Introduction
Colorectal cancer (CRC) is the third most common
malignant disease in the world (1). Despite many advances in medicine,
nearly 50% of CRC patients show tumor recurrence, which is leading
to poor prognosis, median survival ratio following recurrence is
only 13.3 months (2). Most
recurrences of CRC are thought to be the result of tumor invasion
and metastasis of cancer cells. Therefore, understanding the
molecular mechanisms in CRC metastasis is of crucial significance
in developing therapeutic strategies to improve CRC patients.
Transmembrane-4-L6-family-1 (TM4SF1) is a 22-kDa
four-transmembrane-domain protein. It was identified in 1986 as a
tumor cell antigen of mouse monoclonal antibody L6, which also has
a low expression in normal vascular endothelium (3,4).
There are five other structurally similar proteins: TM4SF4/IL-TMP,
TM4SF5/L6H, TM4SF18/L6D, TM4SF19/OCTM4 and TM4SF20/TCCE518
(5). Recently, studies have shown
that TM4SF1 is associated with tumor growth, motility, invasion and
metastasis with high expression in human lung, breast, colon,
ovarian, renal and prostate carcinomas (3,6–10).
In particular, TM4SF1 has high expression in CRC tissues, and
downregulation of TM4SF1 can decrease the progression and
metastasis of CRC (11).
Therefore, TM4SF1 inhibition might provide a strategy for treating
CRC.
MicroRNAs (miRNAs) are a new classification of
endogenous, small, single-stranded RNAs composed of 19–24
nucleotides. miRNAs modulate gene expression by binding to the
3′-untranslated region (3′-UTR) of the target mRNA, resulting in
downregulation of the mRNA transcript or inhibition of the protein
translation process (12). In
addition, miRNAs regulate many cellular processes, including
apoptosis, cell cycle progression, proliferation, differentiation,
invasion and migration and affect tumorigenesis (13–19).
Thus, understanding the underlying molecular mechanisms of miRNA in
malignant tumors is critical to CRC therapy. In this study, we
focused on miRNA-9 (miR-9) because downregulation of its expression
has been observed in several cancer types, such as cervical
adenocarcinoma, breast, gastric, ovarian and hepatocellular
carcinoma (20–24). In CRC, miR-9 expression is also
downregulation by binding to the 3′-UTR, and it has the potential
to suppress Cdx2 (caudal-type homeobox 2) and UHRF1 (ubiquitin-like
with plant homeodomain and ring finger domain 1), leading to
proliferation, apoptosis, migration and invasion (25–27).
However, little is known about the factors that modulate TM4SF1 in
CRC invasion and metastasis. We hypothesized that miRNAs are
associated with TM4SF1 expression in CRC motility.
We analyzed the expression of TM4SF1 in 60 paired
CRC tissues and found that the expression level of TM4SF1 was
significantly higher in CRC tumors than in normal tissues, and the
expression level of TM4SF1 was associated with clinical
pathological stage and lymph node metastasis. Moreover, miR-9
directly targeted its binding site in the TM4SF1 3′-UTR, which has
a critical role in regulating CRC cell migration and invasion.
Furthermore, miR-9 regulated cell motility via suppressing MMP-2,
MMP-9 and VEGF expression in CRC cell lines. Taken together, miR-9
is associated with the motility of CRC and can be used for
molecular targeted therapies in CRC.
Materials and methods
Cell culture
Human colorectal cancer cell lines SW480, Caco2,
LS174T, SW620 and HCT116 were purchased from the American Type
Culture Collection (ATCC; Manassas, VA, USA). The cells were grown
in RPMI-1640 (Invitrogen, Carlsbad, CA, USA) according to the
manufacturer's recommendations, with 10% fetal bovine serum (FBS)
and 1% streptomycin at a humidified 5% CO2 at 37°C.
MicroRNAs transfection and siRNA
treatment
Both miR-9 (hsa-miR-9a-5p; Pre-miRNA, miRNA
Precursor AM17100; Product ID: PM 10022) and anti-miR-9
(anti-hsa-miR-9a-5p; anti-miRNA, miRNA inhibitor AM17100; Product
ID: AM10022) were commercially synthesized (Ambion, Austin, TX,
USA). The miR control and anti-miR control were purchased from
Shanghai GenePharma Co., Ltd., (Shanghai, China) and siRNA-TM4SF1
was commercially synthesized (Thermo Fisher Scientific, Rockford,
IL, USA). SW480 and HCT116 cells (2–5×105) were plated
in 6-well plates and cultured for one day before transfection. For
transfection, miRNAs or siRNA were used at working concentrations
of 50 or 20 nM using Lipofectamine 2000 reagent (Invitrogen). Cells
were harvested at 24, 48 and 72 h after transfection for miRNA,
mRNA and protein, respectively.
Patients and tissue specimens
Sixty of CRC tissues and paired normal tissues were
obtained through the Biobank of Chonbuk National University
Hospital, a member of the National Biobank of Korea. All patients
had a pathological diagnosis of CRC, each paired sample was
classified according to TNM Classification of Malignant Tumours
(TNM) classification and were frozen in liquid nitrogen and stored
at −80°C. The characteristics of patients are shown in Table I. This study consisted of 25
(41.7%) females and 35 (58.3%) males with a mean age of 63.1 years.
The study protocol was approved by the Institutional Review Boards
of Chonbuk National University Hospital (IRB no. 2014-10-05).
 | Table ICharacteristics of the 60 CRC
patients. |
Table I
Characteristics of the 60 CRC
patients.
| Variables | N (%) |
|---|
| Gender |
| Male | 35 (58.3) |
| Female | 25 (41.7) |
| Age (years) |
| <60 | 24 (40.0) |
| ≥60 | 36 (60.0) |
| Histological
differentiation |
| Well | 8 (13.3) |
| Moderate | 46 (76.7) |
| Poor | 6 (10.0) |
| Tumor status
(T) |
| T1–T2 | 20 (33.3) |
| T3–T4 | 40 (66.7) |
| Lymph node
metastasis (N) |
| N0 | 33 (55.0) |
| N1–N2 | 14 (23.3) |
| N3–N4 | 13 (21.7) |
| Distant metastasis
status (M) |
| M0 | 57 (95.0) |
| M1 | 3 (5.0) |
| AJCC |
| I + II | 30 (50.0) |
| III + IV | 30 (50.0) |
RNA isolation and real-time quantitative
polymerase chain reaction (RTQ-PCR) for quantification of miR-9 and
TM4SF1
Total RNA from cells or human normal tissue/matched
tumor samples was extracted using TRIzol reagent (Invitrogen,
Carlsbad, CA, USA). Reverse transcription was performed using M-MLV
Reverse Transcriptase (Promega, Madison, WI, USA), according to the
manufacturer's protocol. RTQ-PCR was performed using an ABI 7500
real-time PCR system (Applied Biosystems, Foster City, CA, USA). In
brief, 20 μl of Master Mix was prepared on ice with 10 μl of 2X
SYBR, 1 μl of primers, 2 μl of DNA and 7 μl of nuclease-free water.
The Master Mix was initially denatured 95°C for 10 min followed by
40 cycels of denaturation at 95°C for 15 sec, annealing and
extension at 60°C for 30 sec. The geometric average Ct value was
used to calculate relative expression of the TM4SF1 using the
method 2−ΔΔCT, which was normalized to
beta-2-microglob-ulin (B2M). Primers used in this experiment were:
5′-TCG CGGCTAATATTTTGCTT-3′ (forward) and 5′-TGCAATT
CCAATGAGAGCAG-3′ (reverse) for TM4SF1; and 5′-CCTG
AATTGCTATGTGTCTGGG-3′ (forward) and 5′-TGATG CTGCTTACATGTCTCGA-3′
(reverse) for B2M. Expression level of miR-9 was determined by the
TaqMan miRNA assay kit (Applied Biosystems) and normalized using
the RNU48. The reaction volume of 20 μl included 2X Master Mix,
each primer 2 μl, cDNA 2 μl, nuclease-free water 6 μl, and
amplification was carried out as follows: 95°C for 10 min, 40
cycles of 95°C for 30 sec, 60°C for 1 min and all the samples were
performed in triplicate.
TM4SF1 target prediction by
bioinformatics methods
To predict the target miRNAs of TM4SF1, we used
bioinformatics software, TargetScan (www.targetscan.org), PicTar (www.mdc-berlin.de), Pita and miRanda-mirSVR
(www.microrna.org), and combined with literature
(27–30), miR-9 was selected for further
study.
Plasmid construction and reporter
assays
TM4SF1 plasmid DNA was kindly donated by Dr R.
Roffler (Academia Sinica, Taipei, Taiwan) and was used to generate
a new construct containing the full open reading frame (ORF) of the
TM4SF1 gene (pcDNA3.1-TM4SF1). The wild-type and mutant TM4SF1
containing the predicted binding site for miR-9 were amplified by
PCR using the primers: 5′-CTCGAGCCCTT TGAACTGCCTTGTGT-3′ (forward)
and 5′-CTCGAGCC CAGTCATCGTAGCCTTTC-3′ (reverse) for TM4SF1-WT:
5′-GGAAAGCCTTTTGTCCTTGAGTACTAGGGATCATG-3′ (forward) and
5′-CATGATCCCTAGTACTCAAGGACAAAAGGCTTTC-3′ (reverse) for TM4SF1-MT.
The PCR product was cloned into the pmirGLO Dual-Luciferase miRNA
target expression vector (Promega), designated TM4SF1-WT after
sequencing. TM4SF1-MT was carried out using a site-directed
mutagenesis kit (Enzynomics, Daejeon, Korea), using TM4SF1-WT as a
template.
For the reporter assay, 5×104 of SW480
cells were seeded in a 24-well plate and transiently transfected
with 200 ng of TM4SF1-WT or TM4SF1-MT reporter plasmid with 50 nM
of miR-9 or miR negative control using Lipofectamine 2000.
Luciferase assays were performed at 24 h after transfection using
the Dual-luciferase assay system (Promega), and they were
normalized with co-transfected Renilla luciferase. All
experiments were performed in triplicate and repeated at least
three times.
Western blot analysis
Protein extraction was prepared according to a
previously described method (31).
Briefly, cells were harvested by resolving in RIPA buffer (50 mM
Tris-HCl, 150 mM NaCl, 1% Triton X-100, 1% sodium deoxycholate,
0.1% SDS and protease inhibitors) and were centrifuged at 13,200
rpm at 4°C for 30 min. After centrifugation, supernatants were used
as whole cell extracts, and 30–35 μg of protein was separated on 8
or 10% polyacrylamide gel and transferred to polyvinylidene
difluoride (PVDF) membranes. The membranes were incubated with 2.5%
non-fat dry milk or 2.5% BSA in TBST for 60 min and then incubated
overnight at 4°C with primary antibodies to TM4SF1 (1:200; Thermo
Fisher Scientific), MMP-2 (1:200; Cell Signaling Technology,
Danvers, MA, USA), MMP-9 (1:200; Cell Signaling Technology) and
VEGF (1:500; Santa Cruz Biotechnology, Santa Cruz, CA, USA). After
washing three times with TBST, the membranes were incubated with
HRP-conjugated rabbit or mouse IgG secondary antibodies for 60 min
at room temperature. After washing three times with TBST, the
proteins were visualized with ECL prime Western blotting substrate
(Amersham, Buckinghamshire, UK) and detected with the
chemiluminescent image system (Fusion Solo S; Vilber Lourmat,
Marne-la-Valle'e Cedex, France). After protein detection, some
membranes were re-probed with an antibody to GAPDH (Bioworld,
Irving, TX, USA) used as a loading control.
Wound healing assay
Cells (5×104) were seeded in 24-well
plates and incubated at 37°C. The confluent cells were scratched
with a 200-μl pipette tip and then transfected as described above.
After 24-h incubation, plates were washed with fresh medium to
remove non-adherent cells and then photographed. Wound area was
determined using an inverted microscope (IX71; Olympus, Center
Valley, PA, USA).
Migration and invasion assay
Cell migration assay was performed using a Transwell
system (24-wells, 8 μm pore size with poly-carbonate membrane; SPL,
Gyeonggi-do Korea) according to the manufacturer's instructions.
Briefly, post-transfected cells were trypsinized, and
1×105 cells were seeded into the upper chamber with
serum free opti-MEM media. The low chamber was filled with 800 μl
medium containing 10% FBS as a chemoattractant. After incubation
for 48 h, cells on the lower side of the filter were fixed in 3.8%
formaldehyde for 20 min and stained with 0.1% crystal violet
solution. The number of cells in five randomly selected fields was
counted under a light microscope and analyzed statistically. For
the invasion assay, the upper chamber was coated with extracellular
matrix (BD Biosciences, Bedford, MA, USA), a soluble basement
membrane matrix. The rest of the assay was performed as the
migration assay.
Statistical analysis
Spearman's correlation analysis was used to
determine the correlation between TM4SF1 and miR-9 expression. All
of the data are shown as mean ± standard deviation (SD).
Statistical differences were analyzed using ANOVA and Student's
t-test, and P-values <0.05 were considered statistically
significant.
Results
TM4SF1 expression is elevated in CRC, and
associated with tumor stage and lymph node metastasis
We first analyzed TM4SF1 expression in human
colorectal cancer specimens, 60 frozen CRC tissues and paired
normal colon tissues using RTQ-PCR. The mRNA level of TM4SF1 in CRC
tissues were significantly higher than the mRNA level in paired
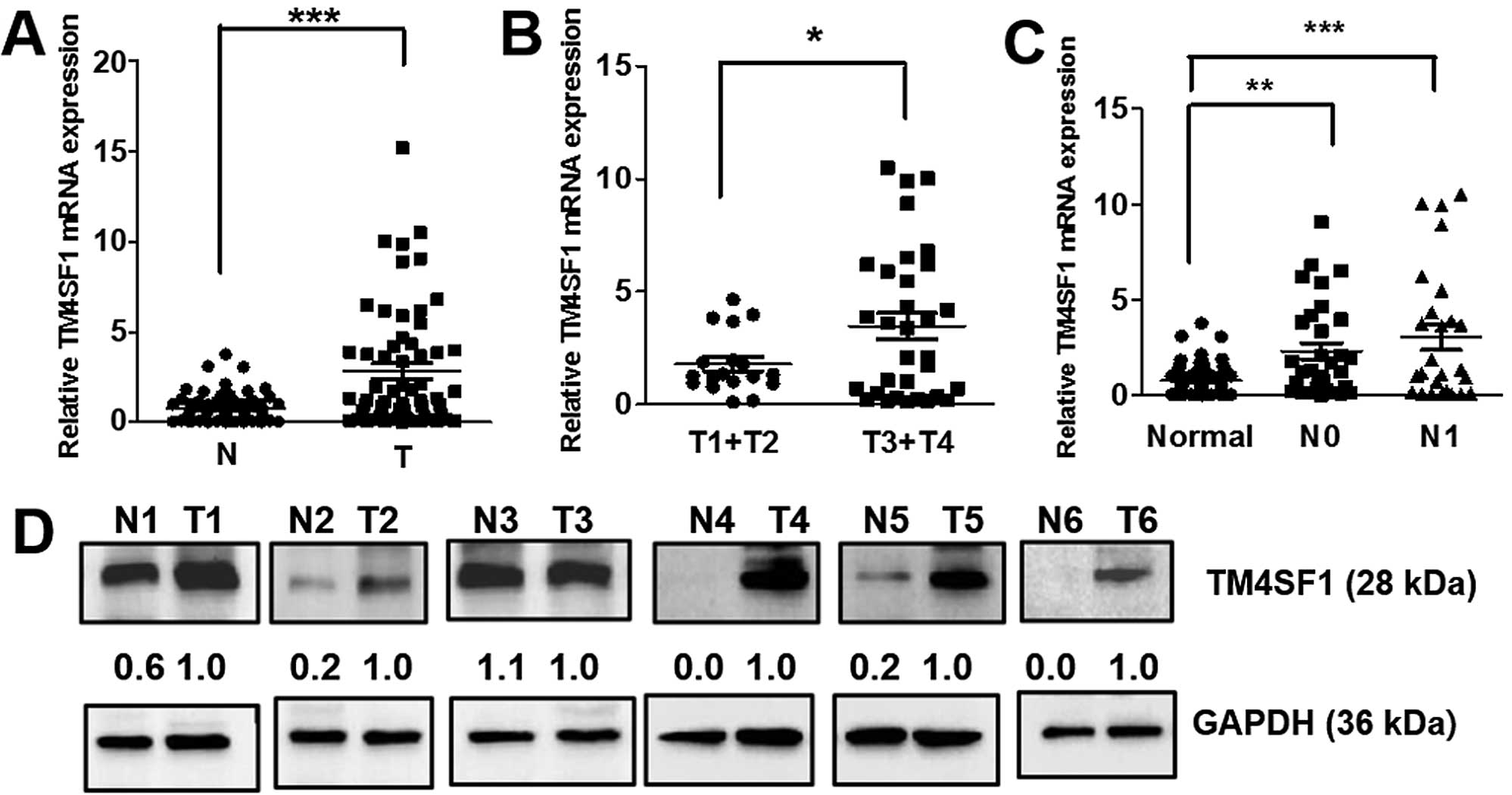
normal tissues (P<0.001 for all comparisons; Fig. 1A). We also found that high level of
TM4SF1 expression was significantly associated with increasing
stage of CRC (P<0.05 for all comparisons; Fig. 1B), suggesting that TM4SF1 has more
important functions in late-stage than early-stage CRC. TM4SF1
expression also was significantly increased with lymph node
metastasis compared with normal tissues (P<0.01 and P<0.001
for all comparisons; Fig. 1C). We
analyzed the level of TM4SF1 protein in 6 CRC tissues and found
that TM4SF1 expression in 5 of 6 were higher than the level in the
matched normal tissues (Fig. 1D).
These results suggest that elevated TM4SF1 expression is associated
with CRC metastasis.
miR-9 is a direct target for TM4SF1 in
CRC and inversely correlated in 5 colorectal cancer cell lines
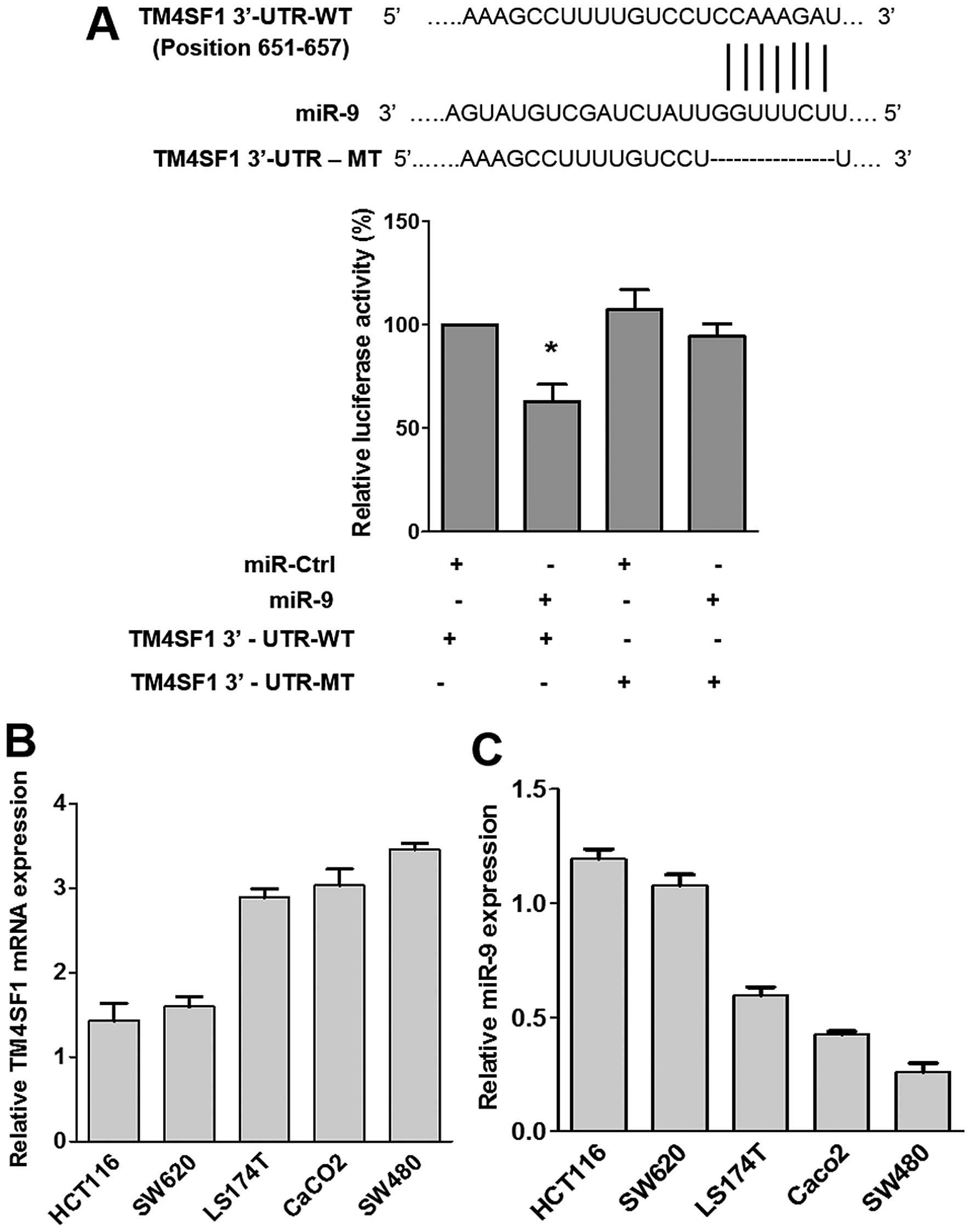
To identify miRNA target sites located in the 3′-UTR
TM4SF1 mRNA, we used bioinformatics software, TargetScan and
miRanda-mirSVR, and the results were combined and compared with
findings in literature (27–30).
miR-9 was found to be conserved in the binding site of TM4SF1
(Fig. 2A) and was therefore
selected for further study.
The luciferase assay showed that co-transfection of
miR-9 and TM4SF1 3′-UTR-WT was significantly reduced by ~40%
compared with the co-transfection of the miR control and TM4SF1
3′-UTR-WT (Fig. 2A). However, no
significant changes in luciferase activity were observed in
co-transfection of either TM4SF1 3′-UTR-MT or the miR-control with
miR-9 (Fig. 2A). These findings
suggest that miR-9 directly targets TM4SF1 via the binding site in
3′-UTR region.
We also examined the expression levels of miR-9 and
TM4SF1 in 5 colorectal cancer cell lines (HCT116, SW620, LS174T,
Caco2 and SW480). As shown that Fig.
2B and C, the mRNA level of TM4SF1 was the highest in SW480 and
the lowest in HCT116 cells. On the contrary, RTQ-PCR results showed
that the expression of miR-9 was highest in HCT116 cells and lowest
in SW480 cells. These findings suggest that the level of TM4SF1
expression is inversely correlated with miR-9 level in CRC cells.
Based on these results, we used SW480 and HCT116 cells to analyze
the gain or loss of TM4SF1.
miR-9 inhibits cell migration and
invasion of CRC cells
To further evaluate the cellular effects of miR-9,
miR-9 alone or miR-9 and TM4SF1 was transfected. MTT assay results
showed that miR-9 significantly inhibited cell growth, but
co-transfection of miR-9 or TM4SF1 had no effect on growth of SW480
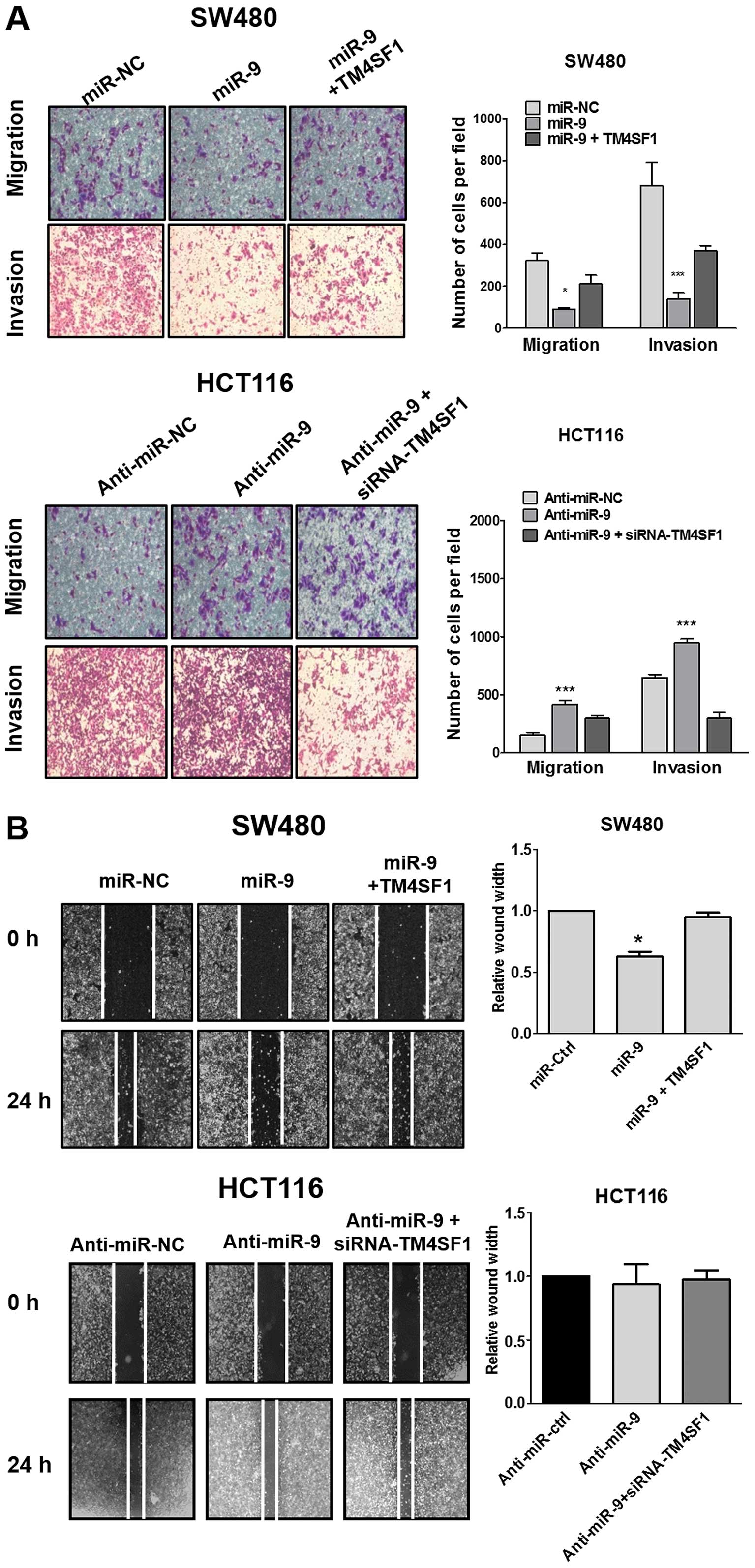
and HCT116 cells (data not shown). To test the cell migration and
invasion potential of CRC Transwell migration, invasion assay and
wound healing in vitro were analyzed. As shown in Fig. 3, overexpression of miR-9 in SW480
cells significantly reduced cell migration (P<0.05) and invasion
(P<0.01), co-transfection of miR-9 with TM4SF1 recovered these
cellular functions. Whereas, overexpression of anti-miR-9 in HCT116
cells significantly increased cell migration (P<0.001) and
invasion compared with the control (P<0.001), co-transfection of
anti-miR-9 with siRNA-TM4SF1 also recovered these effects. Wound
healing assay results showed that miR-9 overexpression
significantly reduced the rate of wound healing (P<0.05), and
co-transfection of miR-9 and TM4SF1 recovered the motility of SW480
cells, however, there was no significant effect of anti-miR-9 and
siRNA-TM4SF1 in HCT116 cells (Fig.
3B). Collectively, miR-9 inhibits CRC cell migration and
invasion in vitro by TM4SF1 expression.
miR-9 inhibits expression of TM4SF1 mRNA
and protein in CRC cells
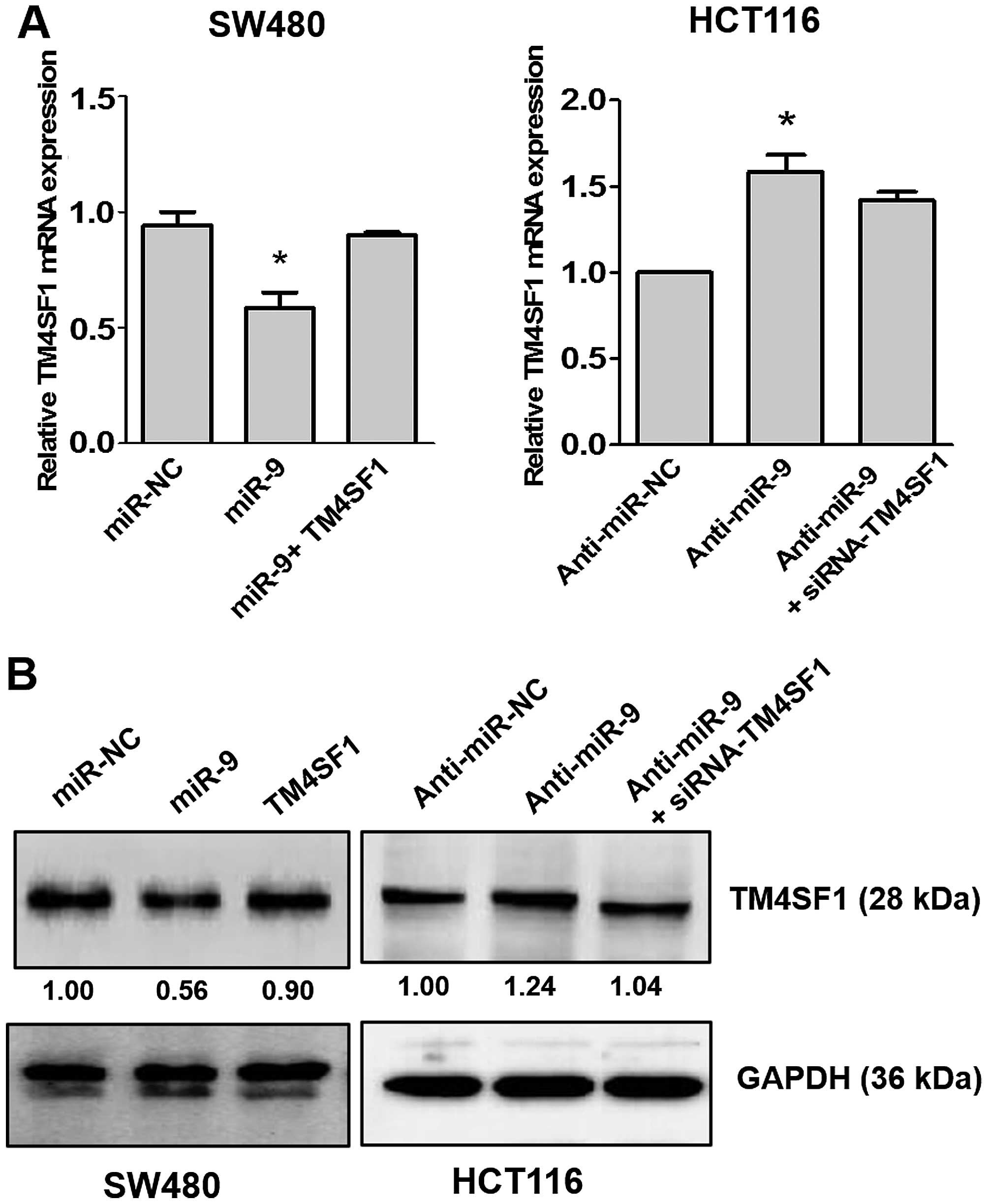
To test whether miR-9 regulates endogenous TM4SF1
expression, miR-9 and anti-miR-9 were transiently transfected into
SW480 and HCT116 cells, and expression levels of TM4SF1 mRNA and
protein were analyzed. As shown in Fig. 4A, transfection of miR-9 in SW480
cells lowered the TM4SF1 mRNA level compared with the level in the
control (P<0.05), and co-transfection of miR-9 and TM4SF1
recovered expression of TM4SF1 mRNA. In contrast, the TM4SF1 mRNA
level in HCT116 cells was enhanced by anti-miR-9 compared with the
control (P<0.05), and the expression level was recovered by
co-transfection with anti-miR-9 and siRNA-TM4SF1. Supporting the
observations, that TM4SF1 protein expression was reduced by miR-9,
and the expression level was also recovered by co-transfection of
miR-9 and TM4SF1. In HCT116 cells, TM4SF1 expression was enhanced
by anti-miR-9, and recovered by co-transfection of anti-miR-9 and
siRNA-TM4SF1 (Fig. 4B). These
results indicate that miR-9 regulates both the transcription and
translation of TM4SFl.
Alteration of miR-9 expression influences
MMP-9, MMP-2 and VEGF activation
To further understand the molecular mechanism of
miR-9/TM4SF1 in inhibiting cell migration and invasion, we
investigated whether it was due to the moderation of adhesion,
which then lead to regulation of MMPs (matrix metalloproteinases)
and the VEGF (vascular endothelial growth factor) signaling pathway
(32,33). Consequently, we analyzed expression
of MMP-9, MMP-2, VEGF, ICAM-1 (intercellular adhesion molecule 1),
VCAM-1 (vascular cell adhesion protein 1) in CRC cells.
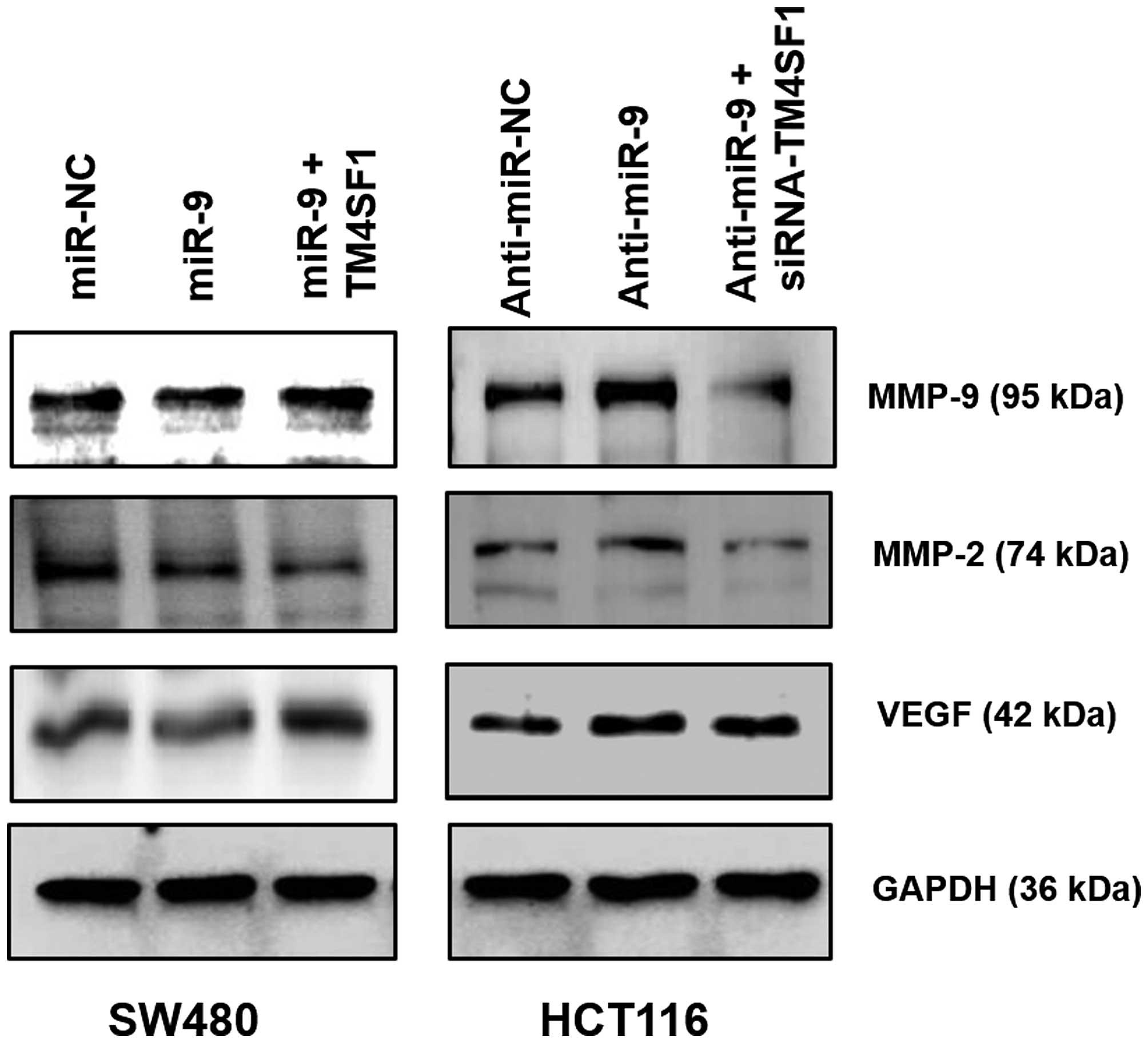
Although miR-9 did not significantly change ICAM-1
or VACM-1 protein levels (data not shown), the expression of MMP-2,
MMP-9 and VEGF was downregulated by miR-9 overexpression, this
expression also recovered with co-transfection of miR-9 and TM4SF1
in SW480 cells (Fig. 5). In
contrast, expression of TM4SF1 protein was upregulated by
anti-miR-9, and this was recovered via co-transfection with
anti-miR-9 and siRNA-TM4SF1 in HCT116 cells. Taken together, miR-9
not only downregulates the expression of TM4SF1, but also
downregulates MMP-2, MMP-9 and VEGF expression in CRC cells.
TM4SF1 inversely correlates with miR-9
expression in CRC
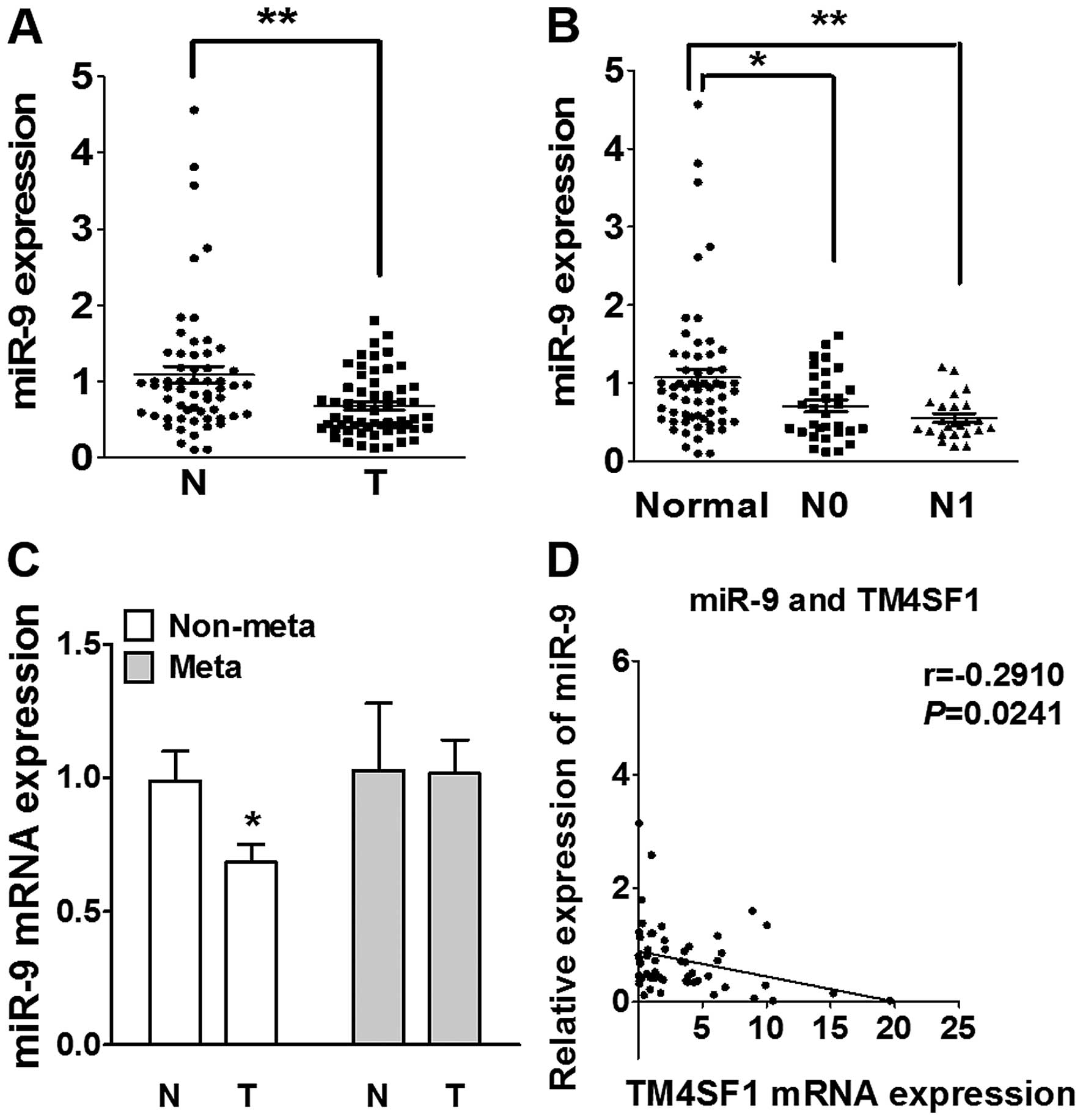
To gain insight into the clinical implications of
miR-9, expression of miR-9 was analyzed in 60 cases of human CRC
using RTQ-PCR. The results revealed that expression level of miR-9
was significantly higher in normal tissues than in CRC tissues
(P-value in each <0.01; Fig.
6A), in normal non-metastasis tissues than in tumor of
non-metastasis (P-value in each <0.05; Fig. 6C). On the contrary, the level of
miR-9 in CRC was significantly decreased with lymph node metastasis
compared with normal tissues (P-value <0.05 and <0.01;
Fig. 6B). We also evaluated that
the correlation between TM4SF1 mRNA and miR-9 expression in the
same CRC tissues. As shown in Fig.
6D, Spearman's rank correlation analysis showed that the
expression levels of miR-9 and TM4SF1 were inverse in the 60 CRC
specimens.
Discussion
TM4SF1 is known to play important roles in growth,
motility, invasion, and metastasis of cancer cells (3,6–8).
Previously, studies have identified aberrant expression of miRNAs
in various types of cancer and analyzed the role of miRNAs in
cancer development, process, metastasis and invasion (34,35).
In the present study, we focused on the expression of TM4SF1 in
human CRC tissues and CRC cell lines along with the miRNA that
regulates TM4SF1 expression. Our results showed that TM4SF1 is
upregulated in 60 paired CRC specimens compared with corresponding
normal tissues and is associated with increased stage and lymph
node metastasis. The expression of TM4SF1 was significantly higher
in late stages compared to early stages.
On the basis of the above findings, bioinformatics
prediction analysis and literature search (27–30),
were performed, miRNA-9 was selected as a potential target of
TM4SF1. This prediction was confirmed by luciferase assay. Our
results also revealed that miR-9 regulates TM4SF1 expression via
the binding site in its 3′-UTR region thereby invasion and
metastasis of CRC cells were reduced. These results suggest that
miR-9 is strongly associated with CRC cell motility via its direct
target gene the TM4SF1.
Clinically, increased expression levels of MMP-2 and
MMP-9 in tumors are significantly associated with metastatic
potential (36–38). Overexpression of MMP-2 and MMP-9
was also identified as prognostic markers of CRC (39). Our results show that miR-9
regulated the protein level of MMP-2/MMP-9 and VEGF via TM4SF1 in
CRC cells. A similar study of VEGF expression found that loss of
TM4SF1 inhibited the regulation of VEGF, thereby inducing
angiogenesis (αV, β3 and β5) in endothelial cells (40). We, therefore, hypothesized that
MMP-2/MMP-9 and VEGF are central mediators in CRC invasion and
metastasis.
Although accumulating evidence has shown that miR-9
is upregulated in several cancers (41–44),
in contrast, many studies have shown downregulation of miR-9 in
cervical adenocarcinoma, gastric, ovarian and hepatocellular
carcinoma (20–24), indicating a diverse role of miR-9
in different cancer types. These results are studies with a small
number of patients. In this study, miR-9 expression level in 60
paired CRC specimens was significantly upregulated in normal
tissues compared with paired tumor tissues. Moreover, our data
showed that level miR-9 expression inversely correlates with the
level of TM4SF1 expression in the same CRC tumor specimens. These
findings suggest that miR-9 expression is an important factor in
the initiation and early stages of the development of CRC.
We could not determine a survival correlation
between miR-9 and TM4SF1 in CRC patients, because survival
information was not provided by Biobank of Korea. However, Shiedeck
et al (45) have reported that overall 3-year survival is
significantly lower in patients with L6-positive blood serum
compared to patients L6-negative. Further studies are warranted to
identify the roles of miR-9 and TM4SF1 in reducing invasion and
metastasis in vivo.
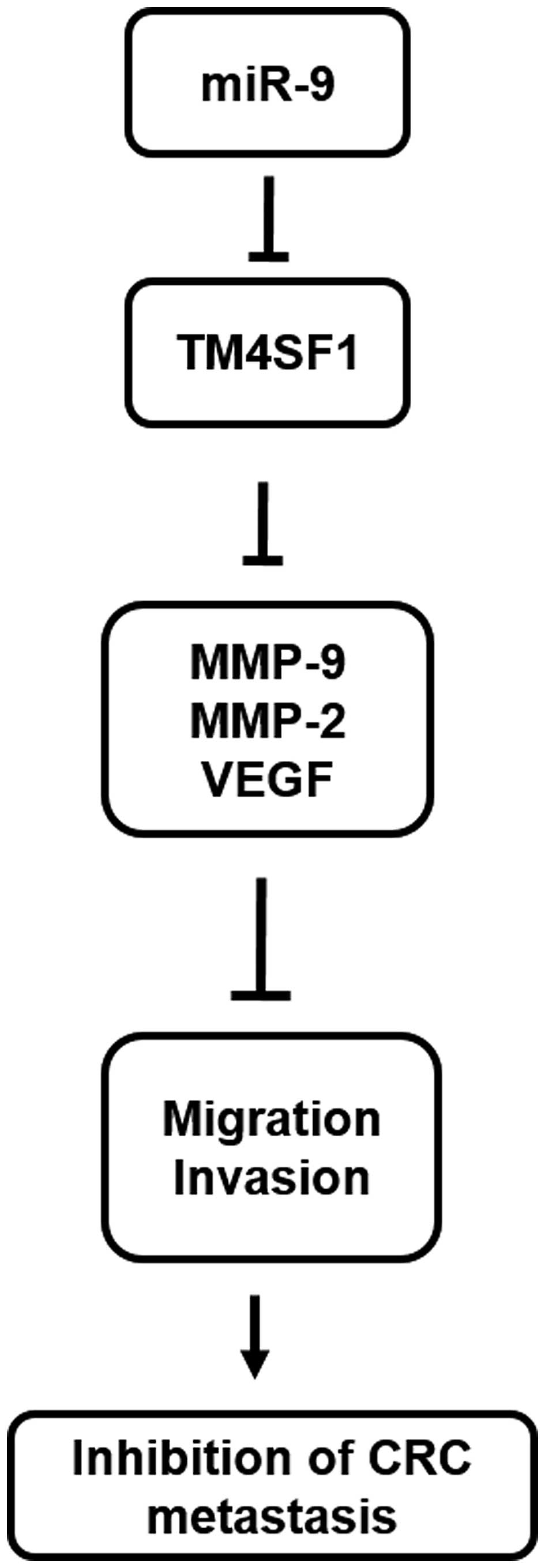
In conclusion, miR-9 is downregulated in CRC
specimens and plays a crucial role in CRC invasion and metastasis
through regulation of TM4SF1 expression. Our results provide
information on a novel mechanism through which miR-9 inhibits the
invasion and metastasis of CRC through inhibiting TM4SF1 expression
regulation of MMP-2/MMP-9/VEGF in vitro (Fig. 7). Collectively, these findings not
only help us understand the molecular mechanism for CRC invasion
and metastasis, but also provide a strong rationale to further
investigate miR-9 as a potential biomarker for CRC.
Acknowledgements
The biospecimens and data used in the present study
were provided by the Biobank of Chonbuk National University
Hospital, a member of the Korea Biobank Network, which is supported
by the Ministry of Health, Welfare and Family Affairs. All samples
derived from the Korea Biobank Network were obtained with informed
consent under institutional review board-approved protocols. This
research was supported by the Basic Science Research Program
through the National Research Foundation of Korea (NRF), funded by
the Ministry of Science, ICT, and the Future Planning
(NRF-2015R1D1A3A01016026) and by the Research Institute of Clinical
Medicine of Chonbuk National University Hospital-Biomedical
Research Institute of Chonbuk National University Hospital.
References
|
1
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
O'Connell MJ, Campbell ME, Goldberg RM,
Grothey A, Seitz JF, Benedetti JK, André T, Haller DG and Sargent
DJ: Survival following recurrence in stage II and III colon cancer:
Findings from the ACCENT data set. J Clin Oncol. 26:2336–2341.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hellstrom I, Horn D, Linsley P, Brown JP,
Brankovan V and Hellstrom KE: Monoclonal mouse antibodies raised
against human lung carcinoma. Cancer Res. 46:3917–3923.
1986.PubMed/NCBI
|
|
4
|
O'Donnell RT, DeNardo SJ, Shi XB, Mirick
GR, DeNardo GL, Kroger LA and Meyers FJ: L6 monoclonal antibody
binds prostate cancer. Prostate. 37:91–97. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wright MD, Ni J and Rudy GB: The L6
membrane proteins - a new four-transmembrane superfamily. Protein
Sci. 9:1594–1600. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chang YW, Chen SC, Cheng EC, Ko YP, Lin
YC, Kao YR, Tsay YG, Yang PC, Wu CW and Roffler SR: CD13
(aminopeptidase N) can associate with tumor-associated antigen L6
and enhance the motility of human lung cancer cells. Int J Cancer.
116:243–252. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lekishvili T, Fromm E, Mujoomdar M and
Berditchevski F: The tumour-associated antigen L6 (L6-Ag) is
recruited to the tetraspanin-enriched microdomains: Implication for
tumour cell motility. J Cell Sci. 121:685–694. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kao YR, Shih JY, Wen WC, Ko YP, Chen BM,
Chan YL, Chu YW, Yang PC, Wu CW and Roffler SR: Tumor-associated
antigen L6 and the invasion of human lung cancer cells. Clin Cancer
Res. 9:2807–2816. 2003.PubMed/NCBI
|
|
9
|
Marken JS, Schieven GL, Hellström I,
Hellström KE and Aruffo A: Cloning and expression of the
tumor-associated antigen L6. Proc Natl Acad Sci USA. 89:3503–3507.
1992. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Allioli N, Vincent S, Vlaeminck-Guillem V,
Decaussin-Petrucci M, Ragage F, Ruffion A and Samarut J: TM4SF1, a
novel primary androgen receptor target gene over-expressed in human
prostate cancer and involved in cell migration. Prostate.
71:1239–1250. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Otsuka M, Kato M, Yoshikawa T, Chen H,
Brown EJ, Masuho Y, Omata M and Seki N: Differential expression of
the L-plastin gene in human colorectal cancer progression and
metastasis. Biochem Biophys Res Commun. 289:876–881. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ying SY, Chang DC, Miller JD and Lin SL:
The microRNA: Overview of the RNA gene that modulates gene
functions. Methods Mol Biol. 342:1–18. 2006.PubMed/NCBI
|
|
13
|
Wang F, Xue X, Wei J, An Y, Yao J, Cai H,
Wu J, Dai C, Qian Z, Xu Z, et al: hsa-miR-520h downregulates ABCG2
in pancreatic cancer cells to inhibit migration, invasion, and side
populations. Br J Cancer. 103:567–574. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhou L, Liu F, Wang X and Ouyang G: The
roles of microRNAs in the regulation of tumor metastasis. Cell
Biosci. 5:322015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Aguda BD, Kim Y, Piper-Hunter MG, Friedman
A and Marsh CB: MicroRNA regulation of a cancer network:
Consequences of the feedback loops involving miR-17–92, E2F, and
Myc. Proc Natl Acad Sci USA. 105:19678–19683. 2008. View Article : Google Scholar
|
|
16
|
Grady WM, Parkin RK, Mitchell PS, Lee JH,
Kim YH, Tsuchiya KD, Washington MK, Paraskeva C, Willson JK, Kaz
AM, et al: Epigenetic silencing of the intronic microRNA
hsa-miR-342 and its host gene EVL in colorectal cancer. Oncogene.
27:3880–3888. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Mo JS, Alam KJ, Kang IH, Park WC, Seo GS,
Choi SC, Kim HS, Moon HB, Yun KJ and Chae SC: MicroRNA 196B
regulates FAS-mediated apoptosis in colorectal cancer cells.
Oncotarget. 6:2843–2855. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hoffman AE, Zheng T, Yi C, Leaderer D,
Weidhaas J, Slack F, Zhang Y, Paranjape T and Zhu Y: microRNA
miR-196a-2 and breast cancer: A genetic and epigenetic association
study and functional analysis. Cancer Res. 69:5970–5977. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Aqeilan RI, Calin GA and Croce CM: miR-15a
and miR-16-1 in cancer: Discovery, function and future
perspectives. Cell Death Differ. 17:215–220. 2010. View Article : Google Scholar
|
|
20
|
Zhang J, Jia J, Zhao L, Li X, Xie Q, Chen
X, Wang J and Lu F: Down-regulation of microRNA-9 leads to
activation of IL-6/Jak/STAT3 pathway through directly targeting
IL-6 in HeLa cell. Mol Carcinog. Mar 25–2015.n/a, 2015. View Article : Google Scholar : Epub ahead of
print.
|
|
21
|
Mohammadi-Yeganeh S, Mansouri A and Paryan
M: Targeting of miR9/NOTCH1 interaction reduces metastatic behavior
in triple-negative breast cancer. Chem Biol Drug Des. 86:1185–1191.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Laios A, O'Toole S, Flavin R, Martin C,
Kelly L, Ring M, Finn SP, Barrett C, Loda M, Gleeson N, et al:
Potential role of miR-9 and miR-223 in recurrent ovarian cancer.
Mol Cancer. 7:352008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Higashi T, Hayashi H, Ishimoto T, Takeyama
H, Kaida T, Arima K, Taki K, Sakamoto K, Kuroki H, Okabe H, et al:
miR-9-3p plays a tumour-suppressor role by targeting TAZ (WWTR1) in
hepatocellular carcinoma cells. Br J Cancer. 113:252–258. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Luo H, Zhang H, Zhang Z, Zhang X, Ning B,
Guo J, Nie N, Liu B and Wu X: Down-regulated miR-9 and miR-433 in
human gastric carcinoma. J Exp Clin Cancer Res. 28:822009.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Tagawa T, Haraguchi T, Hiramatsu H,
Kobayashi K, Sakurai K, Inada K and Iba H: Multiple microRNAs
induced by Cdx1 suppress Cdx2 in human colorectal tumour cells.
Biochem J. 447:449–455. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhu M, Xu Y, Ge M, Gui Z and Yan F:
Regulation of UHRF1 by microRNA-9 modulates colorectal cancer cell
proliferation and apoptosis. Cancer Sci. 106:833–839. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sarver AL, French AJ, Borralho PM,
Thayanithy V, Oberg AL, Silverstein KA, Morlan BW, Riska SM,
Boardman LA, Cunningham JM, et al: Human colon cancer profiles show
differential microRNA expression depending on mismatch repair
status and are characteristic of undifferentiated proliferative
states. BMC Cancer. 9:4012009. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Slattery ML, Herrick JS, Mullany LE,
Valeri N, Stevens J, Caan BJ, et al: An evaluation and replication
of miRNAs with disease stage and colorectal cancer-specific
mortality. Int J Cancer. 137:428–438. 2015. View Article : Google Scholar
|
|
29
|
Bandres E, Agirre X, Bitarte N, Ramirez N,
Zarate R, Roman-Gomez J, Prosper F and Garcia-Foncillas J:
Epigenetic regulation of microRNA expression in colorectal cancer.
Int J Cancer. 125:2737–2743. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Rotkrua P, Akiyama Y, Hashimoto Y, Otsubo
T and Yuasa Y: MiR-9 downregulates CDX2 expression in gastric
cancer cells. Int J Cancer. 129:2611–2620. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kim SL, Liu YC, Park YR, Seo SY, Kim SH,
Kim IH, Lee SO, Lee ST, Kim DG and Kim SW: Parthenolide enhances
sensitivity of colorectal cancer cells to TRAIL by inducing death
receptor 5 and promotes TRAIL-induced apoptosis. Int J Oncol.
46:1121–1130. 2015.
|
|
32
|
Liu LP, Liang HF, Chen XP, Zhang WG, Yang
SL, Xu T and Ren L: The role of NF-kappaB in Hepatitis b virus X
protein-mediated upregulation of VEGF and MMPs. Cancer Invest.
28:443–451. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kim Y, Kang H, Jang SW and Ko J: Celastrol
inhibits breast cancer cell invasion via suppression of
NF-κB-mediated matrix metalloproteinase-9 expression. Cell Physiol
Biochem. 28:175–184. 2011. View Article : Google Scholar
|
|
34
|
Lu J, Getz G, Miska EA, Alvarez-Saavedra
E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA,
et al: MicroRNA expression profiles classify human cancers. Nature.
435:834–838. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Landgraf P, Rusu M, Sheridan R, Sewer A,
Iovino N, Aravin A, Pfeffer S, Rice A, Kamphorst AO, Landthaler M,
et al: A mammalian microRNA expression atlas based on small RNA
library sequencing. Cell. 129:1401–1414. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Duffy MJ, Maguire TM, Hill A, McDermott E
and O'Higgins N: Metalloproteinases: Role in breast carcinogenesis,
invasion and metastasis. Breast Cancer Res. 2:252–257. 2000.
View Article : Google Scholar
|
|
37
|
Hanemaaijer R, Verheijen JH, Maguire TM,
Visser H, Toet K, McDermott E, O'Higgins N and Duffy MJ: Increased
gelatinase-A and gelatinase-B activities in malignant vs. benign
breast tumors. Int J Cancer. 86:204–207. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Chambers AF and Matrisian LM: Changing
views of the role of matrix metalloproteinases in metastasis. J
Natl Cancer Inst. 89:1260–1270. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Mc Donnell S, Chaudhry V, Mansilla-Soto J,
Zeng ZS, Shu WP and Guillem JG: Metastatic and non-metastatic
colorectal cancer (CRC) cells induce host metalloproteinase
production in vivo. Clin Exp Metastasis. 17:341–349. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Shih SC, Zukauskas A, Li D, Liu G, Ang LH,
Nagy JA, Brown LF and Dvorak HF: The L6 protein TM4SF1 is critical
for endothelial cell function and tumor angiogenesis. Cancer Res.
69:3272–3277. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Nass D, Rosenwald S, Meiri E, Gilad S,
Tabibian-Keissar H, Schlosberg A, Kuker H, Sion-Vardy N, Tobar A,
Kharenko O, et al: MiR-92b and miR-9/9* are specifically expressed
in brain primary tumors and can be used to differentiate primary
from metastatic brain tumors. Brain Pathol. 19:375–383. 2009.
View Article : Google Scholar :
|
|
42
|
Ma L, Young J, Prabhala H, Pan E, Mestdagh
P, Muth D, Teruya-Feldstein J, Reinhardt F, Onder TT, Valastyan S,
et al: miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin
and cancer metastasis. Nat Cell Biol. 12:247–256. 2010.PubMed/NCBI
|
|
43
|
Shiiyama R, Fukushima S, Jinnin M,
Yamashita J, Miyashita A, Nakahara S, Kogi A, Aoi J, Masuguchi S,
Inoue Y, et al: Sensitive detection of melanoma metastasis using
circulating microRNA expression profiles. Melanoma Res. 23:366–372.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Song Y, Li J, Zhu Y, Dai Y, Zeng T, Liu L,
Li J, Wang H, Qin Y, Zeng M, et al: MicroRNA-9 promotes tumor
metastasis via repressing E-cadherin in esophageal squamous cell
carcinoma. Oncotarget. 5:11669–11680. 2014. View Article : Google Scholar : PubMed/NCBI
|