Gastric cancer (GC) is the fifth most common
malignant tumor worldwide and has the third highest mortality rate;
its development is strongly associated with chronic infection with
Helicobacter pylori (H. pylori), a high-salt diet and
genetic susceptibility (such as a cadherin-1 mutation) (1-3).
Although neoadjuvant chemotherapy centered on the fluorouracil,
leucovorin, oxaliplatin and docetaxel (FLOT) regimen and targeted
therapies (such as trastuzumab against human epidermal growth
factor receptor 2) have significantly improved clinical outcomes,
the 5-year survival rate of patients is still <30% and drug
resistance-related recurrence and molecular heterogeneity [such as
differences in Epstein-Barr virus (EBV)-positive/genome-stable
subtypes] limit the efficacy of existing targeted strategies
(4,5). Therefore, analysis of the molecular
regulatory network of GC progression and exploration of novel
targets are urgently needed.
Post-transcriptional regulation of gene expression
is a key feature of cancer that precisely shapes the tumor
phenotype through the dynamic control of RNA metabolism (6,7).
As core effectors in post-transcriptional regulation, RNA-binding
proteins (RBPs) bind to single- or double-stranded RNA in a
sequence-specific manner and regulate mRNA splicing,
polyadenylation, mRNA stabilization and localization (8,9).
GC cells are also involved in ribonucleoprotein complex (RNP)
synthesis on the basis of their interactions with other
biomolecules, especially coding and noncoding RNAs (10). The binding of RBPs to structural
motifs or RNA sequences occurs through a set of structurally
defined RNA binding domains (RBDs). Depending on the presence or
absence of RBDs, they can be categorized as conventional/typical or
unconventional/atypical RBPs. Conventional RBPs include RNA
recognition motifs, cold shock structural domains, K homology
structural domains, DEAD/DEAH deconjugation enzymes and zinc finger
structural domains (11,12). Notably, RBPs bind to both DNA and
RNA and serve a role in DNA replication and the DNA damage response
(13,14).
RBPs are involved in the development of numerous
human diseases, including cancer neurodegenerative, renal and
cardiovascular diseases (15-17). RBPs can drive stemness
maintenance, metabolic reprogramming, immune escape and treatment
tolerance in GC (9,18-22). The present review focused on the
role of RBPs as post-transcriptional regulatory hubs to integrate
cutting-edge advances and provide a theoretical basis for targeted
therapeutic strategies (Table
I).
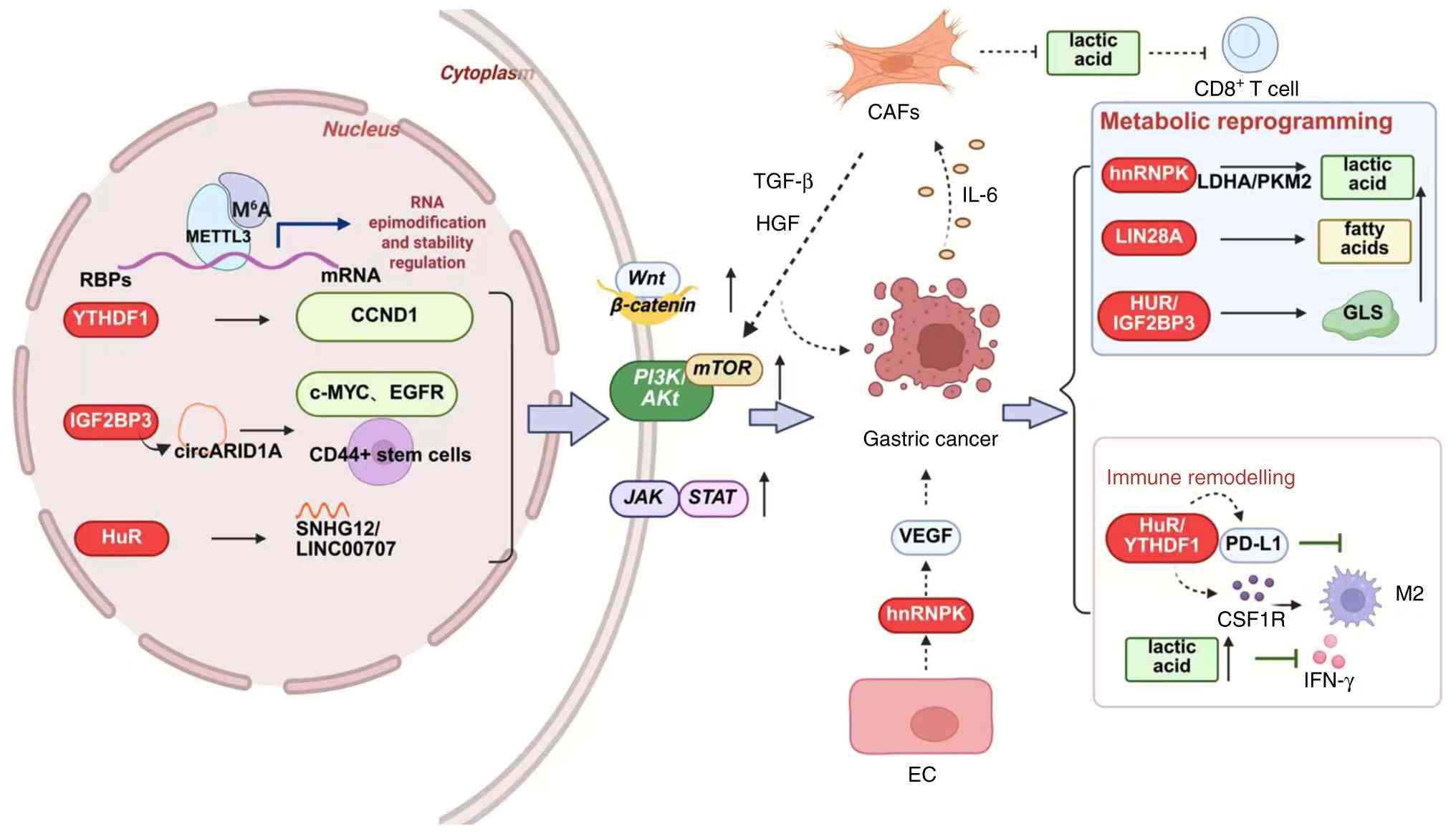
In cancer cells, RBPs serve important roles in
regulating the expression of tumor suppressor genes and
oncoproteins involved in various cell signaling pathways (9,23). In GC, RBPs are classified into
two categories according to their functional properties: i)
Oncogenic RBPs, including LIN28A/B, insulin-like growth factor 2
mRNA-binding protein (IGF2BP1/3), human antigen R (HuR) and YTH
N6-methyladenosine RNA binding protein F1/2 (YTHDF1/2) (24-28), which primarily drive GC
progression by stabilizing oncogenic mRNAs [such as c-MYC and
cyclin D1 (CCND1)], mediating N6-methyladenosine (m6A)
modification, activating pro-tumor signaling pathways (including
PI3K/AKT and Wnt/β-catenin), and facilitating metabolic
reprogramming or immune escape. ii) Tumor-suppressive RBPs,
including poly (rC)-binding protein 1 (PCBP1), zinc finger protein
36, RBM5 and QKI (29,30), which inhibit GC malignant
phenotypes through mechanisms such as degrading pro-metastatic
mRNAs (such as MMP9 and IL-33), regulating alternative splicing to
generate tumor-suppressive isoforms (for example, SMAD2ΔE3 and
caspase-9a), inducing cell cycle arrest or apoptosis, and
suppressing inflammatory niche formation.
The IGF2BP family comprises key post-transcriptional
regulators (IGF2BP1, IGF2BP2 and IGF2BP3) that regulate
tumor-associated gene expression by mediating RNA epigenetic
modifications. IGF2BP1 and IGF2BP3, as m6A reader
proteins, both stabilize target mRNAs via m6A
modification mediated by the methyltransferase-like 3 (METTL3), yet
exhibit subtype differences in target gene preference (26,33-35).
IGF2BP1 drives GC cell proliferation and metastasis
primarily by stabilizing target molecules [such as nuclear factor
erythroid 2-related factor 2 (NRF2) and c-MYC] via METTL3-mediated
m6A modification (36,37). At the noncoding RNA level, it
also forms complexes with pro-oncogenic long noncoding RNAs
(lncRNAs), such as ABL, GLCC1 and GHET1 to promote GC cell
migration and chemotherapy resistance, these complexes act by
inhibiting cytochrome c/APAF1 interactions (blocking caspase-9/3
activation), stabilizing c-MYC mRNA or activating the PI3K/AKT
pathway (37-41). The core innovative mechanism of
IGF2BP1 in GC lies in enhancing NRF2 mRNA stability via the
methyltransferase-like 5 (METTL5)/m6A/NRF2 axis. This
simultaneously inhibits ferroptosis whilst upregulating the
mitochondrial complex protein NDUFA4 to activate the
glycolysis-oxidation metabolic coupling. This energy metabolism
regulatory pattern remains unreported in colorectal cancer (CRC)
and breast cancer (BC), representing a unique strategy for GC cells
to sustain their malignant phenotype (42-45).
However, functional controversy persists: A previous
small-sample study reported that IGF2BP1 exhibits low expression
and tumor-suppressive effects (by downregulating MYC) in
EBV-positive GC (46), which is
in contrast to its pro-oncogenic role in genomically stable
subtypes. Furthermore, IGF2BP1 exerts tumor suppression in
hepatocellular carcinoma (HCC) by degrading the lncRNA HULC
(47), and the molecular
mechanism underlying this tissue-specific functional divergence
(such as dependence on distinct RBD targets) remains unclear.
Clinically, the specific inhibitor AVJ16 can efficiently reduce the
RNA-binding activity of IGF2BP1and the viability of GC cell lines
with high IGF2BP1 expression, showing potential for targeted
therapy (48).
YTH structural domain family proteins (including
YTHDF1-3) are core effector proteins of m6A
modification, and YTHDF1 and YTHDF2 serve key roles in gastric
carcinogenesis (24,61,62).
YTHDF1 drives malignant progression and therapeutic
resistance in GC through multidimensional mechanisms. Within
tumor-autonomous regulation, YTHDF1 promotes c-MYC translation to
activate the PI3K/AKT/mammalian target of rapamycin (mTOR) pathway,
upregulates FZD7/TCF7 to maintain tumor stemness, induces
SNAIL-mediated epithelial-mesenchymal transition (EMT), while
simultaneously stabilizing PARP1 mRNA to preserve CD133+
stem cell properties and enhances SPHK2/USP14 translation to
mediate oxaliplatin resistance (63-68). By contrast, within the tumor
immune microenvironment, YTHDF1 exerts a dual regulatory role
within the immune microenvironment: Suppressing T-cell activity via
m6A-dependent programmed death-ligand 1 (PD-L1)
translation, whilst simultaneously recruiting mature dendritic
cells upon its depletion, upregulates MHC II/IL-12 and activates
the JAK/STAT1 pathway to enhance CD4+/CD8+ T
cell infiltration and IFN-γ secretion (22,69-71). This regulatory pattern exhibits a
more multifaceted phenotype compared to the unidimensional action
solely dependent on the circMAP2K4/miR-139-5p axis observed in
hepatocellular carcinoma (72).
Furthermore, a clinical study indicated that
elevated YTHDF1 expression is significantly associated with
oxaliplatin resistance and poor prognosis in patients with GC. Its
inhibitor SAC could reverse drug resistance in MGC-803 cells by
inhibiting SPHK2/USP14 translation according to preclinical data
(73). Patients with high YTHDF1
expression exhibited higher PD-L1 positivity rates and reduced
CD8+ T-cell infiltration, suggesting its potential as a
predictive biomarker for immunotherapy response. The precision
treatment strategy combining SAC with PD-1 antibodies is currently
being preclinically validated in GC organoids (69). This suggests that combining SAC
with PD-1 antibodies may constitute a precision treatment strategy
for YTHDF1-overexpressing GC.
However, the bidirectional regulatory function of
YTHDF2 remains a subject of core controversy. Within the same GC
cell line, the spatiotemporal coexistence of progression-promoting
mechanisms (stabilizing JAK1/PRKAA1) and tumor growth-inhibiting
mechanisms (upregulating PPP2CA, regulating FOXC2) remains
unclarified. The molecular switches governing this functional
switch (such as m6A modification levels or differential
interactions with long non-coding RNAs) remain unidentified.
Furthermore, YTHDF2 exhibits tissue-specific functionality; whilst
it induces PD-L1 expression via JAK1-dependent pathways in GC,
YTHDF2 primarily regulates ARHGEF2 translation in CRC (80). The tissue-specific variations in
its immunomodulatory functions remain under-explored, potentially
limiting the universality of targeted therapies.
In GC, RBPs can specifically recognize and bind to
conserved sequences or structural elements of target mRNAs,
significantly enhancing the stability of these mRNAs and thereby
regulating the expression levels and translation efficiency of
downstream genes. The regulatory effects of these RBPs permeate key
stages of GC initiation and progression, covering multiple
dimensions including the maintenance of cancer stem cell stemness,
EMT, metabolic reprogramming, chemoresistance and immune escape
(81-83).
The hnRNP family serves as a key participant in the
post-transcriptional regulatory network of GC. As core members of
this family, hnRNPA1 and hnRNPK, regulate the initiation and
progression of GC in a multidimensional manner by stabilizing the
expression of target mRNAs. Although both proteins exert their
functions relying on mRNA stabilization, there are significant
differences in their regulatory focuses and functional
characteristics (81,82).
Among hnRNPs, hnRNPA1, which serves a central role
in hnRNPA1, promotes GC metastasis by stabilizing WNT1 inducible
signaling pathway protein 2 mRNA to activate Wnt/β-catenin
signaling, upregulate SNAIL/Vimentin expression and induce EMT
(84). Furthermore, hnRNPA1
regulates ferroptosis inhibition and chemoresistance in GC. hnRNPA1
binds downstream arachidonate 15-lipoxygenase and inhibits lipid
peroxidation; levels of reactive oxygen species (ROS) increase in
metastatic GC cells compared with that in primary tumor cells,
ultimately contributing to ferroptosis resistance and
chemoresistance (85); it also
upregulates stearoyl coenzyme A desaturase 1, which maintains the
oncogenicity of GC stem cells (86) and enhances solute carrier family
7 member 11 (SLC7A11) and glutathione peroxidase 4 (GPX4, an
antioxidant enzyme) expression, thus inhibiting lipid peroxidation
and promoting GC progression (87). Additionally, in the regulation of
metabolic reprogramming in GC, the lncRNA OIP5-AS1 drives
glycolysis-dependent proliferation and metastasis by inhibiting
Trim21-mediated ubiquitination degradation of hnRNPA1, which
promotes the expression of LDHA and PKM2 (57).
Notably, cancer-associated fibroblasts (CAFs) serve
as a core stromal component within the TME. Through multiple
pathways, CAFs interact with hnRNPA1, IGF2BP3, YTHDF2 and other
RBPs to form an interactive network that reinforces the malignant
phenotype of GC epithelial cells: i) miR-522 from CAF-derived
exosomes targets the ubiquitination site of hnRNPA1, inhibiting its
degradation to enhance the stabilization of SLC7A11 and GPX4
(87); ii) paracrine TGF-β
activates epithelial SMAD2/3 signaling to directly upregulate
IGF2BP3 transcription (88,89); and iii) secreted hepatocyte
growth factor activates epithelial c-MET signaling, inducing YTHDF2
expression. This protein stabilizes JAK1 mRNA via m6A
modification, activates the JAK/STAT pathway and upregulates PD-L1.
It also forms a positive feedback loop with CAF-secreted IL-6 to
suppress CD8+ T-cell infiltration (70). Such regulatory circuits involving
CAFs-TGF-β-IGF2BP3-epithelial cells not only expand the functional
dimensions of RBPs within the microenvironment but also demonstrate
their pivotal role as epithelial transcriptome coordinators linking
stromal signals to epithelial malignant transformation.
hnRNPK exhibits functional dynamics in GC and its
role is regulated by microenvironmental signals and genetic
background, such as p53 status. First, hnRNPK is involved in the
maintenance of cancer stem cell properties. hnRNPK increases
β-catenin stability and synergistically activates Wnt signaling to
drive GCSC self-renewal and chemoresistance (90), promotes the expression of the
prometastatic isoform CD44v6 and enhances invasiveness and stemness
through the modulation of the selective splicing of CD44 precursor
mRNAs (91). Binding to the
lncRNA ELF3-AS1 activates the signal transducer and STAT3 pathway
and induces the release of thrombopoietin, promoting
tumor-associated thrombocytosis and the formation of a metastatic
microenvironment (92). hnRNPK
upregulates key enzymes involved in glycolysis, hexokinase 2 (HK2)
and LDHA, which drive abnormal energy metabolism (93). Furthermore, hnRNPK promotes the
TPT1-OCT1/CDX2 transcriptional axis and induces gastrointestinal
epithelial chemotaxis, a precancerous stage of GC (94). However, the functional
controversy surrounding hnRNPK poses challenges for clinical
targeted therapies; in p53-mutant gastric carcinomas (GC), hnRNPK
exhibits significant oncogenic effects (stabilizing β-catenin and
driving glycolysis), whereas in p53-wild-type GC, hnRNPK exerts
tumor-suppressing effects by stabilizing p53 (95). Currently, p53 mutations account
for 40-60% of clinical gastric carcinoma cases (3). The primary controversy and
technical challenge in current research lies in designing
conditionally active inhibitors based on p53 status, such as those
selectively blocking hnRNPK-β-catenin interactions only in mutant
p53 contexts, to avoid compromising its tumor-suppressing
function.
LIN28A/B is a class of RBPs that regulate stem cell
properties and tumorigenesis. LIN28A/B drives the malignant
progression of GC by inhibiting lethal-7 (let-7) microRNA (miRNA)
family biosynthesis and directly binding to target mRNAs (82). Blocking let-7 maturation and
deregulating its inhibitory effects on proto-oncogenes (such as MYC
and RAS) by LIN28A/B maintains stem cell pluripotency and promotes
gastric carcinogenesis (82,96,97). In GC, the NF-κB/LIN28A/let-7a
axis accelerates carcinogenesis by activating telomerase human
telomerase reverse transcriptase (98). LIN28A is also involved in stem
cell and proliferation regulation, binding Oct4 (a stem cell
factor), insulin-like growth factor 2 and cell cycle-related mRNAs
(such as cyclin B1) to increase their stability or translational
efficiency and drive proliferation and dedifferentiation (96,99-102). In addition, the lin28A/B/let-7
axis regulates cellular metabolic reprogramming and enhances
glucose uptake and glycolysis through inhibition of the
insulin-PI3K-mTOR pathway (103,104). Components of this axis bind to
sterol-regulatory element binding proteins (SREBP-1)/SREBP
cleavage-activating protein mRNAs, which promotes fatty acid ab
initio synthesis and desaturation (saturated to unsaturated fatty
acid conversion), providing raw materials and energy for membrane
synthesis (105). Therefore,
targeting the LIN28A/B-let-7 axis may reverse GC stemness and drug
resistance.
HuR, also known as ELAVL1, is an RNA-binding
protein. It regulates the stability and translation of target mRNAs
by recognizing their AU-rich regions (AREs) and exerts pro-cancer
effects in GC via multiple mechanisms (83). HuR stabilizes VEGF mRNA, induces
tumor neovascularization, enhances the expression of the
antiapoptotic protein Bcl-2, ubiquitin-specific protease 1 (USP1)
and autophagy-related gene 4B, and mediates resistance to
chemotherapy (5-FU/cisplatin) (106-108). HuR also upregulates SNAIL and
vimentin at the mRNA level, and promotes EMT and metastasis
(109). HuR activates
pro-oncogenic signaling pathways by forming complexes with multiple
lncRNAs. The LINC00707/LINC00324 complex stabilizes vav guanine
nucleotide exchange factor 3/F11 receptor (VAV3/F11R) mRNA and
activates β-catenin. The small nucleolar RNA host gene 12 complex
stabilizes catenin b 1/tyrosine 3-monooxygenase/tryptophan
5-monooxygenase activation protein ζ (CTNNB1/YWHAZ) mRNA and
synergistically activates the Wnt and AKT/GSK-3β pathways, in
addition to the VCAN-AS1/RP11-138J23.1 complex, which upregulates
FAM83B (a metastasis-associated protein) expression (110-116). Furthermore, in synergy with
IGF2BP3, GLS drives glutamine catabolism (58) and activates the p38
MAPK-COX-2/IL-8 axis, promoting gastrin-dependent
inflammation-associated tumor progression (119). In addition, HuR inhibits CD8
T-cell activity and promotes immune escape by stabilizing PD-L1
mRNA (118,119).
The HuR inhibitor CMLD-2 combined with oxaliplatin
achieved a tumor suppression rate of 62% in patient-derived
xenograft (PDX) models of GC, significantly outperforming
monotherapy (120).
Furthermore, a nanoparticle-mediated CRISPR/Cas9-HuR knockout
system achieved tumor-specific delivery in head and neck cancer
models (121). This system has
been adapted for GC cell lines (SGC-7901), with in vitro
experiments confirming it significantly reduces VEGF and PD-L1
expression, potentially providing a viable delivery strategy for
subsequent GC clinical trials.
A mammalian target of rapamycin complex 1 (mTORC1)
pathway-dependent immunosuppressive factor. PUM1 drives tumor
progression in GC through multidimensional mechanisms: It activates
the DEPTOR-mediated glycolytic pathway; and promotes GC cell
development via circGMPS carried by exosomes through the
miR-144-3p/PUM1 axis, while simultaneously regulating the NPM3/NPM1
axis to facilitate PD-L1-mediated immune evasion (122-124). Furthermore, PUM1 activates the
mTORC1-HIF-1α axis by inhibiting the mRNA translation of the mTORC1
inhibitor DEPTOR. This not only upregulates glycolytic enzymes such
as HK2 and LDHA but also directly binds to the PD-L1 promoter to
promote its transcription, thereby regulating the GC immune
microenvironment (122).
MSI1/2 regulates the GC immunosuppressive
microenvironment by stabilizing Notch1 and Wnt3a mRNA: It binds to
the 3'-UTR regions of Notch1 and Wnt3a mRNA, prolonging their
half-lives (from 8.7 and 7.9 h to 14.2 and 13.5 h, respectively).
The activated Notch1 pathway promotes Treg cell differentiation
(increasing IL-10 secretion by 2.3-fold), while the Wnt3a pathway
upregulates CCL2 to recruit M2 macrophages. Single-cell sequencing
demonstrated that in MSI1/2-high GC tissues, the proportion of Treg
cells (FOXP3+CD4+) and M2 macrophages
(CD206+CD68+) was significantly higher
compared with that in the low-expression group (Treg cells: 15.7%
vs. 6.2%; M2 macrophages: Not specified vs. 8.3%, respectively;
P<0.001) (125,126). Clinically, patients with high
MSI1/2 expression demonstrated significantly shorter
progression-free survival (4.2 months) following anti-PD-1 therapy
compared with those with low expression (9.5 months; P=0.008)
(127,128).
FMR1 inhibits GC ferroptosis by stabilizing
suppressor of cytokine signaling 2 (SOCS2) mRNA. SOCS2 promotes the
ubiquitin-mediated degradation of the cystine transporter SLC7A11,
thereby reducing glutathione synthesis. FMR1 deficiency is reported
to cause a 40% reduction in SLC7A11 expression, significantly
increasing GC cell susceptibility to ferroptosis (129). A preclinical study demonstrated
that the FMR1 inhibitor Sc1-VHLL, by degrading FMR1, increases
CD8+ T cell infiltration in GC tumor tissue by 2.4-fold
and reduces cell proportions of Tregs by 35%, thereby converting
'cold tumors' to 'hot tumors', which achieved a tumor growth
inhibition rate of 58.7% (P<0.01) (130). In summary, oncogenic RBPs
promote GC progression by regulating the
epithelial-immune-metabolic axis, thereby providing novel targets
for GC prevention and treatment (Fig. 1).
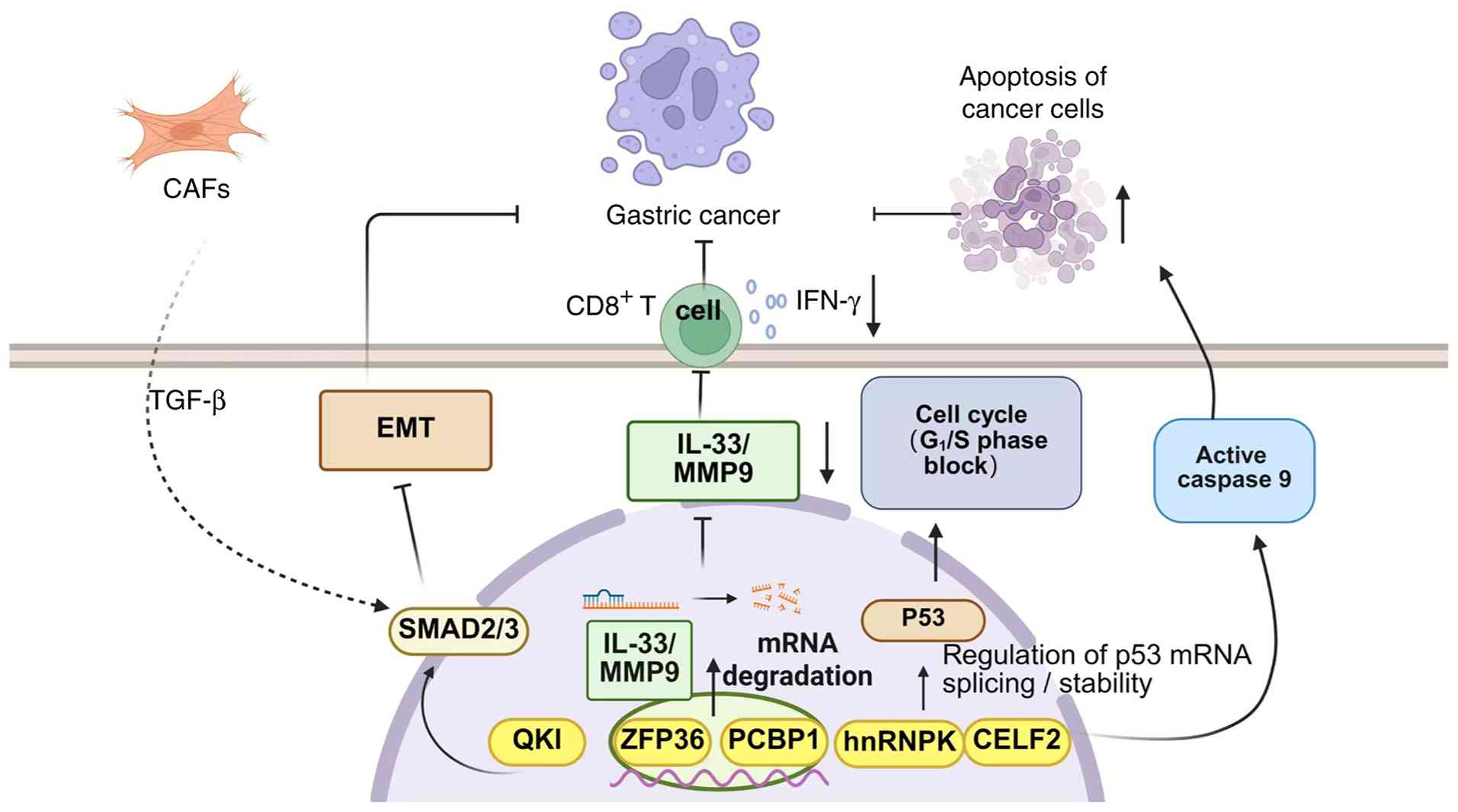
In GC, oncogenic RBPs inhibit tumor progression by
regulating key nodes of RNA metabolism (stability, splicing and
translation) (29,30), which regulate the malignant
phenotype of GC through different mechanisms of action.
PCBP1(αCP1), as a tumor suppressor RBPs, inhibits
the malignant progression of GC through multidimensional
mechanisms. Its functional abnormalities are closely associated
with GC metastasis, drug resistance and prognosis (131,132). C12orf48 depletion also inhibits
metastasis by upregulating PCBP1 to suppress MMP9/CXCR4 regulation
(133). In hypoxic
microenvironments, circPRELID2 binds to PCBP1 and promotes its
O-GlcNAcylation modification, thereby reducing its RNA-binding
activity and releasing its inhibition on GC transfer (134). Regarding chemotherapy
resistance, PCBP1 forms a complex with Siva-1 to enhance BCL-2 mRNA
degradation, thereby increasing cisplatin sensitivity. Inhibition
of Siva-1 leads to its inactivation and subsequent resistance
development, which can be reversed by PCBP1 supplementation
(135,136).
In a clinical cohort of 128 patients with GC
(including 43 with peritoneal metastasis), PCBP1 exhibited a lower
positive rate in metastatic sites (23.3%) than that in primary
tumors (68.5%). As aforementioned, the oncogenic lncRNAs ABL, GLCC1
and GHET1 are closely associated with GC metastasis via interacting
with IGF2BP1, with a median overall survival (14.2 months) shorter
in low-expression patients compared with high-expression patients
(28.6 months; P<0.001). Low expression of PCBP1 was also an
independent prognostic risk factor for peritoneal metastasis
(hazard ratio=2.87; P=0.003) (132). Furthermore, EGCG-lys fibrils
silence circMAP2K2 to upregulate PCBP1 and inhibit GC
proliferation, providing a direction for targeted therapy (137). In summary, PCBP1 exerts its
effects through the pathway of mRNA degradation-protein
modification-drug sensitivity regulation. PCBP1 expression serves
as a potential biomarker for predicting GC metastasis, assessing
prognosis and determining treatment response, thereby providing a
target for precision medicine.
QKI inhibits GC metastasis by regulating the
alternative splicing of key molecules in the TGF-β pathway. QKI
specifically binds to splicing sites on SMAD2/3 precursor mRNA,
promoting the generation of the tumor-suppressor subtypes SMAD2ΔE3
and SMAD3ΔE4 while reducing expression of the pro-metastatic
subtypes SMAD2-FL and SMAD3-FL, thereby blocking TGF-β-mediated
EMT. Analysis of clinical samples demonstrated that QKI expression
levels in GC tissue were negatively correlated with lymph node
metastasis (r=-0.42; P<0.01). Patients with low QKI expression
levels exhibited a significantly higher lymph node metastasis rate
(62.7%) compared with those with high expression (28.3%) (138-141).
TTP inhibits the formation of the GC inflammatory
microenvironment by targeting IL-33 mRNA degradation. As a
pro-cancer cytokine, reduced IL-33 expression diminishes M2
macrophage recruitment. Clinically, patients with high TTP
expression demonstrated significantly longer median overall
survival (31.5 months) compared with those with low expression
(18.2 months; P<0.01), indicating its prognostic value (142). CELF2 promotes the generation of
the pro-apoptotic subtype caspase-9a, by regulating the alternative
splicing of the caspase-9 precursor mRNA, thereby increasing GC
apoptosis by 2.3-fold (P<0.05) (143). In summary, in GC, tumor
suppressor RBPs inhibit tumor progression through differential RNA
metabolic regulatory mechanisms (Fig. 2).
In cancer research, RBPs serve crucial roles in
post-transcriptional regulation. A comparison of their functions in
GC with those in other common tumors, such as lung cancer, BC and
CRC, demonstrated both similarities and differences. In terms of
similarity, the IGF2BP family (such as IGF2BP1/3) stabilizes target
mRNAs through m6A modification across multiple tumors,
thereby promoting malignant behaviors including tumor cell
proliferation and migration (18,27). In GC, BC, HCC and CRC, IGF2BPs
bind to and stabilize the mRNA of key target molecules such as
c-MYC, activating downstream oncogenic pathways to confer a growth
advantage to tumors (40,144-149).
HuR promotes PD-L1 expression in both BC and GC, thereby inducing
tumor metastasis and immune evasion (106,150). LIN28A/B promotes tumorigenesis
in both CRC and GC by inhibiting let-7 miRNA synthesis (25,151).
In terms of differences, IGF2BP1 promotes
proliferation in BC via the estrogen receptor signaling pathway and
influences endocrine therapy sensitivity in cancer cells; this
mechanism has not been reported in GC (152). The function of HuR in BC is
also regulated by estrogen receptor signaling, unlike in GC
(153). Furthermore, YTHDF1
maintains tumor stemness and EMT in GC by enhancing c-MYC
translation and activating the PI3K/AKT/mTOR pathway, whilst
exerting bidirectional regulation on the immune microenvironment
(its deletion enhances antitumor immunity). In BC, YTHDF1
expression correlates with estrogen receptor status and influences
tumor progression by regulating the translation of proliferation-
and invasion-related genes, lacking the immune regulatory function
observed in GC (154-156).
In summary, the specificity of RBPs in GC manifests
in three aspects: i) IGF2BP1/3 maintains an energy advantage by
regulating ferroptosis and glycolysis-oxidation metabolic coupling,
an immune-metabolic crosstalk mechanism rarely observed in other
gastrointestinal tumors; ii) YTHDF1 regulates dendritic cell
recruitment and the MHC II/IL-12 axis, representing an immune
regulatory feature unique to GC; and iii) HuR synergizes with the
gastrin-dependent p38 MAPK-COX-2/IL-8 axis, distinctly differing
from the estrogen receptor-mediated regulatory pathways observed in
BC. These GC-specific mechanisms provide core targets for
developing tumor-specific targeted therapeutic strategies.
RBPs serve a pivotal role in post-transcriptional
regulation of GC and its progression, with their therapeutic and
clinical translational value attracting significant attention. Cui
et al (157)
demonstrated that RBPs and miRNAs jointly constitute the core
mechanism of cancer drug resistance, confirming that targeting
their complexes represents a potential strategy to overcome
treatment failure in GC. Furthermore, the nanomedicine delivery
systems reviewed by Molinaro et al (158) and Li et al (159) enhance targeting through surface
modification and combination with phototherapy, nano-drug delivery
systems, chemotherapy and immunotherapy. Notably, it was recently
reported that differentially expressed circRNAs in GC promote
carcinogenesis by inhibiting the ubiquitination pathway of RBPs
such as G3BP1 (160). This
uncovered circRNA-RBP interaction provides novel directions for
post-transcriptional regulatory mechanisms and therapeutic target
development (160).
Furthermore, gastrointestinal organoid models provide a
physiologically relevant research platform for investigating RBP
function, evaluating RBP-related pathways and detecting drug
responses preclinically (161,162). In summary, integrating
RBP-targeting strategies with nanomedicine, molecular diagnostics
and organoid technologies holds promise for enhancing the precision
of GC treatment, overcoming drug resistance and advancing the
clinical translation of novel RBP-related interventions.
RBPs have GC-specific regulatory features (such as
the metabolic coupling of IGF2BP1/3, DC-MHC II/IL-12 regulation of
YTHDF1 and gastrin pathway synergy of HuR) and clinical utility
(such as PCBP1 for metastasis/prognosis and PUM1 for immunotherapy
prediction). RBPs act as molecular switches for GC development
through a multilevel and multidimensional post-transcriptional
regulatory network. Intervention strategies targeting
cancer-promoting RBPs (such as inhibitor development and
combination therapies) have demonstrated translational potential,
whereas the activation or restoration of cancer-suppressing RBPs
has provided new ideas for reversing drug resistance. Under the
framework of precision medicine, future studies may combine
multi-omics technology and preclinical models to resolve the
spatial and temporal specificity of RBP regulatory networks and
facilitate the development of individualized treatment options. The
study of RBPs in GC not only deepens the understanding of tumor
biology but also provides hope for overcoming existing therapeutic
challenges.
Not applicable.
XL and TL conceived and designed the review, and
supervised the entire research process. SL drafted the initial
manuscript and organized the core literature. BJ contributed to
topic selection, participated in the design of the review
framework, and provided critical insights into the immune-metabolic
crosstalk section. JZ performed the literature review. ZM
constructed figures. SY and ST performed the literature review. BT,
TL and XL revised the manuscript critically for important
intellectual content, including mechanism validation and clinical
significance discussion. Data authentication is not applicable. All
authors read and approved the final version of the manuscript.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
Not applicable.
The present study was supported by the National Natural Science
Foundation of China (grant nos. 82070536, 82470540, 32460215,
82160505, 81660098 and 82073087), Guizhou Province International
Science and Technology Cooperation (Gastroenterology) Base [grant
nos. Qian Ke He Platform Talents-HZJD (2021) 001 and Qian Ke He
basic research-ZK (2023) major project 059], Guizhou Innovative
Talent Team on Ion Channels and Malignant Tumors of Epithelial
Origin [grant no. Qian Ke He Platform Talents-CXTD (2023) 001] and
the Collaborative Innovation Center of the Chinese Ministry of
Education (grant no. 2020-39).
|
1
|
Smyth EC, Nilsson M, Grabsch HI, Van
Grieken NC and Lordick F: Gastric cancer. Lancet. 396:635–648.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Machlowska J, Baj J, Sitarz M, Maciejewski
R and Sitarz R: Gastric cancer: Epidemiology, risk factors,
classification, genomic characteristics and treatment strategies.
Int J Mol Sci. 21:40122020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yang WJ, Zhao HP, Yu Y, Wang JH, Guo L,
Liu JY, Pu J and Lv J: Updates on global epidemiology, risk and
prognostic factors of gastric cancer. World J Gastroenterol.
29:2452–2468. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sinnamon AJ, Savoldy M, Mehta R, Dineen
SP, Peña LR, Lauwers GY and Pimiento JM: Tumor regression grade and
overall survival following gastrectomy with preoperative therapy
for gastric cancer. Ann Surg Oncol. 30:3580–3589. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Guan WL, He Y and Xu RH: Gastric cancer
treatment: Recent progress and future perspectives. J Hematol
Oncol. 16:572023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mehta M, Raguraman R, Ramesh R and Munshi
A: RNA binding proteins (RBPs) and their role in DNA damage and
radiation response in cancer. Adv Drug Deliv Rev. 191:1145692022.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Pereira B, Billaud M and Almeida R:
RNA-binding proteins in cancer: Old players and new actors. Trends
Cancer. 3:506–528. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wurth L: Versatility of RNA-binding
proteins in cancer. Comp Funct Genomics. 2012:1785252012.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Qin H, Ni H, Liu Y, Yuan Y, Xi T, Li X and
Zheng L: RNA-binding proteins in tumor progression. J Hematol
Oncol. 13:902020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Mallam AL, Sae-Lee W, Schaub JM, Tu F,
Battenhouse A, Jang YJ, Kim J, Wallingford JB, Finkelstein IJ,
Marcotte EM and Drew K: Systematic discovery of endogenous human
ribonucleoprotein complexes. Cell Rep. 29:1351–1368.e55. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Mohibi S, Chen X and Zhang J: Cancer
the'RBP' eutics-RNA-binding proteins as therapeutic targets for
cancer. Pharmacol Ther. 203:1073902019. View Article : Google Scholar
|
|
12
|
Hentze MW, Castello A, Schwarzl T and
Preiss T: A brave new world of RNA-binding proteins. Nat Rev Mol
Cell Biol. 19:327–341. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Nishida K, Kuwano Y, Nishikawa T, Masuda K
and Rokutan K: RNA binding proteins and genome integrity. Int J Mol
Sci. 18:13412017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Hudson WH and Ortlund EA: The structure,
function and evolution of proteins that bind DNA and RNA. Nat Rev
Mol Cell Biol. 15:749–760. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Nutter CA and Kuyumcu-Martinez MN:
Emerging roles of RNA-binding proteins in diabetes and their
therapeutic potential in diabetic complications. Wiley Interdiscip
Rev RNA. 9:e14592018. View Article : Google Scholar
|
|
16
|
Gebauer F, Schwarzl T, Valcarcel J and
Hentze MW: RNA-binding proteins in human genetic disease. Nat Rev
Genet. 22:185–198. 2021. View Article : Google Scholar
|
|
17
|
Kechavarzi B and Janga SC: Dissecting the
expression landscape of RNA-binding proteins in human cancers.
Genome Biol. 15:R142014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Korn SM, Ulshofer CJ, Schneider T and
Schlundt A: Structures and target RNA preferences of the
RNA-binding protein family of IGF2BPs: An overview. Structure.
29:787–803. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li W, Deng X and Chen J: RNA-binding
proteins in regulating mRNA stability and translation: Roles and
mechanisms in cancer. Semin Cancer Biol. 86:664–677. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Gerstberger S, Hafner M and Tuschl T: A
census of human RNA-binding proteins. Nat Rev Genet. 15:829–845.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Dai S, Huang Y, Liu T, Xu ZH, Liu T, Chen
L, Wang ZW and Luo F: Development and validation of RNA binding
protein-applied prediction model for gastric cancer. Aging (Albany
NY). 13:5539–5552. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
You Q, Wang F, Du R, Pi J, Wang H, Huo Y,
Liu J, Wang C, Yu J, Yang Y and Zhu L: m6A reader
YTHDF1-targeting engineered small extracellular vesicles for
gastric cancer therapy via epigenetic and immune regulation. Adv
Mater. 35:e22049102023. View Article : Google Scholar
|
|
23
|
Zhao Y, Mir C, Garcia-Mayea Y, Paciucci R,
Kondoh H and Leonart MEL: RNA-binding proteins: Underestimated
contributors in tumorigenesis. Semin Cancer Biol. 86:431–444. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Chen D, Cheung H, Lau HC, Yu J and Wong
CC: N6-methyladenosine RNA-binding protein YTHDF1 in
gastrointestinal cancers: Function, molecular mechanism and
clinical implication. Cancers (Basel). 14:34892022. View Article : Google Scholar
|
|
25
|
Zhang YW, Huang Q, Wu YY, Sun Y and Wei
YL: Progress on the role of LIN28A/B in tumor development and
progression. Yi Chuan. 46:452–465. 2024.PubMed/NCBI
|
|
26
|
Lederer M, Bley N, Schleifer C and
Huttelmaier S: The role of the oncofetal IGF2 mRNA-binding protein
3 (IGF2BP3) in cancer. Semin Cancer Biol. 29:3–12. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhu TY, Hong LL and Ling ZQ: Oncofetal
protein IGF2BPs in human cancer: Functions, mechanisms and
therapeutic potential. Biomark Res. 11:622023. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wu X and Xu L: The RNA-binding protein HuR
in human cancer: A friend or foe? Adv Drug Deliv Rev.
184:1141792022. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wan L, Jia Y, Chen N and Zheng S:
Circ_0003789 knockdown inhibits tumor progression by
miR-429/ZFP36L2 axis in gastric cancer. Biochem Genet.
62:2504–2521. 2024. View Article : Google Scholar
|
|
30
|
Lu XC, Zhou HY, Wu J, Jin Y, Yao XM and Wu
XY: LncRNA LINP1 promotes proliferation and inhibits apoptosis of
gastric cancer cells by repressing RBM5. Eur Rev Med Pharmacol Sci.
24:137–144. 2020.PubMed/NCBI
|
|
31
|
An Y and Duan H: The role of
m6A RNA methylation in cancer metabolism. Mol Cancer.
21:142022. View Article : Google Scholar
|
|
32
|
Maity A and Das B:
N6-methyladenosine modification in mRNA: Machinery,
function and implications for health and diseases. FEBS J.
283:1607–1630. 2016. View Article : Google Scholar
|
|
33
|
Mancarella C and Scotlandi K: IGF2BP3 from
physiology to cancer: Novel discoveries, unsolved issues, and
future perspectives. Front Cell Dev Biol. 7:3632019. View Article : Google Scholar
|
|
34
|
Bell JL, Wachter K, Muhleck B, Pazaitis N,
Köhn M, Lederer M and Hüttelmaier S: Insulin-like growth factor 2
mRNA-binding proteins (IGF2BPs): Post-transcriptional drivers of
cancer progression? Cell Mol Life Sci. 70:2657–2675. 2013.
View Article : Google Scholar :
|
|
35
|
Cao J, Mu Q and Huang H: The roles of
insulin-like growth factor 2 mRNA-binding protein 2 in cancer and
cancer stem cells. Stem Cells Int. 2018:42172592018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wang X, Guan D, Wang D, Liu H, Wu Y, Gong
W, Du M, Chu H, Qian J and Zhang Z: Genetic variants in
m6A regulators are associated with gastric cancer risk.
Arch Toxicol. 95:1081–1088. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Xiao J, Lin L, Luo D, Shi L, Chen W, Fan
H, Li Z, Ma X, Ni P, Yang L and Xu Z: Long noncoding RNA TRPM2-AS
acts as a microRNA sponge of miR-612 to promote gastric cancer
progression and radioresistance. Oncogenesis. 9:292020. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Liu HT, Zou YX, Zhu WJ, Sen-Liu, Zhang GH,
Ma RR, Guo XY and Gao P: lncRNA THAP7-AS1, transcriptionally
activated by SP1 and post-transcriptionally stabilized by
METTL3-mediated m6A modification, exerts oncogenic
properties by improving CUL4B entry into the nucleus. Cell Death
Differ. 29:627–641. 2022. View Article : Google Scholar
|
|
39
|
Yang F, Xue X, Zheng L, Bi J, Zhou Y, Zhi
K, Gu Y and Fang G: Long non-coding RNA GHET1 promotes gastric
carcinoma cell proliferation by increasing c-Myc mRNA stability.
FEBS J. 281:802–813. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wang Q, Chen C, Xu X, Shu C, Cao C, Wang
Z, Fu Y, Xu L, Xu K, Xu J, et al: APAF1-binding long noncoding RNA
promotes tumor growth and multidrug resistance in gastric cancer by
blocking apoptosome assembly. Adv Sci (Weinh). 9:e22018892022.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Yang DL, Dong LF, Qiu YB and Luo GY: An
oncogenic lncRNA, GLCC1, promotes tumorigenesis in gastric
carcinoma by enhancing the c-Myc/IGF2BP1 interaction. Neoplasma.
68:1052–1062. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Jiang T, Xia Y, Li Y, Lu C, Lin J, Shen Y,
Lv J, Xie L, Gu C, Xu Z and Wang L: TRIM29 promotes antitumor
immunity through enhancing IGF2BP1 ubiquitination and subsequent
PD-L1 downregulation in gastric cancer. Cancer Lett.
581:2165102024. View Article : Google Scholar
|
|
43
|
Tang B, Bi L, Xu Y, Cao L and Li X:
N6-methyladenosine (m6A) reader IGF2BP1
accelerates gastric cancer development and immune escape by
targeting PD-L1. Mol Biotechnol. 66:2850–2859. 2024. View Article : Google Scholar
|
|
44
|
Li X, Yang G, Ma L, Tang B and Tao T:
N6-methyladenosine (m6A) writer METTL5
represses the ferroptosis and antitumor immunity of gastric cancer.
Cell Death Discov. 10:4022024. View Article : Google Scholar
|
|
45
|
Xu W, Lai Y, Pan Y, Tan M, Ma Y, Sheng H
and Wang J: m6A RNA methylation-mediated NDUFA4 promotes
cell proliferation and metabolism in gastric cancer. Cell Death
Dis. 13:7152022. View Article : Google Scholar
|
|
46
|
Ding N, Cao G, Wang Z, Xu S and Chen W:
Tumor suppressive Function of IGF2BP1 in gastric cancer through
decreasing MYC. Cancer Sci. 115:427–438. 2024. View Article : Google Scholar :
|
|
47
|
Hämmerle M, Gutschner T, Uckelmann H,
Ozgur S, Fiskin E, Gross M, Skawran B, Geffers R, Longerich T,
Breuhahn K, et al: Post-transcriptional destabilization of the
liver-specific long noncoding RNA HULC by the IGF2 mRNA-binding
protein 1 (IGF2BP1). Hepatology. 58:1703–1712. 2013. View Article : Google Scholar
|
|
48
|
Singh A, Singh V, Wallis N, Abis G,
Oberman F, Wood T, Dhamdhere M, Gershon T, Ramos A and Yisraeli J:
Development of a specific and potent IGF2BP1 inhibitor: A promising
therapeutic agent for IGF2BP1-expressing cancers. Eur J Med Chem.
263:1159402023. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhou Y, Huang T, Siu HL, Wong CC, Dong Y,
Wu F, Zhang B, Wu WK, Cheng AS, Yu J, et al: IGF2BP3 functions as a
potential oncogene and is a crucial target of miR-34a in gastric
carcinogenesis. Mol Cancer. 16:772017. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Ma Q, Yang F, Huang B, Pan X, Li W, Yu T,
Wang X, Ran L, Qian K, Li H, et al: CircARID1A binds to IGF2BP3 in
gastric cancer and promotes cancer proliferation by forming a
circARID1A-IGF2BP3-SLC7A5 RNA-protein ternary complex. J Exp Clin
Cancer Res. 41:2512022. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Yang F, Ma Q, Huang B, Wang X, Pan X, Yu
T, Ran L, Jiang S, Li H, Chen Y, et al: CircNFATC3 promotes the
proliferation of gastric cancer through binding to IGF2BP3 and
restricting its ubiquitination to enhance CCND1 mRNA stability. J
Transl Med. 21:4022023. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Hong Y, Qin H, Li Y, Zhang Y, Zhuang X,
Liu L, Lu K, Li L, Deng X, Liu F, et al: FNDC3B circular RNA
promotes the migration and invasion of gastric cancer cells via the
regulation of E-cadherin and CD44 expression. J Cell Physiol.
234:19895–19910. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Yu T, Ran L, Zhao H, Yin P, Li W, Lin J,
Mao H, Cai D, Ma Q, Pan X, et al: Circular RNA circ-TNPO3
suppresses metastasis of GC by acting as a protein decoy for
IGF2BP3 to regulate the expression of MYC and Snail. Mol Ther
Nucleic Acids. 26:649–664. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Xiao Y, Liu X, Xie K, Luo J, Zhang Y,
Huang X, Luo J and Tan S: Mitochondrial dysfunction induced by
HIF-1α under hypoxia contributes to the development of gastric
mucosal lesions. Clin Transl Med. 14:e16532024. View Article : Google Scholar
|
|
55
|
Wang Q, Chen C, Ding Q, Zhao Y, Wang Z,
Chen J, Jiang Z, Zhang Y, Xu G, Zhang J, et al: METTL3-mediated
m6A modification of HDGF mRNA promotes gastric cancer
progression and has prognostic significance. Gut. 69:1193–1205.
2020. View Article : Google Scholar
|
|
56
|
Huangfu L, Zhu H, Wang G, Chen J, Wang Y,
Fan B, Wang X, Yao Q, Guo T, Han J, et al: The deubiquitinase USP15
drives malignant progression of gastric cancer through glucose
metabolism remodeling. J Exp Clin Cancer Res. 43:2352024.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Xie R, Liu L, Lu X, He C, Yao H and Li G:
N6-methyladenosine modification of OIP5-AS1 promotes
glycolysis, tumorigenesis, and metastasis of gastric cancer by
inhibiting Trim21-mediated hnRNPA1 ubiquitination and degradation.
Gastric Cancer. 27:49–71. 2024. View Article : Google Scholar :
|
|
58
|
Fang L, Huang H, Lv J, Chen Z, Lu C, Jiang
T, Xu P, Li Y, Wang S, Li B, et al: m5C-methylated lncRNA NR_033928
promotes gastric cancer proliferation by stabilizing GLS mRNA to
promote glutamine metabolism reprogramming. Cell Death Dis.
14:5202023. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Ge L, Rui Y, Wang C, Wu Y, Wang H and Wang
J: The RNA m6A reader IGF2BP3 regulates NFAT1/IRF1
axis-mediated anti-tumor activity in gastric cancer. Cell Death
Dis. 15:1922024. View Article : Google Scholar
|
|
60
|
Lin K, Lin X and Luo F: IGF2BP3 boosts
lactate generation to accelerate gastric cancer immune evasion.
Apoptosis. 29:2147–2160. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Sikorski V, Selberg S, Lalowski M,
Karelson M and Kankuri E: The structure and function of YTHDF
epitranscriptomic m6A readers. Trends Pharmacol Sci.
44:335–353. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Zou Z and He C: The YTHDF proteins display
distinct cellular functions on m6A-modified RNA. Trends
Biochem Sci. 49:611–621. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Chen W, He Q, Liu J, Li N, Xiao K and Chen
H: PLAGL2 promotes Snail expression and gastric cancer progression
via UCA1/miR-145-5p/YTHDF1 axis. Carcinogenesis. 44:328–340. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Wang A, Huang H, Shi JH, Yu X, Ding R,
Zhang Y, Han Q, Ni ZY, Li X, Zhao R and Zou Q: USP47 inhibits
m6A-dependent c-Myc translation to maintain regulatory T
cell metabolic and functional homeostasis. J Clin Invest.
133:e1693652023. View Article : Google Scholar
|
|
65
|
Pi J, Wang W, Ji M, Wang X, Wei X, Jin J,
Liu T, Qiang J, Qi Z, Li F, et al: YTHDF1 promotes gastric
carcinogenesis by controlling translation of FZD7. Cancer Res.
81:2651–2665. 2021. View Article : Google Scholar
|
|
66
|
Li Y, Guo X, Liang X and Wang Z: YTHDF1
promotes proliferation and inhibits apoptosis of gastric cancer
cells via upregulating TCF7 mRNA translation. Front Biosci
(Landmark Ed). 29:1172024. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Huo FC, Zhu ZM, Zhu WT, Du QY, Liang J and
Mou J: METTL3-mediated m6A methylation of SPHK2 promotes
gastric cancer progression by targeting KLF2. Oncogene.
40:2968–2981. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Chen XY, Liang R, Yi YC, Fan HN, Chen M,
Zhang J and Zhu JS: The m6A reader YTHDF1 facilitates
the tumorigenesis and metastasis of gastric cancer via USP14
translation in an m6A-dependent manner. Front Cell Dev
Biol. 9:6477022021. View Article : Google Scholar
|
|
69
|
Xu Z, Chen Q, Shu L, Zhang C, Liu W and
Wang P: Expression profiles of m6A RNA methylation
regulators, PD-L1 and immune infiltrates in gastric cancer. Front
Oncol. 12:9703672022. View Article : Google Scholar
|
|
70
|
Jang D, Hwa C, Kim S, Oh J, Shin S, Lee
SJ, Kim J, Lee SE, Yang Y, Kim D, et al: RNA
N6-methyladenosine-binding protein YTHDFs redundantly
attenuate cancer immunity by downregulating IFN-γ signaling in
gastric cancer. Adv Sci (Weinh). 12:e24108062025. View Article : Google Scholar
|
|
71
|
Bai X, Wong CC, Pan Y, Chen H, Liu W, Zhai
J, Kang W, Shi Y, Yamamoto M, Tsukamoto T, et al: Loss of YTHDF1 in
gastric tumors restores sensitivity to antitumor immunity by
recruiting mature dendritic cells. J Immunother Cancer.
10:e0036632022. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Chi F, Cao Y and Chen Y: Analysis and
validation of circRNA-miRNA network in regulating m6A RNA
methylation modulators reveals CircMAP2K4/miR-139-5p/YTHDF1 axis
involving the proliferation of hepatocellular carcinoma. Front
Oncol. 11:5605062021. View Article : Google Scholar
|
|
73
|
Zou Z, Wei J, Chen Y, Kang Y, Shi H, Yang
F, Shi Z, Chen S, Zhou Y, Sepich-Poore C, et al: FMRP
phosphorylation modulates neuronal translation through YTHDF1. Mol
Cell. 83:4304–4317.e8. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Fan X, Han F, Wang H, Shu Z, Qiu B, Zeng
F, Chen H, Wu Z, Lin Y, Lan Z, et al: YTHDF2-mediated
m6A modification of ONECUT2 promotes stemness and
oxaliplatin resistance in gastric cancer through transcriptionally
activating TFPI. Drug Resist Updat. 79:1012002025. View Article : Google Scholar
|
|
75
|
Ren M, Pan H, Zhou X, Yu M and Ji F:
KIAA1429 promotes gastric cancer progression by destabilizing RASD1
mRNA in an m6A-YTHDF2-dependent manner. J Transl Med.
22:5842024. View Article : Google Scholar
|
|
76
|
Fang Y, Wu X, Gu Y, Shi R, Yu T, Pan Y,
Zhang J, Jing X, Ma P and Shu Y: LINC00659 cooperated with ALKBH5
to accelerate gastric cancer progression by stabilising JAK1 mRNA
in an m6A-YTHDF2-dependent manner. Clin Transl Med.
13:e12052023. View Article : Google Scholar
|
|
77
|
Zhang Y, Zhou X, Cheng X, Hong X, Jiang X,
Jing G, Chen K and Li Y: PRKAA1, stabilized by FTO in an
m6A-YTHDF2-dependent manner, promotes cell proliferation
and glycolysis of gastric cancer by regulating the redox balance.
Neoplasma. 69:1338–1348. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Zhou Y, Fan K, Dou N, Li L, Wang J, Chen
J, Li Y and Gao Y: YTHDF2 exerts tumor-suppressor roles in gastric
cancer via up-regulating PPP2CA independently of m6A
modification. Biol Proced Online. 25:62023. View Article : Google Scholar
|
|
79
|
Shen X, Zhao K, Xu L, Cheng G, Zhu J, Gan
L, Wu Y and Zhuang Z: YTHDF2 inhibits gastric cancer cell growth by
regulating FOXC2 signaling pathway. Front Genet. 11:5920422020.
View Article : Google Scholar
|
|
80
|
Wang S, Gao S, Zeng Y, Zhu L, Mo Y, Wong
CC, Bao Y, Su P, Zhai J, Wang L, et al:
N6-methyladenosine reader YTHDF1 promotes ARHGEF2
translation and RhoA signaling in colorectal cancer.
Gastroenterology. 162:1183–1196. 2022. View Article : Google Scholar
|
|
81
|
Kedzierska H and Piekielko-Witkowska A:
Splicing factors of SR and hnRNP families as regulators of
apoptosis in cancer. Cancer Lett. 396:53–65. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Nguyen LH, Robinton DA, Seligson MT, Wu L,
Li L, Rakheja D, Comerford SA, Ramezani S, Sun X, Parikh MS, et al:
Lin28b is sufficient to drive liver cancer and necessary for its
maintenance in murine models. Cancer Cell. 26:248–261. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Yang F, Hu A, Li D, Wang J, Guo Y, Liu Y,
Li H, Chen Y, Wang X, Huang K, et al: Circ-HuR suppresses HuR
expression and gastric cancer progression by inhibiting CNBP
transactivation. Mol Cancer. 18:1582019. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Jiang C, Xu D, Feng H, Ren Z, Li X, Chen
Y, Yu J and Cang S: hnRNPA1 promotes the metastasis and
proliferation of gastric cancer cells through WISP2-guided
Wnt/β-catenin signaling pathway. Discov Oncol. 15:4652024.
View Article : Google Scholar
|
|
85
|
Zhang H, Deng T, Liu R, Ning T, Yang H,
Liu D, Zhang Q, Lin D, Ge S, Bai M, et al: CAF secreted miR-522
suppresses ferroptosis and promotes acquired chemo-resistance in
gastric cancer. Mol Cancer. 19:432020. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Zhang H, Wang M, He Y, Deng T, Liu R, Wang
W, Zhu K, Bai M, Ning T, Yang H, et al: Chemotoxicity-induced
exosomal lncFERO regulates ferroptosis and stemness in gastric
cancer stem cells. Cell Death Dis. 12:11162021. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Duan Y, Yan Y, Fu H, Dong Y, Li K, Ye Z,
Dou Y, Huang B, Kang W, Wei GH, et al: SNHG15-mediated feedback
loop interplays with HNRNPA1/SLC7A11/GPX4 pathway to promote
gastric cancer progression. Cancer Sci. 115:2269–2285. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Fuyuhiro Y, Yashiro M, Noda S, Kashiwagi
S, Matsuoka J, Doi Y, Kato Y, Hasegawa T, Sawada T and Hirakawa K:
Upregulation of cancer-associated myofibroblasts by TGF-β from
scirrhous gastric carcinoma cells. Br J Cancer. 105:996–1001. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Wang Q, Tjokrodirijo RTN, Mei H, van
Veelen PA, Ten Dijke P and Fan C: circTGFBR2(3-6) acts as an
assembly platform for RNA-binding protein IGF2BP3 and TGFBR1 mRNA
to enhance breast cancer cell plasticity. Cell Death Differ. Oct
27–2025. View Article : Google Scholar : Epub ahead of
print.
|
|
90
|
Xia Y, Lv J, Jiang T, Li B, Li Y, He Z,
Xuan Z, Sun G, Wang S, Li Z, et al: CircFAM73A promotes the cancer
stem cell-like properties of gastric cancer through the
miR-490-3p/HMGA2 positive feedback loop and HNRNPK-mediated
beta-catenin stabilization. J Exp Clin Cancer Res. 40:1032021.
View Article : Google Scholar
|
|
91
|
Peng WZ, Liu JX, Li CF, Ma R and Jie JZ:
hnRNPK promotes gastric tumorigenesis through regulating CD44E
alternative splicing. Cancer Cell Int. 19:3352019. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Song S, He X, Wang J, Wang R, Wang L, Zhao
W, Wang Y, Zhang Y, Yu Z, Miao D and Xue Y: ELF3-AS1 contributes to
gastric cancer progression by binding to hnRNPK and induces
thrombocytosis in peripheral blood. Cancer Sci. 112:4553–4569.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Lu Y, Cheng J, Cai W, Zhuo H, Wu G and Cai
J: Inhibition of circRNA circVPS33B reduces warburg effect and
tumor growth through regulating the miR-873-5p/HNRNPK axis in
infiltrative gastric cancer. Onco Targets Ther. 14:3095–3108. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Zheng S, Wang Y, Ni C, Guo R, Qiu X, Chen
J, Wang L, Sun X, Chen M, Liu Y, et al: Helicobacter pylori SlyD
stabilizes TPT1 via hnRNPK and enhances OCT1-mediated CDX2
transcriptional activation to drive gastric intestinal metaplasia.
BMC Med. 23:712025. View Article : Google Scholar
|
|
95
|
Huang H, Han Y, Yang X, Li M, Zhu R, Hu J,
Zhang X, Wei R, Li K and Gao R: HNRNPK inhibits gastric cancer cell
proliferation through p53/p21/CCND1 pathway. Oncotarget.
8:103364–103374. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Heo I, Joo C, Cho J, Ha M, Han J and Kim
VN: Lin28 mediates the terminal uridylation of let-7 precursor
MicroRNA. Mol Cell. 32:276–284. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Hennchen M, Stubbusch J, Abarchan-El
Makhfi I, Kramer M, Deller T, Pierre-Eugene C, Janoueix-Lerosey I,
Delattre O, Ernsberger U, Schulte JB and Rohrer H: Lin28B and let-7
in the control of sympathetic neurogenesis and neuroblastoma
development. J Neurosci. 35:16531–16544. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Shen L, Zeng J, Ma L, Li S, Chen C, Jia J
and Liang X: Helicobacter pylori induces a Novel
NF-kB/LIN28A/let-7a/hTERT axis to promote gastric carcinogenesis.
Mol Cancer Res. 19:74–85. 2021. View Article : Google Scholar
|
|
99
|
Newman MA, Thomson JM and Hammond SM:
Lin-28 interaction with the let-7 precursor loop mediates regulated
microRNA processing. RNA. 14:1539–1549. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Rybak A, Fuchs H, Smirnova L, Brandt C,
Pohl EE, Nitsch R and Wulczyn FG: A feedback loop comprising lin-28
and let-7 controls pre-let-7 maturation during neural stem-cell
commitment. Nat Cell Biol. 10:987–993. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
West RC, McWhorter ES, Ali A, Goetzman LN,
Russ JE, Gonzalez-Berrios CL, Anthony RV, Bouma GJ and Winger QA:
HMGA2 is regulated by LIN28 and BRCA1 in human placental cells.
Biol Reprod. 100:227–238. 2019. View Article : Google Scholar
|
|
102
|
Wang XW, Li Q, Liu CM, Hall PA, Jiang JJ,
Katchis CD, Kang S, Dong BC, Li S and Zhou FQ: Lin28 signaling
supports Mammalian PNS and CNS Axon regeneration. Cell Rep.
24:2540–2552.e6. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Zhu H, Shyh-Chang N, Segre AV, Shinoda G,
Shah SP, Einhorn WS, Takeuchi A, Engreitz JM, Hagan JP, Kharas MG,
et al: The Lin28/let-7 axis regulates glucose metabolism. Cell.
147:81–94. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Shinoda G, Shyh-Chang N, Soysa TY, Zhu H,
Seligson MT, Shah SP, Abo-Sido N, Yabuuchi A, Hagan JP, Gregory RI,
et al: Fetal deficiency of lin28 programs life-long aberrations in
growth and glucose metabolism. Stem Cells. 31:1563–1573. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Zhang Y, Li C, Hu C, Wu Q, Cai Y, Xing S,
Lu H, Wang L, Huang D, Sun L, et al: Lin28 enhances de novo fatty
acid synthesis to promote cancer progression via SREBP-1. EMBO Rep.
20:e481152019. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Xie M, Yu T, Jing X, Ma L, Fan Y, Yang F,
Ma P, Jiang H, Wu X, Shu Y and Xu T: Exosomal circSHKBP1 promotes
gastric cancer progression via regulating the miR-582-3p/HUR/VEGF
axis and suppressing HSP90 degradation. Mol Cancer. 19:1122020.
View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Li Q, Tong D, Guo C, Wu F, Li F, Wang X,
Jiang Q, Wei Y, Liu L, Ni L, et al: MicroRNA-145 suppresses gastric
cancer progression by targeting Hu-antigen R. Am J Physiol Cell
Physiol. 318:C605–C614. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Li R, Wang J, Xie Z, Tian X, Hou J, Wang
D, Qian H, Shen H and Xu W: CircUSP1 as a novel marker promotes
gastric cancer progression via stabilizing HuR to upregulate USP1
and Vimentin. Oncogene. 43:1033–1049. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Liu N, Jiang F, Ye M, Wang B, Ge D and
Chang S: HuR confers IL-17a-induced migration and invasion of
gastric cancer cells via upregulation of Snail translation.
Cytokine. 153:1558302022. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Xie M, Ma T, Xue J, Ma H, Sun M, Zhang Z,
Liu M, Liu Y, Ju S, Wang Z and De W: The long intergenic
non-protein coding RNA 707 promotes proliferation and metastasis of
gastric cancer by interacting with mRNA stabilizing protein HuR.
Cancer Lett. 443:67–79. 2019. View Article : Google Scholar
|
|
111
|
Xu Y, Yu X, Xu J, Lu J, Jiang H, Lou N, Lu
W, Xu J, Ye G, Dong S and Nie F: LncRNA RP11-138J23.1 contributes
to gastric cancer progression by interacting with RNA-binding
protein HuR. Front Oncol. 12:8484062022. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Zou Z, Ma T, He X, Zhou J, Ma H, Xie M,
Liu Y, Lu D, Di S and Zhang Z: Long intergenic non-coding RNA 00324
promotes gastric cancer cell proliferation via binding with HuR and
stabilizing FAM83B expression. Cell Death Dis. 9:7172018.
View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Zhang T, Beeharry MK, Zheng Y, Wang Z, Li
J, Zhu Z and Li C: Long noncoding RNA SNHG12 promotes gastric
cancer proliferation by binding to HuR and stabilizing YWHAZ
expression through the AKT/GSK-3β pathway. Front Oncol.
11:6458322021. View Article : Google Scholar
|
|
114
|
Xu W, Zuo H, Miao X, Zhu J, Tang W, Yuan
Z, Gu M and Gu X: Long non-coding RNA VCAN-AS1 promotes gastric
cancer progression via the HuR/F11R pathway. Am J Transl Res.
16:6489–6499. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Li J, Dong W, Jiang Q, Zhang F and Dong H:
LINC00668 cooperated with HuR dependent upregulation of PKN2 to
facilitate gastric cancer metastasis. Cancer Biol Ther. 22:311–323.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Zhang T, Beeharry MK, Wang Z, Zhu Z, Li J
and Li C: YY1-modulated long non-coding RNA SNHG12 promotes gastric
cancer metastasis by activating the miR-218-5p/YWHAZ axis. Int J
Biol Sci. 17:1629–1643. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Subramaniam D, Ramalingam S, May R,
Dieckgraefe BK, Berg DE, Pothoulakis C, Houchen CW, Wang TC and
Anant S: Gastrin-mediated interleukin-8 and cyclooxygenase-2 gene
expression: Differential transcriptional and post-transcriptional
mechanisms. Gastroenterology. 134:1070–1082. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Liu Y, Li X, Zhang H, Zhang M and Wei Y:
HuR up-regulates cell surface PD-L1 via stabilizing CMTM6
transcript in cancer. Oncogene. 40:2230–2242. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Zhang Q, Yang Z, Hao X, Dandreo LJ, He L,
Zhang Y, Wang F, Wu X and Xu L: Niclosamide improves cancer
immunotherapy by modulating RNA-binding protein HuR-mediated PD-L1
signaling. Cell Biosci. 13:1922023. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Majumder M, Chakraborty P, Mohan S,
Mehrotra S and Palanisamy V: HuR as a molecular target for cancer
therapeutics and immune-related disorders. Adv Drug Deliv Rev.
188:1144422022. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Wang CS, Chang CH, Tzeng TY, Lin AMY and
Lo YL: Gene-editing by CRISPR-Cas9 in combination with
anthracycline therapy via tumor microenvironment-switchable,
EGFR-targeted, and nucleus-directed nanoparticles for head and neck
cancer suppression. Nanoscale Horiz. 6:729–743. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Yin S, Liu H, Zhou Z, Xu X, Wang P, Chen
W, Deng G, Wang H, Yu H, Gu L, et al: PUM1 promotes tumor
progression by activating DEPTOR-Meditated glycolysis in gastric
cancer. Adv Sci (Weinh). 10:e23011902023. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Zhang Y, Xie W, Zheng W, Qian X and Deng
C: Exosomemediated circGMPS facilitates the development of gastric
cancer cells through miR-144-3p/PUM1. Cytotechnology. 76:53–68.
2023. View Article : Google Scholar
|
|
124
|
Wang H, Zhou Z, Zhang J, Hao T, Wang P, Wu
P, Su R, Yang H, Deng G, Chen S, et al: Pumilio1 regulates
NPM3/NPM1 axis to promote PD-L1-mediated immune escape in gastric
cancer. Cancer Lett. 581:2164982023. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Wang T, Ong CW, Shi J, Srivastava S, Yan
B, Cheng CL, Yong WP, Chan SL, Yeoh KG, Iacopetta B and
Salto-Tellez M: Sequential expression of putative stem cell markers
in gastric carcinogenesis. Br J Cancer. 105:658–665. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Zhu Y, Zhou B, Hu X, Ying S, Zhou Q, Xu W,
Feng L, Hou T, Wang X, Zhu L and Jin H: LncRNA LINC00942 promotes
chemoresistance in gastric cancer by suppressing MSI2 degradation
to enhance c-Myc mRNA stability. Clin Transl Med. 12:e7032022.
View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Zhao J, Li X, Sun X, Xiao R, Xue J, Sui K
and Liu Z: Combination of cadonilimab (PD-1/CTLA-4 bispecific
antibody) and apatinib as salvage therapy achieves partial response
in MSI-H advanced gastric cancer: A case report. Front Immunol.
16:15337002025. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Dong Z, Ni B, Yang L, Guan Y, Zhu C, Zhao
E, Zhao G, Xia X and Zhang Z: Efficacy and safety of camrelizumab
in combination with docetaxel + S-1 sequenced by camrelizumab + S-1
for Stage III (PD-1+/MSI-H/EBV+/dMMR) gastric cancer: study
protocol for a single-center, prospective, open-label, single-arm
trial. Front Surg. 9:9173522022. View Article : Google Scholar
|
|
129
|
Wang J, Jia Q, Jiang S, Lu W and Ning H:
POU6F1 promotes ferroptosis by increasing lncRNA-CASC2
transcription to regulate SOCS2/SLC7A11 signaling in gastric
cancer. Cell Biol Toxicol. 40:32024. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Peng R, Huang Q, Wang L, Qiao G, Huang X,
Jiang J and Chu X: G-Quadruplex RNA based PROTAC enables targeted
degradation of RNA binding protein FMRP for tumor immunotherapy.
Angew Chem Int Ed Engl. 63:e2024027152024. View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Zhang ZZ, Shen ZY, Shen YY, Zhao EH, Wang
M, Wang CJ, Cao H and Xu J: HOTAIR long noncoding RNA promotes
gastric cancer metastasis through suppression of Poly r(C)-Binding
protein (PCBP) 1. Mol Cancer Ther. 14:1162–1170. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Ji FJ, Wu YY, An Z, Liu XS, Jiang JN, Chen
FF and Fang XD: Expression of both poly r(C) binding protein 1
(PCBP1) and miRNA-3978 is suppressed in peritoneal gastric cancer
metastasis. Sci Rep. 7:154882017. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Lin L, Li H, Shi D, Liu Z, Wei Y, Wang W,
Wu D, Li B and Guo Q: Depletion of C12orf48 inhibits gastric cancer
growth and metastasis via up-regulating Poly r(C)-Binding Protein
(PCBP) 1. BMC Cancer. 22:1232022. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Zhang P, Luo Z, Xu Y, Zhang Y, Zhang R,
Paerhati N, Zhang S, Cai Q, Qiu Z and Huang C: Hypoxia-Induced
circPRELID2 promotes gastric cancer metastasis by facilitating ZEB2
translation via PCBP1 O-GlcNAcylation. Adv Sci (Weinh).
12:e053962025. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Kong FB, Shi ZY, Huang YL, Chen HH, Deng
QM, Wu K, Zhu Z, Li L, Xu S, Zhong XG, et al: SIVA-1 interaction
with PCBP1 serves as a predictive biomarker for cisplatin
sensitivity in gastric cancer and its inhibitory effect on tumor
growth in vivo. J Cancer. 15:4301–4312. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Kong F, Wu K, Pang L, Huang Y, Li L, Xu J,
Li F, Qing Y, Wang Z, Huang X, et al: Inhibition of
apoptosis-regulatory protein Siva-1 reverses multidrug resistance
in gastric cancer by targeting PCBP1. Oncol Res. 30:277–288. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Dong J, Zheng Z, Zhou M, Wang Y, Chen J,
Cen J, Cao T, Yang T, Xu Y, Shu G, et al: EGCG-LYS Fibrils-Mediated
CircMAP2K2 silencing decreases the proliferation and metastasis
ability of gastric cancer cells in vitro and in vivo. Adv Sci
(Weinh). 10:e23040752023. View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Liu HQ, Shu X, Ma Q, Wang R, Huang MY, Gao
X and Liu YN: Identifying specific miRNAs and associated mRNAs in
CD44 and CD90 cancer stem cell subtypes in gastric cancer cell line
SNU-5. Int J Clin Exp Pathol. 13:1313–1323. 2020.PubMed/NCBI
|
|
139
|
Tang G, Song Q, Dou J, Chen Z, Hu X, Li Z,
Li X, Wang T, Dong S and Zhang H: Neutrophil-centric analysis of
gastric cancer: Prognostic modeling and molecular insights. Cell
Mol Life Sci. 81:4522024. View Article : Google Scholar : PubMed/NCBI
|
|
140
|
Feng H, Jin Z, Liu K, Peng Y, Jiang S,
Wang C, Hu J, Shen X, Qiu W, Cheng X and Zhao R: Identification and
validation of critical alternative splicing events and splicing
factors in gastric cancer progression. J Cell Mol Med.
24:12667–12680. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
141
|
He Z, Duan Z, Chen L, Li B and Zhou Y:
Long non-coding RNA Loc490 inhibits gastric cancer cell
proliferation and metastasis by upregulating RNA-binding protein
quaking. Aging (Albany NY). 12:17681–17693. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
142
|
Deng K, Wang H, Shan T, Chen Y, Zhou H,
Zhao Q and Xia J: Tristetraprolin inhibits gastric cancer
progression through suppression of IL-33. Sci Rep. 6:245052016.
View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Wang J, Liu L, Sun Y, Xue Y, Qu J, Pan S,
Li H, Qu H, Wang J and Zhang J: miR-615-3p promotes proliferation
and migration and inhibits apoptosis through its potential target
CELF2 in gastric cancer. Biomed Pharmacother. 101:406–413. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
144
|
Huang H, Weng H, Sun W, Qin X, Shi H, Wu
H, Zhao BS, Mesquita A, Liu C, Yuan CL, et al: Recognition of RNA
N6-methyladenosine by IGF2BP proteins enhances mRNA
stability and translation. Nat Cell Biol. 20:285–295. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
145
|
Xiong DD, Chen ZD, Li JD, Deng YL, He RQ,
Huang ZG, An SQ, Dang YW and Chen G: Nitidine chloride inhibits the
progression of hepatocellular carcinoma by suppressing IGF2BP3 and
modulates metabolic pathways in an m6A-dependent manner.
Mol Med. 31:472025. View Article : Google Scholar
|
|
146
|
Zhu Q, Zhang C, Qu T, Lu X, He X, Li W,
Yin D, Han L, Guo R and Zhang E: MNX1-AS1 promotes phase separation
of IGF2BP1 to drive c-myc-mediated cell-cycle progression and
proliferation in lung cancer. Cancer Res. 82:4340–4358. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
147
|
Shen Q, Xu Z, Sun G, Wang H and Zhang L:
TFAP4 activates IGF2BP1 and promotes progression of non-small cell
lung cancer by stabilizing TK1 expression through m6A
modification. Mol Cancer Res. 20:1763–1775. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
148
|
Liu X, He H, Zhang F, Hu X, Bi F, Li K, Yu
H, Zhao Y, Teng X and Li J: m6A methylated EphA2 and
VEGFA through IGF2BP2/3 regulation promotes vasculogenic mimicry in
colorectal cancer via PI3K/AKT and ERK1/2 signaling. Cell Death
Dis. 13:4832022. View Article : Google Scholar
|
|
149
|
Gao H, Shi L, Liu J, Zhao Y, Du F, He Y,
Yang X, Song N, Wen J and Zheng G: FOXM1-activated IGF2BP3 promotes
cell malignant phenotypes and M2 macrophage polarization in
hepatocellular carcinoma by inhibiting ferroptosis via stabilizing
RRM2 mRNA in an m6A-dependent manner. Mol Cell Biochem.
480:3051–3066. 2025. View Article : Google Scholar
|
|
150
|
Li J, Dong X, Kong X, Wang Y, Li Y, Tong
Y, Zhao W, Duan W, Li P, Wang Y and Wang C: Circular RNA
hsa_circ_0067842 facilitates tumor metastasis and immune escape in
breast cancer through HuR/CMTM6/PD-L1 axis. Biol Direct. 18:482023.
View Article : Google Scholar : PubMed/NCBI
|
|
151
|
Pretzsch E, Max N, Kirchner T, Engel J,
Werner J, Klauschen F, Angele MK and Neumann J: LIN28 promotes
tumorigenesis in colorectal cancer but is not associated with
metastatic spread. Pathol Res Pract. 228:1536692021. View Article : Google Scholar : PubMed/NCBI
|
|
152
|
Gong EY, Jung D, Woo H, Song J, Choi E, Jo
SG, Eyun SI, Kim S and Park YY: Genomic analysis uncovers that
cold-inducible RNA binding protein is associated with estrogen
receptor in breast cancer. Genes Genomics. 46:899–907. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
153
|
Calaluce R, Gubin MM, Davis JW, Magee JD,
Chen J, Kuwano Y, Gorospe M and Atasoy U: The RNA binding protein
HuR differentially regulates unique subsets of mRNAs in estrogen
receptor negative and estrogen receptor positive breast cancer. BMC
Cancer. 10:1262010. View Article : Google Scholar : PubMed/NCBI
|
|
154
|
Yin H, Zhang X, Yang P, Zhang X, Peng Y,
Li D, Yu Y, Wu Y, Wang Y, Zhang J, et al: RNA m6A
methylation orchestrates cancer growth and metastasis via
macrophage reprogramming. Nat Commun. 12:13942021. View Article : Google Scholar
|
|
155
|
Jing L, Zhou K, Wang Z, Li Y, Fan Y, Liu
T, Shan Z and Lin Y: YTHDF1 shapes 'cold' tumor and inhibits
CD8+ T cells infiltration and function in breast cancer.
Exp Cell Res. 432:1137782023. View Article : Google Scholar
|
|
156
|
Zhuang X, Yin S, Cheng J, Sun W, Fang Z,
Xiang Y, Peng EY, Yao Y, Li Y, He X, et al: METTL14-mediated
m6A modification enhances USP22-ERalpha axis to drive
breast cancer malignancy. Pharmacol Res. 210:1075092024. View Article : Google Scholar
|
|
157
|
Cui S, Peng Q, Ma Q, Xu X, Zhang W, Jiang
X, Tan S, Yang W, Han Y, Oyang L, et al: Crosstalk between
RNA-binding proteins and non-coding RNAs in tumors: Molecular
mechanisms, and clinical significance. Int J Biol Sci.
21:2991–3010. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
158
|
Molinaro M, Skrodzki D and Pan D:
Chemoradiotherapy and nanomedicine: Drug mechanisms and delivery
systems. Wiley Interdiscip Rev Nanomed Nanobiotechnol.
16:e19842024. View Article : Google Scholar : PubMed/NCBI
|
|
159
|
Li Z, Zhang X, Liu C, Wu Y, Wen Y, Zheng
R, Xu C, Tian J, Peng Q, Zheng X, et al: Engineering a nano-drug
delivery system to regulate m6A modification and enhance
immunotherapy in gastric cancer. Acta Biomater. 191:412–427. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
160
|
Wang J, Wang S, Wang Y and Xu L, Wu C,
Zhang X, Liang C, Wan S, Xia Y, Huang X and Xu L: Hsa_circ_0000479
promotes gastric cancer progression by inhibiting BTRC-mediated
ubiquitination of G3BP1. Exp Cell Res. 449:1145852025. View Article : Google Scholar : PubMed/NCBI
|
|
161
|
Banerjee P and Senapati S: Translational
utility of organoid models for biomedical research on
gastrointestinal diseases. Stem Cell Rev Rep. 20:1441–1458. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
162
|
Carvalho MR, Yan LP, Li B, Zhang CH, He
YL, Reis RL and Oliveira JM: Gastrointestinal organs and
organoids-on-a-chip: Advances and translation into the clinics.
Biofabrication. 15:2023. View Article : Google Scholar : PubMed/NCBI
|