Introduction
Renal cell carcinoma (RCC) is a disease in which
cells in the kidney tubules undergo oncogenic transformation. RCC
has multiple subtypes and may occur in hereditary (2–3% of RCC) or
sporadic forms (1,2). RCC is the third most common
urological cancer and accounts for 3% of all adult neoplasias.
Clear cell RCC (ccRCC) is the most common subtype of sporadic RCC
(~80%) (1). The standard curative
treatment for localized diseases remains surgical excision with
total nephrectomy. In contrast, at diagnosis, ~30% of RCCs have
already metastasized. The 5-year survival rate in patients with
advanced stage RCC is poor (5–10%) due to recurrence or distant
metastasis (3,4). Recent molecularly targeted therapy
has improved the survival rate of patients with advanced RCC
(5,6). However, almost all patients
eventually relapse or show distant metastasis due to acquired
resistance to molecularly targeted therapy. Identifying molecular
pathways responsible for RCC metastasis could provide novel
approaches for the development of therapies that block the RCC
metastatic pathways.
The discovery of microRNA (miRNA) in the human
genome provided new directions in cancer research. The miRNAs are
endogenous small RNA molecules (19–22 bases long) that regulate
protein coding gene expression by repressing translation or
cleaving RNA transcripts in a sequence-specific manner (7,8).
Numerous studies have shown that miRNAs are aberrantly expressed in
many human cancers, and they have significant roles in the
initiation, development and metastasis of those cancers (9–11).
Moreover, normal regulatory mechanisms can be disrupted by the
aberrant expression of tumor-suppressive or oncogenic miRNAs in
cancer cells. Therefore, identifying aberrantly expressed miRNAs is
an important first step toward elucidating miRNA-mediated oncogenic
pathways.
Using miRNA expression signatures, we have
identified molecular pathways in RCC that are mediated by
aberrantly expressed miRNAs (12–15).
For example, downregulation of tumor-suppressive miR-218 promoted
cancer cell migration and invasion through dysregulation of the
focal adhesion pathway. In this regard, caveolin-2 has an oncogenic
function in RCC cells (13). The
epithelial-mesenchymal transition (EMT)-related miR-200
family (miR-200a/b/c, miR-141 and miR-429) is
significantly downregulated in RCC where they act as tumor
suppressors that target the focal adhesion and ErbB signaling
pathways (14). The
miR-143/145 cluster was frequently reduced in RCC tissues;
restoration of these miRNAs significantly inhibited RCC cell
proliferation and invasion through targeting of hexokinase-2
(16). More recently, expression
of the miR-23b/27b cluster was significantly decreased in
ccRCC tissues and associated with pathological grade and stage of
the disease (17).
Our miRNA expression signatures of human cancers
revealed that miR-26a and miR-26b were frequently
downregulated in various types of cancer tissues (10,18,19),
suggesting that these miRNAs act as tumor suppressors targeting
several oncogenic pathways. Database searches revealed that there
were few reports of functional analyses of miR-26a or
miR-26b in RCC. The aim of the present study was to
investigate the functional significance of miR-26a and
miR-26b and to identify molecular targets and pathways
contributing to metastasis in RCC cells by miR-26a or
miR-26b regulation. We expect that this analysis will
provide important insights into the potential molecular mechanisms
of RCC oncogenesis and metastasis and will facilitate the
development of therapeutic strategies for the treatment of the
disease.
Materials and methods
RCC clinical specimens and cell
culture
A total of 15 pairs of ccRCC specimens and
corresponding non-cancerous specimens were collected from patients
who had undergone radical nephrectomy at Chiba University Hospital
(Chiba, Japan) from 2012 to 2015. These specimens were staged
according to the General Rule for Clinical and Pathological Studies
on Renal Cell Carcinoma based on the American Joint Committee on
Cancer (AJCC)-UICC TNM classification. The clinicopathological
characteristics of the patients are summarized in Table I. Before tissue collection, written
informed consent of tissue donation for research purposes was
obtained from all the patients.
 | Table ICharacteristics of ccRCC clinical
specimens. |
Table I
Characteristics of ccRCC clinical
specimens.
| No. | Pathology | Grade | pT | INF | v | ly | eg or ig | fc | im | rc | rp | s |
|---|
| 1 | Clear cell | G2 | T1a | a | 0 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 2 | Clear cell | G1>G2 | T1a | a | 0 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 3 | Clear cell | G3>G2 | T1b | a | 0 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 4 | Clear cell | G2>G3>G1 | T1a | a | 0 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 5 | Clear cell | G2>G3 | T1b | a | 0 | 0 | eg | 1 | 1 | 0 | 0 | 0 |
| 6 | Clear cell | G2>G3 | T3a | a | 1 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 7 | Clear cell | G2>G3>G1 | T3a | b | 1 | 0 | ig | 0 | 1 | 1 | 0 | 0 |
| 8 | Clear cell | G2>G3>G1 | T3a | b | 1 | 0 | ig | 1 | 0 | 0 | 0 | 0 |
| 9 | Clear cell | G3 | T3a | b | 1 | 0 | ig | 0 | 0 | 0 | 0 | 0 |
| 10 | Clear cell | G1>G2 | T1b | a | 0 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 11 | Clear cell | G2>G1>G3 | T3a | b | 1 | 0 | ig | 0 | 0 | 0 | 0 | 0 |
| 12 | Clear cell | G2 | T1a | a | 0 | 0 | eg | 0 | 0 | 0 | 0 | 0 |
| 13 | Clear cell |
G2>G1>>G3 | T1b | b | 0 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 14 | Clear cell | G2>G1 | T1a | b | 0 | 0 | eg | 1 | 0 | 0 | 0 | 0 |
| 15 | Clear cell | G2 | T1b | a | 0 | 0 | eg | 0 | 0 | 0 | 0 | 0 |
We used two human RCC cell lines (786-O and A498)
obtained from the American Type Culture Collection (ATCC; Manassas,
VA, USA) as previously described (12–14).
Quantitative real-time reverse
transcription polymerase chain reaction (qRT-PCR)
The procedure for PCR quantification was previously
described. TaqMan probes and primers for LOXL2 (P/N:
Hs00158757_ml; Applied Biosystems, Foster City, CA, USA),
PLOD2 (P/N: Hs01118190_ml; Applied Biosystems) and
GUSB (the internal control; P/N: Hs00939627_ml; Applied
Biosystems) were assay-on-demand gene expression products. The
expression levels of miR-26a (assay ID: 000405; Applied
Biosystems) and miR-26b (assay ID: 000407; Applied
Biosystems) were analyzed by TaqMan quantitative real-time RT-PCR
(TaqMan MicroRNA assay; Applied Biosystems) and normalized to the
expression of RNU48 as previously described (12,20,21).
Transfection with mature miRNAs and
siRNAs
The following mature miRNAs were used: Ambion
Pre-miR miRNA precursor for hsa-miR-26a-5p (product ID:
PM10249; Applied Biosystems) and for hsa-miR-26b-5p (product
ID: PM12899; Applied Biosystems). The following siRNAs were used:
Stealth Select RNAi si-RNA, si-PLOD2 (cat nos. HSS108124 and
HSS182371; Invitrogen) and negative control miRNA/siRNA (P/N:
AM17111; Applied Biosystems). RNAs were incubated with Opti-MEM
(Invitrogen) and Lipofectamine RNAiMax transfection reagent
(Invitrogen) as previously described (12,20,21).
Cell proliferation, migration and
invasion assays
786-O and A498 cells were transfected with 10 nM
miRNAs or si-RNAs by reverse transfection. Cell proliferation,
migration and invasion assays were performed as previously
described (12,20,21).
Western blotting
Cells were harvested 72 h after transfection, and
lysates were prepared. Protein lysates (20 μg) were separated on
Mini-PROTEAN TGX gels (Bio-Rad Laboratories, Hercules, CA, USA) and
transferred to PVDF membranes. Immunoblotting was performed with
rabbit anti-LOXL2 antibodies (1:1000; ab96233; Abcam, Cambridge,
UK) and rabbit anti-PLOD2 antibodies (1:300; 21214-1-AP;
Proteintech Group, Inc., Chicago, IL, USA). Anti-GAPDH antibodies
(1:1,000; ab8245; Abcam) were used as an internal loading control.
The membranes were washed and incubated with anti-rabbit IgG
horseradish peroxidase (HRP)-linked antibodies (#7074; Cell
Signaling Technology). Complexes were visualized with Clarity
Western Substrate (Bio-Rad Laboratories).
Screening of miR-26a and miR-26b target
genes using in silico analysis and gene expression data
To identify miR-26a/b target genes, we used
in silico analysis and genome-wide gene expression analysis.
First, we screened genes using TargetScan release 6.2 (http://www.targetscan.org/). Next, to identify
upregulated genes in ccRCC clinical specimens, we analyzed publicly
available gene expression profiles in the GEO database (accession
nos. GSE22541 and GSE36895). Our strategies for miRNA target
screening were previously described (12,20,21).
Plasmid construction and dual-luciferase
reporter assay
Partial wild-type sequences of the LOXL2 and
PLOD2 3′-untranslated region (UTR) or those with deleted
miR-26a/b binding sites were inserted between the
XhoI-PmeI restriction sites in the 3′-UTR of the
hRluc gene in the psiCHECK-2 vector (C8021; Promega,
Madison, WI, USA). The procedure for the dual-luciferase reporter
assay was previously described (12,20,21).
Statistical analysis
The relationships between the two groups and the
numerical values obtained by real-time RT-PCR were analyzed using
the Mann-Whitney U-test. The relationships among the three
variables and numerical values were analyzed using the
Bonferroni-adjusted Mann-Whitney U test. Spearman's rank test was
used to evaluate the correlations between the expression of
(miR-26a and LOXL2), (miR-26a and
PLOD2), (miR-26b and LOXL2) and
(miR-26b and PLOD2). All analyses were performed
using Expert StatView (version 5; SAS Institute, Inc., Cary, NC,
USA).
Results
Expression levels of miR-26a and miR-26b
in ccRCC clinical specimens and cell lines
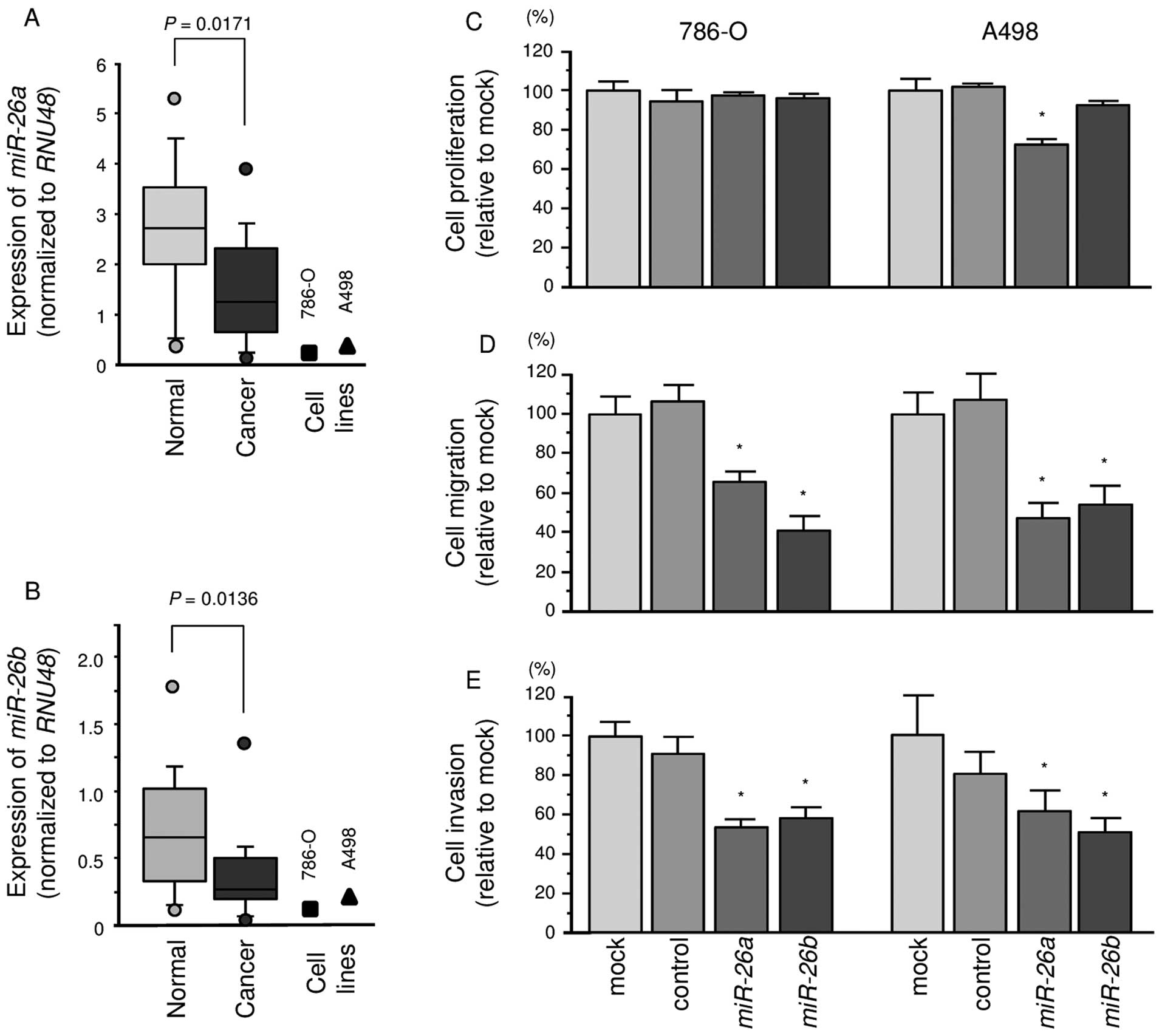
The expression levels of miR-26a and
miR-26b were significantly lower in ccRCC specimens than in
corresponding non-cancerous specimens (P=0.0171 and P=0.0136,
respectively; Fig. 1A and B). In
786-O and A498 cells, the expression levels of miR-26a or
miR-26b were lower than in non-cancerous specimens.
Effects of miR-26a and miR-26b
restoration on cell proliferation, migration and invasion
activities in ccRCC cells
To investigate the functional effects of
miR-26a or miR-26b, we performed gain-of-function
studies using mature miRNA transfection of 786-O and A498
cells.
The XTT assays demonstrated that cell proliferation
was not inhibited in miR-26a or miR-26b transfectants
in comparison with the mock or miR-control transfectants (Fig. 1C).
In contrast, the migration assays demonstrated that
cell migration activity was significantly inhibited in
miR-26a or miR-26b transfectant cells in comparison
with the mock or miR-control transfectants (Fig. 1D). The Matrigel invasion assays
demonstrated that cell invasion activity was significantly
inhibited in miR-26a or miR-26b transfectant cells in
comparison with the mock or miR-control transfectants (Fig. 1E).
Identification of candidate target genes
of miR-26a and miR-26b in ccRCC cells
To identify target genes of miR-26a and
miR-26b (the seed sequences of the two miRNAs are
identical), we used in silico analysis and genome-wide gene
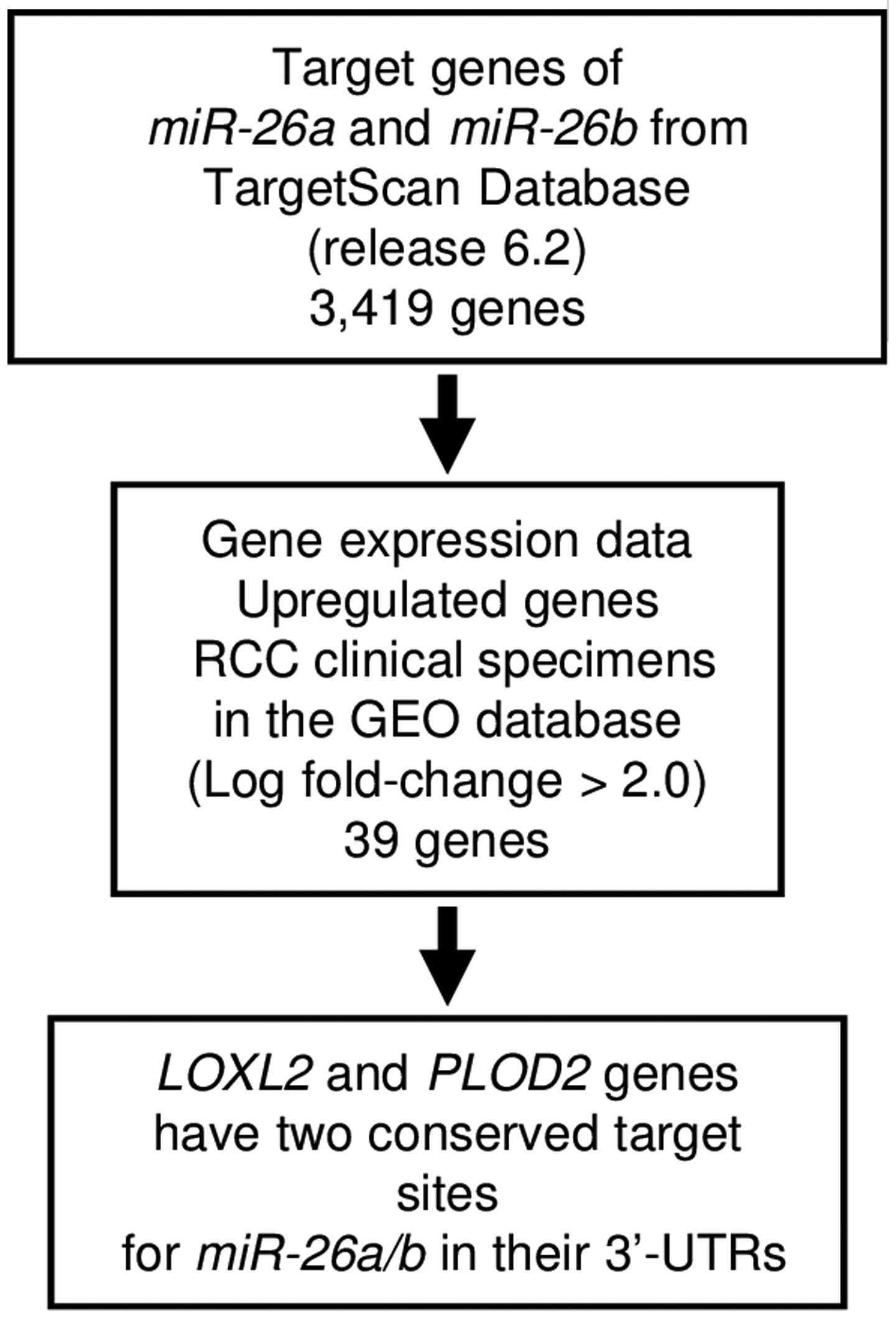
expression data. First, we searched the TargetScan database
(release 6.2: http://www.targetscan.org/) and identified 3,419 genes
that had putative target sites for miR-26a and
miR-26b in their 3′-UTRs. Next, we pared down the list of
putative candidate genes based on upregulated genes determined by
the gene expression data set of RCC clinical specimens in the GEO
(Gene Expression Omnibus) database (accession numbers: GSE36895,
GSE22541). The flow chart outlining our strategy for identification
of candidate target genes of miR-26a and miR-26b is
shown in Fig. 2.
From this selection, 39 candidate genes were
identified as targets of miR-26a and miR-26b
(Table II). Among these candidate
genes, we focused on LOXL2 and PLOD2 genes because
these genes have two conserved target sites for miR-26a and
miR-26b in their 3′-UTRs, and function as collagen
cross-linking enzymes associated with extracellular matrix (ECM)
stiffness. Recent studies showed that aberrantly expressed ECM
contributes to cancer cell metastasis (22,23).
Therefore, these two genes were chosen for further analysis.
 | Table IIPutative candidate target genes
regulated by miR-26a and miR-26b in RCC cells. |
Table II
Putative candidate target genes
regulated by miR-26a and miR-26b in RCC cells.
| Entrez gene ID | Symbol | Gene name | Location | No. of conserved
target sites | No. of poorly
conserved target sites | GEO (GSE36895,
GSE22541 average fold-change |
|---|
| 5352 | PLOD2 | Procollagen-lysine,
2-oxoglutarate 5-dioxygenase 2 | 3q24 | 2 | 0 | 2.2220507 |
| 4017 | LOXL2 | Lysyl oxidase-like
2 | 8p21.3 | 2 | 0 | 2.7719142 |
| 2146 | EZH2 | Enhancer of zeste
homolog 2 (Drosophila) | 7q35-q36 | 1 | 0 | 2.0032272 |
| 3625 | INHBB | Inhibin, β B | 2cen-q13 | 1 | 0 | 3.7558112 |
| 3678 | ITGA5 | Integrin, α 5
(fibronectin receptor, α polypeptide) | 12q11-q13 | 1 | 0 | 2.8391342 |
| 23023 | TMCC1 | Transmembrane and
coiled-coil domain family 1 | 3q22.1 | 1 | 1 | 2.226072 |
| 1404 | HAPLN1 | Hyaluronan and
proteoglycan link protein 1 | 5q14.3 | 1 | 1 | 2.7813237 |
| 7903 | ST8SIA4 | ST8
α-N-acetyl-neuraminide
α-2,8-sialyltransferase 4 | 5q21 | 1 | 0 | 3.1741676 |
| 1846 | DUSP4 | Dual specificity
phosphatase 4 | 8p12-p11 | 1 | 0 | 2.1518986 |
| 6890 | TAP1 | Transporter 1,
ATP-binding cassette, sub-family B (MDR/TAP) | 6p21.3 | 0 | 1 | 2.0403051 |
| 7272 | TTK | TTK protein
kinase | 6q14.1 | 0 | 1 | 2.3837836 |
| 170384 | FUT11 | Fucosyltransferase
11 (α (1,3) fucosyltransferase) | 10q22.2 | 0 | 1 | 2.0443428 |
| 22974 | TPX2 | TPX2,
microtubule-associated, homolog (Xenopus laevis) | 20q11.2 | 0 | 1 | 2.662108 |
| 2210 | FCGR1B | Fc fragment of IgG,
high affinity Ib, receptor (CD64) | 1p11.2 | 0 | 1 | 2.294377 |
| 4747 | NEFL | Neurofilament,
light polypeptide | 8p21 | 0 | 1 | 2.1319628 |
| 5836 | PYGL | Phosphorylase,
glycogen, liver | 14q21-q22 | 0 | 1 | 2.0643747 |
| 1234 | CCR5 | Chemokine (C-C
motif) receptor 5 | 3p21.31 | 0 | 1 | 3.3846455 |
| 55165 | CEP55 | Centrosomal protein
55 kDa | 10q23.33 | 0 | 1 | 2.0711598 |
| 10288 | LILRB2 | Leukocyte
immunoglobulin-like receptor, subfamily B (with TM and ITIM
domains), member 2 | 19q13.4 | 0 | 1 | 2.454539 |
| 1356 | CP | Ceruloplasmin
(ferroxidase) | 3q23-q25 | 0 | 1 | 3.9467278 |
| 3910 | LAMA4 | Laminin, α 4 | 6q21 | 0 | 1 | 2.2182174 |
| 163404 | LPPR5 | Lipid phosphate
phosphatase-related protein type 5 | 1p21.3 | 0 | 1 | 2.450066 |
| 5027 | P2RX7 | Purinergic receptor
P2X, ligand-gated ion channel, 7 | 12q24 | 0 | 3 | 3.0084689 |
| 330 | BIRC3 | Baculoviral IAP
repeat containing 3 | 11q22 | 0 | 1 | 2.2927191 |
| 6507 | SLC1A3 | Solute carrier
family 1 (glial high affinity glutamate transporter), member 3 | 5p13 | 0 | 1 | 2.1052346 |
| 2335 | FN1 | Fibronectin 1 | 2q34 | 0 | 1 | 2.4469628 |
| 8701 | DNAH11 | Dynein, axonemal,
heavy chain 11 | 7p21 | 0 | 1 | 2.2785249 |
| 79850 | FAM57A | Family with
sequence similarity 57, member A | 17p13.3 | 0 | 1 | 2.2900116 |
| 1462 | VCAN | Versican | 5q14.3 | 0 | 1 | 2.524361 |
| 128346 |
C1orf162 | Chromosome 1 open
reading frame 162 | 1p13.2 | 0 | 1 | 2.2255776 |
| 4015 | LOX | Lysyl oxidase | 5q23.2 | 0 | 1 | 3.3194032 |
| 115761 | ARL11 | ADP-ribosylation
factor-like 11 | 13q14.2 | 0 | 1 | 2.4013827 |
| 286336 | FAM78A | Family with
sequence similarity 78, member A | 9q34 | 0 | 1 | 2.1942985 |
| 6664 | SOX11 | SRY (sex
determining region Y)-box 11 | 2p25 | 0 | 1 | 2.577679 |
| 9770 | RASSF2 | Ras association
(RalGDS/AF-6) domain family member 2 | 20p13 | 0 | 1 | 2.619857 |
| 57823 | SLAMF7 | SLAM family member
7 | 1q23.1-q24.1 | 0 | 1 | 2.063896 |
| 58475 | MS4A7 | Membrane-spanning
4-domains, subfamily A, member 7 | 11q12 | 0 | 1 | 2.0315962 |
| 79742 | CXorf36 | Chromosome X open
reading frame 36 | Xp11.3 | 0 | 1 | 2.3148956 |
| 146857 | SLFN13 | Schlafen family
member 13 | 17q12 | 0 | 1 | 2.6972997 |
Direct regulation of LOXL2 and PLOD2 by
miR-26a and miR-26b in ccRCC cells
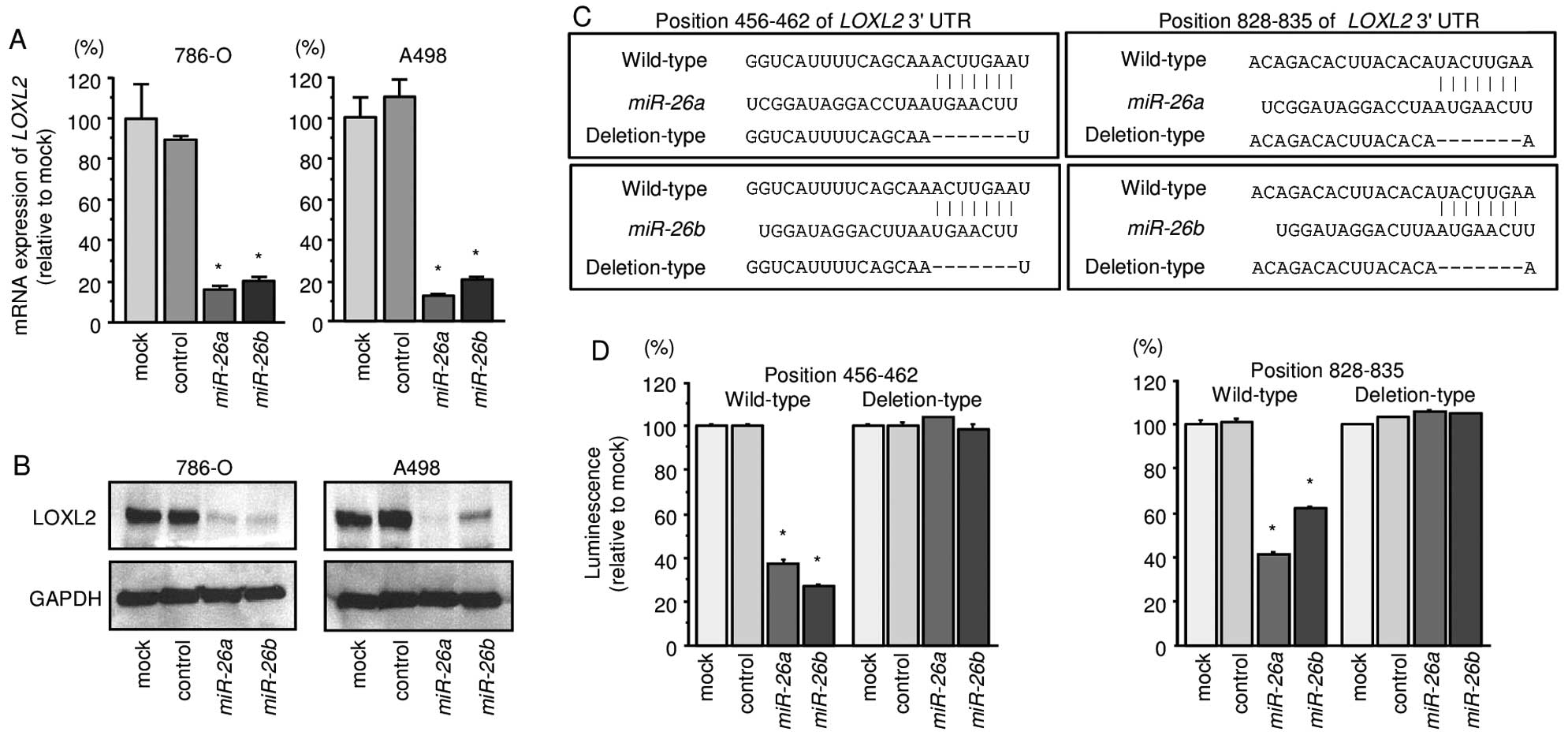
We first performed qRT-PCR and Western blotting to
investigate whether expression of the LOXL2 gene and protein
were reduced by restoration of miR-26a or miR-26b in
786-O and A498 cells. We found that the mRNA and protein expression
levels of LOXL2/LOXL2 were significantly repressed in
miR-26a or miR-26b transfectant cells in comparison
with mock or miR-control transfectants (Fig. 3A and B).
Next, to investigate whether LOXL2 mRNA had
target sites for miR-26a or miR-26b, we performed
luciferase reporter assays in 786-O cells. We used vectors encoding
either the partial wild-type sequence of the 3′-UTR of
LOXL2, including the predicted miR-26a/b target
sites, or deletion vectors lacking the miR-26a/b target
sites. We found that the luminescence intensities were
significantly reduced by transfection with miR-26a or
miR-26b and vectors carrying the wild-type 3′-UTR of
LOXL2, whereas transfection with deletion vectors blocked
the decrease in luminescence. These data suggested that
miR-26a or miR-26b bound directly to specific sites
in the 3′-UTR of LOXL2 (Fig. 3C
and D).
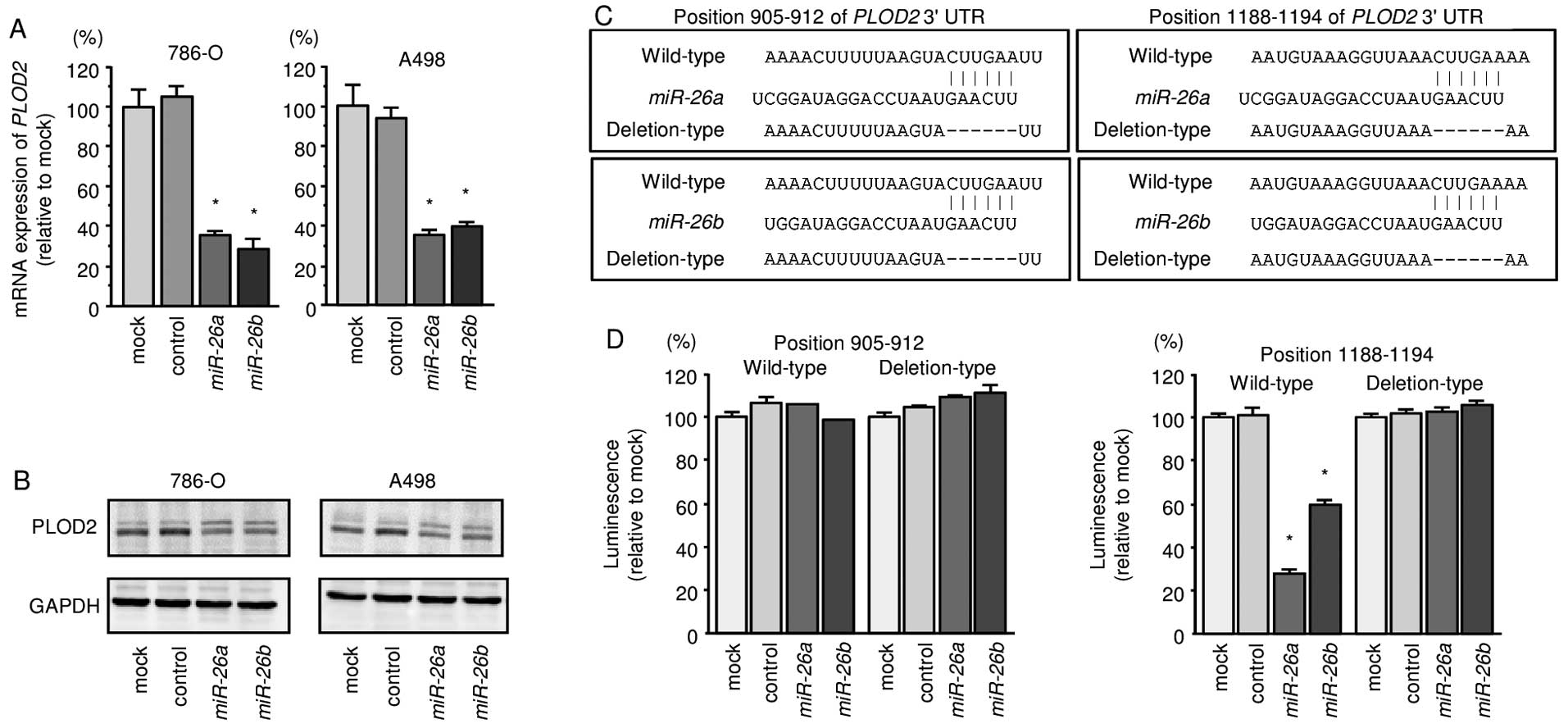
We also found that the mRNA and protein expression
levels of PLOD2/PLOD2 were significantly repressed in
miR-26a or miR-26b transfectant cells in comparison
with mock or miR-control transfectants (Fig. 4A and B). We also observed that the
luminescence intensities were significantly reduced by transfection
with miR-26a or miR-26b and vectors carrying the
wild-type 3′-UTR of PLOD2, whereas transfection with
deletion vectors blocked the decrease in luminescence. These data
suggested that miR-26a or miR-26b bound directly to
specific sites in the 3′-UTR of PLOD2 (Fig. 4C and D).
Silencing PLOD2 affected cell
proliferation, migration and invasion activities in ccRCC
cells
We recently presented a loss-of-function study of
LOXL2 in RCC cells (786-O and A498) by using two siRNAs
(786-O and A498) (12). Those data
showed that the silencing of LOXL2 significantly suppressed
cancer cell migration and invasion activities in RCC cells.
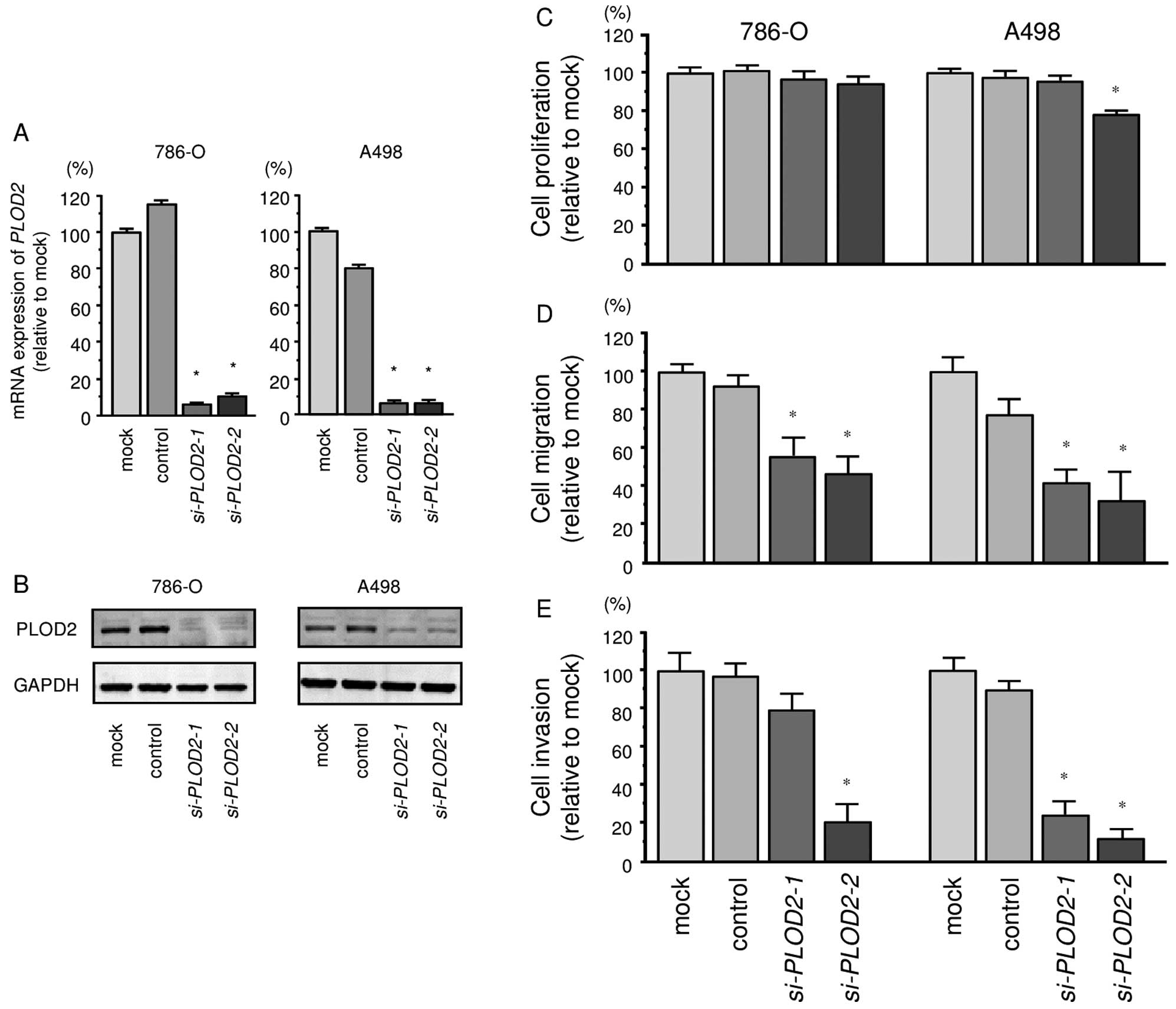
To investigate the functional role of PLOD2
in ccRCC cells, we performed a loss-of-function study using
si-PLOD2 transfected cells. First, we evaluated the
knockdown efficiency of si-PLOD2 transfection in 786-O and
A498 cells. qRT-PCR and western blotting indicated that
si-PLOD2 transfection effectively downregulated PLOD2
expression in both cell lines (786-O, P<0.0001; A498,
P<0.0001; Fig. 5A and B).
The XTT assay demonstrated that cell proliferation
was not inhibited significantly in si-PLOD2 transfectant
cells in comparison with the mock or negative control transfectants
(Fig. 5C).
In contrast, the migration assay demonstrated that
cell migration activity was significantly inhibited in
si-PLOD2 transfectants in comparison with the mock or
negative control transfectants (Fig.
5D). The Matrigel invasion assay demonstrated that invasive
activity was significantly inhibited in si-PLOD2
transfectants in comparison with the mock or negative control
transfectants (Fig. 5E).
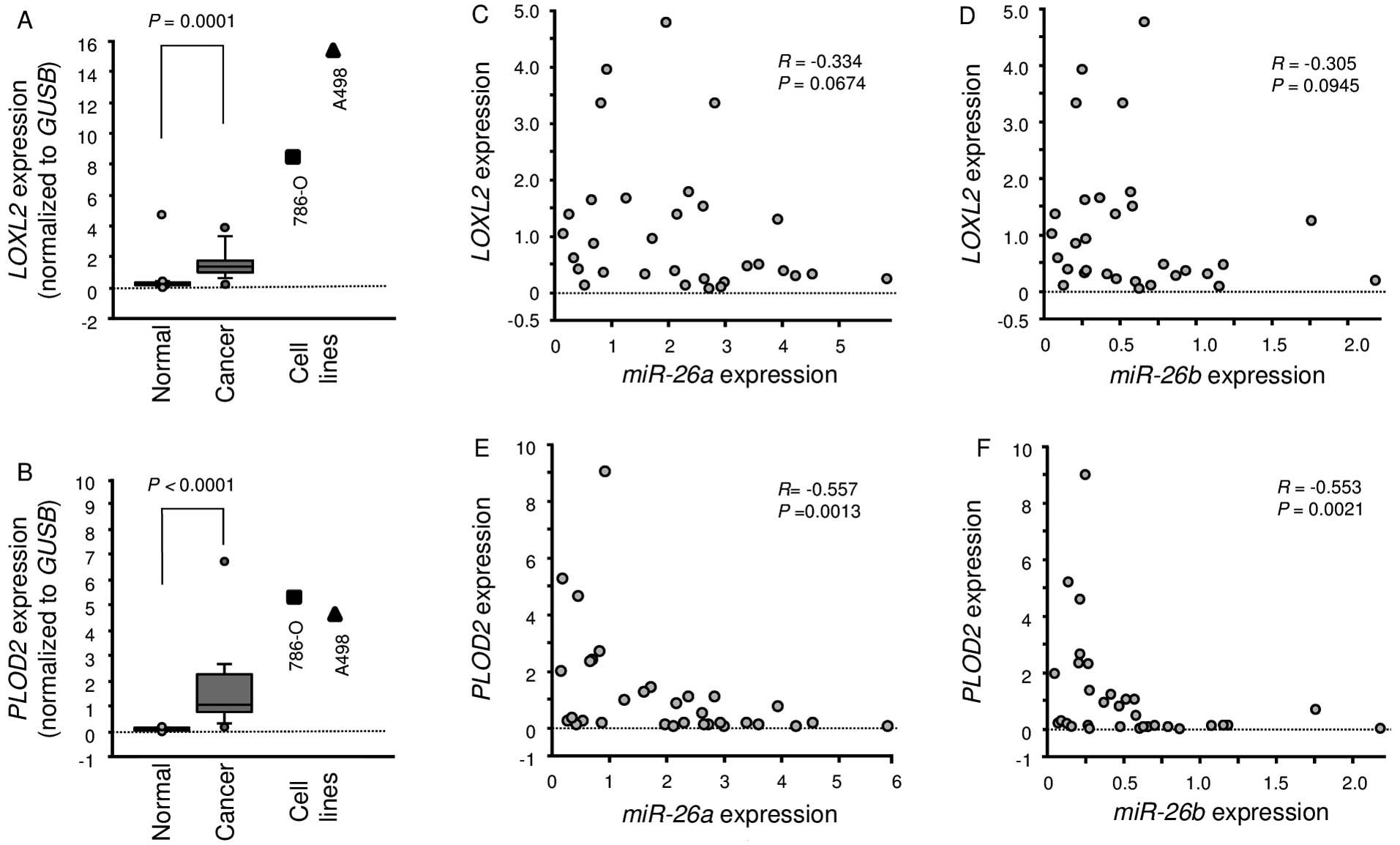
Expression of LOXL2 and PLOD2 in ccRCC
clinical specimens
A total of 15 pairs of ccRCC specimens and
corresponding non-cancerous specimens were used for expression
studies of LOXL2 and PLOD2 using RT-PCR. We showed
that LOXL2 and PLOD2 were significantly upregulated
in cancer tissues compared with normal tissues (P=0.0001 and
P<0.0001, respectively; Fig. 6A and
B). Furthermore, Spearman's rank test showed a negative
correlation between the expression of miR-26a/PLOD2 and
miR-26b/PLOD2 (Fig. 6E and
F).
Discussion
A growing body of evidence has shown that aberrantly
expressed miRNAs can disrupt tightly regulated RNA networks in
cancer cells and promote human oncogenesis and metastasis (7,9,24–26).
Recently, our studies identified a variety of novel RCC molecular
pathways regulated by tumor-suppressive miRNAs (12–15).
In the present study, we focused on miR-26a and
miR-26b because the expression levels of these miRNAs were
reduced in the miRNA signatures of various types of cancers
(10,18,19,27).
Moreover, the functional roles of these miRNAs in RCC cells are not
clear. Our present data showed that miR-26a and
miR-26b act as tumor suppressors that modulate cancer cell
migration and invasion in RCC cells. Our previous studies of oral
cancer and prostate cancer demonstrated the tumor-suppressive roles
of these miRNAs (19,20), and those findings support the
present results obtained with RCC cells. Downregulation and
tumor-suppressive roles of miR-26a or miR-26b have
been reported in several types of cancer, such as bladder, breast,
hepatocellular carcinoma and oral cancer (19,28–30).
In the human genome, the miR-26 family
consists of three subtypes of miRNAs: miR-26a-1,
miR-26a-2 and miR-26b. The mature sequences of
miR-26a-1 and miR-26a-2 are identical, whereas the
two nucleotides differ from that of miR-26b (miRBase release
21; http://www.mirbase.org/). The molecular
mechanisms responsible for silencing the expression of the
miR-26 family are still unclear. A recent study indicated
that MYC oncogene directly bound to the promoter regions of
miR-26a-1, miR-26a-2 and miR-26b and
negatively regulated expression of these miRNAs in prostate cancer
cells (31). Overexpression of
MYC was observed in RCC clinical specimens (15,32),
suggesting MYC might be a mediator for expression control of
tumor-suppressive miRNAs in cancer cells.
A single miRNA may regulate multiple protein-coding
genes; indeed, bioinformatics studies have shown that miRNAs
regulate >30–60% of the protein-coding genes in the human genome
(7,33). Reduced expression of
tumor-suppressive miRNAs may cause overexpression of oncogenic
genes in cancer cells. To better understand RCC oncogenesis and
metastasis, we identified miR-26a and miR-26b target
genes using in silico analysis. Recent miRNA studies in our
laboratory have utilized this strategy to identify novel molecular
targets and pathways regulated by tumor-suppressive miRNAs in
several cancers, including RCC (12,20).
A total of 39 putative target genes of
miR-26a and miR-26b were identified in the present
study. Among these genes, we focused on LOXL2 and
PLOD2 because they function as collagen cross-linking
enzymes. Numerous studies have shown that aberrant expression of
collagen cross-linking enzymes promotes extracellular matrix (ECM)
stiffening, resulting in enhanced cancer cell migration and
invasion (22,34–39).
Overexpression of ECM components has been observed in several
cancers (21,23,40).
Recently, a number of studies indicated that several miRNAs
regulated ECM component genes, and aberrantly expressed miRNAs have
contributed to cancer cell progression by dysregulation of cell
adhesion, polarity and ECM remodeling (21,23).
Our past studies found that the tumor-suppressive
miR-29-family (miR-29a, miR-29b and
miR-29c) and miR-218 directly regulated laminins
(LAMC2 and LAMB3) and integrins (ITGA6 and
ITGB3), such that restoration of these miRNAs inhibited
cancer cell migration and invasion (21,41,42).
Once collagen is secreted, collagen cross-linking
occurs on lysine and hydroxylysine residues by the lysyl oxidase
(LOX) family of enzymes (22,43).
More recently, we showed that the miR-29s-family directly
targeted LOXL2 in RCC and lung cancers (12). Overexpression of LOXL2 was
observed in RCC clinical specimens and silencing of LOXL2
inhibited cancer cell migration and invasion in ccRCC cell lines
(12). Other research groups found
that increased expression of LOXL2 is correlated with
disease progression, including RCC (34,44).
The function of the LOX-family is covalent crosslinking of collagen
and/or elastin in the ECM (35,36).
Aberrant expression of LOX-family proteins has been reported in
several diseases, including cancers (34–39).
Interestingly, LOXL2 is a direct transcriptional target of
HIF-1. Moreover, nuclear LOXL2 interacts with transcription factor
SNAIL1 and represses E-cadherin as well as induces EMT (45,46).
In this study, we demonstrated direct regulation of LOXL2 by
miR-26a and miR-26b in RCC cells as observed with the
miR-29s-family. These findings showed that tumor-suppressive
miR-26a/b-LOXL2 is the pivotal pathway contributing to
cancer cell migration and invasion in RCC.
In this study, we also focused on the PLOD2
(procollagen-lysine 2-oxyglutarate-dioxygenase) gene as a target of
miR-26a and miR-26b and demonstrated the direct
regulation of these miRNAs by luciferase reporter assays.
PLOD2 encodes an enzyme that mediates collagen lysine
hydroxylation. Collagen cross-linking that are derived from
hydroxylated lysine residues have increased stability compared with
non-hydroxylated lysine residues (22,47).
Overexpression of PLOD2 in ccRCC clinical specimens and
promoting migration and invasion in cancer cells were observed in
the present study. In breast cancer, Kaplan-Meier curves of
disease-specific survival stratified by PLOD2 expression
revealed that high PLOD2 expression was significantly
associated with decreased disease-specific survival (48). Moreover, PLOD2 expression
promoted tumor stiffness and was required for metastasis to lymph
nodes and lungs (22,48).
In conclusion, miR-26a and miR-26b
were significantly downregulated in ccRCC clinical specimens and
appeared to function as tumor suppressors through regulation of
collagen cross-linking enzymes, LOXL2 and PLOD2, both
of which function as oncogenes in this disease. The identification
of novel molecular targets and pathways regulated by the
tumor-suppressive miR-26a and miR-26b may lead to a
better understanding of ccRCC and the development of new
therapeutic strategies to treat this disease.
Acknowledgements
The present study was supported by the Japanese
Society for the Promotion of Science (KAKENHI), grant numbers (C)
15K10801, (C) 15K20070, (C) 15K20071, and (B) 25293333.
References
|
1
|
Rini BI, Campbell SC and Escudier B: Renal
cell carcinoma. Lancet. 373:1119–1132. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Randall JM, Millard F and Kurzrock R:
Molecular aberrations, targeted therapy, and renal cell carcinoma:
Current state-of-the-art. Cancer Metastasis Rev. 33:1109–1124.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Gupta K, Miller JD, Li JZ, Russell MW and
Charbonneau C: Epidemiologic and socioeconomic burden of metastatic
renal cell carcinoma (mRCC): A literature review. Cancer Treat Rev.
34:193–205. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Motzer RJ and Russo P: Systemic therapy
for renal cell carcinoma. J Urol. 163:408–417. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Motzer RJ, Hutson TE, Tomczak P,
Michaelson MD, Bukowski RM, Oudard S, Negrier S, Szczylik C, Pili
R, Bjarnason GA, et al: Overall survival and updated results for
sunitinib compared with interferon alfa in patients with
meta-static renal cell carcinoma. J Clin Oncol. 27:3584–3590. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Motzer RJ, Hutson TE, Tomczak P,
Michaelson MD, Bukowski RM, Rixe O, Oudard S, Negrier S, Szczylik
C, Kim ST, et al: Sunitinib versus interferon alfa in metastatic
renal-cell carcinoma. N Engl J Med. 356:115–124. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Filipowicz W, Bhattacharyya SN and
Sonenberg N: Mechanisms of post-transcriptional regulation by
microRNAs: Are the answers in sight? Nat Rev Genet. 9:102–114.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Nelson KM and Weiss GJ: MicroRNAs and
cancer: Past, present, and potential future. Mol Cancer Ther.
7:3655–3660. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Goto Y, Kurozumi A, Enokida H, Ichikawa T
and Seki N: Functional significance of aberrantly expressed
microRNAs in prostate cancer. Int J Urol. 22:242–252. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Esquela-Kerscher A and Slack FJ: Oncomirs
- microRNAs with a role in cancer. Nat Rev Cancer. 6:259–269. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Nishikawa R, Chiyomaru T, Enokida H,
Inoguchi S, Ishihara T, Matsushita R, Goto Y, Fukumoto I, Nakagawa
M and Seki N: Tumour-suppressive microRNA-29s directly regulate
LOXL2 expression and inhibit cancer cell migration and invasion in
renal cell carcinoma. FEBS Lett. 589:2136–2145. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yamasaki T, Seki N, Yoshino H, Itesako T,
Hidaka H, Yamada Y, Tatarano S, Yonezawa T, Kinoshita T, Nakagawa
M, et al: MicroRNA-218 inhibits cell migration and invasion in
renal cell carcinoma through targeting caveolin-2 involved in focal
adhesion pathway. J Urol. 190:1059–1068. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yoshino H, Enokida H, Itesako T, Tatarano
S, Kinoshita T, Fuse M, Kojima S, Nakagawa M and Seki N:
Epithelial-mesenchymal transition-related microRNA-200s regulate
molecular targets and pathways in renal cell carcinoma. J Hum
Genet. 58:508–516. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yamada Y, Hidaka H, Seki N, Yoshino H,
Yamasaki T, Itesako T, Nakagawa M and Enokida H: Tumor-suppressive
microRNA-135a inhibits cancer cell proliferation by targeting the
c-MYC oncogene in renal cell carcinoma. Cancer Sci. 104:304–312.
2013. View Article : Google Scholar
|
|
16
|
Yoshino H, Enokida H, Itesako T, Kojima S,
Kinoshita T, Tatarano S, Chiyomaru T, Nakagawa M and Seki N:
Tumor-suppressive microRNA-143/145 cluster targets hexokinase-2 in
renal cell carcinoma. Cancer Sci. 104:1567–1574. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ishihara T, Seki N, Inoguchi S, Yoshino H,
Tatarano S, Yamada Y, Itesako T, Goto Y, Nishikawa R, Nakagawa M,
et al: Expression of the tumor suppressive miRNA-23b/27b cluster is
a good prognostic marker in clear cell renal cell carcinoma. J
Urol. 192:1822–1830. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Kikkawa N, Hanazawa T, Fujimura L, Nohata
N, Suzuki H, Chazono H, Sakurai D, Horiguchi S, Okamoto Y and Seki
N: miR-489 is a tumour-suppressive miRNA target PTPN11 in
hypopharyngeal squamous cell carcinoma (HSCC). Br J Cancer.
103:877–884. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Fukumoto I, Hanazawa T, Kinoshita T,
Kikkawa N, Koshizuka K, Goto Y, Nishikawa R, Chiyomaru T, Enokida
H, Nakagawa M, et al: MicroRNA expression signature of oral
squamous cell carcinoma: Functional role of microRNA-26a/b in the
modulation of novel cancer pathways. Br J Cancer. 112:891–900.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kato M, Goto Y, Matsushita R, Kurozumi A,
Fukumoto I, Nishikawa R, Sakamoto S, Enokida H, Nakagawa M,
Ichikawa T, et al: MicroRNA-26a/b directly regulate La-related
protein 1 and inhibit cancer cell invasion in prostate cancer. Int
J Oncol. 47:710–718. 2015.PubMed/NCBI
|
|
21
|
Kinoshita T, Hanazawa T, Nohata N, Kikkawa
N, Enokida H, Yoshino H, Yamasaki T, Hidaka H, Nakagawa M, Okamoto
Y, et al: Tumor suppressive microRNA-218 inhibits cancer cell
migration and invasion through targeting laminin-332 in head and
neck squamous cell carcinoma. Oncotarget. 3:1386–1400. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Gilkes DM, Semenza GL and Wirtz D: Hypoxia
and the extracellular matrix: Drivers of tumour metastasis. Nat Rev
Cancer. 14:430–439. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kurozumi A, Goto Y, Matsushita R, Fukumoto
I, Kato M, Nishikawa R, Sakamoto S, Enokida H, Nakagawa M, Ichikawa
T, et al: Tumor-suppressive microRNA-223 inhibits cancer cell
migration and invasion by targeting ITGA3/ITGB1 signaling in
prostate cancer. Cancer Sci. 107:84–94. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wiemer EA: The role of microRNAs in
cancer: No small matter. Eur J Cancer. 43:1529–1544. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Singh SK, Pal Bhadra M, Girschick HJ and
Bhadra U: MicroRNAs - micro in size but macro in function. FEBS J.
275:4929–4944. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Iorio MV and Croce CM: MicroRNAs in
cancer: Small molecules with a huge impact. J Clin Oncol.
27:5848–5856. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Fuse M, Kojima S, Enokida H, Chiyomaru T,
Yoshino H, Nohata N, Kinoshita T, Sakamoto S, Naya Y, Nakagawa M,
et al: Tumor suppressive microRNAs (miR-222 and miR-31) regulate
molecular pathways based on microRNA expression signature in
prostate cancer. J Hum Genet. 57:691–699. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Lin Y, Chen H, Hu Z, Mao Y, Xu X, Zhu Y,
Xu X, Wu J, Li S, Mao Q, et al: miR-26a inhibits proliferation and
motility in bladder cancer by targeting HMGA1. FEBS Lett.
587:2467–2473. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhang B, Liu XX, He JR, Zhou CX, Guo M, He
M, Li MF, Chen GQ and Zhao Q: Pathologically decreased miR-26a
antagonizes apoptosis and facilitates carcinogenesis by targeting
MTDH and EZH2 in breast cancer. Carcinogenesis. 32:2–9. 2011.
View Article : Google Scholar
|
|
30
|
Zhu Y, Lu Y, Zhang Q, Liu JJ, Li TJ, Yang
JR, Zeng C and Zhuang SM: MicroRNA-26a/b and their host genes
cooperate to inhibit the G1/S transition by activating the pRb
protein. Nucleic Acids Res. 40:4615–4625. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Koh CM, Iwata T, Zheng Q, Bethel C,
Yegnasubramanian S and De Marzo AM: Myc enforces overexpression of
EZH2 in early prostatic neoplasia via transcriptional and
post-transcriptional mechanisms. Oncotarget. 2:669–683. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tang SW, Chang WH, Su YC, Chen YC, Lai YH,
Wu PT, Hsu CI, Lin WC, Lai MK and Lin JY: MYC pathway is activated
in clear cell renal cell carcinoma and essential for proliferation
of clear cell renal cell carcinoma cells. Cancer Lett. 273:35–43.
2009. View Article : Google Scholar
|
|
33
|
Friedman RC, Farh KK, Burge CB and Bartel
DP: Most mammalian mRNAs are conserved targets of microRNAs. Genome
Res. 19:92–105. 2009. View Article : Google Scholar :
|
|
34
|
Barker HE, Cox TR and Erler JT: The
rationale for targeting the LOX family in cancer. Nat Rev Cancer.
12:540–552. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Kagan HM and Li W: Lysyl oxidase:
Properties, specificity, and biological roles inside and outside of
the cell. J Cell Biochem. 88:660–672. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Vadasz Z, Kessler O, Akiri G,
Gengrinovitch S, Kagan HM, Baruch Y, Izhak OB and Neufeld G:
Abnormal deposition of collagen around hepatocytes in Wilson's
disease is associated with hepatocyte specific expression of lysyl
oxidase and lysyl oxidase like protein-2. J Hepatol. 43:499–507.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Kim YM, Kim EC and Kim Y: The human lysyl
oxidase-like 2 protein functions as an amine oxidase toward
collagen and elastin. Mol Biol Rep. 38:145–149. 2011. View Article : Google Scholar
|
|
38
|
Fong SF, Dietzsch E, Fong KS, Hollosi P,
Asuncion L, He Q, Parker MI and Csiszar K: Lysyl oxidase-like 2
expression is increased in colon and esophageal tumors and
associated with less differentiated colon tumors. Genes Chromosomes
Cancer. 46:644–655. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Peinado H, Moreno-Bueno G, Hardisson D,
Pérez-Gómez E, Santos V, Mendiola M, de Diego JI, Nistal M,
Quintanilla M, Portillo F, et al: Lysyl oxidase-like 2 as a new
poor prognosis marker of squamous cell carcinomas. Cancer Res.
68:4541–4550. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Liu LX, Jiang HC, Liu ZH, Zhou J, Zhang
WH, Zhu AL, Wang XQ and Wu M: Integrin gene expression profiles of
human hepatocellular carcinoma. World J Gastroenterol. 8:631–637.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Kinoshita T, Nohata N, Hanazawa T, Kikkawa
N, Yamamoto N, Yoshino H, Itesako T, Enokida H, Nakagawa M, Okamoto
Y, et al: Tumour-suppressive microRNA-29s inhibit cancer cell
migration and invasion by targeting laminin-integrin signalling in
head and neck squamous cell carcinoma. Br J Cancer. 109:2636–2645.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Yamamoto N, Kinoshita T, Nohata N, Itesako
T, Yoshino H, Enokida H, Nakagawa M, Shozu M and Seki N: Tumor
suppressive microRNA-218 inhibits cancer cell migration and
invasion by targeting focal adhesion pathways in cervical squamous
cell carcinoma. Int J Oncol. 42:1523–1532. 2013.PubMed/NCBI
|
|
43
|
Gordon MK and Hahn RA: Collagens. Cell
Tissue Res. 339:247–257. 2010. View Article : Google Scholar :
|
|
44
|
Hase H, Jingushi K, Ueda Y, Kitae K, Egawa
H, Ohshio I, Kawakami R, Kashiwagi Y, Tsukada Y, Kobayashi T, et
al: LOXL2 status correlates with tumor stage and regulates integrin
levels to promote tumor progression in ccRCC. Mol Cancer Res.
12:1807–1817. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Schietke R, Warnecke C, Wacker I, Schödel
J, Mole DR, Campean V, Amann K, Goppelt-Struebe M, Behrens J,
Eckardt KU, et al: The lysyl oxidases LOX and LOXL2 are necessary
and sufficient to repress E-cadherin in hypoxia: Insights into
cellular transformation processes mediated by HIF-1. J Biol Chem.
285:6658–6669. 2010. View Article : Google Scholar :
|
|
46
|
Moon HJ, Finney J, Xu L, Moore D, Welch DR
and Mure M: MCF-7 cells expressing nuclear associated lysyl
oxidase-like 2 (LOXL2) exhibit an epithelial-to-mesenchymal
transition (EMT) phenotype and are highly invasive in vitro. J Biol
Chem. 288:30000–30008. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
van der Slot AJ, Zuurmond AM, van den
Bogaerdt AJ, Ulrich MM, Middelkoop E, Boers W, Karel Ronday H,
DeGroot J, Huizinga TW and Bank RA: Increased formation of
pyridinoline cross-links due to higher telopeptide lysyl
hydroxylase levels is a general fibrotic phenomenon. Matrix Biol.
23:251–257. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Gilkes DM, Bajpai S, Wong CC, Chaturvedi
P, Hubbi ME, Wirtz D and Semenza GL: Procollagen lysyl hydroxylase
2 is essential for hypoxia-induced breast cancer metastasis. Mol
Cancer Res. 11:456–466. 2013. View Article : Google Scholar : PubMed/NCBI
|