Introduction
Tears to the rotator cuff are a common shoulder
joint injury, and postoperative chronic inflammation frequently
results in fibrotic scarring, which significantly elevates the risk
of re-tear and compromises the restoration of shoulder joint
function (1). Due to the unique
gradient structure and limited regenerative capacity of the
tendon-bone interface (TBI), healing at the insertion site
typically occurs via fibrovascular scar tissue formation, even
after successful surgical repair (2). This fibrovascular scar connection
between the tendon and bone exhibits significant mechanical
impairment, resulting in high re-tear rates, particularly in
elderly patients, where the re-tear rate can reach 20-90% (3). Therefore, inflammation plays a key
role in the excessive scar formation in various tissues (4), and reducing excessive inflammation
and scar formation may lead to improved clinical outcomes.
Neutrophils are a critical component of the
inflammatory response. In the early postoperative stage,
inflammatory cells rapidly accumulate at the TBI and attract
monocytes/macrophages to aggregate and proliferate (5). Subsequently, most neutrophils
rapidly undergo apoptosis and begin to accumulate as uncleared
apoptotic cells (ACs). However, if not promptly cleared, ACs can
progress to secondary necrosis, releasing intracellular
self-antigens and cytotoxic compounds, thereby exacerbating the
inflammatory response and leading to excessive scar formation
(6). Apoptosis and the
phagocytic clearance of ACs (efferocytosis) are evolutionarily
conserved processes that occurs in all tissues throughout the life
cycle of an organism, and they are crucial for organism
development, tissue renewal and regeneration, as well as the
development and function of the immune system (7). However, an increasing number of
chronic inflammatory diseases, such as autoimmune diseases and
atherosclerosis, are caused by macrophage efferocytosis dysfunction
(8). Efferocytosis not only
prevents tissue necrosis and inflammation caused by secondary
necrosis of dead cells but also activates anti-inflammatory signals
in macrophages, which is crucial for tissue dissolution and repair
after injury or inflammation (6,9).
Therefore, exploring the molecular mechanisms required for
efferocytosis may uncover novel therapeutic targets for the
treatment or prevention of inflammatory diseases.
E-box binding homeobox-1 (ZEB1), a zinc finger
transcription factor, is well-known for its role in triggering the
epithelial-mesenchymal transition (EMT) program in cancer cells,
which is crucial for embryonic development (neural crest migration)
as well as a range of pathological conditions, such as tumor
metastasis and organ fibrosis (10,11). In recent years, ZEB1 has emerged
as a pivotal regulator of cellular plasticity in non-cancer cells
(12,13). For instance, in macrophages
infiltrating injured tissues, ZEB1 facilitates the phenotypic
transition from pro-inflammatory to anti-inflammatory phenotypes
(14). Additionally, a recent
study demonstrated that a reduction in ZEB1 levels in macrophages
increases atherosclerotic plaque formation, and targeted
upregulation of Zeb1 expression can reverse the high lipid
accumulation in macrophages (15). Meanwhile, Zhang et al
(16) revealed that ZEB1 links
fragmented mitochondrial morphology with the self-renewal of
hematopoietic stem cells (HSC) by inhibiting mitofusin-2
(MFN2)-mediated mitochondrial fusion. Specifically, ZEB1 represses
MFN2 to induce mitochondrial fragmentation, which leads to reduced
mitochondrial mass and lower reactive oxygen species (ROS) levels,
which is essential to prevent HSC exhaustion and maintain their
long-term self-renewal capacity.
The significant role of mitochondrial morphology
changes in mitochondrial homeostasis has been highlighted in
previous studies. Mitochondrial morphology alterations, also known
as mitochondrial dynamics, mainly include mitochondrial fusion and
fission. Wang et al (17)
reported that mitochondrial fission plays a crucial role in
promoting the continuous clearance of ACs by macrophages. Silencing
the fusion mediator Mfn1 leads to mitochondrial hyper-fragmentation
and enhances efferocytosis at an ACs:macrophage ratio of 10:1,
mainly due to the increase in cytoplasmic calcium concentration
induced by ACs. Moreover, Hu et al (18) also indicated that a reduction in
MFN2 leads to an increase in mitochondrial fission. The dynamic
changes in mitochondrial fission and fusion are mainly regulated by
fusion proteins such as MFN1 and MFN2, and fission proteins,
including dynamin-related protein 1 (DRP1) and mitochondrial
fission factor, which are essential for cellular functions under
different conditions (8).
However, the precise mechanism by which ZEB1 coordinates
mitochondrial dynamics to regulate efferocytosis in macrophages
remains to be elucidated. Therefore, the present study hypothesizes
that ZEB1 plays a significant role in the efferocytosis process of
macrophages by regulating MFN2-mediated mitochondrial dynamics,
which may be closely related to the improvement of the inflammatory
microenvironment at the TBI.
Therefore, exploring the potential role of ZEB1 in
macrophages within the chronic inflammatory microenvironment of
rotator cuff injuries is of notable importance. The present study
aimed to elucidate the regulatory mechanism of ZEB1 on
mitochondrial dynamics and macrophage efferocytosis, specifically
focusing on the ZEB1-MFN2 axis. To achieve this, a rat rotator cuff
injury repair model and in vitro macrophage systems were
used to investigate how ZEB1-mediated mitochondrial remodeling
influences the inflammatory response and the quality of tendon-bone
healing.
Materials and methods
Animal models and gene knockdown
Bone marrow-derived macrophages (BMDMs) were
extracted and characterized from 10, 6-week-old male C57BL/6 mice
(weighing 20-22 g). Following a one-week adaptation period, the
mice were euthanized by cervical dislocation.
A total of 30 12-week-old male Sprague-Dawley (SD)
rats (weighing 300-350 g) were used to establish the rotator cuff
injury model for in vivo experiments. All animals were
purchased from the Experimental Animal Center of Xi'an Jiaotong
University and housed in a specific pathogen-free environment
(temperature, 20-25°C; humidity, 50-60%; 12-h light/dark cycle),
with ad libitum access to food and water, with five rats per
cage. After a 7 day adaptation period, the rats were randomly
divided into two groups using a random number table method.
Group 1: Rotator cuff injury negative control group
(NC group; n=15); a rotator cuff injury repair model was
established as described below and rats were injected with a
non-targeting virus [recombinant lentivirus (rLV)-scramble short
hairpin (sh)RNA]. SD rats were anesthetized via intraperitoneal
injection of 0.3% pentobarbital sodium (40 mg/kg), and then shaved
and disinfected. A 1-cm longitudinal incision was made in the
surgical area (anterolateral aspect of the shoulder), and the
subcutaneous tissue was bluntly separated. The connection between
the deltoid muscle and the acromion was transected, and the
supraspinatus tendon was lifted with a retractor. A 6-0 prolene
suture (Ethicon, Inc.) was placed at the end of the supraspinatus
tendon, and the residual fibrocartilage at the implantation site
was removed. A total of two 0.5 mm bone tunnels were drilled at the
joint margin of the implantation site. The suture was passed
through the bone tunnels, tightened and knotted. The in situ
fixation of the supraspinatus tendon was confirmed by exploration.
The deltoid muscle and skin were sutured layer by layer.
Group 2: Rotator cuff injury control group +
Zeb1 knockdown group (shZEB1 group; n=15); in addition to
the aforementioned treatment, Zeb1 gene knockdown
experiments were conducted. rLV-shRNA1-mZeb1 suspension
(1×107 TU/ml; 100 µl; Ribo-Bio) was injected locally
into the affected shoulder joint immediately after surgery and at
the 1st, 2nd and 3rd weeks after surgery (rats scheduled for 1-week
euthanasia received injections only at the first two time points),
while the NC group was injected with the same amount of
rLV-scramble shRNA at the same time points. The knockdown
efficiency was verified by reverse transcription-quantitative
(RT-q)PCR at 48 h post-surgery (using dedicated validation samples;
n=3 per group) and immunohistochemistry (IHC) at 1 week
post-surgery. At 1, 4 and 8 weeks after surgery, the rats were
euthanized by intraperitoneal injection of an overdose of
pentobarbital sodium (150 mg/kg). Subsequently, the rotator
cuff-humeral complex specimens (the tendon-bone interface
specimens, comprising the supraspinatus tendon and the greater
tuberosity of the humerus) were collected for analysis. This animal
experiment protocol was approved by the Animal Experiment Ethics
Committee of Xi'an Jiaotong University (approval no.
XJTUAE2025-2322).
Cell culture
BMDMs were generated as described in reference
through the differentiation of bone marrow cells (18). Male C57BL/6 mice (6-weeks old)
were euthanized by cervical dislocation and used as the source of
bone marrow. The humeri, femurs and tibiae were aseptically
harvested and surface-sterilized by brief immersion in 75% ethanol
for 30 sec at 20-25°C. Bone marrow was flushed from the isolated
bones using ice-cold phosphate-buffered saline (PBS), and the
resulting cell suspension was passed through a 70 µm cell strainer
to remove debris and obtain a single-cell suspension. The collected
cells (1-10×105 /cm2) were seeded in a medium
(DMEM; Gibco; Thermo Fisher Scientific, Inc.) containing 15% fetal
bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc.) and 20
ng/ml recombinant macrophage colony-stimulating factor (M-CSF;
PeproTech, Inc.). After 3 days of culture (37°C with 5%
CO2), the medium was replaced with fresh medium. After a
total of 6 days, the cells were firmly attached to the surface of
the culture plate. The expression of F4/80, a well-established and
specific cell-surface marker for mature murine macrophages, was
verified by immunofluorescence (IF) to verify the successful
differentiation and purity of the BMDMs. Briefly, the cultured
cells were fixed with 4% paraformaldehyde (PFA) for 15 min at room
temperature. Following PBS washes, the cells were blocked with 5%
normal goat serum (Sigma-Aldrich; Merck KGaA) for 1 h at room
temperature. The cells were then incubated overnight at 4°C with a
primary antibody against F4/80 (1:500; cat. no. ab300421; Abcam).
Subsequently, the cells were incubated with an Alexa Fluor
488-conjugated goat anti-rabbit IgG secondary antibody (1:1,000;
cat. no. ab150081; Abcam) for 1 h at room temperature in the dark.
Nuclei were counterstained with DAPI for 5 min at room temperature.
Finally, the cells were observed and imaged using a fluorescence
microscope (Olympus BX53) at ×200 and ×400 magnifications. Image
analysis was performed using ImageJ software (version 1.53k;
National Institutes of Health).
The in vitro experiment divided BMDMs into
two groups: i) Control group (NC group) where cells were treated
with an equal amount of rLV-scramble shRNA as the negative control
group; and ii) Zeb1-knockdown group (sh-ZEB1 group) where 50
µl of lentivirus (1×107 TU/ml,MOI=20,
rLV-shRNA1-mZeb1) was added to each well for Zeb1
gene knockdown, along with polybrene (8 µg/ml) to assist infection
(19). Third-generation
lentiviral particles were produced in 293T cells using a
four-plasmid packaging system. The shRNA constructs were cloned
into the pLKO.1-U6-Puro lentiviral vector system (Ribo-Bio). For a
10-cm dish, a total of 20 µg of plasmid DNA was transfected,
including the transfer plasmid, packaging plasmids (pMDLg/pRRE and
pRSV-Rev), and the envelope plasmid (pMD2.G) at a mass ratio of
4:2:1:1. Transfection was performed at 37°C for 6-8 h using
Lipofectamine 3000, followed by replacement with fresh complete
medium. Lentiviral particles were harvested from the cell
supernatant at 48 and 72 h post-transfection, filtered through a
0.45 µm filter, and concentrated. The core target sequences
were as follows: Sh-Zeb1 target sequence, 5′-ATA GAG GCT ACA
AGC GCT TTA-3′; scramble shRNA target sequence, 5′-CAA CAA GAT GAA
GAG CAC CAA-3′. After 12 h of infection, the medium was replaced
with fresh medium and the cells were further cultured for 48 h. To
establish stable cell lines, transduced cells were selected with 2
µg/ml puromycin for 7 days and subsequently maintained in
0.5 µg/ml puromycin. After verifying the Zeb1
knockdown efficiency by RT-qPCR and western blotting (as described
below), the cells were seeded at a density of 5×105
cells/well in six-well plates for subsequent experiments.
The human lymphocyte line Jurkat (clone E6-1;
American Type Culture Collection) was selected as the apoptotic
cell model. Logarithmic growth phase Jurkat cells (1×106
cells/ml) were seeded in six-well plates and exposed to ultraviolet
(UV) light under uncovered conditions. A 254 nm UV light was used
with an energy dose of 200 mJ/cm2 for 15-20 min. Fresh
RPMI-1640 medium (Gibco; Thermo Fisher Scientific, Inc.)
[supplemented with 10% fetal bovine serum (FBS; Gibco; Thermo
Fisher Scientific, Inc.)] was then added and the cells were further
cultured for 6 h, with an apoptosis rate of ~85% (17). Then, apoptotic Jurkat cells were
labeled with propidium iodide (PI, 1 µg/ml, incubated for 15
min, at 20-25°C, protected from light), and set aside at 37°C for
30 min for later use.
Efferocytosis assay
In vitro experiments were conducted using the
phagocytosis of ACs assay, following the previously reported method
(16). To investigate the
dynamic expression of Zeb1 during efferocytosis, BMDMs were
co-cultured with ACs at a ratio of 5:1 (ACs:BMDMs) for varying
durations (0, 1, 3 and 6 h). At each time point, total RNA was
extracted, and Zeb1 mRNA levels were determined by RT-qPCR and
western blotting. Briefly, the aforementioned UV pre-treated
apoptotic Jurkat cells were co-cultured with BMDMs that had been
labeled with Mito-Tracker Green (cat. no. C1996s; Beyotime
Biotechnology) for mitochondrial staining, and the mixture was
incubated for the designated time period (37°C with 5%
CO2 for 6 h). Non-ingested Jurkat cells were
subsequently removed by washing three times with PBS. Phagocytic
activity was visualized via fluorescence microscopy, and the
efferocytic index was quantified as the percentage of PI-positive
macrophages as follows: Efferocytic Index (%)=(number of
PI-positive macrophages/total number of macrophages) ×100% (ImageJ
software; version 1.53k; National Institutes of Health).
RNA isolation and RT-qPCR
Total RNA was extracted from BMDM cells using
TRIzol® reagent (Thermo Fisher Scientific, Inc.), and
reverse transcribed using a AffinityScript QPCR cDNA Synthesis Kit
(Agilent Technologies, Inc.) according to the manufacturers
instructions. RNA samples were analyzed by qPCR using SYBR Green
PCR premix (Applied Biosystems; Thermo Fisher Scientific, Inc.),
with GAPDH as the internal reference gene. The sequences of the
primers used are as follows: GAPDH forward, 5′-CTG CAC CAC CAA CTG
CTT AG-3′, and reverse, 5′-GTC TGG GAT GGA AAT TGT GA-3′; inducible
nitric oxide synthase (iNOS) forward, 5′-CAA GCA CCT TGG AAG AGG
AG-3′, and reverse, 5′-AAG GCC AAA CAC AGC ATA CC-3′; TNF-α
forward, 5′-CTG TAG CCC ACG TCG TAG C-3′, and reverse, 5′-TTG AGA
TCC ATG CCG TTG-3′; arginase 1 (Arg-1) forward, 5′-CTC CAA GCC AAA
GTC CTT AGAG-3′, and reverse, 5′-AGG AGC TGT CAT TAG GGA CATC-3′;
CD206 forward, 5′-CAA GGA AGG TTG GCA TTT GT-3′, and reverse,
5′-CCT TTC AGT CCT TTG CAA GC-3′; ZEB1 forward, 5′-GCT GGC AAG ACA
ACG TGA AAG-3′, and reverse, 5′-GCC TCA GGA TAA ATG ACG GC-3′. The
quantification of gene expression was carried out using the
2-ΔΔCq method relative to the internal control (20).
Western blotting
Protein extraction was performed using a standard
protocol, and protein quantification was carried out with a BCA
Protein Assay Kit (Beyotime Biotechnology). Mitochondrial proteins
were extracted using the Cell Mitochondria Isolation Kit (cat. no.
C3601; Beyotime Biotechnology). The procedure was performed
strictly according to the manufacturer's instructions, involving
cell homogenization followed by differential centrifugation to
separate the mitochondrial fraction from the cytosolic fraction.
The proteins were diluted to the same concentration with loading
buffer (Beyotime Biotechnology) and then loaded onto 10% SDS-PAGE
gels. After electrophoresis, the proteins (30 µg per lane)
were transferred to PVDF membranes (MilliporeSigma) by wet
transfer. The PVDF membranes were blocked with 5% skim milk in TBST
(0.05% Tween) for 2 h at room temperature and incubated with
primary antibodies overnight at 4°C. After washing, the membranes
were incubated with secondary antibodies (HRP-conjugated goat
anti-rabbit IgG) at room temperature for 2 h (1:10,000; cat. no.
SA00001-2; Proteintech Group, Inc.). Protein detection was
performed using an ultra-sensitive enhanced chemiluminescence kit
(Shanghai Yamei Biomedical Technology Co Ltd.) and using ImageJ
software (version 1.53k; National Institutes of Health). The
primary antibodies used were as follows: Anti-β-Actin antibody
(1:5,000; cat. no. 81115-1-RR; Proteintech Group, Inc.), anti-ZEB1
antibody (1:1,000; cat. no. ab203829; Abcam), anti-MFN2 antibody
(1:5,000; cat. no. ab124773; Abcam), and anti-DRP1 antibody
(1:5,000; cat. no. 5391; Cell Signaling Technology, Inc.).
Flow cytometry
For purity identification, BMDMs were harvested and
resuspended (5-10×106 cells/ml) in Immunol Staining
Buffer (Beyotime Biotechnology) to prevent non-specific binding and
maintain cell viability. Cells were incubated with an
APC-conjugated anti-mouse F4/80 antibody (cat. no. AC0988; Beyotime
Biotechnology) for 30 min at 4°C in the dark. The stained cells
were resuspended in PBS and analyzed immediately to prevent signal
loss and ensure accurate detection of surface markers on live
cells. Unstained cells served as a negative control to determine
the gating strategy. The purity of BMDMs was defined as the
percentage of F4/80-positive cells (number of F4/80-positive
macrophages/total number of macrophages) ×100%). The flow cytometry
analysis was performed using a Sino-Cyte Flow Cytometer (Beijing
Sino-Cyte Scientific Co., Ltd.). Data acquisition and analysis were
performed using FlowJo software (version 10; Tree Star, Inc.).
For apoptosis analysis, Jurkat cell death (early +
late apoptosis) was evaluated using an Annexin V-FITC/PI Apoptosis
Detection Kit (cat. no. C1062S; Beyotime Biotechnology). Briefly,
after UV treatment (detailed above), Jurkat cells were collected,
washed with cold PBS, and resuspended in 1× Binding Buffer
(Beyotime Biotechnology) at a density of 5-10×106
cells/ml. Annexin V-FITC and PI were added to the cell suspension
and incubated for 15 min at room temperature in the dark.
Histological analysis
The TBI specimens collected at 4 and 8 weeks
post-surgery were processed for histological analysis. Serial
sections were obtained from each specimen and stained with
hematoxylin and eosin (H&E), safranin O-fast green (SO/FG) and
Sirius Red to evaluate structural healing, fibrocartilage formation
and collagen organization, respectively. Specimens were fixed in
10% neutral formalin solution for 48 h at room temperature
(20-25°C). Subsequently, decalcification was performed in 10% EDTA
buffer at 37°C water bath, with the EDTA solution being replaced
every 3 days for ~4 weeks until the samples became soft. After
decalcification, the samples were embedded in paraffin and cut into
5-µm thick sections using an ultrathin sectioning machine
along the coronal plane parallel to the shoulder joint. The
interface region between the tendon graft and bone tissue was
observed by H&E staining (sections were stained with
hematoxylin for 5 min and eosin for 2 min at room temperature).
SO/FG staining was performed according to the Safranin O-Fast Green
Stain Kit (cat. no. C0621S; Beyotime Biotechnology) instructions to
evaluate the formation and integration of fibrocartilage and bone
tissue. For Sirius Red staining, the sections were incubated with
Sirius Red solution at room temperature (20-25°C) for 1 h (cat. no.
C0190S; Beyotime Biotechnology), followed by rapid dehydration in
absolute ethanol and clearing in xylene to assess collagen
organization. The histological scoring system applied in this study
was adopted the TBI healing assessment method described by Ide
et al (21). The TBI
scoring system assigned a maximum score of 16 points for normal
tissue morphology, with higher scores indicating better healing
outcomes. To ensure objectivity and reduce subjective bias, the
histological grading was conducted by two independent observers who
were blinded to the experimental groups. The inter-rater
reliability was assessed using Cohen's κ coefficient (with κ>0.8
considered as excellent agreement). The H&E and SO/FG stained
sections were scanned using a Pannoramic Digital Slide Scanner
(3DHistech, Ltd.), and digital images were analyzed using
CaseViewer 2.4 software (3DHistech, Ltd.). Subsequently, Sirius
Red-stained sections were examined under a polarizing light
microscope (Eclipse E800; Nikon Corporation) to assess collagen
fiber composition. Strong yellow-to-red birefringence was
indicative of type I collagen fibers, whereas weak green
birefringence represented the presence of type III collagen.
Digital images were captured under polarized light, and ImageJ
software (version 1.53k; National Institutes of Health) was
utilized to quantify the mean intensity (calculated as the
arithmetic mean of pixel gray values) of type I collagen across
experimental groups for comparative analysis.
Immunofluorescence, immunohistochemical
(IHC) and TUNEL staining
The preparation process of immunofluorescence (IF)
staining sections is consistent with that of H&E staining. The
sections were deparaffinized in xylene and rehydrated through a
descending ethanol series (100, 95, 80 and 75%). Antigen retrieval
was performed by heating the sections in citrate buffer (pH 6.0) at
95-100°C for 15 min. After cooling to room temperature, the
sections were permeabilized with 0.1% Triton X-100 for 15 min to
facilitate antibody penetration for intracellular antigens.
Subsequently, the sections were treated with normal goat serum
(cat. no. C0265; Beyotime Biotechnology) at room temperature for 30
min to reduce non-specific staining. Then, the sections were
incubated overnight at 4°C with primary antibodies against iNOS
(1:50; cat. no. 18985-1-AP), CD206 (1:50; cat. no. 18704-1-AP) and
CD68 (1;50; cat. no. 28058-1-AP) (Proteintech Group, Inc.)
according to the manufacturer's instructions. The sections were
incubated with the corresponding fluorescent secondary antibodies
for 2 h at room temperature in the dark. Specifically,
FITC-conjugated goat anti-rabbit IgG (1:200; cat. no. GB22404;
Wuhan Servicebio Technology Co., Ltd.) was used for iNOS detection,
while Cy3-conjugated goat anti-rabbit IgG (1:300; cat. no. GB21303;
Wuhan Servicebio Technology Co., Ltd.) was used for CD206 and CD68.
Finally, the sections or smears were treated with DAPI staining
agent at room temperature for 5 min to visualize nuclei. Digital
images were acquired using a Pannoramic Digital Slide Scanner
(3DHistech, Ltd.) and analyzed using CaseViewer 2.4 software
(3DHistech, Ltd.).
For IHC, the procedures for slice preparation,
dewaxing and antigen retrieval were performed in accordance with
the protocols used for immunofluorescence (IF). The tissue sections
were treated with 3% hydrogen peroxide (H2O2)
solution at room temperature for 15 min to suppress endogenous
peroxidase activity. Following this, the sections were incubated
with 5% normal goat serum for 30 min at room temperature (20-25°C)
to block non-specific binding sites. Subsequently, the sections
were incubated overnight at 4°C with appropriately diluted primary
antibodies against SOX9 (1:50; cat. no. GB14171), RUNx2 (1:50;
GB125631) and α-SMA (1:500; cat. no. GB111364). After thorough
washing with PBS, the sections were incubated with the
corresponding HRP-conjugated goat anti-rabbit IgG secondary
antibodies (1:300; cat. no. GB23303; Wuhan Servicebio Technology
Co., Ltd.) at 37°C for 30 min. Thereafter, the sections were
stained with diaminobenzidine (DAB) at room temperature (20-25°C)
for 2-5 min, with the degree of color development strictly
monitored under a microscope. After washing with distilled water,
the sections were counterstained with hematoxylin at room
temperature for 3 min to visualize nuclear morphology.
Subsequently, the sections were differentiated in 1% acid alcohol,
blued in running tap water, dehydrated through an ascending graded
ethanol series, cleared in xylene, and mounted with a neutral
resinous medium. Finally, the dried sections were examined under a
light microscope, and representative images were captured.
Apoptosis detection was carried out using the
One-Step TUNEL Apoptosis Assay Kit (cat. no. C1086; Beyotime
Biotechnology), following the manufacturer's instructions. Briefly,
samples were fixed in 4% paraformaldehyde at 4°C for 24 h and
permeabilized. After incubation with the TUNEL reaction mixture,
nuclei were counterstained with DAPI (1 µg/ml) at room
temperature for 5 min. Sections were mounted with Antifade Mounting
Medium (Beyotime Biotechnology). The sections were then scanned
using a Pannoramic Digital Slide Scanner (3DHistech, Ltd.). Digital
images from 5 random fields of view per sample were captured and
analyzed using CaseViewer 2.4 software. Each section was incubated
with 50 µl of TUNEL detection reagent at 37°C in the dark
for 60 min. Following staining, the proportion of positively
stained areas for each marker was quantitatively analyzed using
Image analysis software.
Micro-CT analysis
The rotator cuff-humeral head complexes were
harvested at 4 and 8 weeks postoperatively. After being fixed in 4%
paraformaldehyde for 24 h at 4°C, they were scanned using a
micro-computed tomography (micro-CT, NMC-200; Pingseng Medical
Technology) with the following parameters: 80 kV, 270 µA and
an effective pixel size of 18 µm. After scanning,
three-dimensional images were reconstructed. A region of interest
(ROI) with a diameter of 1.6 mm was selected in the greater
tuberosity area of the humerus. Bone mineral density (BMD),
trabecular bone volume fraction (BV/TV), trabecular thickness
(Tb.Th), and trabecular number (Tb.N) of the selected ROI were
analyzed.
Biomechanical test
At 4 and 8 weeks post-operation, 3 rats were
randomly selected from each group and sacrificed. The rotator
cuff-humerus complex specimens were retrieved, with only the intact
connection structure between the supraspinatus tendon and the
humerus retained. The cross-sectional area of the middle part of
the supraspinatus tendon was measured using a digital caliper
(Tajima Industries, Ltd.). The proximal end of the humerus was
fixed with polymethyl methacrylate at room temperature (20-25°C)
for ~15 min, and the supraspinatus tendon was covered with
polyester cloth and fixed by a lockstitch suture. The biomechanical
testing was performed using a Shimadzu AG-IS Universal Testing
Machine (Shimadzu Corporation). After a 0.1 Newtons (N)
pretreatment (20-25°C), the specimens were subjected to uniaxial
tensile loading at a constant speed of 10 mm/min until the tendon
failed at the bone repair site. The ultimate failure load was
determined as the peak force recorded during the tensile test.
Stiffness (N/mm) was calculated as the slope of the linear portion
of the load-deformation curve.
Grip strength test and animal gait
analysis
Grip strength tests were conducted at 4 and 8 weeks
postoperatively in each group of rats to assess the biomechanical
recovery following rotator cuff repair. During the test, the rat's
tail was gently held and its forelimbs were placed on the metal
grid of the grip strength meter. As the rat instinctively grasped
the grid, the tester maintained the animal's body parallel to the
grid and gradually pulled the tail until the forelimbs released.
The peak force exerted at the moment of release was recorded by the
device. Each rat performed the test in triplicate, and the mean
value was calculated for subsequent statistical analysis.
At 4 and 8 weeks postoperatively, gait analysis was
conducted on each group of rats using the Greenwalk system
(22) to assess the gait
condition after rotator cuff repair. The experimental setup
consisted of a transparent glass runway and a high-resolution
camera below. The rats were placed at one end of the runway in a
quiet environment and walked freely through the entire channel. The
camera simultaneously captured the footprints and pressure
distribution. Each rat completed ≥3 effective walks, and the
average value was used for analysis. The analysis parameters
included: i) Maximum contact area-the largest area of contact
between the forepaws and the glass plate; ii) stride length-the
distance between consecutive positions of the forepaws during
movement; iii) running speed-the length of the glass plate divided
by the time taken to pass through; and iv) mean intensity-the
average pressure exerted by the forelimbs on the glass plate. A
larger maximum contact area, longer stride length, faster speed and
greater pressure indicated a greater healing effect.
Statistical analysis
All data were analyzed using IBM SPSS Statistics
24.0 software (IBM Corp.) and GraphPad Prism 8.0 software
(GraphPad; Dotmatics). Experimental data were expressed as (X̄ ± s)
or percentages. All in vitro experiments were performed with
at least three independent biological replicates. Normality tests
(Shapiro-Wilk test) and homogeneity of variance tests (Levene's
test) were first conducted for each group of data. If the data were
normally distributed and had homogeneity of variance, independent
sample Students t-tests were used for inter-group comparisons; For
data that did not meet the assumptions of normality, the
Mann-Whitney U test was utilized. P<0.05 was considered to
indicate a statistically significant difference.
Results
Identification of BMDMs
After 6 days of M-CSF-induced differentiation, the
identity and purity of the obtained BMDMs were verified.
Immunofluorescence staining showed robust expression of the
macrophage-specific marker F4/80 (green) (Fig. 1A). Consistently, flow cytometry
analysis further confirmed that the proportion of F4/80-positive
cells reached a high level of purity (>85%), ensuring that the
cells were suitable for subsequent functional assays (Fig. 1B).
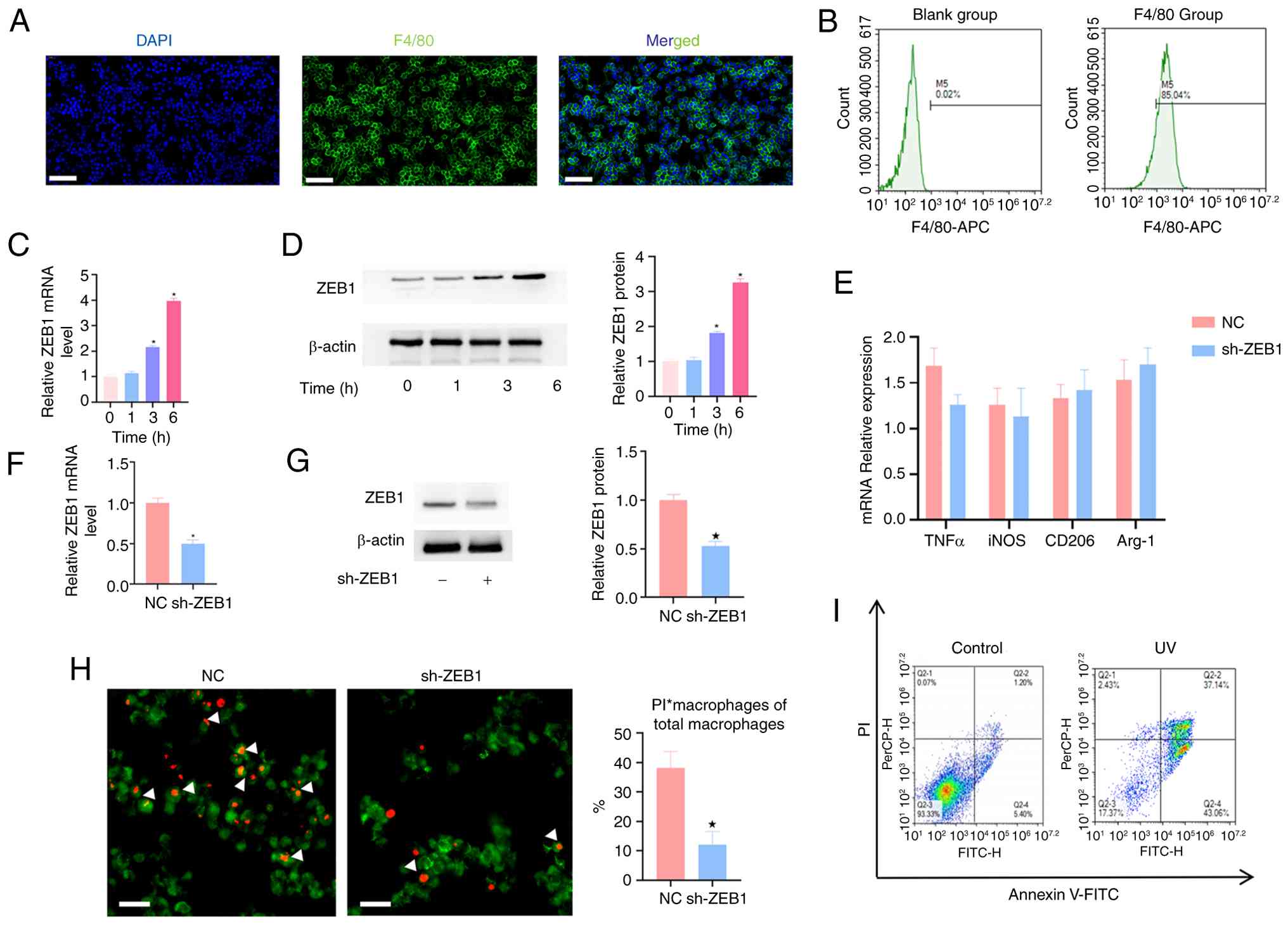 | Figure 1ZEB1 knockdown affects the cell
efferocytosis efficiency of macrophages. (A) After 6 days of
induction culture with M-CSF, the expression level of F4/80 (green)
was detected by immunofluorescence staining (scale bar, 20
µm). (B) Flow cytometry was performed to detect the
proportion of F4/80-positive cells to verify the purity of
macrophages. (C) The mRNA expression levels of ZEB1 in macrophages
at different time points (0, 1, 3 and 6 h) of co-culture were
determined by RT-qPCR. (D) After co-incubation of BMDMs with ACs
for 0, 1, 3 and 6 h, the protein expression of ZEB1 was detected by
western blotting and semi-quantitative analysis of ZEB1 expression
was conducted. (E) The mRNA expressions of TNFα, iNOS, CD206 and
Arg-1 were detected by RT-qPCR. (F) The expression of ZEB1 mRNA was
detected by RT-qPCR to evaluate the knockout efficiency. (G)
Detection of ZEB1 protein expression by WB and semi-quantitative
analysis of ZEB1 expression. (H) BMDMs were labeled with
Mito-Tracker (green label), and then co-cultured with PI-labeled
apoptotic Jurkat cells at a ratio of 5:1 of apoptotic cells to
macrophages for 6 h. After removing the non-phagocytic apoptotic
cells, the proportion of PI-positive macrophages (indicated by
arrows) among the total macrophages was determined by fluorescence
microscopy (Scale bar, 20 µm). (I) The apoptosis rate of
cells induced by UV radiation was detected by flow cytometry
(Annexin V-FITC/PI double staining) to prepare target cells for the
efferocytosis assay. n=3. *P<0.05 vs. NC or 0 h
groups. NC, negative control; sh, short hairpin; ZEB1, zinc finger
E-box binding homeobox 1; BMDMs, bone marrow-derived macrophages;
ACs, apoptotic cells; M-CSF, macrophage colony-stimulating factor;
RT-qPCR, reverse transcription-quantitative PCR; iNOS, inducible
nitric oxide synthase. |
ZEB1 is involved in the regulation of
macrophage efferocytosis
To explore the potential role of ZEB1 in
efferocytosis, BMDMs were co-cultured with ACs at a ratio of 5:1
for varying durations (0, 1, 3, and 6 h). RT-qPCR results
demonstrated that Zeb1 mRNA levels increased in a time-dependent
manner during the clearance of ACs (Fig. 1C). This trend was further
confirmed at the protein level using western blotting; as shown in
Fig. 1D, ZEB1 protein expression
significantly increased following co-culture. Specifically, while
no significant change was observed at 1 h, ZEB1 levels increased
~1.5-fold at 3 h and 3-fold at 6 h compared with the baseline.
These data suggested that ZEB1 was involved in the biological
response of macrophages during efferocytosis.
ZEB1-knockdown impairs macrophage
efferocytosis
To investigate whether ZEB1 regulates macrophage
polarization-a process closely linked to mitochondrial metabolism
(23)-the present study first
evaluated the effect of Zeb1-knockdown on macrophage markers
under baseline conditions. BMDMs were transfected with shRNA
targeting Zeb1, and the knockdown efficiency was validated.
Compared with the NC group, Zeb1 expression was reduced by
~53% at the mRNA level (Fig. 1F)
and 51% at the protein level (Fig.
1G). Notably, RT-qPCR analysis showed no significant
differences in the mRNA levels of M1 (TNFα, iNOS) and M2 (CD206,
Arg-1) markers between the NC and sh-ZEB1 groups under baseline
conditions (Fig. 1E), indicating
that Zeb1-knockdown does not spontaneously alter macrophage
polarization.
Next, the impact of ZEB1 on the phagocytic capacity
of macrophages was assessed. Target apoptotic Jurkat cells were
prepared via UV radiation; Fig.
1I shows a representative flow cytometry plot confirming that
the majority of Jurkat cells were in the early/late apoptotic stage
(Annexin V-positive) prior to co-culture. BMDMs (labeled with
Mito-Tracker Green) were then co-incubated with PI-labeled ACs
(red) for 6 h. Fluorescence microscopy revealed that in the NC
group, numerous macrophages successfully engulfed or bound to ACs
(indicated by arrows showing overlapping signals). By contrast, the
sh-ZEB1 group exhibited a significant reduction in phagocytic
activity; the percentage of PI-positive macrophages was
significantly lower in the Zeb1 knockdown group compared to
the NC group (Fig. 1H).
Collectively, these results demonstrate that ZEB1 knockdown
significantly impaired the efferocytosis efficiency of
macrophages.
ZEB1 knockdown leads to the reduction of
mitochondrial fission
Since mitochondrial fission promotes the continuous
clearance of ACs by macrophages, and it has been reported that its
inhibition can reduce macrophage efferocytosis (16), the present study investigated
whether ZEB1 knockdown affects macrophage efferocytosis by
inhibiting mitochondrial fission. To explore the effect of
Zeb1-knockdown on mitochondrial morphology, mitochondria
were stained with Mito Tracker Deep Red, images were collected with
a confocal microscope and the mitochondrial morphology was
analyzed. Mitochondria in the NC group exhibited a more punctuated
and fragmented network with fewer branches compared with the
sh-ZEB1 group (Fig. 2A),
indicating that Zeb1-knockdown inhibited the fission of
mitochondria.
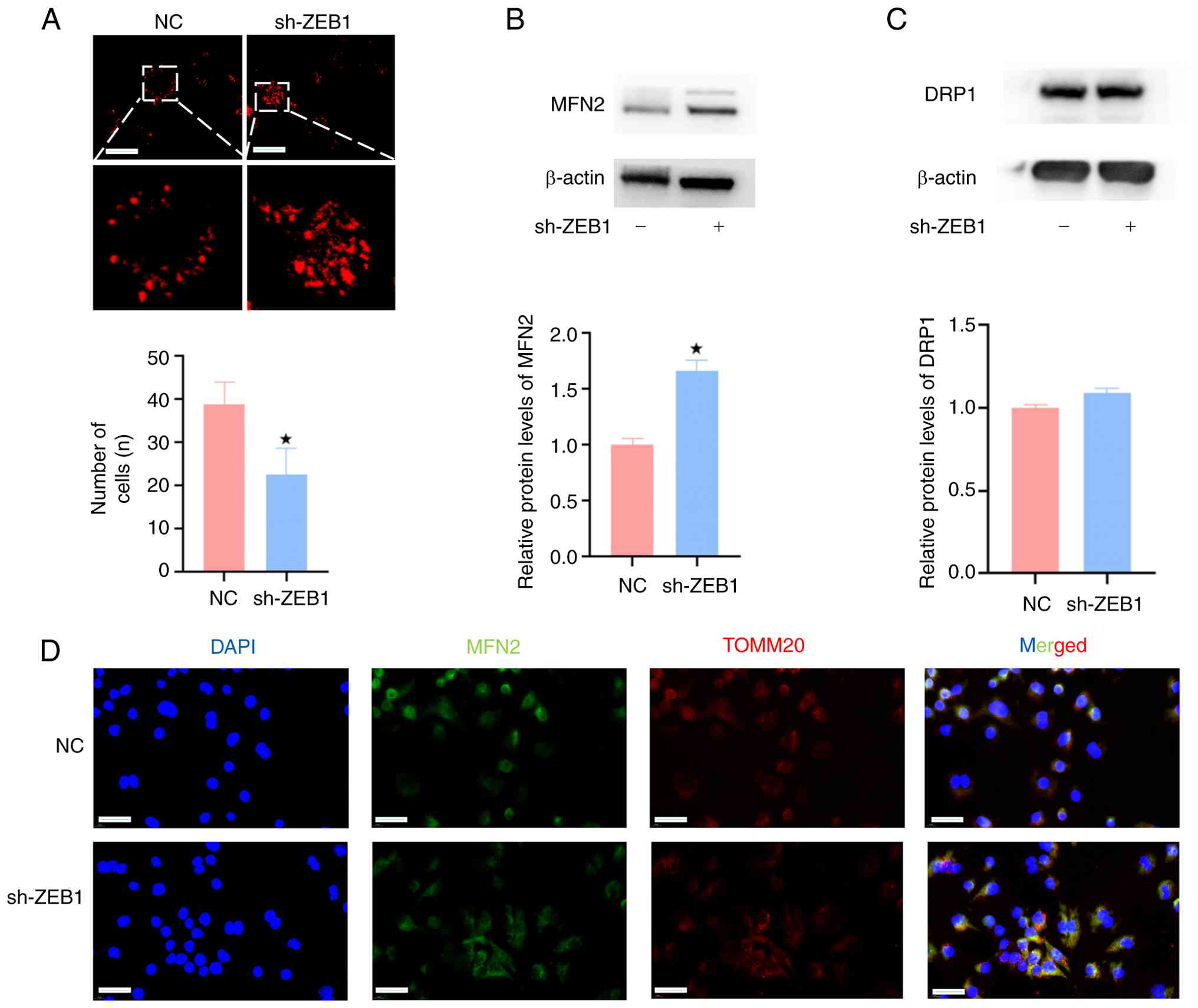 | Figure 2ZEB1 affects mitochondrial fission by
regulating MFN2. (A) Confocal microscopy results of mitochondrial
morphology in macrophages. Mitochondria were labeled with
Mito-Tracker Red fluorescent probe (scale bar, 10 µm; lower
panels, ×200 magnification). Western blotting analysis of (B) MFN2
and (C) DRP1 protein expression in macrophages after ZEB1
knockdown. Quantitative density analysis data are presented as mean
± standard deviation (n=3 independent experiments). (D)
Immunofluorescence colocalization analysis of MFN2 localization in
mitochondria. Nuclei were stained with DAPI (blue), MFN2 protein
was labeled with Alexa Fluor 488 (green) and mitochondrial membrane
protein TOMM20 was labeled with Alexa Fluor 594 (red) (scale bar,
10 µm). n=3, *P<0.05 vs. NC. DRP1,
dynamin-related protein 1; MFN2, Mitofusin-2; ZEB1, zinc finger
E-box binding homeobox 1; NC, negative control; sh, short
hairpin. |
ZEB1-knockdown leads to an increase in
the levels of MFN2 in mitochondria
Mitochondrial morphology is closely related to the
process of fusion and fission, mainly regulated by fusion and
fission proteins. A previous study has shown that ZEB1 is a
transcriptional repressor of MFN2 (15). However, it is also known that
DRP1 plays a dominant role in mitochondrial fission (24). Therefore, the present study
extracted mitochondrial proteins from Zeb1-knockdown primary
macrophages using a cell mitochondrial isolation kit and mainly
detected the protein levels of MFN2 and DRP1 by western blotting.
The results showed that after Zeb1-knockdown, compared with
the NC group, the expression level of DRP1 protein did not change
significantly, but the expression level of MFN2 protein increased
significantly (Fig. 2B and C).
Therefore, the expression levels of MFN2 in mitochondria were
further detected by immunofluorescence colocalization technology.
The results showed that MFN2 exhibited distinct colocalization with
the mitochondrial marker Tomm20 in the sh-ZEB1 group (Fig. 2D), confirming its mitochondrial
localization. These results indicated that ZEB1 affected macrophage
efferocytosis by regulating the expression of MFN2 on mitochondria.
In summary, ZEB1 regulated mitochondrial dynamics by inhibiting
MFN2 expression, thereby regulating macrophage efferocytosis, which
is characterized by the inhibition of mitochondrial fusion.
ZEB1-knockdown exacerbates the early
inflammatory response at the TBI
To evaluate the effect of ZEB1 on the
microenvironment of the TBI post-surgery, the knockdown efficiency
in vivo was first verified. At 48 h post-operation, RT-qPCR
analysis of the tendon tissues confirmed a significant reduction in
Zeb1 mRNA levels in the sh-ZEB1 group compared with the NC
group (Fig. 3A). Consistently,
IHC staining performed 1 week post-surgery demonstrated that ZEB1
protein expression was significantly decreased at the interface
(Fig. 3B). Furthermore, ZEB1
deficiency led to a upregulation of MFN2 at both mRNA (Fig. 3C) and protein levels (Fig. 3D), as evidenced by RT-qPCR and
IHC analysis, respectively.
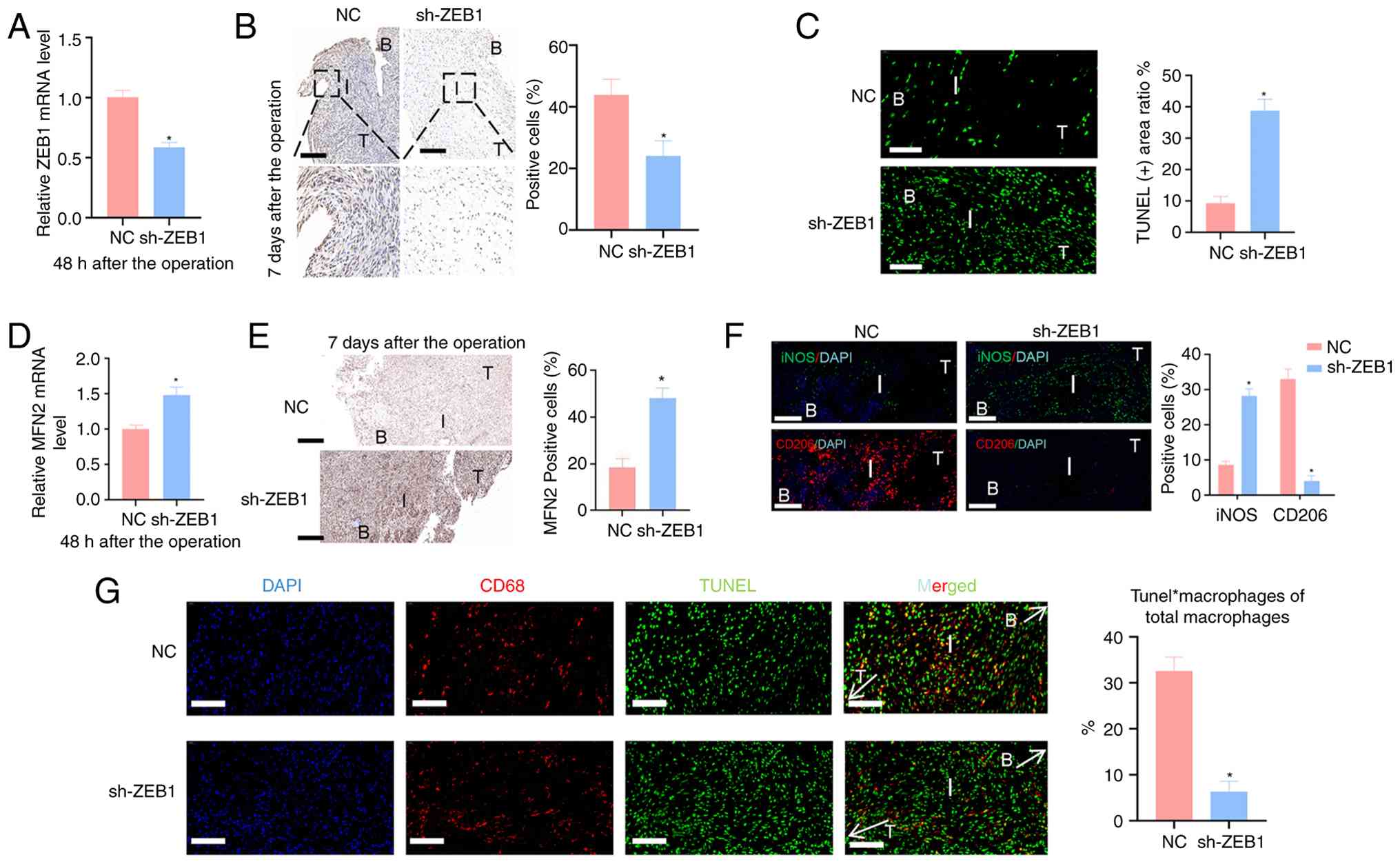 | Figure 3ZEB1 knockdown aggravates the
inflammatory response at the TBI. (A) The expression level of ZEB1
was detected by RT-qPCR 48 h after the operation. (B) The
expression of ZEB1 protein was detected by IHC 1 week after the
operation, and the optical density value was semi-quantitatively
analyzed by ImageJ software (n=3 high-power fields) (scale bar, 50
µm; lower panels, ×50 magnification, scale bar, 20
µm). (C) Apoptotic cells at the TBI were detected by TUNEL
method in the early postoperative period (1 week), with green
fluorescence (FITC labeled) indicating apoptotic cells with DNA
fragmentation (scale bar, 20 µm). (D) The expression level
of MFN2 was detected by RT-qPCR 48 h after the operation. (E) The
expression of MFN2 protein was detected by IHC 1 week after the
operation, and the optical density value was semi-quantitatively
analyzed by ImageJ software (scale bar, 50 µm). (F) At 1
week after the operation, double immunofluorescence staining was
used to detect the polarization phenotype of macrophages. The cell
nuclei were stained with DAPI (red), M1 macrophages were labeled
with iNOS (green), and M2 macrophages were labeled with CD206 (red)
(scale bar, 20 µm) (G) At 1 week after the operation,
co-localization analysis of apoptotic cells (Tunel, green) and
macrophages (CD68, red) at the TBI was performed by
immunofluorescence. The cell nuclei were stained with DAPI (blue)
(scale bar, 20 µm). The labels in the figure are as follows:
T=tendon, B=bone tissue, I=tendon-bone interface. n=3.
*P<0.05 vs. NC group. TBI, tendon-bone interface;
ZEB1, zinc finger E-box binding homeobox 1; RT-qPCR, reverse
transcription-quantitative PCR; IHC, immunohistochemistry; MFN2,
Mitofusin-2; iNOS, inducible nitric oxide synthase; NC, negative
control; sh, short hairpin. |
The impact of Zeb1 knockdown on cellular
apoptosis and macrophage behavior was then investigated. TUNEL
staining at 1 week post-operation revealed a significantly higher
density of ACs (green fluorescence) at the TBI in the sh-ZEB1 group
compared with the NC group (Fig.
3E), indicating that the efficiency of recognition and
clearance of ACs by macrophages at the TBI was significantly
reduced after Zeb1 knockdown. To further characterize the
macrophage response, polarization phenotypes were examined using
double immunofluorescence staining. At 1 week post-surgery, the
sh-ZEB1 group exhibited a significant increase in the positive area
of the M1 marker iNOS (green), accompanied by a marked decrease in
the M2 marker CD206 (red) compared to the NC group (Fig. 3F). These results indicate that
underexpression of ZEB1 shifts the macrophage population toward a
pro-inflammatory M1 phenotype in the early postoperative
period.
Finally, to clarify whether this inflammatory shift
was linked to defective efferocytosis, co-localization analysis of
macrophages (CD68, red) and ACs (TUNEL, green) was performed.
Immunofluorescence results showed that ZEB1 deficiency
significantly impaired the physical association and clearance of
ACs by macrophages at the TBI (Fig.
3G). Collectively, these findings suggest that ZEB1 is needed
to maintain efficient efferocytosis and to modulate macrophage
polarization during the early stages of tendon-to-bone healing.
ZEB1 deficiency delays the healing of the
TBI tissue
At 4 and 8 weeks postoperatively, the structure of
the TBI was examined by H&E staining and SO/FG staining
(Fig. 4A and B) in order to
capture the remodeling and maturation phases of healing.
Histological healing of the TBI was evaluated using Ide
histological scoring system. A maximum score of 16 indicates normal
TBI, with higher scores reflecting superior healing quality. As the
healing process of the rotator cuff injury progressed, the
continuity of the TBI was relatively well restored at 8 weeks
postoperatively. Compared with the 4-week time point, there was a
marked increase in fibrocartilage cells and proteoglycan matrix at
the interface, and the fiber arrangement exhibited a more organized
and aligned pattern. In comparison to the sh-ZEB1 group, the NC
group demonstrated a significantly thicker fibrocartilage layer,
along with improved chondrocyte polarity and maturation. The
inter-rater reliability analysis demonstrated excellent agreement
between the two independent observers, with a Cohen's κ coefficient
of 0.83. Based on the Ide scoring system, the histological
quantification score in the NC group was significantly higher than
that in the sh-ZEB1 group (Fig.
4D), with statistically significant differences at 4 and 8
weeks.
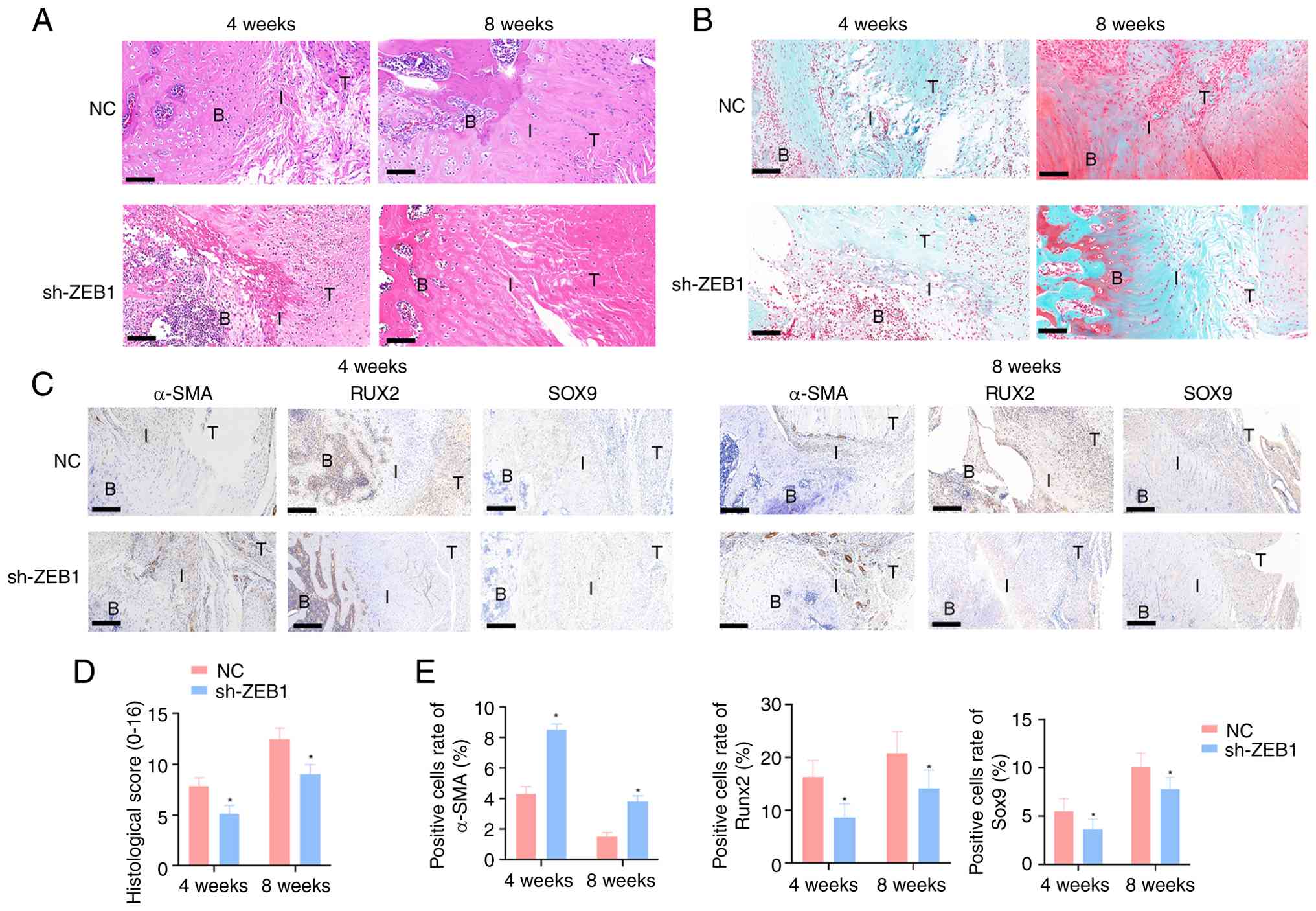 | Figure 4ZEB1 deficiency impairs the
morphological healing of the TBI tissue. (A) H&E and (B) SO/FG
staining were used to show the histological changes at the TBI at 4
weeks and 8 weeks postoperatively (scale bar, 50 µm). (C)
Immunohistochemical detection of the expression and distribution of
α-SMA, RUNX2 and SOX9 at the TBI. Positive signals are
brownish-yellow granules, and the cell nuclei are counterstained
with hematoxylin (blue) (scale bar, 50 µm). (D) Results of
Ide histological quantitative scoring. (E) Bar charts showing the
histological scores and quantitative analysis of the positive cell
rates of α-SMA, RUNX2 and SOX9 at 4 and 8 weeks postoperatively.
The labels in the figure are as follows: T=tendon, B=bone tissue,
I=tendon-bone interface. Data are presented as mean ± standard
deviation. n=3. *P<0.05 vs. NC. TBI, tendon-bone
interface; ZEB1, zinc finger E-box binding homeobox 1; H&E,
hematoxylin and eosin; SO/FG, safranin O-fast green; NC, negative
control; sh, short hairpin. |
To investigate the formation of fibrous scar tissue
and the expression of osteogenic and chondrogenic markers during
the repair of rotator cuff injuries in rats, IHC staining was
performed to assess the expression levels of α-SMA (a fibrotic scar
marker), RUNx2 (an osteogenic marker) and SOX9 (a chondrogenic
marker) (Fig. 4C). The results
demonstrated that ZEB1 deficiency during the healing process
significantly promoted the formation of fibrous scar in the TBI
while suppressing the development of bone and cartilage tissues,
with statistically significant differences (Fig. 4E).
Furthermore, at 4 weeks post-surgery, collagen
fibers in the sh-ZEB1 group exhibited a more disorganized
arrangement compared to those in the NC group, and the yellow-red
birefringent areas indicative of type I collagen were reduced.
Semi-quantitative analysis based on the intensity of birefringence
under polarized light microscopy revealed no significant difference
between the two groups. By 8 weeks post-surgery, collagen fibers in
both groups displayed a more regular and aligned structure with
enhanced birefringence, although the NC group showed superior
healing outcomes. Semi-quantitative analysis further confirmed that
the NC group had significantly higher levels of type I collagen
compared to the sh-ZEB1 group (Fig.
5D).
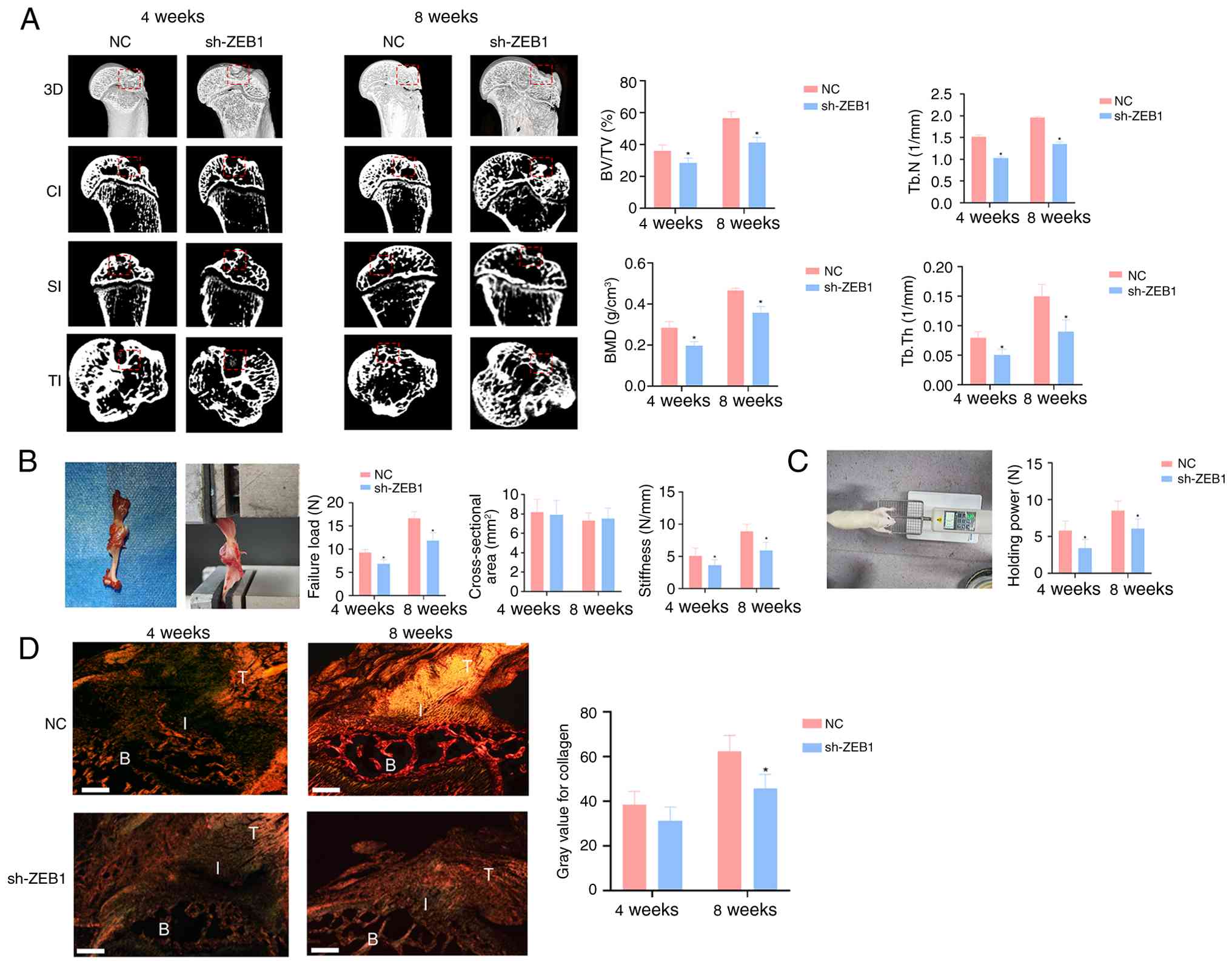 | Figure 5ZEB1 deficiency impairs bone
regeneration and biomechanical properties at the tendon-bone
interface. (A) Representative Micro-CT images and quantitative
analysis of BV/TV, Tb.N, BMD and Tb.Th. The red dashed boxes
indicate the regions of interest within the necrotic lesions of the
weight-bearing area for quantitative analysis and morphological
observation. (B) Gross observation of biomechanical testing along
with quantitative analysis of failure load, cross-sectional area,
and stiffness. (C) Gross observation and quantitative analysis of
grip strength. (D) Picrosirius Red staining showing the recovery of
the tendon-bone junction and semi-quantitative analysis of type I
collagen content based on optical density (scale bar, 50
µm). The labels in the figure are as follows: T=tendon,
B=bone tissue, I=tendon-bone interface. n=3. *P<0.05
vs. NC. TBI, tendon-bone interface; ZEB1, zinc finger E-box binding
homeobox 1; BV/TV, bone volume fraction; Tb.N, trabecular number;
BMD, bone mineral density; Tb.Th, trabecular thickness; NC,
negative control; sh, short hairpin; CI, coronal interface; SI,
sagittal interface; TI, transverse interface. |
Micro-CT scanning was performed at 4 and 8 weeks
postoperatively to evaluate bone formation at the tendon-bone
healing interface. Three-dimensional reconstruction of the TBI
specimens was conducted, and quantitative analysis of the ROI was
carried out (Fig. 5A). The
three-dimensional morphological data revealed the growth and
remodeling of subchondral bone at the TBI in rats. At 4 weeks
post-surgery, minimal bone ingrowth into the subchondral region was
observed, whereas by 8 weeks, new bone formation was evident at the
interface. ROI-based quantitative analysis demonstrated that, at
both 4 and 8 weeks postoperatively, the NC group exhibited
significantly higher values of BV/TV, Tb.Th, BMD, and Tb.N in the
subchondral bone at the TBI compared with the sh-ZEB1 group.
As the healing period progressed, biomechanical
parameters such as failure load and stiffness gradually increased.
Biomechanical analysis (Fig. 5B)
revealed no significant differences in the cross-sectional area of
the TBI among groups at 4 and 8 weeks post-surgery. However, at
both time points, the NC group exhibited significantly higher
failure load and stiffness at the TBI compared to the sh-ZEB1
group. In parallel, grip strength measurements (Fig. 5C) demonstrated that the NC group
displayed significantly greater grip strength than the sh-ZEB1
group at both 4 and 8 weeks postoperatively.
Collectively, these findings indicated that ZEB1
deficiency during the healing of rotator cuff injuries in rats
impaired histological repair of the TBI and compromised the
regeneration and remodeling of subchondral bone, ultimately
resulting in reduced biomechanical integrity and functional grip
performance.
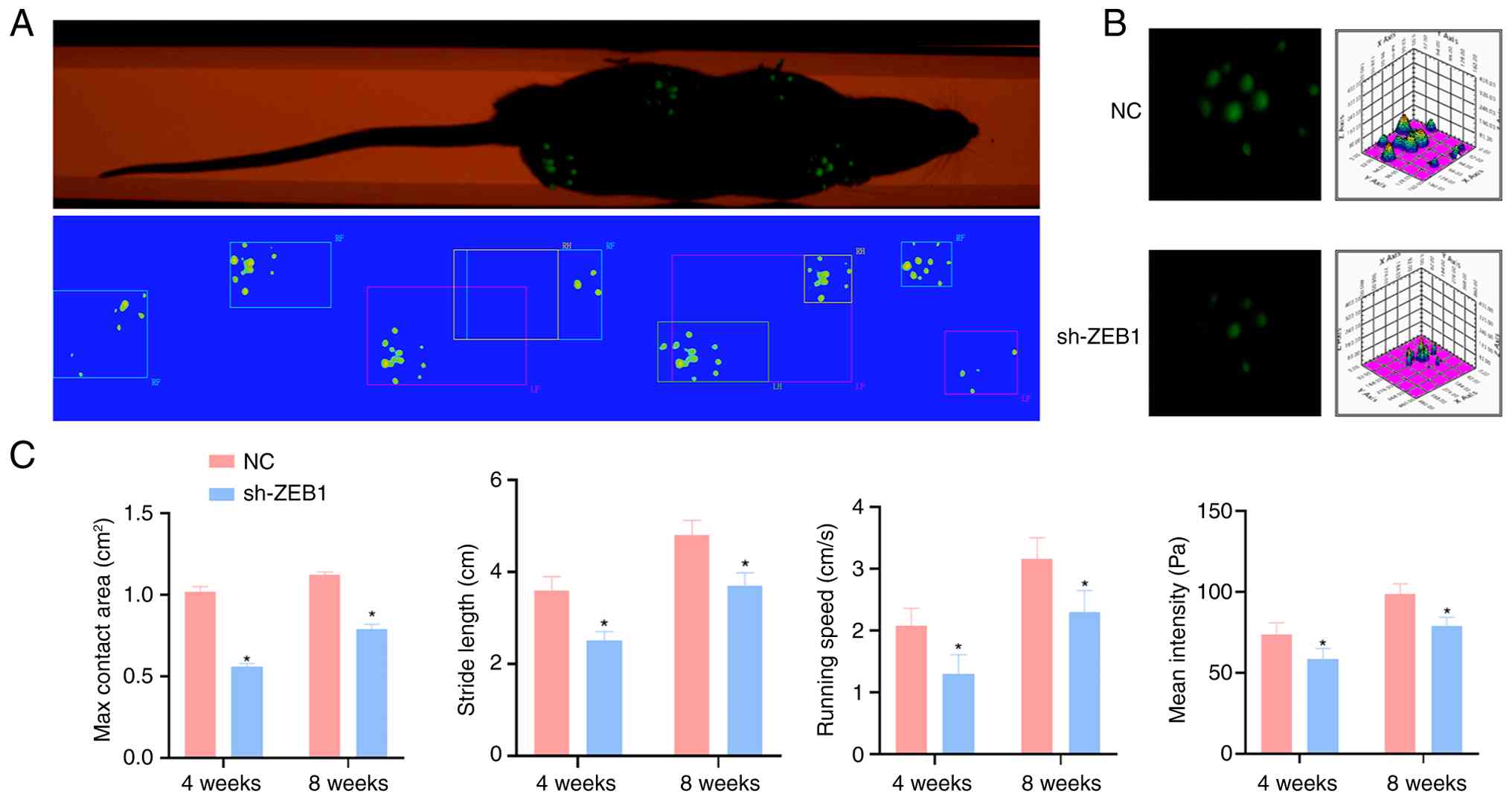
ZEB1-knockdown impairs postoperative gait
recovery in rats
The Greenwalk system was used to evaluate the
walking movement ability of rats after rotator cuff injury repair,
in order to obtain detailed quantitative analysis of motor
function. The footprints of the rats were successfully captured by
a high-resolution camera placed under a transparent glass plate
(Fig. 6A), and a
three-dimensional volcano plot was generated based on pressure
intensity (Fig. 6B). At 4 and 8
weeks after surgery, the maximum contact area of swing, stride
length, running speed and average pressure of the NC group were all
significantly higher than those of the sh-ZEB1 group (Fig. 6C).
Discussion
The present study aimed to investigate the impact of
ZEB1 deficiency on macrophage efferocytosis and its subsequent
effects for rotator cuff healing. To the best of our knowledge,
this is the first study to systematically elucidate the
interrelationships among macrophage ZEB1 expression, efferocytosis,
polarization and tendon-bone junction repair within an animal model
of rotator cuff injury. The key findings demonstrated that ZEB1
deficiency significantly impaired macrophage efferocytosis.
Mechanistically, our results demonstrate that ZEB1 modulates
MFN2-mediated mitochondrial dynamics, a process critically involved
in the regulation of efferocytosis. These effects may be closely
associated with the modulation of the inflammatory microenvironment
at the TBI. Therefore, targeting ZEB1 may represent a promising
therapeutic strategy for enhancing tendon-bone healing.
During the process of tendon-bone healing,
inflammatory microenvironment disorder, cellular senescence and the
persistent activation of myofibroblasts are all important factors
for the occurrence and progression of the disease (25). As central mediators of
inflammatory infiltration, macrophages play an essential role in
tissue injury and repair. Therefore, the ability to effectively
reduce excessive inflammatory responses and prevent excessive scar
formation, and restore the gradient structure at the tendon-bone
junction as much as possible (26) has become a focus of current
academic research. Efferocytosis is the specialized process by
which ACs are recognized, engulfed and digested by phagocytes,
particularly macrophages, guided by chemokine gradients. This
mechanism is mainly responsible for the clearance of ACs (27). In the absence of effective
efferocytosis, ACs may release cytotoxic metabolic byproducts that
can initiate excessive inflammatory responses, ultimately resulting
in secondary necrosis (6,28).
Efferocytosis is a highly regulated biological process that
encompasses the recognition, binding, internalization and
degradation of ACs. Beyond merely preventing secondary
necrosis-induced damage, this process actively reprograms
macrophages to secrete pro-reparative signaling molecules, thereby
playing a pivotal role in tissue regeneration following injury or
inflammatory insult.
In recent years, growing evidence has highlighted
the critical role of macrophage polarization in tendon-bone healing
(29,30). However, the relationship between
macrophage efferocytosis and tendon-bone healing remains poorly
understood, with significant gaps still present in current
research. The present study revealed that the expression level of
ZEB1 in macrophages was time-dependent with macrophage
efferocytosis. Meanwhile, knockdown of ZEB1 in macrophages led to a
decrease in their efferocytic efficiency, indicating that ZEB1 was
involved in the process of macrophage efferocytosis. ZEB1 has been
widely studied for its role in triggering the EMT program in cancer
cells and regulating stem cell functions (10,15). However, the latest research
evidence indicates that it can also promote the plasticity of
non-malignant cells from various tissue sources, including
macrophages (14). In the
context of skeletal physiology, ZEB1 serves as a pivotal regulator
of mesenchymal stem cell differentiation and bone development,
playing a crucial role in balancing osteogenesis and adipogenesis
(13). Furthermore, the loss of
ZEB1 impairs the formation of type H vessels, thereby exacerbating
osteonecrosis (31). Meanwhile,
a study has indicated that the downregulation of ZEB1 enhances
atherosclerotic plaque formation in mice and is closely associated
with the aggravation of chronic inflammation (15). The present study indicated that
knocking down the homeostatic level of ZEB1 in macrophages led to
poor healing at the TBI. The underlying mechanism involves the
regulation of MFN2-mediated mitochondrial dynamics, which directly
affects macrophage efferocytosis. The present study revealed that
ZEB1 expression is essential for maintaining mitochondrial fission
and preserving the efficiency of macrophage efferocytosis.
Mitochondria play a crucial role in cellular
functions such as energy metabolism, calcium homeostasis regulation
and reactive oxygen species (ROS) generation (32,33). Mitochondrial homeostasis is
governed by a finely tuned equilibrium between fission and fusion
processes. This dynamic balance is primarily orchestrated by DRP1
and mitochondrial fusion proteins 1 and 2 (MFN1/2), which serve as
master regulators in sustaining mitochondrial structural and
functional integrity (32). A
number of studies have shown that efferocytosis is a process
mediated by mitochondria, and mitochondrial metabolism,
mitochondrial dynamics, and communication between mitochondria and
other organelles all affect the clearance of ACs by macrophages
(17,34). The present study demonstrated
that Zeb1 knockdown in macrophages resulted in the formation
of a distinct tubular mitochondrial network, a morphological change
that aligned with previous observations by Zhang et al
(16); this study revealed that
ZEB1, functioning as a transcriptional repressor, directly binds to
the promoter region of MFN2 and suppresses its expression (16). This downregulation of MFN2 by
ZEB1 disrupts mitochondrial fusion, leading to a fragmented
mitochondrial morphology characterized by small, rounded
structures. By contrast, the absence of ZEB1 promotes the formation
of elongated, tubular mitochondrial networks (15). Collectively, these findings
indicate that ZEB1 depletion enhances mitochondrial fusion through
the upregulation of MFN2 expression. Emerging evidence has firmly
established a link between mitochondrial dynamics and phagocytic
activity (17). In this study,
reduction of the mitochondrial fission protein DRP1 in macrophages
was shown to diminish phagocytic capacity, whereas depletion of the
mitochondrial fusion protein MFN1 enhanced phagocytosis. These
findings indicate a positive association between mitochondrial
fission and phagocytic efficiency. Our data demonstrated that MFN2
expression was significantly upregulated in ZEB1-knockdown
macrophages, and increased MFN2 levels promoted mitochondrial
fusion, thereby attenuating phagocytosis. Notably, the present
study observed that ZEB1 deficiency did not significantly affect
DRP1 expression levels. We hypothesized that this is due to ZEB1
directly binding to the promoter region of MFN2 as a
transcriptional repressor, directly inhibiting the expression of
MFN2 (16). Secondly, the
process of mitochondrial fission itself is positively associated
with the ability of phagocytosis. When macrophages phagocytize ACs,
it further enhances mitochondrial fission. Although DRP1 expression
remains unchanged, enhanced mitochondrial fusion may shift the
fission-fusion equilibrium, resulting in a more tubular,
network-like mitochondrial morphology. Collectively, these
observations support the notion that ZEB1 modulates macrophage
phagocytosis-most plausibly via suppression of MFN2
expression-thereby limiting mitochondrial fusion and supporting
efferocytosis.
The present study found that ZEB1 plays a crucial
role in maintaining macrophage efferocytosis, and this effect could
in turn promote M2 polarization, thereby forming a positive
feedback cycle of an anti-inflammatory microenvironment. A previous
study has indicated that macrophage polarization phenotypes are
closely associated with their capacity to clear ACs (35). M1 macrophages primarily drive the
early inflammatory response by secreting IL-1β and TNF-α. Although
they possess a certain degree of phagocytic function, their
efficiency in recognizing and clearing ACs is generally lower than
that of the M2 phenotype. By contrast, M2 macrophages express high
levels of receptors such as MerTK and CD206, exhibiting a more
efficient efferocytic capacity. They are specialized in the timely
clearance of ACs to prevent secondary necrosis and inflammatory
leakage (36). More importantly,
efferocytosis is not merely a clearance process but a key driving
force for inducing M2 polarization. After macrophages engulf ACs,
they activate intracellular signaling pathways that inhibit the
release of pro-inflammatory mediators and instead secrete TGF-β and
IL-10, thereby further promoting the conversion of macrophages to
the M2 anti-inflammatory phenotype (37). Das et al (38) suggests that this efficient
efferocytosis acts as an important 'signal' for the conversion of
M0 to the anti-inflammatory M2 phenotype. Our in vivo
experimental data show that ZEB1-knockdown leads to a significant
increase in ACs at the TBI, a marked decrease in efferocytosis
efficiency, a significant increase in the area of iNOS positivity
and a significant decrease in the area of CD206 positivity. It is
worth noting that, although our in vitro experimental
results show that Zeb1 knockdown does not affect macrophage
polarization, this does not present a conflict. This is because our
in vitro experiments were conducted in an environment
without apoptotic cell accumulation, using isolated macrophages.
The direct impact of Zeb1 knockdown on intrinsic
polarization signaling pathways may be minimal. However, in the
in vivo tendon-bone injury environment, there is a large
number of ACs. In this context, the impaired efferocytosis due to
ZEB1 deficiency leads to the accumulation of uncleared ACs
(secondary necrosis), which in turn drives the inflammatory
response and alters macrophage polarization. Taken together, all
these results indicate that Zeb1 knockdown impairs
macrophage efferocytosis, reduces M2 polarization and aggravates
inflammation at the TBI. Moreover, previous studies have confirmed
that inducing M2 macrophage aggregation at the TBI through
biological intervention in the early postoperative period is
beneficial for tendon-bone healing (39,40). In summary, these findings suggest
that ZEB1 plays a key role in regulating the inflammatory
microenvironment during tendon-bone healing by modulating
macrophage efferocytosis and M2 polarization.
The natural gradient structure of the TBI is
composed of tendon, uncalcified fibrocartilage, calcified
fibrocartilage and bone (41).
Following injury, this delicate architecture is frequently replaced
by fibrovascular scar tissue, which possesses inferior mechanical
properties. Achieving an ideal, site-specific transition interface
remains a significant challenge. Currently, researchers believe
that the biomechanical properties of this interface are closely
related to bone ingrowth and bone integration. Notably, enhanced M2
polarization is associated with a reduced TBI width and improved
fiber alignment. (29,42). Our micro-CT analysis demonstrated
that all groups exhibited an increase in microstructural parameters
between 4 and 8 weeks post-surgery. However, the ZEB1-knockdown
group displayed significantly impaired bone formation and reduced
bone ingrowth around the tunnels. This is also similar to the
previous research results of our research group, that is, the
absence of ZEB1 has an important impact on vascular-dependent bone
regeneration (31). Efficient
clearance of ACs is pivotal for maintaining bone homeostasis and
immune equilibrium. Dysfunction in this process not only disrupts
normal bone metabolism but also amplifies inflammatory responses,
potentially contributing to the development of inflammatory bone
disorders (43). Histological
evaluation revealed that at 4 weeks post-operation, the TBI in all
groups was predominantly occupied by disorganized fibrovascular
tissue. By the 8th week, however, the continuity of the TBI was
markedly improved. Notably, compared to the sh-ZEB1 group, the NC
group exhibited a significantly thicker fibrocartilage layer, with
more organized and mature chondrocyte arrangement. Furthermore, the
histological scoring system confirmed a significantly higher tissue
quality score in the NC group relative to the sh-ZEB1 group.
Furthermore, immunohistochemical and Sirius Red staining analyses
demonstrated that Zeb1 knockdown significantly exacerbated
scar formation at the TBI, markedly reduced osteogenesis and
chondrogenesis, and led to more disorganized collagen architecture
and tissue structure. These pathological alterations ultimately
resulted in a significant decline in both the maximum failure load
and stiffness in the experimental animals. In addition, functional
assessments of postoperative grasping force and gait recovery
revealed that the sh-ZEB1 group exhibited significantly decreased
grasping force, maximum contact area, stride length, running speed
and average pressure compared with the control group. These
findings suggest that favorable structural healing is closely
associated with improved functional outcomes, highlighting the
potential of targeting ZEB1 to enhance TBI regeneration as a
promising therapeutic strategy for rotator cuff injuries.
The present study has several limitations that
warrant consideration. First, due to technical constraints
associated with surgical procedures, rats were selected as the
experimental model. Future studies should employ genetically
modified models, such as ZEB1-deficient or macrophage-depleted
rats, to more rigorously validate the observed findings. Second,
given that rodents do not fully recapitulate human pathophysiology,
further investigations in large mammalian models are necessary to
enhance translational relevance. Third, TUNEL staining lacks
specificity for polymorphonuclear neutrophils (PMNs). To provide
more definitive evidence, dual fluorescence labeling combining
TUNEL with PMN-specific immunofluorescent markers should be
employed for precise identification of apoptotic neutrophils.
Nonetheless, the TUNEL results obtained in the present study do
indicate a general trend toward improved apoptotic cell clearance.
Finally, the present study focused solely on the regulatory effects
of ZEB1-knockdown on macrophages and did not assess the potential
benefits of ZEB1 overexpression. This was primarily due to the
suboptimal transfection efficiency of the ZEB1 plasmid in
macrophages. Despite testing multiple transfection reagents and
optimizing DNA concentration (500-1,000 ng/µl), transfection
efficiency remained low, and higher reagent concentrations induced
notable cytotoxicity. Therefore, in our subsequent research, we
plan to explore its upstream regulation, with the aim of perfecting
the regulatory network of ZEB1-mitochondrial dynamics-
efferocytosis. In summary, the absence of ZEB1 disrupts the
delicate balance of the inflammatory microenvironment at the
tendon-bone healing interface, thereby impeding the physiological
trajectory of tissue repair. This impairment is mechanistically
driven by the dysregulation of MFN2-mediated mitochondrial dynamics
which subsequently compromises macrophage efferocytosis. Therefore,
targeting ZEB1 may be a potential strategy to improve tendon-bone
healing.
Availability of data and materials
Data supporting the findings of this study are
available from the corresponding author upon reasonable
request.
Authors' contributions
LHF conceived and supervised the study. YZ and YKZ
suggested experiments. YZ, TL and SQZ performed experiments. SQZ,
XZ and TL performed statistical analysis. YZ, YKZ and KN did animal
experimental procedures. YKZ, KN and XZ provided critical advice.
YZ, YKZ and LHF wrote the paper; YZ, KN and LHF confirm the
authenticity of all the raw data. All authors read and approved the
final reviewed the final manuscript.
Ethics approval and consent to
participate
This animal experiment protocol was approved by the
Animal Experiment Ethics Committee of Xi'an Jiaotong University
(approval no. XJTUAE2025-2322).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Abbreviations:
|
TBI
|
tendon-bone interface
|
|
ZEB1
|
zinc finger E-box binding homeobox
1
|
|
BMDMs
|
bone marrow-derived macrophages
|
|
ACs
|
apoptotic cells
|
|
MFN2
|
Mitofusin-2
|
|
DRP1
|
dynamin-related protein 1
|
Acknowledgments
Not applicable.
Funding
The APC was funded by the National Natural Science Foundation of
China (grant no. 81772355).
References
|
1
|
Bedi A, Bishop J, Keener J, Lansdown DA,
Levy O, MacDonald P, Maffulli N, Oh JH, Sabesan VJ, Sanchez-Sotelo
J, et al: Rotator cuff tears. Nat Rev Dis Primers. 10:82024.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Font Tellado S, Balmayor ER and Van
Griensven M: Strategies to engineer tendon/ligament-to-bone
interface: Biomaterials, cells and growth factors. Adv Drug Deliv
Rev. 94:126–140. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Tokunaga T, Karasugi T, Tanimura S and
Miyamoto T: Association of severe histological degeneration of the
torn supraspinatus tendon and retear after arthroscopic repair of
full-thickness rotator cuff tears using the suture bridge
technique. Am J Sports Med. 51:2411–2421. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Muscat S, Nichols AEC, Gira E and Loiselle
AE: CCR2 is expressed by tendon resident macrophage and T cells,
while CCR2 deficiency impairs tendon healing via blunted
involvement of tendon-resident and circulating
monocytes/macrophages. FASEB J. 36:e226072022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wynn TA and Vannella KM: Macrophages in
tissue repair, regeneration, and fibrosis. Immunity. 44:450–462.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mantovani A, Cassatella MA, Costantini C
and Jaillon S: Neutrophils in the activation and regulation of
innate and adaptive immunity. Nat Rev Immunol. 11:519–531. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Elliott MR, Koster KM and Murphy PS:
Efferocytosis signaling in the regulation of macrophage
inflammatory responses. J Immunol. 198:1387–1394. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Tang X, Huang Z, Wang F, Chen J, Qin D,
Peng D and Yu B: Macrophage-specific deletion of MIC26 (APOO)
mitigates advanced atherosclerosis by increasing efferocytosis.
Atherosclerosis. 386:1173742023. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Trzeciak A, Wang YT and Perry JSA: First
we eat, then we do everything else: The dynamic metabolic
regulation of efferocytosis. Cell Metab. 33:2126–2141. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang Y, Zhang Q, He T, Wang Y, Lu T, Wang
Z, Wang Y, Lin S, Yang K, Wang X, et al: The transcription factor
Zeb1 controls homeostasis and function of type 1 conventional
dendritic cells. Nat Commun. 14:66392023. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Jiang H, Wei H, Wang H, Wang Z, Li J, Ou
Y, Xiao X, Wang W, Chang A, Sun W, et al: Zeb1-induced metabolic
reprogramming of glycolysis is essential for macrophage
polarization in breast cancer. Cell Death Dis. 13:2062022.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cortés M, Brischetto A,
Martinez-Campanario MC, Ninfali C, Domínguez V, Fernández S, Celis
R, Esteve-Codina A, Lozano JJ, Sidorova J, et al: Inflammatory
macrophages reprogram to immunosuppression by reducing
mitochondrial translation. Nat Commun. 14:74712023. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhu L, Tang Y, Li XY, Kerk SA, Lyssiotis
CA, Feng W, Sun X, Hespe GE, Wang Z, Stemmler MP, et al: A
Zeb1/MtCK1 metabolic axis controls osteoclast activation and
skeletal remodeling. EMBO J. 42:e1111482023. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chen J, Guan X, Chen L, Zheng B, Li F,
Fang C, Fu Y, Li X, Wang H and Zhou Y: Customized hydrogel system
for the spatiotemporal sequential treatment of periodontitis
propelled by ZEB1. Adv Sci (Weinh). 12:e25033382025. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Martinez-Campanario MC, Cortés M,
Moreno-Lanceta A, Han L, Ninfali C, Domínguez V, Andrés-Manzano MJ,
Farràs M, Esteve-Codina A, Enrich C, et al: Atherosclerotic plaque
development in mice is enhanced by myeloid ZEB1 downregulation. Nat
Commun. 14:83162023. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhang K, Zhao H, Sheng Y, Chen X, Xu P,
Wang J, Ji Z, He Y, Gao WQ and Zhu HH: Zeb1 sustains hematopoietic
stem cell functions by suppressing mitofusin-2-mediated
mitochondrial fusion. Cell Death Dis. 13:7352022. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang Y, Subramanian M, Yurdagul A Jr,
Barbosa-Lorenzi VC, Cai B, de Juan-Sanz J, Ryan TA, Nomura M,
Maxfield FR and Tabas I: Mitochondrial fission promotes the
continued clearance of apoptotic cells by macrophages. Cell.
171:331–345.e22. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hu L, Ding M, Tang D, Gao E, Li C, Wang K,
Qi B, Qiu J, Zhao H, Chang P, et al: Targeting mitochondrial
dynamics by regulating Mfn2 for therapeutic intervention in
diabetic cardiomyopathy. Theranostics. 9:3687–3706. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Davis HE, Morgan JR and Yarmush ML:
Polybrene increases retrovirus gene transfer efficiency by
enhancing receptor-independent virus adsorption on target cell
membranes. Biophys Chem. 97:159–172. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2 (-Delta Delta C (T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
21
|
Ide J, Kikukawa K, Hirose J, Iyama K,
Sakamoto H, Fujimoto T and Mizuta H: The effect of a local
application of fibroblast growth factor-2 on tendon-to-bone
remodeling in rats with acute injury and repair of the
supraspinatus tendon. J Shoulder Elbow Surg. 18:391–398. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ai Y, Hu C, Wang Y, Liu Y, Liu R, Xu H, Li
H, Zhao Y, Wang Y, Ning R, et al: Core-shell hydrogel microspheres
with sequential delivery of cerium oxide nanoparticles and spinal
white matter extracellular matrix for improved functional recovery
in spinal cord injury. Chem Eng J. 508:1608612025. View Article : Google Scholar
|
|
23
|
Li P, Fan Z, Huang Y, Luo L and Wu X:
Mitochondrial dynamics at the intersection of macrophage
polarization and metabolism. Front Immunol. 16:15208142025.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Mishra SR, Mishra P, Senapati PK,
Mahapatra KK and Bhutia SK: Intricate role of DRP1 and associated
mitochondrial fission signaling in carcinogenesis and cancer
progression. Biochim Biophys Acta Rev Cancer. 1880:1894532025.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Han Q, Bai L, Qian Y, Zhang X, Wang J,
Zhou J, Cui W, Hao Y and Yang X: Antioxidant and anti-inflammatory
injectable hydrogel microspheres for in situ treatment of
tendinopathy. Regen Biomater. 11:rbae0072024. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Rossetti L, Kuntz LA, Kunold E, Schock J,
Müller KW, Grabmayr H, Stolberg-Stolberg J, Pfeiffer F, Sieber SA,
Burgkart R and Bausch AR: The microstructure and micromechanics of
the tendon-bone insertion. Nat Mater. 16:664–670. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kumar S and Birge RB: Efferocytosis. Curr
Biol. 26:R558–R559. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gheibi Hayat SM, Bianconi V, Pirro M and
Sahebkar A: Efferocytosis: Molecular mechanisms and
pathophysiological perspectives. Immunol Cell Biol. 97:124–133.
2019. View Article : Google Scholar
|
|
29
|
Li J, Ke H, Lei X, Zhang J, Wen Z, Xiao Z,
Chen H, Yao J, Wang X, Wei Z, et al: Controlled-release hydrogel
loaded with magnesium-based nanoflowers synergize immunomodulation
and cartilage regeneration in tendon-bone healing. Bioact Mater.
36:62–82. 2024.PubMed/NCBI
|
|
30
|
He Y, Lu S, Chen W, Yang L, Li F, Zhou P,
Chen Z, Wan R, Zhang Z, Sun Y, et al: Exosomes derived from tendon
stem/progenitor cells enhance tendon-bone interface healing after
rotator cuff repair in a rat model. Bioact Mater. 40:484–502.
2024.PubMed/NCBI
|
|
31
|
Zhang G, Cai Y, Liang J, Jing Z, Wei W, Lv
L, Dang X and Song Q: The decrease in zinc-finger E-box-binding
homeobox-1 could accelerate steroid-induced osteonecrosis of the
femoral head by repressing type-H vessel formation via
Wnt/β-catenin pathway. Animal Model Exp Med. 7:802–815. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tábara LC, Segawa M and Prudent J:
Molecular mechanisms of mitochondrial dynamics. Nat Rev Mol Cell
Biol. 26:123–146. 2025. View Article : Google Scholar
|
|
33
|
Chen W, Zhao H and Li Y: Mitochondrial
dynamics in health and disease: Mechanisms and potential targets.
Signal Transduct Target Ther. 8:3332023. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
No authors listed. Mitochondrial fission
in macrophages fuels phagocytosis of tumor cells. Nat Cancer.
3:384–385. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Doran AC, Yurdagul A Jr and Tabas I:
Efferocytosis in health and disease. Nat Rev Immunol. 20:254–267.
2020. View Article : Google Scholar
|
|
36
|
Li Y, Miao J, Liu C, Tao J, Zhou S, Song
X, Zou Y, Huang Y and Zhong L: Kushenol O regulates GALNT7/NF-κB
axis-mediated macrophage M2 polarization and efferocytosis in
papillary thyroid carcinoma. Phytomedicine. 138:1563732025.
View Article : Google Scholar
|
|
37
|
Zhang Y, Zang H and Wen Q: Macrophage
polarization in tumors: The impact of efferocytosis. Int
Immunopharmacol. 168:1158072026. View Article : Google Scholar
|
|
38
|
Das A, Ghatak S, Sinha M, Chaffee S, Ahmed
NS, Parinandi NL, Wohleb ES, Sheridan JF, Sen CK and Roy S:
Correction of MFG-E8 resolves inflammation and promotes cutaneous
wound healing in diabetes. J Immunol. 196:5089–5100. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Lu J, Chamberlain CS, Ji ML, Saether EE,
Leiferman EM, Li WJ and Vanderby R: Tendon-to-bone healing in a rat
extra-articular bone tunnel model: A comparison of fresh autologous
bone marrow and bone marrow-derived mesenchymal stem cells. Am J
Sports Med. 47:2729–2736. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Gao H, Wang L, Lin Z, Jin H, Lyu Y, Kang
Y, Zhu T, Zhao J and Jiang J: Bi-lineage inducible and
immunoregulatory electrospun fibers scaffolds for synchronous
regeneration of tendon-to-bone interface. Mater Today Bio.
22:1007492023. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Voleti PB, Buckley MR and Soslowsky LJ:
Tendon healing: Repair and regeneration. Annu Rev Biomed Eng.
14:47–71. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Dagher E, Hays PL, Kawamura S, Godin J,
Deng XH and Rodeo SA: Immobilization modulates macrophage
accumulation in tendon-bone healing. Clin Orthop Relat Res.
467:281–287. 2009. View Article : Google Scholar
|
|
43
|
Wang H, Zhang Y, Zhang Y, Li C, Zhang M,
Wang J, Zhang Y, Du Y, Cui W and Chen W: Activating macrophage
continual efferocytosis via microenvironment biomimetic short
fibers for reversing inflammation in bone repair. Adv Mater.
36:e24029682024. View Article : Google Scholar : PubMed/NCBI
|