Introduction
microRNAs (miRNAs, miRs) are a class of small
non-coding RNAs (18–25 nucleotides in length) that are widely
expressed in the cells of several organisms such as invertebrates,
vertebrates, plants and fungi. miRs typically bind to the 3′
untranslated region (UTR) of an mRNA target and direct the
translational repression and/or degradation of this mRNA. miRNAs
are involved in the control of many physiological processes, such
as embryonic development, cell differentiation, proliferation and
apoptosis. Additionally, miRs have been associated with several
pathological conditions, including cancer, neurodegenerative
diseases and autoimmune diseases (1–4).
To date, there are 2,042 annotated human mature miRs
in the official registry (miRBase, http://microrna.sanger.ac.uk/sequences/) (5). Several authors suggest that
additional miRs remain to be identified. Thus, the complete
annotation of miRs is still an ongoing process. Many of the known
miRs were discovered by examining miRNA libraries, whereby the
cDNAs obtained following the size selection of 18–24 nucleotide
RNAs were cloned and then sequenced (6,7).
This method provided direct evidence for the existence and
expression of miRs. miR libraries are also suitable for the
determination of miR levels, which correlate with the cloning
frequency. A limitation in the discovery of miRs by cloning is that
it is difficult to find miRs that are expressed at a low level or
only under specific conditions. This limitation could be overcome
by deep sequencing, which allows for the detection of even poorly
expressed miRs in the transcriptome as well as those not conserved
across species (8). Further tools
that aid in the search for new miRs are based on the bioinformatic
prediction of miR sequences in the genomes (9–11).
Briefly, these tools consider the conservation of the miR sequences
across species and the intrinsic characteristics of their predicted
hairpin structures such as the presence of a fold-back secondary
structure and the thermodynamic stability of the hairpins; however,
such computational methods do not predict whether a given miR is
expressed in certain tissues or cells. Therefore, predicted miRs
have to be experimentally validated (12).
Human hepatocellular carcinoma (HCC) is one of the
cancers characterized by an extremely unfavorable prognosis and is
the third leading cause of cancer-related death worldwide (13). In the present study, to gather
novel information concerning miRs in HCC and with the aim to clone
new miRs, we developed a small RNA expression library in the HCC
cell line HA22T/VGH. We analyzed the expression profile of the miRs
most frequently cloned (miR-24, miR-27a and miR-21) in the tumor
and peritumoral tissues from biopsy specimens of patients
presenting with HCC.
Materials and methods
Cell cultures
Undifferentiated HCC-derived cells, HA22T/VGH, were
maintained in RPMI-1640 (Invitrogen, Carlsbad, CA, USA)
supplemented with 10% fetal bovine serum at 37°C in a 5%
CO2 incubator. These cells were kindly provided by
Professor N. D’Alessandro (University of Palermo, Italy).
SKHep1Clone3 (SKHep1C3) (14),
selected from human HCC-derived cells (SKHep1, ATCC HTB-52) were
maintained in Earle’s MEM (Invitrogen) supplemented with 10% fetal
bovine serum (Invitrogen) at 37°C in a 5% CO2
incubator.
Tissues and clinicopathological features
of HCC
All of the human HCC samples (n=41) as well as the
corresponding peritumoral (PT) non-tumor samples (resected 1–2 cm
from the malignant tumor) were obtained from HCC patients for
pathological examination. Each biopsy specimen was obtained
following patient informed consent under standard conditions of
sampling and processing as previously described (15). Each specimen was determined to be
HCC or PT by pathological examination. In this study, 41 HCC
subjects underwent surgical resection. The subjects consisted of 28
men and 13 women (40 Italian and 1 Chinese) ranging from 38 to 82
years of age (mean age, 67.9±8.9 years). The subjects did not have
any apparent distant metastases, and none had been previously
treated for HCC. The PT tissues revealed the presence of different
background diseases (25 cirrhosis, 15 hepatitis, 1 steatosis), and
the patients were analyzed for the presence of the hepatitis B
(HBV) and hepatitis C (HCV) viruses. Fifteen patients were positive
for HCV, 10 were positive for HBV, 4 were positive for both HBV and
HCV and 7 were found to be negative for both HBV and HCV; for 5
patients no information was available (Table I).
 | Table I.Clinical and pathological
characteristics of the studied population. |
Table I.
Clinical and pathological
characteristics of the studied population.
| Case | Gender | Age | Grade | TNM | Background
disease | HBV | HCV |
|---|
| 137 | M | 69 | G3 | T3bN0M0 | Active
cirrhosis | NA | NA |
| 139 | M | 65 | G2 | T2N0M0 | Active
cirrhosis | + | + |
| 140 | M | 69 | G2 | T1N0M0 | Aspecific reactive
hepatitis | − | + |
| 145 | M | 65 | G2 | T1N0M0 | Steatotic hepatitis
with portal and periportal fibrosis | NA | NA |
| 185 | M | 66 | G1 | T1N0M0 | Mildly active
chronic hepatitis with steatosis of moderate level | + | − |
| 188 | M | 73 | G1 | T3bN0M0 | Cirrhosis with
active chronic hepatitis | + | + |
| 191 | F | 63 | G1 | T1N0M0 | Cirrhosis with
active chronic hepatitis | − | + |
| 197 | M | 70 | G2 | T1N0M0 | Cirrhosis with
microvesicular and macrovesicular steatosis | − | − |
| 205 | M | 73 | G2 | T1N0M0 | Active chronic
hepatitis | − | + |
| 211 | M | 51 | G2 | T1N0M0 | Cirrhosis with
active chronic hepatitis with foci of macrovesicular steatosis and
presence of iperplastic and regenerative macronodules | + | + |
| 218 | M | 64 | G2 | T2N0M0 | Cirrhosis with
active chronic hepatitis | + | − |
| 219 | M | 57 | G1 | T1N0M0 | Cirrhosis with
active chronic hepatitis | + | − |
| 224 | M | 55 | G3 | T3bN0M0 | Cirrhosis with
active chronic hepatitis | + | + |
| 225 | M | 49 | G3 | T3bN0M0 | Microvesicular
steatosis; focal lipofuscinosis; cholestasis | − | − |
| 227 | F | 72 | G2/G3 | T1N0M0 | Cirrhosis with
active chronic hepatitis | − | + |
| 228 | M | 59 | G2 | T1N0M0 | Active chronic
hepatitis of severe level with necrosis and bridging porto-portal
fibrosis (HBsAG) | + | − |
| 229 | F | 79 | G2/G3 | T3bN0M0 | Cirrhosis with
active chronic hepatitis | NA | NA |
| 235 | F | 82 | G3 | T2N0M0 | Cirrhosis with
active chronic hepatitis | − | + |
| 236 | F | 76 | G1 | T1N0M0 | Cirrhosis with
active chronic hepatitis | − | + |
| 237 | M | 68 | G2/G3 | T1N0M0 | Mildly active
chronic hepatitis | − | + |
| 240 | M | 71 | G3 | T3bN0M0 | Active chronic
hepatitis with necrosis and fibrosis ponte-portale | + | − |
| 242 | F | 63 | G2 | T2N0M0 | Active chronic
hepatitis with focal and bridging porto-portal fibrosis | − | + |
| 241 | F | 38 | G2 | T3N0M0 | Reactive
hepatitis | + | − |
| 257 | M | 69 | G1/G2 | T1N0M0 | Cirrhosis and
hemochromatosis | − | − |
| 268 | F | 68 | G1 | T1N0M0 | Cirrhosis with
active chronic hepatitis | + | − |
| 271 | F | 71 | G2/G3 | T1N0M0 | Cirrhosis with
active chronic hepatitis | − | + |
| 272 | M | 65 | G1 | T2N0M0 | Cirrhosis with
active chronic hepatitis and macrovesicular and microvesicular
steatosis (30% of parenchyma) | − | − |
| 273 | M | 73 | G2 | T1N0M0 | Cirrhosis with
active chronic hepatitis and mild macrovesicular and microvesicular
steatosis | − | + |
| 274 | F | 81 | G2 | T1N0M0 | Mildly active
chronic hepatitis with microvesicular and macrovesicular steatosis
(30% of parenchyma) | NA | NA |
| 276 | M | 72 | G2 | T1N0M0 | Cirrhosis with
active chronic hepatitis | − | + |
| 277 | F | 75 | G2 | T2N0M0 | Mildly active
chronic hepatitis | − | − |
| 280 | F | 74 | G2/G3 | T1N0M0 | Mildly active
chronic hepatitis | − | + |
| 281 | M | 74 | G2/G3 | T3N0M0 | Cirrhosis | − | − |
| 283 | M | 78 | G2 | T1N0M0 | Mildly active
chronic hepatitis | + | − |
| 284 | M | 76 | G2 | T1N0M0 | Active chronic
hepatitis | + | − |
| 285 | M | 77 | G2 | T2N0M0 | Active chronic
hepatitis with necrosis ponte-portal of moderate/severe level | − | + |
| 286 | M | 69 | G3 | T4N0M0 | Active
cirrhosis | − | + |
| 287 | M | 63 | G2 | T2N0M0 | Active
cirrhosis | NA | NA |
| 288 | F | 64 | G2 | T1N0M0 | Active
cirrhosis | − | − |
| 289 | M | 75 | G2 | T1N0M0 | Cirrhosis with
iperplastic-displastic macronodules | − | + |
| 290 | M | 65 | G2/G3 | T1N0M0 | Active
cirrhosis | + | − |
Small RNA isolation
Total RNA was extracted from HA22T/VGH cells using
the miRNeasy Mini kit (Qiagen) and the small RNAs (<200
nucleotides) were further separated using the RNeasy MinElute
Cleanup kit (Qiagen, Gaithersburg, MD, USA) according to the
manufacturer’s instructions. Six full-confluence 10-cm plates of
HA22T/VGH cells were used to extract the small RNAs.
Construction and screening of a cDNA
library of small RNAs
Small RNAs were polyadenylated at 37°C for 60 min in
a 100-μl reaction volume with 1.5 μg of RNA and 8 U
of poly(A) polymerase (Ambion, Austin, TX, USA). Poly(A)-tailed
small RNAs were recovered using the miRNeasy Mini kit. A 5′ adapter
(5′-CGA CUG GAG CAC GAG GAC ACU GAC AUG GAC UGA AGG AGU AGA AA-3′)
was ligated to the poly(A)-tailed RNAs using T4 RNA ligase
(Promega) in a 40-μl reaction volume at 37°C for 30 min. The
ligation products were recovered by the miRNeasy Mini kit. Reverse
transcription was performed using 1.5 μg RNA and 55 pmol of
the RT primer (5′-GTA CAG CCG GCG GAG CCG GAG ATC TTA -d(T)30 (A,
G, or C) (A, G, C, or T)-3′ with 200 U of SuperScript III
reverse-transcriptase (Invitrogen). Poly(A)-tailed small RNAs (20
μl total volume) were incubated with 0.55 μl of the
RT primer and 5 μl of a dNTP mix (2 mM) at 65°C for 5 min to
remove any RNA secondary structure. The reactions were chilled on
ice for at least 1 min and the remaining reagents [5X buffer,
dithiothreitol (DTT), RNaseout, SuperScript III] were added as
specified in the SuperScript III protocol. The reaction was allowed
to proceed for 60 min at 55°C. Finally, the reverse transcriptase
was inactivated by a 15-min incubation at 70°C. The cDNA
amplification was carried out for 30 cycles at a final annealing
temperature of 50°C using the forward primer 5′-GGA CAC TGA CAT GGA
CTG AAG GAG TA-3′ and the reverse primer 5′-ATT CTA GAG GCC GAG GCG
GCC GAC ATG T-3′. The PCR was performed using GoTaq DNA polymerase
(Promega, Madison, WI, USA). The PCR product was separated on a 2%
agarose gel with ethidium bromide staining and gel slices
containing ∼109 nucleotide-long DNA were excised. The DNA was
purified using the Wizard SV® Gel and PCR Clean-Up
System kit (Promega). The DNA fragment was directly subcloned into
the pGEM-T vector (Promega) which was subsequently used to
transform JM109 competent cells (Promega). Colony PCR was performed
using primers for the T7 promoter (5′-TAA TAC GAC TCA CTA TAG
GG-3′) and SP6 promoter (5′-ATT TAG GTG ACA CTA TAG AA-3′) and the
clones resulting in PCR products of ∼271 bp in length were
sequenced using the ABI PRISM 310 Genetic Analyser (Applied
Biosystems, Foster City, CA, USA).
Real-time quantification of mature
miR-24, miR-27a and miR-21 by stem-loop RT-PCR
Total RNA from tissue samples was isolated using the
TRIzol reagent (Invitrogen) according to the manufacturer’s
instructions. For a quantitative analysis of mature miRs, a
two-step Taq-Man real-time PCR analysis was performed using primers
and probes obtained from Applied Biosystems. cDNA was synthesized
from total RNA (50 ng) in 15-μl reactions using reverse
transcriptase and the stem-loop primers for miR-24 (Applied
Biosystems; assay ID 000402), miR-27a (assay ID 000408), miR-21
(assay ID 000397), or RNU66 (internal control; assay ID 001002)
provided with the TaqMan MicroRNA Reverse Transcription kit
(Applied Biosystems). The reverse transcriptase reaction was
performed by incubating the samples at 16°C for 30 min, 42°C for 30
min, and 85°C for 5 min. Each PCR reaction (20 μl) contained
1.3 μl of reverse transcriptase product, 10 μl of
Taq-Man 2X Universal PCR Master Mix, and 1 μl of the
appropriate TaqMan MicroRNA Assay solution containing primers and
probes for each miR of interest. The PCR mixtures were incubated at
95°C for 10 min followed by 40 cycles of 95°C for 15 sec and 60°C
for 60 sec. PCRs were performed in triplicate using a 7500
Real-Time PCR system. The expression level calculations for the 3
miRNAs were based on the ΔΔCT method, using RNU66 as an
internal control. For each case the ratio between the relative
levels in HCC and those in PT was assessed. The level of expression
of the miRNAs was considered to be decreased for a value <0.7
and increased for a value >1.3. A value between 0.7 and 1.3 was
defined as having no change in expression level.
Bioinformatic analyses
The cloned RNA sequences were compared to the
sequences in miRBase (http://www.mirbase.org/). Sequences not found in the
miRNA database were subjected to BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi) or BLAT
(http://genome.ucsc.edu/) analyses against the
human genome (http://www.ncbi.nlm.nih.gov/blast). The mFold Web
server (http://mfold.rna.albany.edu/?q=mfold) was used to
evaluate the ability of the candidate miRNA sequence to form
thermodynamically stable hairpin structures. The tool Clustal W2
(http://www.ebi.ac.uk/Tools/msa/clustalw2/) was used to
evaluate miRNA candidate sequence conservation between different
species.
Validation of the miRNA candidate
sequence by northern blotting
The total RNA samples (7.5 μg each) and the
miR markers (New England Biolabs, UK) (17, 21 and 25 nucleotides in
length) were electrophoresed on 15% TBE/urea precast gels
(Invitrogen) and transferred to a Nylon+ membrane
(Invitrogen). The hybridi zation buffer, wash buffer, blocking
buffer and detection buffer were supplied by Signosis (Sunnyvale,
CA, USA). Membranes were hybridized (42°C for 16 h) with
5′-biotin-labelled mercury LNA detection probes (Exiqon, Vedbaek,
Denmark) corresponding to the complementary sequences of the mature
miRNA candidate (hsa-miR-1199 probe: 5′-CTG CGC GGC CCG GGC TCA
GG-3′, hsa-miR-1199* probe: 5′-GTT GAG CAC CGG CCG CAC
GC-3′). As a control, the blots were hybridized with an
oligonucleotide probe complementary to the U6 RNA (U6 probe: 5′-CAC
GAA TTT GCG TGT CAT CCT T-3′). The blots were incubated with
streptavidin-coupled horseradish peroxidase (Signosis) and signals
were detected by enhanced chemiluminescence (Thermo Scientific,
Rockford, IL, USA). The results were visualized on X-ray film
(Thermo Scientific).
Statistical analysis
The histograms represent the mean values, and the
bars indicate standard errors of the mean. The statistical
significance of the results was determined using the Student’s
t-test. The linear correlation between miR expression and overall
survival was determined by the Pearson correlation test. The data
were considered significant at p<0.05. The statistical analysis
was performed with KyPlot (v.2.0b15, http://www.woundedmoon.org/win32/kyplot.html).
Results
miRNAs identified by a small RNA
expression library in HA22T/VGH cells
To uncover new miRNAs and to study global miR
expression in HCC, a small RNA library was generated in the HCC
cell line HA22T/VGH. A total of 200 bacterial clones were sequenced
and 118 clones corresponded to 31 known miRNAs (Tables II and III). The most frequently isolated miRNAs
were miR-24, miR-27a and miR-21 (Table
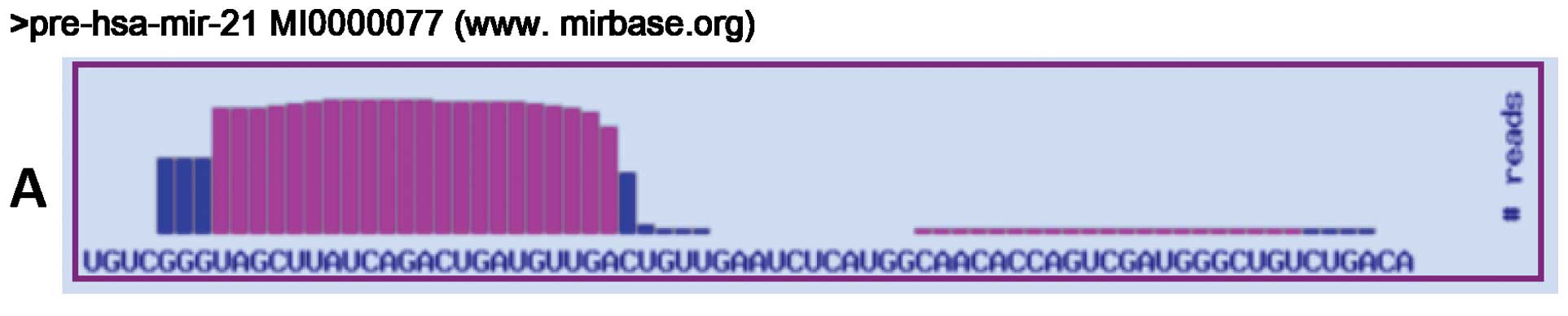
III). For miR-21, we found sequence differences located at the
3′ end of the mature miR. The most frequent variations resulted
from C→U (5/51; 9.8%) and A→I editing (4/51; 7.8%) (Fig. 1). Three clones corresponded to the
precursor stem-loop of the hsa-miR-1308, and 1 clone was partially
homologous to the mmu-miR-1199 precursor stem-loop. Twenty-five
clones corresponded to rRNAs, tRNA, snRNAs, mitochondrial RNAs and
mRNA fragments, 31 clones were empty and 19 clones could not be
sequenced.
 | Table II.Composition of the small RNA
population cloned in the HA22T/VGH library. |
Table II.
Composition of the small RNA
population cloned in the HA22T/VGH library.
| RNA class | No. |
|---|
| miRNA | 118 |
| miRNA
stem-loopa | 4 |
| Partially
homologous miRNAsb | 3 |
| mRNA | 10 |
| rRNA | 8 |
| snRNA | 2 |
| Y RNA | 1 |
| Match with
genomec | 3 |
| Mitochondrial | 1 |
| Empty vector | 31 |
| Uncorrected
sequencing | 19 |
| Total | 200 |
 | Table III.List of the known miRNAs identified
in the library. |
Table III.
List of the known miRNAs identified
in the library.
| miRNA | Frequency | Genomic locus
(Homo sapiens) |
|---|
| hsa-miR-let7a | 1 | 9q22.31 |
| | 11q24.1 |
| | 22q13.31 |
| hsa-miR-let7i | 2 | 12q14.1 |
| hsa-miR-10a | 1 | 17q21.31 |
| hsa-miR-17 | 1 | 13q31.3 |
| hsa-miR-19b | 1 | 13q31.3 |
| | Xq26.2 |
| hsa-miR-20a | 2 | 13q31.3 |
| hsa-miR-21 | 51 | 17q22 |
| hsa-miR-22 | 2 | 17p13.3 |
| hsa-miR-23a | 2 | 19p13.2 |
| hsa-miR-24 | 7 | 9q22.32 |
| | 19p13.2 |
| hsa-miR-25 | 1 | 7q22.1 |
| hsa-miR-26a | 3 | 3p22.2 |
| | 12q13.2 |
| hsa-miR-26b | 2 | 2q35 |
| hsa-miR-27a | 17 | 19p13.2 |
| hsa-miR-27b | 1 | 9q22.32 |
| hsa-miR-29b | 1 | 7q32.2 |
| | 1q32.1 |
| hsa-miR-30b | 3 | 8q24.22 |
| hsa-miR-30c | 1 | 1p34.2 |
| | 6q13 |
| hsa-miR-30e | 1 | 1p34.2 |
| hsa-miR-34b | 1 | 11q23.1 |
|
hsa-miR-92a/ba | 1 | 92a: 13q31.3;
Xq26.2 |
| | 92b: 1q22 |
| hsa-miR-92a | 3 | 13q31.3 |
| | Xq26.2 |
| hsa-miR-93 | 1 | 7q22.1 |
| has-miR-99a | 1 | 21q21.1 |
| hsa-miR-181a | 2 | 1q31.3 |
| | 9q33.3 |
| hsa-miR-224 | 1 | Xq28 |
| hsa-miR-324-5p | 1 | 17p13.1 |
| hsa-miR-365 | 3 | 16p13.12 |
| | 17q11.2 |
| hsa-miR-424 | 2 | Xq26.2 |
| hsa-miR-1308 | 1 | Xp22.11 |
|
hcmv-mir-US25-2 | 1 | Genoma CMV |
miR-24, miR-27a and miR-21 differential
expression in HCC tissues from human biopsy specimens
The expression levels of the most frequently cloned
miRNAs, miR-24, miR-27a and miR-21, were evaluated using real-time
PCR in the tumor and corresponding PT tissues from the biopsy
specimens of 41 HCC patients. The ratio (R) was determined between
the miR expression level (RQHCC) in the HCC sample
versus the expression level (RQPT) detected in the PT.
We assumed miR upregulation when R>1.3, miR downregulation when
R<0.7 and no variation when 0.7≤R≤1.3.
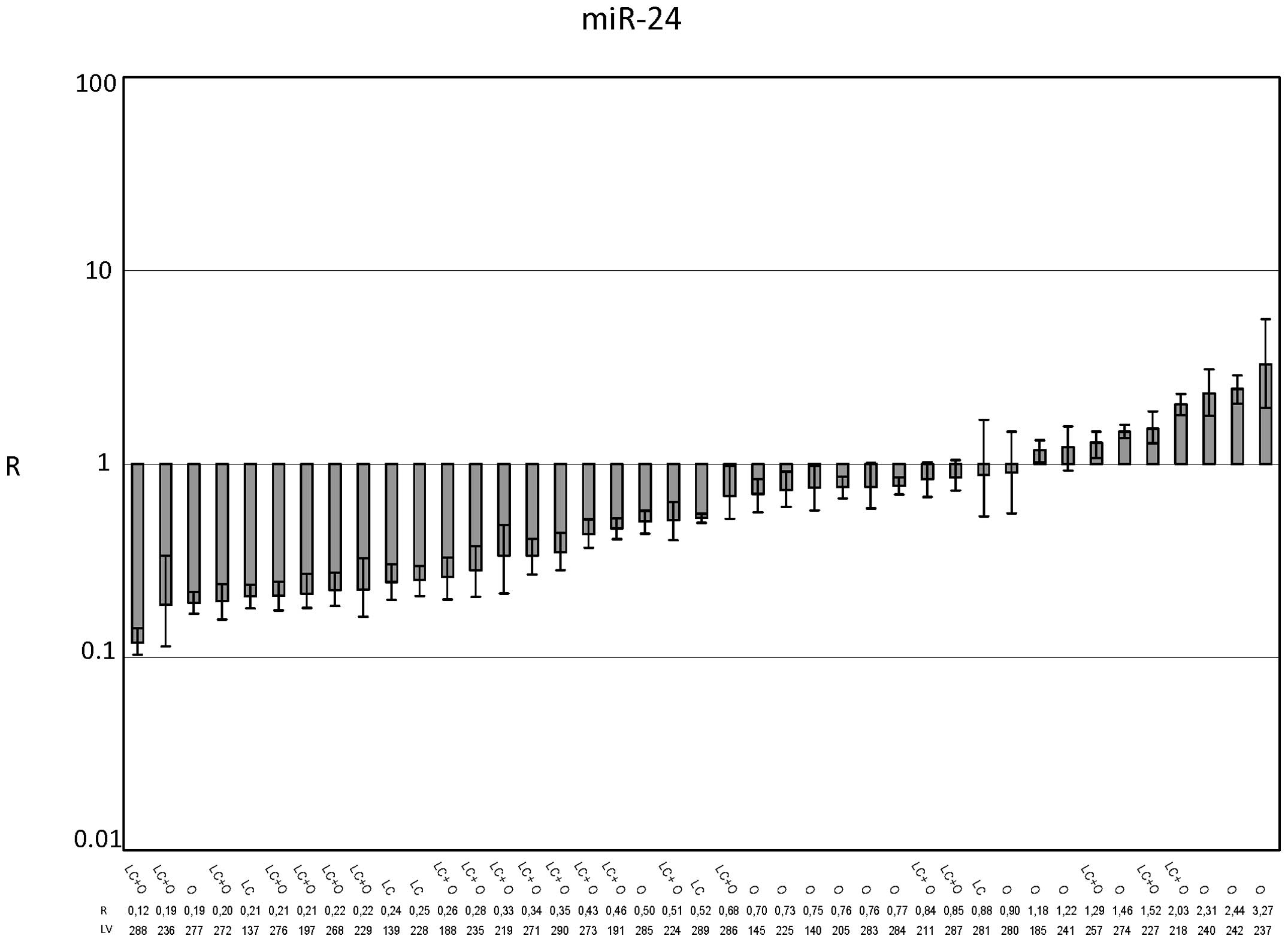
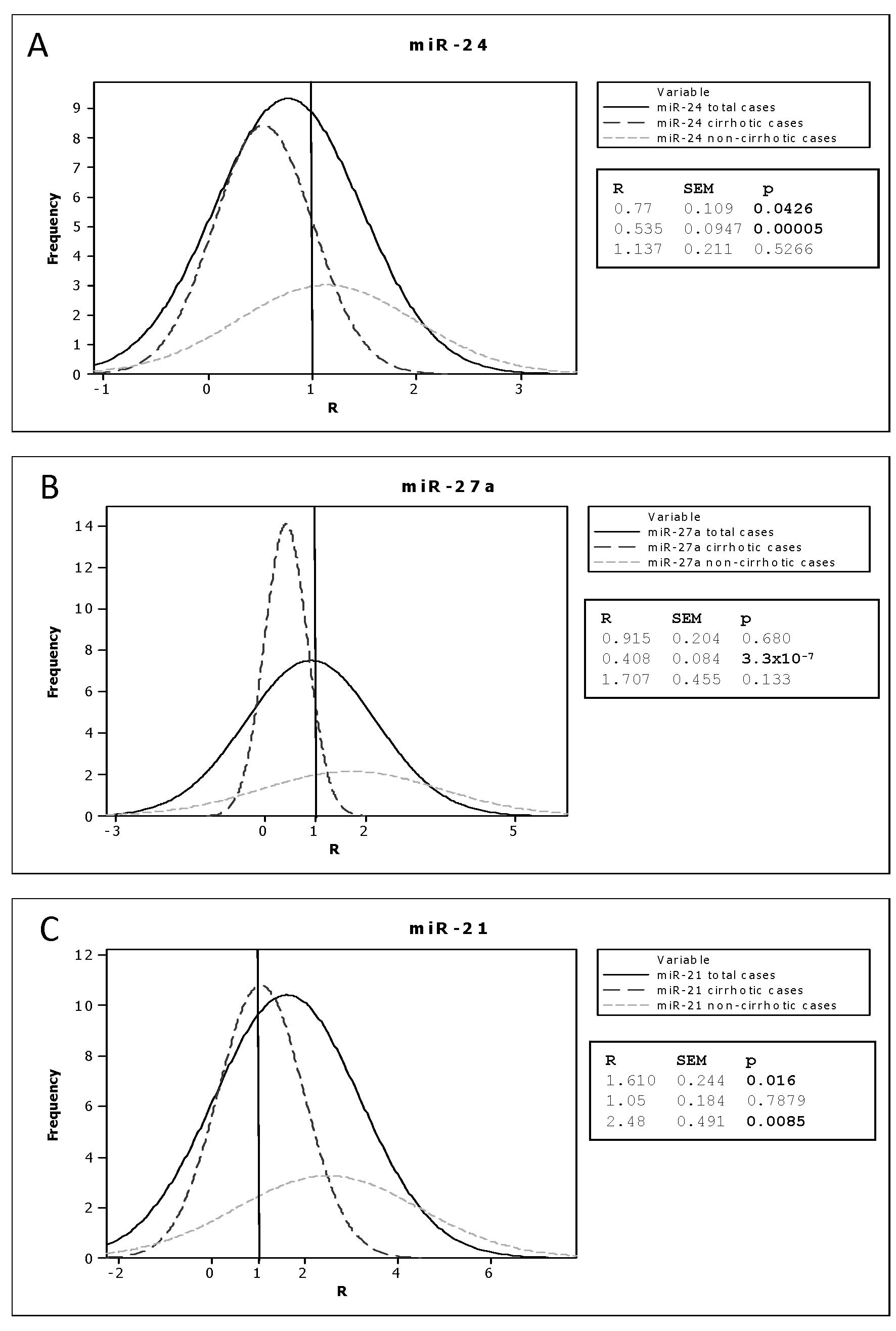
miR-24 (Fig. 2) was
not dysregulated, based on the average R value for the 41 examined
cases (Fig. 5, R=0.77±0.109). The
stratification of the samples on the basis of the presence (25/41)
or absence (16/41) of cirrhosis as the background liver disease
changed the R values. In particular, the R value in the cirrhosis
subgroup was 0.535±0.0947 (p<0.0001), suggesting that miR-24 was
downregulated in HCC with respect to cirrhotic PT tissue. The R
value in the non-cirrhotic liver subgroup was 1.137±0.211,
suggesting no variation in the expression level (Fig. 5).
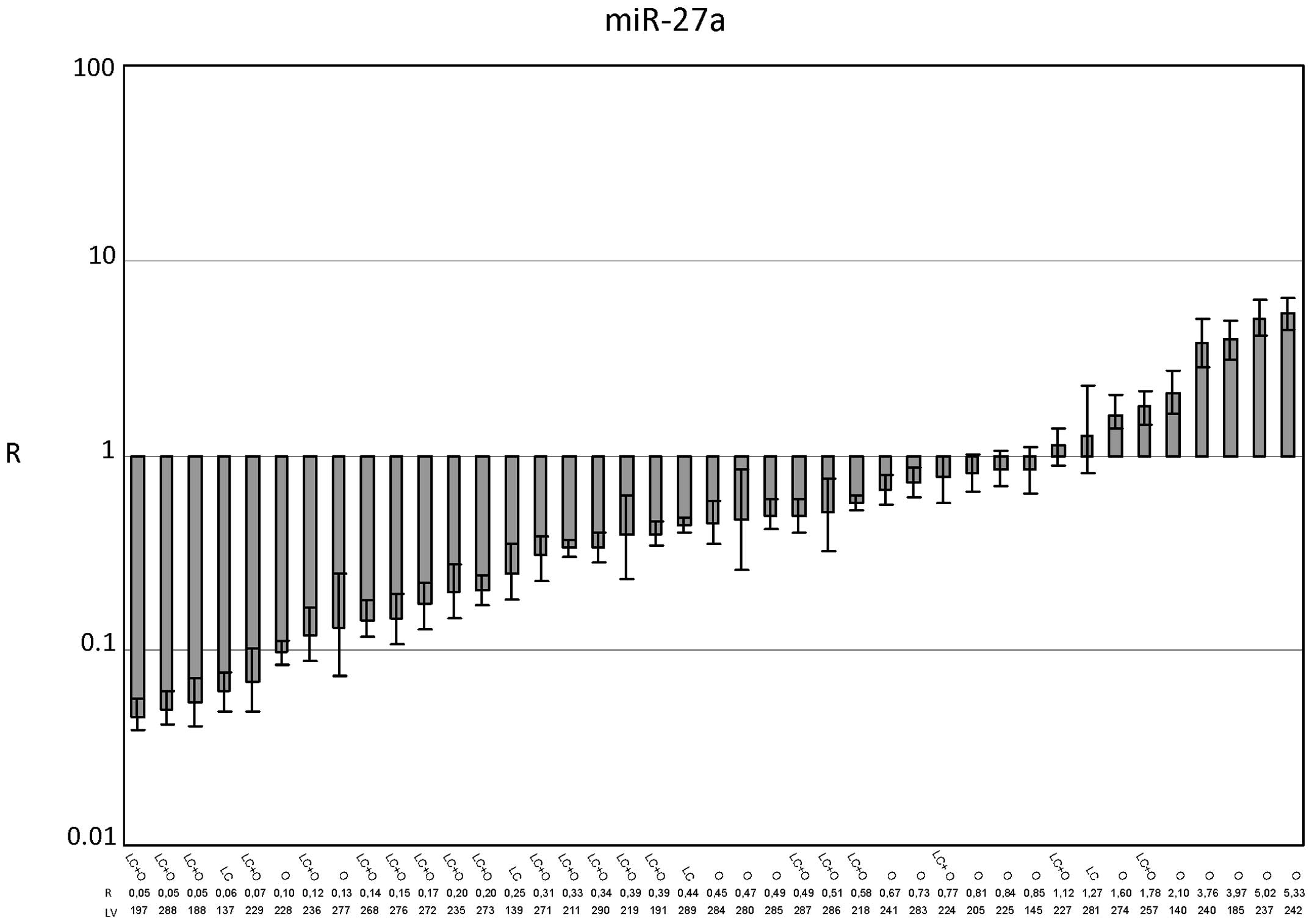
Similar to miR-24, miR-27a (Figs. 3 and 5) did not show dysregulation among the 41
cases, as the average R value was 0.915±0.204. In the cirrhosis
subgroup, the R value of 0.408±0.084 (p<0.0001) underlines the
downregulation of miR-27a in HCC with respect to cirrhotic PT
tissues. In the noncirrhotic subgroup, the R value was 1.707±0.455,
indicating an upregulation of miR-27a expression. This result,
however, was not statistically significant (Fig. 5).
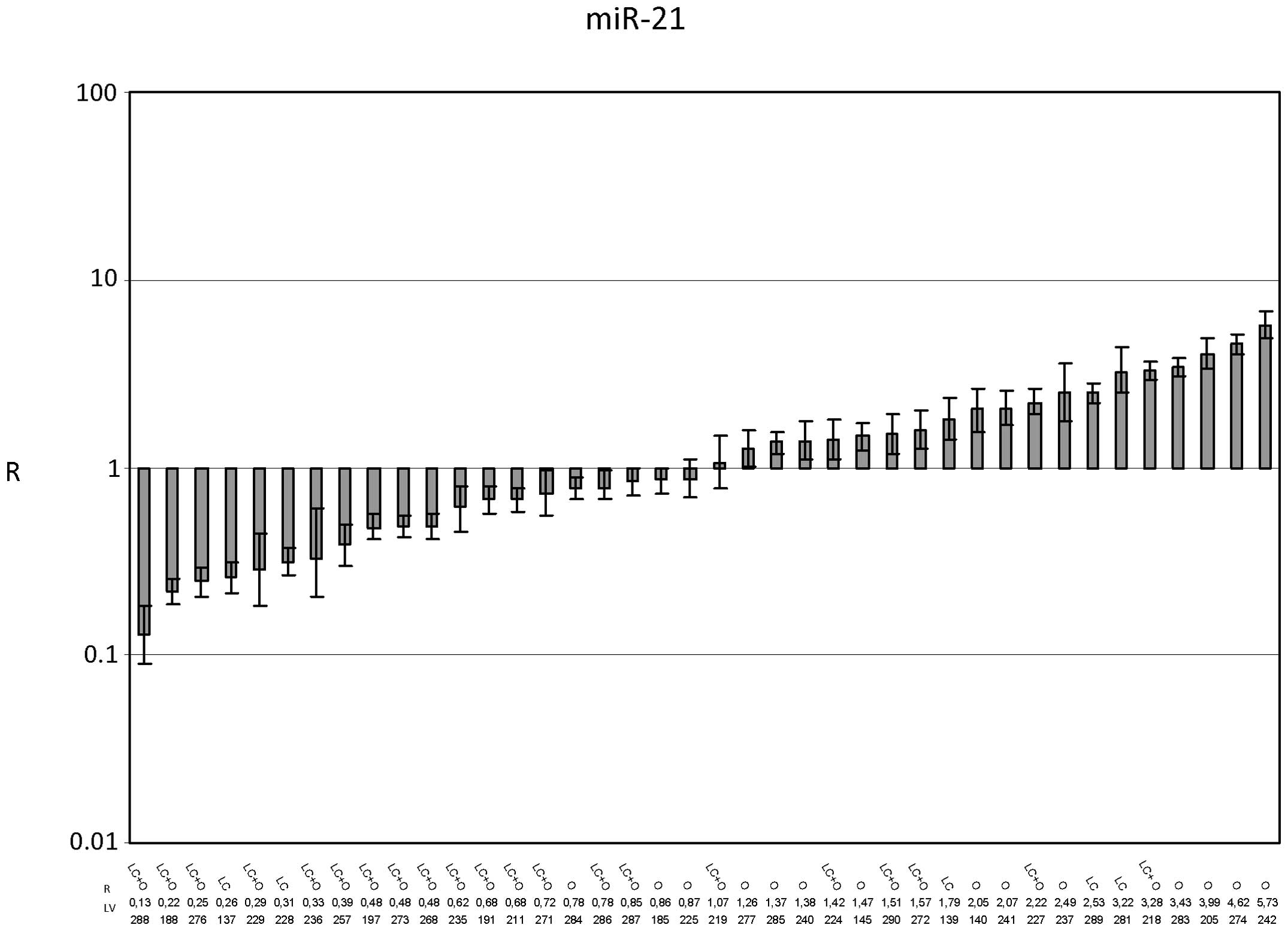
For miR-21 (Fig.
4), the average R value was 1.610±0.244 (p=0.016), indicating
the upregulation of miR-21 expression in HCC tissues with respect
to PT tissues. The R value in the presence of cirrhosis was
1.05±0.184, and the R value in its absence was 2.48±0.491
(p=0.0085) (Fig. 5). This result
suggested that miR-21 expression was unchanged in HCC that
developed in cirrhotic livers but was increased in HCC that
developed in noncirrhotic livers.
To determine whether the presence of the hepatitis
virus influenced miR expression levels, we stratified the cirrhotic
HCCs with respect to HBV/HCV infection status and we calculated the
mean R values for each miR considered (Table IV). The patient clinical
information concerning HBV/HCV infections and overall survival (OS)
was available in 18 of 25 cases of cirrhosis.
 | Table IV.R values in cirrhotic HCCs
subclassified in respect to hepatitis viral infections and the
correlations between R values and overall survival. |
Table IV.
R values in cirrhotic HCCs
subclassified in respect to hepatitis viral infections and the
correlations between R values and overall survival.
| HBV (n=3) | HCV (n=7) | HBV/HCV (n=4) | −/− (n=4) | Pearson correlation
(R vs OS) |
|---|
|
|
|
|
|
|---|
| R value (mean) | p-value | R value (mean) | p-value | R value (mean) | p-value | R value (mean) | p-value | Correlation
coefficient | p-value |
|---|
| miR-24 | 0.707 | 0.554 | 0.523 | 0.0184 | 0.462 | 0.0311 | 0.645 | 0.276 | 0.988 | 0.00945 |
| miR-27a | 0.302 | 0.0084 | 0.412 | 0.0024 | 0.35 | 0.0234 | 0.817 | 0.694 | 0.373 | 0.694 |
| miR-21 | 1.285 | 0.705 | 1.155 | 0.74 | 1.027 | 0.72 | 1.41 | 0.57 | 0.406 | 0.724 |
For miR-24, a significant decrease in expression was
observed in the HCV and HBV/HCV subclasses (R=0.523, p=0.0184;
R=0.462, p=0.0311 respectively). No changes in expression levels
were observed for the HBV and −/− subclasses. The mean R values
directly correlated with the OS, expressed in months (Pearson
correlation coefficient=0.988, p=0.00945). The mean OS rates were
59.5, 44.3, 39.7 and 57.5 months for the subclasses HBV, HCV,
HBV/HCV and −/−, respectively.
For miR-27a, a significant decrease in expression
was detected in the HBV, HCV and HBV/HCV subclasses (R=0.302,
p=0.0084; R=0.412, p=0.0024; R=0.35, p=0.0234, respectively). No
change in the expression level was observed for the −/− subclass
(R=0.817).
No miR-21 expression level variation was detected
among the 4 classes considered: HBV, HCV, HBV/HCV or no infection
(−/−). In fact, the R values ranged from 0.7 to 1.3 with the
exception of the −/− class, which displayed a slight increase in
miR-21 expression in comparison to the other samples (R=1.41).
Identification of a novel miRNA by
bioinformatic tools and the verification of its expression by
northern blotting
We cloned a sequence of 36 nucleotides that was
homologous to a repeating element on chromosomes 1 and 6 as
indicated by a matching of the sequence against the human genome
(BLAT). However, if one performs an alignment of the sequence
against the stem-loop sequences present in the miRBase database
(http://www.mirbase.org) it is partially
homologous (78%) to the precursor of the Mus musculus miR-1199
(Fig. 6A). To our knowledge, the
human miR orthologue of mmu-miR-1199 has not been yet discovered,
as it is not present within the miRBase official registry. The
sequence coding for mmu-miR-1199 is located on chromosome 8 between
the Prkaca and Rln3 genes. Based on an analysis of the synteny
between mice and humans (www.ensembl.org),
the corresponding homologous human sequence (79 nucleotides) is
located on chromosome 19p13.2 in the 2nd exon of a 1424
nucleotide-long transcript (ENST00000269720) coding an
uncharacterized 361 aa protein (ENSP00000269720). The human
sequence (79 nucleotides) plus an additional 72 nucleotides
flanking the 5′ and 3′ ends (Fig.
6B) is predicted to form a hairpin secondary structure, as
evidenced by mFOLD 3.2 (Fig. 6C)
and the sequence is well conserved among different mammalian
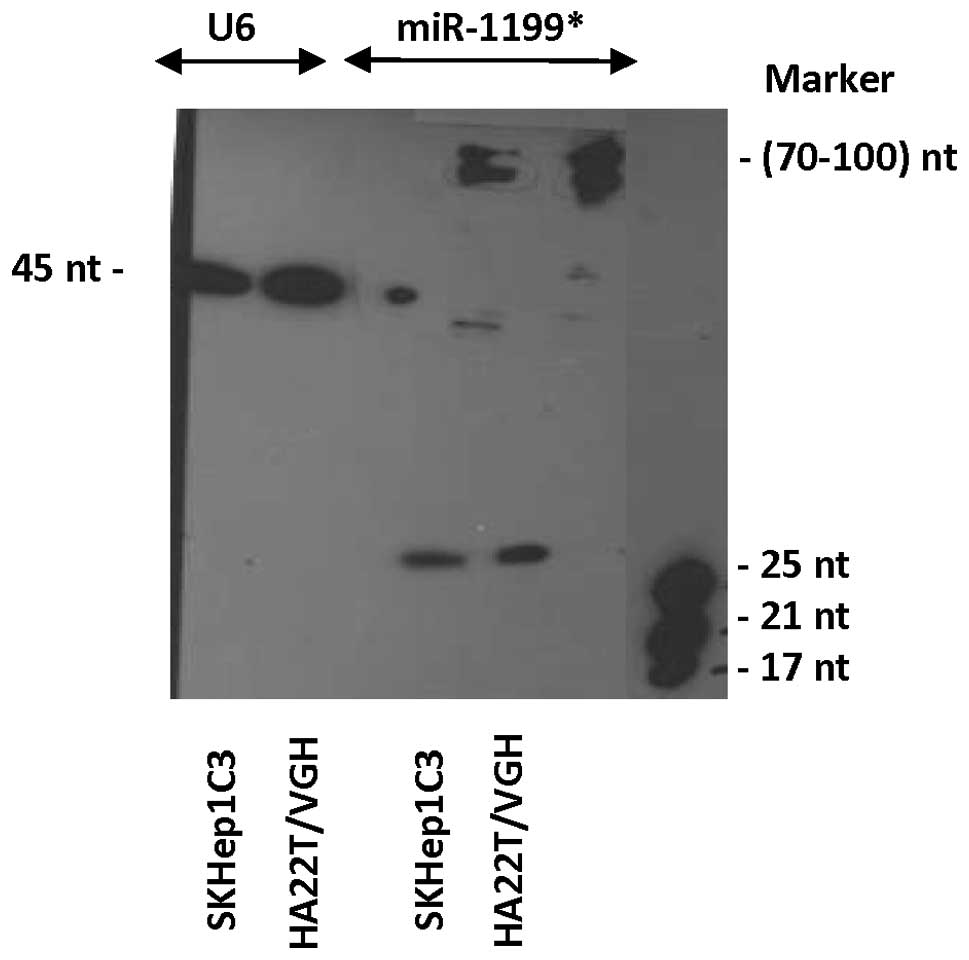
species (Fig. 6E). Fig. 7 shows the expression of the novel
candidate miR, as detected by northern blot analysis using
biotinylated probes. The expression of the miR, here denominated
hsa-miR-1199* (* indicates the strand that matures from
the 3′ arm of the candidate pre-miRNA), was evaluated in 2 HCC cell
lines, HA22T/VGH and SKHep1C3. The mature form (∼22–25 nucleotides)
and the precursor form (∼70–100 nucleotides) were clearly
detectable. The strand that matures from the 5′ arm of the
candidate pre-miRNA was not detected in the cell lines studied
(data not shown).
Discussion
In the present study we developed a small RNA
expression library in the HCC cell line HA22T/VGH to identify new
miRNAs and to study the profile of the expressed miRNAs. Among the
200 bacterial clones sequenced, 118 clones corresponded to 31 known
miRs cloned with different frequencies and the miR-24, miR-27a,
miR-21 were cloned with the highest frequency. Some of the 31
miRNAs cloned have been described previously to play a key role in
cancer pathogenesis. miR-17, miR-19b, miR-20a and miR 92a belong to
the cluster miR-17-92 known to be upregulated in several solid
tumors (16,17). The miR-25 and miR-93 belong to the
miR-106b-93-25 cluster that acts as oncomiRs (18). The miR-21 is well studied. It is
commonly upregulated in several types of malignancies, included HCC
(19–22). The miRs let7a and miR-7i belong to
the let-7 family that is involved in apoptosis induction and
inhibition of tumorigenicity (23–25).
The miR-29b regulates epigenetic changes and triggers apoptosis and
loss of tumorigenicity. It is downmodulated in colon and lung
cancer, cholangiocarcinoma and in chronic lymphocytic leukemia
(26,27). The miR-34b is a member of the
miR-34 family known to be implicated in inhibition of the
aggressive properties in several tumor cell lines by targeting MET,
bcl2 and CDK4/6 (4). Although
miR-122 has been identified as the most abundant liver-specific
microRNA, clones carrying the miR-122 were not observed in the
HA22T/VGH library. This may be due firstly to evidence that miR-122
is either silent or expressed at a very low level in most HCCs and
transformed cell lines (28).
Secondly, the sensitivity of the technique may have resulted in its
lack of identification.
To date, few studies have been published concerning
small RNA expression libraries in liver or in HCC cells/tissues. Fu
et al(29) cloned 27
different miRs in a human fetal liver identifying 5 novel miRs.
Among the most abundant, miR-24 was noted. Mizuguchi et
al(30) cloned miRs in HCC and
adjacent normal liver tissues by conventional cloning and 454
sequencing and they identified 7 novel miRs and several miR
sequence modifications. As corroborated by our results, they found
that miR-21 was expressed frequently as the edited form.
Considering the results obtained from a small RNA
expression library in the HPV16+ cervical cancer cell
line CaSki (31), with the
identification of 46 different miRs among a total of 174 clones
sequenced, notably several miRs cloned were the same found in our
study. In particular, the miRs cloned with the highest frequency
were miR-21, miR-27a and miR-24 as noted in our study. These and
our findings may indicate a relevant role of miR-21, miR-24 and
miR-27a in the malignant behavior of cervical cancer and HCC cell
lines; for this reason we monitored their expression levels in
human HCC tissues and their PT counterparts and then matched the
miR expression levels with clinical patient features.
miR-24 and miR-27a displayed the same expression
trend in 66.7% of the cases examined; this may have occurred due to
the fact that they are clustered in 1 transcript on chromosome 19.
To our knowledge, few data exist concerning miR expression during
the progression from normal liver to HCC through cirrhosis. In this
context, considering the subclass of HCC tumors developed in
cirrhotic liver, miR-24 and miR-27a were downregulated in HCC in
respect to PT tissues. This suggests that downregulation of miR-24
and miR-27a influences the hepatocyte transformation of cirrhotic
tissues. Our data revealed miR-24 and miR-27a dysregulation in HCC
in respect to their corresponding PT tissues and distinguished a
profile in cirrhotic but not in non-cirrhotic tissues. For miR-24
in cirrhotic HCCs, the data obtained indicate a linear correlation
between mean overall survival and miR-24 expression. The subclass
of cirrhotic HCC with HBV/HCV infection displayed the worst
outcome. It has been confirmed that miR expression can vary in
response to HBV/HCV virus infections and their combination but the
mechanisms are still poorly investigated.
miR-27a modification was different in cases without
cirrhosis (but with other background diseases). In this tumor
subclass miR-27a was upregulated in HCC tissues with respect to PT.
It has been reported that a given miR may act as an oncomiR or
tumor suppressor or that it may display pleiotropic properties in
different cells, tissues or stages of cancer progression due also
to the fact that a miR can control several targets (32). miR-27a was found to function as a
tumor suppressor by targeting the anti-apoptotic protein FADD in
human embryonic kidney cells (33). On the contrary, it acted as an
oncomiR favoring the MDR1 (multi-drug resistance 1) protein
upregulation in human ovarian cancer cells (34). miR-24 was described as an
anti-oncomiR by regulating c-myc and E2F2 in the HCC-derived cell
line HepG2 and causing inhibition of cell proliferation (35). In another study miR-24 acted as an
oncomiR negatively regulating p16 in cervical carcinoma cells and
the pro-apoptotic FAF1 protein in prostate, gastric and HeLa cancer
cells (36,37). More studies are necessary to better
explore the biological role of miR-24 and miR-27a in HCC and in
other cancers. The observation that miR-24 expression is correlated
with OS of cirrhotic HCC patients will be further validated in a
larger group of patients.
miR-21, a well known wide-broad oncomiR, known to be
overexpressed in HCC vs. normal liver was upregulated in HCC
respect to PT tissues, and this became more evident in the subclass
of HCC developed in non-cirrhotic liver. In the subclass of
cirrhotic HCC the average R value was very near to 1 indicating no
differences in the miR-21 expression level between cirrhotic PT and
HCC tissues, thus supporting the hypothesis that miR-21
upregulation may be an early event during hepatocarcinogenesis.
This is in line with the consideration that the tumor suppressor
PTEN, one of the miR-21 targets, is already downmodulated in the
pathological stages that often preceed the onset of HCC such as
steatosis and cirrhosis (21). As
a novel findings, our present data revealed miR-21 overexpression
in HCC without cirrhosis as a background disease and found no
expression variation between HCC and PT cirrhotic tissues (30). Jiang et al(38) reported miR-21 upregulation when the
tumor tissue expression data were compared with the adjacent PT
tissues, that were not stratified according to the background
disease.
Another novel finding of the present report is the
identification of a new human miR. During the sequence analyses we
identified a 36-nt sequence that was partially homologous to the
Mus musculus pre-miR-1199 and we realized that the corresponding
human miR ortholog was not yet annotated in the miR registry. Three
annotation criteria (expression and biogenesis criteria) (10) based on bioinformatic tools and
experimental validation confirmed that miR-1199 was a new
hsa-miR.
The expression analysis verified by northern blot
analysis conducted in two human HCC cell lines showed that the
expressed miR was miR-1199* contained in the 3′ arm of
the candidate hairpin-precursor miR. In contrast, in the mouse the
miR strand cloned with the highest frequency was that matured from
the the 5′ arm of the pre-miR-1199. However, to date no information
is available concerning the mmu-miR-1199 role and target
validations and further studies are necessary to identify them both
in the human and mouse.
The annotation criteria also suggest that close
homologs in other species can be annotated as miR orthologues
without experimental validation, based on the criterion concerning
the phylogenetic conservation of the ∼22 nt miR sequence and its
predicted fold-back precursor secondary structure. In our case,
miR-1199 may be annotated as a new miR also in Pongo
pygmaeus, Pan troglodytes, Canis familiaris and
Rattus norvegicus.
In conclusion, we identified the new human miR-1199,
also phylogenetically conserved in other mammalian species. We also
found that miR-24 and miR-27a were downregulated in HCC cancer
developed in cirrhotic liver. Thus, these findings may contribute
to advances in the discovery and validation of novel early
molecular biomarkers of HCC progression from cirrhosis to
cancer.
Acknowledgements
The authors thank V. Mazzaferro and S.
Toffanin for the critical reading of the manuscript. The authors
would like to thank Dr E. Frassi for collecting clinical data of
HCC patients. The English text was edited by the NPGLE service.
This study was partially supported by the Ministero
dell’Istruzione, dell’Università e della Ricerca (MIUR), MIUR PRIN
2007 (MWCEAL_003), Fondazione Cariplo, by MIUR local funds of the
University of Brescia, by Regione Lombardia NEDD. Research
fellowships were awarded to A.S. and E.A. by Lega Italiana per la
lotta contro i Tumori di Brescia (LILT-Brescia, Italy).
References
|
1.
|
Pauli A, Rinn JL and Schier AF: Non-coding
RNAs as regulators of embryogenesis. Nat Rev Genet. 12:136–149.
2011. View
Article : Google Scholar : PubMed/NCBI
|
|
2.
|
Iorio MV and Croce CM: MicroRNAs in
cancer: small molecules with a huge impact. J Clin Oncol.
27:5848–5845. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3.
|
O’Connell MR, Rao DS, Chaudhuri AA and
Baltimore D: Physiological and pathological roles for microRNAs in
the immune system. Nat Rev Immunol. 10:111–122. 2010.PubMed/NCBI
|
|
4.
|
Garofalo M and Croce CM: microRNAs: master
regulators as potential therapeutics in cancer. Annu Rev Pharmacol
Toxicol. 51:25–43. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5.
|
Griffiths-Jones S, Saini HK, van Dongen S
and Enright AJ: miRBase: tools for microRNA genomics. Nucleic Acids
Res. 36:D154–D158. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Lagos-Quintana M, Rauhut R, Meyer J,
Borkhardt A and Tuschl T: New microRNAs from mouse and human. RNA.
9:175–179. 2003. View Article : Google Scholar
|
|
7.
|
Lagos-Quintana M, Rauhut R, Lendeckel W
and Tuschl T: Identification of novel genes coding for small
expressed RNAs. Science. 294:853–858. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
8.
|
Vaz C, Ahmad HM, Sharma P, Gupta R, Kumar
L, Kulshreshtha R and Bhattacharya A: Analysis of microRNA
transcriptome by deep sequencing of small RNA libraries of
peripheral blood. BMC Genomics. 11:2882010. View Article : Google Scholar : PubMed/NCBI
|
|
9.
|
Huang Y, Zou Q, Wang SP, Tang SM, Zhang GZ
and Shen XJ: The discovery approaches and detection methods of
microRNAs. Mol Biol Rep. 38:4125–4135. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10.
|
Ambros V, Bartel B, Bartel DP, et al: A
uniform system for microRNA annotation. RNA. 9:277–279. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
11.
|
Berezikov E, Cuppen E and Plasterk RHA:
Approaches to microRNA discovery. Nat Genet. 38:S2–S7. 2006.
View Article : Google Scholar
|
|
12.
|
Takane K, Fujishima K, Watanabe Y, Sato A,
Saito N, Tomita M and Kanai A: Computational prediction and
experimental validation of evolutionarily conserved microRNA target
genes in bilaterian animals. World J Gastroenterol. 17:1910–1914.
2011.PubMed/NCBI
|
|
13.
|
McKillop IH, Moran DM, Jin X and Koniaris
LG: Molecular pathogenesis of hepatocellular carcinoma. J Surg Res.
136:125–135. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
14.
|
Barlati S, Zoppi N, Copeta A, Tavian D, De
Petro G and Colombi M: Quantitative in situ hybridization for the
evaluation of gene expression in asynchronous and synchronized cell
cultures and in tissue sections. Histol Histopathol. 14:1231–1240.
1999.PubMed/NCBI
|
|
15.
|
Salvi A, Sabelli C, Moncini S, Venturin M,
Arici B, Riva P, Portolani N, Giulini SM, De Petro G and Barlati S:
MicroRNA-23b mediates urokinase and c-met downmodulation and a
decreased migration of human hepatocellular carcinoma cells. FEBS
J. 276:2966–2982. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16.
|
Hayashita Y, Osada H, Tatematsu Y, Yamada
H, Yanagisawa K, Tomida S, Yatabe Y, et al: A polycistronic
microRNA cluster, miR-17-92, is overexpressed in human lung cancers
and enhances cell proliferation. Cancer Res. 65:9628–9632. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
17.
|
Inomata M, Tagawa H, Guo YM, Kameoka Y,
Takahashi N and Sawada K: MicroRNA-17-92 down-regulates expression
of distinct targets in different B-cell lymphoma subtypes. Blood.
113:396–402. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
18.
|
Petrocca F, Visone R, Onelli MR, Shah MH,
Nicoloso MS, de Martino I, Iliopoulos D, et al: E2F1-regulated
microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis
in gastric cancer. Cancer Cell. 13:272–286. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
19.
|
Si ML, Zhu S, Wu HS, Lu Z, Wu F and Mo Y:
miR-21-mediated tumor growth. Oncogene. 26:2799–2803. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
20.
|
Selcuklu SD, Donoghue MT and Spillane C:
miR-21 as a key regulator of oncogenic processes. Biochem Soc
Trans. 37:918–925. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21.
|
Meng F, Henson R, Wehbe-Janek H, Ghoshal
K, Jacob ST and Patel T: MicroRNA-21 regulates expression of the
PTEN tumor suppressor gene in human hepatocellular cancer.
Gastroenterology. 133:647–658. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22.
|
Gaur AB, Holbeck SL, Colburn NH and Israel
MA: Downregulation of Pdcd4 by miR-21 facilitates glioblastoma
proliferation in vivo. Neurooncology. 13:580–590. 2011.PubMed/NCBI
|
|
23.
|
Johnson SM, Grosshans H, Shingara J, Byrom
M, Jarvis R, Cheng A, Labourier E, et al: RAS is regulated by the
let-7 microRNA family. Cell. 120:635–647. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
24.
|
Sampson VB, Rong NH, Han J, Yang Q, Aris
V, Soteropoulos P, Petrelli NJ, et al: MicroRNA let-7a
down-regulates MYC and reverts MYC-induced growth in Burkitt
lymphoma cells. Cancer Res. 67:9762–9770. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25.
|
Osada H and Takahashi T: let-7 and
miR-17-92: small-sized major players in lung cancer development.
Cancer Sci. 102:9–17. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26.
|
Mott JL, Kobayashi S, Bronk SF and Gores
GJ: miR-29 regulates Mcl-1 protein expression and apoptosis.
Oncogene. 26:6133–6140. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27.
|
Garzon R, Heaphy CE, Havelange V, Fabbri
M, Volinia S, Tsao T, Zanesi N, et al: MicroRNA 29b functions in
acute myeloid leukemia. Blood. 114:5331–5341. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
28.
|
Bai S, Nasser MW, Wang B, Hsu SH, Datta J,
Kutay H, Yadav A, et al: MicroRNA-122 inhibits tumorigenic
properties of hepato-cellular carcinoma cells and sensitizes these
cells to sorafenib. J Biol Chem. 284:32015–32027. 2009. View Article : Google Scholar
|
|
29.
|
Fu H, Tie Y, Xu C, Zhang Z, Zhu J, Shi Y,
Jiang H, et al: Identification of human fetal liver miRNAs by a
novel method. FEBS Lett. 579:3849–3854. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
30.
|
Mizuguchi Y, Mishima T, Yokomuro S, Arima
Y, Kawahigashi Y, Shigehara K, Kanda T, et al: Sequencing and
bioinformatics-based analyses of the microRNA transcriptome in
hepatitis B-related hepatocellular carcinoma. PLoS One.
6:e153042011. View Article : Google Scholar : PubMed/NCBI
|
|
31.
|
Wang X, Tang S, Le SY, Lu R, Rader JS,
Meyers C and Zheng ZM: Aberrant expression of oncogenic and
tumor-suppressive microRNAs in cervical cancer is required for
cancer cell growth. PLoS One. 3:e25572008. View Article : Google Scholar : PubMed/NCBI
|
|
32.
|
Kwak PB, Iwasaki S and Tomari Y: The
microRNA pathway and cancer. Cancer Sci. 101:2309–2315. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
33.
|
Chhabra R, Adlakha YK, Hariharan M, Scaria
V and Saini N: Upregulation of miR-23a-27a-24-2 cluster induces
caspase-dependent and -independent apoptosis in human embryonic
kidney cells. PLoS One. 4:e58482009. View Article : Google Scholar : PubMed/NCBI
|
|
34.
|
Li Z, Hu S, Wang J, Cai J, Xiao L, Yu L
and Wang Z: miR-27a modulates MDR1/P-glycoprotein expression by
targeting HIPK2 in human ovarian cancer cells. Gynecol Oncol.
119:125–130. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
35.
|
Lal A, Navarro F, Maher CA, Maliszewski
LE, Yan N, O’Day E, Chowdhury D, et al: miR-24 inhibits cell
proliferation by targeting E2F2, MYC, and other cell-cycle genes
via binding to ‘seedless’ 3′UTR microRNA recognition elements. Mol
Cell. 35:610–625. 2009.PubMed/NCBI
|
|
36.
|
Lal A, Kim HH, Abdelmohsen K, Kuwano Y,
Pullmann R Jr, Srikantan S, Subrahmanyam R, et al: p16(INK4a)
translation suppressed by miR-24. PLoS One. 3:e18642008. View Article : Google Scholar : PubMed/NCBI
|
|
37.
|
Qin W, Shi Y, Zhao B, Yao C, Jin L, Ma J
and Jin Y: miR-24 regulates apoptosis by targeting the open reading
frame (ORF) region of FAF1 in cancer cells. PLoS One. 5:e94292010.
View Article : Google Scholar : PubMed/NCBI
|
|
38.
|
Jiang J, Gusev Y, Aderca I, Mettler TA,
Nagorney DM, Brackett DJ, Roberts LR and Schmittgen TD: Association
of microRNA expression in hepatocellular carcinomas with hepatitis
infection, cirrhosis, and patient survival. Clin Cancer Res.
14:419–427. 2008. View Article : Google Scholar : PubMed/NCBI
|