Introduction
Optical microscopy (OM) of tissues has developed in
various ways leading to great discoveries and advances in biology
and medicine (1,2). Tissue samples are either prepared by
time-consuming paraffin embedding at room temperature (RT) and cut,
or quickly frozen and cut at cryo-temperatures. Paraffin-embedding
is widely used in basic biological studies, applied biology fields
and medicine. In the quick cryo-sectioning method, excised tissues
are frozen in OCT compound, cryo-sectioned at a thickness of 3–5
μm, fixed, stained by hematoxylin-eosin (H&E) and inspected
during surgery performed to remove tissue suffering from cancer
invasion and metastasis (3,4). The
size of cell nuclei is the most significant indicator used for the
intraoperative cancer diagnosis, the nucleus of cancer cells
generally being larger than that of normal cells. However,
cryo-thin-sectioning is quite difficult and takes ~15–30 min for
each sample, thus, the procedure can prolong the whole operation,
which is a burden for patients.
The surface level of tissue blocks excised from
various organs (heart and kidney) (5), kidney (6), brain tumor (7,8),
liver, sciatic nerve, spinal cord (9), colon (10) has been successfully observed in an
environmental capsule by scanning electron microscopy (SEM).
Environmental capsules allow ‘wet’ samples to be examined in the
vacuum of the microscope, avoiding dehydration artifacts. However,
the samples are not accessible for additional staining or
manipulation. The Atmospheric SEM (ASEM) ClairScope™ was developed
to realize SEM of a sample at atmospheric pressure in a
readily-accessible, open container (ASEM dish) (11). The microscope has an inverted SEM
configuration, the column being closed by the ASEM dish at the top.
An optical microscope (OM) positioned above the ASEM dish can be
used to observe wide areas of the sample, and the SEM to image
specified smaller regions through a thin SiN film in the base of
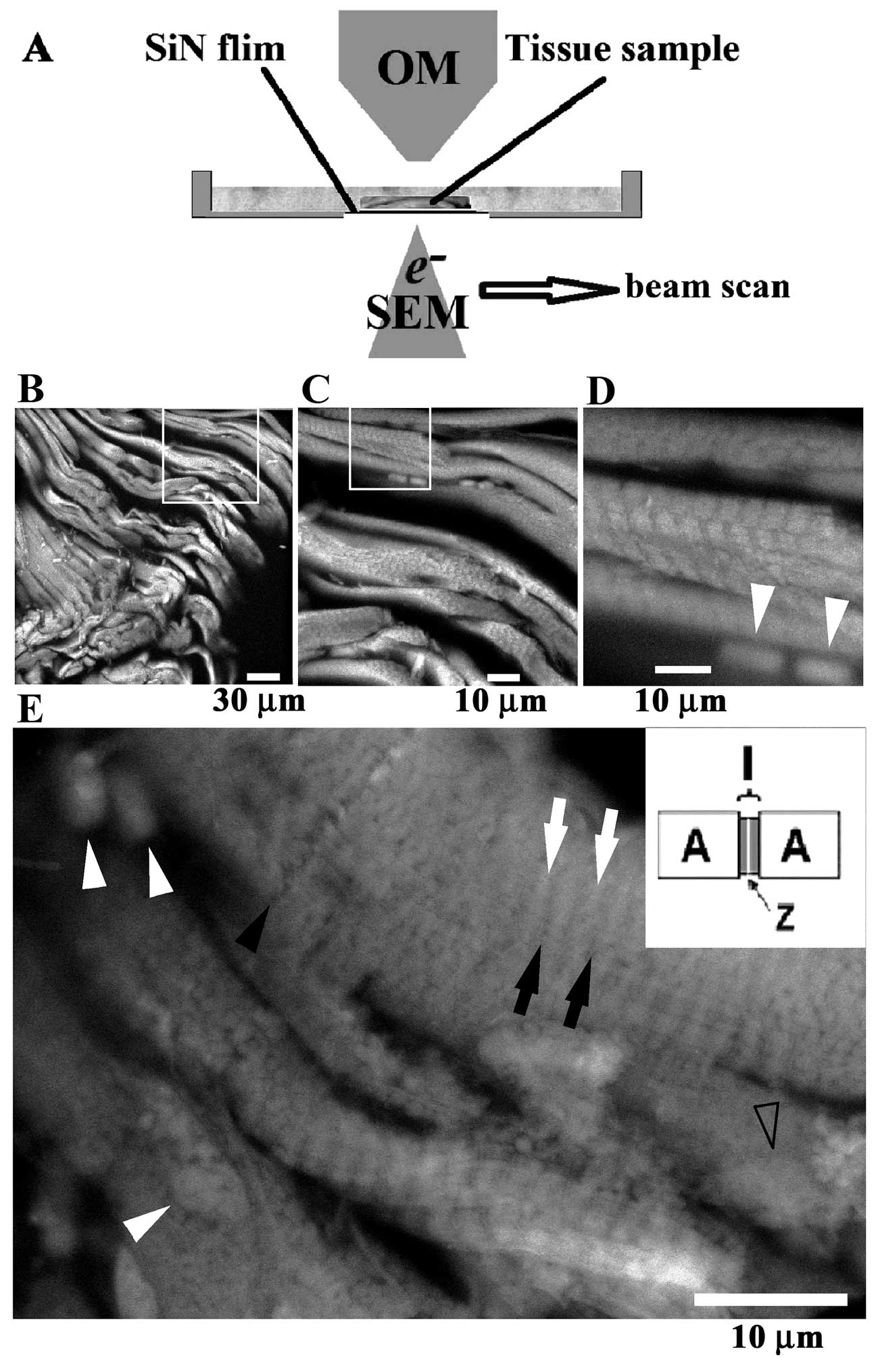
the dish (Fig. 1A). The optical
axes of both microscopes are aligned and fixed to ensure that
correlative images are recorded, and the specimen stage can move
two-dimensionally (x–y) for targeting. The specimen depth
observable by ASEM is 2–3 μm at 30 kV (12,13)
and the resolution is 8 nm near the SiN membrane, which is
advantageous for the observation of fine subcellular structures.
The detachable 35-mm ASEM dish allows the culture of various
primary cells and subsequent efficient staining-washing cycles
(12,14–16).
We have cultured primary cells sampled from various mammalian
(12,14) and insect organs (14,15),
including mouse hippocampus (12),
cortex and cerebellum (14), and
from prokaryotes, including coccus, bacillus (15) and mycoplasma (17). Cultured cells were successfully
stained with different heavy metals and imaged (11). Because the epitopes of the cells
were preserved in the aqueous solution, we were also able to
successfully immunolabel different cells with various antibodies,
including mouse monoclonal antibodies (12,14,15,17–20).
Furthermore, we recently used ASEM to observe TI-Blue-stained
tissue from a gold fish brain, as briefly outlined in the study of
Nishiyama et al (11)
(Fig. 9).
In the present study, we report the observation of
various mouse tissues by ASEM after staining with heavy metals, and
indicate the possible use of ASEM for the diagnosis of cancer and
kidney diseases. Staining with phosphotungstic acid (PTA) or
platinum blue (Pt-blue) made cell nuclei very prominent, while
staining with uranyl acetate (UA) or by a modified NCMIR method
clearly revealed cell outlines, organelles and extracellular
structures. The difference between normal and cancer-metastasized
tissue was successfully visualized for lung and spinal cord
specimens. Importantly, the area observable by ASEM was enlarged
both by the use of a new multi-windowed sample dish and by
exploiting the open ASEM dish configuration to slide tissue across
the windows.
Materials and methods
Animals
Six to fifteen week-old male and female ICR mice and
8–12-week-old female BALB/c mice (Japan Clea, Tokyo, Japan) were
sacrificed to obtain normal tissue for observation. Tumor bearing
mice were obtained 10 days after the intravenous injection of
4T1E/M3 breast cancer cells (1×106/mouse) or 30 days
after the subcutaneous injection of the cancer cells
(1×106/mouse) to 8-week-old female BALB/c mice (21–23).
The animal studies were in compliance with the national institute
rules of conduct and adhered to the principles of Institutional
Animal Care and Use Committee Guidebook. All experiments were
approved by the Animal Care and Use Committee of the National
Institute of Advanced Industrial Science and Technology (AIST).
Tissue sample preparation
Animals were anesthetized using isoflurane (Abbott,
Maidenhead, UK) and sacrificed by intracardiac perfusion of 4%
paraformaldehyde (PFA; Wako Pure Chemicals, Osaka, Japan) in
phosphate-buffered saline (PBS, pH 7.4) (24). Tissues were either cut with a
scalpel to obtain 1–2-mm thick tissue slabs, or sliced with a PRO7
linear slicer (Dosaka, Kyoto, Japan) to obtain 100–200 μm thick
slabs. Samples were washed several times in PBS, and further fixed
with 4% PFA or 1% glutaraldehyde (GA) (Nisshin EM, Tokyo, Japan) or
with a solution containing both fixatives, for 30 min at RT; the
fixative volume was 15–20 times the sample volume.
Histology for OM
Lungs and spines harvested from tumor bearing mice
were fixed in 10% neutral buffered formalin (Wako Pure Chemicals)
at RT. The spines were then decalcified in decalcifying solution A
(Wako Pure Chemicals) following the manufacturer’s protocol. Fixed
lungs, spleens or decalcified fixed spines were then embedded in
paraffin, cut into 5–6 μm sections, stained with H&E and
inspected and photographed by OM.
Heavy metal staining
PTA or Pt-blue (TI-Blue; Nisshin EM), 6% solution;
Pt4(NH3)8(C6H13O5)4)
staining: the fixed tissues were perforated with 0.5% Triton X-100
(MP Biomedicals, LLC, Illkirch, France) in PBS at RT for 15 min,
and washed with DDW twice. They were stained with 2% PTA (TAAB
Laboratories Equipment Ltd., Aldermaston, UK) or 0.6% TI-Blue or
double stained with both, for 3 h at RT or overnight at 4°C.
UA staining: the tissues were incubated in filtered
1% tannic acid (Polysciences Inc., Warrington, FL, USA), 1% GA for
30 min at RT, and washed with DDW twice. They were stained with
filtered 2% uranyl acetate (UA) for 30 min at RT.
Quick method: without prior perforation, fixed
tissues were incubated with 2% UA for 15 min at RT.
NCMIR staining method: tissues were stained using a
slight modification of the NCMIR method developed by the Ellisman
group (25). The fixed tissues
were perforated with 0.5% Triton X-100 in PBS at RT for 15 min,
washed with DDW, and further fixed with 2.5% GA, 2% PFA in filtered
0.1 M phosphate buffer (PB, pH 7.4) containing 2 mM
CaCl2 at RT for 15 min. They were then washed using
filtered 0.15 M PB containing 2 mM CaCl2, and further
fixed/stained with the same buffer supplemented with 1.5% potassium
ferricyanide (Sigma-Aldrich, St. Louis, MO, USA), 2% aqueous osmium
tetroxide (OsO4) (Nisshin EM) at RT for 60 min. After
washing with DDW, tissues were then incubated with filtered 1%
thiocarbohydrazide (TCH; Tokyo Chemical Industry, Co., Ltd., Tokyo,
Japan) (25) at RT for 20 min,
rinsed with DDW, further stained with 2% aqueous OsO4 at
RT for 30 min, rinsed with DDW, stained with 2% UA in DDW and kept
at 4°C overnight. Finally, after rinsing with DDW, the tissue
samples were stained with 0.4% lead citrate (TAAB Laboratories
Equipment) at RT for 2 min.
Once fully stained, all samples were placed ‘cut’
face down in the ASEM dish immersed in radical scavenger solution
and imaged as soon as possible by OM and SEM, as the contrast of
the most stains fades on storage.
ASEM imaging
The ClairScope ASEM system (JASM-6200; JEOL Ltd.,
Tokyo, Japan) was used to record SEM images (11). Unless otherwise stated, the
standard 35-mm ASEM dish was used; the 100-nm thick (250×250)-μm
SiN film window built into its base separates the sample immersed
in aqueous liquid from the vacuum inside the SEM column. A
newly-developed dish, with eight 100-nm thick (250×250)-μm SiN film
windows was employed to observe larger areas. After fixation and
staining, tissues were placed on the SiN film window, and observed
in 10 mg/ml (w/v) D-glucose (Dextrose; MP Biomedicals LLC) or
ascorbic acid (L-Ascorbic acid; Sigma-Aldrich) in DDW, using the
inverted SEM of the ASEM. When the uneven surface of tissue
prevented it from attaching well to the film, 150–250 Pascal
pressure was applied during imaging. To achieve this a cover glass
was positioned on the upper surface of the tissue and loaded with a
lead weight. The acceleration voltage of the SEM was 30 kV,
electrons backscattered from the cells were recorded for
visualization. As the stain scatters more electrons than tissue,
more heavily stained regions are bright in the ASEM images.
Sliding a stained tissue slab across the
ASEM dish
A stained tissue slab was placed on the
multi-windowed ASEM dish and imaged in 10 mg/ml (w/v) D-glucose or
ascorbic acid in DDW from underneath by the inverted SEM. The
tissue was then pushed slightly to the side with tweezers under
monitoring by OM from above. The tissue was again imaged by SEM,
and the images before and after the movement were merged.
Results
As detailed below, ASEM was successfully employed to
image a range of mouse tissue samples. In each case tissue
slabs were stained with heavy metal solution, laid cut face down on
the SiN film of the ASEM dish, and observed in liquid by ASEM (see
Materials and methods).
Cardiac muscles
Hearts of mice sacrificed by intracardiac perfusion
of 4% PFA in PBS were dissected into 1- to 2-mm slabs, and either
further fixed with PFA alone or doubly with PFA and GA in PBS.
After perforation with Triton X-100, each tissue slab was stained
with PTA and imaged by ASEM (Fig.
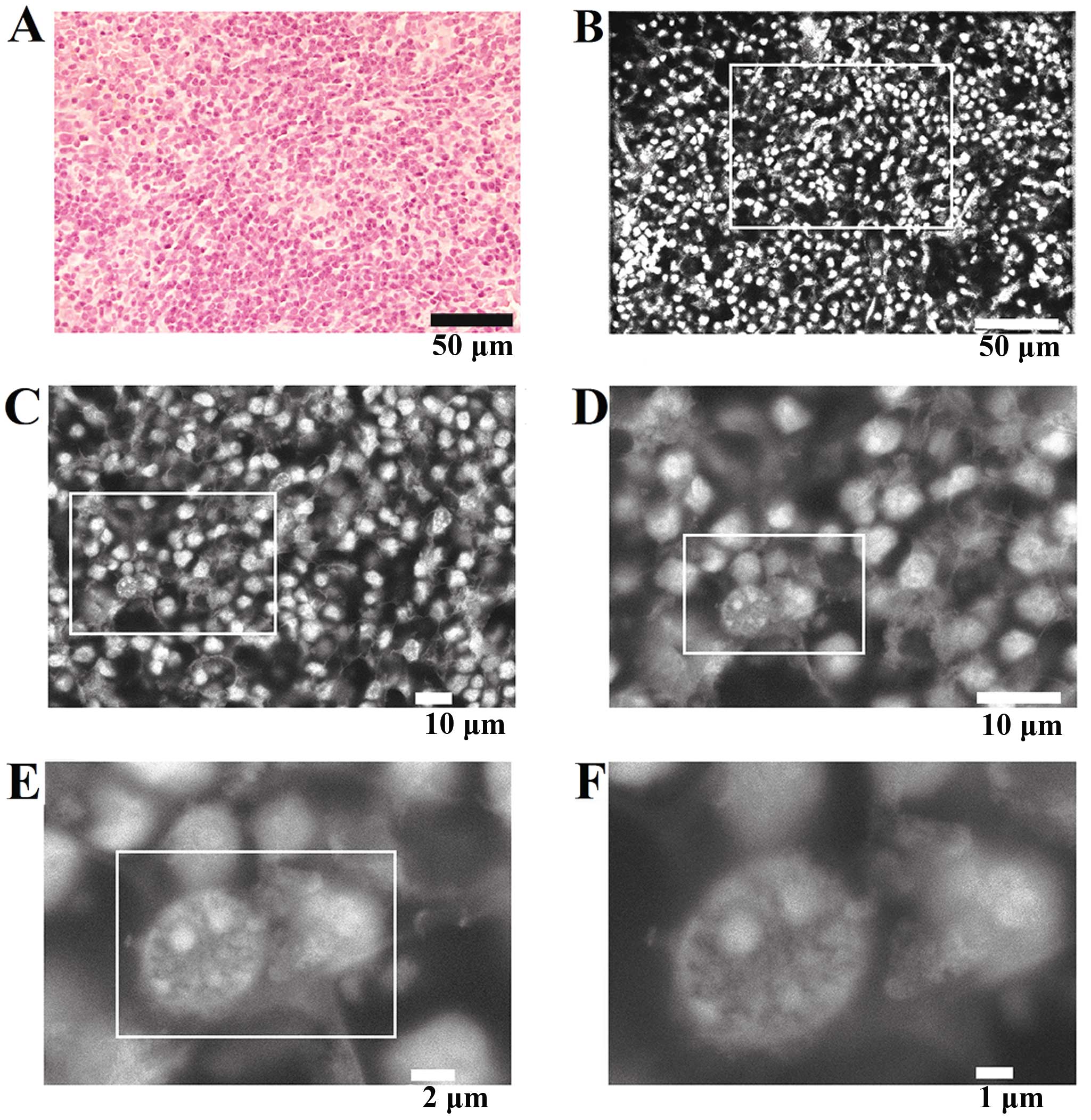
1A). Cardiomyocytes were evident (Fig. 1B–E) and their more highly stained
nuclei were observed. The nuclei of what are presumably surrounding
cells, possibly attached fibroblasts, were observed as blurred
white ellipses (Fig. 1D and E).
This blurring is probably attributable to the distant location of
the nuclei from the supporting SiN film, as demonstrated earlier
for gold particles (11,15). Intercalated discs could be clearly
distinguished. The alternating dark and bright bands imaged by ASEM
can be interpreted as thin dark zones (I-bands) and broad bright
zones (A-bands) of myofibril sarcomeres. Z-lines were also faintly
imaged, each appearing as a thin white line in the center of a dark
I-band.
Neural tissues
Nerve tissues from ICR mice were stained with PTA
and similarly imaged by ASEM. Observation of the surface level of
the tissues showed that neurons extend their nerve fibers to form
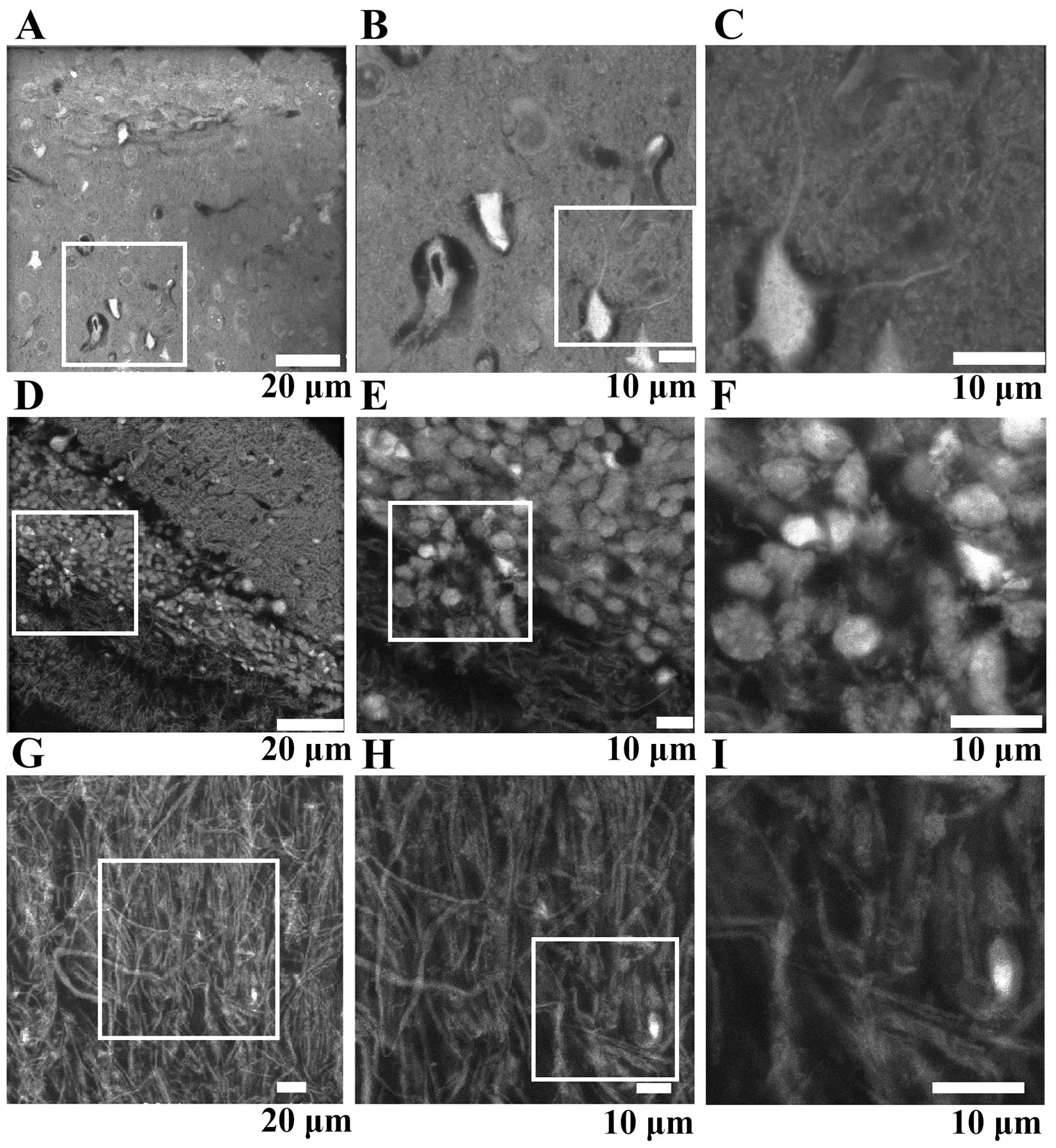
fine networks surrounded by glia cells (Fig. 2). Imaging the cerebrum revealed a
highly ordered structure of cells and a systematic network of
neurons including delicate connections (Fig. 2A–C). In the cerebellum, three
clearly different layers were observed (Fig. 2D), presumably the molecular layer,
the granular layer and cerebral white matter. Again, neurons were
connected by delicate systematic networks formed by nerve fibers
(Fig. 2E and F). At higher
magnification, nuclei were seen to include brightly stained patches
that were presumably chromatin and nucleoli (Fig. 2F).
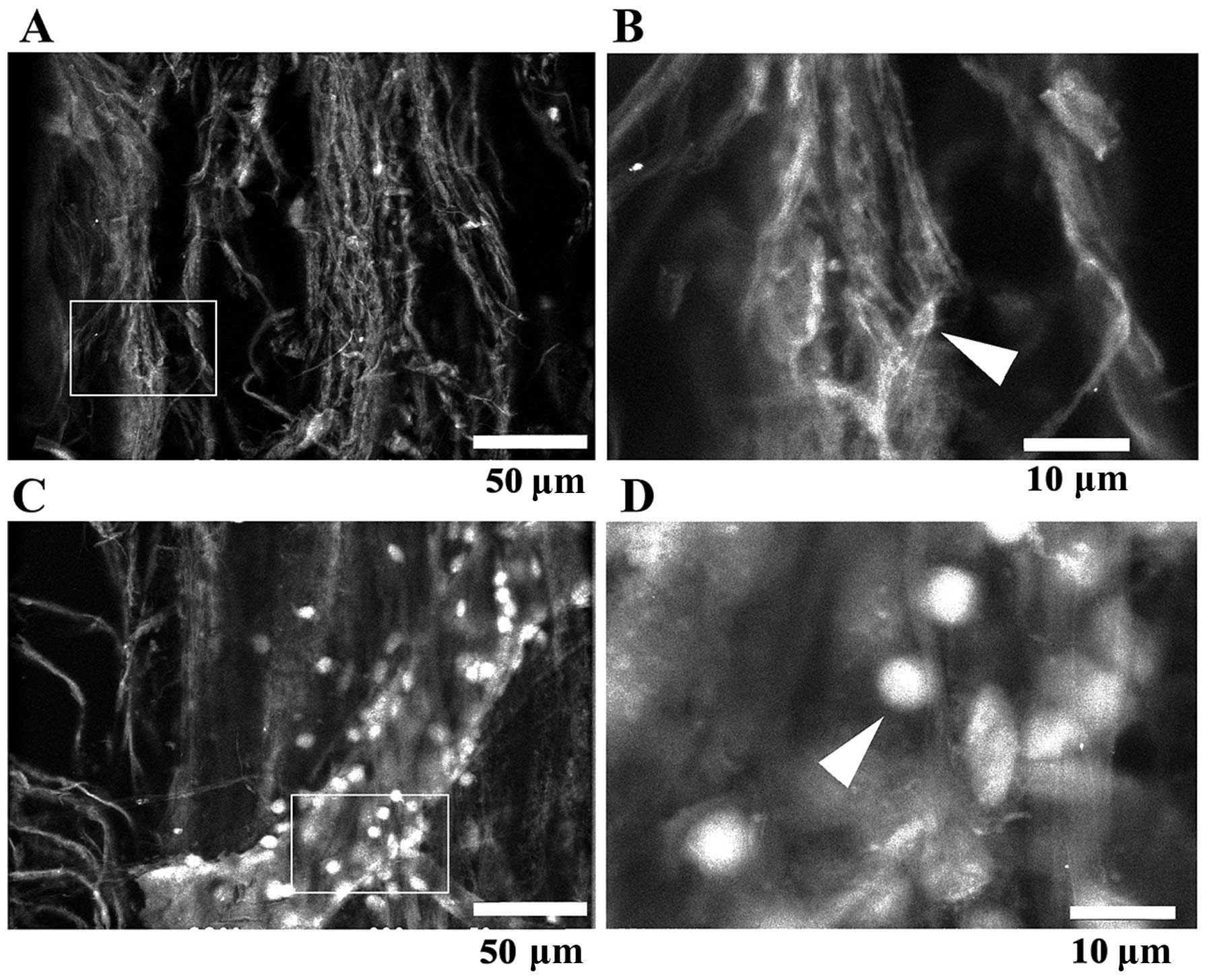
The spinal cord contains a larger number of fibrous
structures than the brain. Imaging vertical sections of spinal cord
by ASEM revealed 200 nm - 3 μm thick, almost perpendicular filament
‘strings’ together with a lower number of filaments running in
various directions, overall forming a complex network. Bright
nuclei were sometimes observed at the end of the strings (Fig. 2G–I).
Liver
When liver tissue slabs were stained with UA,
hepatocytes, erythrocytes and probably sinusoids in the surface
layers were visualized by ASEM (Fig.
3A–C). The nuclei of hepatocytes were also sometimes
distinguishable (Fig. 3C). When
other tissue slabs were stained with PTA and imaged, collagen
fibers and fat droplet-like structures were also prominent
(Fig. 3D). However, it is not easy
to evaluate the staining preferences of UA and PTA using different
samples.
Kidney
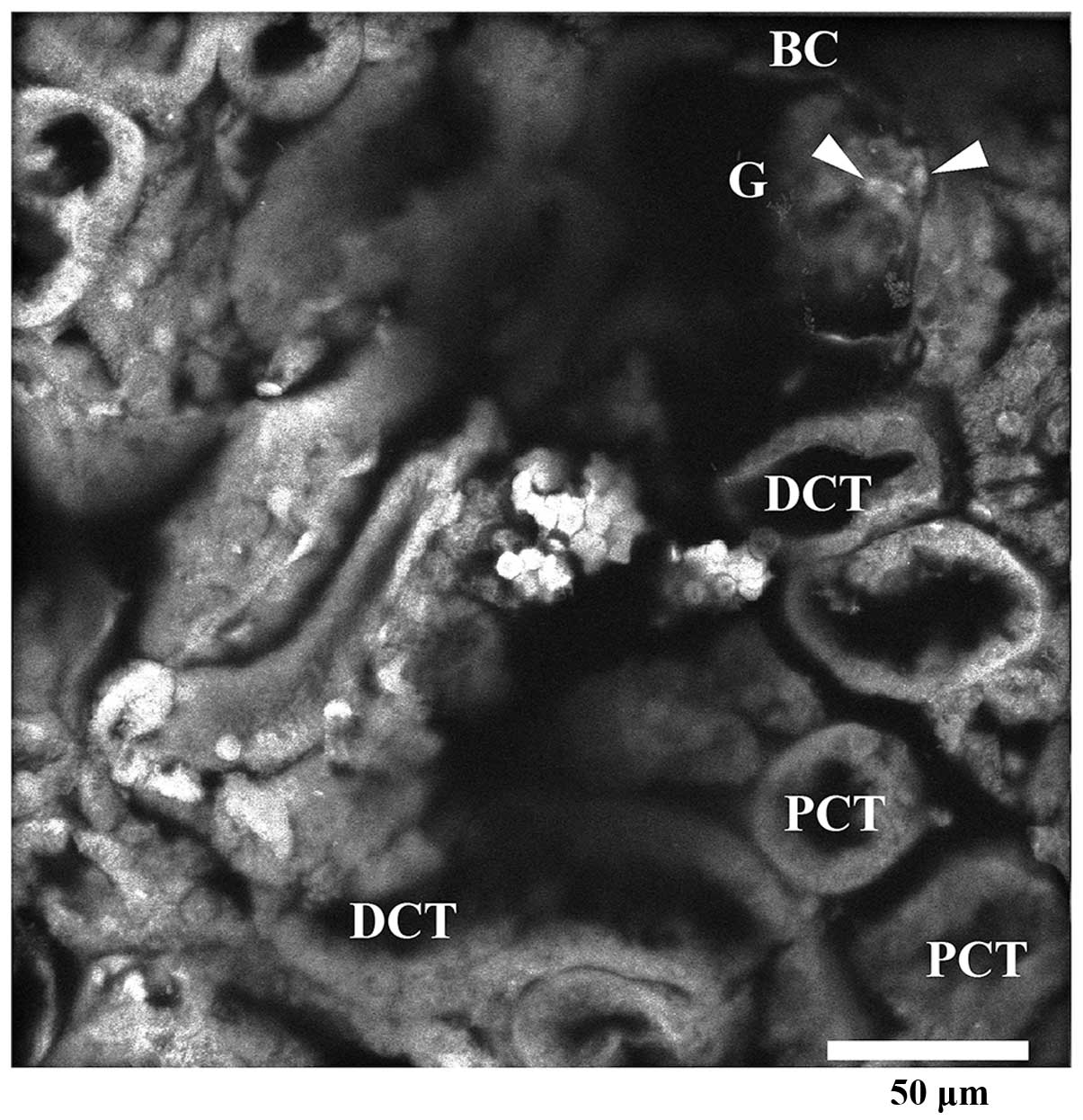
A slab excised from a mouse kidney was imaged by
ASEM without any pretreatment other than fixation, permeation and
PTA staining (Fig. 4). Bowman’s
capsules, glomeruluses, proximal or distal convoluted tubules and
also podocytes were visualized (Fig.
4). The overviews are consistent with images obtained when
traditional Epon-thin sections of kidney biopsies are inspected by
transmission electron microscopy (TEM). Importantly, sample
preparation took much less time, indicating the potential of the
ASEM method for quick diagnosis or to rapidly obtain a second
opinion. Accelerating the sample preparation further by omitting
the perforation step (see Materials and methods; quick method),
delivered images of basically the same quality (data not shown) as
those obtained by the standard ASEM staining protocol including
perforation.
Skeletal muscle
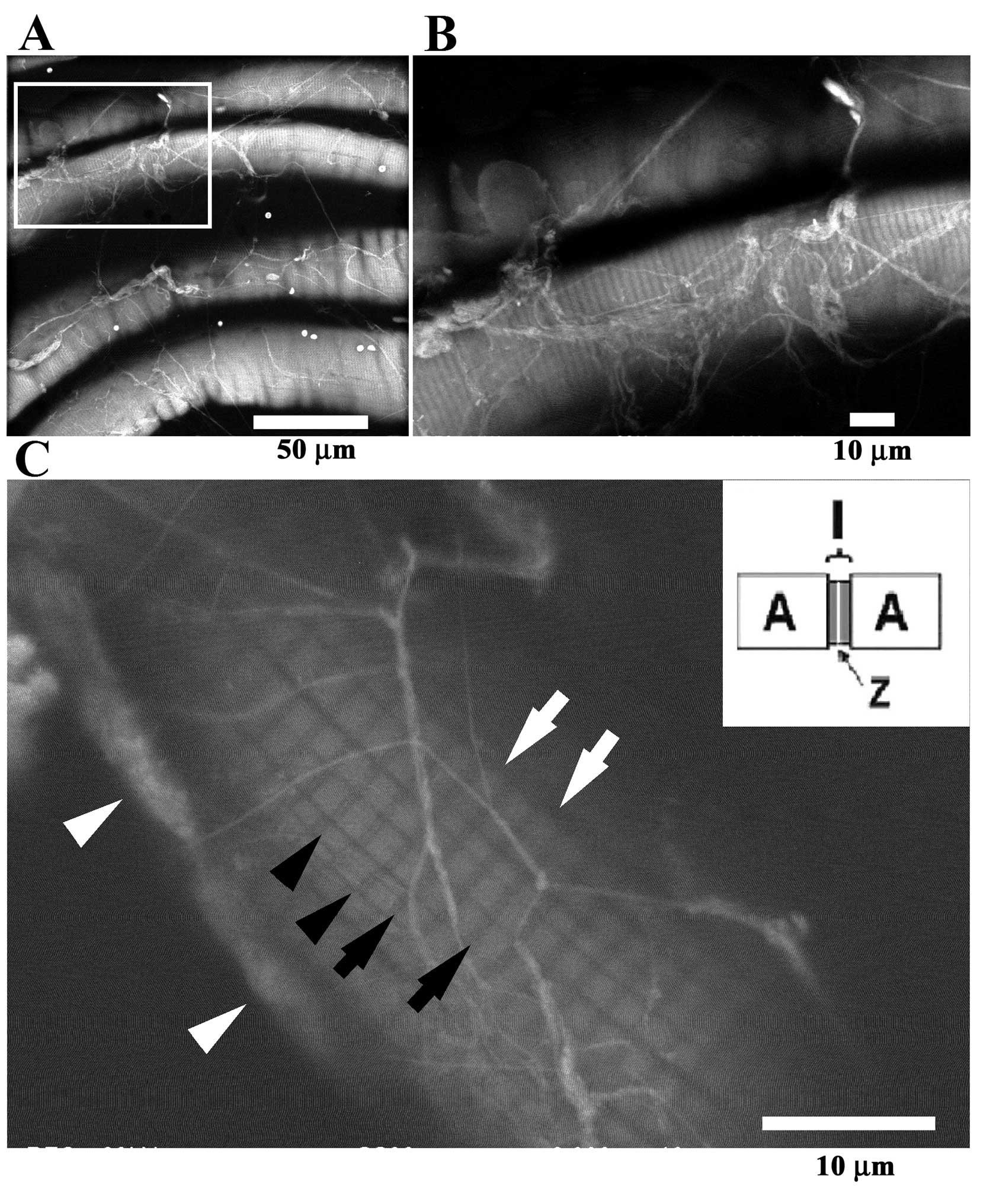
Muscle fibers were clearly visible when PTA-stained
tissue slabs of gastrocnemius muscle were observed by ASEM
(Fig. 5). The myocytes were found
to be surrounded by filaments (Fig. 5A
and B). In myocytes, I-bands and A-bands were visualized as
dark zones and broader bright zones, respectively, and Z-lines were
frequently distinguished as a fine thin line in the center of dark
I-bands (Fig. 5C). These results
suggest that myocytes can be easily imaged at high resolution using
the ASEM.
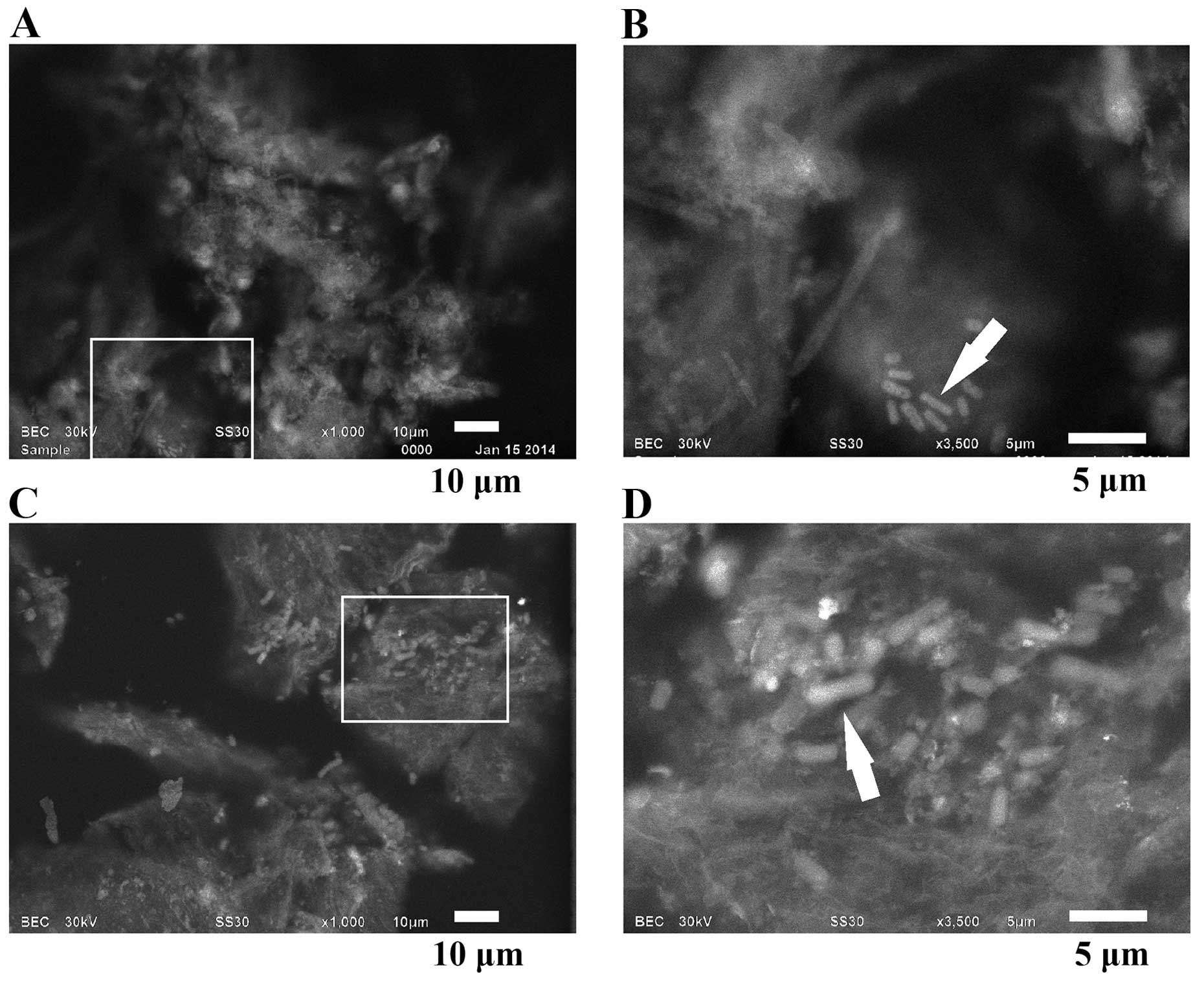
Stomach
Mucosa of the stomach lumen were observed together
with symbiotic bacteria (Fig. 6A and
B); a bacteria colony can be seen in Fig. 6B (arrow). Staining by the modified
NCMIR method instead of with PTA, delivered clear images of the
symbiotic bacteria revealing their different shapes (Fig. 6C and D).
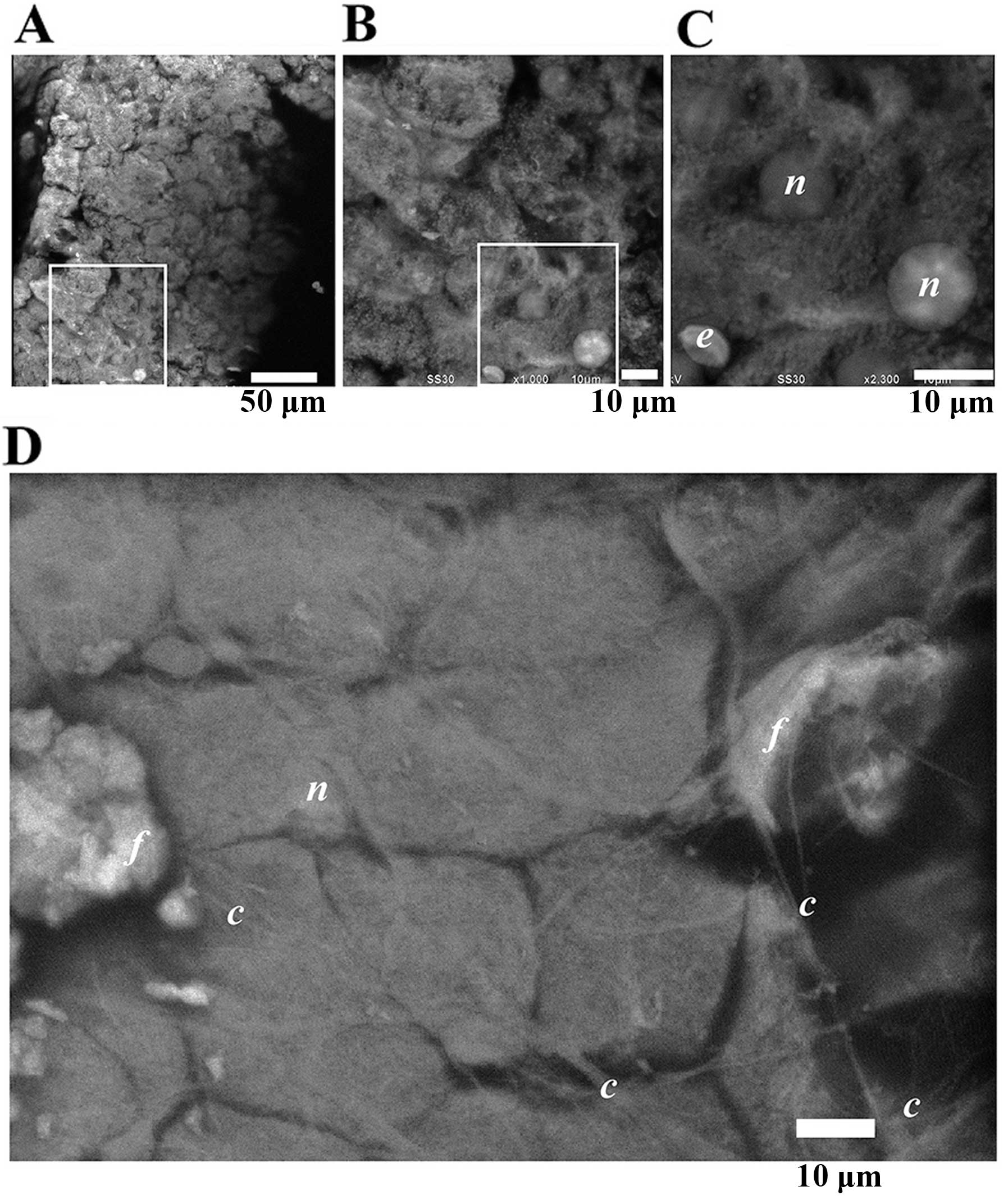
Spleen
H&E-stained thin-slices of spleen were observed
by OM, revealing a large number of round cells with and without
filopodia, presumably hematopoietic cells (Fig. 7A). The surface layers of
independently cut, thicker slabs of spleen tissue stained with PTA
looked similar when imaged at low magnification by ASEM (Fig. 7B). In a typical area there were
many differently shaped cells. From their shapes and nuclei, some
of them were probably lymphocytes, e.g., B cells, T cells or plasma
cells (Fig. 7C–F). At higher
magnification, nuclei were found to have brightly stained patches,
presumably nucleoli and chromatin (Fig. 7F compared to Figs. 2F and 3C). Fibrous structures <2 μm in thick,
were also observed. These results suggest that ASEM can be employed
to study lymph cells, including their development and immunity in
the spleen.
Comparative observation of normal lung
and the lung metastasized by breast cancer
The lungs are known to be organs easily metastasized
by cancer. Lung tissues excised from normal mice and from mice with
tumors induced by the injection of breast cancer cells, were
stained with both Pt-blue and PTA and observed by ASEM. OM of
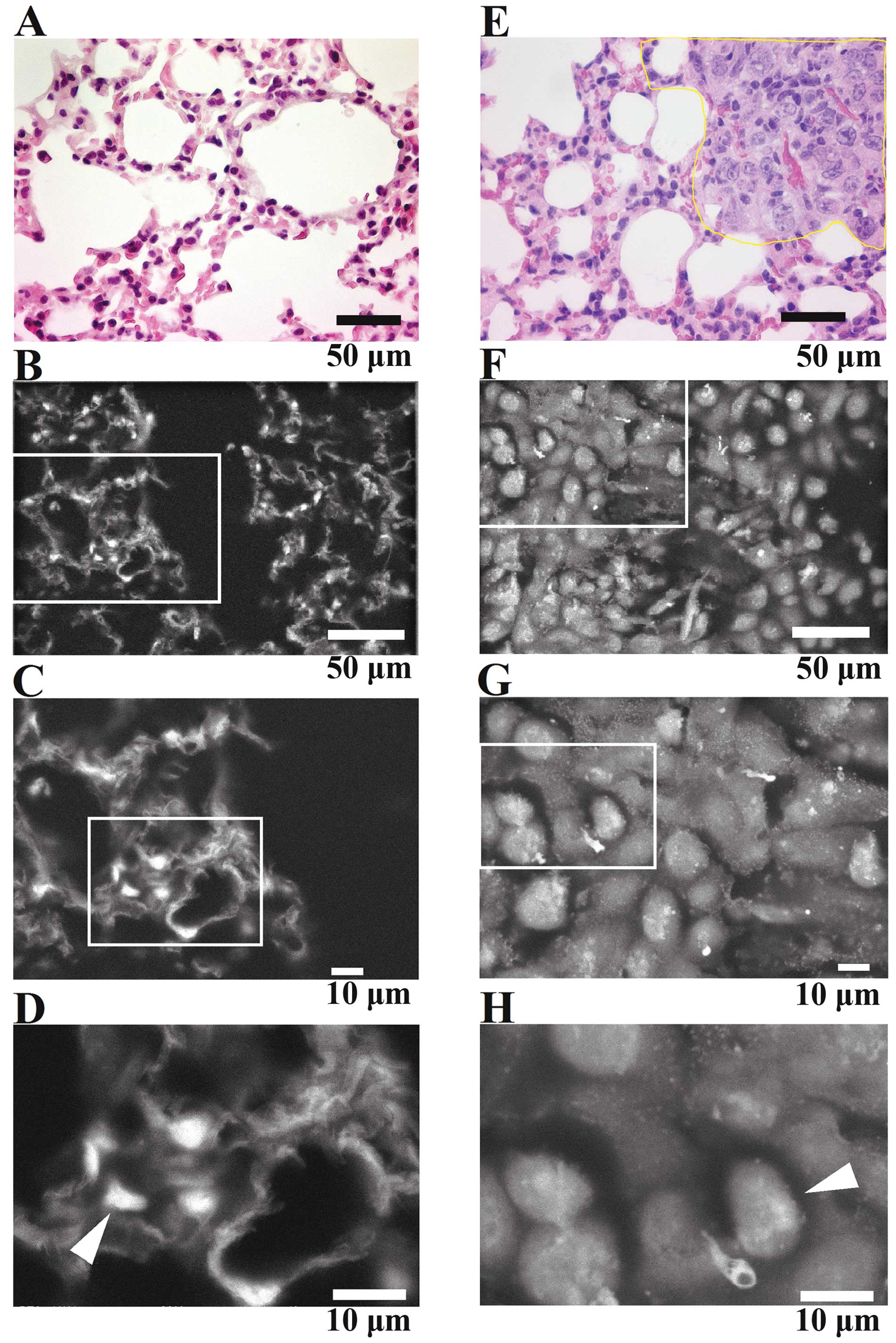
H&E-stained thin lung sections from other normal (Fig. 8A) and metastasized mice (Fig. 8E) was independently performed for
comparison. Observed by ASEM, the normal lung tissue showed typical
thin-wall structures with alveoli, alveolar ducts, a vein system
and trachea (Fig. 8B–D). In the
cells, nuclei close to the surface of the tissue slabs were
observed as bright features in the images. Their outlines were
clear, facilitating the measurement of their diameters. By
contrast, only faint traces of the regular alveoli and alveolar
ducts were discernable for metastasized lung (Fig. 8F–H); most of the alveolar system
had been replaced by differently shaped cells with larger nuclei,
i.e., breast cancer cells (Fig.
8H). These results are consistent with those obtained by
H&E staining and OM (Fig. 8A and
E), suggesting that ASEM could be used for cancer
diagnosis.
Normal spinal cord and the spinal cord
metastasized by breast cancer
Spinal cord excised from the mice with tumors
induced by the injection of breast cancer cells, were stained with
both Pt-blue and PTA and compared with those from the normal mice
using the ASEM. While nuclei seen at the end of the fibers in
normal spinal cord were all the correct size (Fig. 9A and B, arrowhead), a thin
film-like tissue of cells with unusually large nuclei was observed
in a metastasized spinal cord (Fig. 9C
and D, arrowhead). The large nuclei were similar in size to
those seen in lung metastasized with breast cancer cells (compare
Fig. 8E–H).
Wide area observation by sliding tissue
on the ASEM dish
To enlarge the area observable by ASEM, a spinal
cord slab stained with PTA was placed in a newly developed 8-window
ASEM dish and the tissue was repeatedly induced to slide very
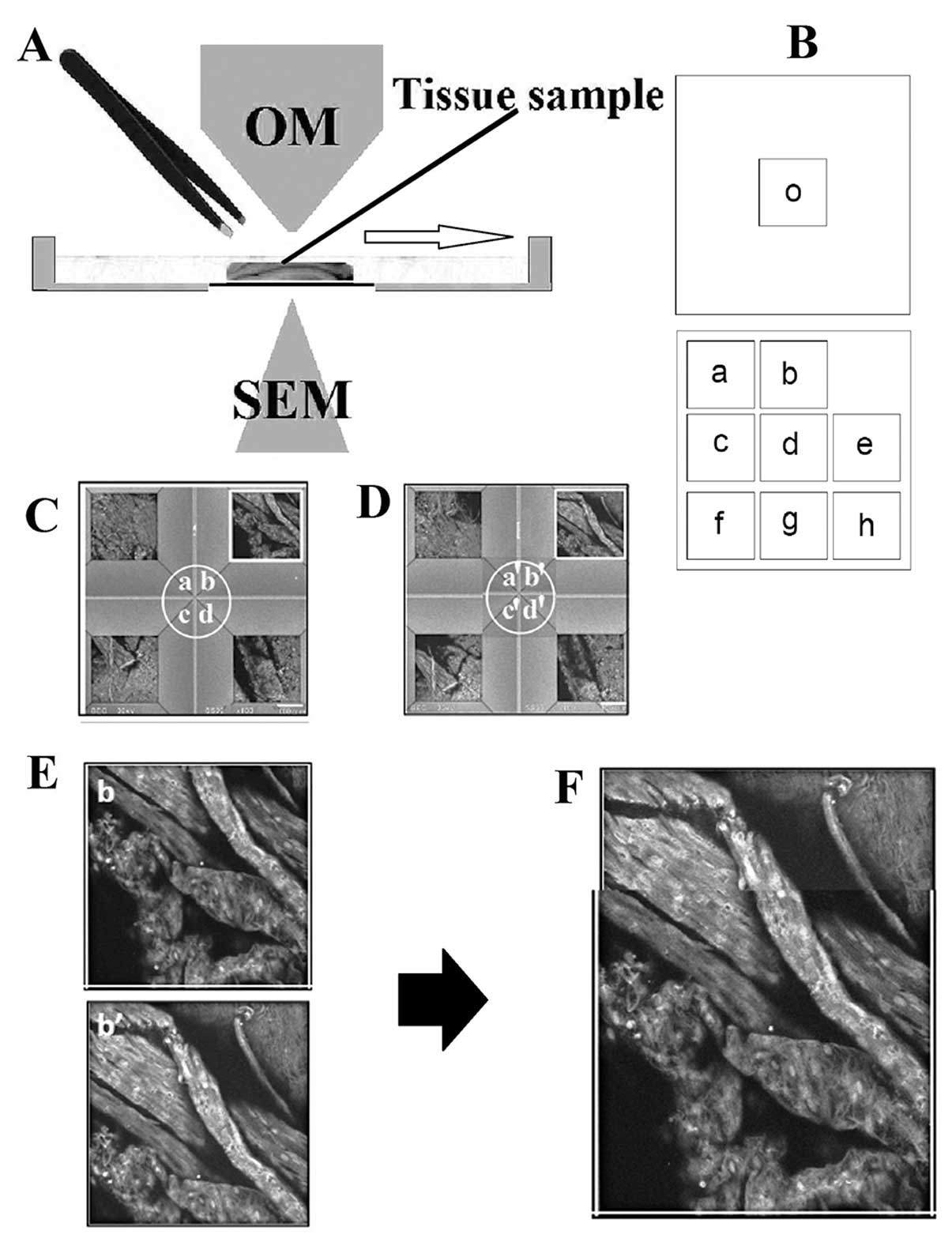
slightly across the thin membrane windows and imaged (Fig. 10A). Although the observable area
in each ASEM imaging step was restricted to the field of the 8
windows, two sequential images partly overlapped and could be
merged, covering a wider area of the spinal cord (Fig. 10C–F). The observable area of the
multi-windowed dish (Fig. 10B)
was thus successfully extended. The shift of the tissue on the flat
base of the dish (Fig. 10A) was
precisely monitored by OM from above. This was made possible by the
open ASEM dish configuration and the axis-aligned OM (Fig. 1A). In the future, the procedure
could be automated by constructing a micro motor-driven manipulator
to realize an accurate shift at regular time intervals synchronized
with ASEM scanning.
Discussion
SEM has proved very important to the study of
morphology, especially the surface morphology of tissues. To
prevent deformation artifacts, tissue samples generally require
time-consuming pretreatment, including dehydration and metal
coating, before they are placed in the vacuum of the microscope.
Dehydration, metal coating and the other tedious sample
pretreatments are not required for ASEM.
ASEM is a unique imaging technique with many special
characteristics. First, samples can be observed in liquid at
atmospheric pressure. This is made possible by the use of an open
dish-shaped sample holder and an inverted SEM. The former seals the
top of the SEM column and has a thin SiN-film window in its base,
allowing SEM images of the sample to be obtained from below.
Second, an OM positioned above the open ASEM dish facilitates the
outside manipulation of the sample during observation. Third, the
electron-conducting aqueous solution employed eliminates charging
of the sample. Fourth, the detection of backscattered electrons
allows a 2- to 3-μm slice of the specimen resting on the flat
SiN-film window to be observed when the SEM is operated at an
accelerating voltage of at 30 kV (12,13,26).
This means that diced and stained tissues can be viewed quickly
without further thin-sectioning, providing structural information
about the connectivity and organization of cells and the fine
intracellular and extracellular features near the SiN film.
The oxidized surface of the SiN film,
SiOx, formed during the fabrication process is
considered to have a glass-like nature. Certain cell types, e.g.,
HEK, COS, HeLa and B16 melanoma cells, can grow directly on this
film and on the glass base of the sample dish. However, the culture
of primary cells directly isolated from mouse tissues is
facilitated if the surfaces are first coated by poly-L-lysine or
proteins (12,14,15).
As a hippocampus slice from mammal brain has been successfully
cultured on cover glass, organotypic cultures (27,28)
are highly expected to grow on the ASEM dish, allowing their study
at high resolution. Because the dish accommodates 2 ml of liquid,
it provides a stable culture environment for organs. It eliminates
the delicate handling of a few tens of microliters of liquid
containing a tiny piece of tissue, required to charge an
environmental capsule. Furthermore, it facilitates the
administration of drugs (29),
making drug screening a further possible application of the ASEM.
The open configuration of the ASEM dish also enables
micro-injection or electrophysiology of tissues, monitored by
OM.
In the present study, the staining reagents UA,
Pt-blue, PTA, OsO4 and lead citrate, were successfully
employed for tissue observation by ASEM. In particular, the bright
images of chromatin and nucleoli observed in nuclei stained with
Pt-blue and/or PTA (Figs. 2F,
3C and 7F) suggest that these moderate and
economical reagents can be used to visualize nucleotide-protein
complexes. This, together with earlier images of chromosomes
(11), raises the possibility of
applying Pt-blue/PTA staining and ASEM to chromatin-related fields,
e.g., spermatology. High resolution karyotyping using ASEM might
help to find slight chromosomal deviations that are difficult to
observe at present.
Simple PTA staining and ASEM visualized a bacteria
colony on the mucosal side of stomach tissue (Fig. 6). If the staining time can be
reduced, ASEM has the potential to visualize the characteristics of
bacteria without further culturing, which would be especially
important for infectious bacteria that are viable but
non-culturable (VNC bacteria), and for anaerobes that take a
relatively long time (several days) to culture for diagnosis.
Furthermore, ASEM could be applied to visualize the infiltration of
bacteria into tissue regions and the inflammation they trigger
during surgery and treatment. The modified NCMIR staining method
involving the use of OsO4 (25) clearly visualized membranous
structures at high contrast, not only for eukaryotic cells but also
for prokaryotes (Fig. 6). This
could be exploited to study the symbiotic bacteria that are
necessary to establish natural immunity, and also commensal
bacteria, including Helicobactor pylori.
Today, most kidney biopsies are examined using a
combination of three microscopies: OM, immunofluorescence (IF)
microscopy and electron microscopy (EM). EM is especially necessary
for the diagnosis of the primary glomerulopathies: thin glomerular
basement membrane disease, Alport syndrome, fibrillary and/or
immunotactoid glomerulonephritis and minimal change disease
(30). The diagnosis is basically
performed by Epon thin-section TEM (31), and takes a relatively long time due
to the required embedding and ultra-thin sectioning procedure. By
contrast, ASEM of kidney tissue can be completed within 30 min, or
less if the quick UA staining method presented here is employed.
With further improvement of the method, ASEM could be an attractive
tool for the diagnosis of kidney diseases or to obtain a second
opinion. The size of renal biopsies that can be taken from patients
is limited. A tissue cube as small as (1×1×5 mm) can be imaged by
embedding and TEM, and deliver meaningful results. The open ASEM
dish configuration in combination with OM, allows tissue to be
manually moved across the flat SiN window using tweezers (Fig. 10). In this way, biopsies of
similar size could be investigated by ASEM. As demonstrated with
spinal cord tissue, the observation efficiency can also be
increased by the use of a new multi-windowed ASEM dish that has an
imaging field eight times that of the previous 250×250 μm
single-windowed version (Fig.
10). Imaging large areas will be accelerated even more in the
future by increasing the number of windows, optimizing their shape
and arrangement, and automating sample shifts by constructing
motor-driven auto-manipulator over the dish. Two lines of windows
with one displaced to give the region a checkered appearance, or a
single long narrow rectangular window, would realize the efficient
monitoring of a large square area when combined with orthogonal
sample shift increments.
During intraoperative cancer diagnosis, the sizes of
cell nuclei are usually determined by cryo-sectioning tissue
excisons, staining them with H&E and observing them by OM.
However, this method takes at least 15–30 min, during which time
the tumor excision surgery may have to be suspended. Among the
procedures involved in the diagnosis, cryo-thin sectioning is the
most difficult, and takes most of the time. Because of the
difficulty of this step diagnosis is sometimes confirmed later by
paraffin-thin sectioning or occasionally by antibody labeling,
involving the possibility of further surgery. When stained with PTA
or Pt-blue, nuclei in various tissues, including metastasized
cancer cells, were clearly imaged by ASEM without the need for
thin-sectioning (Figs. 8 and
9). ASEM observes a 2- to 3-μm
specimen layer at 30 kV (12,13,26),
which is comparable to the thickness of the cryo-thin sections used
for intraoperative diagnosis. Equipped with a fail-safe system
against film-window breakage (29), the ASEM has the potential to make
diagnoses both quicker and easier, and may be a suitable way to
obtain a second opinion. Larger tissue areas could be inspected by
moving the sample and by the use of multi-windowed ASEM dishes, as
detailed above. To aid diagnosis, the colored OM images of
H&E-stained tissue sections more familiar to pathologists
should be mimicked by artificially coloring nuclei and cytoplasm in
ASEM images blue and red using image recognition algorithms.
Furthermore, the immunohistological identification of cancer cells
should also be accelerated by the use of immuno-gold labeling and
ASEM, as successfully performed for cells cultured on an ASEM dish
(12,14,15).
In addition, the cathodoluminescence (CL) of luminescent probes is
a further option that could be exploited to obtain multi-colored
high-resolution images by ASEM (29).
In conclusion, in the present study, we stained
different mouse tissues with various heavy metals and imaged them
immersed in an open aqueous solution at atmospheric pressure using
the ASEM. Not only intercellular connections but also cell
organelles were clearly observed in tissues at high resolution.
Staining with UA could be achieved in as little as 15 min. After
further development of both the staining procedure and the ASEM
dish to allow the quick observation of a wider area, this method is
expected to be used for rapid intraoperative cancer diagnosis and
for the diagnosis of kidney diseases. Furthermore, ASEM can be
applied to study the host-microbe interaction that is critical for
carcinogenesis in the stomach and also for the body’s homeostasis,
especially for the establishment of our natural immunity.
The results presented clearly demonstrate that ASEM
is widely applicable to study the basic biology of cells and
tissues, to zymology and to food science and industry in general
and, perhaps most importantly, to pathology, especially for the
diagnosis of cancer and the detection of pathogens during surgery
and treatment.
Acknowledgements
We thank Dr Toshihiko Ogura at the National
Institute of Advanced Industrial Science and Technology (AIST) for
valuable discussions concerning the development of the ClairScope,
and also Dr Tatsuo Ushiki (Niigata University) for critical reading
of the manuscript, Dr Hirofumi Matsui (Tsukuba University), Dr
Shinya Sugimoto and Dr Ken-ichi Okuda (The Jikei University School
of Medicine), Dr Tom Kouki (Jichi Medical University), Dr Yuusuke
Maruyama and Dr Toshio Moriya (AIST) for discussions during the
course of the research. The present study was supported by
Grant-in-Aid for Scientific Research on Innovative Areas,
Structural Cell Biology (to C.S.), by CREST (to C.S.), by grants
from the Ministry of Education, Culture, Sports, Science, and
Technology (MEXT) (to C.S.), by grants from the Kambayashi
Zaydan-in-Aid for Scientific Research and by AIST. This manuscript
was prepared with the assistance of a scientific editing
service.
Abbreviations:
|
ASEM
|
atmospheric SEM
|
|
DDW
|
double distilled water
|
|
EM
|
electron microscope/microscopy
|
|
GA
|
glutaraldehyde
|
|
H&E
|
hematoxylin and eosin
|
|
NCMIR
|
National Center for Microscopy and
Imaging Research
|
|
OM
|
optical microscope/microscopy
|
|
PB
|
phosphate buffer
|
|
PBS
|
phosphate-buffered saline
|
|
PFA
|
paraformaldehyde
|
|
PTA
|
phosphotungstic acid
|
|
Pt-blue
|
platinum-blue,
Pt4(NH3)8(C6H13O5)4,
also known as TI-blue
|
|
RT
|
room temperature
|
|
SEM
|
scanning electron
microscope/microscopy
|
|
SiN
|
silicon nitride
|
|
TEM
|
transmission electron
microscope/microscopy
|
|
THC
|
tetrahydrocannabinol
|
|
UA
|
uranyl acetate
|
References
|
1
|
Golgi C: Intorno alla struttura delle
cellule nervose. Bollettino della Società Medico-Chirurgica di
Pavia. 13:1–14. 1898.(In Italian).
|
|
2
|
Hell SW: Far-field optical nanoscopy.
Science. 316:1153–1158. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wilson LB: A method for the rapid
preparation of fresh tissues for the microscope. JAMA. 45:17371905.
View Article : Google Scholar
|
|
4
|
Ferreiro JA, Myers JL and Bostwick DG:
Accuracy of frozen section diagnosis in surgical pathology: review
of a 1-year experience with 24,880 cases at Mayo Clinic Rochester.
Mayo Clin Proc. 70:1137–1141. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Thiberge S, Nechushtan A, Sprinzak D, et
al: Scanning electron microscopy of cells and tissues under fully
hydrated conditions. Proc Natl Acad Sci USA. 101:3346–3351. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Nyska A, Cummings CA, Vainshtein A, et al:
Electron microscopy of wet tissues: a case study in renal
pathology. Toxicol Pathol. 32:357–363. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Barshack I, Polak-Charcon S, Behar V, et
al: Wet SEM: a novel method for rapid diagnosis of brain tumors.
Ultrastruct Pathol. 28:255–260. 2004. View Article : Google Scholar
|
|
8
|
Behar V: Applications of a novel SEM
technique for the analysis of hydrated samples. Microsc Microanal.
19:21–23. 2005.
|
|
9
|
Kristt D and Nyska A: The wet tissue SEM -
a new technology with applications in drug development and safety.
J Toxicol Pathol. 20:1–11. 2007. View
Article : Google Scholar
|
|
10
|
Barshack I, Kopolovic J, Chowers Y, et al:
A novel method for ‘wet’ SEM. Ultrastruct Pathol. 28:29–31. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Nishiyama H, Suga M, Ogura T, Maruyama Y,
Koizumi M, Mio K, Kitamura S and Sato C: Atmospheric scanning
electron microscope observes cells and tissues in open medium
through silicon nitride film. J Struct Biol. 172:191–202. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Maruyama Y, Ebihara T, Nishiyama H, Suga M
and Sato C: Immuno EM-OM correlative microscopy in solution by
atmospheric scanning electron microscopy (ASEM). J Struct Biol.
180:259–270. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Suga M, Nishiyama H, Ebihara T, Ogura T
and Sato C: Atmospheric electron microscope: limits of observable
depth. Microsc Microanal. 15:924–925. 2009. View Article : Google Scholar
|
|
14
|
Hirano K, Kinoshita T, Uemura T, Motohashi
H, Watanabe Y, Ebihara T, Nishiyama H, Sato M, Suga M, Maruyama Y,
Tsuji NM, Yamamoto M, Nishihara S and Sato C: Electron microscopy
of primary cell cultures in solution and correlative optical
microscopy using ASEM. Ultramicroscopy. 143:52–66. 2014. View Article : Google Scholar
|
|
15
|
Kinoshita T, Mori Y, Hirano K, Sugimoto S,
Okuda K, Matsumoto S, Namiki T, Ebihara T, Kawata M, Nishiyama H,
Sato M, Suga M, Higashiyama K, Sonomoto K, Mizunoe Y, Nishihara S
and Sato C: Immuno-electron microscopy of primary cell cultures
from genetically modified animals in liquid by atmospheric scanning
electron microscopy. Microsc Microanal. 20:469–483. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sato C: Atmospheric scanning electron
microscopy (ASEM) realizes direct EM-OM linkage in solution:
aqueous immunocytochemistry. JEOLnews. 46:17–22. 2011.
|
|
17
|
Sato C, Manaka S, Nakane D, Nishiyama H,
Suga M, Nishizaka T, Miyata M and Maruyama Y: Rapid imaging of
mycoplasma in solution using atmospheric scanning electron
microscopy (ASEM). Biochem Biophys Res Commun. 417:1213–1218. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Maruyama Y, Ebihara T, Nishiyama H,
Konyuba Y, Senda M, Numaga-Tomita T, Senda T, Suga M and Sato C:
Direct observation of protein microcrystals in crystallization
buffer by atmospheric scanning electron microscopy. Int J Mol Sci.
13:10553–10567. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Akita M, Tanaka K, Murai N, Matsumoto S,
Fujita K, Takaki T and Nishiyama H: Detection of CD133 (prominin-1)
in a human hepatoblastoma cell line (HuH-6 clone 5). Microsc Res
Tech. 76:844–852. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Murai T, Maruyama Y, Mio K, Nishiyama H,
Suga M and Sato C: Low cholesterol triggers membrane
microdomain-dependent CD44 shedding and suppresses tumor cell
migration. J Biol Chem. 286:1999–2007. 2011. View Article : Google Scholar :
|
|
21
|
Sakai H, Furihata M, Matsuda C, et al:
Augmented autocrine bone morphogenic protein (BMP) 7 signaling
increases the metastatic potential of mouse breast cancer cells.
Clin Exp Metastasis. 29:327–338. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Takahashi M, Furihata M, Akimitsu N, et
al: A highly bone marrow metastatic murine breast cancer model
established through in vivo selection exhibits enhanced
anchorage-independent growth and cell migration mediated by ICAM-1.
Clin Exp Metastasis. 25:517–529. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Takahashi M, Miyazaki H, Furihata M, et
al: Chemokine CCL2/MCP-1 negatively regulates metastasis in a
highly bone marrow-metastatic mouse breast cancer model. Clin Exp
Metastasis. 26:817–828. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gage GJ, Kipke DR and Shain W: Whole
animal perfusion fixation for rodents. J Vis Exp.
65:35642012.PubMed/NCBI
|
|
25
|
Deerinck TJ, Bushong EA, Thor A and
Ellisman MH: NCMIR methods for 3D EM: a new protocol for
preparation of biological specimens for serial blockage scanning
electron microscopy. Microscopy. pp. 6–8. 2010, Available at:
http://ncmir.ucsd.edu/sbfsem-protocol.pdf.
|
|
26
|
Suga M, Nishiyama H, Konyuba Y, Iwamatsu
S, Watanabe Y, Yoshiura C, Ueda T and Sato C: The atmospheric
scanning electron microscope with open sample space observes
dynamic phenomena in liquid or gas. Ultramicroscopy. 111:1650–1658.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Gahwiler BH: Organotypic monolayer
cultures of nervous tissue. J Neurosci Methods. 4:329–342. 1981.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gähwiler BH, Capogna M, Debanne D,
McKinney RA and Thompson SM: Organotypic slice cultures: a
technique has come of age. Trends Neurosci. 20:471–477. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Nishiyama H, Koizumi M, Ogawa K, Kitamura
S, Konyuba Y, Watanabe Y, Ohbayashi N, Fukuda M, Suga M and Sato C:
Atmospheric scanning electron microscope system with an open sample
chamber: Configuration and applications. Ultramicroscopy.
147:86–97. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Amann K and Haas CS: What you should know
about the work-up of a renal biopsy. Nephrol Dial Transplant.
21:1157–1161. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Mochizuki S, Moriya T, Naganuma H, et al:
Significance of fat stains in serial sections from Epon-embedded
tissue samples for electron microscopy in renal diseases. Clin Exp
Nephrol. 5:240–245. 2001. View Article : Google Scholar
|