Introduction
Cervical cancer is the fourth most common cancer
type among women worldwide, with 570,000 new cases and 311,000
deaths estimated in 2018; however, the incidence and mortality
rates have decreased over the last few decades due to effective
integration of human papillomavirus (HPV)-based testing through
screening and HPV vaccination programs in a number of developed
countries (1). Persistent
infection with HPV is a primary cause of cervical cancer and its
precursor lesions, and HPV testing is used for primary screening
(2). HPV infection is common in
sexually active women, with an estimated incidence rate of 74-80%,
but cervical cancer is comparatively rare, as 90% of HPV infections
clear within two years (3,4). Currently, it is not possible to
predict which women who test positive for HPV may develop cervical
cancer. Therefore, improved understanding of the underlying
molecular mechanisms may facilitate the development of novel
diagnostic markers and effective therapeutic targets for
HPV-associated cancer.
Our previous study performed RNA sequencing to
analyze the transcriptomes in healthy cervical tissues without HPV
infection and cervical cancer tissues with high-risk HPV (HR-HPV)
infection, and identified family with sequence similarity 83 member
A (FAM83A) as one of the most upregulated genes in cervical cancer
compared with healthy tissues (5).
FAM83A is one of the eight-member FAM83 family of proteins; this
434-amino acid protein shares a conserved N-terminal domain of
unknown function, termed the DUF1669 domain, with other FAM83
members and also contains serine-rich and proline-rich domains
(6,7). FAM83A was originally identified as a
novel tumor-specific gene BJ-TSA-9, located on chromosome 8q24, and
was demonstrated to be highly expressed in lung cancer using
bioinformatics (8). Additionally,
FAM83A upregulation was identified in multiple human tumor types,
including breast (6,9,10),
pancreatic (11) and ovarian
(12) cancer, as well as lung
adenocarcinoma (13). Using a 3D
laminin-rich extracellular matrix culture system, Lee et al
(6) revealed that FAM83A was able
to confer resistance to epidermal growth factor receptor tyrosine
kinase inhibitors (EGFR-TKIs) via its putative interactions with
proto-oncogene c-RAF and PI3K p85 in breast cancer. FAM83A
depletion in breast cancer cells leads to suppressed proliferation
and invasiveness in vitro as well as to suppressed tumor
growth in vivo (14). In
addition, analysis of a published breast cancer gene expression
dataset reported that patients with breast cancer with high
expression of FAM83A exhibited a poor clinical prognosis (15).
Based on the aforementioned studies, FAM83A is
considered to be a candidate oncogene. However, the cellular
mechanism of action of FAM83A in human cancer is not fully
understood. The present study aimed to identify the functional
roles and mechanisms of FAM83A in cervical cancer cells. The
results of the present study may provide a potential target for
cervical cancer therapeutics.
Materials and methods
Human samples
A total of 153 human cervical samples without
preoperative treatment were obtained by surgical resection from
patients with a mean age of 45 (range 28-66) years at the
Department of Gynecologic Oncology at the Women's Hospital,
Zhejiang University between January 2012 and December 2013. The
Ethics Committee of the Women's Hospital approved the collection
and use of human samples. All patients signed informed consent.
Diagnosis of all samples was confirmed independently by two
pathologists before further analysis.
RNA extraction and reverse
transcription-quantitative PCR (RT-qPCR)
RNA extraction and RT-qPCR was performed as
previously described (16).
Briefly, total RNA was extracted from tissues or cells using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.). cDNA synthesis was performed using a PrimeScript RT reagent
kit (Takara Biotechnology Co., Ltd.) according to the
manufacturer's instructions. The samples were analyzed on an
ABI/ViiA7 Real-Time PCR System (Applied Biosystems; Thermo Fisher
Scientific, Inc.) using a SYBR® Premix Ex Taq kit
(Takara Biotechnology Co., Ltd.). The thermocycling conditions were
as follows: Denaturation at 95°C for 30 sec, followed by 40 cycles
of 95°C for 5 sec and 60°C for 30 sec, and the melting curve stage
at 95°C for 15 sec, 60°C for 1 min and 95°C for 15 sec. Gene
quantification was performed using the 2−∆∆Cq method
(16). GAPDH was used as an
internal control. The primers used are presented in Table SI.
Immunohistochemistry (IHC)
IHC was performed on paraffin-embedded human
cervical tissue sections and cervical tumor xenografts with the use
of the primary antibodies. Briefly, each 4 μm-section was
deparaffinized in xylene and rehydrated with 100, 85 and 75%
ethanol in deionized water. The slides were placed in antigen
retrieval solution, boiled at 100°C for 15 min and subsequently
incubated with primary antibodies including anti-FAM83A (1:800;
cat. no. bs-16014R; BIOSS), integrin α1 (1:100; cat. no.
22146-I-Ap; ProteinTech Group, Inc.), integrin α3 (1:1,000; cat.
no. 66070-I-Ig; ProteinTech Group, Inc.), integrin α5 (1:100; cat.
no. ET1701-58; Hangzhou HuaAn Biotechnology Co., Ltd.), integrin β4
(1:50; cat. no. ET1703-52; Hangzhou HuaAn Biotechnology Co., Ltd.)
and integrin β5 (1:400; cat. no. bs-23987R; BIOSS) at 4°C
overnight, followed by incubation with Dako Envision Peroxidase
(Dako; Agilent Technologies, Inc.) for 1 h at room temperature. The
antibody staining was visualized with DAB (Dako; Agilent
Technologies, Inc.). All section slides were counterstained in
hematoxylin. The IHC results were evaluated and scored based on the
intensity of staining and the proportion of stained positive cells
in a blinded manner by a technician (17). The frequency of positive cells were
defined as follows: <5%, 0 points; 5-30%, 1 point; 31-70%, 2
points; >71%, 3 points. The staining intensity was scored as
follows: No staining, 0 points; light brown staining, 1 point;
brown staining, 2 points; dark brown staining, 3 points. The
staining result was determined by multiplying the score for the
frequency of positive cells with the score for staining intensity.
The expression of FAM83A were graded as 'none' (score, 0), 'weak'
(score, 1-3), 'moderate' (score, 4-6) or 'strong' (score, 7-9).
'Strong' staining was defined as 'high expression', whereas 'weak'
or 'moderate' staining was considered 'low expression'.
Cell lines
Human cervical cancer cell lines HeLa and CaSki were
purchased from the American Type Cell Culture and cultured in DMEM
(Thermo Fisher Scientific, Inc.) supplemented with 10% FBS (Gibco;
Thermo Fisher Scientific, Inc.). Cells were cultured at 37°C with
5% CO2. Both cell lines were negative for
mycoplasma.
Small interfering (si)RNA
transfection
FAM83A siRNA pool (si-FAM83A; cat. no.
M-015040-00-0005) and non-targeting siRNA pool (si-NS; cat. no.
D-001206-13-05) were purchased from GE Healthcare Dharmacon, Inc.
HeLa and CaSki cells (3×105) were plated in 6-well
plates and transfected with 50 μM si-NS or si-FAM83A using
the DharmaFECT 1 transfection reagent (cat. no. T-2001-03; GE
Healthcare Dharmacon, Inc) and incubated at 37°C in a humidified
CO2 incubator for 24 h according to the manufacturer's
instructions. The RNA was extracted at 48 h post-transfection, and
protein was obtained at 72 h post-transfection. The transfection
efficiency was confirmed by RT-qPCR and western blotting.
Cell proliferation, migration and
invasion assays
To determine cell proliferation, 5,000 HeLa and
CaSki cells were plated in triplicate in 96-well plates, and a Cell
Counting Kit-8 (CCK-8) assay (Dojindo Molecular Technologies, Inc.)
was performed at 24, 48, 72 and 96 h post-transfection. All
experiments were performed in triplicate and repeated three times
independently.
Migration assays were performed using Transwell
filter chambers (Corning, Inc.). HeLa and CaSki cells were
trans-fected with 50 nM siRNA in 12-well plates for 24 h, and cell
viability was determined with a Trypan blue assay prior to the
migration assay. A total of 1×105 cells with similar
viability in 200 μl serum-free DMEM in each group were then
seeded in the upper chamber of each Transwell, and 500 μl
medium supplemented with 10% FBS was added to the lower chamber.
After 16 h incubation for HeLa and 24 h for CaSki cells, the
non-migrated cells were removed by cotton swabs, and cells that
migrated through the membrane were fixed by 4% paraformaldehyde for
20 min at room temperature, stained with 0.1% crystal violet for 30
min at room temperature, and counted in five random visual fields
per well under a light microscope (×100 magnification; Leica
Microsystems GmbH).
Invasion assays were performed using Transwell
filter chambers pre-coated with Matrigel for 30 min at 37°C. The
detection was performed following incubation for 24 h of HeLa and
36 h for CaSki cells. All migration and invasion experiments were
performed in duplicate and repeated independently three times.
Xenograft models
BALB/c female nude mice (n=12; age, 4 weeks) were
purchased from Shanghai Laboratory Animal Center. All mice were
housed in a dedicated SPF facility (temperature, 22±1°C; humidity,
50±10%; light/dark cycle, 12:12 h) with standard laboratory food
and water, and used with the approval of the Animal Care and Use
Committee of Zhejiang University. The mice were randomly divided in
two groups in a blinded manner. Human cervical cancer HeLa cells
transfected with scrambled short hairpin (sh)RNA (nc-shRNA) or
FAM83A-shRNA were injected subcutaneously (1×107
cells/inoculum) into the right-side flank of each mouse (n=10).
Tumor growth was detected with calipers every 4 days, and tumor
volume was calculated using the following formula: Volume = (length
× width2)/2. To further evaluate the metastatic
potential of FAM83A, mice (n=2) were intraperitoneally transplanted
with 1×107 tumor cells (HeLa/nc-shRNA and
HeLa/FAM83A-shRNA) suspended in 200 μl PBS. Five weeks after
transplantation, all mice were euthanized by cervical dislocation.
Death was confirmed through close observation of the mice. The mice
were assessed for any movement, including the respiratory movement
of the chest, and checked for heartbeat by a stethoscope. Once the
muscles of the mice stiffened and evidence of animal death was
observed, the tumors were harvested and imaged.
Immunofluorescence staining
Experiments were performed on 4-μm
paraffin-embedded tumor tissues mounted on slides. Paraffin was
removed, and the sections were serially rehydrated by descending
ethanol series as described above. Antigen retrieval was performed
by heating the slides in EDTA buffer at 100°C for 15 min. Slides
were blocked by incubating with 3% BSA (Wuhan Servicebio Technology
Co., Ltd.) for 30 min at 37°C and then incubated with primary
antibodies against FAM83A (1:500; BIOSS), integrin α1 (1:200;
ProteinTech Group, Inc.), integrin α3 (1:1,000; ProteinTech Group,
Inc.), integrin α5 (1:100; Hangzhou HuaAn Biotechnology Co., Ltd.),
integrin β4 (1:100; ProteinTech Group, Inc.) and integrin β5
(1:100; BIOSS) at 4°C overnight, followed by the secondary
antibodies Alexa Fluor 488-conjugated goat anti-mouse (1:400; cat.
no. GB25303) or Cy-3-conjugated goat anti-rabbit (1:300; cat. no.
GB21303; both from Wuhan Servicebio Technology Co., Ltd.) for 1 h
at room temperature. The cell nuclei were stained with DAPI. Nikon
Eclipse C1 fluorescence microscope with an integrated camera was
used to visualize the fluorescence and acquire images from five
representative fields of each sections (×400).
RNA sequencing (RNA-seq)
After transiently transfecting HeLa cells with
si-FAM83A or si-NS for 48 h, total RNA of each group was extracted
and subjected to poly(A) selection using Oligo(dT) beads (Thermo
Fisher Scientific, Inc.) according to the manufacturer's protocol.
RNA libraries were sequenced on the Illumina HiSeq/MiSeq platform
at Novogene Co., Ltd. RNA-seq reads in fasta format were mapped to
the human genome GRCh38 using TopHat (version 2.0.13) (18). DESeq was used for differential
expression (DE) analysis (19). In
DE tests, a gene was considered significantly differentially
expressed when the adjusted P<0.05. Integrative Genomics Viewer
(20) was used to visualize the
expression of FAM83A in each group.
Western blot analysis
At 72 h post-transfection, HeLa and CaSki cells were
lysed in RIPA buffer in presence of protease and phosphatase
inhibitors (ThermoFisher Scientific, Inc.). The protein
concentration was quantified using a BCA protein assay kit (Thermo
Fisher Scientific, Inc.). Western blotting was performed following
a standard procedure (21). The
membranes were blocked with 5% skimmed milk for 1 h at room
temperature and incubated with the primary antibodies overnight at
4°C, washed with TBS-Tween-20 and incubated with the secondary
antibodies for 1 h at room temperature. The following antibodies
were used: FAM83A (1:1,000; cat. no. SAB1103067; Sigma-Aldrich;
Merck KGaA), integrin α1 (1:500; ProteinTech Group, Inc.), integrin
α2 (1:1,000; cat. no. ET1611-57; Hangzhou HuaAn Biotechnology Co.,
Ltd.), integrin α3 (1:1,000; ProteinTech Group, Inc.), integrin α5
(1:1,000; Hangzhou HuaAn Biotechnology Co., Ltd.), integrin β4
(1:1,000; Hangzhou HuaAn Biotechnology Co., Ltd.), integrin β5
(1:1,000; BIOSS), phosphorylated (p)-PI3K (1:1,000; cat. no. 4228;
Cell Signaling Technology, Inc.), p-AKT (1:1,000; cat. no. 4060;
Cell Signaling Technology, Inc.), GAPDH (1:4,000; cat. no.
sc-47724; Santa Cruz Biotechnology, Inc.), horseradish peroxidase
(HRP)-conjugated anti-mouse IgG (1:1,000; Cell Signaling
Technology, Inc.; cat. no. 7076) and anti-rabbit IgG (1:1,000; cat.
no. 7074; Cell Signaling Technology, Inc.). Blots were visualized
with SuperSignal West Pico Chemiluminescent Substrate (Pierce;
Thermo Fisher Scientific, Inc.) and analyzed on a LAS-4000 Imaging
System (Fuji Photo Film, Inc.). Band intensities were determined by
Quantity One software v4.6.6 (Bio-Rad Laboratories, Inc.).
Statistical analysis
The data obtained using tissue samples are presented
as the mean ± SEM; all other data are presented as the mean ± SD.
The experiments were performed ≥3 times unless stated otherwise.
Analyses were performed using GraphPad Prism 6.0 software (GraphPad
Software, Inc.). Testing involving two groups in cervical tissues
were performed using the nonparametric Mann-Whitney U test.
Association between FAM83A expression and patient
clinicopathological parameters were analyzed by the χ2
test and Fisher's exact probability test. Kaplan-Meier curves and
the log-rank test were used to examine overall survival. For
experiments in cell lines and xenograft models, two-tailed
Student's t-test was used. P<0.05 was considered to indicate a
statistically significant difference.
Results
FAM83A is upregulated in human cervical
cancer tissues, but low expression is associated with a poor
prognosis
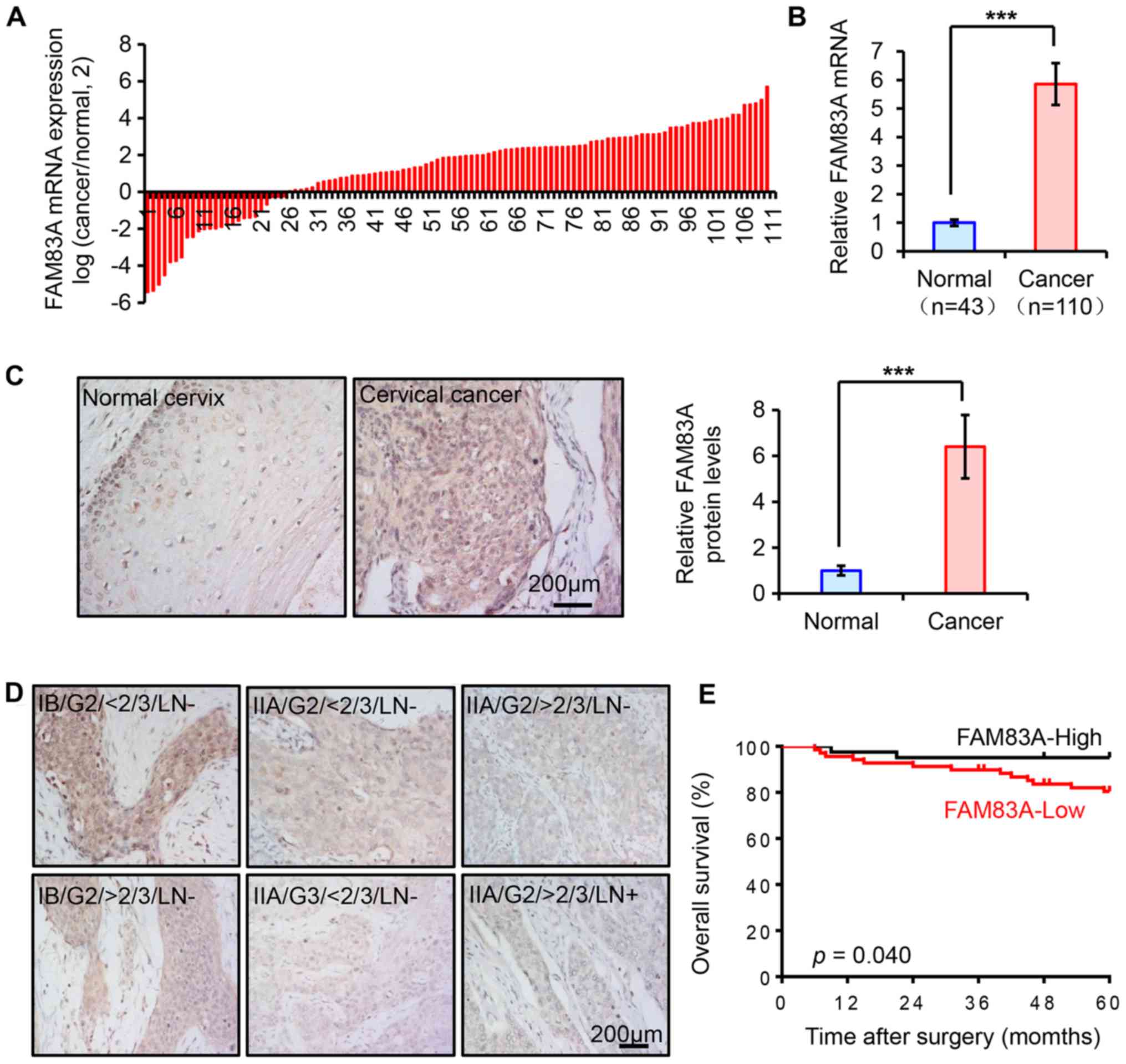
In our previous study, FAM83A mRNA was demonstrated
to be upregulated in cervical cancer by RNA sequencing (5). In the present study, the expression
pattern of FAM83A was assessed in another cohort of human cervical
tissue samples from the Women's Hospital of Zhejiang University.
The mRNA expression levels of FAM83A were quantified using RT-qPCR
in 153 cervical specimens, including 43 healthy and 110 cancerous
cervical tissues. Compared with the healthy cervix, FAM83A mRNA
expression was significantly increased in cancer tissues (Fig. 1A and B).
FAM83A IHC staining was performed in the
aforementioned human cervical samples to determine whether the
protein expression exhibited similar trends. The expression of
FAM83A protein was weak in the healthy cervical epithelia, but
significantly higher in cervical cancer tissues (Fig. 1C). As FAM83A is considered to be an
oncogene, it was hypothesized that its upregulation may be
associated with poor outcomes in patients with cervical cancer.
However, in the present study, the upregulation of FAM83A protein
expression was more likely to occur in patients with favorable
clinical parameters, as the results demonstrated that FAM83A
protein expression was inversely associated with advanced FIGO
stage, deep stromal invasion, high pathological grade and lymph
node metastasis, but not with other clinicopathological parameters
including age, parametrium, margins, lymphovascular space invasion,
tumor size or the presence of squamous cell carcinoma antigen in
patients with cervical cancer (Fig.
1D; Table I). As demonstrated
in Fig. 1D, the FAM83A protein was
located both in the cytoplasm and the nucleus in cervical tissues,
and differences in the localization were observed among groups with
different FIGO stage, stomal invasion and lymph node metastasis .
The intensity of FAM83A protein staining was similar in the cell
nucleus and the cytoplasm of healthy cervical epithelium; in
cervical cancer samples, high FAM83A expression was located both in
nucleus and cytoplasm, but the localization was mainly in cytoplasm
in patients with low FAM83A expression. Kaplan-Meier survival
analysis results indicated that patients with low FAM83A protein
expression exhibited a shorter overall survival time (Fig. 1E). Thus, FAM83A expression was
frequently upregulated in human cervical cancer types, but
negatively associated with poor prognostic parameters, suggesting
the possibility that FAM83A may serve a suppressive role in
cervical cancer.
 | Table IAssociation between FAM83A protein
expression and clinicopathologic parameters of patients with
cervical cancer. |
Table I
Association between FAM83A protein
expression and clinicopathologic parameters of patients with
cervical cancer.
| Characteristic | Total (n=110) | FAM83A protein
expression
| P-value |
|---|
| Low | High |
|---|
| Age, years | | | | 0.369 |
| ≤45 | 61 | 36 | 25 | |
| >45 | 49 | 33 | 16 | |
| Pelvic nodes | | 0.003a | | |
| Negative | 92 | 52 | 40 | |
| Positive | 18 | 17 | 1 | |
| Parametrium | | | | 1.000a |
| Negative | 99 | 62 | 37 | |
| Positive | 11 | 7 | 4 | |
| Margins | | | | 1.000a |
| Negative | 107 | 67 | 40 | |
| Positive | 3 | 2 | 1 | |
| FIGO stageb | | | 0.046 | |
| IA/IB | 76 | 43 | 33 | |
| IIA | 34 | 26 | 8 | |
| Pathological
grade | | | 0.046a | |
| 1/2 | 95 | 56 | 39 | |
| 3 | 15 | 13 | 2 | |
| Lymphovascular
space invasion | | | 0.741 | |
| Negative | 73 | 45 | 28 | |
| Positive | 37 | 24 | 13 | |
| Deep stromal
invasion | | 0.035 | | |
| <2/3 | 58 | 31 | 27 | |
| ≥2/3 | 52 | 38 | 14 | |
| Tumor size, cm | | 0.554 | | |
| ≤4 | 99 | 63 | 36 | |
| >4 | 11 | 6 | 5 | |
| SCC-Ag, ng/ml | | | 0.245a | |
| <4 | 96 | 58 | 38 | |
| ≥4 | 14 | 11 | 3 | |
Knockdown of FAM83A promotes aggressive
phenotypes in cervical cancer cells
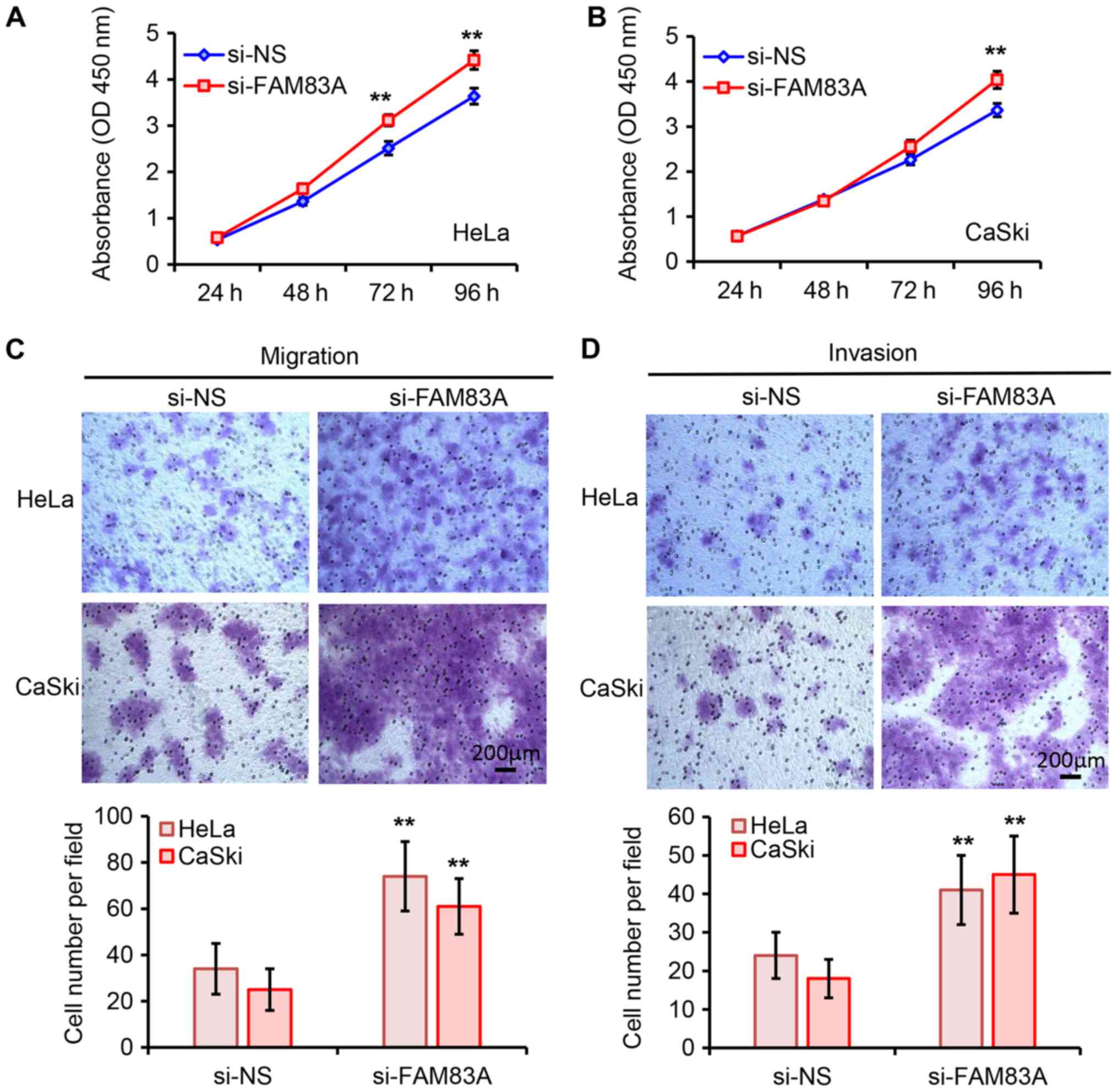
To further elucidate the biological roles of FAM83A
in cervical cancer cells, FAM83A expression in HeLa and CaSki cells
was silenced by transfection with si-FAM83A (Fig. S1). As demonstrated in Fig. 2A and B, knockdown of FAM83A
significantly promoted HeLa and CaSki cell proliferation compared
with that in the negative controls groups. The tumor cell migratory
and invasive abilities were assessed using Transwell assays.
Knockdown of FAM83A significantly increased the migratory and
invasive abilities of HeLa and CaSki cells (Fig. 2C and D). Collectively, these
results suggested that knockdown of FAM83A promoted cervical cancer
cell proliferation and invasion in vitro.
Knockdown of FAM83A promotes tumor growth
and invasion in vivo
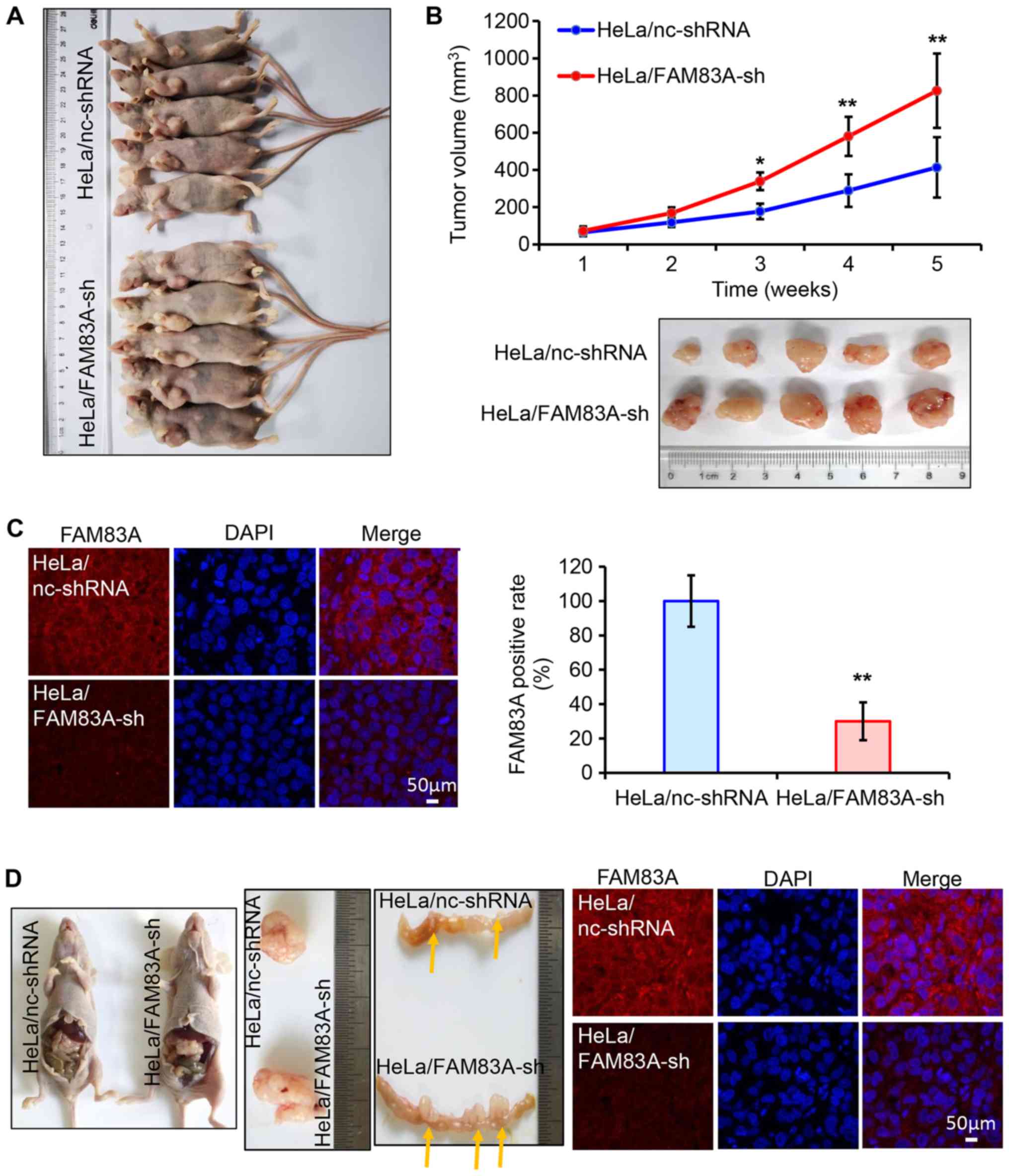
Subsequently, the role of FAM83A knockdown in
cervical tumor growth and invasion was evaluated in vivo.
BALB/c nude mice (n=10) were randomly injected subcutaneously with
HeLa cells stably expressing either control or FAM83A shRNA (each
n=5). Of note, HeLa/FAM83A-sh tumors exhibited more progressive
growth compared with that of HeLa/nc-shRNA tumors, as the volumes
of the tumors generated from HeLa/FAM83A-sh cells were higher
compared with those of tumors from HeLa/nc-shRNA cells (Fig. 3A and B). Immunofluorescence
staining in tumors dissected from the mice identified that tumors
transfected with FAM83A shRNA exhibited lower FAM83A protein
expression compared with those from the control group (Fig. 3C).
The invasive potential of FAM83A was determined
in vivo. HeLa/FAM83A-sh cells and HeLa/nc-shRNA cells were
intraperitoneally transplanted into nude mice (n=2). The volume of
the abdominal HeLa/FAM83A-sh tumor was larger compared with that of
the HeLa/nc-shRNA tumor (Fig. 3D).
In addition, HeLa/FAM83A-sh cells significantly increased the
formation of tumor nodules in the colon compared with that in the
HeLa/nc-shRNA group (Fig. 3D). The
expression of FAM83A in these tumors was also confirmed by
immunofluorescence. These results demonstrated that knockdown of
FAM83A potentiated cervical tumor growth and invasion.
RNA-seq demonstrates that FAM83A
knockdown alters key oncogenic signaling pathways
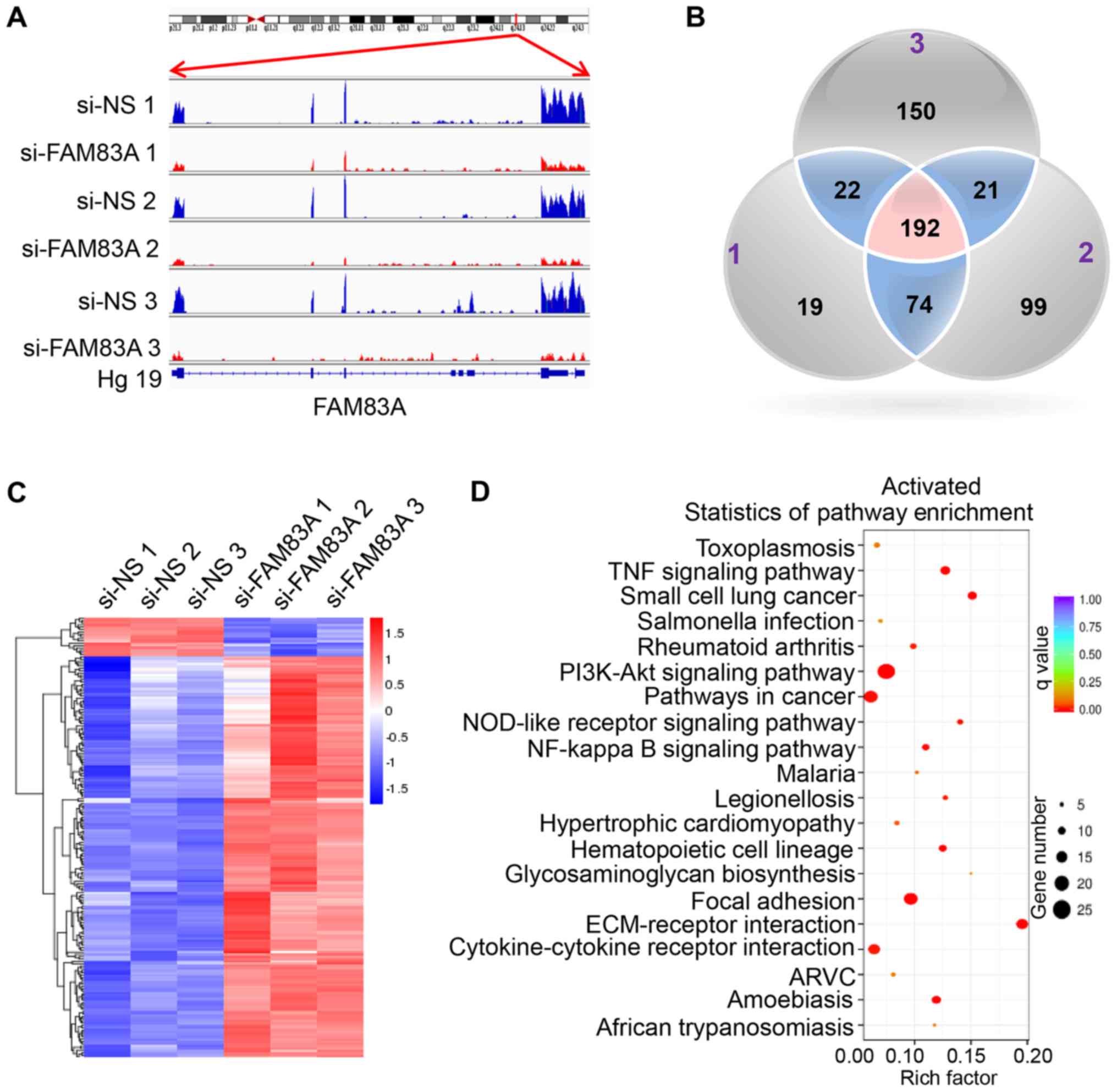
To identify the underlying mechanisms behind the
observed effects of FAM83A, the present study investigated the
signaling pathways regulated by FAM83A. RNA-seq was conducted in
HeLa cells transiently transfected with si-FAM83A and the negative
control groups (Table SII). The
expression of FAM83A was significantly inhibited in HeLa cells
transfected with FAM83A siRNA (Fig.
4A).
In total, 192 differentially expressed genes were
identified in si-FAM83A-transfected cells compared with the control
groups, with 175 upregulated and 17 downregulated genes (Fig. 4B and C; Table SIII). KEGG pathway analysis of the
192 differentially expressed genes identified a significant
activation of key oncogenesis-associated pathways following FAM83A
knockdown, including the 'PI3K/AKT signaling pathway', 'TNF
signaling pathway', 'focal adhesion' and 'ECM-receptor interaction'
(Fig. 4D), while the
'proteoglycans in cancer' and 'microRNAs in cancer' pathways were
suppressed (Fig. S2). To confirm
the KEGG pathway analysis results, the activated PI3K/AKT pathways
was validated in vitro. The results demonstrated that
knockdown of FAM83A significantly increased the expression of
p-PI3K and p-AKT proteins in HeLa and CaSki cells (Fig. S3).
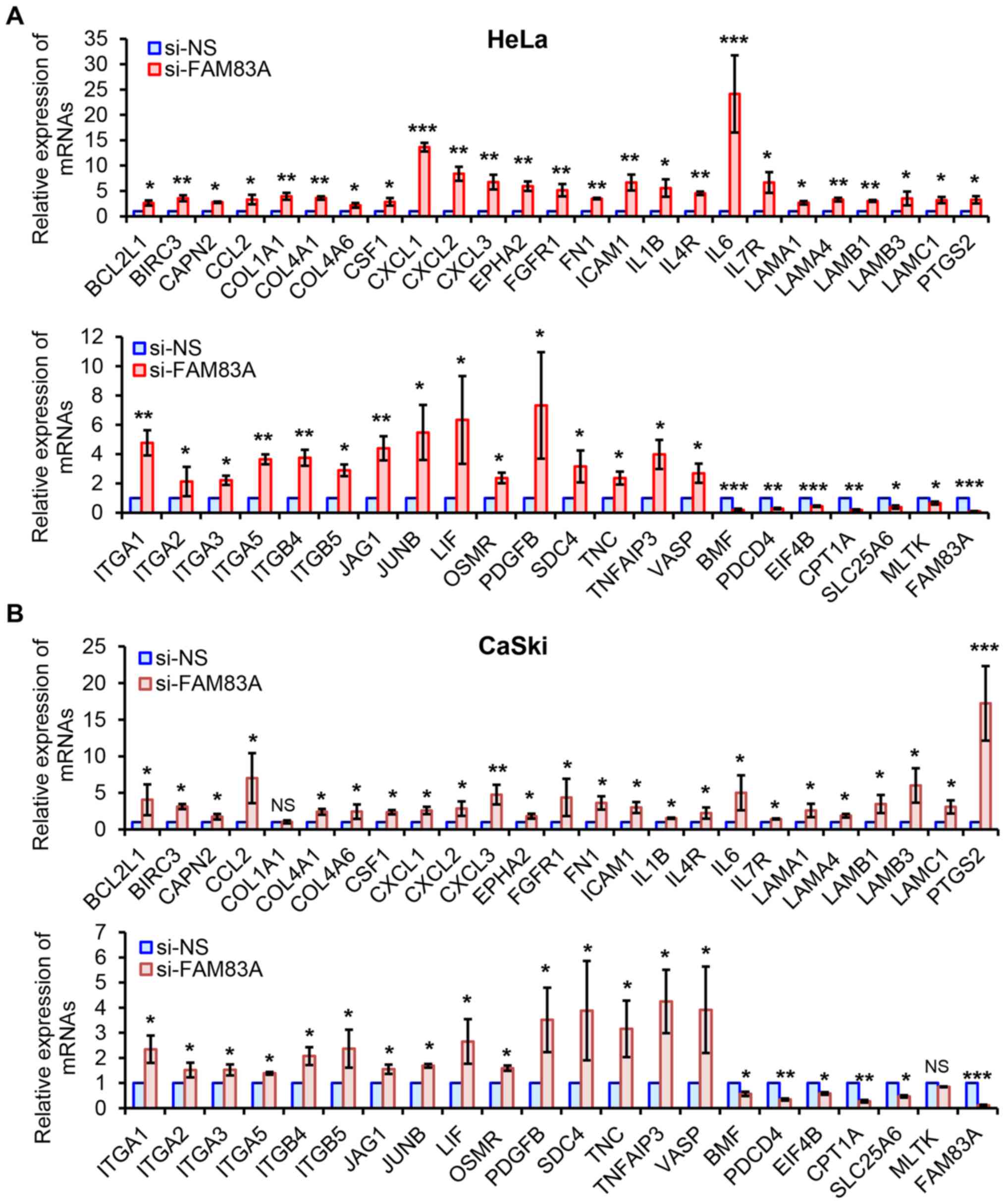
A total of 46 candidate genes, including 40 genes
from the top four activated pathways and six from the main
suppressed pathways in both HeLa and CaSki cells, were analyzed by
RT-qPCR. In line with the RNA-seq data, all 46 genes were similarly
regulated following FAM83A inhibition in HeLa cells (Fig. 5A). Although each expression pattern
of the 46 genes was different compared with those in HeLa cells,
similar expression patterns of 44 candidates were observed in CaSki
cells following FAM83A knockdown, with the exception of collagen
type 1 α1 and mitogen-activated protein kinase kinase kinase 20
(Fig. 5B). Therefore, FAM83A
inhibition may activate multiple downstream genes mainly involved
in oncogenesis-related pathways.
FAM83A knockdown activates integrins in
vitro and in vivo
The main cell adhesion receptors, known as
integrins, are essential for migration and invasion and have been
implicated in almost every step of cancer progression, from primary
tumor to metastasis (22,23). Dysregulated integrin-mediated
adhesion to the extracellular matrix (ECM) and signaling is a
precursor in the majority of invasive tumors (24-26).
To validate these candidate integrins, additional in vitro
and in vivo experiments were performed. The effects of
FAM83A knockdown on the protein expression levels of integrins α1,
α2, α3, α5, β4 and β5 in HeLa and CaSki cells were determined. As
presented in Fig. 6A and B, FAM83A
knockdown significantly increased the integrin α1, α3, α5, β4 and
β5 protein expression levels in HeLa and CaSki cells compared with
that in the corresponding si-NS groups, whereas integrin α2 protein
expression was not significantly affected.
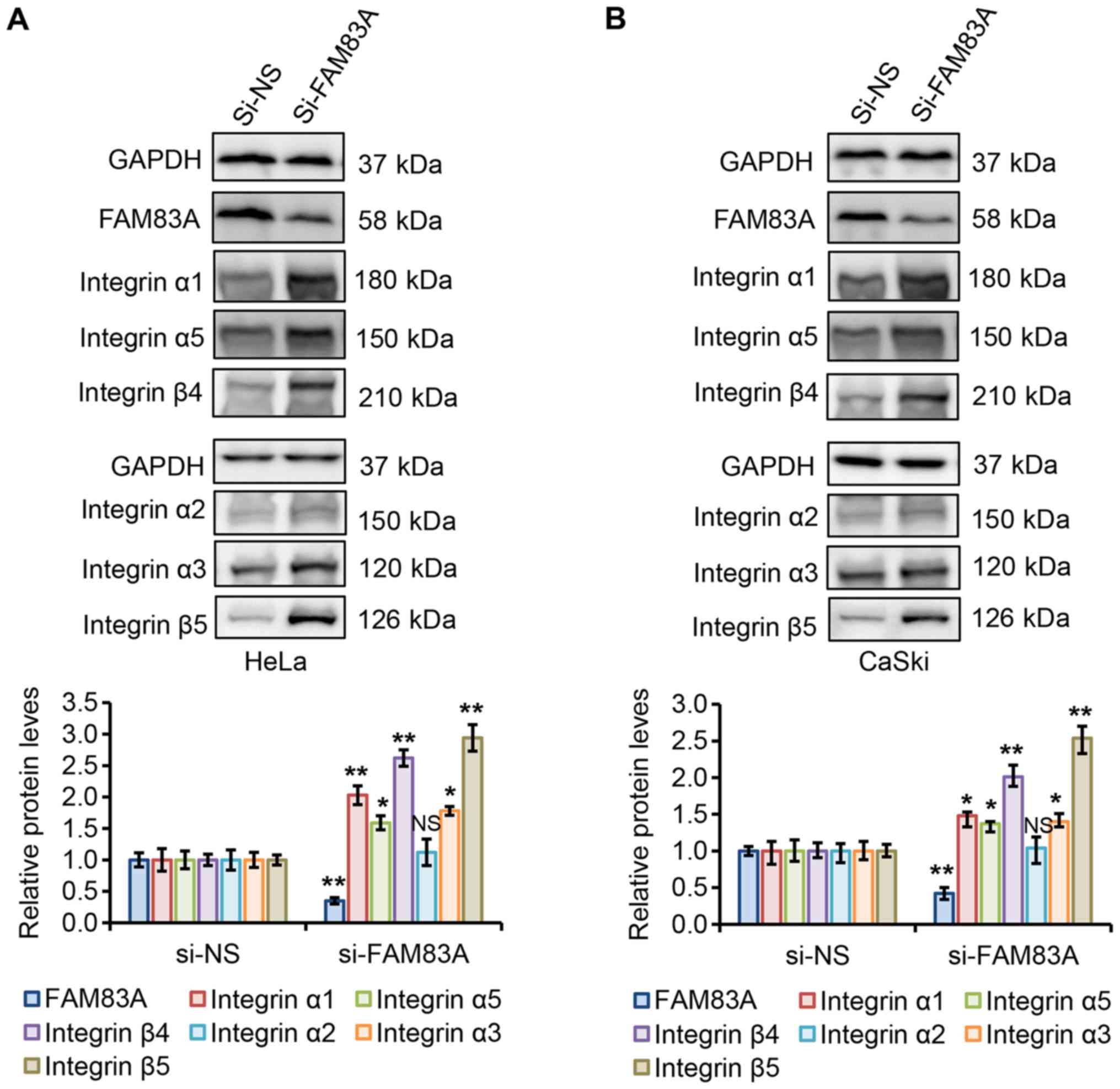 | Figure 6FAM83A knockdown activates integrins
α1, α3, α5, β4 and β5 in cervical cancer cells. (A and B) Western
blot analysis of FAM83A and integrins α1, α2, α3, α5, β4 and β5 in
(A) HeLa and (B) CaSki cells with or without FAM83A knockdown.
GAPDH protein level was used as the loading control. NS, not
significant; *P<0.05, **P<0.01 vs.
si-NS. FAM83A, family with sequence similarity 83 member A; si,
small interfering RNA; si-NS, scrambled control. |
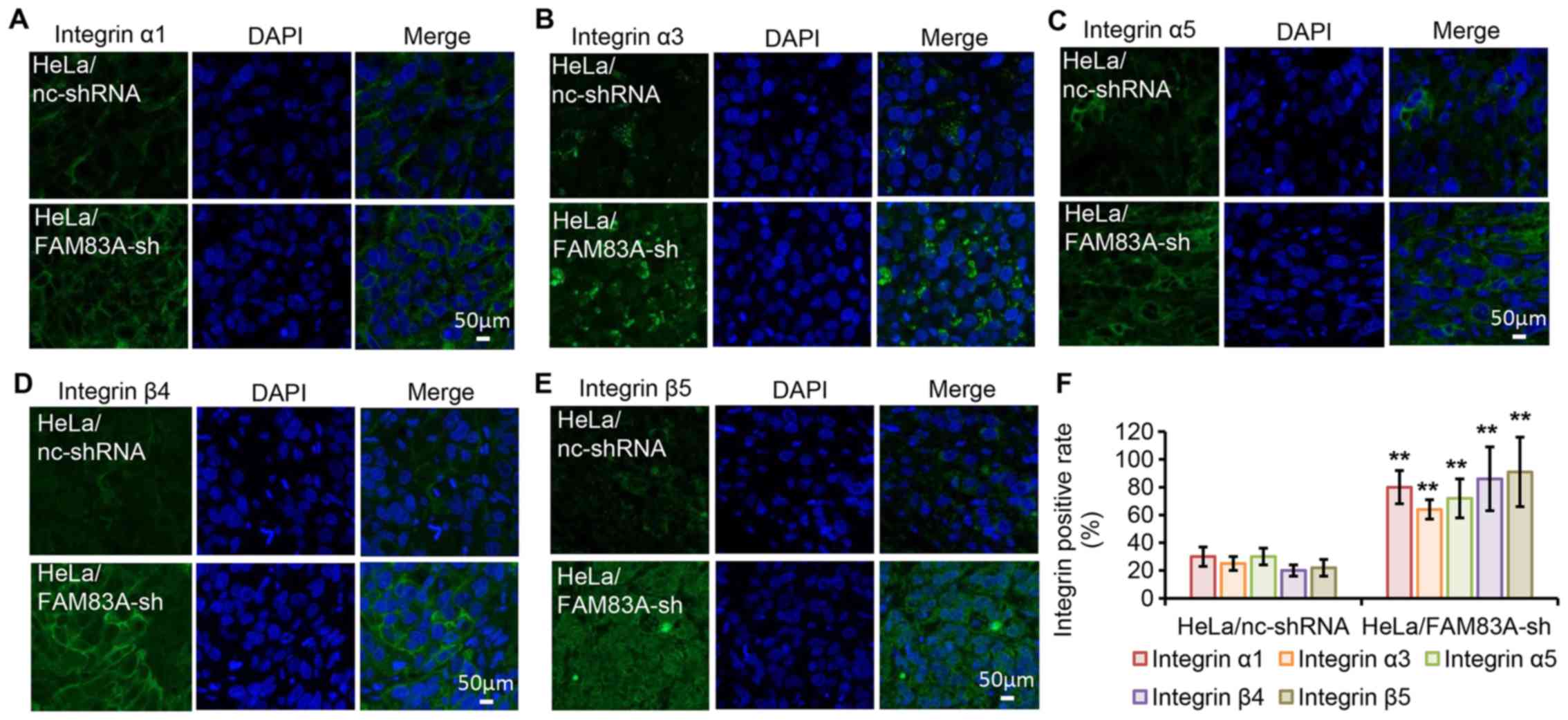
To evaluate the effect in vivo, the protein
expression levels of integrins α1, α3, α5, β4 and β5 were
investigated in tumor xenografts harvested from nude mice by
immunofluorescence staining (Fig.
7A-F). Consistent with the in vitro findings, tumors
expressing FAM83A shRNA exhibited increased protein expression
levels of the five tested integrins. These results further suggest
that integrins α1, α3, α5, β4 and β5 may be involved in the
tumor-promoting effects of FAM83A knockdown.
Clinical association among FAM83A and
integrins α1, α3, α5, β4 and β5 in human cervical cancer
tissues
The clinical relevance of the associations among
FAM83A and integrins α1, α3, α5, β4 and β5 were determined in
clinical samples from 20 patients with cervical cancer (10 with
high and 10 with low FAM83A expression) by IHC. As presented in
Fig. 8A and B, the protein
expression levels of integrins α1, α3, α5, β4 and β5 were
negatively associated with FAM83A protein expression level. For
example, the protein expression of integrin α1 was higher in the
cervical cancer tissue samples with low FAM83A expression compared
with that in samples with high FAM83A expression, whereas the
expression of integrin α1 was lower in cancer tissues with high
FAM83A expression. In addition, the protein expression levels of
integrin α3, α5, β4 and β5 were higher in cervical cancer samples
with low FAM83A expression compared with those in tissues with high
FAM83A expression. Collectively, the results suggested an
association between FAM83A and certain integrin subtypes in human
cervical cancer samples.
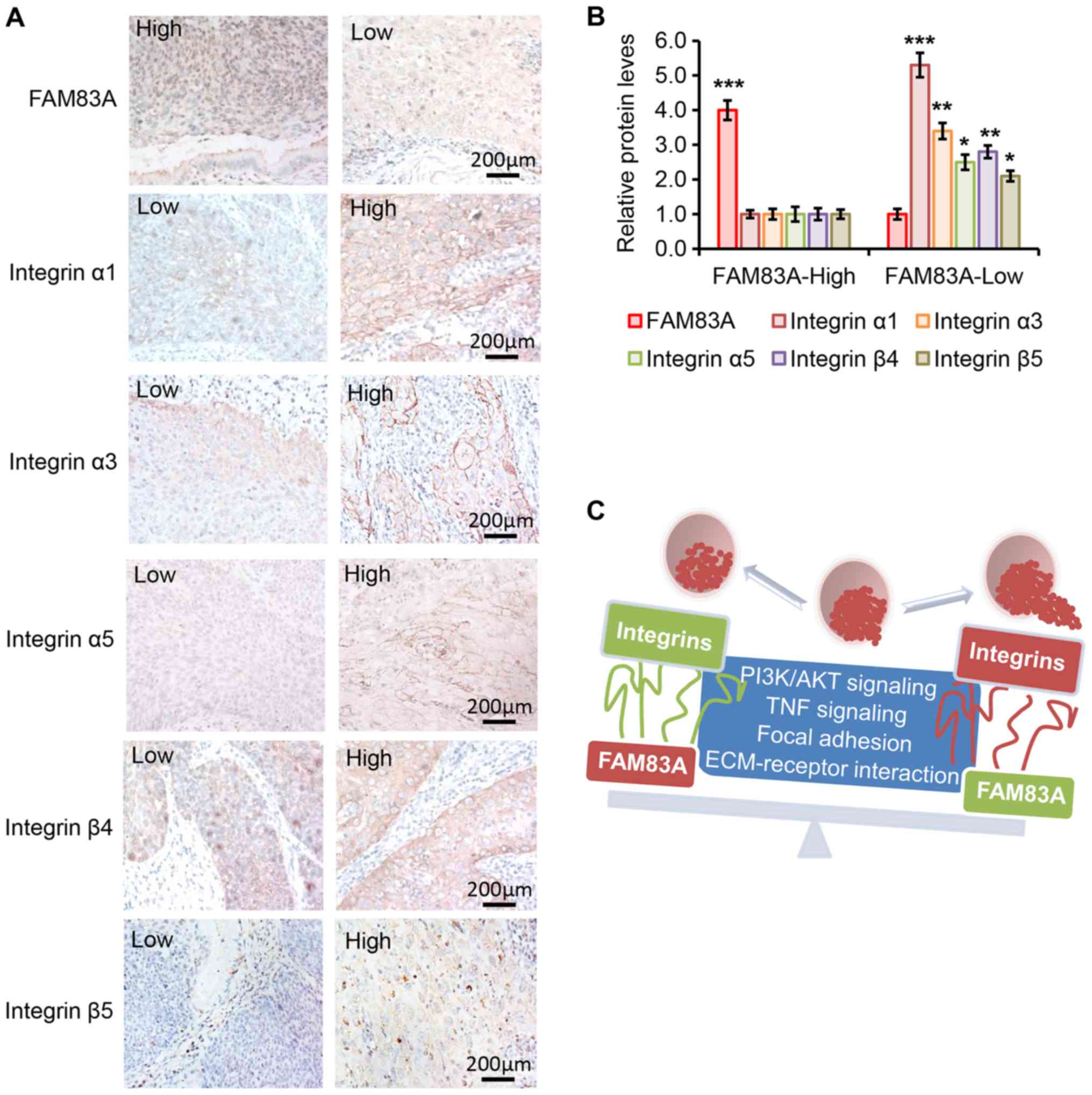 | Figure 8Protein levels of FAM83A and
integrins α1, α3, α5, β4 and β5 in human cervical cancer specimens.
(A) Representative images of the staining of FAM83A and integrins
α1, α3, α5, β4 and β5 in tissues from patients with cervical cancer
in the high and low FAM83A expression groups. 'High' and 'low' in
each image indicate each protein level separately. (B) Relative
protein expressions of FAM83A and integrins in cervical cancer
tissues from patients in the low and high FAM83A expression groups
determined by immunohistochemistry analysis (n=10 patients per
group). (C) Schematic diagram demonstrating that upregulation of
FAM83A inhibits the PI3K/AKT and TNF signaling, focal adhesion and
ECM-receptor interaction oncogenic pathways, followed by inhibition
of certain integrins to attenuate cervical cancer progression.
FAM83A, family with sequence similarity 83 member A; TNF, tumor
necrosis factor; ECM, extracellular matrix. |
Discussion
FAM83A belongs to a recently discovered oncogenic
FAM83 family and has been demonstrated to be significantly
upregulated in a variety of types of cancer, such as breast and
lung cancer (6,10,12,14,27,28).
Thus, FAM83A is presumed to be a novel oncogene. However, the
results of the present study suggested that FAM83A upregulation may
serve a tumor-suppressing role in cervical cancer in vitro
and in vivo, at least in part by suppressing integrins. It
was speculated that the contrasting results identified in different
types of cancer may indicate the context dependency of FAM83A
functions.
The present clinical observations in human cervical
samples have suggested complex roles for FAM83A. FAM83A mRNA
expression was profiled across all human tumor and paired healthy
tissues from TCGA and GTEx projects using GEPIA (29). As presented in Fig. S4, the mRNA expression of FMA83A
was upregulated in bladder urothelial carcinoma, cervical squamous
cell carcinoma, endocervical adenocarcinoma, head and neck squamous
cell carcinoma, lung adenocarcinoma, lung squamous cell carcinoma,
ovarian serous cystadenocarcinoma and pancreatic adenocarcinoma,
but suppressed in esophageal carcinoma, acute myeloid leukemia and
skin cutaneous melanoma. The mRNA expression of FAM83A was not
altered in other types of tumors, indicating the tissue specificity
of FAM83A expression. The present results demonstrated that FAM83A
expression was upregulated in human cervical cancer compared with
the normal cervix. Chen et al (11) have reported that elevated FAM83A
expression was attributed to the genomic amplification of
chromosome 8q24.13 and c-MYC gene amplification. FAM83A is
located near the HPV integration hot spots (30,31),
which may affect the expression of FAM83A. In the present study,
FAM83A upregulation was identified to be negatively associated with
advanced FIGO stage, deep stromal invasion, poor differentiation,
lymph node metastasis and poor prognosis. Although FAM83A was
upregulated in cervical cancer tissues, it is likely that FAM83A
serves tumor-suppressive role. The results of the present study
suggested that FAM83A may be used as a potential biomarker for
cervical cancer progression.
As FAM83A overexpression could drive HMEC
transformation and confer resistance to EGFR-TKIs, and knockdown of
FAM83A suppressed growth and tumorigenicity in breast cancer
(6,14,27),
it was initially hypothesized that FAM83A may also serve similar
functions in cervical cancer. However, in the present study,
knockdown of FAM83A promoted cell proliferation, migration,
invasion and tumorigenicity in vitro and in vivo. By
examining how FAM83A knockdown promoted tumor growth and invasion,
the present study identified 192 altered genes that associated
FAM83A with four key signaling pathways using RNA-seq. It was
demonstrated that FAM83A knockdown resulted in the activation of
the PI3K/AKT and TNF signaling pathways, focal adhesion and
ECM-receptor interaction. Opposite roles were reported by Lee et
al (6), who demonstrated that
FAM83A activated the PI3K/AKT pathways in breast cancer. Although
the results of the present study were different from the reported
data, both studies observed that FAM83A exerted its roles via the
same pathway. The differences in the results may have occurred due
to the different function of FAM83A in the different types of
cancer. To assess the present findings, 46 genes from the top
pathways in HeLa and CaSki cells were further validated, and it was
demonstrated that FAM83A knockdown upregulated the expression of
these pathway-related genes compared with that in the
si-NS-transfected cells. It is possible that FAM83A, despite being
generally considered to be an oncogene, may exert tumor-suppressive
activities in cervical cancer. Similar results have been observed
with other genes; for instance, polo-like kinase 1 (Plk1) has been
considered to be a classical oncogene in a number of tumor types
(32). However, de Cárcer et
al (33) revealed that Plk1
overexpression exerted a tumor suppressor role, and the increased
levels of Plk1 expression was associated with improved prognosis in
patients with breast cancer. In addition, similar observations have
been made for viral E7-induced expression of tumor-suppressive
p16Ink4a (34). High-level of
p16Ink4a expression is the clinical biomarker for high-risk
HPV-associated lesions and cancers (35). Thus, one reasonable assumption may
be that the HPV viral oncoproteins may also contribute to the
altered roles of FMA83A in cervical cancer.
The results of the present study also suggested that
FAM83A knockdown upregulated a set of integrin subunits. Integrins
are members of a diverse family of glycoproteins that form
heterodimeric receptors for ECM molecules (36). In total, 24 transmembrane
heterodimers are generated from a combination of its 18 α and eight
β subunits (36). Integrins are
essential for cell migration, invasion and tumor metastasis, not
only because they directly alter adhesion to the ECM, but also as
they respond to intracellular cues that control pro-survival
signaling pathways (37). Aberrant
integrin expression is frequently detected in various types of
cancer (24-26). For example, most β1 integrins are
necessary for mammary tumorigenesis (38,39).
In addition, integrin α5β1 upregulation contributes to poor tumor
prognosis and progression in lung cancer (40). Integrin αvβ3 is significantly
upregulated at the invasive front of melanoma cells and angiogenic
blood vessels (41). The
laminin-binding integrin α6β4 regulates the
mesenchymal-to-epithelial transition oncogenic signaling, activates
PI3K and correlates with the progression to invasive carcinoma,
such as breast carcinoma, thyroid cancer and gastric carcinoma
(42-45). The present study demonstrated that
FAM83A knockdown upregulated integrins α1, α3, α5, β4 and β5 in
cervical cancer cells and animal models. Thus, the results of the
present study provided evidence that cervical cancer samples with
low FAM83A expression exhibited high protein expression of
integrins α1, α3, α5, β4 and β5. In addition, it was indicated that
FAM83A acted as a tumor suppressor and exerted its suppressive role
at least partially via the inhibition of integrins α1, α3, α5, β4
and β5. In addition, laminins and collagens have been reported to
be closely associated with integrins to promote tumor progression
(40,46). The results of the present study
demonstrated FAM83A knockdown also significantly increased the
expression of several members of the laminin and collagen family,
indicating a potential association among integrins, collagens and
laminins.
In conclusion, the present study identified novel
potential roles of FAM83A in cervical cancer. Understanding the
exact role of FAM83A in cervical cancer and the molecular
mechanisms by which FAM83A knockdown activates integrins may offer
new insight into the biological basis of cervical cancer and lay
the groundwork for future studies.
Supplementary Data
Funding
This study was supported by the National Natural
Scientific Foundation of China (grant no. 81702552 to JX), the
National Key Research and Development Program of China (grant no.
2016YFC1302900 to WL) and the Fundamental Research Funds for the
Central Universities (grant no. 2019QNA7035 to JX).
Availability of data and materials
The datasets used and analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
JX and WL conceived the project, designed the study,
developed the methodology and analyzed the data. JX performed
experiments. JX wrote the manuscript. WL revised the manuscript and
supervised the study. Both authors read and approved the final
manuscript.
Ethics approval and consent to
participate
Ethics approval was obtained from the Ethics
Committee of Women's Hospital, Zhejiang University School of
Medicine (approval no. IRB-20190070). Written informed consent was
obtained from each patient. The animal study was approved by the
Animal Care and Use Committee of Zhejiang University (approval no.
ZJU20170031).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgments
Not applicable.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Walboomers JM, Jacobs MV, Manos MM, Bosch
FX, Kummer JA, Shah KV, Snijders PJ, Peto J, Meijer CJ and Muñoz N:
Human papillomavirus is a necessary cause of invasive cervical
cancer worldwide. J Pathol. 189:12–19. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hutchinson DJ and Klein KC: Human
papillomavirus disease and vaccines. Am J Health Syst Pharm.
65:2105–2112. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Markowitz LE, Dunne EF, Saraiya M, Lawson
HW, Chesson H and Unger ER: Centers for Disease Control and
Prevention (CDC); Advisory Committee on Immunization Practices
(ACIP): Quadrivalent human papillomavirus vaccine: Recommendations
of the advisory committee on immunization practices (ACIP). MMWR
Recomm Rep. 56:1–24. 2007.PubMed/NCBI
|
|
5
|
Xu J, Liu H, Yang Y, Wang X, Liu P, Li Y,
Meyers C, Banerjee NS, Wang HK, Cam M, et al: Genome-wide profiling
of cervical RNA-binding proteins identifies human papillomavirus
regulation of RNASEH2A expression by viral E7 and E2F1. MBio.
10:2019. View Article : Google Scholar
|
|
6
|
Lee SY, Meier R, Furuta S, Lenburg ME,
Kenny PA, Xu R and Bissell MJ: FAM83A confers EGFR-TKI resistance
in breast cancer cells and in mice. J Clin Invest. 122:3211–3220.
2012. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Fulcher LJ, Bozatzi P, Tachie-Menson T, Wu
KZL, Cummins TD, Bufton JC, Pinkas DM, Dunbar K, Shrestha S, Wood
NT, et al: The DUF1669 domain of FAM83 family proteins anchor
casein kinase 1 isoforms. Sci Signal. 11:eaao23412018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Li Y, Dong X, Yin Y, Su Y, Xu Q, Zhang Y,
Pang X, Zhang Y and Chen W: BJ-TSA-9, a novel human tumor-specific
gene, has potential as a biomarker of lung cancer. Neoplasia.
7:1073–1080. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bartel CA and Jackson MW: HER2-positive
breast cancer cells expressing elevated FAM83A are sensitive to
FAM83A loss. PLoS One. 12:pp. e01767782017, View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Grant S: FAM83A and FAM83B: Candidate
oncogenes and TKI resistance mediators. J Clin Invest.
122:3048–3051. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen S, Huang J, Liu Z, Liang Q, Zhang N
and Jin Y: FAM83A is amplified and promotes cancer stem cell-like
traits and chemore-sistance in pancreatic cancer. Oncogenesis.
6:pp. e3002017, View Article : Google Scholar
|
|
12
|
Snijders AM, Lee SY, Hang B, Hao W,
Bissell MJ and Mao JH: FAM83 family oncogenes are broadly involved
in human cancers: An integrative multi-omics approach. Mol Oncol.
11:167–179. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li Y, Xiao X, Ji X, Liu B and Amos CI:
RNA-seq analysis of lung adenocarcinomas reveals different gene
expression profiles between smoking and nonsmoking patients. Tumour
Biol. 36:8993–9003. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Cipriano R, Miskimen KL, Bryson BL, Foy
CR, Bartel CA and Jackson MW: Conserved oncogenic behavior of the
FAM83 family regulates MAPK signaling in human cancer. Mol Cancer
Res. 12:1156–1165. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Pawitan Y, Bjöhle J, Amler L, Borg AL,
Egyhazi S, Hall P, Han X, Holmberg L, Huang F, Klaar S, et al: Gene
expression profiling spares early breast cancer patients from
adjuvant therapy: Derived and validated in two population-based
cohorts. Breast Cancer Res. 7:R953–R964. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
17
|
Zhou Q, Wang X, Yu Z, Wu X, Chen X, Li J,
Zhu Z, Liu B and Su L: Transducin (β)-like 1 X-linked receptor 1
promotes gastric cancer progression via the ERK1/2 pathway.
Oncogene. 36:1873–1886. 2017. View Article : Google Scholar
|
|
18
|
Kim D, Pertea G, Trapnell C, Pimentel H,
Kelley R and Salzberg SL: TopHat2: Accurate alignment of
transcriptomes in the presence of insertions, deletions and gene
fusions. Genome Biol. 14:R362013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Anders S and Huber W: Differential
expression analysis for sequence count data. Genome Biol.
11:R1062010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Robinson JT, Thorvaldsdóttir H, Winckler
W, Guttman M, Lander ES, Getz G and Mesirov JP: Integrative
genomics viewer. Nat Biotechnol. 29:24–26. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Fang Y, Yu H, Liang X, Xu J and Cai X:
Chk1-induced CCNB1 overexpression promotes cell proliferation and
tumor growth in human colorectal cancer. Cancer Biol Ther.
15:1268–1279. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Winograd-Katz SE, Fässler R, Geiger B and
Legate KR: The integrin adhesome: From genes and proteins to human
disease. Nat Rev Mol Cell Biol. 15:273–288. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hamidi H and Ivaska J: Every step of the
way: Integrins in cancer progression and metastasis. Nat Rev
Cancer. 18:533–548. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Seguin L, Desgrosellier JS, Weis SM and
Cheresh DA: Integrins and cancer: Regulators of cancer stemness,
metastasis, and drug resistance. Trends Cell Biol. 25:234–240.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Raab-Westphal S, Marshall JF and Goodman
SL: Integrins as therapeutic targets: Successes and cancers.
Cancers (Basel). 9. pp. 1102017, View Article : Google Scholar
|
|
26
|
Hamidi H, Pietilä M and Ivaska J: The
complexity of integrins in cancer and new scopes for therapeutic
targeting. Br J Cancer. 115:1017–1023. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Cipriano R, Graham J, Miskimen KL, Bryson
BL, Bruntz RC, Scott SA, Brown HA, Stark GR and Jackson MW: FAM83B
mediates EGFR- and RAS-driven oncogenic transformation. J Clin
Invest. 122:3197–3210. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Bartel CA, Parameswaran N, Cipriano R and
Jackson MW: FAM83 proteins: Fostering new interactions to drive
oncogenic signaling and therapeutic resistance. Oncotarget.
7:52597–52612. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res. 45:W98–W102.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ojesina AI, Lichtenstein L, Freeman SS,
Pedamallu CS, Imaz-Rosshandler I, Pugh TJ, Cherniack AD, Ambrogio
L, Cibulskis K, Bertelsen B, et al: Landscape of genomic
alterations in cervical carcinomas. Nature. 506:371–375. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Hu Z, Zhu D, Wang W, Li W, Jia W, Zeng X,
Ding W, Yu L, Wang X, Wang L, et al: Genome-wide profiling of HPV
integration in cervical cancer identifies clustered genomic hot
spots and a potential microhomology-mediated integration mechanism.
Nat Genet. 47:158–163. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ito Y, Miyoshi E, Sasaki N, Kakudo K,
Yoshida H, Tomoda C, Uruno T, Takamura Y, Miya A, Kobayashi K, et
al: Polo-like kinase 1 overexpression is an early event in the
progression of papillary carcinoma. Br J Cancer. 90:414–418. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
de Cárcer G, Venkateswaran SV, Salgueiro
L, El Bakkali A, Somogyi K, Rowald K, Montañés P, Sanclemente M,
Escobar B, de Martino A, et al: Plk1 overexpression induces
chromosomal instability and suppresses tumor development. Nat
Commun. 9:30122018. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Munger K, Gwin TK and McLaughlin-Drubin
ME: p16 in HPV-associated cancers. Oncotarget. 4:1864–1865. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Malhone C and Longatto-Filho A: Cervical,
ovarian and endometrial tumor markers: Potential clinical value.
Semin Ultrasound CT MR. 40:350–357. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
van der Flier A and Sonnenberg A: Function
and interactions of integrins. Cell Tissue Res. 305:285–298. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Hood JD and Cheresh DA: Role of integrins
in cell invasion and migration. Nat Rev Cancer. 2:91–100. 2002.
View Article : Google Scholar
|
|
38
|
Cagnet S, Faraldo MM, Kreft M, Sonnenberg
A, Raymond K and Glukhova MA: Signaling events mediated by α3β1
integrin are essential for mammary tumorigenesis. Oncogene.
33:4286–4295. 2014. View Article : Google Scholar
|
|
39
|
White DE, Kurpios NA, Zuo D, Hassell JA,
Blaess S, Mueller U and Muller WJ: Targeted disruption of
beta1-integrin in a transgenic mouse model of human breast cancer
reveals an essential role in mammary tumor induction. Cancer Cell.
6:159–170. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Roman J, Ritzenthaler JD, Roser-Page S,
Sun X and Han S: alpha5beta1-integrin expression is essential for
tumor progression in experimental lung cancer. Am J Respir Cell Mol
Biol. 43:684–691. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Brooks PC, Clark RA and Cheresh DA:
Requirement of vascular integrin alpha v beta 3 for angiogenesis.
Science. 264:569–571. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Trusolino L, Bertotti A and Comoglio PM: A
signaling adapter function for alpha6beta4 integrin in the control
of HGF-dependent invasive growth. Cell. 107:643–654. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Serini G, Trusolino L, Saggiorato E,
Cremona O, De Rossi M, Angeli A, Orlandi F and Marchisio PC:
Changes in integrin and E-cadherin expression in neoplastic versus
normal thyroid tissue. J Natl Cancer Inst. 88:442–449. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Khwaja A, Rodriguez-Viciana P, Wennström
S, Warne PH and Downward J: Matrix adhesion and Ras transformation
both activate a phosphoinositide 3-OH kinase and protein kinase
B/Akt cellular survival pathway. EMBO J. 16:2783–2793. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Shaw LM, Rabinovitz I, Wang HH, Toker A
and Mercurio AM: Activation of phosphoinositide 3-OH kinase by the
alpha6beta4 integrin promotes carcinoma invasion. Cell. 91:949–960.
1997. View Article : Google Scholar
|
|
46
|
Rathinam R and Alahari SK: Important role
of integrins in the cancer biology. Cancer Metastasis Rev.
29:223–237. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
FIGO Committee on Gynecologic Oncology:
FIGO staging for carcinoma of the vulva, cervix, and corpus uteri.
Int J Gynaecol Obstet. 125:97–98. 2014. View Article : Google Scholar : PubMed/NCBI
|