Introduction
Hepatocyte growth factor (HGF) is a multifunctional
molecule that acts as mitogen or morphogen in variety of cells
(1). It is known that HGF acts as
paracrine mediator that promotes proliferation, survival, migration
and morphogenesis of epithelial cells (1,2). HGF
also induces angiogenesis and inhibits apoptosis through c-MET
(1,2). A number of reports have demonstrated
that the overexpression of HGF and c-MET is associated with cancer
invasion, metastasis and poor prognosis in various types of cancer
(3–8). Increased serum levels of HGF were
reported in breast and lung cancer patients (9,10),
and an association between increased serum HGF level and poor
prognosis was observed in ovarian, colorectal, hepatocellular,
renal cell and bladder cancer (1,2,11–13).
Ma et al have reported a possible molecular
mechanism for aberrant HGF expression in human breast cancer
(14). They showed that the
HGF promoter element harbors a mononucleotide repeat of
approximately 30 deoxyadenosines, which has been termed the
deoxyadenosine tract element (DATE), and the repeat length mutation
within the DATE had a significant effect on HGF promoter
activity in breast cancer cell lines (14). Several studies have shown the
mechanism by which the poly-deoxyadenosine repeats affect gene
expression levels (14–19). Namely, long deoxyadenosine repeats
have been shown to prevent accessibility to promoter regions and to
increase the equilibrium accessibility of other DNA target sites
buried inside nucleosomes, while short deoxyadenosine tracts were
suggested to stimulate transcription by improving accessibility to
promoters and decreasing the stability of the DNA wrapping
(18–20). While the HGF DATE may
influence the malignant phenotype and/or the progression of breast
and gastric cancers (14,21), there has been no report evaluating
the significance of the HGF DATE in bladder cancer.
In the present study, we investigated the influence
of germ line variants of the HGF DATE on bladder cancer
risk. Furthermore, we assessed the association of
clinicopathological factors and HGF mRNA expression with
somatic variants of the HGF DATE in bladder cancer, which
may represent a mutational effect of DATE alteration.
Materials and methods
Subjects
A total of 140 patients with bladder cancer treated
at Akita University Medical Center were enrolled in this study. All
the patients were histologically diagnosed with urothelial
carcinoma of the bladder with specimens obtained from transurethral
biopsy or surgical resection. Clinical and histopathological
information was obtained from patients’ medical charts, imaging
studies and pathological reports. A total of 95 healthy native
Japanese men and women over the age of 60 years from Akita
Prefecture, who underwent routine community health checkups, were
recruited as controls. They were checked by routine urinalysis to
rule out urinary tract diseases. The tumor grading system conformed
to the World Health Organization grading system (22) and the tumor staging system was
based on the American Joint Committee on Cancer TNM classification
system (23). This study was
approved by the Institutional Review Board of Akita University
School of Medicine, and all the subjects provided written informed
consent and were asked to provide clinical information, blood and
tissue samples for genetic analysis.
HGF genotyping analysis
Germ line DNA was extracted from peripheral blood
lymphocyte (PBL) samples collected from individuals using a QIAamp
Blood Kit (Qiagen, Hilden, Germany). Tumor DNA was extracted from
bladder tumor tissues obtained at transurethral resection (TURBT).
The length of the DATE in the HGF promoter region was
determined by a polymerase chain reaction (PCR). The PCR primers
(forward, 5′-GGGACAGGTATTGTGGGGCCAAAATAAG-3′; and reverse,
5′-GGGTGTGGTATTGTGGGGCCAAAATAAG-3′) generated a 247-bp product when
the length of DATE was 30 deoxyadenosines. PCR reactions were
performed in a 25 μl volume containing 20 ng of genomic DNA,
1X PCR buffer (PE Applied Biosystems, Branchburg, NJ, USA), 0.2 mM
of each dNTP, 2.5 mM MgCl2, 50 pM of each primer, and
one unit of Ampli-Taq Gold DNA polymerase (PE Applied Biosystems).
The PCR amplification conditions were as follows: 10 min at 94°C,
followed by 35 cycles of 30 sec at 94°C, 30 sec at 62°C and 60 sec
at 72°C, with a final extension of 10 min at 72°C in a thermal
cycler (GeneAmp PCR System 9700, Perkin-Elmer, Branchburg, NJ). The
samples were purified using the Illustra AutoSeq G-50 Kit (GE
Healthcare Life Sciences, Little Chalfont, UK). All purified
fragments were subjected to DNA sequencing with forward and reverse
primers using the BigDye Terminator Cycle Sequencing Ready Reaction
Kit (Applied Biosystems, Foster City, CA, USA), and analyzed with a
Genetic Analyzer (ABI PRISM 310 Genetic Analyzer, Applied
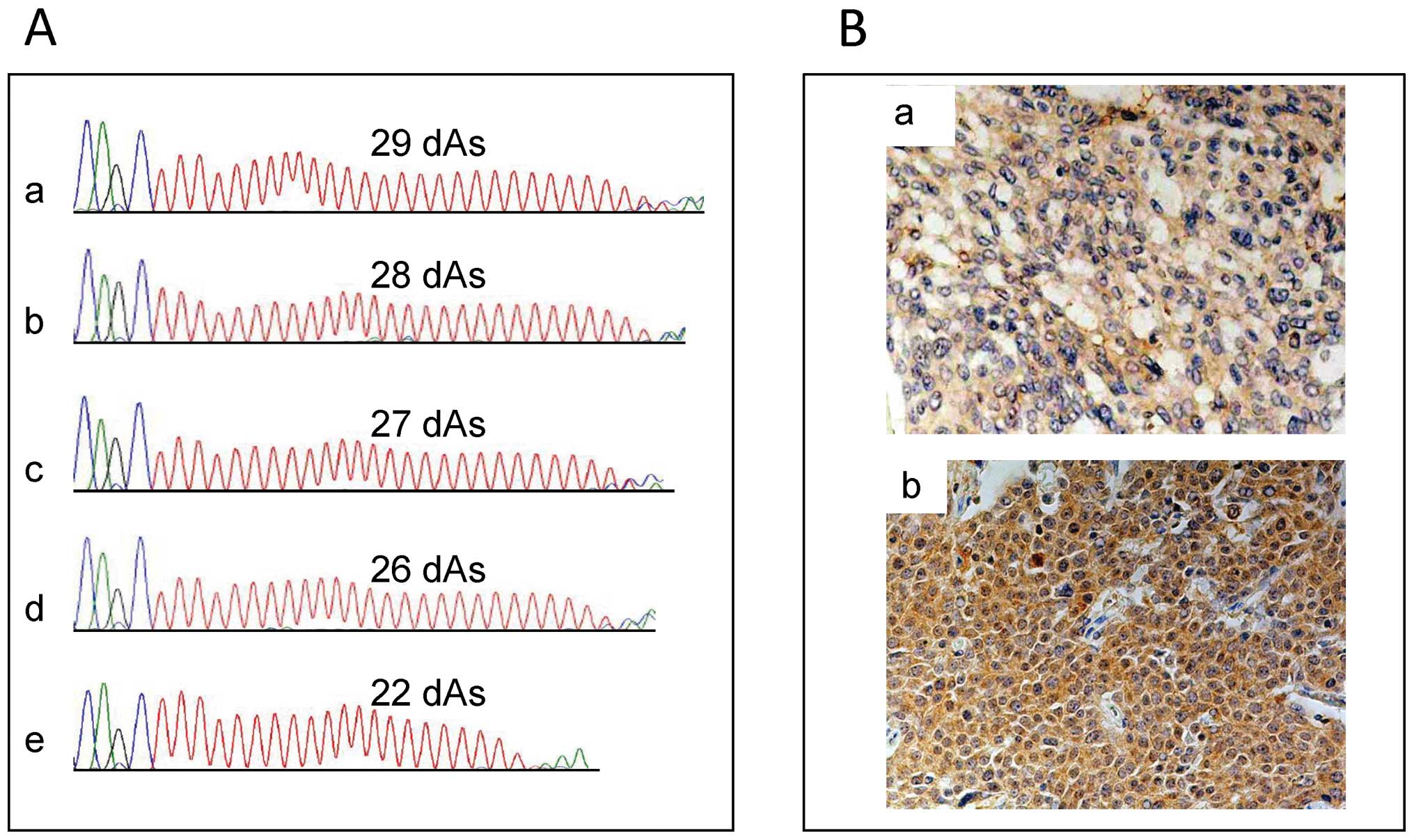
Biosystems). Because we could clearly define only the length of the
shorter DATE owing to the direct sequencing method, the data of the
shorter DATE length were used in further statistical analyses
(Fig. 1A). The short DATE was
defined as 27 deoxyadenosine repeats or fewer, while the long DATE
was defined as more than 27 repeats, as described in the
Results.
Expression analysis of HGF mRNA using
RT-PCR
Total RNA was extracted from approximately 30 mg of
tumor tissues obtained by TURBT using TRIzol RNA Isolation Reagent
(Life Technologies Inc., Rockville, MD, USA) and was
reverse-transcribed to cDNA using a SuperScript VILO cDNA Synthesis
Kit (Life Technologies Inc.).
Real-time RT-PCR amplication mixtures (20 μl)
contained 2 μl template cDNA, 2X SYBR-Green Master mix
buffer (10 μl) (Takara Bio Inc., Otsu, Japan) and 100
μM forward and reverse primers (0.8 μl). Reactions
were run on a Thermal Cycler Dice Real-Time System (Takara Bio
Inc.) with cycling conditions of 30 sec at 95°C, 40 cycles at 95°C
for 5 sec, and 60°C for 60 sec. The primers used were as follows:
5′-ATGATGATGCTCATGGACCCT-3′ (forward) and
5′-CTGGCAAGCTTCATTAAAACC-3′ (reverse) for HGF and
5′-TGATGACATCAAGAAGGTGGTGAAG-3′ (forward) and
5′-TCCTTGGAGGCCATGTGGGCCAT-3′ (reverse) for GAPDH. PCR
reactions were simultaneously performed for HGF and the
reference GAPDH. The HGF expression levels normalized
by the GAPDH were used for further analyses.
HGF immunohistochemical analysis
Ninety-one bladder cancer specimens obtained at
radical cystectomy were subjected to immunohistochemical analysis.
The specimens were fixed in 10% buffered formalin and embedded in
paraffin. Serial sections (5-μm thick) were deparaffinized
in xylene, rehydrated in a graded series of decreasing ethanol
concentration, and then rinsed in Tris-buffered saline. For HGF
immnostaining, HGF was detected with a goat polyclonal anti-human
HGF antibody (1:50; R&D Systems, Minneapolis, MN, USA) as
primary antibody. Antigen retrieval treatment was performed at
121°C for 10 min in 10 mM sodium citric acid (pH 6.0), and
endogenous peroxidases were blocked using 0.3% hydrogen
peroxidase/methanol for 30 min. After washing in phosphate-buffered
saline (PBS) for 5 min and 10% bovine serum albumin/PBS for 30 min,
the sections were exposed to a primary antibody diluted 1:50
overnight at 4°C. After washing in PBS, a secondary antibody
conjugated with anti-goat IgG was applied, followed by incubation
at room temperature for 30 min. Immunoreactions were visualized
using 3,3′-diaminobenzidine as chromogen. Positive control was
represented by a normal hepatic cell showing strong cytoplasmic
expression for HGF.
To assess HGF immunoreactivity, we used the
following scoring systems. Cytoplasmic HGF staining intensity was
scored on a semi-quantitative scale as: 1, weak; 2, moderate; 3,
strong; and 4, very strong. The percent of cytoplasmic HGF-positive
cells was divided into four groups: 1 (<25%), 2 (25–50%), 3
(50–75%) and 4 (>75%). Total immunoreactivity was finally
calculated by multiplying the two scores and was classified into
two groups: low expression (≤9) and high expression (>9)
(Fig. 1B).
Statistical analysis
The data was analyzed using SPSS software (version
19.0J, SPSS Inc., Chicago, IL, USA). Two group comparisons were
performed with the Mann-Whitney U test for continuous variables and
χ2 test for categorical variables. A probability of
<0.05 was required for statistical significance. Kaplan-Meier
survival curves with the log-rank test were used to compare overall
survival and disease-specific survival between patients with high
and low HGF expression.
Results
Comparison of the length of the HGF DATE
in germ-line DNA
We evaluated the length of the DATEs in PBLs of 140
bladder cancer patients and 95 healthy controls. Distributions of
the DATE lengths in bladder cancer patients and controls were as
follows: in bladder cancer patients, the frequency of 30, 29, 28,
27, 26, 25, 24 and 22 doxyadenosine repeats in the DATE were 3
(2.1%), 15 (10.7%), 52 (37.1%), 48 (34.3%), 13 (9.3%), 5 (3.6%), 2
(1.4%) and 2 (1.4%), respectively. In the controls, the frequency
of 30, 29, 28, 27, 26 and 25 doxyadenosine repeats in the DATE were
4 (4.2%), 32 (33.7%), 36 (37.9%), 18 (18.9%), 4 (4.2%) and 1
(1.1%), respectively. Because the median length of DATE was 28 in
both groups, we classified the length of DATE into two categories
as short (≤27) and long (>27). When we applied our
classification, the frequency of individuals with a short DATE was
significantly higher in bladder cancer patients than controls (49.3
vs. 24.2%, P<0.001) (Table
I).
 | Table I.Bladder cancer patient and control
demographics and status of short deoxyadenosine repeats in the
HGF DATE. |
Table I.
Bladder cancer patient and control
demographics and status of short deoxyadenosine repeats in the
HGF DATE.
| Bladder cancer | Control | P-value |
|---|
| Total | 140 | 95 | |
| Age (Range) | 68.4±7.8 (16–94) | 68.1±4.5 (60–75) | 0.694 |
| Gender | | | 0.399 |
| Male | 107 | 77 | |
| Female | 33 | 18 | |
| Grade | | | |
| G1–2 | 48 (34.3) | | |
| G3 | 92 (65.7) | | |
| pT | | | |
| pTa-1 | 63 (45) | | |
| >pT1 | 77 (55) | | |
| HGF DATE | | | <0.001 |
| Short DATE
(−) | 71 (50.7) | 72 (75.8) | |
| Short DATE
(+) | 69 (49.3) | 23 (24.2) | |
Comparison of the frequency of short DATE
between bladder tumor tissue and PBLs
We assessed the difference in the frequency of short
DATEs between bladder tumor tissue and PBLs. In addition, we
evaluated the association between the clinicopathological factors
of bladder cancer and the presence of a short DATE in bladder tumor
tissue. The patients comprised 55 males and 15 females with a mean
age of 73.2 years. There were 51 non-muscle invasive tumors (NMIT)
(pTa-1) and 19 muscle invasive tumors (MIT) (pT2), and 42 low grade
tumors (grade 1 or 2) and 28 high grade tumors (grade 3). The
frequency of patients with a short DATE in their PBLs was 41.4%
(29/70), while that in their bladder tumor tissue was 52.9% (37/70)
(Table II). The frequency of
patients with a short DATE in their bladder tumor tissue was
significantly higher than that in their PBLs (p=0.008) (Table II). In 25 (35.7%) patients, the
length of the DATE in their bladder tumor tissues was shorter than
that in their PBLs. In 12 (17.1%) patients, the length of the DATE
in their bladder tumor tissues was longer than that in their PBLs.
The frequency of the mutation to long DATE was significantly higher
than that to short DATE (p=0.047 by binominal test). The results
suggested that the DATE was a frequent target of somatic mutation
(37/70, 52.9%) in bladder cancers and the mutation from long DATE
to short DATE might be dominant and play a role in the progression
of bladder cancer. There was no significant difference between NMIT
and MIT in the frequency of patients with a short DATE either in
PBLs or bladder tumor tissue (Table
II). No significant association was observed between the
presence of a short DATE in PBLs and tumor grade, while tumor grade
was significantly higher in bladder tumor tissue with a short DATE
than those without a short DATE (p=0.015, Table II).
 | Table II.Comparison of clinicopathological
factors with the status of the HGF DATE in bladder tumor
tissues and peripheral blood lymphocytes. |
Table II.
Comparison of clinicopathological
factors with the status of the HGF DATE in bladder tumor
tissues and peripheral blood lymphocytes.
| | BT tissue
| PBL
|
|---|
| Total | Short DATE (−) | Short DATE (+) | P-value | Short DATE (−) | Short DATE (+) | P-value |
|---|
| 70 | 33 | 37 | | 41 | 29 | 0.008 |
| Age (Range) | 73.2±10.5 | 73.2±10.6
(43–88) | 73.1±10.5
(53–94) | 0.949 | 71.6±11.0
(53–89) | 75.3±9.5
(56–94) | 0.144 |
| Gender | | | | 0.784 | | | 0.780 |
| Male | 55 | 26 | 29 | | 31 | 23 | |
| Female | 15 | 7 | 8 | | 10 | 6 | |
| pT | | | | 0.178 | | | 0.786 |
| pTa-1 | 51 | 27 (82) | 24 (66) | | 29 (71) | 22 (76) | |
| pT2 | 19 | 6 (18) | 13 (34) | | 12 (29) | 7 (24) | |
| Grade | | | | 0.015 | | | 0.621 |
| G1–2 | 42 | 25 (76) | 17 (46) | | 26 (65) | 16 (55) | |
| G3 | 28 | 8 (24) | 20 (54) | | 15 (35) | 13 (45) | |
Association of HGF mRNA expression levels
with DATE length in bladder tumor tissue
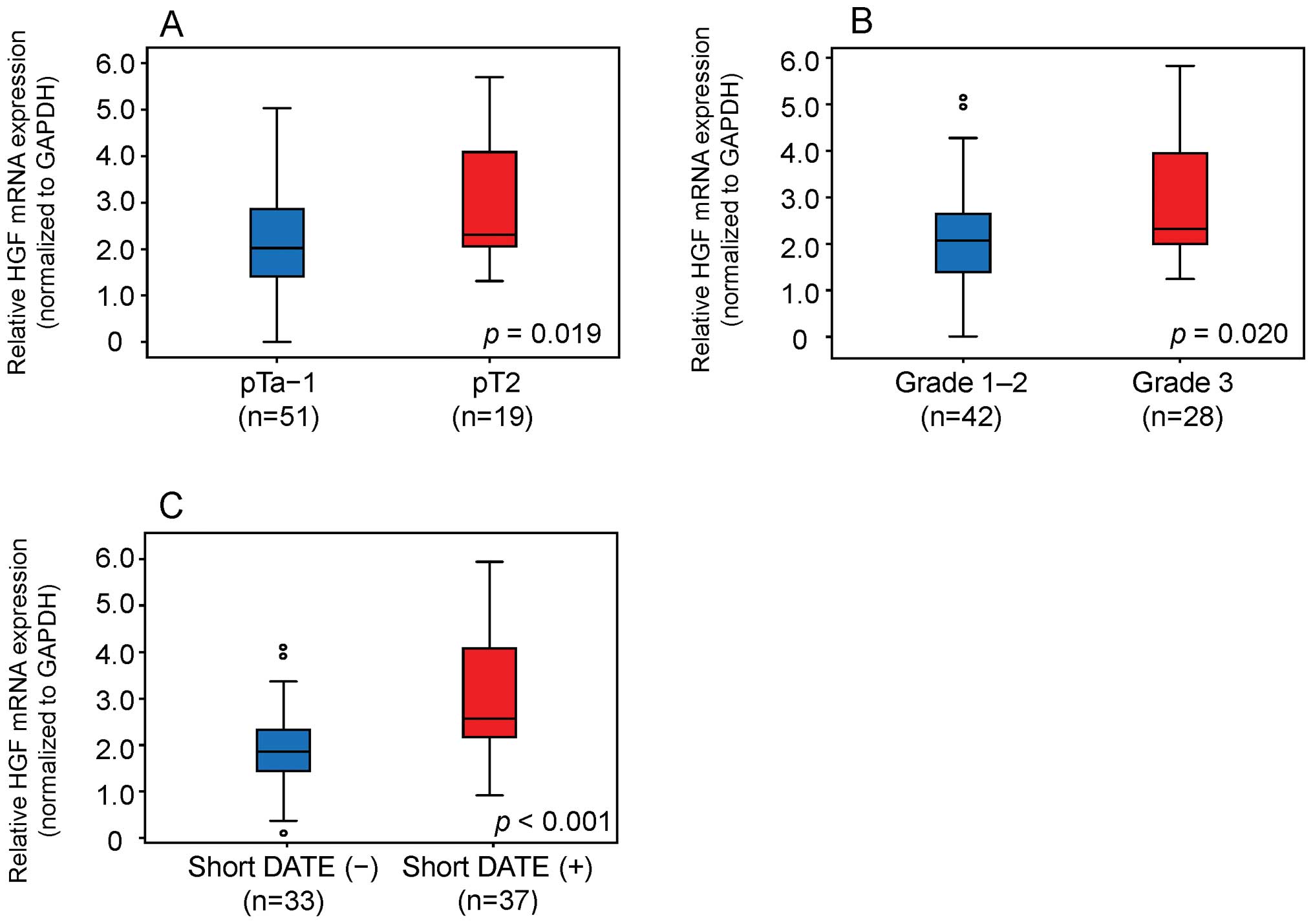
We compared HGF mRNA levels between bladder
tumor tissue with and without a short DATE. MIT showed
significantly higher HGF mRNA expression levels (p=0.019)
than NMIT (Fig. 2A). High grade
tumors showed significantly higher HGF mRNA expression
levels than low grade tumors (p=0.020) (Fig. 2B). HGF mRNA expression was
significantly higher in bladder tumor tissue with a short DATE than
that without a short DATE (p<0.001) (Fig. 2C).
HGF immunohistochemical analysis
We evaluated the relationship between HGF
immunohistochemical expression in bladder cancer and
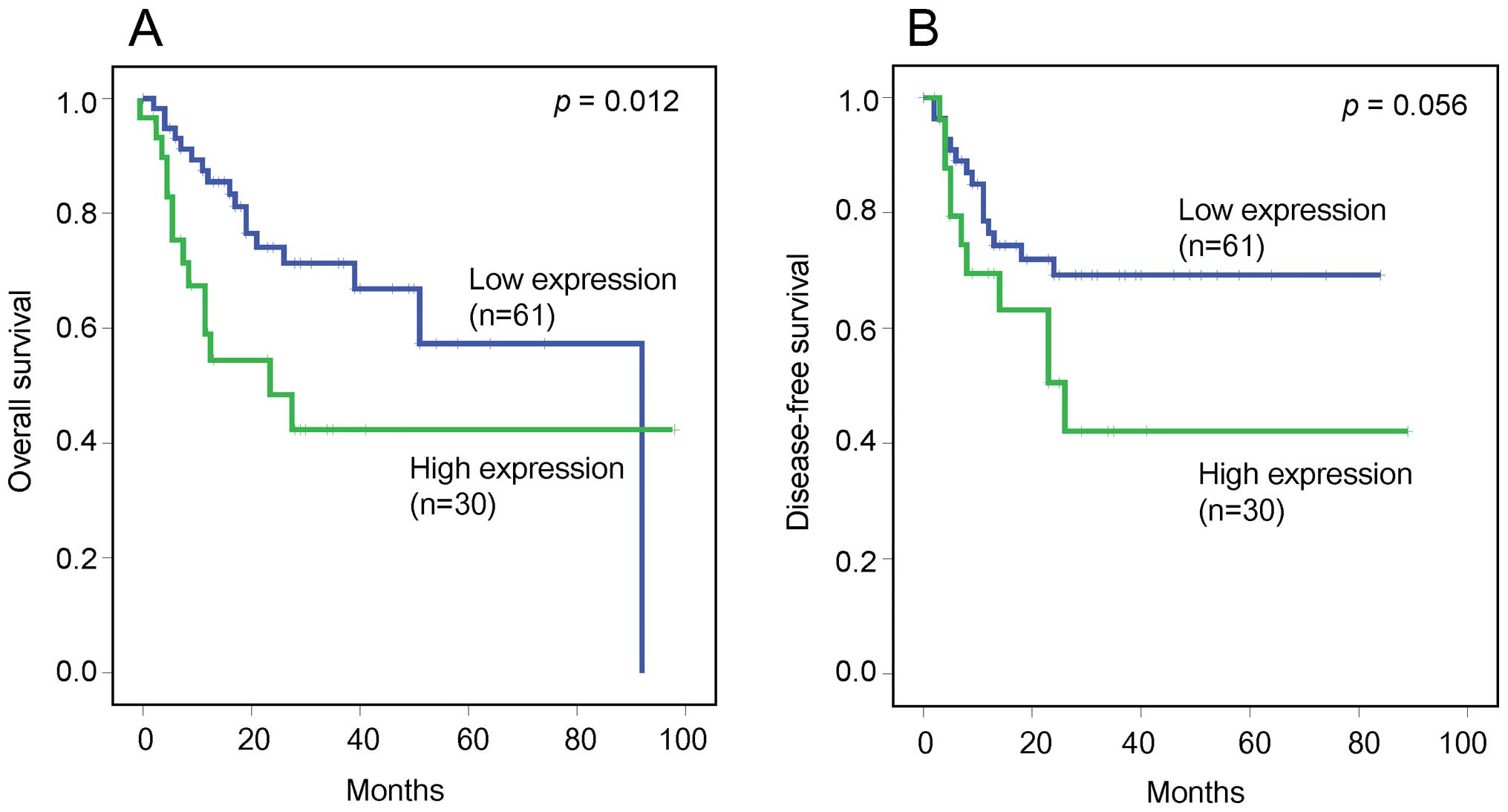
clinicopathological factors. Overall and disease-free survival
rates were compared according to the HGF expression scores in 91
patients who underwent radical cystectomy. The patients comprised
70 males and 21 females with a mean age of 69.8 years. The mean
follow-up duration was 19 months (range 0–98). Low and high HGF
expression scores were observed in 61 (67.0%) and 30 (33.0%)
patients, respectively (Table
III). The high HGF expression group had a significantly worse
overall survival than the low HGF expression group (p=0.012).
Furthermore, although not statistically significant, the high HGF
expression group had a tendency towards worse disease-free survival
than the low HGF expression group (p=0.056) (Fig. 3).
 | Table III.Comparison of bladder tumor HGF
expression score with the clinical characteristics of patients who
had undergone radical cystectomy. |
Table III.
Comparison of bladder tumor HGF
expression score with the clinical characteristics of patients who
had undergone radical cystectomy.
| Total | Low | High | P-value |
|---|
| Number | 91 | 61 | 30 | |
| Age (Range) | 69.8±11.0
(25–91) | 67.5±11.6
(25–85) | 74.5±7.7
(56–91) | 0.004 |
| Gender | | | | 0.298 |
| Male | 70 | 49 | 21 | |
| Female | 21 | 12 | 9 | |
| pT | | | | 0.502 |
| ≤pT2 | 51 | 36 | 15 | |
| >pT2 | 40 | 25 | 15 | |
| pN | | | | 0.395 |
| pN0 | 67 | 47 | 20 | |
| pN1-3 | 17 | 10 | 7 | |
| pNx | 7 | 4 | 3 | |
Discussion
In this study, we showed that the frequency of the
short HGF DATE in germ line DNA was significantly higher in
bladder cancer patients than that of healthy controls, suggesting
that the HGF DATE plays a role in the carcinogenesis of
bladder cancer. Ma et al reported that the frequency of
healthy individuals with a short DATE, defined as 25 or fewer
deoxyadenosine repeats, was 26% in African Americans and 3% in
Caucasians (14). In our study,
because the frequency of healthy controls having 25 or fewer
repeats of the DATE was only 1% and the median was 28 repeats among
all subjects, we defined 27 or fewer deoxyadenosine repeats as the
short DATE. Based on our criterion, the frequency of Japanese
healthy controls having the short DATE was 24.2% (23/95). It is
interesting to note that there is an ethnic difference in the
distribution of the length of the HGF DATE. A distinct role
of the HGF DATE in the carcinogenesis of bladder cancer
requires validation in an epidemiological study designed to assess
gene-environment interaction in a large population. Additionally,
our study showed that the DATE was a frequent target for somatic
mutation in bladder cancer and the frequency of the short DATE in
bladder tumor tissues was significantly higher than that in matched
PBLs. The results suggested that the somatic mutational shortening
of the DATE is possibly involved in the genesis and progression of
bladder cancer. The results may further strengthen the significance
of the DATE in bladder cancer carcinogenesis.
A previous study demonstrated that HGF protein
expression in both breast cancer and normal epithelium was
significantly increased with decreased length of the HGF
DATE (14). In our study, the
tumor grade was significantly higher in bladder tumor tissues with
a short DATE than those without a short DATE (Table II). Furthermore, HGF mRNA
expression in bladder tumor tissues with the short DATE was
significantly higher than in those without the short DATE (Fig. 2C). Our results were consistent with
a report by Ma et al in which HGF mRNA expression in
human breast cancer tissues increased with decreased length of the
DATE (14). Our study also
demonstrated that MIT showed significantly higher HGF mRNA
expression levels than NMIT, and high grade tumors showed
significantly higher HGF mRNA expression levels than low
grade tumors. In a previous study, HGF was reported to stimulate
invasion of tumor cells and to induce angiogenesis in vivo
(1). It was also reported that HGF
signaling might be involved in tumor progression and invasion by
directly regulating the transcription of downstream functional
molecules (1,2). Taken together, the short DATE is
suggested to be associated with higher malignant potential and
tumor progression by regulating the transcriptional activity of the
HGF gene in bladder cancer cells.
The association between prognosis and high serum
level of HGF was previously reported in several types of cancer,
and serum HGF was suggested to be a useful marker for
discriminating a malignant tumor from benign disease and as a
potential therapeutic target (3–5,10).
In our study, patients with high HGF expression in their bladder
cancer tissue had a significantly worse overall survival than those
with low HGF expression, and patients with high HGF expression
tended to have worse disease-free survival than those with low HGF
expression. These results suggest that high HGF expression in
bladder cancer tissue is a predictor of survival and recurrence
after radical treatment of bladder cancer.
The results of this study should be interpreted with
some caution. One limitation was that we genotyped only the length
of deoxyadenosine repeat in the shorter DATE allele because the
number of repeats in the longer DATE allele could not be clearly
defined owing to the use of a direct sequencing method. According
to a previous study, however, only the presence of shorter
deoxyadenosine repeats has clinical significance and longer
deoxyadenosine repeats may be ignored in the analysis (14). Second, our studies indicated a
significant relationship between HGF protein expression and the
clinical outcome of patients with bladder cancer in
immunohistochemical analysis. However, we failed to genotype the
HGF DATE in the same patients group using paraffin-embedded
tumor tissues obtained by radical cystectomy, and could not
directly analyze the relationship between the short DATE and the
prognosis of bladder cancer patients. Further studies are required
to clarify whether the short DATE may be a prognostic marker for
invasive bladder cancer.
In conclusion, our results suggested that the DATE
in the HGF promoter region is associated with carcinogenesis
and aggressiveness of bladder cancer by aberrantly activating HGF
expression. Further investigation is warranted to evaluate the
clinical usefulness of the short DATE as a marker of poor prognosis
in invasive bladder cancer patients.
Acknowledgements
The authors are grateful to Ms. Yoko
Mitobe and Ms. Yuka Izumida (Department of Urology, Akita
University Graduate School of Medicine, Akita, Japan) for their
excellent technical assistance.
References
|
1.
|
Trusolino L and Comoglio PM:
Scatter-factor and semaphorin receptors: cell signalling for
invasive growth. Nat Rev Cancer. 2:289–300. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
2.
|
Birchmeier C, Birchmeier W, Gherardi E and
Vande Woude GF: Met, metastasis, motility and more. Nat Rev Mol
Cell Biol. 4:915–925. 2003. View
Article : Google Scholar : PubMed/NCBI
|
|
3.
|
Aune G, Lian AM, Tingulstad S, Torp SH,
Forsmo S, Reseland JE, Stunes AK and Syversen U: Increased
circulating hepatocyte growth factor (HGF): a marker of epithelial
ovarian cancer and an indicator of poor prognosis. Gynecol Oncol.
121:402–406. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4.
|
Gohji K, Nomi M, Niitani Y, Kitazawa S,
Fujii A, Katsuoka Y and Nakajima M: Independent prognostic value of
serum hepatocyte growth factor in bladder cancer. J Clin Oncol.
18:2963–2971. 2000.PubMed/NCBI
|
|
5.
|
Saigusa S, Toiyama Y, Tanaka K, Yokoe T,
Fujikawa H, Matsushita K, Okugawa Y, Inoue Y, Uchida K, Mohri Y and
Kusunoki M: Inhibition of HGF/cMET expression prevents distant
recurrence of rectal cancer after preoperative chemoradiotherapy.
Int J Oncol. 40:583–591. 2012.PubMed/NCBI
|
|
6.
|
Wu CW, Li AF, Chi CW, Chung WW, Liu TY,
Lui WY and P’Eng FK: Hepatocyte growth factor and Met/HGF receptors
in patients with gastric adenocarcinoma. Oncol Rep. 5:817–822.
1998.PubMed/NCBI
|
|
7.
|
Ueki T, Fujimoto J, Suzuki T, Yamamoto H
and Okamoto E: Expression of hepatocyte growth factor and its
receptor c-met proto-oncogene in hepatocellular carcinoma.
Hepatology. 25:862–866. 1997. View Article : Google Scholar
|
|
8.
|
Yamashita J, Ogawa M, Yamashita S, Nomura
K, Kuramoto M, Saishoji T and Shin S: Immunoreactive hepatocyte
growth factor is a strong and independent predictor of recurrence
and survival in human breast cancer. Cancer Res. 54:1630–1633.
1994.PubMed/NCBI
|
|
9.
|
Bharti A, Ma PC, Maulik G, Singh R, Khan
E, Skarin AT and Salgia R: Haptoglobin alpha-subunit and hepatocyte
growth factor can potentially serve as serum tumor biomarkers in
small cell lung cancer. Anticancer Res. 24:1031–1038.
2004.PubMed/NCBI
|
|
10.
|
Coskun U, Bukan N, Sancak B, Gunel N,
Ozenirler S, Unal A and Yucel A: Serum hepatocyte growth factor and
interleukin-6 levels can distinguish patients with primary or
metastatic liver tumors from those with benign liver lesions.
Neoplasma. 51:209–213. 2004.PubMed/NCBI
|
|
11.
|
Toiyama Y, Miki C, Inoue Y, Okugawa Y,
Tanaka K and Kusunoki M: Serum hepatocyte growth factor as a
prognostic marker for stage II or III colorectal cancer patients.
Int J Cancer. 125:1657–1662. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12.
|
Tanimoto S, Fukumori T, El-Moula G,
Shiirevnyamba A, Kinouchi S, Koizumi T, Nakanishi R, Yamamoto Y,
Taue R, Yamaguchi K, Nakatsuji H, Kishimoto T, et al: Prognostic
significance of serum hepatocyte growth factor in clear cell renal
cell carcinoma: comparison with serum vascular endothelial growth
factor. J Med Invest. 55:106–111. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13.
|
Chau GY, Lui WY, Chi CW, Chau YP, Li AF,
Kao HL and Wu CW: Significance of serum hepatocyte growth factor
levels in patients with hepatocellular carcinoma undergoing hepatic
resection. Eur J Surg Oncol. 34:333–338. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
14.
|
Ma J, DeFrances MC, Zou C, Johnson C,
Ferrell R and Zarnegar R: Somatic mutation and functional
polymorphism of a novel regulatory element in the HGF gene promoter
causes its aberrant expression in human breast cancer. J Clin
Invest. 119:478–491. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
15.
|
Shimizu M, Mori T, Sakurai T and Shindo H:
Destabilization of nucleosomes by an unusual DNA conformation
adopted by poly(dA) small middle dotpoly(dT) tracts in vivo. EMBO
J. 19:3358–3365. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
16.
|
Koch KA and Thiele DJ: Functional analysis
of a homopolymeric (dA-dT) element that provides nucleosomal access
to yeast and mammalian transcription factors. J Biol Chem.
274:23752–23760. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
17.
|
Fox KR: Wrapping of genomic polydA.polydT
tracts around nucleosome core particles. Nucleic Acids Res.
20:1235–1242. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
18.
|
Iyer V and Struhl K: Poly(dA:dT), a
ubiquitous promoter element that stimulates transcription via its
intrinsic DNA structure. EMBO J. 14:2570–2579. 1995.PubMed/NCBI
|
|
19.
|
Anderson JD and Widom J: Poly(dA-dT)
promoter elements increase the equilibrium accessibility of
nucleosomal DNA target sites. Mol Cell Biol. 21:3830–3839. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
20.
|
Reardon BJ, Winters RS, Gordon D and
Winter E: A peptide motif that recognizes A.T tracts in DNA. Proc
Natl Acad Sci USA. 90:11327–11331. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
21.
|
Graziano F, Galluccio N, Lorenzini P,
Ruzzo A, Canestrari E, D’Emidio S, Catalano V, Sisti V, Ligorio C,
Andreoni F, Rulli E, Di Oto E, et al: Genetic activation of the MET
pathway and prognosis of patients with high-risk, radically
resected gastric cancer. J Clin Oncol. 29:4789–4795. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
22.
|
Lopez-Beltran A and Montironi R:
Non-invasive urothelial neoplasms: according to the most recent WHO
classification. Eur Urol. 46:170–176. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
23.
|
Edge SB, Byrd DR, Compton CC, Fritz AG,
Greene FL and Trotti A: Urinary bladder. AJCC Cancer Staging
Manual. 7th edition. Springer; New York, NY: pp. 497–505. 2010
|