Introduction
Colorectal cancer (CRC) is responsible for ~930,000
deaths annually worldwide, making it the second most common cause
of cancer-related death globally (1). Metastasis and local progression are
the major limiting factors in the efficacy of CRC treatment. The
stemness of cancer cells and mitochondrial stability play pivotal
roles in the invasion and metastasis of CRC (2-4).
Cancer cell stemness is closely associated with
tumor progression and recurrence. CRC cells with enhanced stemness
are more likely to breach the basement membrane and enter the
circulatory system, thereby facilitating metastasis to distant
organs (5). These stem-like tumor
cells also exhibit greater resistance to conventional therapies,
presenting a notable challenge in CRC treatment (6). Mitochondrial stability is central to
the metabolic adaptability and the anti-apoptotic capacity of CRC
cells (7,8). Through mitochondrial metabolic
reprogramming, tumor cells can adapt to various microenvironments,
thereby increasing their survival rate. Thus, the regulation of
stemness and mitochondrial stability influences the invasiveness
and metastatic potential of CRC cells. Intervening in the
mechanisms governing stemness and mitochondrial stability may offer
novel strategies to enhance the therapeutic outcomes of CRC.
Ubiquitin-specific protease 4 (USP4) plays crucial
regulatory roles in multiple cellular processes implicated in
cancer development and progression. Accumulating evidence
demonstrates that USP4 regulates key players (namely WNT3A, RAD50,
TGFB1 and PRL3) in cancer-relevant pathways, including cell
proliferation, apoptosis and DNA damage responses (9-12).
Given these multifaceted roles in cancer biology, understanding how
USP4 activity is regulated and how its function can be modulated
through interaction with other regulatory molecules, such as long
non-coding RNAs (lncRNAs), represents an important avenue for
cancer research and therapeutic development.
lncRNAs are a class of RNAs >200 nucleotides in
length and lacking protein-coding sequences; they play crucial
roles in chromatin dynamics, gene expression regulation and cell
growth and differentiation (13-15). Whole-genome association studies of
tumor samples have confirmed that lncRNAs are key factors in the
initiation and progression of various cancer types. For instance,
lncRNA-H19 promotes the development of gastric cancer, CRC and
gliomas by encoding and producing microRNA-675 (16,17). Moreover, linc02273 drives breast
cancer metastasis by upregulating the transcription of AGR2
(18). In addition, the oncogene
c-Myc promotes tumor formation by silencing p53 through the lncRNA
MILIP (19). Beyond their
well-established roles in transcriptional and post-transcriptional
regulation, lncRNAs can serve as molecular scaffolds that
facilitate or modulate protein-protein interactions, thereby
affecting the enzymatic activity, substrate specificity or
stability of their protein partners (20,21). Several studies have demonstrated
that lncRNAs can directly bind to deubiquitinating enzymes and
influence their deubiquitinase activity or subcellular
localization, adding an additional layer of complexity to
post-translational regulatory networks in cancer cells (22,23).
AL445238.2, located on the long arm of chromosome 13
(q21.31), consists of two exons and is 1,581 bp in length. At
present and to the best of our knowledge, there are no published
reports on AL445238.2. Targeting the cell survival and metastasis
mechanisms, such as by inhibiting the interaction between
AL445238.2 and its downstream signaling pathways, may provide novel
strategies to improve CRC treatment outcomes.
Materials and methods
Cell lines, cell culture and
transfection
The CRC cell lines SW480 and DLD1 were purchased
from the American Type Culture Collection. Cells were cultured in
Dulbecco's Modified Eagle's Medium (Gibco; Thermo Fisher
Scientific, Inc.) containing 10% fetal bovine serum (FBS;
HyClone™; Cytiva) and incubated at 37°C with 5%
CO2. Overexpression or knockdown lentiviral vectors
encoding AL445238.2, USP4 or the corresponding empty vector were
constructed using the pLVX-IRES-Puro plasmid backbone by Shanghai
GeneChem Co., Ltd. Lentiviral particles were also produced and
concentrated to a titer of 1×108 TU/ml by Shanghai
GeneChem Co., Ltd. For lentiviral transduction, cells were seeded
at 30-40% confluency and transduced with lentiviral particles at a
multiplicity of infection of 10-20 in the presence of 5
μg/ml polybrene (MilliporeSigma) at 37°C for 24 h. Stable
transfected cells were selected and maintained with 2 μg/ml
puromycin (MilliporeSigma) starting 72 h post-transfection. All of
the cell lines were confirmed to be free from mycoplasma
contamination and were authenticated by an expert before being used
for experimentation. All shRNA sequences are listed in Table SI.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from SW480 and DLD1 cells
using TRIzol reagent (MilliporeSigma) and RNA purity and integrity
were assessed by electrophoresis. RT was performed using random
primers at 42°C for 1 h (RevertAid First Strand cDNA Synthesis Kit;
Thermo Fisher Scientific, Inc.). The target gene was then amplified
in a 20 μl reaction system using SYBR-Green qPCR Mix (Thermo
Fisher Scientific, Inc.). The PCR conditions were as follows: 50°C
for 2 min, 95°C for 2 min, followed by 40 cycles of 95°C for 15 sec
and 60°C for 32 sec. Relative gene expression levels were
normalized to the reference gene β-actin and quantified using the
2−ΔΔCq method (24).
The primer sequences for qPCR are provided in Table SII.
Western blot analysis
CRC cells were lysed in Cell Lysis Buffer for
Western and IP without Inhibitors (NCM Biotech; Suzhou Xinsaimei
Biotechnology Co., Ltd.) containing a protease and phosphatase
inhibitor mixture (Beyotime Biotechnology). The protein
concentration was determined using the BCA assay. Denatured
proteins (20 μg) were separated by sodium dodecyl
sulfate-polyacrylamide gel electrophoresis on a 4-20% gel and
transferred to polyvinylidene fluoride membranes (MilliporeSigma).
Membranes were blocked with 5% non-fat milk (BD Biosciences) in
Tris-buffered saline containing 0.1% Tween-20 at room temperature
for 1 h. Membranes were incubated overnight at 4°C with primary
antibodies (dilution 1:1,000) and then washed and incubated at room
temperature for 1 h with HRP-conjugated Goat Anti-Rabbit IgG (H+L)
secondary antibody (dilution 1:5,000). Enhanced chemiluminescence
(Thermo Fisher Scientific, Inc.) was used to detect protein bands.
GAPDH or β-actin were used as the internal control. Protein band
intensities were quantified by densitometry using ImageJ software
(version 1.53k; National Institutes of Health). Antibody
information is provided in Table
SIII.
Co-immunoprecipitation (Co-IP) assay
Cells (1×107) were harvested and lysed in
IP lysis buffer (Pierce IP Lysis Buffer; Thermo Fisher Scientific,
Inc.; cat. no. 87787) supplemented with protease inhibitor cocktail
(1:100 dilution; Thermo Fisher Scientific, Inc.; cat. no. 78430)
and phosphatase inhibitor cocktail (1:100 dilution; Thermo Fisher
Scientific, Inc.; cat. no. 78428) on ice for 30 min with
intermittent vortexing. Cell lysates were centrifuged at 14,000 × g
for 15 min at 4°C, and the supernatant was collected. Protein
concentration was determined using the BCA Protein Assay Kit
(Thermo Fisher Scientific, Inc.; cat. no. 23225). For each IP
reaction, 500 μg of total protein lysate was pre-cleared
with 20 μl of Protein A/G Magnetic Beads (Thermo Fisher
Scientific, Inc.; cat. no. 88802) at 4°C for 1 h with rotation to
reduce non-specific binding. After magnetic separation, the
pre-cleared lysate was incubated with 2 μg of anti-USP4
antibody (cat. no. ab108931; Abcam) or normal rabbit IgG control
antibody (cat. no. 2729; Cell Signaling Technology, Inc.) overnight
at 4°C with gentle rotation. Subsequently, 30 μl of Protein
A/G Magnetic Beads were added to each sample and incubated for 2 h
at 4°C with rotation. The beads were then collected using a
magnetic separator and washed five times with IP wash buffer (25 mM
Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% NP-40 and 5% glycerol).
After the final wash, beads were resuspended in 30 μl of 2X
SDS loading buffer and boiled at 95°C for 5 min to elute the
immunocomplexes. The supernatant was collected by centrifugation at
1,000 × g for 1 min at room temperature and subjected to western
blot analysis.
Cell Counting Kit-8 (CCK-8) cell
proliferation assay
CRC cells (3×103 cells/well) were seeded
into a 96-well plate. After 24, 48 or 72 h of incubation, 10
μl CCK-8 reagent (Beijing Solarbio Science & Technology
Co., Ltd.) was added to each well and incubated at 37°C for 60 min.
Cell proliferation was measured at 450 nm using a Varioskan LUX
microplate reader.
Lactate dehydrogenase (LDH) release
assay
Cytotoxicity was assessed by measuring LDH release
into the culture medium using the CytoTox 96®
Non-Radioactive Cytotoxicity Assay (Promega Corporation; cat. no.
G1780) according to the manufacturer's instructions. Briefly, cells
were seeded in 96-well plates at a density of 5×103
cells per well and cultured for 24, 48 or 72 h. At each time point,
50 μl of culture supernatant was transferred to a new
96-well plate and mixed with 50 μl of CytoTox 96®
Reagent. After incubating at room temperature for 30 min in the
dark, the reaction was stopped by adding 50 μl of Stop
Solution. Absorbance was measured at 490 nm using a microplate
reader (BioTek Instruments, Inc.; Agilent Technologies, Inc.). LDH
release was calculated as a percentage of maximum LDH release
(determined by lysing cells with 1% Triton X-100). Background
absorbance from culture medium alone was subtracted from all
measurements. Each experiment was performed in triplicate and
repeated three times independently.
RNA pull-down assay
Cells (1×107) were collected and lysed in
RIPA buffer (Beyotime Biotechnology) with a protease inhibitor
mixture (NCM Biotech; Suzhou Xinsaimei Biotechnology Co., Ltd.).
For each 1 ml of cell lysate, 10% was used as the input control.
Biotinylated AL445238.2 sense probe and a scrambled negative
control probe with no homology to any known transcripts were
synthesized by GenePharma Co., Ltd. AL445238.2 sense and scrambled
probes were incubated with the cell lysate at 4°C for 4 h. Next,
streptavidin magnetic beads were added and the mixture was
incubated at 4°C for 1 h. After boiling the magnetic beads in 0.1%
sodium dodecyl sulfate solution for 3 min, protein interactions
with AL445238.2 were detected by western blot. The control cells
were treated with an equal volume of beads with scrambled probes to
assess non-specific protein binding.
Flow cytometry
Apoptosis was detected using an Annexin V-FITC/PI
dual-staining apoptosis detection kit (Absin Bioscience, Inc.)
according to the manufacturer's instructions. In brief, cells were
harvested, washed with pre-chilled phosphate buffered saline (PBS),
resuspended in 100 μl binding buffer and sequentially
stained with Annexin V-FITC (5 μl) for 10 min in the dark at
room temperature, followed by PI (5 μl) for 5 min in the
dark. After washing with PBS, the cells were resuspended and
analyzed by flow cytometry.
Stemness was assessed by staining with a
FITC-conjugated CD133 flow cytometric monoclonal antibody (cat. no.
372803; BioLegend, Inc.). After collection, the cells were
resuspended in staining buffer (BioLegend, Inc.) at a concentration
of 1.25×106 cells/100 μl, incubated with 5
μl CD133 antibody in the dark for 20 min at room
temperature, washed with PBS and resuspended in PBS for flow
cytometry analysis.
Mitochondrial activity was assessed using the
tetramethylrhodamine (TMRM) probe (cat. no. I34361 Invitrogen;
Thermo Fisher Scientific, Inc.). After collecting the cells, the
supernatant was discarded and the cells were incubated with DMEM
containing a 1:1,000 dilution of the TMRM probe at 37°C with 5%
CO2 for 30 min. After washing with PBS, the cells were
resuspended for flow cytometry analysis.
All flow cytometry data were collected using BD
FACSCelesta and analyzed using FlowJo software (version 10; FlowJo
LCC; BD Biosciences). Median fluorescence intensity was calculated
using the median fluorescence value for each population.
Transwell migration assay
Transwell migration assays were performed using
Corning, Inc. Transwell chambers. In brief, cells (1×106
cells/well) were seeded in the upper chambers of 24-well plates
with 100 μl serum-free medium. The lower chambers were
filled with 600 μl medium containing 30% FBS. After 24 h of
incubation at 37°C, cells that migrated to the lower chambers were
fixed with 4% paraformaldehyde at room temperature for 15 min,
washed with PBS and stained with 0.1% crystal violet at room
temperature for 20 min. The number of migrated cells was counted
under a light microscope (Zeiss Axio Observer; Carl Zeiss AG) and
quantified using Image-Pro Plus software (version 6.0; Media
Cybernetics, Inc.). Images were acquired using ZEN imaging software
(version 2.3 blue edition; Carl Zeiss AG).
Cell immunofluorescence
Cells were seeded on glass slides in 24-well plates
and cultured for 24 h. After washing with PBS, the cells were fixed
with 4% paraformaldehyde at room temperature for 10 min, followed
by permeabilization with 0.5% Triton X-100 (PBS) for 15 min at room
temperature. After blocking with PBS containing 5% normal goat
serum (MilliporeSigma) at room temperature for 1 h, the cells were
incubated with the primary antibody (Anti-USP4; 1:500; cat. no.
ab181105; Abcam) overnight at 4°C, followed by incubation with
Alexa Fluor 488-conjugated goat anti-rabbit IgG (H+L) secondary
antibody (1:500; cat. no. A11008; Thermo Fisher Scientific, Inc.)
for 1 h at room temperature in the dark. For FISH staining of
AL445238.2, after permeabilization, cells were incubated with
pre-hybridization buffer (Abnova Corporation; cat. no. U0028) at
37°C for 30 min, then hybridized with Alexa Fluor 555-labeled FISH
probe provided by GenePharma Co., Ltd. at 37°C overnight in the
dark. Following hybridization, cells were washed with SSC buffer
(Thermo Fisher Scientific, Inc.; cat. no. J60839.K2; 2X SSC for 5
min, 1X SSC for 5 min and 0.5X SSC for 5 min) at 37°C. The slides
were mounted using an anti-fade mounting medium containing
4',6-diamidino-2-phenylindole and analyzed under a fluorescence
microscope.
Spheroid formation assay
Cells were dissociated into a single-cell suspension
and seeded at a density of 5×103 cells/well in a 6-well
ultra-low attachment plate (Corning, Inc.). Cells were cultured in
serum-free DMEM/F12 (Gibco; Thermo Fisher Scientific, Inc.)
supplemented with epidermal growth factor (20 ng/ml; PeproTech,
Inc.; Thermo Fisher Scientific, Inc.) and B27 (2%; Gibco; Thermo
Fisher Scientific, Inc.). After 1-2 weeks of culture, the tumor
spheroids were observed under a light microscope and the size and
number of spheroids were measured using Image-Pro Plus
software.
In vivo tumor xenograft model
A nude mouse xenograft model was established using
female BALB/c-nude immunodeficient mice aged 5-6 weeks (initial
body weight: 18-20 g; GemPharmatech, Co., Ltd.). Mice were randomly
assigned to 3 groups using a random number generator, with each
mouse given a unique identification number before group allocation
to ensure unbiased distribution. The groups were subcutaneously
injected with SW480 cells transfected with empty vector, USP4
overexpression vector or AL445238.2 overexpression vector (n=4 per
group). The mice were housed under a 12-h light/dark cycle with
free access to food and water. Each mouse was subcutaneously
injected with 5×106 SW480 cells suspended in 100
μl PBS into the right abdominal flank. Animal health and
behavior were monitored daily for signs of distress, weight loss or
abnormal behavior. Tumor size was measured twice weekly using
digital calipers and the tumor volume was calculated using the
formula: Volume (mm3)=(length × width2)/2,
where length is the longest diameter and width is the perpendicular
diameter. After 12 weeks, the mice were euthanized using
CO2 following the AVMA Guidelines for the Euthanasia of
Animals (25). Mice were placed
in a clear euthanasia chamber and 100% CO2 from a
compressed gas cylinder was introduced using a gradual fill method
with a displacement rate of 30% of the chamber volume per min. The
CO2 flow was maintained for at least 1 min after
respiratory arrest and death was confirmed by physical examination
(absence of heartbeat and respiration for at least 10 mi with
graying of mucous membranes) before tumor harvest.
The sample size for the animal study was determined
following the recommendations of the Animal Ethics Committee of
Wenzhou Medical University (Wenzhou, China) to ensure the
establishment of well-behaved animal models with adequate
statistical power. During the experiment, the tumor diameter was
not allowed to exceed 1.5 cm, nor was the tumor volume allowed to
exceed 15% of the total body volume of the mouse. No animals
reached these humane endpoints during the study.
Statistical analysis
Data from three independent experiments were
analyzed using R (version 4.3.1; R Foundation for Statistical
Computing). Results are expressed as the mean ± standard deviation.
Comparisons between two groups were analyzed using unpaired
Student's t-test. Multiple group comparisons were performed using
one-way ANOVA followed by Tukey's post hoc test. P<0.05 was
considered to indicate a statistically significant difference.
Results
AL445238.2 promotes CRC proliferation and
inhibits apoptosis
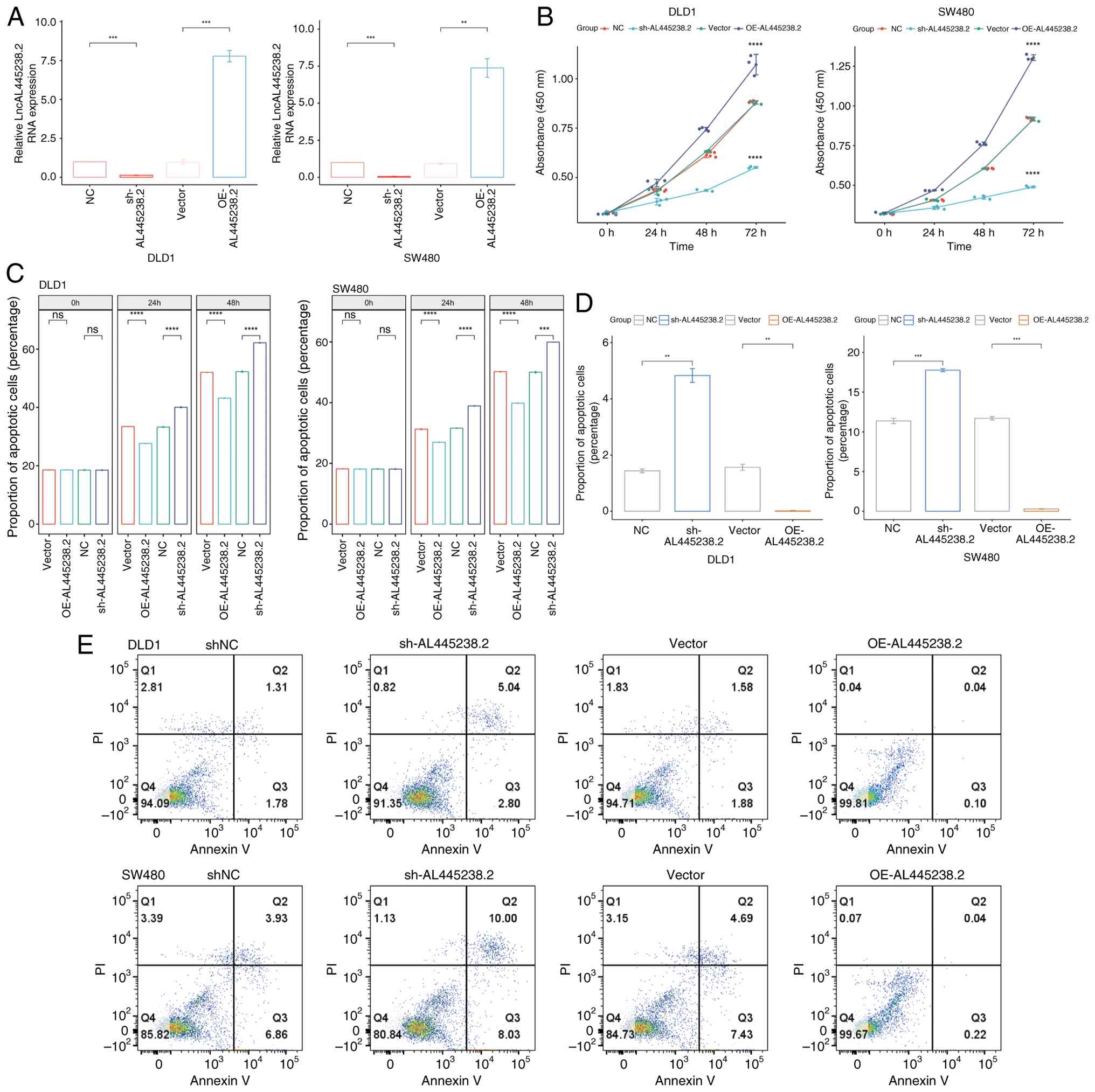
After confirming the overexpression or knockdown
efficiency of AL445238.2 using qPCR (Fig. 1A), the impact of AL445238.2 on CRC
cell proliferation was assessed using the CCK-8 assay and detection
of LDH in the supernatant. AL445238.2 overexpression significantly
enhanced the proliferation of DLD1 and SW480 CRC cells and reduced
LDH levels in the supernatant, suggesting decreased cytotoxicity
and cell membrane damage (Fig. 1B and
C). Conversely, AL445238.2 knockdown weakened the proliferation
of DLD1 and SW480 cells and increased LDH levels in the
supernatant, suggesting enhanced cytotoxicity. Apoptosis analysis
by flow cytometry revealed that overexpression of AL445238.2
reduced overall apoptosis in both DLD1 and SW480 cells, whereas
knockdown of AL445238.2 increased overall apoptosis (Fig. 1D and E). Taken together, these
results demonstrated that AL445238.2 promoted CRC cell
proliferation while protecting cells from both cytotoxic damage and
apoptotic cell death.
AL445238.2 enhances the stemness and
mitochondrial activity of CRC cells
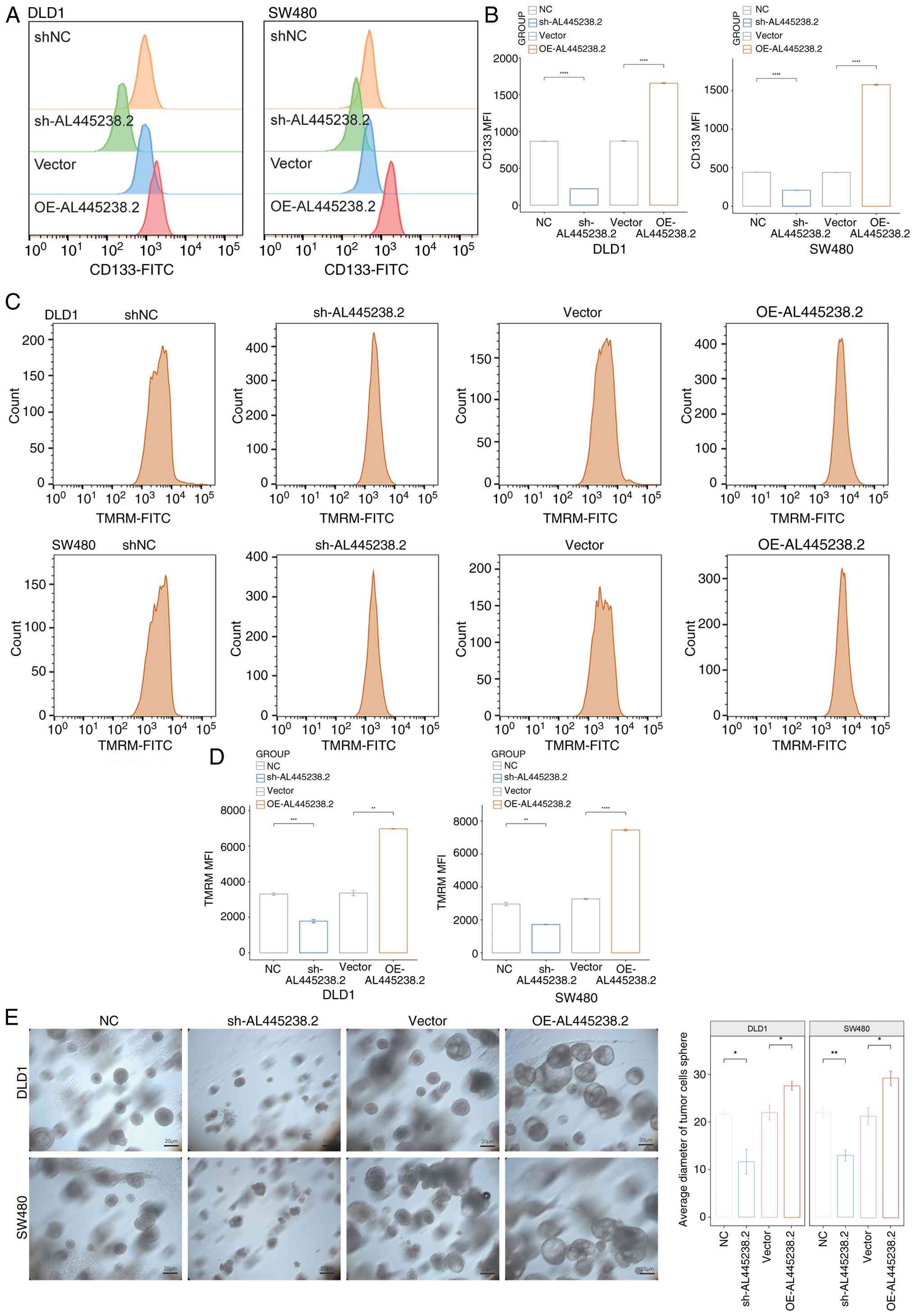
Flow cytometry analysis of CD133 (a stem cell
marker) expression on DLD1 and SW480 cells showed that
overexpression of AL445238.2 increased CD133 expression on the cell
surface compared with the empty vector group, while knockdown of
AL445238.2 decreased CD133 expression (Fig. 2A and B). These results suggest
that AL445238.2 promotes stemness in CRC cells. Subsequently, TMRM
(a probe for assessing mitochondrial membrane potential) flow
cytometry analysis revealed that AL445238.2 overexpression enhanced
TMRM fluorescence in DLD1 and SW480 cells, indicating increased
mitochondrial activity, whereas AL445238.2 knockdown decreased
mitochondrial activity (Fig. 2C and
D).
To further explore the role of AL445238.2 in
regulating CRC cell stemness, a spheroid formation assay was
conducted. AL445238.2 overexpression in DLD1 and SW480 cells
increased the diameter of tumor spheroids, whereas AL445238.2
knockdown reduced the spheroid size, suggesting a role in stemness
enhancement (Fig. 2E).
AL445238.2-USP4 complex stabilizes Bcl2
to promote stemness and the survival of CRC cells
RNA pull-down assays were performed to explore the
molecular interactors of AL445238.2, which revealed that AL445238.2
bound to USP4 (Fig. 3A).
Immunofluorescence confirmed the co-localization of AL445238.2 with
USP4, indicating an interaction (Fig.
3B). The overexpression or knockdown efficiency of USP4 was
confirmed using qPCR (Fig. 3C).
Overexpression of AL445238.2 in SW480 cells upregulated Bcl2 and
downregulated BAX and cleaved CASP3 (C-CASP3) expression, whereas
knockdown of AL445238.2 reduced Bcl2 expression and increased BAX
and C-CASP3 expression, suggesting that AL445238.2 contributed to
cell survival (Fig. 3D).
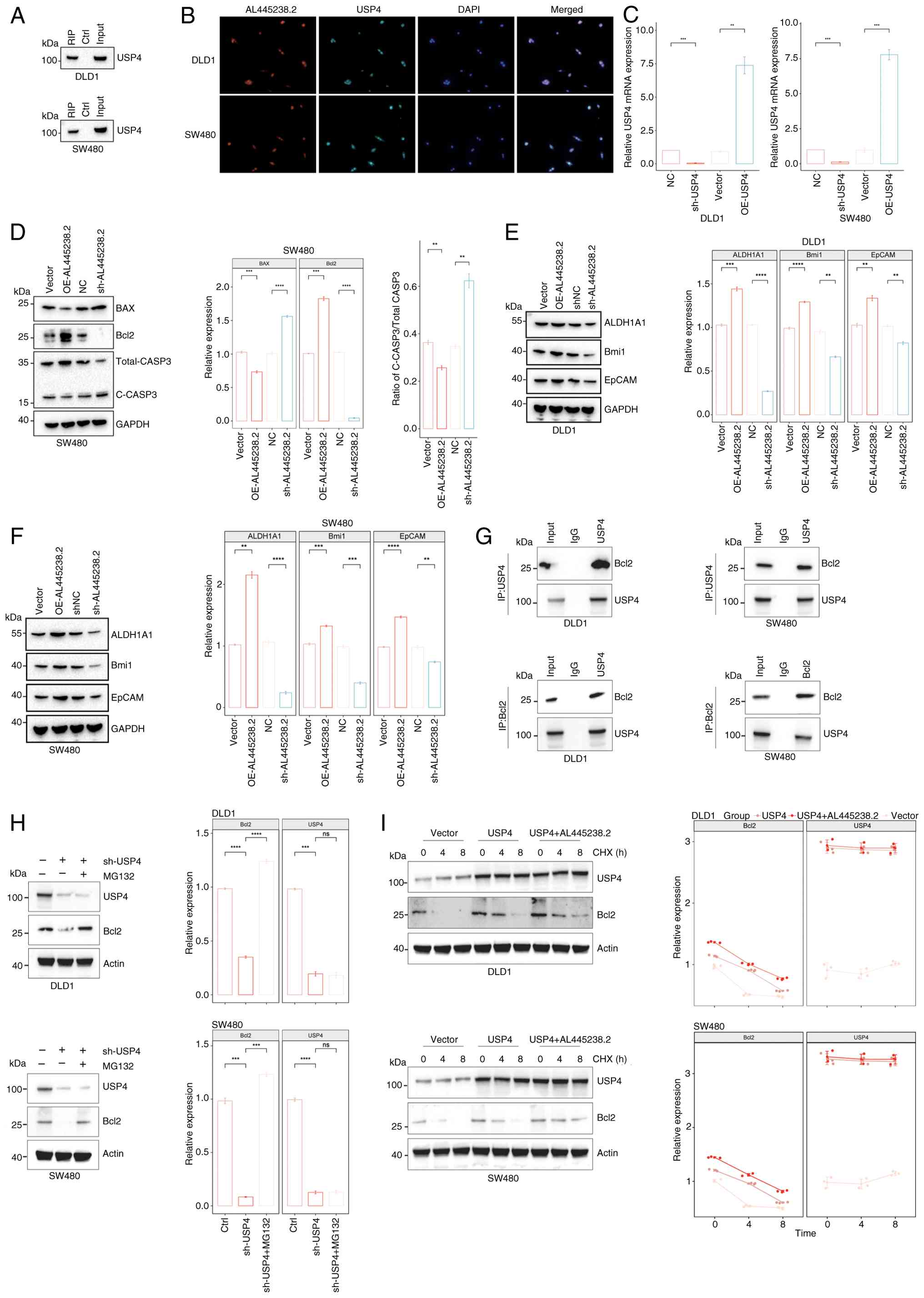 | Figure 3AL445238.2 binding to USP4 stabilizes
Bcl2 to regulate colon cancer cell apoptosis and stemness. (A)
Pulldown assay with AL445238.2 and western blot detection of USP4.
(B) Gene colocalization was detected by immunofluorescence
(magnification, ×200). (C) Validation of gene manipulation
efficiency in SW480 and DLD1 cell lines with stable overexpression
and knockdown of USP4. (D) Western blotting was used to evaluate
the expression of BAX, Bcl2 and C-CASP3 in stable SW480 cells; BAX
and Bcl2 were normalized against GAPDH, C-CASP3 was normalized
against total CASP3. Western blotting was used to evaluate the
expression of ALDH1A1, Bmi1 and EpCAM in stable (E) DLD1 and (F)
SW480 cells. (G) Co-immunoprecipitation experiments were used to
verify the interaction between USP4 and Bcl2. (H) Western blotting
was used to evaluate the effects of USP4 knockdown and MG132 on
Bcl2 protein expression. (I) Protein half-life experiments were
used to evaluate the effects of USP4 and AL445238.2 on the
stability of Bcl2 protein. **P<0.01,
***P<0.001, ****P<0.0001. USP4,
ubiquitin specific peptidase 4; RIP, RNA immunoprecipitation; Ctrl,
control; ALDH1A1, aldehyde dehydrogenase 1 family member A1; Bmi1,
B lymphoma Mo-MLV insertion region 1; EpCAM, epithelial cell
adhesion molecule; CHX, cycloheximide; OE, overexpression; sh,
short hairpin; NC, negative control; ns, not significant; C-CASP3,
cleaved caspase-3. |
The expression of the stemness markers aldehyde
dehydrogenase 1 family member A1 (ALDH1A1), B lymphoma Mo-MLV
insertion region 1 (Bmi1) and epithelial cell adhesion molecule
(EpCAM) in stable SW480 and DLD1 cell lines was further examined.
Overexpression of AL445238.2 increased the levels of ALDH1A1, Bmi1
and EpCAM, whereas knockdown of AL445238.2 reduced their
expression, suggesting that AL445238.2 enhanced stemness in CRC
cells (Fig. 3E and F). Co-IP
experiments were performed to further how the AL445238.2-USP4
complex modulates the apoptotic pathway. It was found that USP4 can
interact with Bcl2 (Fig. 3G).
Knockdown of USP4 expression reduced Bcl2 expression in colon
cancer cells, which could be reversed by MG132 (Fig. 3H). Further protein half-life
experiments confirmed that USP4 overexpression increased the
stability of Bcl2 protein and prolonged its half-life (Fig. 3I). Moreover, the co-expression of
USP4 and AL445238.2 could further enhance the effect of USP4,
extending the intracellular lifespan of Bcl2. Therefore, we
conclude that AL445238.2-USP4 regulates the apoptotic pathway and
exerts its subsequent functions by stabilizing Bcl2 protein.
USP4 promotes CRC Cell proliferation and
inhibits apoptosis
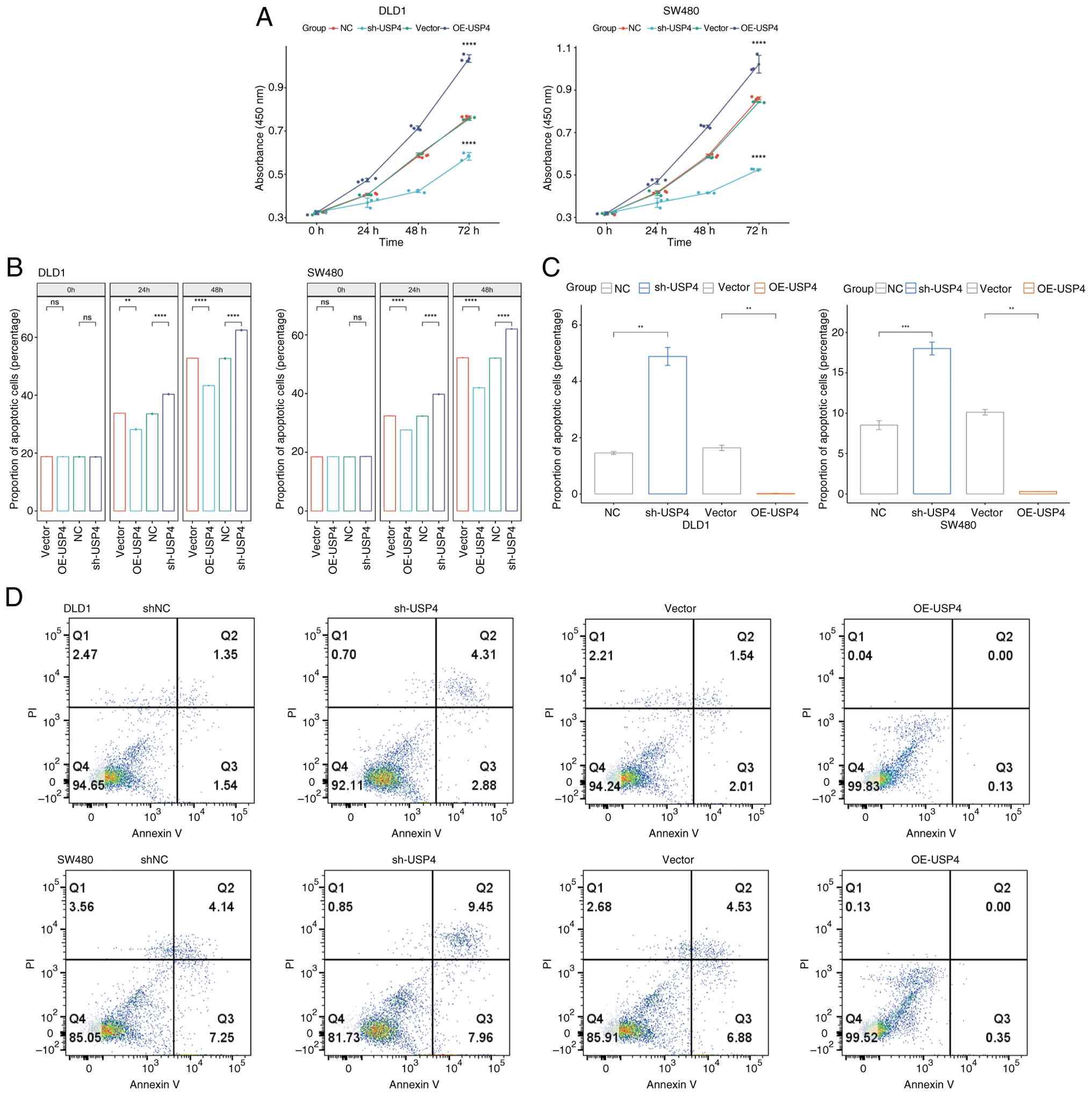
The impact on CRC cell proliferation was evaluated
using CCK-8 assays and LDH detection (Fig. 4A and B). USP4 overexpression
significantly enhanced the proliferation of DLD1 and SW480 cells
and reduced LDH levels in the supernatant, indicating reduced
apoptosis. Conversely, USP4 knockdown weakened cell proliferation
and increased LDH levels, suggesting an increase in apoptosis. Flow
cytometry analysis confirmed that USP4 overexpression reduced
apoptosis in DLD1 and SW480 cells, whereas USP4 knockdown increased
apoptosis (Fig. 4C and D).
Consistent with AL445238.2, USP4 overexpression in SW480 cells
upregulated Bcl2 and downregulated BAX and C-CASP3 expression,
whereas USP4 knockdown reduced Bcl2 expression and increased BAX
and C-CASP3 expression, suggesting that USP4 suppresses cell
apoptosis (Fig. 5A).
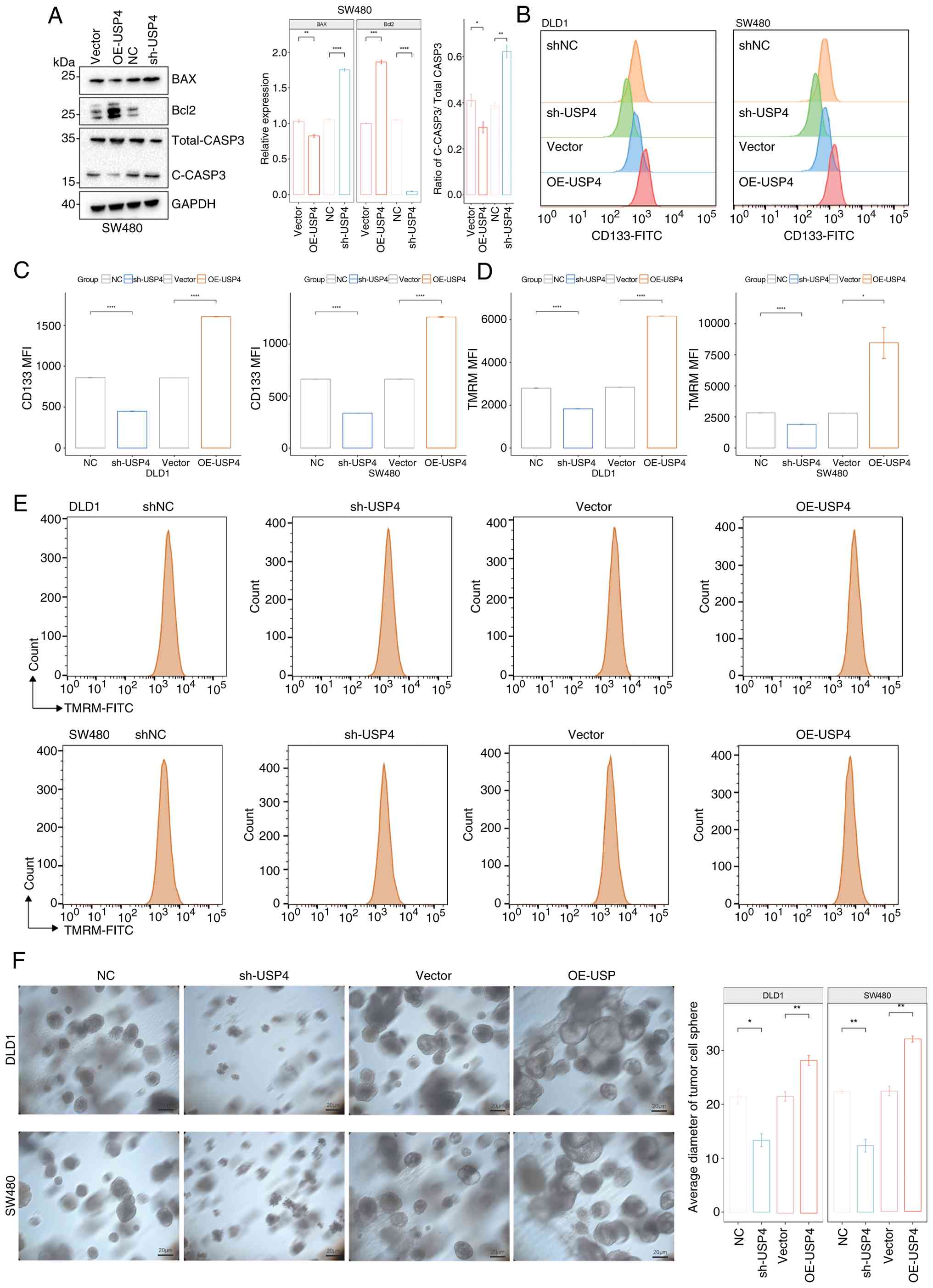 | Figure 5USP4 promotes stemness and
mitochondrial stability of colon cancer cells. (A) Western blotting
was used to evaluate the expression of BAX, Bcl2 and C-CASP3 in
stable SW480 cells; BAX and Bcl2 were normalized against GAPDH,
C-CASP3 was normalized against total CASP3. (B) The expression of
CD133 in the SW480 and DLD1 stable cell lines was detected with
flow cytometry. (C) Statistical analysis of the CD133 MFI. (D)
Statistical analysis of the TMRM MFI. (E) TMRM staining of SW480
and DLD1 stable cell lines. (F) Tumor spheroid formation assay of
SW480 and DLD1 stable cell lines. *P<0.05,
**P<0.01, ***P<0.001,
****P<0.0001. MFI, mean fluorescence intensity; TMRM,
tetramethylrhodamine; C-CASP3, cleaved caspase-3; NC, negative
control; USP4, ubiquitin specific peptidase 4; sh, short hairpin;
OE, overexpression. |
USP4 enhances stemness and mitochondrial
activity in CRC cells
Flow cytometry analysis revealed that USP4
overexpression increased CD133 expression in DLD1 and SW480 cells,
while USP4 knockdown decreased CD133 expression, suggesting that
USP4 may promote stemness (Fig. 5B
and C). TMRM flow cytometry showed that USP4 overexpression
increased mitochondrial activity, whereas USP4 knockdown decreased
mitochondrial activity (Fig. 5D and
E). Spheroid formation assays confirmed that USP4
overexpression in DLD1 and SW480 cells increased the tumor spheroid
diameter, whereas USP4 knockdown reduced the spheroid size
(Fig. 5F).
AL445238.2 relies on USP4 to promote CRC
cell migration
Transwell migration assays demonstrated that both
AL445238.2 and USP4 overexpression enhanced the migratory capacity
of CRC cells (Fig. 6A-D). By
contrast, AL445238.2 or USP4 knockdown impaired cell migration.
Furthermore, when USP4 was knocked down while AL445238.2 was
overexpressed, no significant increase in cell migration was
observed compared with the control group (Fig. 6E and F). These findings suggest
that AL445238.2 promotes CRC cell migration in a USP4-dependent
manner.
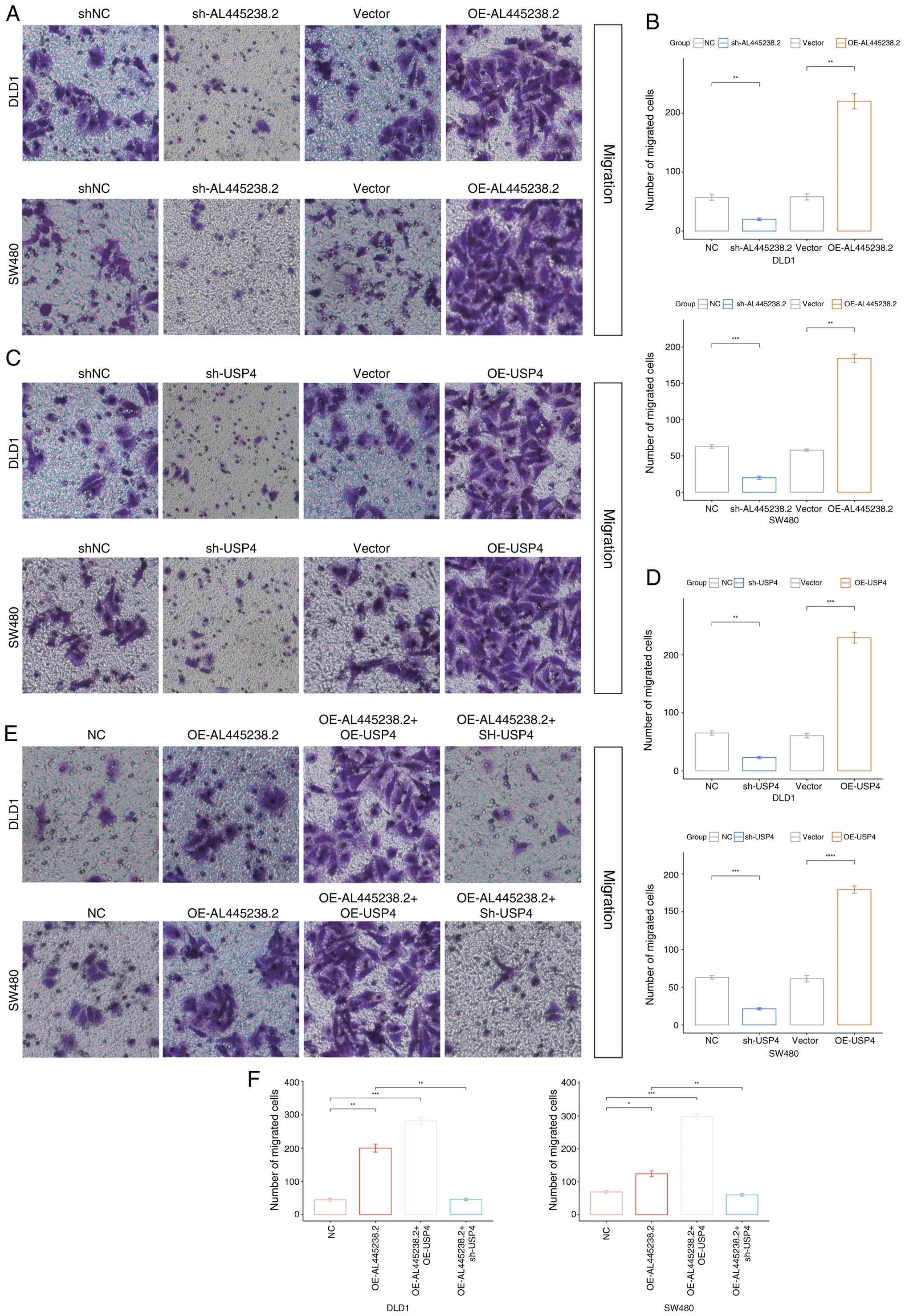 | Figure 6AL445238.2 controlled USP4 promotes
colon cancer cell migration. Migration assay of DLD1 and SW480
cells treated with shNC, sh-AL445238.2, Vector or OE-AL445238.2.
(A) Representative images show migrated cells stained with crystal
violet. (B) Quantitative analysis of migrated cell numbers in DLD1
(upper panel) and SW480 (lower panel) cells following AL445238.2
knockdown or overexpression. Migration assay of DLD1 and SW480
cells treated with shNC, sh-USP4, Vector or OE-USP4. (C)
Representative images show migrated cells stained with crystal
violet. (D) Quantitative analysis of migrated cell numbers in DLD1
(upper panel) and SW480 (lower panel) cells following USP4
knockdown or overexpression. Migration assay of DLD1 and SW480
cells with combined treatments: NC, OE-AL445238.2, OE-AL445238.2 +
OE-USP4 or OE-AL445238.2 + sh-USP4. (E) Representative images show
migrated cells. (F) Quantitative analysis of migrated cell numbers
in DLD1 (left panel) and SW480 (right panel) cells under combined
treatment conditions. *P<0.05,
**P<0.01, ***P<0.001,
****P<0.0001. sh, short hairpin; NC, negative
control; USP4, ubiquitin specific peptidase 4; OE,
overexpression. |
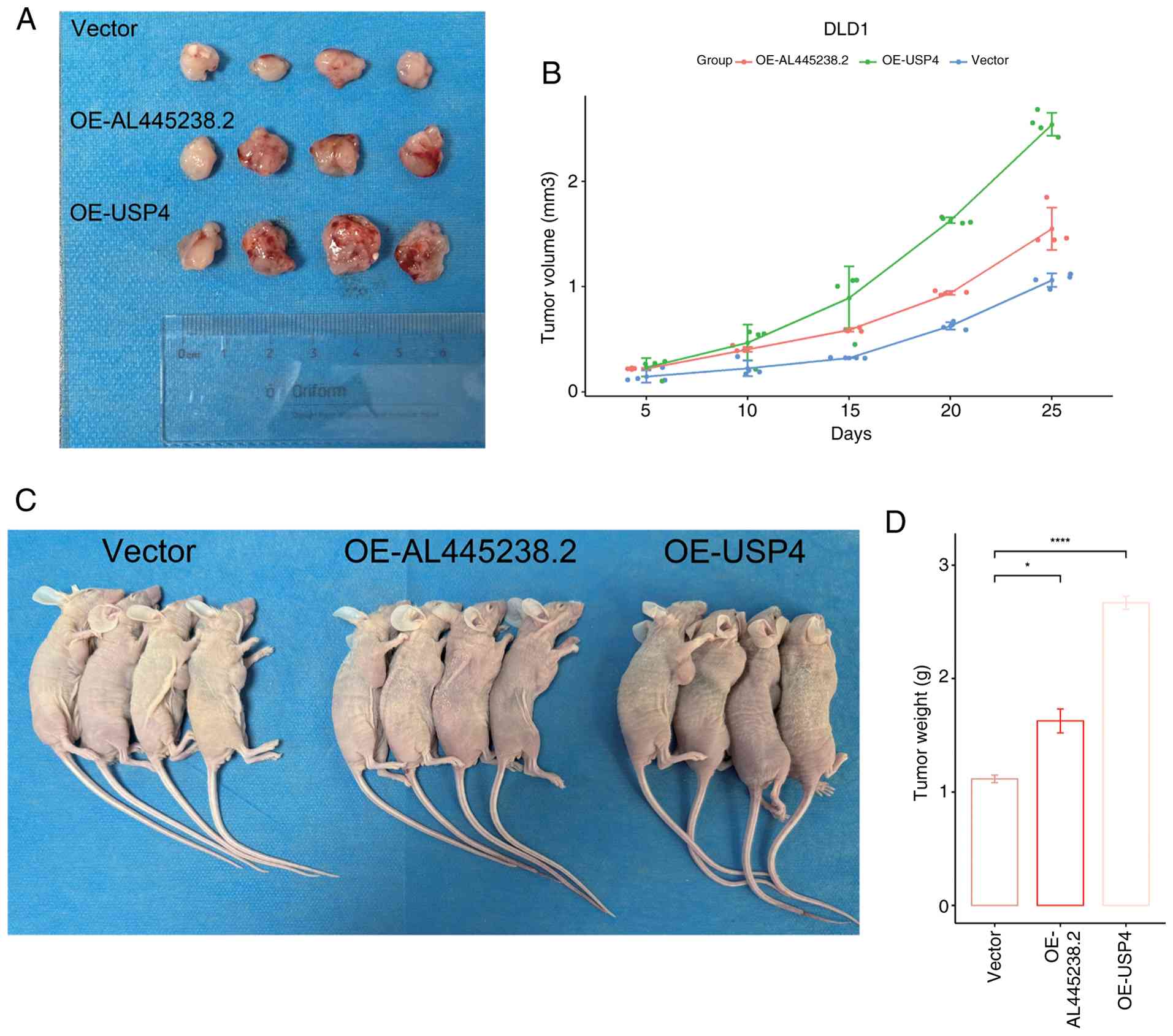
AL445238.2 and USP4 promote CRC cell
proliferation in vivo
In vivo tumorigenesis experiments using SW480
cells demonstrated that the overexpression of AL445238.2 or USP4
increased the growth rate, final tumor size and tumor weight of CRC
cells in mice (Fig. 7A-D). These
results suggest that both AL445238.2 and USP4 contribute to CRC
cell proliferation in vivo.
Discussion
CRC is the third most prevalent malignancy
worldwide, with ~1.8 million new cases diagnosed annually (26,27). Local recurrence or distant
metastasis remains the primary cause of treatment failure in CRC
(28,29). Consequently, there is an urgent
need to deepen our understanding of the mechanisms underlying colon
cancer progression and metastasis. The role of lncRNAs in tumor
biology has attracted increasing attention in recent years.
Mounting evidence suggests that lncRNAs not only participate in
transcriptional regulation and chromatin remodeling but also
modulate critical biological processes such as cell cycle
progression, apoptosis, stemness maintenance and migration, thereby
playing a notable role in tumor initiation and progression
(30-32). Numerous studies have reported that
the aberrant expression of various lncRNAs in solid tumors,
including CRC, is closely linked to clinicopathological features
and prognosis (33-37).
The present study demonstrated that lncRNA
AL445238.2 may be associated with colon cancer progression and
metastasis. The experimental results demonstrated that AL445238.2
promoted the proliferation, survival, stemness maintenance and
migration of CRC cells through multiple pathways, with its function
largely dependent on the deubiquitinase, USP4. This discovery
enriches our understanding of the molecular underpinnings of CRC
and offers a promising target for developing novel therapeutic
strategies.
In the present study, initial in vitro
experiments revealed that overexpression of AL445238.2
significantly enhanced the proliferative capacity of DLD1 and SW480
cells, simultaneously reducing extracellular LDH levels and overall
apoptosis rates, indicating a marked protective effect on cell
survival. Conversely, knocking down AL445238.2 expression led to
diminished cell proliferation, increased LDH levels and heightened
apoptosis, confirming its critical role in maintaining cell
viability. It was also found that AL445238.2 promoted both the
stem-like characteristics and mitochondrial activity of CRC cells.
Flow cytometry analysis of CD133 expression, TMRM staining and
sphere formation assays consistently indicated that elevated levels
of AL445238.2 enhanced cellular stemness and metabolic activity,
potentially underpinning the long-term survival and chemoresistance
of tumor cells. Notably, similar regulatory effects were observed
with changes in USP4 expression, suggesting a synergistic interplay
between AL445238.2 and USP4 in modulating cell stemness and
mitochondrial function.
In the present study, mechanistic investigations
using RNA pull-down assays and immunolocalization experiments
revealed a direct interaction between AL445238.2 and USP4, thereby
providing a molecular basis for their cooperative role in signal
transduction. High expression of either AL445238.2 or USP4 was
associated with increased levels of the anti-apoptotic protein Bcl2
and decreased levels of the pro-apoptotic protein BAX, offering a
plausible mechanism for their inhibition of apoptosis. Cell
migration assays demonstrated that the pro-migratory effect of
AL445238.2 was dependent on the presence of USP4, further
reinforcing the functional connection between these molecules.
Furthermore, in vivo tumorigenesis experiments using SW480
cells showed that the overexpression of either AL445238.2 or USP4
promoted tumor growth, confirming that the AL445238.2-USP4 axis may
play a critical role in CRC tumorigenesis and validating the
physiological relevance of the in vitro findings.
Collectively, these data suggest that AL445238.2 and USP4 may form
a complex regulatory network that governs cell proliferation,
apoptosis, stemness maintenance and migration, ultimately driving
the development and progression of CRC.
The findings of the present study align with and
extend the growing body of evidence regarding lncRNA involvement in
CRC pathogenesis. Consistent with previous reports demonstrating
that lncRNAs regulate critical cellular processes in various cancer
types (38,39), the present study showed that
AL445238.2 modulates multiple hallmarks of cancer progression,
including proliferation, apoptosis resistance, stemness and
migration in CRC cells. Similar to other studies linking aberrant
lncRNA expression with adverse clinicopathological features in
solid tumors (40,41), the results indicated that
AL445238.2 may serve as a prognostic indicator and therapeutic
target in colon cancer. However, the present study provides novel
mechanistic insights by identifying USP4 as a key mediator of
AL445238.2's oncogenic functions. While deubiquitinases have been
implicated in cancer stem cell maintenance and chemoresistance in
other contexts (42), the
specific AL445238.2-USP4 axis represents a previously
uncharacterized regulatory mechanism in CRC. These findings not
only corroborate the functional importance of lncRNAs in CRC
biology but also reveal a specific molecular pathway that may be
exploited for targeted therapeutic intervention.
Despite offering new insights into the molecular
mechanisms of CRC, the present study has certain limitations.
Although both the in vitro and in vivo experiments
supported the cooperative functions of AL445238.2 and USP4, further
elucidation of the precise molecular regulatory mechanisms and
downstream signaling pathways is required. In addition, the limited
sample size and experimental models may restrict the
generalizability of these findings; thus, future studies should
validate these observations across a broader range of clinical
samples and diverse cancer subtypes.
In summary, the present study revealed that
AL445238.2, through its interaction with USP4, promoted CRC cell
proliferation, inhibited apoptosis and enhanced both stemness and
migratory capabilities, thereby providing a novel molecular target
and theoretical foundation for the development of precision
therapies against CRC. Future research should focus on further
delineating the intricacies of this regulatory network and
evaluating its potential clinical applications.
Supplementary Data
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
HL, LZ and SY conceived and designed the study,
including revision. JF and JH performed all the experiments and
bioinformatics analysis. HL and PC performed the statistical
analysis and drafted the manuscript. All the authors read and
approved the final version of the manuscript. LZ and SY confirm the
authenticity of all the raw data.
Ethics approval and consent to
participate
Animal welfare and ethical review for experiments
were conducted by the Institutional Animal Care and Use Committee
of the Wenzhou Medical University Animal Experimentation Ethics
Committee (approval no. xmsq2023-0671).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not appliable.
Funding
The present study was supported by the Medical and Health
Science and Technology Project of Zhejiang Province (grant no.
2024KY1635), the Traditional Chinese Medicine Health Research
Project of Zhejiang Province (grant no. 2024ZL165) and the Wenzhou
Science and Technology Program (grant no. Y2023392).
References
|
1
|
Siegel RL, Wagle NS, Cercek A, Smith RA
and Jemal A: Colorectal cancer statistics, 2023. CA Cancer J Clin.
73:233–254. 2023.PubMed/NCBI
|
|
2
|
Arcos M, Goodla L, Kim H, Desai SP, Liu R,
Yin K, Liu Z, Martin DR and Xue X: PINK1-deficiency facilitates
mitochondrial iron accumulation and colon tumorigenesis. Autophagy.
21:737–753. 2025. View Article : Google Scholar :
|
|
3
|
Chen YJ, Hong WF, Liu ML, Guo X, Yu YY,
Cui YH, Liu TS and Liang L: An integrated bioinformatic
investigation of mitochondrial solute carrier family 25 (SLC25) in
colon cancer followed by preliminary validation of member 5
(SLC25A5) in tumorigenesis. Cell Death Dis. 13:2372022. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hu JL, Wang W, Lan XL, Zeng ZC, Liang YS,
Yan YR, Song FY, Wang FF, Zhu XH, Liao WJ, et al: CAFs secreted
exosomes promote metastasis and chemotherapy resistance by
enhancing cell stemness and epithelial-mesenchymal transition in
colorectal cancer. Mol Cancer. 18:912019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhao H, Ming T, Tang S, Ren S, Yang H, Liu
M, Tao Q and Xu H: Wnt signaling in colorectal cancer: Pathogenic
role and therapeutic target. Mol Cancer. 21:1442022. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Tran TQ, Hanse EA, Habowski AN, Li H,
Ishak Gabra MB, Yang Y, Lowman XH, Ooi AM, Liao SY, Edwards RA, et
al: α-Ketoglutarate attenuates Wnt signaling and drives
differentiation in colorectal cancer. Nat Cancer. 1:345–358. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ren L, Meng L, Gao J, Lu M, Guo C, Li Y,
Rong Z and Ye Y: PHB2 promotes colorectal cancer cell proliferation
and tumorigenesis through NDUFS1-mediated oxidative
phosphorylation. Cell Death Dis. 14:442023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Chen J, Duan S, Wang Y, Ling Y, Hou X,
Zhang S, Liu X, Long X, Lan J, Zhou M, et al: MYG1 drives
glycolysis and colorectal cancer development through
nuclear-mitochondrial collaboration. Nat Commun. 15:49692024.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yun SI, Kim HH, Yoon JH, Park WS, Hahn MJ,
Kim HC, Chung CH and Kim KK: Ubiquitin specific protease 4
positively regulates the WNT/β-catenin signaling in colorectal
cancer. Mol Oncol. 9:1834–1851. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wijnhoven P, Konietzny R, Blackford AN,
Travers J, Kessler BM, Nishi R and Jackson SP: USP4
auto-deubiquitylation promotes homologous recombination. Mol Cell.
85:6652025. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang L, Zhou F, Drabsch Y, Gao R,
Snaar-Jagalska BE, Mickanin C, Huang H, Sheppard KA, Porter JA, Lu
CX and ten Dijke P: USP4 is regulated by AKT phosphorylation and
directly deubiquitylates TGF-β type I receptor. Nat Cell Biol.
14:717–726. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xing C, Lu XX, Guo PD, Shen T, Zhang S, He
XS, Gan WJ, Li XM, Wang JR, Zhao YY, et al: Ubiquitin-specific
protease 4-mediated deubiquitination and stabilization of PRL-3 is
required for potentiating colorectal oncogenesis. Cancer Res.
76:83–95. 2016. View Article : Google Scholar
|
|
13
|
Toden S, Zumwalt TJ and Goel A: Non-coding
RNAs and potential therapeutic targeting in cancer. Biochim Biophys
Acta Rev Cancer. 1875:1884912021. View Article : Google Scholar
|
|
14
|
Yan H and Bu P: Non-coding RNA in cancer.
Essays Biochem. 65:625–639. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Nemeth K, Bayraktar R, Ferracin M and
Calin GA: Non-coding RNAs in disease: From mechanisms to
therapeutics. Nat Rev Genet. 25:211–232. 2024. View Article : Google Scholar
|
|
16
|
Wang J, Ma X, Si H, Ma Z, Ma Y, Wang J and
Cao B: Role of long non-coding RNA H19 in therapy resistance of
digestive system cancers. Mol Med. 27:12021. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Shirvaliloo M: LncRNA H19 promotes tumor
angiogenesis in smokers by targeting anti-angiogenic miRNAs.
Epigenomics. 15:61–73. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Xiu B, Chi Y, Liu L, Chi W, Zhang Q, Chen
J, Guo R, Si J, Li L, Xue J, et al: LINC02273 drives breast cancer
metastasis by epigenetically increasing AGR2 transcription. Mol
Cancer. 18:1872019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Feng YC, Liu XY, Teng L, Ji Q, Wu Y, Li
JM, Gao W, Zhang YY, La T, Tabatabaee H, et al: c-Myc inactivation
of p53 through the pan-cancer lncRNA MILIP drives cancer
pathogenesis. Nat Commun. 11:49802020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
He X, Yang T, Lu YW, Wu G, Dai G, Ma Q,
Zhang M, Zhou H, Long T, Yan Y, et al: The long noncoding RNA
CARDINAL attenuates cardiac hypertrophy by modulating protein
translation. J Clin Invest. 134:e1691122024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yao ZT, Yang YM, Sun MM, He Y, Liao L,
Chen KS and Li B: New insights into the interplay between long
non-coding RNAs and RNA-binding proteins in cancer. Cancer Commun
(Lond). 42:117–140. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Xu Y, Wang J, Deng X, Gao H, Xing F, Ding
H, Qu L and Xi Z: USPs and non-coding RNAs: Unraveling their
interplay within tumor microenvironment for targeted cancer
therapy. Pharmacol Res. 218:1078532025. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xu P, Xiao H, Yang Q, Hu R, Jiang L, Bi R,
Jiang X, Wang L, Mei J, Ding F and Huang J: The USP21/YY1/SNHG16
axis contributes to tumor proliferation, migration, and invasion of
non-small-cell lung cancer. Exp Mol Med. 52:41–55. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
25
|
American Veterinary Medical Associationn
(AVMA): AVMA guidelines for the euthanasia of animals: 2020
edition. AVMA; Schaumburg, IL: 2020
|
|
26
|
Morton D, Seymour M, Magill L, Handley K,
Glasbey J, Glimelius B, Palmer A, Seligmann J, Laurberg S, Murakami
K, et al: Preoperative chemotherapy for operable colon cancer:
Mature results of an international randomized controlled trial. J
Clin Oncol. 41:1541–1552. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Chalabi M, Verschoor YL, Tan PB, Balduzzi
S, Van Lent AU, Grootscholten C, Dokter S, Büller NV, Grotenhuis
BA, Kuhlmann K, et al: Neoadjuvant immunotherapy in locally
advanced mismatch repair-deficient colon cancer. N Engl J Med.
390:1949–1958. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Chen X, Chen J, Feng W, Huang W, Wang G,
Sun M, Luo X, Wang Y, Nie Y, Fan D, et al: FGF19-mediated ELF4
overexpression promotes colorectal cancer metastasis through
transactivating FGFR4 and SRC. Theranostics. 13:1401–1418. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Bertocchi A, Carloni S, Ravenda PS,
Bertalot G, Spadoni I, Lo Cascio A, Gandini S, Lizier M, Braga D,
Asnicar F, et al: Gut vascular barrier impairment leads to
intestinal bacteria dissemination and colorectal cancer metastasis
to liver. Cancer Cell. 39:708–724.e11. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Tan YT, Lin JF, Li T, Li JJ, Xu RH and Ju
HQ: LncRNA-mediated posttranslational modifications and
reprogramming of energy metabolism in cancer. Cancer Commun (Lond).
41:109–120. 2021. View Article : Google Scholar :
|
|
31
|
Bhan A, Soleimani M and Mandal SS: Long
noncoding RNA and cancer: A new paradigm. Cancer Res. 77:3965–3981.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ebrahimi N, Parkhideh S, Samizade S,
Esfahani AN, Samsami S, Yazdani E, Adelian S, Chaleshtori SR,
Shah-Amiri K, Ahmadi A and Aref AR: Crosstalk between lncRNAs in
the apoptotic pathway and therapeutic targets in cancer. Cytokine
Growth Factor Rev. 65:61–74. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang Y, Lu JH, Wu QN, Jin Y, Wang DS, Chen
YX, Liu J, Luo XJ, Meng Q, Pu HY, et al: LncRNA LINRIS stabilizes
IGF2BP2 and promotes the aerobic glycolysis in colorectal cancer.
Mol Cancer. 18:1742019. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wu N, Jiang M, Liu H, Chu Y, Wang D, Cao
J, Wang Z, Xie X, Han Y and Xu B: LINC00941 promotes CRC metastasis
through preventing SMAD4 protein degradation and activating the
TGF-β/SMAD2/3 signaling pathway. Cell Death Differ. 28:219–232.
2021. View Article : Google Scholar
|
|
35
|
Guo Y, Jiang Z, Chen Q, Xie D, Zhou Y, Yin
W, Wang Z, Wang B, Ren C and Jiang X: Construction and experimental
validation of a signature for predicting prognosis and immune
infiltration analysis of glioma based on disulfidptosis-related
lncRNAs. Front Immunol. 14:12913852023. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Lang X, Huang C and Cui H: Prognosis
analysis and validation of fatty acid metabolism-related lncRNAs
and tumor immune microenvironment in cervical cancer. J Immunol
Res. 2022:49544572022. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Pan J, Huang Z, Lin H, Cheng W, Lai J and
Li J: M7G-Related lncRNAs predict prognosis and regulate the immune
microenvironment in lung squamous cell carcinoma. BMC Cancer.
22:11322022. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Shin CH, Kim K, Ho CW, Lee JW, Jo MJ, Min
KW and Yoon JH: Long noncoding RNAs regulating enzymatic reactions
in cancer. Exp Mol Med. 57:1641–1650. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Segal D and Dostie J: The talented
LncRNAs: Meshing into transcriptional regulatory networks in
cancer. Cancers (Basel). 15:34332023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Mehrad-Majd H, Akhtari J, Haerian MS and
Ravanshad Y: Clinicopathological and prognostic value of lncRNA
PANDAR expression in solid tumors: Evidence from a systematic
review and meta-analysis. J Cell Physiol. 234:4206–4216. 2019.
View Article : Google Scholar
|
|
41
|
Liang C, Liu J, Ge H, Xu Y, Li G and Wu J:
The clinicopathological and prognostic value of long non-coding RNA
ZEB1-AS1 in solid tumors: A meta-analysis. Clin Chim Acta.
484:91–98. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Qiu GZ, Sun W, Jin MZ, Lin J, Lu PG and
Jin WL: The bad seed gardener: Deubiquitinases in the cancer
stem-cell signaling network and therapeutic resistance. Pharmacol
Ther. 172:127–138. 2017. View Article : Google Scholar
|