Introduction
Gastric cancer (GC) is the fifth leading main cause
of cancer-associated morbidity and mortality worldwide, accounting
for 5.6% of all new cancer cases (1,089,103 cases) and 7.7% of all
cancer deaths (768,793 deaths) globally in 2020 (1). Although emerging therapies such as
immunotherapy (2) and targeted
therapy (3) have been developed,
the prognosis remains poor due to late-stage diagnosis, limiting
treatment efficacy. Globally, ~50% of GC patients are diagnosed at
advanced (unresectable or metastatic) stages as of 2020, with an
overall 5-year survival rate of only around 20% for all stages
combined (1). Thus, elucidating
the molecular mechanisms underlying GC initiation and progression
is key.
RNA-binding proteins (RBP) are key regulators of
gene expression through their roles in RNA metabolism (4,5).
Most RBPs interact with mRNA and pre-mRNA, participating in
splicing, stabilization and transport (5,6).
Dysregulated RBP expression is implicated in numerous human
diseases (6,7). For example, poly(A) binding protein
cytoplasmic 1 (PABPC1) stabilizes p21-activated kinase 1 mRNA,
promoting epithelial-mesenchymal transition (EMT) and GC
migration/invasion (7).
DAZ-associated protein 1 (DAZAP1), a part of the heterogeneous
nuclear ribonucleoprotein (hnRNP) family, is an RBP (8,9).
Abnormal expression of DAZAP1 regulates cancer cells (8,10).
For example, DAZAP1 is significantly upregulated in oral squamous
cell carcinoma (OSCC), promoting cell migration, invasion and the
EMT process (10). It regulates
post-transcriptional gene expression by binding target mRNAs via
the 'UAGKWWR' motif (9,11,12). For example, DAZAP1 enhances
hepatocellular carcinoma progression and modulates ferroptosis
through interaction with SLC7A11 mRNA (12) and facilitates alternative splicing
of kit ligand (KITLG) to trigger numerous myeloma cell
proliferation (13). However, its
role in regulating mRNA expression in GC remains poorly
understood.
Notch is a canonical signaling pathway that
comprises four Notch receptors (Notch 1, 2, 3 and 4) and five Notch
ligands [Δ-like 1, 3 and 4 and Jagged (JAG)1 and 2] in mammals
(14,15). NOTCH1 is most commonly found in
cancer tissue (15,16). For example, elevated expression of
NOTCH1 is associated with tumor invasion and lymph node metastasis
of GC (16). Moreover, RBP
insulin-like growth factor 2 mRNA-binding protein 2 directly binds
T cell acute lymphoblastic leukemia oncogene NOTCH1 mRNA (17). JAG1 is highly expressed in GC and
colorectal and breast cancer (18-21). JAG1 promotes cell
migration/invasion and is associated with poor prognosis in breast
cancer (20). Furthermore, RBP
zinc finger protein 36 (ZFP36) directly binds JAG1 mRNA and is
involved in mRNA decay (21). In
addition, the Notch/JAG pathway is key in EMT during embryonic
development, fibrotic disorder and cancer metastasis (22,23). Activation of Notch signaling
induces EMT in tumor cells and JAG1-mediated Notch signaling
triggers the EMT process, migration and invasion of malignant
cells; these effects are abrogated by Notch silencing (23). To the best of our knowledge,
however, the mechanism by which DAZAP1 post-transcriptionally
regulates NOTCH1 or JAG1 expression to affect GC progression has
not been examined.
The present study aimed to explore DAZAP1 expression
in GC, elucidate its regulatory role in GC cell behavior and EMT
progression, verify its effect on the mRNA stability of NOTCH1 and
JAG1 and define the role of the DAZAP1-NOTCH1/JAG1 axis in GC
development and its potential as a therapeutic target.
Materials and methods
Cell culture
Normal human gastric mucosal GES-1 and GC cell lines
(HGC-27, MKN-74, SNU-216, MKN-45, AGS and SNU-719) were obtained
from American Type Culture Collection and cultured in RPMI-1640
medium enriched with 10% FBS (both purchased from Gibco (Thermo
Fisher Scientific). All cells were cultivated in a 37°C incubator
with 5% CO2.
Data acquisition and bioinformatics
analysis
The microarray or RNA sequencing datasets GSE70880
(ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE70880),
GSE99416 (ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE99416) and
GSE109476 (ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE109476) were
downloaded from the Gene Expression Omnibus (GEO) database and
analysis was conducted to obtain the differentially expressed genes
(DEGs) between GC and healthy tissue samples, with the screening
criterion set as a fold change >1.2. DEGs were integrated with
RNA-binding protein (RBP) prediction data downloaded online from
the RBPmap website (rbpmap.technion.ac.il/) (24). Bioinformatics analysis, including
the construction of Venn diagrams, was performed using R software
(v4.0.2; r-project.org/). The present study used
expression levels of DAZAP1 from the FireBrowse (firebrowse.org/) and UALCAN (ualcan.path.uab.edu/) databases to further validate
the results obtained from bioinformatics analysis. Gene searches
were performed using the PubMed database (pubmed.ncbi.nlm.nih.gov/). To obtain EMT-related
genes, the gene set from the EMT database (emtome.org)
(25). The Cancer Genome Atlas
(cancer.gov/tcga) database was used for
bioinformatics analysis, focusing on the STAD (stomach
adenocarcinoma) cohort. Kyoto Encyclopedia of Genes and Genomes
(KEGG) pathway enrichment analysis for target genes was conducted
with the WebGestalt database (webgestalt.org/). Analysis of potential transcription
factors of DAZAP1 was performed using the AnimalTFDB v4.0 database
(guolab.wchscu.cn/AnimalTFDB4/#/).
Tissue samples
Gastric adenocarcinoma tissue microarray (TMA; cat.
no. HStm-Ade180Sur-04) was acquired from Shanghai Outdo Biotech Co
Ltd. The TMA analysis included 90 GC samples and 90 matched
adjacent healthy tissue samples (70 male; 20 female; median age, 65
years; age range, 34-83 years; distance, 5 cm). The use of tissue
chips was approved by the Ethics Committee of Shanghai Outdo
Biotech Co. Ltd. (approval no. SHYJS-CP-1804015).
A total of 10 formalin-fixed and paraffin-embedded
GC specimens and adjacent healthy tissue specimens were collected
from patients (seven male; three female; median age, 63 years; age
range, 42-81 years) with GC at Nanfang Hospital (Guangzhou, China)
between December 2023 and January 2024. Normal tissues were
collected 5 cm from the tumor edge and immediately stored in liquid
nitrogen at −196°C for subsequent experimentation. The diagnosis of
primary GC was validated by two pathologists. Patients who had
received chemotherapy, radiotherapy or molecular-targeted therapy
before surgery were excluded. These tissues were used for western
blotting, reverse transcription-quantitative (RT-q) PCR and RNA
in situ hybridization (ISH). The study received ethical
approval from Nanfang Hospital, Southern Medical University
(approval no. NFEC-202105-K18; Nanfang, China). All patients
provided written informed consent.
Immunohistochemistry (IHC) and TMA
analysis
A total of 10 paired GC tumor and adjacent
non-cancerous gastric mucosa samples were surgically obtained in
2020 from the Department of Surgery, Nanfang Hospital, Southern
Medical University, for analysis by IHC and western blotting.
Table SI presents the antibodies
used. All experimental protocols received approval from the Ethics
Committee of Southern Medical University, China.
Following 24 h fixation with 4% paraformaldehyde at
room temperature, tumor specimens were embedded in paraffin wax.
Sections of 4-mm thickness were cut from paraffin-embedded tissue
blocks and placed on glass slides. The slides were deparaffinized
using xylene, rehydrated using ethanol. For antigen retrieval,
sections were microwaved in 0.01 mol/l sodium citrate buffer
(pH=6.0) at 95°C for 15 min. After immersing the sections in 3%
hydrogen peroxide for 10 min at room temperature to inhibit
endogenous peroxidase activity. Blocking was performed at room
temperature for 20 min using 5% BSA; cat. no. A8850, Solarbio
Science & Technology Co., Ltd.) to eliminate non-specific
binding. The sections were then incubated overnight at 4°C with the
primary antibody against DAZAP1(Santa Cruz Biotechnology Co.,
Ltd.). Subsequently, they were incubated with the biotin-linked
anti-Mouse IgG (cat. no. ab6823, dilution 1:1,000; Abcam) at room
temperature for 30 min. Following another three PBS washes,
chromogen detection was performed using DAB. The staining results
were observed under a light microscope and analyzed using Image J
1.8 software (National Institutes of Health). Normal rabbit or
mouse IgG (Sigma-Aldrich; Merck KGaA) was used as the isotype
controls. The TMA was stained for DAZAP1 (Santa Cruz Biotechnology
Co., Ltd.) according to the same protocol.
DAZAP1 staining intensity in carcinoma cells was
scored as follows: 0 (negative), 1 (weak), 2 (moderate), and 3
(intense). Cells with scores of 0-1 were classified as low
expression, while scores of 2-3 indicated high expression. The
final IHC score for each sample was the average of evaluations by
two independent observers.
Western blotting
Total protein from tissue or cells was extracted via
RIPA lysis buffer (Beijing Solarbio Science & Technology Co.,
Ltd.) containing a protease inhibitor (MedChemExpress). The protein
concentration of lysates was measured using the BCA assay. A total
of 20 μg total protein were separated by 10% SDS-PAGE and
were transferred to PVDF membranes. The membranes were treated with
5% non-fat dry milk at room temperature for 30 min. The membranes
were blotted with primary antibodies (Table SI) overnight at 4°C, followed by
incubation with HRP-conjugated secondary anti-rabbit or anti-mouse
antibodies (Abcam; cat. nos. ab7090 and ab97046; 1:6,000) for 1 h
at room temperature. We confirm that β-tubulin (cat. no.
10094-1-AP, 1:5,000; Proteintech) was used as the reference
protein. Enhanced chemiluminescence was utilized for visualization
with the Pierce ECL Western Blotting Substrate (Thermo Fisher
Scientific, Waltham, MA, USA), and densitometric analysis was
performed using Image J 1.8 (National Institutes of Health,
Bethesda, MD, USA).
Plasmid construct, transfection,
lentiviral production and infection
DAZAP1-pEnter, DAZAP1-flag, DAZAP1-RNA recognition
motif 1 (RRM1)-flag, DAZAP1-RRM2-flag, DAZAP1-C-terminal domain
(CTD)-flag, NOTCH1-pEnter and JAG1-pEnter plasmids were constructed
by Kidan Biosciences. Human DAZAP1 small interfering RNA (siRNA),
including siRNA1, siRNA2 and siRNA pool (siRNAp; a 1:1 mixture of
siRNA1 and siRNA2 with the total siRNA amount constant in the
transfection system). NOTCH1-siRNAp, JAG1-siRNAp and scrambled
siRNA non-specific control (Scr-siRNA) were synthesized by Shanghai
GenePharma Co., Ltd. DAZAP1, NOTCH1 and Scramble shRNA were all
purchased from GenePharma Co., Ltd. Both short hairpin (sh)RNAs and
siRNAs were used. The siRNAs were applied for short-term functional
assays, while shRNAs were used to establish stable cell lines and
xenograft assays. Briefly, cells were plated in 6-well culture
plates at a density of 3×105 cells per well. At 30-50%
confluency, the cells were transfected with 50 nM target siRNA or
Scr-siRNA via Lipo8000 (Beyotime Institute of Biotechnology)
transfection reagent, at 37°C for 6 h. Subsequent experiments were
conducted on the cells at 48 h post-transfection, and the
transfection efficiency was validated by RT-qPCR) and western blot
assays.
To establish stable AGS cell lines, the cells were
transduced with lentiviruses for target gene manipulation
(2nd-generation lentiviral system) purchased from GeneChem, Inc.
The lentiviral vectors were engineered to express green fluorescent
protein (GFP) as a reporter gene; with overexpression and knockdown
constructed via GV348 and GV493 vectors (1 μg/μl,
GenePharma, Inc). Lentiviral particles were produced by
co-transfecting 293T cells (ATCC, USA) with 8 μg lentiviral
target plasmid, 4 μg packaging and 2 μg envelope
plasmid at 37°C for 6 h; viral supernatants were collected at 48-72
h post-transfection. AGS cells were seeded at 5×105
cells/well in 6-well plates and cultured to 80% confluency, then
transduced with the lentivirus at MOI=100 with polybrene and
transfection reagent A (GenePharma, Inc.) at 37°C for 24-48 h
following the manufacturer's protocol. Puromycin (10 μg/ml)
was added at 72 h post-transduction for stable cell selection over
7 days; stably infected cells were maintained in 10 μg/ml
puromycin and subjected to subsequent experiments 2-3 days after
the completion of selection. Table
SII lists the siRNA and shRNA target sequences.
RT-qPCR
Total RNA was isolated from AGS and MKN-45 cells, as
well as 10 pairs of human GC and their adjacent normal tissues. The
extraction was performed using TRIGene Plus reagent (cat. no.
ZP108-101S, Genestar) in accordance with the manufacturer's
instructions. RT was performed in strict accordance with the
manufacturer's instructions for the was conducted via
Hieff® Fast Cell SYBR Green RT-qPCR Kit (cat. no.
11172ES40, Shanghai Yeasen Biotechnology Co., Ltd.). RT-qPCR was
conducted via SYBR Green Master Mix (Shanghai Yeasen Biotechnology
Co., Ltd.) according to the manufacturer's guidelines. The
thermocycling conditions were as follows: Initial denaturation at
95°C for 2 min, followed by 40 cycles of denaturation at 95°C for
15 sec and combined annealing/extension at 60°C for 30 sec
(two-step PCR). Quantification of relative levels of genes was
conducted via the 2−ΔΔCq method (26), and then normalization to the
internal reference gene GAPDH was conducted. Table SIII illustrates the primer
sequences.
Colony-forming assay
A total of 1,000 transfected cells were plated into
12-well plates and cultivated in a 37°C, 5% CO2
incubator for 10-12 days. All liquids were removed, 4%
paraformaldehyde was used to fix them for 30 min at room
temperature, then crystal violet was used for staining at room
temperature for 15 min. Colonies were defined as cell populations
large enough to be observed with the naked eye and were counted
manually directly under a Zeiss microscope.
EdU assay
MKN-45, AGS and MKN-74 cells were cultured in 10
μM EdU-supplemented medium (Guangzhou RiboBio Co., Ltd.;
cat. no. #C10310) for 4 h at 37°C. Following removal of the
EdU-supplemented medium, 4% paraformaldehyde was used to fix the
cells at room temperature for 60 min. After washing with PBS
solution and subsequent incubation with Apollo and Hoechst 33258
dyes at 37°C, images were obtained via a fluorescence microscope.
Proportion of EdU-positive staining was calculated as follows:
EdU-stained cells/Hoechst-stained cells) ×100%.
Wound healing assay
Serum starved MKN-45, AGS and MKN-74 cells were
seeded into a 6-well culture plate at a density of 5×105
cells/well and grown to 100% confluence. After cells were rinsed
with PBS, they were scratched. Images were captured under an
inverted light microscope. For MKN-45 cells, imaging was performed
at 0, 24 and 48 h. For AGS and MKN-74 cells, imaging was conducted
from 0 to 24 h at 12 h intervals. Migration index was calculated as
follows: (Original wound width-wound width after healing)/original
wound width ×100%.
Migration and invasion assay
Migration assays were performed via Transwell
chambers in 24-well plates (BD Biosciences; cat. no. #353092).
MKN-45 (8×104), AGS (2×104 cells) or MKN-74
cells (2×104 cells) were suspended in 100 μl
serum-free RPMI 1640 medium (Gibco, Thermo Fisher Scientific) and
plated in the top chambers, while the bottom chambers contained 700
μl RPMI-1640 medium with 20% FBS. After incubating for 48 h
at 37°C, non-migrated cells on the top membrane surface were
removed with a cotton swab. Then, 4% paraformaldehyde was utilized
to fix the migrated cells on the bottom surface for 60 min at room
temperature, 0.1% crystal violet was used to stain them for 60 min
at 37°C and counting was conducted in four randomly selected
high-power fields of view/sample under a light microscope (Zeiss
microscope). For invasion assays, the top chambers were coated with
Matrigel (cat. no. #358248, Corning, Inc.) at 37°C for 30 min to
allow for solidification; all other procedures were identical to
the migration assay.
F-actin cytoskeleton staining
MKN-45 (1×104 cells) were cultured in a
24-well plate overnight at 37°C. Cells were fixed with 4%
paraformaldehyde at room temperature for 15 min, permeabilized with
0.1% Triton X-100 at room temperature for 10 min, and blocked with
5% FBS at room temperature for 30 min. F-actin staining was
performed with rhodamine-conjugated phallotoxin (5 U/ml, Molecular
Probes; Thermo Fisher Scientific, Inc.) according to the
manufacturer's guidelines. Nuclei were stained with Hoechst33258 at
room temperature for 10 min in the dark. F-actin cytoskeleton
staining was assessed using a fluorescence microscope.
Immunofluorescence staining
Following washing in PBS, 4% paraformaldehyde was
used to fix the MKN-45 cells for 15 min at room temperature.
Following permeabilization with PBS containing 0.2% Triton X-100
for 5 min, cells were blocked with 10% goat serum (cat. no.
ZLI-9056, Beijing Zhongshan Jinqiao Biotechnology Co., Ltd.) at
37°C for 30 min. Cells were incubated with primary antibodies
against E-cadherin (1:200, Proteintech Group, Inc.) and Vimentin
(1:200, Cell Signaling Technology, Inc.) at 4°C overnight. After
three rinses with PBS, the cells were incubated with
fluorochrome-conjugated secondary antibodies (1:100, cat. no.
ZF-0512/ZF-0513, Beijing Zhongshan Jinqiao Biotechnology Co., Ltd.,
Alexa Fluor® 488/Alexa Fluor® 594 Conjugate)
at 37°C for 1 h in the dark. Nuclear counterstaining was performed
with Hoechst 33258 dye at room temperature for 10 min in the dark,
followed by two washes with PBS to remove excess dye. The stained
cells were observed under a fluorescence microscope and all
fluorescent images were acquired and analyzed using Image J 1.8
(National Institutes of Health, Bethesda, MD, USA).
RNA immunoprecipitation (RIP)
Magna RIP RBPs Immunoprecipitation kit
(MilliporeSigma) was used according to the manufacturer's
instructions. In brief, cells were subjected to centrifugation at
300 × g for 5 min at 4°C, the cells were suspended in 200 μl
RIP lysis buffer (MilliporeSigma) provided in the kit that
contained 1 μl protease inhibitor cocktail and 0.5 μl
RNase inhibitor. A total of 200 μl lysate was used per IP
reaction. A total of 5 μg anti-mouse DAZAP1 and mouse IgG
isotype control antibody (MilliporeSigma) were added to the
magnetic bead suspension, and the mixtures were gently shaken at
room temperature for 1 h to form bead-antibody complexes. The
bead-antibody complexes were allowed to sit at 4°C overnight.
Magnetic bead-RNA-protein complexes were washed three times with
the RIP wash buffer 1 and 2 provided in the kit using a magnetic
stand for magnetic separation of the complexes. After the
RNA-protein complexes coupled to the beads were isolated, total RNA
was extracted from the isolated complexes and subjected to
sequencing analysis for target gene detection.
RIP sequencing (RIP-seq)
Anti-DAZAP1 antibody was utilized to
immunoprecipitate DAZAP1-RNA complexes, with input as the control
(IgG). DAZAP1-bound RNAs were extracted via TRIzol, and a cDNA
library was created via the TruSeq RNA library preparation kit
(Illumina, Inc. cat. no. RS-122-2201), the quality and integrity of
the processed RNA samples were verified using a Agilent 2100
Bioanalyzer (Agilent Technologies) prior to library construction.
Sequencing was performed on the Illumina HiSeq X Ten platform by
ABLife Inc. with 150 bp paired-end sequencing; the sequencing
reaction was conducted using the HiSeq X Ten Reagent kit v2.5 (300
cycles; cat. no. FC-510-1003; Illumina, Inc.). The final library
was loaded at a concentration of 10 pM, and the molar concentration
was measured via Qubit 4 Fluorometer with Qubit dsDNA HS Assay kit
(Thermo Fisher Scientific, Inc.; cat. no. Q32854). Raw sequencing
data were analyzed using HISAT2 (v2.2.1; daehwankimlab.github.io/hisat2/), StringTie (v2.2.1;
ccb.jhu.edu/software/stringtie/) and DESeq2 (v1.38.3;
bioconductor.org/packages/release/bioc/html/DESeq2.html)
for read alignment, transcript assembly and differential expression
analysis, respectively.
mRNA stability assay
Scr-siRNA, DAZAP1-siRNAp, DAZAP1-siRNA1 and
DAZAP1-siRNA2 were transfected into cells. After 24 h at 37°C, the
medium was substituted with medium (Gibco; Thermo Fisher
Scientific; cat. no. 11875093). enriched with actinomycin D (10
μg/ml, MedChemExpress) or DMSO. Total RNA was collected at
0, 2, 4, 6 and 8 h post-actinomycin D treatment using TRIGene Plus
reagent (cat. no. ZP108-101S, Genestar) and mRNA half-life was
estimated by determining NOTCH1 and JAG1 mRNA levels at each time
point via RT-qPCR (normalized to GAPDH RNA) and curve fitting with
the one-phase decay model in GraphPad Prism 9 (Dotmatics). NOTCH1
or JAG1 mRNA relative levels were assessed by RT-qPCR as
aforementioned. mRNA levels were normalized to those of GAPDH
RNA.
RNA in situ hybridization (ISH)
ISH was conducted to observe NOTCH1 or JAG1
expression in GC specimens with locked nucleic acid-based ISH via
digoxigenin-labeled miRCURY miRNA probes (Exiqon; Qiagen GmbH). DNA
probes designed for the spatial expression analysis of mRNA were
derived from the cDNA plasmid pGEM-T Easy (Promega, cat. no.
A1360). Both probes had a length of 36 nucleotides, and the
specific sequences of the probes used for ISH are illustrated in
Table SIV. GC tissue was fixed
with 4% paraformaldehyde at 4°C for 24 h, embedded in paraffin and
cut into 4 μm-thick serial sections on poly-L-lysine-coated
glass slides (Thermo Fisher Scientific; cat. no. 12-550-15). The
slides were deparaffinized, rehydrated and incubated with
Proteinase K (Roche Diagnostics; cat. no. 3115887001) at 37°C for
15 min, slides were then subjected to post-fixation with 4%
paraformaldehyde at room temperature for 10 min and followed by
washing with PBS. Pre-hybridization was performed with 5X
saline-sodium citrate buffer (SSC; Sigma-Aldrich; cat. no. S6639)
hybridization buffer (without probe) at room temperature for 15
min, followed by probe denaturation at 95°C for 1 min. NOTCH1 mRNA
and JAG1 mRNA probes (final concentration, 50 nM) were applied for
hybridization in 5X SSC hybridization buffer with probe at 50°C for
1 h. The slides were rinsed with 2.0 and 0.1X SSC (at 50°C for 15
min each (30 min total) to remove excess probes, then blocked with
10% goat serum (Thermo Fisher Scientific; cat. no. 16210064) at RT
for 30 min. An overnight incubation was performed with an
anti-digoxigenin alkaline phosphatase-conjugated primary antibody
[1:1,000, Roche Diagnostics (Shanghai) Co,. Ltd.; cat. no.
11093274910] at 4°C.BCIP/NBT Chromogenic Detection System (Roche
Diagnostics; cat. no. 11681451001) was used to visualize the
hybridization signals. Nuclear fast red (Sigma-Aldrich; cat. no.
N3020) was used to counterstain the slides at room temperature for
1 min, and mounting was performed via an aqueous solution. For each
patient, four slides were hybridized with the NOTCH1 mRNA or JAG1
mRNA probe. Hybridization signals were observed under a light
microscope and ImageJ v1.8 (National Institutes of Health) was used
for signal intensity analysis.
Xenograft assay
A total of 24 nude female BALB/c mice (nu/nu; age,
4-6 weeks; weight, 20-22 g) were obtained from the Guangdong
Medical Laboratory Animal Center (Guangzhou, China) and housed in
specific-pathogen-free conditions were as follows: Temperature,
20-26°C; humidity, 50-70%; light/dark cycle, 12/12-h; free access
to food/water. The Animal Ethics Committee of Southern Medical
University and Nanfang Hospital approved the animal experiments
(approval no. IACUC-LAC-20250111-002).
For the tail vein-lung metastasis model, two groups
of mice were utilized to assess the metastatic capability of
malignant cells. The first group comprised the shRNA and
DAZAP1-shRNA groups, while the second group included Vector +
Scr-shRNA, DAZAP1 + Scr-shRNA, Vector + NOTCH1-shRNA, DAZAP1 +
NOTCH1-shRNA, DAZAP1 + NOTCH1-shRNA + Vector and DAZAP1 +
NOTCH1-shRNA + JAG1. An injection of 5×106 lentivirus
(LV)-transfected cells was administered into the tail veins of nude
mice, which were euthanized after 30 days via CO2
asphyxiation using a gradual-fill approach, with the gas introduced
at a rate of 30% of the chamber volume/min. Cervical dislocation
was performed to confirm death. Death was confirmed by
disappearance of pain response, no reaction when the toes were
pressed with hands or tweezers and cessation of heartbeat and
respiration. The humane endpoint was body weight loss >20%; no
animals met this endpoint during the experimental period.
Subsequently, the lung metastases were counted. Paraffin-embedded
lung tissues were sectioned into 4 μm sections and stained
with hematoxylin and eosin and vimentin antibody as
aforementioned.
Statistical analysis
Statistical analyses were performed using SPSS
version 25.0 (IBM Corp.) and GraphPad Prism 10.1.2 (Dotmatics). All
experiments were conducted in triplicate. All data are presented as
the mean ± SD. For normally distributed continuous variables,
Student's t-test (paired) or one-way ANOVA with Dunnett's post hoc
test was applied as appropriate. For non-parametric comparisons,
the Mann-Whitney U test was used. The Pearson correlation
coefficient was used to assess the correlation between variables.
Kaplan-Meier survival analysis was performed to evaluate overall or
progression-free survival and the log-rank test was used to compare
survival differences between groups. A two-tailed P-value <0.05
was considered to indicate a statistically significant
difference.
Results
DAZAP1 is upregulated in GC
To identify novel protein coding genes involved in
the onset and progression of GC, the present study assessed the
microarray data of GSE70880, GSE99416 and GSE109476 downloaded from
the GEO database and visualized the DEGs in GC and normal tissue
samples (Fig. 1A). A total of 19
genes were up- and 17 were downregulated in the GSE109476 dataset,
81 genes were up- and 30 downregulated in the GSE70880 dataset and
48 genes were up- and 37 downregulated in the GSE99416 dataset. The
intersection of the datasets demonstrated that only DAZAP1 was
upregulated, and three genes (PPARGC1A, CIRBP, PABPC4) were
downregulated. DAZAP1 was selected for further study due to its
enhanced potential for use as an early diagnostic marker or
intervention target (8-12). DAZAP1 was highly expressed in most
tumors compared with adjacent healthy tissues based on FireBrowse
and UALCAN databases (Fig. S1A and
B). Expression of DAZAP1 was increased in GC compared with
normal samples (Fig. 1B).
Compared with normal gastric cell line GES-1, DAZAP1 protein
expression was markedly upregulated in HGC-27, MKN-74, SNU-216,
MKN-45, AGS and SNU-719 cells (Fig.
1C). IHC was performed to determine the intensity of the DAZAP1
staining (Fig. 1D). TMA showed
upregulation of DAZAP1 in tumor tissue (Fig. 1Ea). DAZAP1 deposition was
displayed in the stroma, with scattered malignant cells in the
peripheral margin of the tumors (tumor-associated stromal cells),
and the cells displayed EMT-like morphological alterations,
including the loss of typical epithelial cobblestone morphology,
acquisition of a fusiform mesenchymal shape and increased cell
scattering and spindle-like polarization. Semiquantitative scoring
illustrated that DAZAP1 protein was significantly upregulated in
cancer compared with adjacent healthy gastric mucosa tissue
(Fig. 1Eb).
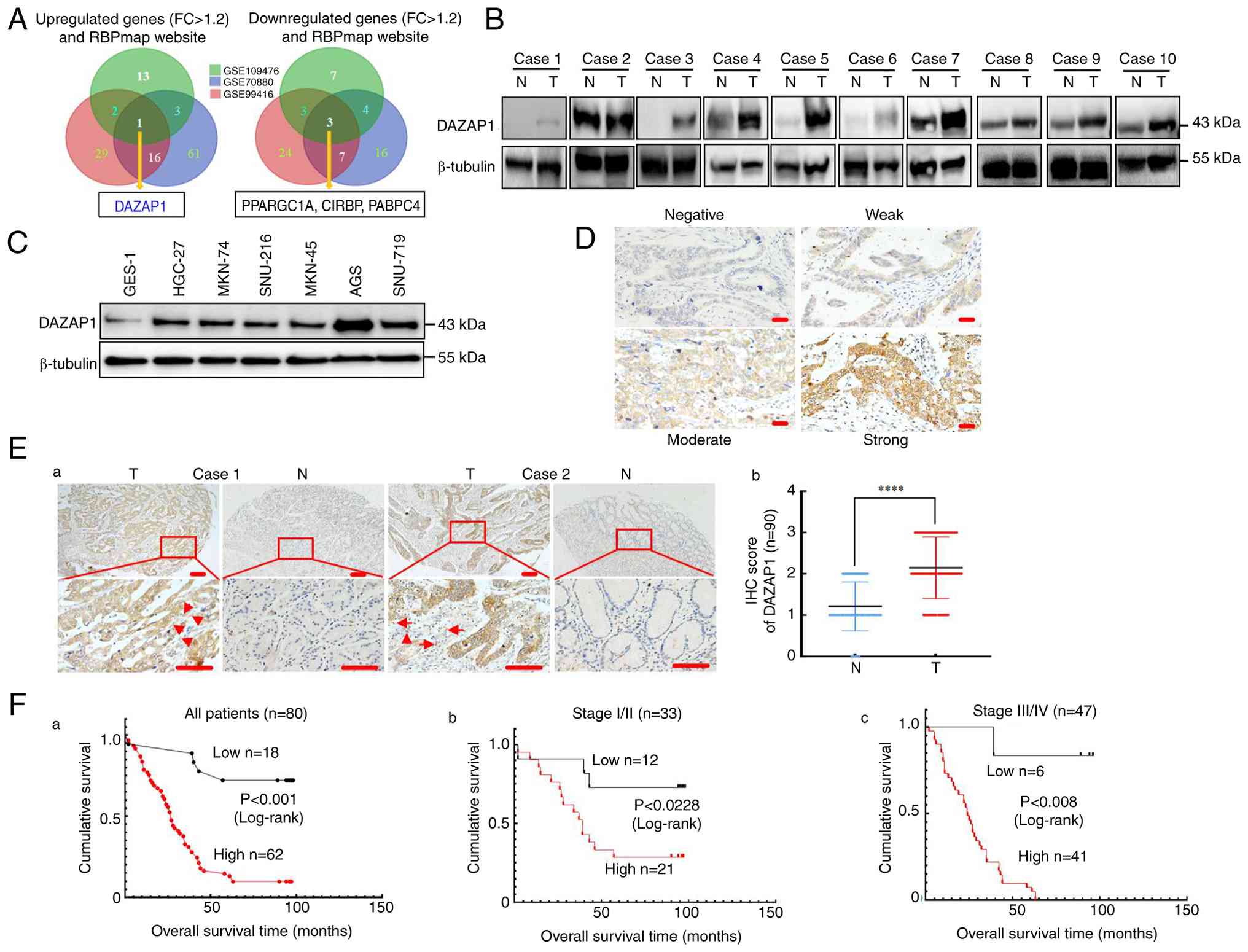 | Figure 1DAZAP1 is upregulated in GC. (A)
Differentially expressed genes in the three Gene Expression Omnibus
datasets and the RBPmap website. (B) Relative expression of DAZAP1
in 10 GC and their adjacent N tissues was examined by western blot
assay. β-tubulin was used as the internal control. (C) Expression
of DAZAP1 protein in six human GC and normal mucous cell line
GES-1. (D) Representative staining of DAZAP1 intensity using IHC.
(Ea) Representative gastric tissue from two cancerous and
non-cancerous patients. Expression of DAZAP1 in normal and
malignant human gastric tissue was detected by tissue microarray.
The arrows indicate the cells with epithelial-to-mesenchymal
transition) morphological alterations. (Eb) Expression of DAZAP1
was determined in N and T gastric tissue. Scale bar, 100 μm.
Kaplan-Meier survival analysis of overall survival in (Fa) all
patients and those with (Fb) early and (Fc) late stage GC
(according to DAZAP1 expression. The log-rank test was used to
calculate P-values. ****P<0.001. DAZAP, deleted in
azoospermia-associated protein 1; GC, gastric cancer; N, normal
tissue; T, tumor tissue; PPARGC1A, peroxisome
proliferator-activated receptor gamma coactivator 1 alpha; CIRBP,
cold inducible RNA-binding protein; PABPC4, poly(A)-binding protein
cytoplasmic; FC, fold-change; IHC, immunohistochemistry. |
The present study analyzed correlations between
DAZAP1 expression and clinicopathological characteristics and
prognosis in 90 GC tissue samples. High DAZAP1 expression was
significantly associated with local invasion (T1/2 vs. T3/4),
American Joint Committee on Cancer (AJCC) stage (I/II vs. III/IV),
lymph node metastasis (N0 vs. N1) and tumor size (<5 vs. ≥5 cm).
However, DAZAP1 was not linked to additional clinical
characteristics, such as age, sex and differentiation in GC
(Table SV). Kaplan-Meier
survival analysis demonstrated a relationship between DAZAP1 levels
and overall survival time (Fig.
1Fa). This correlation detected in individuals with late-stage
GC (AJCC stage III/IV; Fig. 1Fc)
was more noticeable than that in early-stage GC (AJCC stage I/II;
Fig. 1Fb). DAZAP1 expression
demonstrated predictive value for overall survival in early-(AJCC
I/II) and late-stage (AJCC III/IV) GC patient subgroups. These data
illustrated that DAZAP1 is upregulated and has an oncogenic
function in GC.
DAZAP1 overexpression promotes the
malignant biological behavior of GC
MKN-45 GC cell line with relatively low and AGS and
MKN-74 cell lines with relatively elevated endogenous DAZAP1 levels
were selected for transfection and functional experiments (Fig. 1C). Stable transfectants were
constructed via DAZAP1-sense plasmids in MKN-45 cells (Fig. S2A) or DAZAP1 was suppressed using
siRNAp in AGS and MKN-74 cells (Fig.
S2B), as demonstrated via western blot analysis and qPCR.
Subsequently, colony formation assays showed that overexpression of
DAZAP1 led to significantly more colonies than the vector in MKN-45
cells (Fig. 2A). EdU assay showed
that overexpression of DAZAP1 significantly induced cell
proliferation compared with the control vector in MKN-45 cells
(Fig. 2B). Proliferation of AGS
and MKN-74 cells was inhibited by the knockdown of DAZAP1 (Fig. 2A and B).
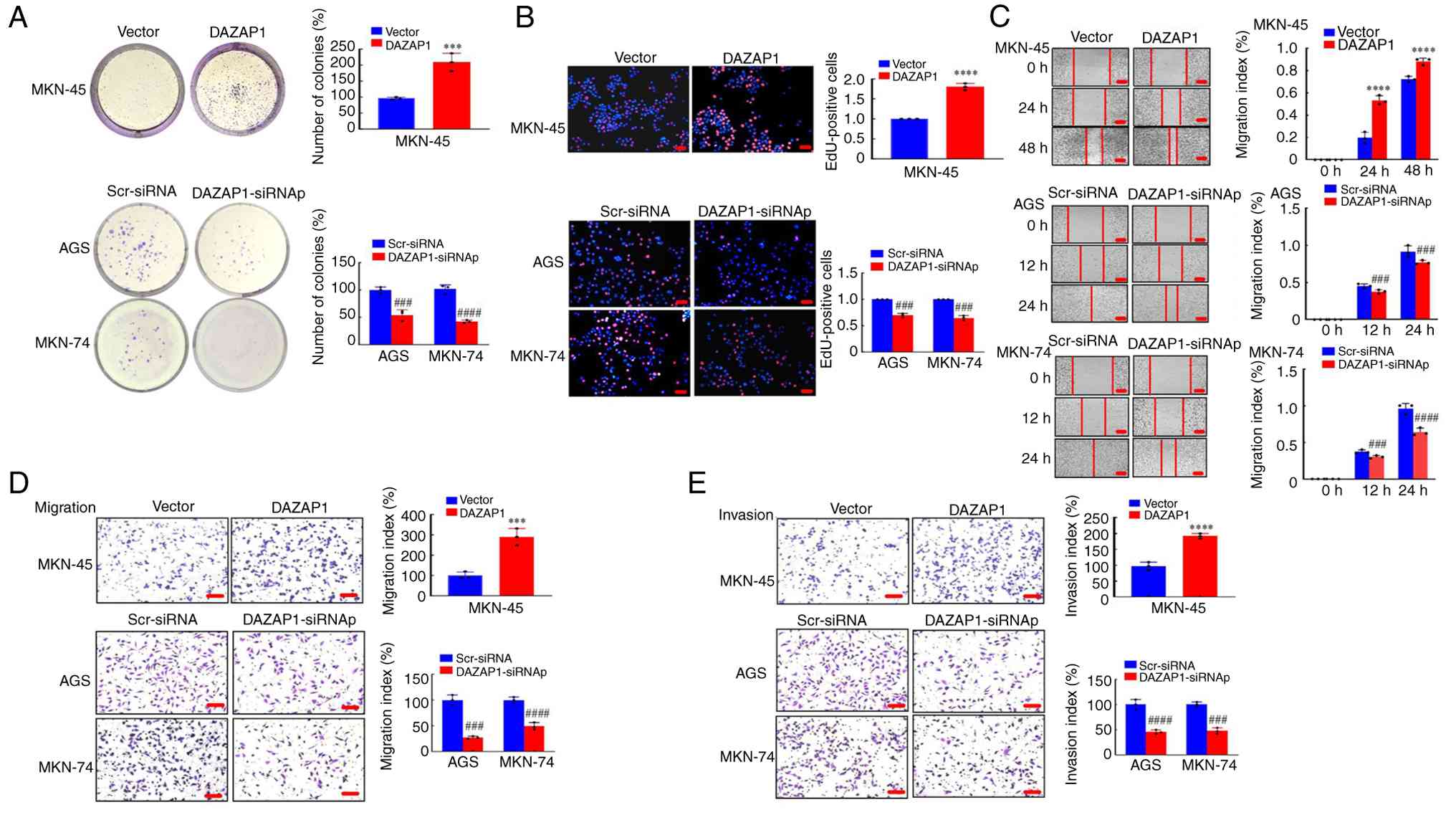 | Figure 2Functional analysis of DAZAP1 in GC
cells in vitro. (A) Colony formation assay was performed to
detect proliferative capability at 48 h. (B) EdU assay was used to
determine the viability of GC cells. Red, EdU-positive cells; blue,
Hoechst33258 (total cells). Scale bar, 100 μm. (C) Motility
was determined wound healing assay. (D) Migration and (E) invasion
assay were performed following transfection in GC cells. Scale bar,
150 μm. ***P<0.01, ****P<0.001
vs. vector; ###P<0.01, ####P<0.001
Scr-siRNA vs. DAZAP1-siRNAp. DAZAP1, deleted in
azoospermia-associated protein 1; GC, gastric cancer; scr,
scramble; si, small interfering; p, pool. |
The present study explored the effect of DAZAP1 on
cell migration and invasion in GC. Wound healing assay demonstrated
that the migration ability was promoted by DAZAP1 overexpression in
MKN-45 cells, whereas DAZAP1 knockdown had the opposite effect on
AGS and MKN-74 cells (Fig. 2C).
Transwell experiments with or without Matrigel demonstrated a
significant increase in migration and invasion following DAZAP1
overexpression and a significant decrease in migration and invasion
in response to DAZAP1 knockdown (Fig.
2D). All these data illustrated that DAZAP1 mediated malignant
phenotypes in GC cells in vitro.
DAZAP1 induces EMT in GC cells
A key step in tumor spreading is EMT, which occurs
when epithelial cells associated with tumors transform into
mesenchymal cells (27,28). The present study assessed cell
morphology to see if DAZAP1 is essential for GC EMT. By contrast
with the control cells, the MKN-45 cells that had DAZAP1
overexpression had an elongated spindle-like phenotype, which was
distinguishable from normal mesenchymal cells (Fig. 3A).
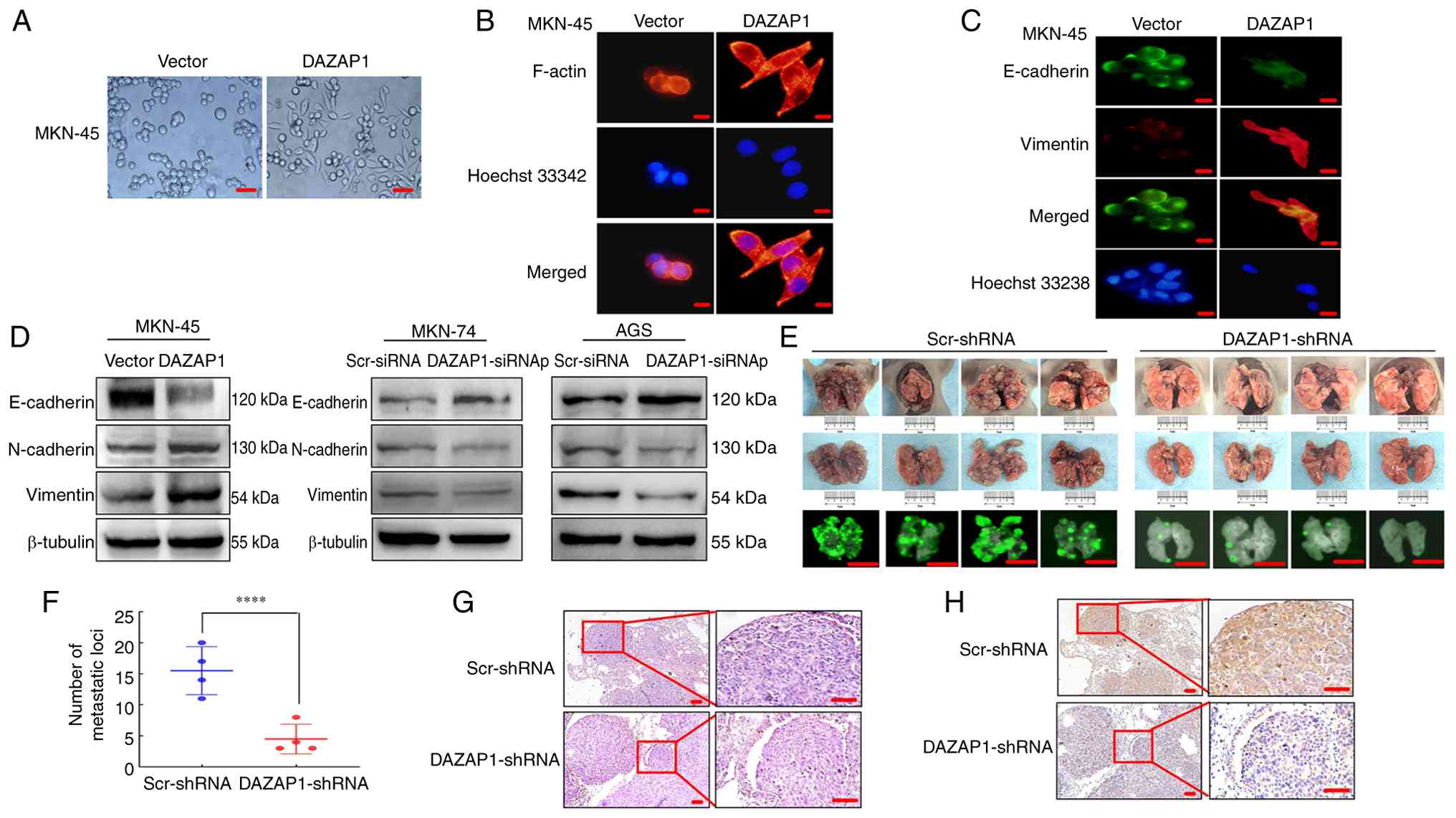 | Figure 3DAZAP1 induces EMT in GC cells. (A)
Phase-contrast microscopy in MKN-45 cells. Scale bar, 50 μm.
(B) F-actin expression and localization were analyzed by
immunocytochemistry. The cytoskeleton and nuclei were stained with
rhodamine-phallotoxin (red) and Hoechst33258 (blue). (C)
Immunofluorescence staining for E-cadherin (green) and vimentin
(red) in MKN-45 cells. Scale bar, 20 μm. (D) EMT biomarkers,
including E-cadherin, N-cadherin and vimentin, were detected by
western blotting in GC cells. (E) Representative metastatic tumors
in the lungs from mice (n=4/group). Scale bar, 1 cm. (F) Number of
metastatic tumors in the lung was counted. (G) Hematoxylin and
eosin staining of lung sections of mice. (H) Immunohistochemical
staining of vimentin. Scale bar, 100 μm.
****P<0.001. DAZAP, deleted in azoospermia-associated
protein 1; EMT, epithelial-mesenchymal transition; GC, gastric
cancer; scr, scramble; si, small interfering; sh, short
hairpin. |
Previous studies have shown that EMT improves
motility and actin reorganization of cells (29,25). Therefore, F-actin was stained with
phalloidin. In DAZAP1-overexpressing cells, F-actin filaments were
more dispersed throughout the cytoplasm and localized at the
protrusion rim compared with control cells (Fig. 3B). Immunofluorescence analysis
revealed that DAZAP1 overexpression in MKN-45 cells downregulated
E-cadherin and upregulated vimentin expression (Fig. 3C). Western blot analysis confirmed
that DAZAP1 transfection altered the expression of EMT-associated
markers. Overexpression of DAZAP1 significantly suppressed
E-cadherin, whereas knockdown of DAZAP1 increased the expression of
E-cadherin in GC cells (Fig. 3D).
Conversely, overexpressing DAZAP1 promoted the expression of
N-cadherin and vimentin, while knockdown of DAZAP1 suppressed their
expression in GC cells (Fig.
3D).
Additionally, to explore the association between
DAZAP1 and migration/invasion, the function of DAZAP1 in metastases
was examined by injecting AGS/LV-pEGFP-DAZAP1-shRNA and
AGS/LV-pEGFP-Scr-shRNA expressing GFP into nude mice (Fig. 3E). The knockdown efficiency of
DAZAP1-shRNA was verified in AGS cells (Fig. S4C). Smaller lung metastatic
nodules were detected in the LV-DAZAP1-shRNA compared with the
LV-pEGFP-Scr-shRNA group (Fig.
3F). Histological examination confirmed the presence of GC cell
metastases in the lung tissue (Fig.
3G). Moreover, IHC analysis demonstrated that vimentin protein
levels were promoted in cancer tissue of the LV-pEGFP-Scr-shRNA
compared with those in the LV-DAZAP1-shRNA group (Fig. 3H). These results indicated that
DAZAP1 in GC cells may be a vital contributor to EMT progression,
thereby mediating the ability of GC cells to migrate and
invade.
DAZAP1 physically binds NOTCH1 or JAG1
mRNA to regulate its stability
As DAZAP1 is an RBP and regulates mRNAs by binding
UAGKWWR motifs (9,11,12) (Fig.
4A), RIP-seq was performed in AGS cells to detect potential RNA
targets. A total of 980 genes (1,376 transcripts) potentially bound
to DAZAP1 (Table SVI).
EMT-related genes were obtained from the EMT database (emtome.org) (Table
SVII) (25,30).
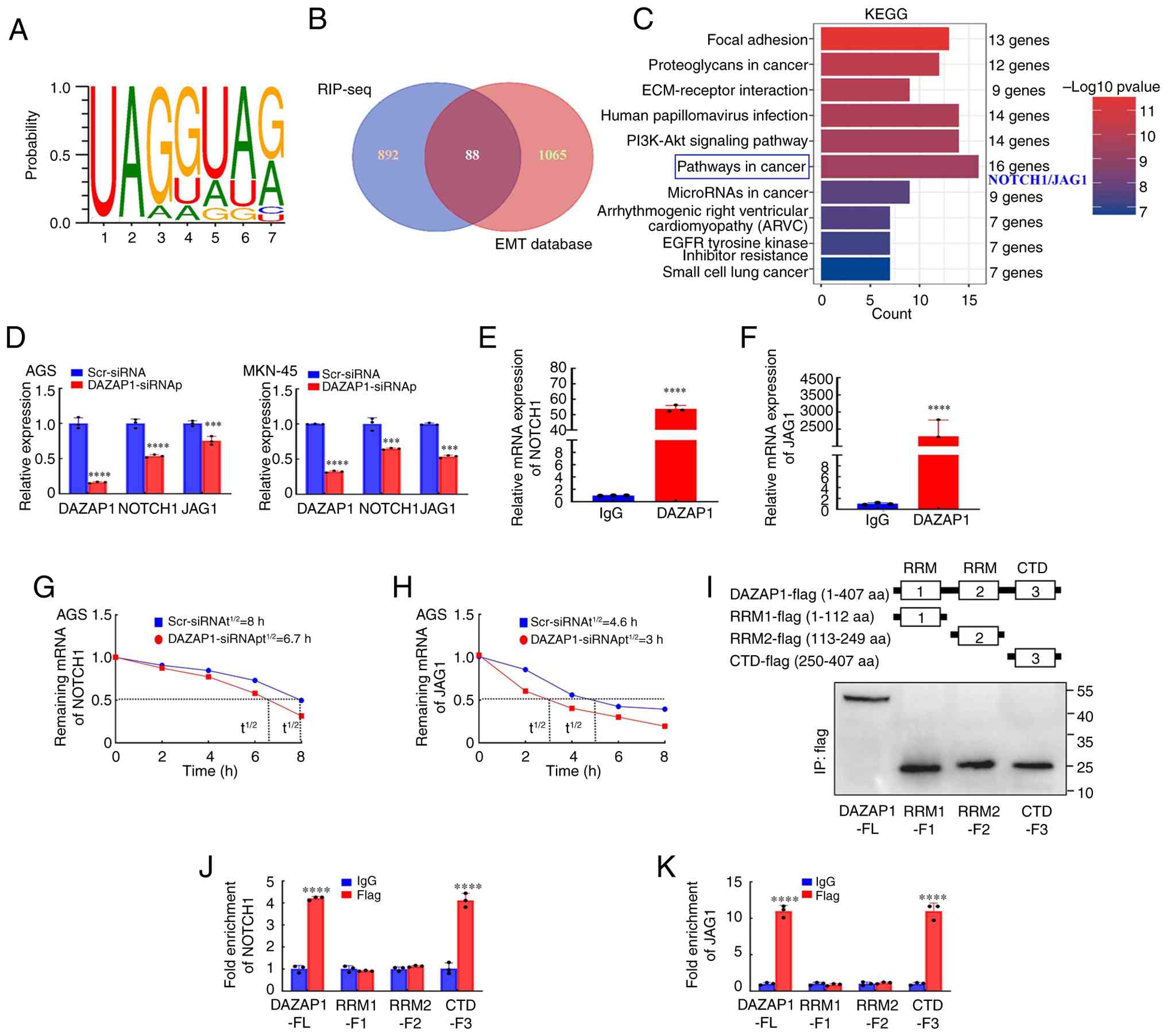 | Figure 4DAZAP1 binds NOTCH1 or JAG1 mRNA in
GC cells. (A) DAZAP1-binding motif was predicted using the RBPmap
database (rbpmap.technion.ac.il/). (B) Overlap between
DAZAP1-interacting genes identified using RIP-seq and EMT-related
genes from the EMT database (emtome.org).
(C) Quantification of target genes involved in the pathways in the
cancer signaling pathway, including NOTCH1 and JAG1, according to
KEGG analysis in the WebGestalt database (webgestalt.org/). (D) RT-qPCR analysis of the mRNA
levels of NOTCH1 and JAG1 following DAZAP1 knockdown in AGS and
MKN-45 cells. RT-qPCR analysis of (E) NOTCH1 and (F) JAG1 with
anti-DAZAP1 and IgG control antibodies in AGS cells. (G) NOTCH1 and
(H) JAG1 mRNA t1/2 in DAZAP1 knockdown AGS cells. n=3.
(I) Verification of full-length or truncated DAZAP1 plasmids using
western blotting and anti-FLAG antibody. RIP assay analysis of (J)
NOTCH1 and (K) JAG1 mRNA binding to the DAZAP1 structure domains.
***P<0.01 and ****P<0.001 vs.
scr-siRNA. RRM, RNA recognition motif; CTD, C-terminal domain;
DAZAP1, deleted in azoospermia-associated protein 1; RIP-seq, RNA
Immunoprecipitation-sequencing; EMT, Epithelial-Mesenchymal
Transition; KEGG, Kyoto Encyclopedia of Genes and Genomes; RT-q,
Reverse Transcription quantitative real-time; t1/2,
half-life; scr, scramble; si, small interfering; F, Fragment; FL,
Full-Length; aa, amino acid. |
The 980 DAZAP1-bound genes and the EMT-related genes
from the database were used to construct a Venn diagram for
identifying common genes (Fig.
4B). The intersection illustrated 88 genes (Table SVIII), among which 44 genes were
associated with GC. These target genes were used for KEGG) pathway
enrichment analyses. A total of 10 significantly enriched KEGG
pathways, including 'pathways in cancer', were identified (Fig. 4C; Table SIX). NOTCH1 and JAG1 are
essential in tumor progression and enriched in 'pathways in cancer'
(16,18). Therefore, NOTCH1 and JAG1 were
selected for subsequent study.
qPCR was performed to determine NOTCH1 and JAG1 mRNA
expression in GC cells with DAZAP1 suppression; DAZAP1 knockdown
group exhibited a significant decrease in NOTCH1 and JAG1 mRNA
expression compared with the negative control group (Fig. 4D). RIP was performed to determine
whether DAZAP1 binds to the NOTCH1 or JAG1 genes. Compared with IgG
control, DAZAP1 antibody demonstrated notable enrichment of NOTCH1
and JAG1 RNA (Fig. 4E and F). The
results demonstrated a physical association between the DAZAP1
protein and either NOTCH1 or JAG1 mRNA in vitro.
The present study explored the effect of DAZAP1 on
NOTCH1 or JAG1 mRNA stability. Actinomycin D assay demonstrated
that the half-life of NOTCH1 or JAG1 mRNA was significantly lower
following DAZAP1 knockdown than in the control group in AGS cells
(Fig. 4G and H). These data
suggest that DAZAP1 can increase NOTCH1 or JAG1 expression by
binding NOTCH1 or JAG1 mRNA and enhancing its stability.
DAZAP1 protein is composed of three primary domains:
RRM1 (10-97 aa), RRM2 (113-190 aa) and CTD (250-407 aa). To
investigate the domain required for DAZAP1-mediated NOTCH1 or JAG1
expression, DAZAP1 [full-length (FL)], including RRM-F1 (1-112 aa),
RRM2-F2 (113-249 aa) and CTD-F3 (250-407 aa) plasmids with a
FLAG-tag, were constructed (Fig.
4I). The in vitro binding assay showed that for NOTCH1
(Fig. 4J) and JAG1 (Fig. 4K) mRNA, the Flag group of
full-length DAZAP1 (-FL) and its CTD domain (-F3) exhibited
markedly high fold enrichment, while RRM1 (-F1) and RRM2 (-F2) only
had weak enrichment. Thus, CTD is key for DAZAP1 to physically bind
NOTCH1/JAG1 mRNA, thereby regulating their stability.
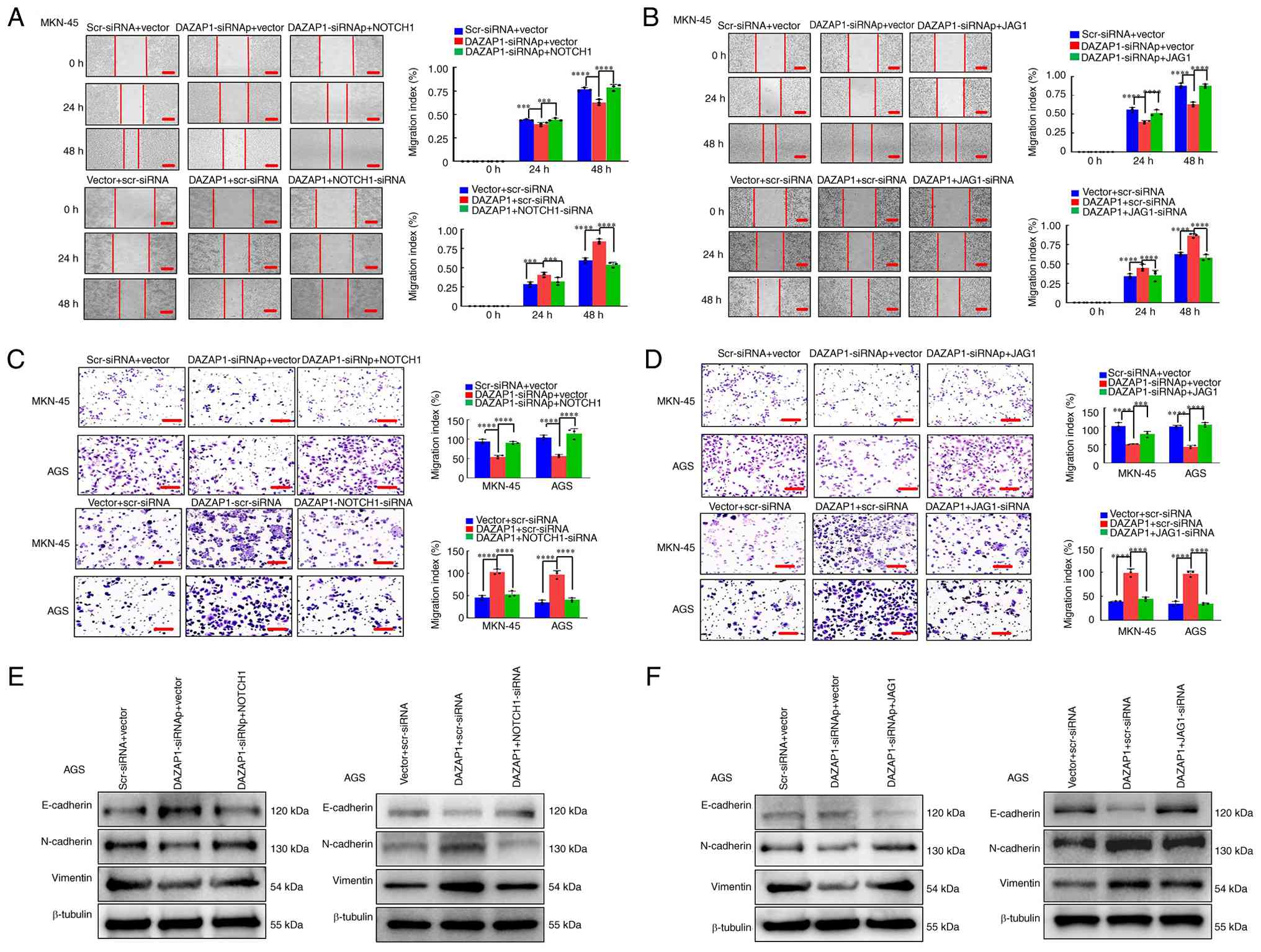
Overexpression of DAZAP1 promotes NOTCH1-
or JAG1-mediated migration/invasion via EMT in GC cells
Ligands for NOTCH1 are transmembrane proteins
expressed on adjacent cells, including JAG and Δ (14,15,23). The present study assessed whether
DAZAP1/NOTCH1 or DAZAP1/JAG1 pathways contribute to GC cell
progression. First, the present study constructed stable
transfectants using NOTCH1- and JAG1-sense plasmids or knockdown of
NOTCH1 and JAG1 with siRNAp in MKN-45 and AGS cells (Fig. S3A), as confirmed by western blot
and RT-qPCR analysis. DAZAP1 knockdown attenuated the migratory and
invasive abilities of GC cells, whereas overexpression of NOTCH1 or
JAG1 partially reversed the phenotypes induced by DAZAP1
suppression compared with the control, as demonstrated by wound
healing and Transwell assays (Figs.
5A-D and S4A-D). Rescue
experiments showed that DAZAP1 overexpression markedly increased
cell motility ability, which was abrogated by downregulation of
NOTCH1 or JAG1 (Figs. 5A-D and
S4A-D).
EMT-associated marker levels were detected by
western blotting, indicating that DAZAP1 suppression elevated the
expression of the epithelial marker (E-cadherin) and decreased
expression of mesenchymal markers (vimentin and N-cadherin) in GC,
Notably, this effect was reversed when NOTCH1 or JAG1 was
introduced alongside DAZAP1 suppression (Figs. 5E and F and S4E-F). Conversely, DAZAP1
overexpression decreased E-cadherin expression and increased
vimentin/N-cadherin levels in GC- and this trend was abolished when
NOTCH1 or JAG1 expression was concurrently reduced. (Figs. 5E-F and S4E-F). These results indicated that
upregulation of DAZAP1 facilitated NOTCH1 or JAG1 expression to
promote motility of GC cells via EMT.
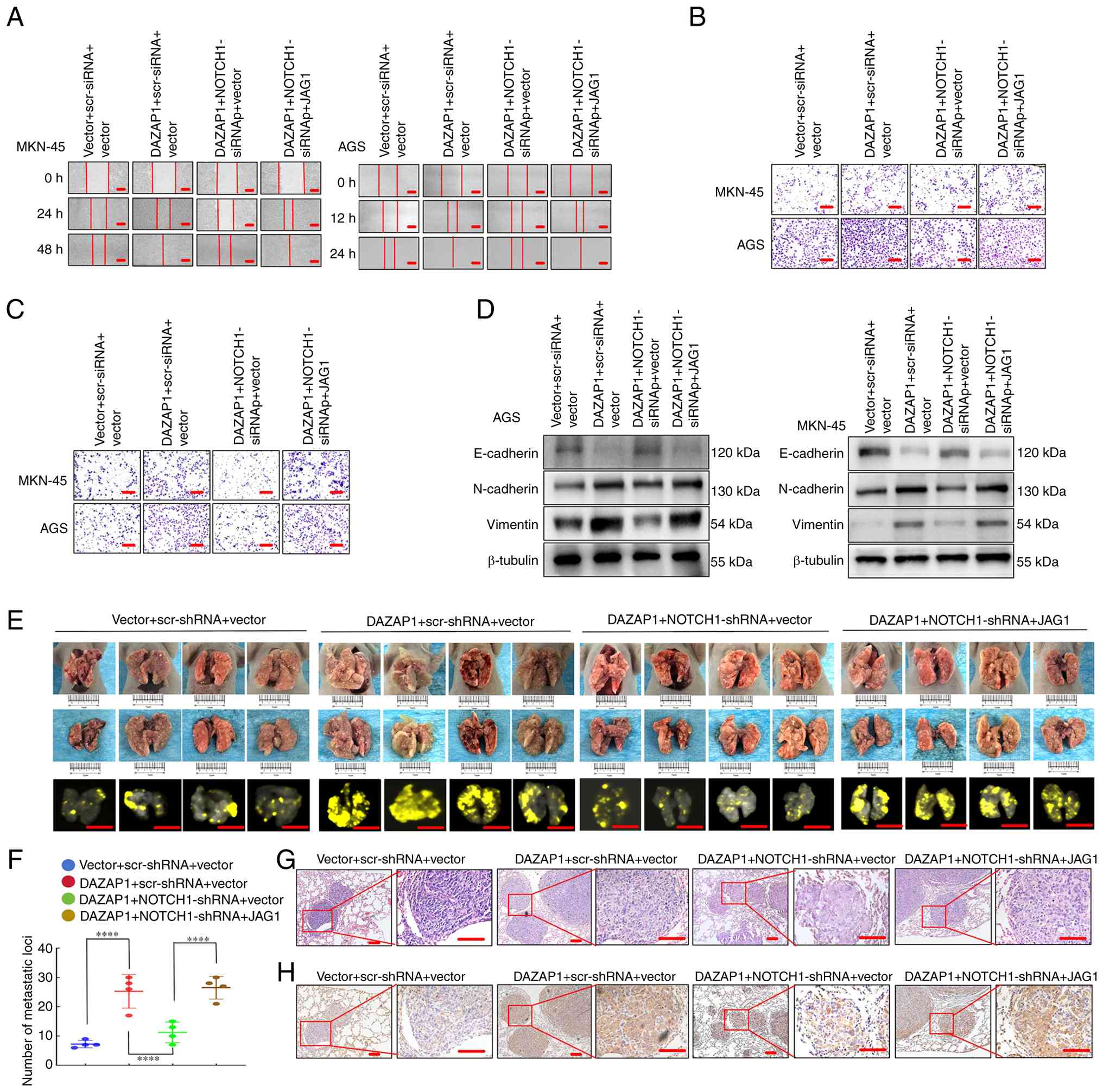
Forced expression of DAZAP1 facilitates
NOTCH1- and JAG1-mediated migration/invasion via EMT in GC
cells
The present study examined whether NOTCH1 and JAG1
mediate the influence of DAZAP1 on GC cell migration and invasion.
DAZAP1 overexpression vector increased motility capacity compared
with the control, while DAZAP1/NOTCH1-siRNA suppressed
migration/invasion capacity compared with DAZAP1 in GC cells
(Figs. 6A-C and S5A and B). Reintroduction of JAG1
partially reversed the suppression compared with
DAZAP1/NOTCH1-siRNA, as demonstrated by wound healing and Transwell
assay (Figs. 6A-C and S5A and B).
Levels of epithelial marker E-cadherin were
downregulated in DAZAP1 compared with control, while
DAZAP1/NOTCH1-siRNAed result in upregulated E-cadherin compared
with DAZAP1. However, E-cadherin upregulation was partly reversed
following DAZAP1/NOTCH1-siRNA/JAG1 transfection. Mesenchymal
markers vimentin and N-cadherin were upregulated in the DAZAP1
overexpression group relative to the vector, downregulated in the
DAZAP1/NOTCH1-siRNA group compared with the DAZAP1 overexpression
group and partially restored following DAZAP1/NOTCH1-siRNA/JAG1
transfection (Fig. 6D).
To investigate the effect of DAZAP1-NOTCH1-JAG1 on
cell metastasis in vivo, the AGS cells expressing the
LV-vector, LV-DAZAP1, DAZAP1/NOTCH1-shRNA or
DAZAP1/NOTCH1-shRNA/JAG1 were implanted into the tail vein of nude
mice, thereby causing lung metastases (Fig. 6E). The knockdown efficiency of
NOTCH1-shRNA was verified in AGS cells (Fig. S3C). More metastatic tumors were
determined in DAZAP1 overexpression compared with the vector group;
DAZAP1/NOTCH1-shRNA caused a significant decrease in the number of
visible tumors than in the control (Fig. 6F). However, the reintroduction of
JAG1 partially reversed the suppression compared with
DAZAP1/NOTCH1-shRNA, which was associated with an increased count
of metastasis loci (Fig. 6F). The
GC metastases in the lung of nude mice were confirmed through
hematoxylin and eosin staining (Fig.
6G), and the difference in cell motility was detected by
vimentin IHC staining. Vimentin is a well-characterized mesenchymal
marker that is associated with cancer cell migratory and invasive
potential as well as metastatic ability (31-32). Vimentin was upregulated in the
DAZAP1 overexpression group relative to the vector, downregulated
in the DAZAP1/NOTCH1-siRNA group compared with the DAZAP1
overexpression group and partially restored following
DAZAP1/NOTCH1-siRNA/JAG1 transfection (Fig. 6H). Therefore, DAZAP1 modulated
NOTCH1/JAG1 signaling in GC cells to promote invasion and
metastasis.
DAZAP1 expression is positively
associated to NOTCH1 or JAG1 expression in GC
The present study assessed the expression of
DAZAP1, NOTCH1 and JAG1 in 10 freshly collected GC biopsy samples.
Western blotting (Fig. 7A) and
RT-qPCR (Fig. 7B and C)
demonstrated that the expression of DAZAP1, NOTCH1 and JAG1 was
significantly upregulated compared with adjacent normal tissue.
DAZAP1 expression was positively associated with NOTCH1 or JAG1
expression (Fig. 7D and E).
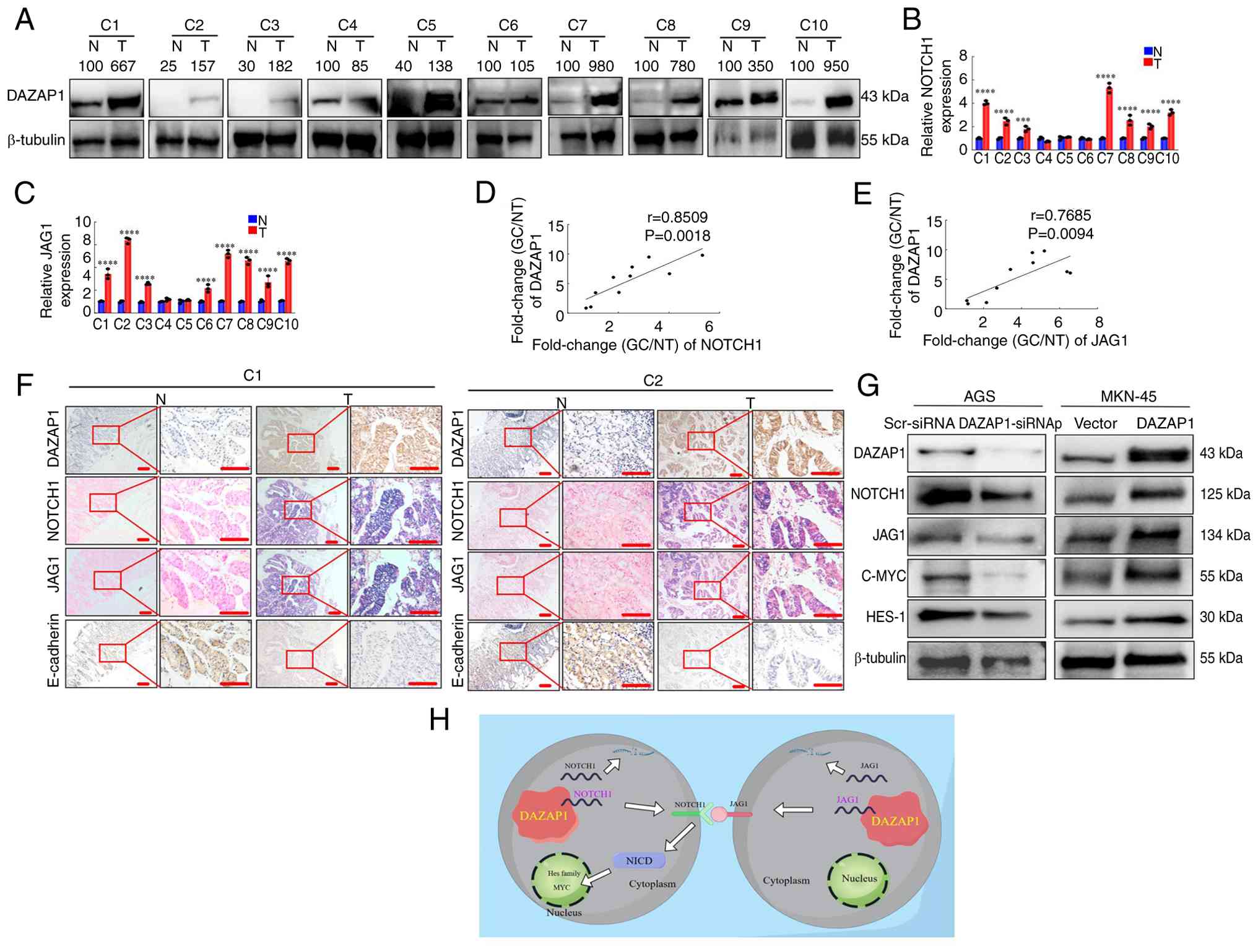 | Figure 7DAZAP1 expression is positively
correlated with NOTCH1 or JAG1 mRNA expression in GC tissues. (A)
Expression of DAZAP1 and (B) Expression levels of NOTCH1 in 10
paired N and GC tissue specimens were detected by qPCR.
***P<0.01, ****P<0.001 (C) Expression
levels of JAG1 in 10 paired N and GC tissue specimens were detected
by qPCR. ****P<0.001 (D) Correlation between DAZAP1
and (D) NOTCH1 in 10 GC tissue samples. (E) Correlation between
DAZAP1 and JAG1 in 10 GC tissue samples. (F) Gene expression in N
and GC tissue was evaluated using immunohistochemistry and in
situ hybridization. (G) Abnormal expression of DAZAP1 modulated
the NOTCH1 signaling pathway, as detected by western blot analysis.
(H) Signal transduction mechanism of DAZAP1. Scale bar, 100
μm. DAZAP1, deleted in azoospermia-associated protein 1; GC,
gastric cancer; N, normal; T, tumor; HES, Hairy and Enhancer of
Split 1; NICD, Notch intracellular domain; C, case; scr, scramble;
si, small interfering. |
IHC and ISH were demonstrated that the expression
of DAZAP1, NOTCH1 and JAG1 was significantly higher, while that of
E-cadherin was significantly lower, in GC compared with healthy
tissue (Fig. 7F).
Aberrant NOTCH signaling activation is typically
linked to its receptors and ligands (33-35). To determine whether DAZAP1 impacts
the NOTCH1 signaling pathway, the present study examined the
expression of key components in the NOTCH1 pathway using western
blot analysis in GC cells. Knockdown of DAZAP1 decreased NOTCH1,
JAG1, MYC and hairy and enhancer of split homolog-1 (HES-1) protein
expression in AGS cells. Conversely, forced expression of DAZAP1
resulted in increased NOTCH1, JAG1, MYC and HES-1 levels in MKN-45
cells.
Collectively, these data demonstrated that DAZAP1
expression was positively associated with levels of NOTCH1 and JAG1
in GC samples and DAZAP1 facilitated the activation of the NOTCH1
signaling pathway.
For the potential transcription factors of DAZAP1,
predictions were made using the AnimalTFDB v4.0 database, which
identified a total of 796 putative transcription factors (Table SX).
Discussion
Numerous investigations have concluded that several
molecular mechanisms participate in the genesis, prognostication
and treatment options in GC (2,7,16,28). However, GC progression remains
uninvestigated. The present study identified a novel target,
DAZAP1, in GC cell progression via the EMT process, which could
promote NOTCH1 and JAG1 mRNA expression and mediate the stimulation
of the NOTCH1 pathway. Hence, the DAZAP1-NOTCH1-JAG1 axis may
participate in GC progression (Fig.
7H).
DAZAP1 gene is positioned on the short arm of human
chromosome 19, region 19p13, and is part of a subfamily of hnRNP
genes (8,9). Its abnormal expression is associated
with certain types of cancers. For example, DAZAP1 is upregulated
in hepatic carcinoma, which indicates a poor prognosis (36). Moreover, DAZAP1 is significantly
upregulated in pancreatic cancer (PC) tissue and cell lines
compared with normal counterparts (37). DAZAP1 suppression inhibits PC cell
proliferation and triggers ferroptosis, while inhibiting
ferroptosis reverses these impacts, promoting PC cell proliferation
and migration/invasion (37).
Nevertheless, the role of DAZAP1 in GC is unknown. Here, high
DAZAP1 expression was significantly associated with poor prognosis
in patients with GC. Furthermore, DAZAP1 overexpression markedly
enhanced GC cell motility and invasion, whereas DAZAP1 knockdown
markedly decreased these. Upregulation of DAZAP1 increased the
levels of the EMT-associated genes N-cadherin and vimentin and
decreased E-cadherin levels. Previous reports have shown that
heterogeneous nuclear ribonucleoprotein F, part of the hnRNP
family, regulates EMT in bladder cancer (38,39), which agrees with the present data.
These outcomes illustrated that DAZAP1 has an oncogenic function
and triggers GC onset and progression.
DAZAP1 is a multifunctional RBP (10,12,13). For example, DAZAP1 enhances
cytochrome c oxidase 16 expression by regulating pre-mRNA
alternative splicing, thereby promoting OSCC invasion and
mitochondrial respiration (10).
DAZAP1 triggers tumor progression in GC stem cells and modulates
the splicing and expression of the mitophagy-associated gene unc-51
like autophagy activating kinase 1 (40). The present study searched for
putative gene targets of DAZAP1 and demonstrated that DAZAP1
physically associated with NOTCH1 or JAG1 mRNA to enhance NOTCH1 or
JAG1 mRNA stability. Moreover, the specific binding domains of
DAZAP1 in NOTCH1 or JAG1 were confirmed. Among the NOTCH receptors,
NOTCH1 has been extensively investigated and identified in tumor
tissue more often than any other member (16,23,35,36). Previous studies have demonstrated
elevated expression of NOTCH1 in GC tissue are associated with
tumor migration/invasion, depth of invasion and vessel invasion
(16,41). JAG1 is a canonical ligand for
Notch receptors, and its upregulation has been observed in numerous
types of cancer, such as GC and breast cancer (BC), where it
facilitates proliferation and migration/invasion (18,20,21) Formerly, reports have shown that
RBP ZFP36 family members ZFP36L1 and ZFP36L2 suppress NOTCH1
expression via interaction with sequences in the NOTCH1 3'
untranslated region (UTR) to modulate tumor development and
progression (42,43). ZFP36 binds the 3' UTR of JAG1 and
post-transcriptionally suppresses JAG1 expression, leading to
decreased JAG1 protein levels and impaired angiogenic sprouting
(21), which were consistent with
the present findings. In addition, wound healing and Transwell
assays revealed that DAZAP1 increased GC cell motility, while
DAZAP1 knockdown abrogated the facilitation of GC cell invasion and
migration mediated by NOTCH1 or JAG1 knockdown, thereby regulating
the EMT process in GC cells. Thus, the present data indicated that
DAZAP1 was a common binding partner of NOTCH1 or JAG1 and
associated with migration and invasion in GC cells.
NOTCH pathways are typically regarded as
ligand-induced (44,45), and the receptors and ligands are
highly co-expressed in malignancy (45,46). NOTCH ligand JAG1 is hypothesized
to enhance the metastatic potential of tumor cells via
NOTCH-dependent induction of EMT (23,47). NOTCH suppression prevents
JAG1-mediated NOTCH signaling from promoting EMT, migration and
invasion of BC cells (23).
Nevertheless, the mechanism by which NOTCH1 and JAG1 are regulated
by DAZAP1 to promote GC cell invasiveness needs further
investigation. Here, overexpression of DAZAP1 caused an increase in
migration/invasion capacity compared with vector, while DAZAP1 +
NOTCH1-siRNAp suppressed motility compared with DAZAP1 in GC cells.
Yet, the reintroduction of JAG1 partially reversed the suppression
compared with DAZAP1 + NOTCH1-siRNAp in GC cells. Western blotting
demonstrated upregulation of these molecules decreased epithelial
marker E-cadherin, with concomitant increased mesenchymal marker
vimentin and N-cadherin expression in GC cells. Hence, it was
hypothesized that DAZAP, NOTCH1 and JAG1 coregulated GC cell
expression and motility via the EMT process.
However, the present study had limitations.
Firstly, the present study did not identify the potential
transcription factors of DAZAP1. Future studies should investigate
the transcription factors and other regulatory mechanisms of
DAZAP1. Secondly, the present study was primarily conducted in cell
lines and animal models, and further validation is required in
larger cohorts of clinical GC samples to confirm the clinical
relevance of the present findings.
In conclusion, DAZAP1 expression was significantly
increased in GC tissue and may be an indicator of poor prognosis
for patients. In addition to mediating the stability of NOTCH1
and/or JAG1 mRNA, which regulates EMT, DAZAP1 is key for GC cell
invasion and metastatic promotion. Therefore, the aforementioned
molecules provide not only a predictor for GC prognosis but also a
potential therapeutic target.
Supplementary Data
Availability of data and materials
The data generated in the present study may be
found in the National Center for Biotechnology Information database
under accession no SRX31515429. or at the following URL: ncbi.nlm.nih.gov/sra/PRJNA1378511.
Authors' contributions
JDW, JW, YFH, AML and SDL and JX conceived and
designed the study. SYP, YDC, JKW, XDH, LJH, YCX, TYL, XYW, PY and
JMZ performed experiments. QY and GNL collected tissue samples and
analyzed and interpreted data. JYL and WYD analyzed data. YFH, AML
and SDL supervised the study. SYP and JDW wrote the manuscript and
confirm the authenticity of all the raw data. All authors have read
and approved the final manuscript.
Ethics approval and consent to
participate
The animal experimental protocols were approved by
the Animal Ethics Committee of Nanfang Hospital, Southern Medical
University, Guangzhou, China (approval no. IACUC-LAC-20250111-002).
The use of tissue chips was approved by the Ethics Committee of
Shanghai Outdo Biotech Co. Ltd, Shanghai, China (approval no.
SHYJS-CP-1804015). Additionally, the research involving human
tissue received ethical approval from the Ethics Committee of
Nanfang Hospital, Guangzhou, China (approval no. NFEC-202105-K18).
Written informed consent was obtained from all participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
The present study was supported by the National Natural Science
Foundation of China (grant nos. 82273354, 82372955, 82073066,
82302891 and 82403543), Guangdong Basic and Applied Basic Research
Foundation (grant no. 2024A1515012891), Science and Technology
Program of Guangzhou (grant nos. 2024A04J5133 and (2025A04J3637)
and the Science and Technology Planning Project of Guangdong
Province (grant no. 2017B030314037).
References
|
1
|
Ferlay J, Colombet M, Soerjomataram I,
Parkin DM, Piñeros M, Znaor A and Bray F: Cancer statistics for the
year 2020: An overview. Int J Cancer. April 5–2021.Epub ahead of
print. View Article : Google Scholar
|
|
2
|
Wang SY, Yang XQ, Wang YX, Shen A, Liang
CC, Huang RJ, Cheng UH, Jian R, An N, Xiao YL, et al:
Overexpression of COX7A1 promotes the resistance of gastric cancer
to oxaliplatin and weakens the efficacy of immunotherapy. Lab
Invest. 104:1020902024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Chen Z, Chen Y, Sun Y, Tang L, Zhang L, Hu
Y, He M, Li Z, Cheng S, Yuan J, et al: Predicting gastric cancer
response to anti-HER2 therapy or anti-HER2 combined immunotherapy
based on multi-modal data. Signal Transduct Target Ther. 9:2222024.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Chen LY, Wang LW, Wen J, Cao JD, Zhou R,
Yang JL, Xiao Y, Su T, Huang Y, Guo Q, et al: RNA-binding protein
YBX3 promotes pparγ-SLC3A2 mediated BCAA metabolism fueling brown
adipogenesis and thermogenesis. Mol Metab. 90:1020532024.
View Article : Google Scholar
|
|
5
|
He S, Valkov E, Cheloufi S and Murn J: The
nexus between RNA-binding proteins and their effectors. Nat Rev
Genet. 24:276–294. 2023. View Article : Google Scholar :
|
|
6
|
Podszywalow-Bartnicka P and Neugebauer KM:
Multiple roles for AU-rich RNA binding proteins in the development
of haematologic malignancies and their resistance to chemotherapy.
RNA Biol. 21:1–17. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Li J, Pei M, Xiao W, Liu X, Hong L, Yu Z,
Peng Y, Zhang J, Yang P, Lin J, et al: The HOXD9-mediated
PAXIP1-AS1 regulates gastric cancer progression through PABPC1/PAK1
modulation. Cell Death Dis. 14:3412023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Choudhury R, Roy SG, Tsai YS, Tripathy A,
Graves LM and Wang Z: The splicing activator DAZAP1 integrates
splicing control into MEK/Erk-regulated cell proliferation and
migration. Nat Commun. 5:30782014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yang HT, Peggie M, Cohen P and Rousseau S:
DAZAP1 interacts via its RNA-recognition motifs with the C-termini
of other RNA-binding proteins. Biochem Biophys Res Commun.
380:705–709. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhang J, Ni Z, Zhang Y, Guo Y, Zhai R,
Wang M, Gong Z, Wang M, Zeng F, Gu Z, et al: Dazap1 phase
separation regulates mitochondrial metabolism to facilitate
invasion and metastasis of oral squamous cell carcinoma. Cancer
Res. 84:3818–3833. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sasaki K, Ono M, Takabe K, Suzuki A and
Kurihara Y: Specific intron-dependent loading of DAZAP1 onto the
cox6c transcript suppresses pre-mrna splicing efficacy and induces
cell growth retardation. Gene. 657:1–8. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang Q, Guo Y, Wang W, Liu B, Yang G, Xu
Z, Li J and Liu Z: RNA binding protein DAZAP1 promotes HCC
progression and regulates ferroptosis by interacting with SLC7A11
mRNA. Exp Cell Res. 399:1124532021. View Article : Google Scholar
|
|
13
|
Zhou Y, Huangfu S, Li M, Tang C, Qian J,
Guo M, Zhou Z, Yang Y and Gu C: DAZAP1 facilitates the alternative
splicing of KITLG to promote multiple myeloma cell proliferation
via ERK signaling pathway. Aging. 14:7972–7685. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Platonova N, Lesma E, Basile A, Bignotto
M, Garavelli S, Palano MT, Moschini A, Neri A, Colombo M and
Chiaramonte R: Targeting notch as a therapeutic approach for human
malignancies. Curr Pharm Des. 23:108–134. 2017. View Article : Google Scholar
|
|
15
|
Yeh TS, Wu CW, Hsu KW, Liao WJ, Yang MC,
Li AF, Wang AM, Kuo ML and Chi CW: The activated Notch1 signal
pathway is associated with gastric cancer progression through
cyclooxygenase-2. Cancer Res. 69:5039–5048. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ma H, Li N and Mo Z: Elevated Notch-1
expression promotes the lymph node metastasis of gastric cancer and
the Notch-1-PTEN-ERK1/2 signalling axis promotes the progression of
gastric cancer. Cytokine. 159:1560132022. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Feng P, Chen D, Wang X, Li Y, Li Z, Li B,
Zhang Y, Li W, Zhang J, Ye J, et al: Inhibition of the m(6)A reader
IGF2BP2 as a strategy against T-cell acute lymphoblastic leukemia.
Leukemia. 36:2180–2188. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Guo J, Zhang CD, An JX, Xiao YY, Shao S,
Zhou NM and Dai DQ: Expression of miR-634 in gastric carcinoma and
its effects on proliferation, migration, and invasion of gastric
cancer cells. Cancer Med. 7:776–787. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sugiyama M, Oki E, Nakaji Y, Tsutsumi S,
Ono N, Nakanishi R, Sugiyama M, Nakashima Y, Sonoda H, Ohgaki K, et
al: High expression of the Notch ligand Jagged-1 is associated with
poor prognosis after surgery for colorectal cancer. Cancer Sci.
107:1705–1716. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Qiao X, Ma B, Sun W, Zhang N, Liu Y, Jia L
and Liu C: JAG1 is associated with the prognosis and metastasis in
breast cancer. Sci Rep. 12:219862022. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sunshine HL, Cicchetto AC,
Kaczor-Urbanowicz KE, Ma F, Pi D, Symons C, Turner M, Shukla V,
Christofk HR, Vallim TA and Iruela-Arispe ML: Endothelial Jagged1
levels and distribution are post-transcriptionally controlled by
ZFP36 decay proteins. Cell Rep. 43:1136272024. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Gao Y, Cheng X and Han M: ZEB1-activated
Notch1 promotes circulating tumor cell migration and invasion in
lung squamous cell carcinoma. Clin Transl Oncol. 25:817–829. 2023.
View Article : Google Scholar
|
|
23
|
Shao S and Zhao X, Zhang X, Luo M, Zuo X,
Huang S, Wang Y, Gu S and Zhao X: Notch1 signaling regulates the
epithelial-mesenchymal transition and invasion of breast cancer in
a Slug-dependent manner. Mol Cancer. 1:282015. View Article : Google Scholar
|
|
24
|
Paz I, Kosti I, Ares M Jr, Cline M and
Mandel-Gutfreund Y: Rbpmap: A web server for mapping binding sites
of RNA-binding proteins. Nucleic Acids Res. 42:W361–W367. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Vasaikar SV, Deshmukh AP, den Hollander P,
Addanki S, Kuburich NA, Kudaravalli S, Joseph R, Chang JT,
Soundararajan R and Mani SA: Emtome: A resource for pan-cancer
analysis of epithelial-mesenchymal transition genes and signatures.
Br J Cancer. 124:259–269. 2021. View Article : Google Scholar :
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
27
|
Xie R, Wang J, Tang W, Li Y, Peng Y, Zhang
H, Liu G, Huang X, Zhao J, Li A, et al: Rufy3 promotes metastasis
through epithelial-mesenchymal transition in colorectal cancer.
Cancer Lett. 390:30–38. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Peng Y, Zhang P, Huang X, Yan Q, Wu M, Xie
R, Wu Y, Zhang M, Nan Q, Zhao J, et al: Direct regulation of FOXK1
by C-jun promotes proliferation, invasion and metastasis in gastric
cancer cells. Cell Death Dis. 7:e24802016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhang P, Tang WM, Zhang H, Li YQ, Peng Y,
Wang J, Liu GN, Huang XT, Zhao JJ, Li G, et al: MiR-646 inhibited
cell proliferation and EMT-induced metastasis by targeting FOXK1 in
gastric cancer. Br J Cancer. 117:525–534. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Liu H, Li Z, Zhang L, Zhang M, Liu S, Wang
J, Yang C, Peng Q, Du C and Jiang N: Necroptosis-related prognostic
model for pancreatic carcinoma reveals its invasion and metastasis
potential through hybrid EMT and immune escape. Biomedicines.
11:17382023. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Coelho-Rato LS, Parvanian S, Modi MK and
Eriksson JE: Vimentin at the core of wound healing. Trends Cell
Biol. 34:239–254. 2024. View Article : Google Scholar
|
|
32
|
Ridge KM, Eriksson JE, Pekny M and Goldman
RD: Roles of vimentin in health and disease. Genes Dev. 36:391–407.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Goenka A, Khan F, Verma B, Sinha P, Dmello
CC, Jogalekar MP, Gangadaran P and Ahn BC: Tumor microenvironment
signaling and therapeutics in cancer progression. Cancer Commun
(London). 43:525–561. 2023. View Article : Google Scholar
|
|
34
|
Teoh SL and Das S: Notch signalling
pathways and their importance in the treatment of cancers. Curr
Drug Targets. 19:128–143. 2018. View Article : Google Scholar
|
|
35
|
Hassan WA and Ito T: Identifying specific
Notch1 target proteins in lung carcinoma cells. Histol Histopathol.
36:69–76. 2021.
|
|
36
|
Deng JJ, Li GP, Lu W, Yan Z and Wang Y:
DAZAP1 overexpression promotes growth of HCC cell lines: A primary
study using CEUS. Clin Transl Oncol. 24:1168–1176. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wang X, Fan H, Ye X, Hu Y, Xiao Y, Zhang
M, Xu Y, Song J and Luo Y: RNA-binding protein DAZAP1 accelerates
the advancement of pancreatic cancer by inhibiting ferroptosis. Eur
J Med Res. 30:32025. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li F, Zhao H, Su M, Xie W, Fang Y, Du Y,
Yu Z, Hou L and Tan W: HnRNP-F regulates EMT in bladder cancer by
mediating the stabilization of Snail1 mRNA by binding to its 3'
UTR. EBioMedicine. 45:208–219. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Li F, Xie W, Fang Y, Xie K, Liu W, Hou L
and Tan W: HnRNP-F promotes the proliferation of bladder cancer
cells mediated by PI3K/AKT/FOXO1. J Cancer. 12:281–291. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Zhang P, Wang W, Xiang H, Zhou Y, Peng Q,
Liu G, Xu ZX and Lu L: DAZAP1 maintains gastric cancer stemness by
inducing mitophagy. JCI Insight. 10:e1754222025. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Zhou L, Yang Y, Ye Y, Qiao Q, Mi Y, Liu H,
Zheng Y, Wang Y, Liu M and Zhou Y: Notch1 signaling pathway
promotes growth and metastasis of gastric cancer via modulating
CDH5. Aging. 16:11893–11903. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Hodson DJ, Janas ML, Galloway A, Bell SE,
Andrews S, Li CM, Pannell R, Siebel CW, MacDonald HR, De
Keersmaecker K, et al: Deletion of the RNA-binding proteins ZFP36L1
and ZFP36L2 leads to perturbed thymic development and T
lymphoblastic leukemia. Nat Immunol. 11:717–724. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Petkau G, Mitchell TJ, Chakraborty K, Bell
SE, D Angeli V, Matheson L, Turner DJ, Saveliev A, Gizlenci O,
Salerno F, et al: The timing of differentiation and potency of CD8
effector function is set by RNA binding proteins. Nat Commun.
13:22742022. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Shi Q, Xue C, Zeng Y, Yuan X, Chu Q, Jiang
S, Wang J, Zhang Y, Zhu D and Li L: Notch signaling pathway in
cancer: From mechanistic insights to targeted therapies. Signal
Transduct Target Ther. 9:1282024. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Wang L, Cao G, Li W, Chen XW, Xiong SS, Mu
YN, Jiang JF, Yang L, Zhang DR and Cao YW: Expressions and
prognostic values of notch3 and DLL4 in human breast cancer.
Technol Cancer Res Treat. 22:153303382211189842023. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Zhao B, Hu S, Xiao Q, Fan S, Yu X, Li C,
Dong P and Zheng J: Expression of NOTCH receptors and ligands and
prognosis of hepatocellular carcinoma. Biomark Med. 14:1631–1639.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Su H, Wang X, Song J, Wang Y, Zhao Y and
Meng J: Microrna-539 inhibits the progression of Wilms' Tumor
through downregulation of JAG1 and Notch1/3. Cancer Biomark.
24:125–133. 2019. View Article : Google Scholar :
|