS)-induced colitis and delineated the precise immune
phenotypes, flow cytometry was subsequently employed to validate
the dynamic changes in CD8+ IFN-γ+ cytotoxic
T lymphocyte cells (CTLs) across disease and therapeutic states
(31). To elucidate the role of
IL-23 as a therapeutic target in chronic inflammation, an
integrated approach combining single-cell transcriptomics and flow
cytometry was applied to characterize IL-23 receptor-expressing
cells in the gut, which identified group 3 ILC-intrinsic cytotoxic
T-lymphocyte-associated antigen-4 (CTLA-4) as a key checkpoint
restraining IL-23-driven pathology, with disruption of these
lymphocytes shown to perpetuate chronic inflammation in IBD
(32). Another study with a
comparative analysis of colonic tissues demonstrated a marked
enrichment of CTLA-4+ T cells in patients with CD
compared to healthy controls, with a notable elevation in the rate
of CD observed both in the CTLA-4+ subpopulations
(33). These findings identify
CTLA-4+ T cells as active contributors to intestinal
inflammation. Through integrated scRNA-seq and mass cytometry
analyses, Cao et al (34)
demonstrated that Crohn's-like disease of the pouch involved a
systemic remodeling of immune and stromal compartments in the pouch
and prepouch ileum, characterized by clonally expanded effector
CD4+ T and CD8+ T cell populations.
By integrating genome-wide association studies with
scRNA-seq for credible causal mapping, the study identified
monogenic and polygenic IBD genes highly expressed in phagocytes
(circulating monocytes, neutrophils, DC2s and macrophages) and
activated T cells (CD8+ IL-17+ and
circulating T cells) which establishes a clinically actionable
framework for classifying and managing monogenic IBD while
delineating shared and distinct features between monogenic and
polygenic forms (35). ScRNA-seq
and RNA scope on terminal ileum biopsies validated upregulated
NOD-like receptor family CARD domain containing 5 and transporter
associated with antigen processing 1 in the intestinal epithelium
of patients with CD, showing co-localization with CD8+ T
cells, suggesting epithelial-lymphocyte crosstalk mediated by
mediated by major histocompatibility complex class (MHC-I)
(36).
The integration of scRNA-seq with complementary
multiomics platforms and functional validation studies (26-28,30) has established a multidimensional
analytical framework. This integrated approach not only
systematically delineates the heterogeneity of CD8+ T
cells, their microenvironmental crosstalk and clinical associations
in IBD but, more importantly, establishes a causal relationship
between cell states, underlying molecular mechanisms, functional
phenotypes and clinical disease manifestations. This framework
provides a novel conceptual and mechanistic dimension to the
understanding of IBD immunopathology and directly informs the
rationale for developing precision therapeutic strategies focused
on CD8+ T cell biology modulation.
Multiple single-cell analyses of CD revealed altered
heterogeneity and subset distribution of intestinal intraepithelial
T cells, identifying unique intraepithelial lymphocytes (IELs)
populations in the terminal ileum of patients with CD, including
NKp30+ γδT cells that express RORγt and produce IL-26
upon NKp30 engagement (46).
Comparative analyses of non-inflamed/inflamed CD tissues vs.
healthy controls showed that expanded T-helper 17 cells but
diminished CD8+ T, γδT, TFH and Treg
populations in inflamed tissues, while parallel lamina propria
findings demonstrated increased CD8+ and decreased
CD4+ T cells, with elevated Th17-to-Treg/TFH
ratios (46). Pseudotemporal
trajectory analyses inferred from transcriptional profiles
uncovered divergent differentiation trajectories among T cell
subsets, with T cell populations in CD preferentially
differentiating from Tregs toward Vδ2 γδ T cells, whereas in
colorectal cancer, the dominant trajectory progressed from
CD8+ T cells to Tregs (47). A previous study indicated that
CEACAM5-stimulated CD8+ T cells markedly inhibit the
proliferation of CD4+ T cells in vitro,
implicating their function as key modulators in maintaining
intestinal immune homeostasis (48).
The widespread application of single-cell sequencing
analyses has identified the heterogeneity of CD8+ T cell
infiltration in numerous human diseases, including malignancies
such as non-small cell lung cancer (49), breast cancer (50), head and neck cancer (51), liver cancer (52), melanoma (53), bladder cancer (54) and colorectal cancer (55), as well as immune-related diseases
such as IBD (56,57) and COVID-19 (58). Research on CD8+ T
cells in IBD, reveal that these cells are primarily categorized
into naïve, cytotoxic, regulatory, proliferative, memory and
exhausted subtypes (Table I).
Corridoni et al (56)
utilized single-cell transcriptomics combined with T cell receptor
repertoire analyses and flow cytometry to establish a comprehensive
map of CD8+ T cells in healthy and UC colonic tissues,
classifying CD8+ T cells into 14 subgroups, including
naïve, memory, effector, TRM, double-positive CD8+
CD4+ cells, mucosal-associated invariant T cells, IELs
and TF-related T cells like IL-26+-expressing
CD8+ T cells.
Tyrobp, an established modulator of
pro-inflammatory mediator synthesis in macrophages and neutrophils,
and has been mechanistically associated with the development of
multiple inflammation-associated disorders (74). Genetic ablation of Tyrobp
markedly attenuates the severity of dextran sulfate sodium
(DSS)-induced colitis in murine models (75), suggesting it may serve as an
upstream regulatory molecule in UC (76). Rosati et al (77) performed comprehensive TCRα and β
chain repertoire analyses using high-throughput sequencing on
peripheral blood and intestinal tissues from patients with IBD and
healthy controls. Subsequent scRNA-seq identified distinct T cell
clonotypes, including a CD-associated invariant T cell, enriched
within the CD8+ effector memory CD161+ subset
and marked by elevated expression of KLRD1 (CD94) and KLRB1
(CD161), molecules indicative of innate-like and NK cell-like
phenotypes (77). Lymphocyte
activation gene-3 (LAG-3) is an immune checkpoint that identifies
activated lymphocytes and may contribute to inflammation (78). Elevated frequencies of
LAG3+ cells, particularly within activated effector
memory T cell populations in the inflamed mucosa, associate
positively with disease activity in UC and genetic ablation of
LAG-3 depletes proliferating T cells, supporting its potential as a
therapeutic target (78).
Immune responses are coordinated by diverse
cellular participants, encompassing classical immunocytes, such as
macrophages, natural killer cells and T lymphocytes, as well as
structural cells including epithelial, endothelial and stromal
elements (81). Several studies
have focused on changes in immune cell composition during UC,
revealing increased infiltration levels of CD8+ T cells,
Tregs, macrophages, DCs, neutrophils and CD4+ T cells in
patients with UC compared with healthy individuals (44,82-84). To investigate immune infiltration
patterns in UC, Huang et al (85) acquired three transcriptional
datasets from the GEO database, identified differentially expressed
genes via linear models for micro-array data, and conducted
functional enrichment analyses to delineate associated biological
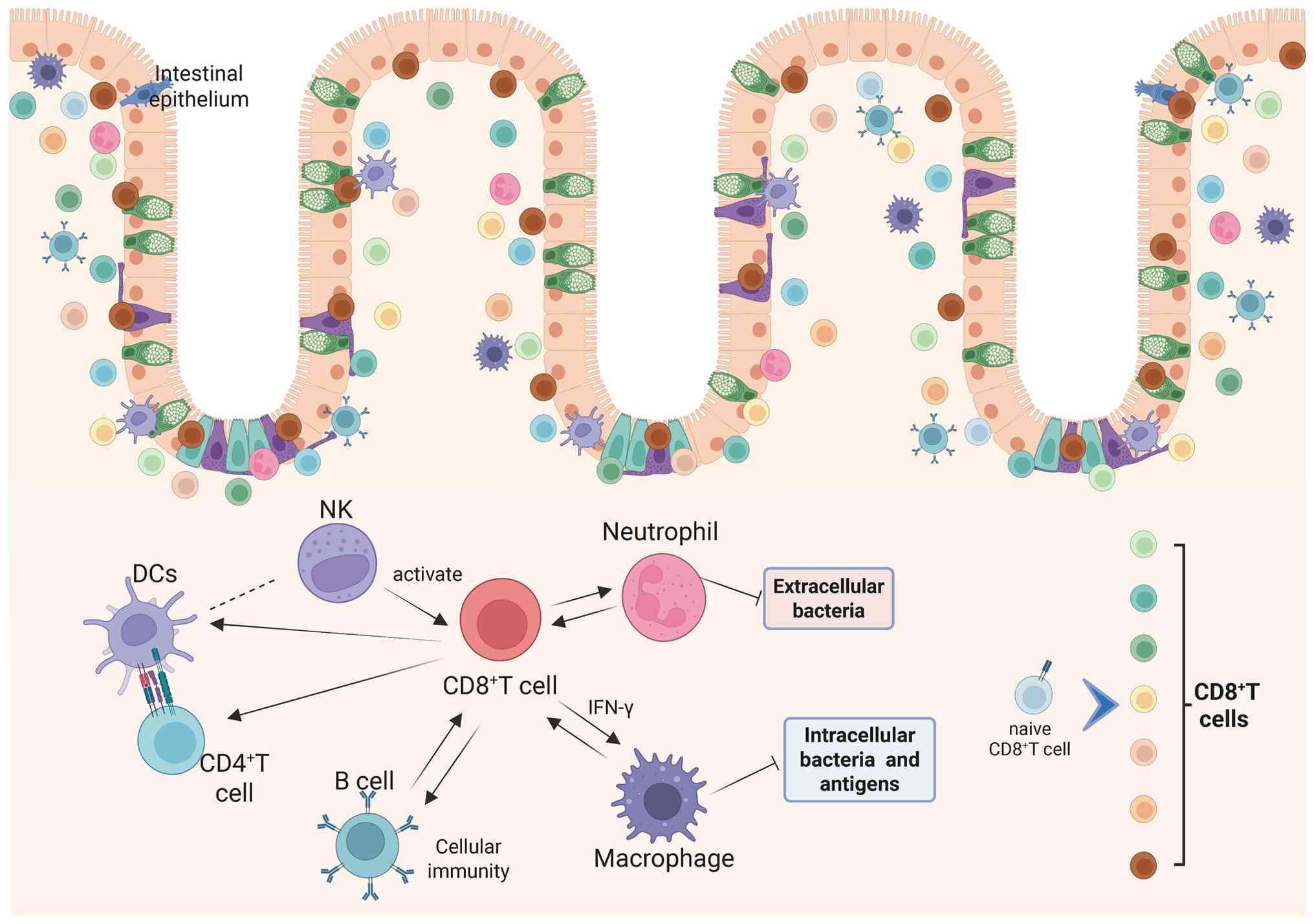
processes. A study comprising a correlation analyses of immune
cells indicated that CD8+ T cells, apart from being
unrelated to immature DCs, showed correlations with activated DCs,
macrophages, neutrophils, NK cells, B cells, Th1, Th2, TILs and
Tregs (85) (Fig. 3).
Patients with active CD exhibit notably elevated
immune scores compared with those with inactive disease,
characterized by a reduced abundance of CD8+ T cells,
alongside increased infiltration of M0 and M1 macrophages and
neutrophils, indicating distinct immune cell profiling between
disease states (89).
Correlation analyses of different immune cells in patients with CD
showed a negative correlation between CD8+ T cells and
immune scores, M0 and M1 macrophages, and neutrophils, whereas
there is a positive correlation between immune scores, M0 and M1
macrophages, and neutrophils (89). Uncontrolled neutrophil
accumulation in the intestinal lumen was associated with IBD
pathogenesis. Bone marrow-derived neutrophils traversed the
epithelium into the mucosal epithelium and subsequently entered the
intestinal lumen, forming an important defensive barrier that can
clear extracellular microorganisms (90). Neutrophils assisted in the
recruitment of other immune cells and promoted mucosal healing by
releasing anti-inflammatory factors, thereby attenuating the
inflammatory response (91).
Conversely, excessive neutrophils can exacerbate inflammation by
compromising the integrity of the intestinal epithelial barrier,
causing physical damage to the epithelium and secreting
pro-inflammatory cytokines (92).
NK group 2 member D (NKG2D) is one of the most
characteristic receptors shared by NK cells and T cells (93). This molecule contributes to IBD
pathogenesis via its expression on intestinal cytotoxic lymphocytes
and upregulated ligand presentation within inflamed mucosal tissues
(94,95). NKG2D inhibition enhanced the
migratory capacity and expansion of CD8+T cells in
co-culture systems with NKG2D-ligand-expressing astrocytes,
underscoring its functional involvement in CD8+ T
cell-astrocyte crosstalk (96).
Evidence indicated that invariant NKT (iNKT) cells and Tregs
suppressed DSS-induced colitis, whereas disease progression was
primarily driven by NK1.1+ CD8+ T cells, with
CD1d-restricted iNKT cells and CD1d-independent NK1.1+
CD8+ T cells interacting to disrupt IFN-γ homeostasis
and promoted intestinal inflammation, identifying NK1.1+
CD8+ T cells as potential therapeutic targets in human
IBD (97).
The study exploring the TRM cell populations in
colonic epithelial and lamina propria tissues adjacent to
mucosa-associated microbes indicated that the fundamental pathology
of human IBD involved a reduced response of CD8+ T cells
to commensal bacteria, leading to a deficiency of TRM cells in the
colon, and this was associated with chronic B cell activation and
excessive IgA secretion, linked to a loss of barrier immunity
(87). In chronic DSS-induced
colitis, pronounced B cell and modest CD8+ T cell
infiltration into enteric ganglia, unlike minimal infiltration in
acute DSS, TNBS or metastatic colitis, may drive visceral pain and
neuronal apoptosis, contributing to hypersensitivity in IBD
(98). IL-35 exerted
anti-inflammatory activity by enhancing regulatory B cell function,
which suppressed proliferation and the production of cytokines,
including IFN-γ, IL-17 and TNF-α, by autologous CD4+
CD25− T cells and CD8+ T cells, however, this
immunoregulatory mechanism was impaired in UC (99).
IBD is an intestinal immune disorder caused by
abnormal immune responses to environmental changes in genetically
predisposed individuals, where innate immunity and adaptive
immunity both play key roles in its onset and progression (6). Research has indicated that immune
regulatory dysfunction may represent the intrinsic core of IBD
pathogenesis, characterized by chronic intestinal inflammation and
tissue damage resulting from aberrant expression of
pro-inflammatory and anti-inflammatory factors in both innate and
adaptive immunity (114). The
medical treatment of IBD primarily encompasses 5-aminosalicylic
acid (5-ASA), glucocorticoids, immunomodulators, biologics,
antibiotics and stem cell transplantation, with ongoing
advancements in immune-related drug research and development driven
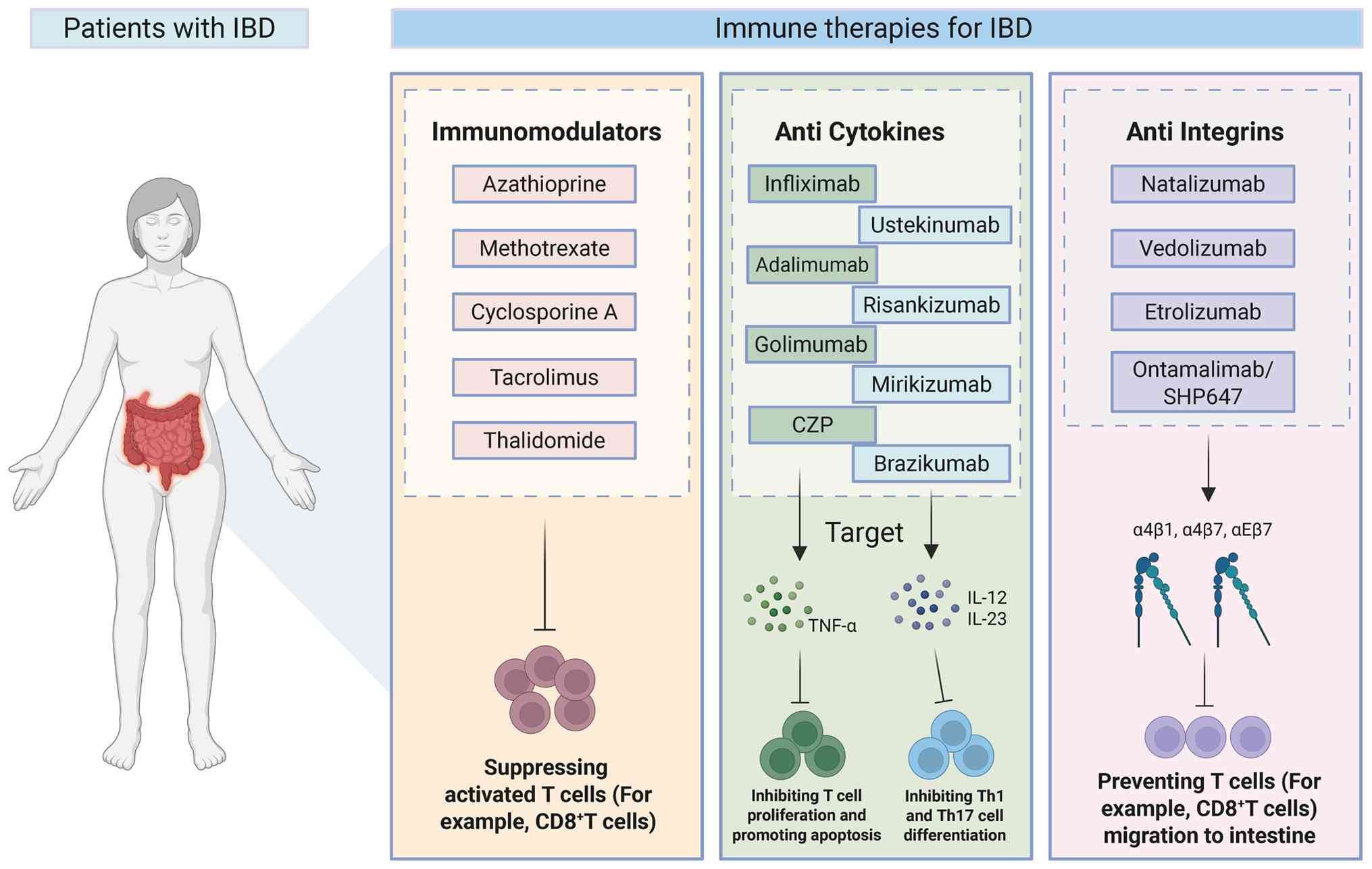
by in-depth studies on IBD pathogenesis (115,116). Immunomodulators and biologics
are currently widely utilized as common therapeutic agents for the
treatment of IBD (115,116) (Fig. 4).
Azathioprine (AZA), methotrexate (MTX),
cyclosporine A (CsA), tacrolimus and thalidomide are commonly used
immunomodulators in IBD (117,118). The active end-product of AZA,
6-thioguanine, incorporates into the DNA and RNA of T cells,
exerting inhibitory effects on cell activity and cytotoxicity;
however, due to its slow induction, AZA is recommended for use in
combination with TNF inhibitors (119). The 2023 European Crohn's and
Colitis Organisation (ECCO) guidelines recommend AZA monotherapy
for maintaining remission in steroid-dependent UC or in patients
intolerant to 5-ASA (120).
Randomized controlled trials have demonstrated that AZA was
superior to placebo in maintaining clinical remission (121,122). Low-dose MTX, while exhibiting
anti-inflammatory properties, is ineffective as a monotherapy for
inducing remission in CD or UC (123), and it is typically combined
with TNF inhibitors to achieve therapeutic efficacy (124). To the best of our knowledge,
there is no evidence supporting the use of MTX monotherapy for
maintaining remission in UC (125). A clinical study demonstrated
that MTX showed no superiority over placebo in achieving
steroid-free clinical remission (126). CsA and tacrolimus, both
calcineurin inhibitors, effectively suppress activated T cells and
demonstrate therapeutic efficacy in UC, although their long-term
use requires monitoring due to potential toxic side effects
(127,128). Thalidomide can serve as an
alternative treatment for pediatric refractory CD in cases of
secondary failure to anti-TNF therapy (129); however, its widespread use in
IBD is limited by adverse effects such as fatigue, somnolence,
peripheral neuropathy and the risk of long-term complications
including deep vein thrombosis and acute pancreatitis (130). According to the 2023 ECCO
guidelines, CsA, tacrolimus and thalidomide were not recommended as
conventional or first-line therapies for either UC or CD (120).
Ustekinumab (UST) selectively binds the common p40
subunit of IL-12 and IL-23, thereby suppressing Th1 and Th17 cell
differentiation and modulating T follicular helper cell
development, contributing to its efficacy in the acute and chronic
phases of inflammatory disease (141-143). UST exhibits a rapid onset of
action, considerably improving patient symptoms ≤7 days, with the
subcutaneous route of administration demonstrating relatively
superior efficacy (144). This
medication has been approved for the treatment of CD and UC. The
2023 ECCO guidelines recommended UST for the treatment of patients
with moderate-to-severe active UC who have an inadequate response
to, or are intolerant to, conventional therapy. A clinical study
demonstrated that UST provided benefit over placebo in both
inducing and maintaining clinical remission, improving clinical
response and ameliorating endoscopic scores in patients with
moderate-to-severe UC (145).
Risankizumab (RIS), mirikizumab and brazikumab, all targeting the
IL-23 p19 subunit, exhibit robust clinical efficacy in the
induction and maintenance of remission in IBD, accompanied by
favorable tolerability and safety profiles (146-148).
Integrins, adhesion molecules and signaling
proteins on the leukocytes' surface are activated through
intracellular signaling triggered by chemokines and other stimuli,
enabling leukocytes to migrate across the vascular wall and
infiltrate tissues such as the intestine, so antagonizing integrins
can block the binding of lymphocytes to adhesion molecules and
their migration into the gut, thereby alleviating local
inflammatory responses (118,149). Natalizumab, which effectively
targets α4β1 and α4β7 integrins, selectively inhibiting lymphocyte
migration to inflammatory sites by blocking α4 integrin signaling,
was the first drug approved for the treatment of CD (150). α4β7 expression facilitates the
infiltration of regulatory T cells into intestinal tissue, while
its inhibition diminishes the intestinal homing of regulatory and
effector T cells (151).
Adoptive transfer of α4-deficient T cells into immunodeficient mice
compromised their trafficking to inflammatory foci, leading to a
marked attenuation of chronic colitis severity (152). Compared with natalizumab,
vedolizumab specifically targets α4β7, selectively inhibiting
lymphocyte trafficking to the gut while exhibiting fewer systemic
adverse effects (153). The
GEMINI I phase 3 induction study demonstrated that a higher
proportion of patients treated with vedolizumab achieved endoscopic
remission compared with those receiving placebo (154). Therefore, it is also
recommended by the ECCO guidelines for inducing remission in
patients with moderate-to-severe active UC (120). Etrolizumab blocks cell adhesion
by targeting α4β7 and αEβ7 integrins (155), demonstrating considerable
efficacy in UC and CD, with favorable safety profiles for
subcutaneous administration. Ontamalimab/SHP647 binds to mucosal
addressin cell adhesion molecule (MAdCAM), preventing lymphocyte
migration to the gut and, thereby, attenuating inflammation, and
has shown potential for long-term maintenance of remission in UC
with a favorable safety profile (156).
With the continuous advancement of clinical and
basic research, the limitations of conventional pharmacological
therapies have become increasingly apparent and the development of
biosimilars and small-molecule drugs has been addressing the
shortcomings of previous treatments in previous years (115). Researchers may leverage
multi-omics technologies to identify potential biomarkers and
explore novel immunotherapeutic targets and drugs, while
integrating disciplines such as immunology and microbiology to
drive innovation and advancement in IBD immune-based therapies.
The integrated colonic barrier system is composed
of four essential elements: An immune barrier mediated by innate
and adaptive immune cells, a mechanical barrier formed by IECs and
tight junctions, a biological barrier dominated by the gut
microbiota, and a chemical barrier comprising the mucus layer (for
example, MUC proteins) and digestive enzymes (157). CD8+ T cells, as
essential immune cells, participate in the pathological damage of
IBD. However, their subpopulation composition, the signaling
pathways they influence, their interactions with other colonic
cells and how they dynamically remodel to impact inflammation in
IBD remain unclear. This could be a viable direction for future
research. Despite the various classifications of CD8+ T
cell subpopulations in IBD, considerable heterogeneity exists among
studies, and there is no unified definition or explanation of their
roles, which poses a challenge for readers. The descriptions in the
present review are relatively simple, and there is hope for more
standardized research in the future.
ScRNA-seq has unveiled novel insights into the
functional diversity and pathogenic mechanisms of CD8+ T
cells, offering promising avenues for elucidating their roles in
immunity and disease. First, the continued development and
refinement of single-cell technologies, particularly multiomics
approaches, will provide a more comprehensive understanding of
CD8+ T cell interactions with higher resolution and
depth. Furthermore, the integration of scRNA-seq with advanced
in vivo imaging techniques, such as intravital microscopy,
will allow for the visualization and real-time analysis of
CD8+ T cell dynamics within their native tissue
microenvironment, offering a clearer understanding of their spatial
and temporal interactions with other colonic cells, particularly in
the context of IBD. Furthermore, combining single-cell data with
ST, proteomics and metabolomics will provide a holistic view of
cellular and molecular interactions in the colon, elucidating the
complex interplay between CD8+ T cells and the colonic
microenvironment. In addition, conducting longitudinal studies to
track changes in CD8+ T cell interactions over time,
particularly during disease progression or therapeutic
interventions, will reveal their dynamic roles and regulatory
mechanisms, offering important insights into disease outcomes and
treatment responses. Finally, incorporating patient-derived samples
and organoid models will strengthen translational relevance by
closing the gap between fundamental research and clinical practice,
thereby accelerating the development of targeted therapies and
personalized treatment approaches for IBD and related
immune-mediated disorders. Together, these advancements will deepen
the understanding of CD8+ T cell biology and pave the
way for innovative diagnostic and therapeutic approaches.
This review summarizes the research advances of
single-cell transcriptomics in IBD over the past decade to unravel
the complex heterogeneity and functional dynamics of
CD8+ T cells, revealing their diverse phenotypes,
interactions with immune and non-immune cells, and pivotal roles in
inflammation and tissue homeostasis. The present review synthesizes
previous findings and highlights the therapeutic potential of
targeting specific CD8+ T cell subsets and their
microenvironmental interactions, offering new avenues for precision
medicine in IBD.
Not applicable.
RZ, JG and ZD wrote the original draft and prepared
the figures. LW and YH contributed to the conceptualization of the
review. Investigation was carried out by QQ and GL. WY and LZ
designed the tables and improved the manuscript. HW participated in
reviewing and editing the manuscript. All the authors have read and
approved the final manuscript. Data authentication not
applicable.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
Not applicable.
The present review was supported by the Young Scientists Project
Fund of the National Key Research and Development Program of China
(2025YFC3510700), the National Natural Science Foundation of China
Excellent Young Scientist Fund (grant no. 82422074), State
Administration of Traditional Chinese Medicine high-level key
discipline construction project (grant no. zyyzdxk-2023068) and
Qi-Huang Chief Scientist Project of National Administration of
Traditional Chinese Medicine (2025).
|
1
|
Bisgaard TH, Allin KH, Keefer L,
Ananthakrishnan AN and Jess T: Depression and anxiety in
inflammatory bowel disease: Epidemiology, mechanisms and treatment.
Nat Rev Gastroenterol Hepatol. 19:717–726. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Glassner KL, Abraham BP and Quigley EMM:
The microbiome and inflammatory bowel disease. J Allergy Clin
Immunol. 145:16–27. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Olfatifar M, Zali MR, Pourhoseingholi MA,
Balaii H, Ghavami SB, Ivanchuk M, Ivanchuk P, Nazari SH, Shahrokh
S, Sabour S, et al: The emerging epidemic of inflammatory bowel
disease in Asia and Iran by 2035: A modeling study. BMC
Gastroenterol. 21:2042021. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Agrawal M, Allin KH, Petralia F, Colombel
JF and Jess T: Multiomics to elucidate inflammatory bowel disease
risk factors and pathways. Nat Rev Gastroenterol Hepatol.
19:399–409. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhang YZ and Li YY: Inflammatory bowel
disease: Pathogenesis. World J Gastroenterol. 20:91–99. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Geremia A, Biancheri P, Allan P, Corazza
GR and Di Sabatino A: Innate and adaptive immunity in inflammatory
bowel disease. Autoimmun Rev. 13:3–10. 2014. View Article : Google Scholar
|
|
7
|
Taniuchi I: CD4 helper and CD8 cytotoxic T
cell differentiation. Annu Rev Immunol. 36:579–601. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Philip M and Schietinger A: CD8+ T cell
differentiation and dysfunction in cancer. Nat Rev Immunol.
22:209–223. 2022. View Article : Google Scholar :
|
|
9
|
Funderburg NT, Stubblefield Park SR, Sung
HC, Hardy G, Clagett B, Ignatz-Hoover J, Harding CV, Fu P, Katz JA,
Lederman MM and Levine AD: Circulating CD4(+) and CD8(+) T cells
are activated in inflammatory bowel disease and are associated with
plasma markers of inflammation. Immunology. 140:87–97. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Rabe H, Malmquist M, Barkman C, Östman S,
Gjertsson I, Saalman R and Wold AE: Distinct patterns of naive,
activated and memory T and B cells in blood of patients with
ulcerative colitis or Crohn's disease. Clin Exp Immunol.
197:111–129. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Stockenhuber K, Hegazy AN, West NR, Ilott
NE, Stockenhuber A, Bullers SJ, Thornton EE, Arnold IC, Tucci A,
Waldmann H, et al: Foxp3+ T reg cells control psoriasiform
inflammation by restraining an IFN-I-driven CD8+ T cell response. J
Exp Med. 215:1987–1998. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Mueller SN and Mackay LK: Tissue-resident
memory T cells: Local specialists in immune defence. Nat Rev
Immunol. 16:79–89. 2016. View Article : Google Scholar
|
|
13
|
Xi NM and Li JJ: Benchmarking
computational doublet-detection methods for single-cell RNA
sequencing data. Cell Syst. 12:176–194.e6. 2021. View Article : Google Scholar
|
|
14
|
Ding J, Sharon N and Bar-Joseph Z:
Temporal modelling using single-cell transcriptomics. Nat Rev
Genet. 23:355–368. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Armingol E, Officer A, Harismendy O and
Lewis NE: Deciphering cell-cell interactions and communication from
gene expression. Nat Rev Genet. 22:71–88. 2021. View Article : Google Scholar
|
|
16
|
Huang Q, Wang F, Hao D, Li X, Li X, Lei T,
Yue J and Liu C: Deciphering tumor-infiltrating dendritic cells in
the single-cell era. Exp Hematol Oncol. 12:972023. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Liu Y, Lan Q, Lu L, Chen M, Xia Z, Ma J,
Wang J, Fan H, Shen Y, Ryffel B, et al: Phenotypic and functional
characteristic of a newly identified CD8+ Foxp3-CD103+ regulatory T
cells. J Mol Cell Biol. 6:81–92. 2014. View Article : Google Scholar
|
|
18
|
Bottois H, Ngollo M, Hammoudi N, Courau T,
Bonnereau J, Chardiny V, Grand C, Gergaud B, Allez M and Le Bourhis
L: KLRG1 and CD103 expressions define distinct intestinal
tissue-resident memory CD8 T cell subsets modulated in Crohn's
Disease. Front Immunol. 11:8962020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Boland BS, He Z, Tsai MS, Olvera JG,
Omilusik KD, Duong HG, Kim ES, Limary AE, Jin W, Milner JJ, et al:
Heterogeneity and clonal relationships of adaptive immune cells in
ulcerative colitis revealed by single-cell analyses. Sci Immunol.
5:eabb44322020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Corridoni D, Chapman T, Antanaviciute A,
Satsangi J and Simmons A: Inflammatory bowel disease through the
lens of single-cell RNA-seq technologies. Inflamm Bowel Dis.
26:1658–1668. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
St Paul M and Ohashi PS: The roles of CD8+
T cell subsets in antitumor immunity. Trends Cell Biol. 30:695–704.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Smith TRF and Kumar V: Revival of CD8+
Treg-mediated suppression. Trends Immunol. 29:337–342. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yu Y, Ma X, Gong R, Zhu J, Wei L and Yao
J: Recent advances in CD8+ regulatory T cell research. Oncol Lett.
15:8187–8194. 2018.PubMed/NCBI
|
|
24
|
Stubbington MJT, Rozenblatt-Rosen O, Regev
A and Teichmann SA: Single-cell transcriptomics to explore the
immune system in health and disease. Science. 358:58–63. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Tanay A and Regev A: Scaling single-cell
genomics from phenomenology to mechanism. Nature. 541:331–338.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Kong L, Pokatayev V, Lefkovith A, Carter
GT, Creasey EA, Krishna C, Subramanian S, Kochar B, Ashenberg O,
Lau H, et al: The landscape of immune dysregulation in Crohn's
disease revealed through single-cell transcriptomic profiling in
the ileum and colon. Immunity. 56:28552023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Devlin JC, Axelrad J, Hine AM, Chang S,
Sarkar S, Lin JD, Ruggles KV, Hudesman D, Cadwell K and Loke P:
Single-cell transcriptional survey of ileal-anal pouch immune cells
from ulcerative colitis patients. Gastroenterology. 160:1679–1693.
2021. View Article : Google Scholar
|
|
28
|
Gupta T, Antanaviciute A, Hyun-Jung Lee C,
Ottakandathil Babu R, Aulicino A, Christoforidou Z,
Siejka-Zielinska P, O'Brien-Ball C, Chen H, Fawkner-Corbett D, et
al: Tracking in situ checkpoint inhibitor-bound target T cells in
patients with checkpoint-induced colitis. Cancer Cell.
42:797–814.e15. 2024. View Article : Google Scholar
|
|
29
|
Kolachala VL, Maddipatla SC, Murthy S,
Hwang Y, Dodd AF, Sharma G, Munasinghe S, Pelia RS, Venkateswaran
S, Anbazhagan M, et al: Altered inflammatory mucosal signatures
within their spatial and cellular context during active ileal
Crohn's disease. JCI Insight. 10:e1717832025. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Mitsialis V, Wall S, Liu P,
Ordovas-Montanes J, Parmet T, Vukovic M, Spencer D, Field M,
McCourt C, Toothaker J, et al: Single-cell analyses of colon and
blood reveal distinct immune cell signatures of ulcerative colitis
and Crohn's disease. Gastroenterology. 159:591–608.e10. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Becker W, Alrafas HR, Busbee PB, Walla MD,
Wilson K, Miranda K, Cai G, Putluri V, Putluri N, Nagarkatti M and
Nagarkatti PS: Cannabinoid receptor activation on haematopoietic
cells and enterocytes protects against colitis. J Crohns Colitis.
15:1032–1048. 2021. View Article : Google Scholar :
|
|
32
|
Ahmed A, Joseph AM, Zhou J, Horn V, Uddin
J, Lyu M, Goc J; JRI Live Cell Bank; Sockolow RE, Wing JB, et al:
CTLA-4-expressing ILC3s restrain interleukin-23-mediated
inflammation. Nature. 630:976–983. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Gordon H, Wichmann K, Lewis A, Sanders T,
Wildemann M, Hoti I, Hornsby E, Kok KB, Silver A, Lindsay JO and
Stagg AJ: Human intestinal dendritic cells can overcome retinoic
acid signaling to generate proinflammatory CD4 T cells with both
gut and skin homing properties. J Immunol. 212:96–106. 2024.
View Article : Google Scholar
|
|
34
|
Cao S, Nguyen KM, Ma K, Du X, Liu X,
Ulezko Antonova A, Rood RP, Gremida A, Chen CH, Gutierrez A, et al:
Mucosal single-cell profiling of Crohn's-like sisease of the pouch
reveals unique pathogenesis and therapeutic targets.
Gastroenterology. 167:1399–1414.e2. 2024. View Article : Google Scholar
|
|
35
|
Bolton C, Smillie CS, Pandey S, Elmentaite
R, Wei G, Argmann C, Aschenbrenner D, James KR, McGovern DPB,
Macchi M, et al: An integrated taxonomy for monogenic inflammatory
bowel disease. Gastroenterology. 162:859–876. 2022. View Article : Google Scholar
|
|
36
|
Dennison TW, Edgar RD, Payne F, Nayak KM,
Ross ADB, Cenier A, Glemas C, Giachero F, Foster AR, Harris R, et
al: Patient-derived organoid biobank identifies epigenetic
dysregulation of intestinal epithelial MHC-I as a novel mechanism
in severe Crohn's Disease. Gut. 73:1464–1477. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Huang B, Chen Z, Geng L, Wang J, Liang H,
Cao Y, Chen H, Huang W, Su M, Wang H, et al: Mucosal profiling of
pediatric-onset colitis and IBD reveals common pathogenics and
therapeutic pathways. Cell. 179:1160–1176.e24. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Kaech SM and Cui W: Transcriptional
control of effector and memory CD8+ T cell differentiation. Nat Rev
Immunol. 12:749–761. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Boles KS, Barchet W, Diacovo T, Cella M
and Colonna M: The tumor suppressor TSLC1/NECL-2 triggers NK-cell
and CD8+ T-cell responses through the cell-surface receptor CRTAM.
Blood. 106:779–786. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Intlekofer AM, Takemoto N, Kao C, Banerjee
A, Schambach F, Northrop JK, Shen H, Wherry EJ and Reiner SL:
Requirement for T-bet in the aberrant differentiation of unhelped
memory CD8+ T cells. J Exp Med. 204:2015–2021. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Joshi NS, Cui W, Chandele A, Lee HK, Urso
DR, Hagman J, Gapin L and Kaech SM: Inflammation directs memory
precursor and short-lived effector CD8(+) T cell fates via the
graded expression of T-bet transcription factor. Immunity.
27:281–295. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Intlekofer AM, Takemoto N, Wherry EJ,
Longworth SA, Northrup JT, Palanivel VR, Mullen AC, Gasink CR,
Kaech SM, Miller JD, et al: Effector and memory CD8+ T cell fate
coupled by T-bet and eomesodermin. Nat Immunol. 6:1236–1244. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Intlekofer AM, Banerjee A, Takemoto N,
Gordon SM, Dejong CS, Shin H, Hunter CA, Wherry EJ, Lindsten T and
Reiner SL: Anomalous type 17 response to viral infection by CD8+ T
cells lacking T-bet and eomesodermin. Science. 321:408–411. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Smillie CS, Biton M, Ordovas-Montanes J,
Sullivan KM, Burgin G, Graham DB, Herbst RH, Rogel N, Slyper M,
Waldman J, et al: Intra- and inter-cellular rewiring of the human
colon during ulcerative colitis. Cell. 178:714–730.e22. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhao Q, Zhang T and Yang H: ScRNA-seq
identified the metabolic reprogramming of human colonic immune
cells in different locations and disease states. Biochem Biophys
Res Commun. 604:96–103. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Jaeger N, Gamini R, Cella M, Schettini JL,
Bugatti M, Zhao S, Rosadini CV, Esaulova E, Di Luccia B, Kinnett B,
et al: Single-cell analyses of Crohn's disease tissues reveal
intestinal intraepithelial T cells heterogeneity and altered subset
distributions. Nat Commun. 12:19212021. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Jiang T, Zheng J, Li N, Li X, He J, Zhou
J, Sun B and Chi Q: Dissecting the mechanisms of intestinal immune
homeostasis by analyzing T-cell immune response in Crohn's disease
and colorectal cancer. Curr Gene Ther. 24:422–440. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Roda G, Jianyu X, Park MS, DeMarte L,
Hovhannisyan Z, Couri R, Stanners CP, Yeretssian G and Mayer L:
Characterizing CEACAM5 interaction with CD8α and CD1d in intestinal
homeostasis. Mucosal Immunol. 7:615–624. 2014. View Article : Google Scholar :
|
|
49
|
Guo X, Zhang Y, Zheng L, Zheng C, Song J,
Zhang Q, Kang B, Liu Z, Jin L, Xing R, et al: Global
characterization of T cells in non-small-cell lung cancer by
single-cell sequencing. Nat Med. 24:978–985. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Bassez A, Vos H, Van Dyck L, Floris G,
Arijs I, Desmedt C, Boeckx B, Vanden Bempt M, Nevelsteen I, Lambein
K, et al: A single-cell map of intratumoral changes during anti-PD1
treatment of patients with breast cancer. Nat Med. 27:820–832.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Eberhardt CS, Kissick HT, Patel MR,
Cardenas MA, Prokhnevska N, Obeng RC, Nasti TH, Griffith CC, Im SJ,
Wang X, et al: Functional HPV-specific PD-1+ stem-like CD8 T cells
in head and neck cancer. Nature. 597:279–284. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zheng CH, Zheng LT, Yoo JK, Guo H, Zhang
Y, Guo X, Kang B, Hu R, Huang JY, Zhang Q, et al: Landscape of
infiltrating T cells in liver cancer revealed by single-cell
sequencing. Cell. 169:1342–1356.e16. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Wang K, Coutifaris P, Brocks D, Wang G,
Azar T, Solis S, Nandi A, Anderson S, Han N, Manne S, et al:
Combination anti-PD-1 and anti-CTLA-4 therapy generates waves of
clonal responses that include progenitor-exhausted CD8+ T cells.
Cancer Cell. 42:1582–1597.e10. 2024. View Article : Google Scholar
|
|
54
|
Oh DY, Kwek SS, Raju SS, Li T, McCarthy E,
Chow E, Aran D, Ilano A, Pai CS, Rancan C, et al: Intratumoral CD4+
T cells mediate anti-tumor cytotoxicity in human bladder cancer.
Cell. 181:1612–1625.e13. 2020. View Article : Google Scholar
|
|
55
|
Li J, Wu C, Hu H, Qin G, Wu X, Bai F,
Zhang J, Cai Y, Huang Y, Wang C, et al: Remodeling of the immune
and stromal cell compartment by PD-1 blockade in mismatch
repair-deficient colorectal cancer. Cancer Cell. 41:1152–1169.e7.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Corridoni D, Antanaviciute A, Gupta T,
Fawkner-Corbett D, Aulicino A, Jagielowicz M, Parikh K, Repapi E,
Taylor S, Ishikawa D, et al: Single-cell atlas of colonic CD8+ T
cells in ulcerative colitis. Nat Med. 26:1480–1490. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Globig AM, Mayer LS, Heeg M, Andrieux G,
Ku M, Otto-Mora P, Hipp AV, Zoldan K, Pattekar A, Rana N, et al:
Exhaustion of CD39-expressing CD8+ T cells in Crohn's disease is
linked to clinical outcome. Gastroenterology. 163:965–981.e31.
2022. View Article : Google Scholar
|
|
58
|
Liao M, Liu Y, Yuan J, Wen Y, Xu G, Zhao
J, Cheng L, Li J, Wang X, Wang F, et al: Single-cell landscape of
bronchoalveolar immune cells in patients with COVID-19. Nat Med.
26:842–844. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Karmele EP, Moldoveanu AL, Kaymak I,
Jugder BE, Ursin RL, Bednar KJ, Corridoni D and Ort T: Single cell
RNA-sequencing profiling to improve the translation between human
IBD and in vivo models. Front Immunol. 14:12919902023. View Article : Google Scholar
|
|
60
|
Sun L, Su Y, Jiao A, Wang X and Zhang B: T
cells in health and disease. Signal Transduct Target Ther.
8:2352023. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Fox CJ, Hammerman PS and Thompson CB: Fuel
feeds function: energy metabolism and the T-cell response. Nat Rev
Immunol. 5:844–852. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Araki K, Morita M, Bederman AG, Konieczny
BT, Kissick HT, Sonenberg N and Ahmed R: Translation is actively
regulated during the differentiation of CD8+ effector T cells. Nat
Immunol. 18:1046–1057. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Stary G, Olive A, Radovic-Moreno AF,
Gondek D, Alvarez D, Basto PA, Perro M, Vrbanac VD, Tager AM, Shi
J, et al: A mucosal vaccine against Chlamydia trachomatis generates
two waves of protective memory T cells. Science. 348:aaa82052015.
View Article : Google Scholar
|
|
64
|
Laidlaw BJ, Craft JE and Kaech SM: The
multifaceted role of CD4+ T cells in CD8+ T cell memory. Nat Rev
Immunol. 16:102–111. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
He JY, Kim YJ, Mennillo E, Rusu I, Bain J,
Rao AA, Andersen C, Law K, Yang H, Tsui J, et al: Dysregulation of
CD4+ and CD8+ resident memory T, myeloid, and stromal cells in
steroid-experienced, checkpoint inhibitor colitis. J Immunother
Cancer. 12:e0086282024. View Article : Google Scholar
|
|
66
|
Qu R, Kluger Y, Yang J, Zhao J, Hafler DA,
Krause DS, Bersenev A, Bosenberg M, Hurwitz M, Lucca L, et al:
Longitudinal single-cell analysis of a patient receiving adoptive
cell therapy reveals potential mechanisms of treatment failure. Mol
Cancer. 21:2192022. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Nagahara N: Multiple role of
3-mercaptopyruvate sulfurtransferase: antioxidative function, H2 S
and polysulfide production and possible SOx production. Br J
Pharmacol. 175:577–589. 2018. View Article : Google Scholar
|
|
68
|
Bourgonje AR, Kloska D, Grochot-Przęczek
A, Feelisch M, Cuadrado A and van Goor H: Personalized redox
medicine in inflammatory bowel diseases: An emerging role for
HIF-1α and NRF2 as therapeutic targets. Redox Biol. 60:1026032023.
View Article : Google Scholar
|
|
69
|
Bálint Š, Müller S, Fischer R, Kessler BM,
Harkiolaki M, Valitutti S and Dustin ML: Supramolecular attack
particles are autonomous killing entities released from cytotoxic T
cells. Science. 368:897–901. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Shi Y, Lu Y and You J: Unfolded protein
response in the activation-induced biological processes of CD8+ T
cells. Pharmacol Res. 169:1056542021. View Article : Google Scholar
|
|
71
|
Kamimura D and Bevan MJ: Endoplasmic
reticulum stress regulator XBP-1 contributes to effector CD8+ T
cell differentiation during acute infection. J Immunol.
181:5433–5441. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Ma X, Bi E, Lu Y, Su P, Huang C, Liu L,
Wang Q, Yang M, Kalady MF, Qian J, et al: Cholesterol induces CD8+
T cell exhaustion in the tumor microenvironment. Cell Metab.
30:143–156.e5. 2019. View Article : Google Scholar
|
|
73
|
Cao Y, Trillo-Tinoco J, Sierra RA, Anadon
C, Dai W, Mohamed E, Cen L, Costich TL, Magliocco A, Marchion D, et
al: ER stress-induced mediator C/EBP homologous protein thwarts
effector T cell activity in tumors through T-bet repression. Nat
Commun. 10:12802019. View Article : Google Scholar
|
|
74
|
Tammaro A, Stroo I, Rampanelli E, Blank F,
Butter LM, Claessen N, Takai T, Colonna M, Leemans JC, Florquin S
and Dessing MC: Role of TREM1-DAP12 in renal inflammation during
obstructive nephropathy. PloS One. 8:e824982013. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Li X, Lee EJ, Gawel DR, Lilja S, Schäfer
S, Zhang H and Benson M: Meta-analysis of expression profiling data
indicates need for combinatorial biomarkers in pediatric ulcerative
colitis. J Immunol Res. 2020:82796192020. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Markov AV, Savin IA, Zenkova MA and
Sen'kova AV: Identification of novel core genes involved in
malignant transformation of inflamed colon tissue using a
computational biology approach and verification in murine models.
Int J Mol Sci. 24:43112023. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Rosati E, Rios Martini G, Pogorelyy MV,
Minervina AA, Degenhardt F, Wendorff M, Sari S, Mayr G, Fazio A,
Dowds CM, et al: A novel unconventional T cell population enriched
in Crohn's disease. Gut. 71:2194–2204. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Slevin SM, Garner LC, Lahiff C, Tan M,
Wang LM, Ferry H, Greenaway B, Lynch K, Geremia A, Hughes S, et al:
Lymphocyte activation gene (LAG)-3 is associated with mucosal
inflammation and disease activity in ulcerative colitis. J Crohns
Colitis. 14:1446–1461. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Dai Z, Zhang J, Xu W, Du P, Wang Z and Liu
Y: Single-cell sequencing-based validation of T cell-associated
diagnostic model genes and drug response in Crohn's disease. Int J
Mol Sci. 24:60542023. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Huang LJ, Mao XT, Li YY, Liu DD, Fan KQ,
Liu RB, Wu TT, Wang HL, Zhang Y, Yang B, et al: Multiomics analyses
reveal a critical role of selenium in controlling T cell
differentiation in Crohn's disease. Immunity. 54:1728–1744.e7.
2021. View Article : Google Scholar
|
|
81
|
Medzhitov R: Recognition of microorganisms
and activation of the immune response. Nature. 449:819–826. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Tom MR, Li J, Ueno A, Fort Gasia M, Chan
R, Hung DY, Chenoo S, Iacucci M, Jijon HB, Kaplan GG, et al: Novel
CD8+ T-cell subsets demonstrating plasticity in patients with
inflammatory bowel disease. Inflamm Bowel Dis. 22:1596–1608. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Holmén N, Lundgren A, Lundin S, Bergin AM,
Rudin A, Sjövall H and Ohman L: Functional CD4+CD25high regulatory
T cells are enriched in the colonic mucosa of patients with active
ulcerative colitis and increase with disease activity. Inflamm
Bowel Dis. 12:447–456. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Huang J, Zhang J, Wang F, Zhang B and Tang
X: Comprehensive analysis of cuproptosis-related genes in immune
infiltration and diagnosis in ulcerative colitis. Front Immunol.
13:10081462022. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Huang J, Zhang J, Wang F, Zhang B and Tang
X: Revealing immune infiltrate characteristics and potential
diagnostic value of immune-related genes in ulcerative colitis: An
integrative genomic analysis. Front Public Health. 10:10030022022.
View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Chen H, Chen C, Yuan X, Xu W, Yang MQ, Li
Q, Shen Z and Yin L: Identification of immune cell landscape and
construction of a novel diagnostic nomogram for Crohn's disease.
Front Genet. 11:4232020. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Noble A, Durant L, Hoyles L, Mccartney AL,
Man R, Segal J, Costello SP, Hendy P, Reddi D, Bouri S, et al:
Deficient resident memory T cell and CD8 T cell response to
commensals in inflammatory bowel disease. J Crohns Colitis.
14:525–537. 2020. View Article : Google Scholar :
|
|
88
|
Corridoni D, Shiraishi S, Chapman T,
Steevels T, Muraro D, Thézénas ML, Prota G, Chen JL, Gileadi U,
Ternette N, et al: NOD2 and TLR2 signal via TBK1 and PI31 to direct
cross-presentation and CD8 T cell responses. Front Immunol.
10:9582019. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Zhong WM, Qian XH and Jin ZW:
Identification of potential predictive biomarkers and biological
pathways and the correction with immune infiltration in the
activation of Crohn's disease. Immunogenetics. 74:527–537. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Brazil JC, Louis NA and Parkos CA: The
role of polymorphonuclear leukocyte trafficking in the perpetuation
of inflammation during inflammatory bowel disease. Inflamm Bowel
Dis. 19:1556–1565. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Zhang Q, Zhang C, Chang F, Liang K, Yin X,
Li X, Zhao K, Niu Q and Tian Z: Wip 1 inhibits intestinal
inflammation in inflammatory bowel disease. Cell Immunol.
310:63–70. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Mantovani A, Cassatella MA, Costantini C
and Jaillon S: Neutrophils in the activation and regulation of
innate and adaptive immunity. Nat Rev Immunol. 11:519–531. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Alkhayer R, Ponath V, Frech M, Adhikary T,
Graumann J, Neubauer A and von Strandmann EP: KLF4-mediated
upregulation of the NKG2D ligand MICA in acute myeloid leukemia: A
novel therapeutic target identified by enChIP. Cell Commun Signal.
21:942023. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Vadstrup K and Bendtsen F: Anti-NKG2D mAb:
A new treatment for Crohn's disease? Int J Mol Sci. 18:19972017.
View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Allez M, Tieng V, Nakazawa A, Treton X,
Pacault V, Dulphy N, Caillat-Zucman S, Paul P, Gornet JM, Douay C,
et al: CD4+NKG2D+ T cells in Crohn's disease mediate inflammatory
and cytotoxic responses through MICA interactions.
Gastroenterology. 132:2346–2358. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Wei L, Xiang Z and Zou Y: The role of
NKG2D and its ligands in autoimmune diseases: new targets for
immunotherapy. Int J Mol Sci. 24:175452023. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Lee SW, Park HJ, Cheon JH, Wu L, Van Kaer
L and Hong S: iNKT cells suppress pathogenic NK1.1+CD8+ T cells in
DSS-induced colitis. Front Immunol. 9:21682018. View Article : Google Scholar
|
|
98
|
Wiese JJ, Manna S, Kühl AA, Fascì A,
Elezkurtaj S, Sonnenberg E, Bubeck M, Atreya R, Becker C, Weixler
B, et al: Myenteric plexus immune cell infiltrations and
neurotransmitter expression in Crohn's disease and ulcerative
colitis. J Crohns Colitis. 18:121–133. 2024. View Article : Google Scholar :
|
|
99
|
Wang S and Qin C: Interleukin 35 rescues
regulatory B cell function, but the effect is dysregulated in
ulcerative colitis. DNA Cell Biol. 36:413–421. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Zundler S, Schillinger D, Fischer A,
Atreya R, López-Posadas R, Watson A, Neufert C, Atreya I and
Neurath MF: Blockade of αEβ7 integrin suppresses accumulation of
CD8+ and Th9 lymphocytes from patients with IBD in the inflamed gut
in vivo. Gut. 66:1936–1948. 2017. View Article : Google Scholar
|
|
101
|
Nancey S, Holvöet S, Graber I, Joubert G,
Philippe D, Martin S, Nicolas JF, Desreumaux P, Flourié B and
Kaiserlian D: CD8+ cytotoxic T cells induce relapsing colitis in
normal mice. Gastroenterology. 131:485–496. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Müller S, Lory J, Corazza N, Griffiths GM,
Z'graggen K, Mazzucchelli L, Kappeler A and Mueller C: Activated
CD4+ and CD8+ cytotoxic cells are present in increased numbers in
the intestinal mucosa from patients with active inflammatory bowel
disease. Am J Pathol. 152:261–268. 1998.PubMed/NCBI
|
|
103
|
Palmen MJ, Wijburg OL, Kunst IH, Kroes H
and van Rees EP: CD4+ T cells from 2,4,6-trinitrobenzene sulfonic
acid (TNBS)-induced colitis rodents migrate to the recipient's
colon upon transfer; down-regulation by CD8+ T cells. Clin Exp
Immunol. 112:216–225. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Teh HS, Phillips RA and Miller RG:
Quantitative studies on the precursors of cytotoxic lymphocytes. V.
The cellular basis for the cross-reactivity of TNP-specific clones.
J Immunol. 121:1711–1717. 1978. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Qiu Y and Yang H: Effects of
intraepithelial lymphocyte-derived cytokines on intestinal mucosal
barrier function. J Interferon Cytokine Res. 33:551–562. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Bhat P, Leggatt G, Waterhouse N and Frazer
IH: Interferon-γ derived from cytotoxic lymphocytes directly
enhances their motility and cytotoxicity. Cell Death Dis.
8:e28362017. View Article : Google Scholar
|
|
107
|
Takashima S, Martin ML, Jansen SA, Fu Y,
Bos J, Chandra D, O'Connor MH, Mertelsmann AM, Vinci P, Kuttiyara
J, et al: T cell-derived interferon-γ programs stem cell death in
immune-mediated intestinal damage. Sci Immunol. 4:eaay85562019.
View Article : Google Scholar
|
|
108
|
Tang F, Sally B, Lesko K, Discepolo V,
Abadie V, Ciszewski C, Semrad C, Guandalini S, Kupfer SS and Jabri
B: Cysteinyl leukotrienes mediate lymphokine killer activity
induced by NKG2D and IL-15 in cytotoxic T cells during celiac
disease. J Exp Med. 212:1487–1495. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Igalouzene R, Hernandez-Vargas H, Benech
N, Guyennon A, Bauché D, Barrachina C, Dubois E, Marie JC and
Soudja SM: SMAD4 TGF-β-independent function preconditions naive
CD8+ T cells to prevent severe chronic intestinal inflammation. J
Clin Invest. 132:e1510202022. View Article : Google Scholar
|
|
110
|
Meresse B, Chen Z, Ciszewski C, Tretiakova
M, Bhagat G, Krausz TN, Raulet DH, Lanier LL, Groh V, Spies T, et
al: Coordinated induction by IL15 of a TCR-independent NKG2D
signaling pathway converts CTL into lymphokine-activated killer
cells in celiac disease. Immunity. 21:357–366. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Bisping G, Lügering N, Lütke-Brintrup S,
Pauels HG, Schürmann G, Domschke W and Kucharzik T: Patients with
inflammatory bowel disease (IBD) reveal increased induction
capacity of intracellular interferon-gamma (IFN-gamma) in
peripheral CD8+ lymphocytes co-cultured with intestinal epithelial
cells. Clin Exp Immunol. 123:15–22. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Nunes NS, Kim S, Sundby M, Chandran P,
Burks SR, Paz AH and Frank JA: Temporal clinical, proteomic,
histological and cellular immune responses of dextran sulfate
sodium-induced acute colitis. World J Gastroenterol. 24:4341–4355.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Ito R, Shin-Ya M, Kishida T, Urano A,
Takada R, Sakagami J, Imanishi J, Kita M, Ueda Y, Iwakura Y, et al:
Interferon-gamma is causatively involved in experimental
inflammatory bowel disease in mice. Clin Exp Immunol. 146:330–338.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Aliberti J: Immunity and tolerance induced
by intestinal mucosal dendritic cells. Mediators Inflamm.
2016:31047272016. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Adamina M, Bonovas S, Raine T, Spinelli A,
Warusavitarne J, Armuzzi A, Bachmann O, Bager P, Biancone L,
Bokemeyer B, et al: ECCO guidelines on therapeutics in Crohn's
disease: surgical treatment. J Crohns Colitis. 14:155–168. 2020.
View Article : Google Scholar
|
|
116
|
Spinelli A, Bonovas S, Burisch J,
Kucharzik T, Adamina M, Annese V, Bachmann O, Bettenworth D,
Chaparro M, Czuber-Dochan W, et al: ECCO guidelines on therapeutics
in ulcerative colitis: surgical treatment. J Crohns Colitis.
16:179–189. 2022. View Article : Google Scholar
|
|
117
|
Löwenberg M, Volkers A, van Gennep S,
Mookhoek A, Montazeri N, Clasquin E, Duijvestein M, van Bodegraven
A, Rietdijk S, Jansen J, et al: Mercaptopurine for the treatment of
ulcerative colitis: A randomized placebo-controlled trial. J Crohns
Colitis. 17:1055–1065. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Vieujean S, Jairath V, Peyrin-Biroulet L,
Dubinsky M, Iacucci M, Magro F and Danese S: Understanding the
therapeutic toolkit for inflammatory bowel disease. Nat Rev
Gastroenterol Hepatol. 22:371–394. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Turbayne AK and Sparrow MP: Low-dose
azathioprine in combination with allopurinol: The past, present and
future of this useful duo. Dig Dis Sci. 67:5382–5391. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Raine T, Bonovas S, Burisch J, Kucharzik
T, Adamina M, Annese V, Bachmann O, Bettenworth D, Chaparro M,
Czuber-Dochan W, et al: ECCO guidelines on therapeutics in
ulcerative colitis: medical treatment. J Crohns Colitis. 16:2–17.
2022. View Article : Google Scholar
|
|
121
|
Sood A, Midha V, Sood N and Kaushal V:
Role of azathioprine in severe ulcerative colitis: one-year,
placebo-controlled, randomized trial. Indian J Gastroenterol.
19:14–16. 2000.PubMed/NCBI
|
|
122
|
Sood A, Kaushal V, Midha V, Bhatia KL,
Sood N and Malhotra V: The beneficial effect of azathioprine on
maintenance of remission in severe ulcerative colitis. J
Gastroenterol. 37:270–274. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Nielsen OH, Steenholdt C, Juhl CB and
Rogler G: Efficacy and safety of methotrexate in the management of
inflammatory bowel disease: A systematic review and meta-analysis
of randomized, controlled trials. EClinicalMedicine. 20:1002712020.
View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Borren NZ, Luther J, Colizzo FP, Garber
JG, Khalili H and Ananthakrishnan AN: Low-dose methotrexate has
similar outcomes to high-dose methotrexate in combination with
anti-TNF therapy in inflammatory bowel diseases. J Crohns Colitis.
13:990–995. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Raine T, Verstockt B, Kopylov U, Karmiris
K, Goldberg R, Atreya R, Burisch J, Burke J, Ellul P, Hedin C, et
al: ECCO topical review: refractory inflammatory bowel disease. J
Crohns Colitis. 15:1605–1620. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Herfarth H, Barnes EL, Valentine JF,
Hanson J, Higgins PDR, Isaacs KL, Jackson S, Osterman MT, Anton K,
Ivanova A, et al: Methotrexate is not superior to placebo in
maintaining steroid-free response or remission in ulcerative
colitis. Gastroenterology. 155:1098–1108.e9. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Weissman S, Chris-Olaiya A, Mehta TI, Aziz
M, Alshati A, Berry R, Fatima R, Kolli S, Hassan A and Sciarra MA:
A novel player: Cyclosporine therapy in the management of
inflammatory bowel disease. Transl Gastroenterol Hepatol. 4:672019.
View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Damião AOMC, de Azevedo MFC, Carlos AS,
Wada MY, Silva TVM and Feitosa FC: Conventional therapy for
moderate to severe inflammatory bowel disease: A systematic
literature review. World J Gastroenterol. 25:1142–1157. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Lazzerini M, Martelossi S, Magazzù G,
Pellegrino S, Lucanto MC, Barabino A, Calvi A, Arrigo S, Lionetti
P, Lorusso M, et al: Effect of thalidomide on clinical remission in
children and adolescents with refractory Crohn disease: A
randomized clinical trial. JAMA. 310:2164–2173. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Zhu Z, Li M, Shu X, Bai A, Long S, Liu D,
Lu N, Zhu X and Liao W: Thalidomide is a therapeutic agent that is
effective in inducing and maintaining endoscopic remission in adult
CD patients. Clin Res Hepatol Gastroenterol. 41:210–216. 2017.
View Article : Google Scholar
|
|
131
|
Atreya R, Zimmer M, Bartsch B, Waldner MJ,
Atreya I, Neumann H, Hildner K, Hoffman A, Kiesslich R, Rink AD, et
al: Antibodies against tumor necrosis factor (TNF) induce T-cell
apoptosis in patients with inflammatory bowel diseases via TNF
receptor 2 and intestinal CD14+ macrophages.
Gastroenterology. 141:2026–2038. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Van den Brande JM, Koehler TC, Zelinkova
Z, Bennink RJ, te Velde AA, ten Cate FJ, van Deventer SJ,
Peppelenbosch MP and Hommes DW: Prediction of antitumour necrosis
factor clinical efficacy by real-time visualisation of apoptosis in
patients with Crohn's disease. Gut. 56:509–517. 2007. View Article : Google Scholar
|
|
133
|
Berns M and Hommes DW: Anti-TNF-α
therapies for the treatment of Crohn's disease: The past, present
and future. Expert Opin Investig Drugs. 25:129–143. 2016.
View Article : Google Scholar
|
|
134
|
ten Hove T, van Montfrans C, Peppelenbosch
MP and van Deventer SJ: Infliximab treatment induces apoptosis of
lamina propria T lymphocytes in Crohn's disease. Gut. 50:206–211.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Mitoma H, Horiuchi T, Hatta N, Tsukamoto
H, Harashima S, Kikuchi Y, Otsuka J, Okamura S, Fujita S and Harada
M: Infliximab induces potent anti-inflammatory responses by
outside-to-inside signals through transmembrane TNF-alpha.
Gastroenterology. 128:376–392. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Oikonomopoulos A, van Deen WK and Hommes
DW: Anti-TNF antibodies in inflammatory bowel disease: Do we
finally know how it works? Curr Drug Targets. 14:1421–1432. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Tsai HH: Editorial: tofacitinib and
biologics for moderate-to-severe ulcerative colitis-what is best in
class? Aliment Pharmacol Ther. 47:539–540. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Singh S, Murad MH, Fumery M, Dulai PS and
Sandborn WJ: First- and second-line pharmacotherapies for patients
with moderate to severely active ulcerative colitis: An updated
network meta-analysis. Clin Gastroenterol Hepatol. 18:2179–2191.e6.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
139
|
Panaccione R, Ghosh S, Middleton S,
Márquez JR, Scott BB, Flint L, van Hoogstraten HJ, Chen AC, Zheng
H, Danese S and Rutgeerts P: Combination therapy with infliximab
and azathioprine is superior to monotherapy with either agent in
ulcerative colitis. Gastroenterology. 146:392–400.e3. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
140
|
Wu S, Liu S, Zhang C, Yu B, Tan J and Liu
X: CD74+CCL5+ effector CD8+ T cells drive mucosal inflammation and
predict biologics response in inflammatory bowel disease. J Transl
Med. 24:222025. View Article : Google Scholar
|
|
141
|
Verstockt B, Salas A, Sands BE, Abraham C,
Leibovitzh H, Neurath MF and Vande Casteele N; Alimentiv
Translational Research Consortium (ATRC): IL-12 and IL-23 pathway
inhibition in inflammatory bowel disease. Nat Rev Gastroenterol
Hepatol. 20:433–446. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
142
|
Feagan BG, Sandborn WJ, Gasink C,
Jacobstein D, Lang Y, Friedman JR, Blank MA, Johanns J, Gao LL,
Miao Y, et al: Ustekinumab as Induction and Maintenance Therapy for
Crohn's Disease. N Engl J Med. 375:1946–1960. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Globig AM, Sommer NP, Wild K, Schardey J,
Zoldan K, Thomann AK, Schulte LA, Schreiner R, Reindl W, Klaus J,
et al: Ustekinumab inhibits T follicular helper cell
differentiation in patients with Crohn's disease. Cell Mol
Gastroenterol Hepatol. 11:1–12. 2021. View Article : Google Scholar
|
|
144
|
Danese S, Sands BE, Abreu MT, O'Brien CD,
Bravatà I, Nazar M, Miao Y, Wang Y, Rowbotham D, Leong RWL, et al:
Early symptomatic improvement after ustekinumab therapy in patients
with ulcerative colitis: 16-week data from the UNIFI trial. Clin
Gastroenterol Hepatol. 20:2858–2867.e5. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
145
|
Sands BE, Sandborn WJ, Panaccione R,
O'Brien CD, Zhang H, Johanns J, Adedokun OJ, Li K, Peyrin-Biroulet
L, Van Assche G, et al: Ustekinumab as induction and maintenance
therapy for ulcerative colitis. N Engl J Med. 381:1201–1214. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
146
|
Sandborn WJ, Ferrante M, Bhandari BR,
Berliba E, Hibi T, D'Haens GR, Tuttle JL, Krueger K, Friedrich S,
Durante M, et al: Efficacy and safety of continued treatment with
mirikizumab in a phase 2 trial of patients with ulcerative colitis.
Clin Gastroenterol Hepatol. 20:105–115.e14. 2022. View Article : Google Scholar
|
|
147
|
Sands BE, Chen J, Feagan BG, Penney M,
Rees WA, Danese S, Higgins PDR, Newbold P, Faggioni R, Patra K, et
al: Efficacy and safety of MEDI2070, an antibody against
interleukin 23, in patients with moderate to severe Crohn's
disease: A phase 2a study. Gastroenterology. 153:77–86.e6. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
148
|
Feagan BG, Panés J, Ferrante M, Kaser A,
D'Haens GR, Sandborn WJ, Louis E, Neurath MF, Franchimont D, Dewit
O, et al: Risankizumab in patients with moderate to severe Crohn's
disease: An open-label extension study. Lancet Gastroenterol
Hepatol. 3:671–680. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
149
|
Slack RJ, Macdonald SJF, Roper JA, Jenkins
RG and Hatley RJD: Emerging therapeutic opportunities for integrin
inhibitors. Nat Rev Drug Discov. 21:60–78. 2022. View Article : Google Scholar
|
|
150
|
Traynor K: FDA advisers endorse
natalizumab for Crohn's disease. Am J Health Syst Pharm.
64:18861888–1890. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
151
|
Fischer A, Zundler S, Atreya R, Rath T,
Voskens C, Hirschmann S, López-Posadas R, Watson A, Becker C,
Schuler G, et al: Differential effects of α4β7 and GPR15 on homing
of effector and regulatory T cells from patients with UC to the
inflamed gut in vivo. Gut. 65:1642–1664. 2016. View Article : Google Scholar :
|
|
152
|
Kurmaeva E, Lord JD, Zhang S, Bao JR,
Kevil CG, Grisham MB and Ostanin DV: T cell-associated α4β7 but not
α4β1 integrin is required for the induction and perpetuation of
chronic colitis. Mucosal Immunol. 7:1354–1365. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
153
|
Sandborn WJ, Baert F, Danese S, Krznarić
Ž, Kobayashi T, Yao X, Chen J, Rosario M, Bhatia S, Kisfalvi K, et
al: Efficacy and safety of vedolizumab subcutaneous formulation in
a randomized trial of patients with ulcerative colitis.
Gastroenterology. 158:562–572.e12. 2020. View Article : Google Scholar
|
|
154
|
Feagan BG, Rutgeerts P, Sands BE, Hanauer
S, Colombel JF, Sandborn WJ, Van Assche G, Axler J, Kim HJ, Danese
S, et al: Vedolizumab as induction and maintenance therapy for
ulcerative colitis. N Engl J Med. 369:699–710. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
155
|
Wiendl M, Dedden M, Liu LJ, Schweda A,
Paap EM, Ullrich KA, Hartmann L, Wieser L, Vitali F, Atreya I, et
al: Etrolizumab-s fails to control E-Cadherin-dependent
co-stimulation of highly activated cytotoxic T cells. Nat Commun.
15:10432024. View Article : Google Scholar : PubMed/NCBI
|
|
156
|
Reinisch W, Sandborn WJ, Danese S,
Hébuterne X, Kłopocka M, Tarabar D, Vaňásek T, Greguš M, Hellstern
PA, Kim JS, et al: Long-term safety and efficacy of the
anti-MAdCAM-1 monoclonal antibody ontamalimab [SHP647] for the
treatment of ulcerative colitis: The open-label study TURANDOT II.
J Crohns Colitis. 15:938–949. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
157
|
Li H, Ye XF, Su YS, He W, Zhang JB, Zhang
Q, Zhan LB and Jing XH: Mechanism of acupuncture and moxibustion on
promoting mucosal healing in ulcerative colitis. Chin J Integr Med.
29:847–856. 2023. View Article : Google Scholar
|