Elevated levels of ENO1 have been observed in
various types of cancer and are associated with a poor prognosis
(1). For example, elevated ENO1
expression has been noted in colorectal cancer (CRC) (2), hepatocellular carcinoma (HCC)
(3), gastric cancer (4), breast cancer (5), glioma (6), non-Hodgkin lymphoma (7,8),
bladder cancer (9) and head and
neck cancers (10).
Overexpression of ENO1 is typically linked to enhanced tumor
proliferation, invasion and metastatic potential (2,4,11-13). Notably, therapies targeting
enolases have shown significant efficacy, with several enolase
inhibitors demonstrating the ability to eliminate tumors (14).
Metabolic reprogramming in tumor cells, particularly
the 'Warburg effect’, is a central focus of cancer research
(15). This process not only
supplies bioenergy and biosynthetic precursors for rapidly dividing
tumor cells but also creates an immunosuppressive microenvironment
that fosters tumor progression (15-18). ENO1, a key catalyst in glycolysis
that converts 2-phosphoglycerate (2-PG) to phosphoenolpyruvate
(PEP), was traditionally considered to function solely in metabolic
pathways. However, emerging evidence suggests that the oncogenic
role of ENO1 extends far beyond its classical glycolytic activity.
As a multifunctional protein, ENO1 promotes extracellular matrix
degradation and tumor metastasis through its plasminogen (PLG)
receptor activity (19,20). It also regulates the translation
and stability of critical messenger RNAs, such as Yes-associated
protein 1 (YAP1), as a nucleic acid-binding protein (21). Furthermore, ENO1 acts as a
signaling scaffold protein, activating key oncogenic pathways like
phosphoinositide 3-kinase (PI3K)/protein kinase B (AKT), while
inhibiting AMP-activated protein kinase (AMPK)/mechanistic target
of rapamycin (mTOR) (22). The
diverse and precisely regulated functions of ENO1 are largely
attributed to its extensive post-translational modifications
(PTMs), including phosphorylation, ubiquitination, acetylation,
methylation and succinylation (23-27). These modifications serve as a
complex molecular code that dynamically governs ENO1's enzymatic
activity, protein stability, subcellular localization and
'functional switching’ across various biological processes.
Moreover, ENO1 plays a pivotal role in mediating chemotherapy
resistance and shaping the immunosuppressive tumor microenvironment
(TME), highlighting its immense potential as a therapeutic target
(9, 28). Currently, various intervention
strategies targeting ENO1, including small-molecule inhibitors,
natural product derivatives and therapeutic vaccines, have
exhibited significant antitumor effects in preclinical models
(29,30). The present review aimed to
systematically summarize the multifaceted oncogenic functions of
ENO1 in tumors and elucidate the multilevel regulatory network that
controls its expression and activity. Additionally, it discussed
the latest therapeutic strategies targeting ENO1 and their
prospects for clinical translation, while identifying key
scientific questions for future exploration in this field.
ENO1 was first identified as a pivotal enzyme in the
glycolytic pathway, catalyzing the conversion of 2-PG to PEP, a
crucial step in glycolysis. Elevated ENO1 expression supports the
'Warburg effect’, meeting the high metabolic demands of rapidly
proliferating tumors (31,32).
Traditionally, glycolytic enzymes were considered functionally
specialized and lacking regulatory signaling capabilities, with
stable expression levels. However, unlike GAPDH, studies have
revealed that ENO1 is not merely a housekeeping gene but a
multifunctional protein. It promotes tumor progression through
multiple mechanisms, including its glycolytic function and various
additional activities, with its expression closely linked to
malignant phenotypes, such as abnormal proliferation, invasion,
drug resistance and immune evasion in tumor cells. Notably, its
functions and involvement in pathophysiological processes are
largely determined by its subcellular localization (6,12,33).
The structural diversity of ENO1 underpins its
functional diversity. The human ENO1 gene spans over 18 kb and
contains 12 exons (34). Its
promoter lacks canonical TATA and CAAT boxes but is GC-rich and
contains potential SP1 binding sites (34), along with a hypoxia response
element (HRE) that mediates transcriptional activation by
hypoxia-inducible factor 1 (HIF-1) (35). An upstream inverted Alu sequence
may act as a transcriptional repressor (36). Through alternative splicing
regulated by the AKT/protein kinase R-like endoplasmic reticulum
kinase/eukaryotic initiation factor 2 α pathway, the ENO1 gene also
produces c-myc promoter-binding protein 1 (MBP-1), a shorter
protein variant with distinct functions (37). The crystal structure of human ENO1
has been resolved at 2.2 Å, revealing its classic dimeric form
(38). Each monomer consists of
434 amino acids (~48 kDa) and comprises two domains: an N-terminal
domain (residues 1-138) with a β-fold and three α-helices and a
larger C-terminal domain (residues 139-432) folding into an α/β
barrel. Mammals possess three tissue-specific enolase isoforms
(ENO1, ENO2 and ENO3) encoded by distinct genes, with ENO1 widely
distributed across tissues, ENO2 confined to neuron-associated
tissues and ENO3 predominantly found in muscle tissue. While
enolases are generally fluoride-sensitive and
Mg2+-dependent, ENO1 possesses distinct surface
properties that underlie its unique moonlighting functions,
including PLG receptor and nucleic acid-binding activities
(38).
ENO1's classic enzymatic function is the catalytic
conversion of 2-PGA to PEP. ENO1 operates as a homodimer and lacks
catalytic activity as a monomer (39). Its catalytic residues are highly
conserved across eukaryotes, with the active site located in a
cleft between the N-terminal and C-terminal domains, containing
both the substrate-binding pocket and the metal ion-binding site.
Catalytic activity requires a divalent metal ion (Mg2+
or Mn2+) to stabilize the substrate conformation and
neutralize the negative charge of the phosphate group, thereby
lowering the reaction energy barrier (40-43). The active site typically
accommodates two metal ions: a high-affinity conformational ion and
a low-affinity catalytic ion (44), with the latter binding only upon
substrate (or analog) engagement (45). The dehydration reaction catalyzed
by ENO1 occurs in three steps: First, the C-terminal domain of ENO1
binds and activates the substrate, 2-PGA (39). At this stage, the domain closes,
creating a hydrophobic environment that encapsulates 2-PGA. This
shields the substrate from water molecules, preventing interference
with the dehydration process. The closure of the domain is
triggered by Mg2+ binding (40,45,46). The second step involves
dehydration and proton transfer. A highly conserved lysine residue,
likely Lys345, acts as a base, abstracting a proton from the C2
position of 2-PGA. This leads to the formation of an unstable
carbocation intermediate, which rapidly isomerizes into an enol
pyruvate intermediate. This enol pyruvate intermediate is a pivotal
transient state in the catalytic cycle (40,47,48). Almost simultaneously, an acidic
residue, likely glutamate 211, donates a proton to the C3 hydroxyl
group of the enol intermediate, thereby completing the dehydration
process (48,49). Subsequently, a double bond forms
upon carbon rearrangement, yielding PEP, the hallmark biochemical
reaction of enolases. Substrates positioned at the ENO1 active site
can interact with two metal ions and the electrostatic interactions
between these ions are crucial for propelling the reaction forward
(40). Lys396 and Ser41 play a
role in stabilizing the negative charge of the transition state and
binding Mg2+, thereby facilitating reaction progression
(50). After the product is
released, ENO1 rapidly reverts to its initial conformation,
preparing to accept the next substrate molecule (20).
ENO1 functions as a cell-surface PLG receptor, its
earliest identified non-glycolytic role (19,51). This activity, independent of its
enzymatic function (38),
involves specific surface lysine residues and a putative binding
motif (FFRSGKY, residues 250-256) that interacts with PLG (Kd ~1.9
μM) (52). This
characteristic is shared with numerous PLG receptors (53) and its binding mode to PLG is
likely dominated by polar interactions arising from complementary
surface conformations (38,54). By concentrating PLG and
facilitating its activation by urokinase plasminogen activator
(uPA) or tissue plasminogenactivator (tPA), ENO1 enhances localized
plasmin generation, promoting extracellular matrix degradation and
tumor cell invasion/metastasis (52). Lys345 plays a vital role in
capturing the R-proton from PLG, with Glu211 becoming protonated
and forming a hydrogen bond with the α-hydroxy group of PLG
(40). Additionally, Lys345,
Glu211 and the metal cation of ENO1 may participate in PLG
activation (55).
ENO1 has been found to directly bind to RNA,
participating in the regulation of RNA metabolism and function. It
binds to the cytosine-uracil-guanine-rich element in YAP1 mRNA,
promoting YAP1 translation (56).
ENO1 also binds to the 3' untranslated regions (3'UTRs) of Klf2 and
FUS mRNA, thereby stabilizing them to inhibit pyroptosis (57). Conversely, ENO1 facilitates the
degradation of IRP1 mRNA to suppress ferroptosis (33). Furthermore, ENO1 functions as a
DNA-binding protein, inhibiting tumorigenicity by binding to the
c-Myc promoter, depending on the binding activity of residues
97-237. This DNA-binding domain is retained in the C-terminal
region shared with its splice variant MBP-1, explaining MBP-1's
antitumor activity (58).
The regulation of ENO1 enzyme activity involves
allosteric control, reversible covalent modifications and
adjustments in enzyme abundance. In practice, ENO1 function is
determined by both its concentration, regulated at the mRNA
transcription and protein stability levels (Fig. 1) and its specific activity (the
rate at which a unit concentration of enzyme catalyzes a specific
reaction), which is dynamically adjusted mainly through diverse
PTMs (Fig. 2).
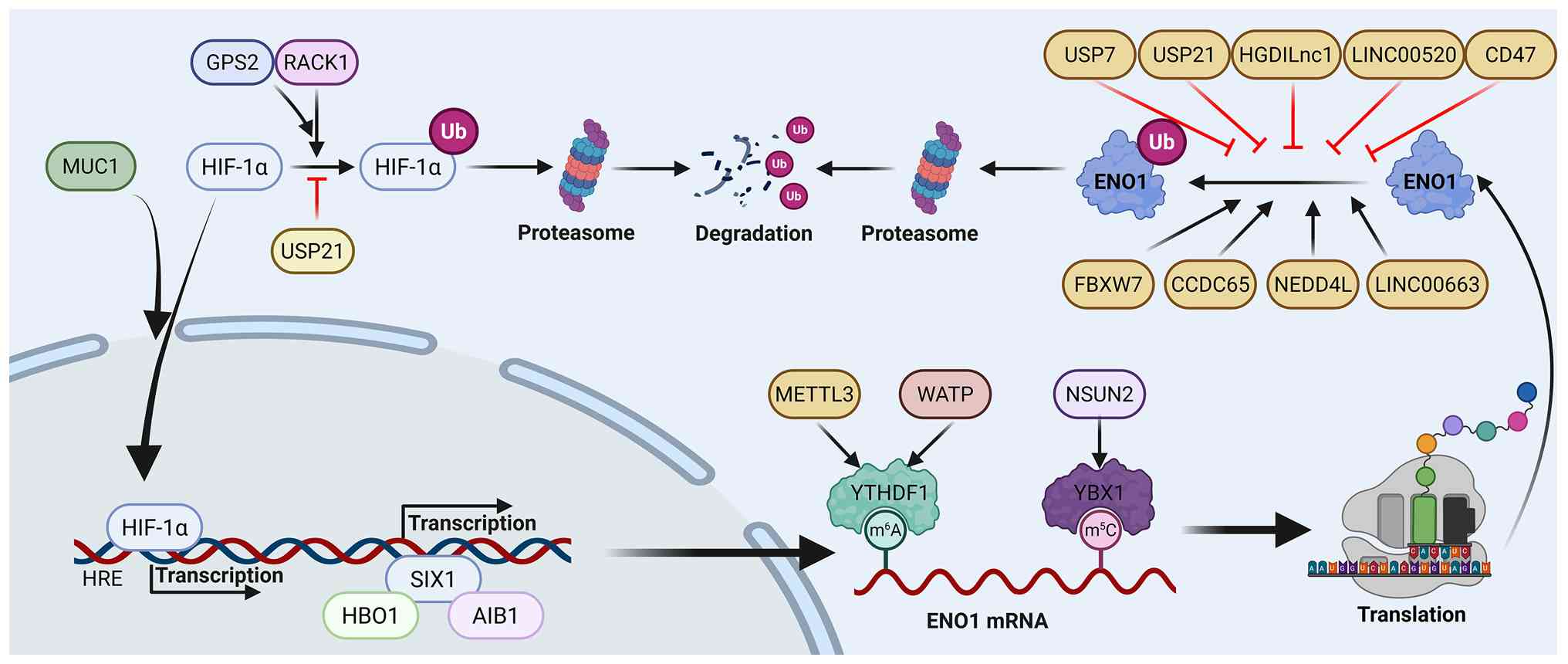
ENO1 mRNA expression is finely regulated through
multiple epigenetic mechanisms, including post-transcriptional
modifications such as 5-methylcytosine (m5C) and
N6-methyladenosine (m6A). The m5C
modification, catalyzed by methyltransferase NOP2/Sun RNA
methyltransferase 2 (NSUN2) and recognized by reader protein Y-box
binding protein 1 (YBX1), is crucial (59). Knockout of NSUN2 in CRC cells
markedly reduces ENO1 mRNA and protein levels, thereby altering
ENO1-dependent glucose metabolism pathways (59). This modification also influences
ENO1 expression via the transcription factor c-Myc (60), while coiled-coil domain containing
65 (CCDC65) suppresses transcription by impairing c-Myc binding to
the ENO1 promoter (61). Notably,
the accumulation of lactate catalyzed by ENO1 induces lactylation
at histone H3K18, activating NSUN2 transcription and forming a
positive feedback loop for m5C modification (59). Beyond m5C modification,
ENO1 mRNA undergoes m6A methylation at adenine position
359. The m6A modification promotes its binding to the
m6A-reading protein YTH N6-methyladenosine RNA binding
protein F1, thereby enhancing ENO1 translation efficiency (62). This process is regulated by the
RNA binding motif protein 15/methyltransferase-like 3 complex
(63) and Wilms tumor
1-associating protein (WTAP) (64), which further enhance ENO1 mRNA
m6A methylation to promote tumor glycolytic activity
(65). Additionally, lysine
methyltransferase 5A catalyzes mono-methylation of lysine 20 on
histone H4 (H4K20me1), which binds to the ENO1 promoter and
inhibits its transcriptional activity (66).
The presence of HREs in the ENO1 promoter enables
its transcriptional upregulation under hypoxia, making ENO1 a key
player in cellular hypoxic responses (35,67,68). Under hypoxic conditions, HIF-1α
stabilizes and translocates to the nucleus, binding to ENO1's HREs
to activate transcription (69,70). Hypoxia preferentially induces
full-length ENO1 over its alternative splice variant MBP-1,
attenuating MBP-1-mediated repression of the c-myc promoter and
leading to c-myc upregulation (71). The regulation of ENO1 expression
under hypoxia primarily relies on the HIF-1α signaling pathway,
with positive regulators such as Mucin 1 and histone H2A histone
family member X (72) stabilizing
HIF-1α to promote its recruitment to the ENO1 promoter (73,74). Conversely, G protein pathway
suppressor 2 binds to receptor for activated C kinase 1 (RACK1),
stabilizing the HIF-1α-RACK1 complex, which triggers HIF-1α
polyubiquitination and degradation, ultimately inhibiting ENO1
transcription (75). By contrast,
SIX homeobox 1 drives glycolytic gene expression independently of
HIF-1α by recruiting histone acetyltransferases histone
acetyltransferase binding to ORC1 and nuclear receptor coactivator
3 to the ENO1 promoter. These enzymes catalyze acetylation
modifications of histone H4K5 and H3K4 (H4K5ac and H3K4ac),
respectively, thereby activating ENO1 expression (76). These studies not only elucidate
the multi-level regulatory network governing ENO1 under hypoxic
conditions but also underscore its pivotal role in tumor metabolic
reprogramming as a key strategy for tumor cells to adapt to hypoxic
microenvironments.
Ubiquitination is a key post-translational
modification process that regulates protein stability and
degradation (77). Although
ubiquitination is a protein modification process, its regulation of
the ENO1 protein ultimately achieves changes in enzyme
concentration by affecting its stability. The E3 ubiquitin ligase
f-box/WD repeat-containing protein 7 (FBXW7) directly facilitates
ENO1 ubiquitination and degradation (61,78), a process finely modulated by
upstream regulators: CCDC65 (61)
recruits FBXW7 to amplify ENO1 ubiquitination, while the long
non-coding RNA LINC00520 binds to ENO1, competitively inhibiting
FBXW7-mediated ubiquitination and thus stabilizing ENO1 (79). Deubiquitination also plays a
pivotal role in ENO1 stability regulation. Ubiquitin specific
peptidase 21 (USP21) influences ENO1 through multiple pathways: On
the one hand, USP21 directly deubiquitinates ENO1 to enhance its
stability; on the other hand, USP21 indirectly upregulates HIF-1α
expression by deubiquitinating and stabilizing the heat shock
protein (HSP)90, thereby promoting ENO1 transcription (80). Additionally, EMC2 recruits
ubiquitin specific peptidase 7, which directly deubiquitinates ENO1
to stabilize it (24).
Beyond the classic ubiquitin-proteasome system,
other factors also target ENO1 for degradation regulation. The E3
ligase neural precursor cell expressed developmentally
downregulated 4-like has been shown to specifically target ENO1,
increasing its ubiquitination levels to promote degradation
(81,82). ENO1 stability is negatively
regulated by LINC00663, which enhances the E6AP-mediated
ubiquitin-proteasome pathway (83). Conversely, the long non-coding RNA
HGDILnc1 (84) and CD47 (85) interact with ENO1 to protect it
from proteasomal degradation. Furthermore, SUMOylation, the
covalent attachment of a small ubiquitin-like modifier (SUMO) to
target proteins (86), is
involved in ENO1 modification. Specifically, the K202 and K343
sites of ENO1 have been demonstrated to undergo SUMO2 and SUMO3
modification, indicating that SUMOylation may play a crucial role
in regulating ENO1 stability and cellular metabolic functions
(84). Collectively, these
multilayered regulatory mechanisms highlight the precise control of
ENO1 protein turnover in cancer cells.
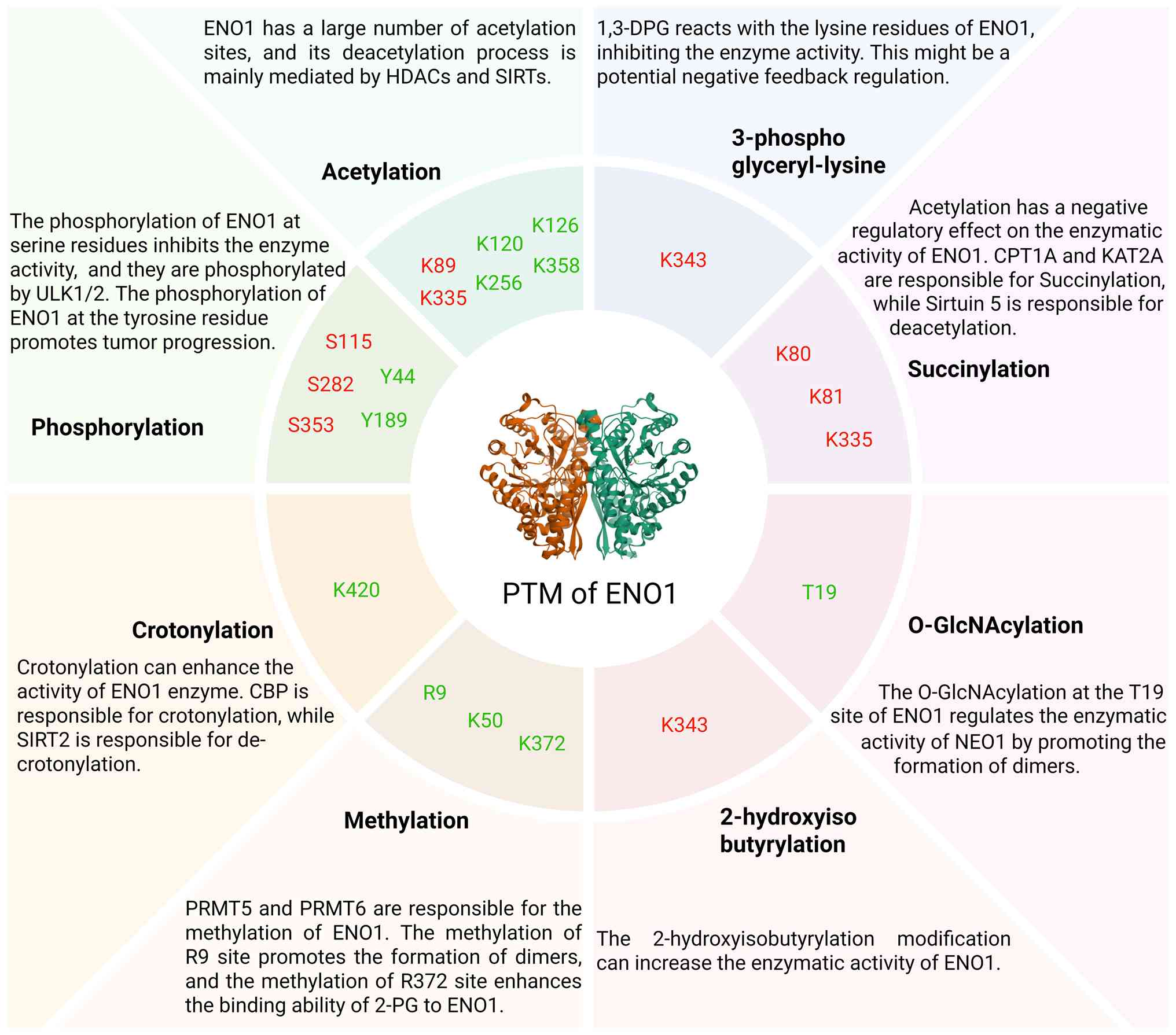
Phosphorylation is one of the most common PTMs of
ENO1, usually exerting a negative regulatory effect on its activity
by adding phosphate groups to specific serine, threonine, or
tyrosine residues (87). Under
conditions of amino acid and growth factor deprivation, Unc-51 like
autophagy activating kinase1/2 directly phosphorylate the Ser115
and Ser282 residues of ENO1. This phosphorylation reduces ENO1's
enzymatic activity and redirects increased carbon flux to the
pentose phosphate pathway to generate nicotinamide adenine
dinucleotide phosphate (reduced coenzyme II). This process helps
maintain cellular energy and redox homeostasis at both cellular and
organismal levels, protecting cells from reactive oxygen species
(ROS)-induced death (88).
Additionally, when exposed to the oxazole compound KB2764, ENO1 can
be phosphorylated at Ser353 by PKM, enhancing mitochondrial
function and reducing cellular reliance on glycolysis (89). High-throughput studies have
identified numerous phosphorylation sites on ENO1, including
multiple serine residues (e.g., S27, S37, S40, S161, S170, S179,
S254, S272, S198, S254, S263, S272, S291 and S419) (88-94) and three tyrosine residues (Y25,
Y44 and Y287) (93,95). However, the specific biological
functions of most of these sites, especially those located at dimer
interfaces, remain to be fully elucidated.
Acetylation typically occurs on lysine residues,
altering protein charge and conformation by adding an acetyl group,
which in turn affects protein function (87,96). This modification is widespread in
glycolysis and the tricarboxylic acid cycle, with traces found on
catalytic residues of ~2/3 of key enzymes (23). Moreover, acetylation modifications
of different glycolytic enzymes show interdependent effects,
suggesting the formation of regulatory networks within bioenergetic
pathways (97). Deacetylation of
ENO1 is mainly mediated by members of the histone deacetylase
(HDAC) family. For instance, histone deacetylase 11 (HDAC11)
inhibits ENO1's glycolytic activity in tumor cells by deacetylating
lysine 335 (K335) (98). Other
studies indicate that HDAC1 and HDAC7 also regulate ENO1's
acetylation status by controlling acetylation at residues K120,
K126 and K256, thereby enhancing enzyme activity (99,100). Furthermore, the deacetylase
sirtuin 4 targets the K358 site of ENO1. Its deacetylation action
increases ENO1's affinity for the substrate 2-PG, thereby boosting
catalytic activity. Notably, this enhanced enzymatic activity is
accompanied by weakened ENO1 binding to RNA, representing a
functional switch partly mediated by acetylation (101). Notably, ENO1 can bind ~2,000
transcript sites. Although the specific functions of these RNA
molecules are not fully understood, they directly inhibit ENO1's
enzymatic activity. Experiments show that exogenously introduced
RNA ligands reduce ENO1-dependent metabolite production while
promoting serine biosynthesis. This RNA-binding capacity is
regulated by acetylation at the K89 site, primarily controlled by
the deacetylase SIRT2. This finding reveals a novel mechanism by
which acetylation modifies metabolic enzyme activity through
nucleosomally regulated pathways (102). In a large-scale acetylation
study of central metabolic enzymes, K339, K390 and K402 of ENO1
were identified as acetylation sites. Although the direct effect of
these modifications on enzyme activity was not validated in that
study, structural analysis revealed that acetylated lysine residues
occupy similar spatial positions and volumes within the active
pocket as native substrates. Thus, it is hypothesized that the
presence of one or more acetyl groups may hinder PEP binding,
thereby inhibiting ENO1's catalytic activity (23).
Succinylation, which involves the covalent
attachment of a succinyl group to lysine residues, is a crucial PTM
regulating ENO1 (103).
Proteomic studies have identified ENO1 as one of the most heavily
succinylated proteins, with key modification sites including K80,
K81 and K335. Succinylation at these sites markedly inhibits ENO1's
glycolytic activity (104,105). Notably, the K335 site, located
near the substrate-binding pocket, may directly impede 2-PG binding
or alter the conformation of the catalytic center. Carnitine
palmitoyltransferase 1a (CPT1A) mediates succinylation at these
sites through its succinyl-CoA transferase (LSTase) function,
rather than its conventional carnitine palmitoyltransferase
function. Under glutamine-depleted conditions, CPT1A exhibits
enhanced LSTase activity, thereby promoting ENO1 succinylation
(105). Additionally, lysine
acetyltransferase 2a (KAT2A) has also been reported to act as a
succinyltransferase for ENO1, while the deacylase Sirtuin 5
mediates its removal (26,104,106).
Although the precise effect of KAT2A-mediated succinylation on
enzymatic kinetics requires further investigation, this
modification is associated with pro-tumor phenotypes, promoting
cell proliferation and migration while inhibiting apoptosis.
ENO1 methylation primarily occurs on arginine
residues and is catalyzed by specific methyltransferases, playing a
critical role in regulating its function. Protein arginine
methyltransferase 5 (PRMT5) mediates symmetric dimethylation at R9
and R50. Methylation at R9, located at the dimer interface, is
essential for stable dimer formation and full enzymatic activity
(25). The R50 modification is
associated with enhanced tumor cell metabolism and invasiveness
(107). Protein arginine
methyltransferase 6 (PRMT6) also methylates ENO1 at two key sites:
Methylation at R9 similarly promotes dimerization, while
methylation at R372 appears to enhance substrate (2-PG) binding
affinity. Therefore, arginine methylation by PRMT5 and PRMT6
fine-tunes ENO1 activity, stability and oncogenic function through
distinct molecular mechanisms (108).
O-GlcNAcylation is a dynamic monosaccharide
modification of intracellular proteins (109,110) that can influence protein
folding, stability, secretion and cell surface localization
(111). O-GlcNAcylation at
threonine 19 (T19) is a key regulatory modification of ENO1,
markedly enhancing its glycolytic function. Mechanistically, this
modification promotes the formation of enzymatically active dimers.
Evidence shows that a T19A mutation severely impairs dimerization,
resulting in a mutant protein with a higher Km for 2-PGA and a
catalytic efficiency reduced by 80% compared to the wild type.
Thus, T19 O-GlcNAcylation acts as a positive regulator of ENO1 by
stabilizing its active dimeric form (112).
Several additional PTMs further diversify the
functional regulation of ENO1. Lysine 2-hydroxyisobutyrylation at
K228 and K281 enhances its activity (113) and the deacylase CobB can reverse
this modification at the conserved K343 site (114). The glycolytic intermediate
1,3-bisphosphoglycerate can spontaneously form
3-phosphoglyceryl-lysine (pgK) at K343, inhibiting ENO1 activity
and constituting a direct metabolic feedback loop (115). This reaction occurs
spontaneously, exploiting the electrophilic nature of
1,3-bisphosphoglycerate without requiring enzymatic catalysis
(116). ENO1 is also modified by
interferon-stimulated gene 15, although the functional consequences
of this ISGylation event, first reported in 2005, remain undefined
(117). Lysine crotonylation,
predominantly at K420 and regulated by CBP/SIRT2, is elevated in
tumors and enhances both enzymatic and oncogenic functions
(118,119). Citrullination, the conversion of
arginine residues to citrulline catalyzed by protein arginine
deiminases (PADs) (120), has
been identified at least 18 citrullination sites in ENO1. Although
its effect on ENO1 activity is unclear, it serves as a diagnostic
marker in rheumatoid arthritis (121).
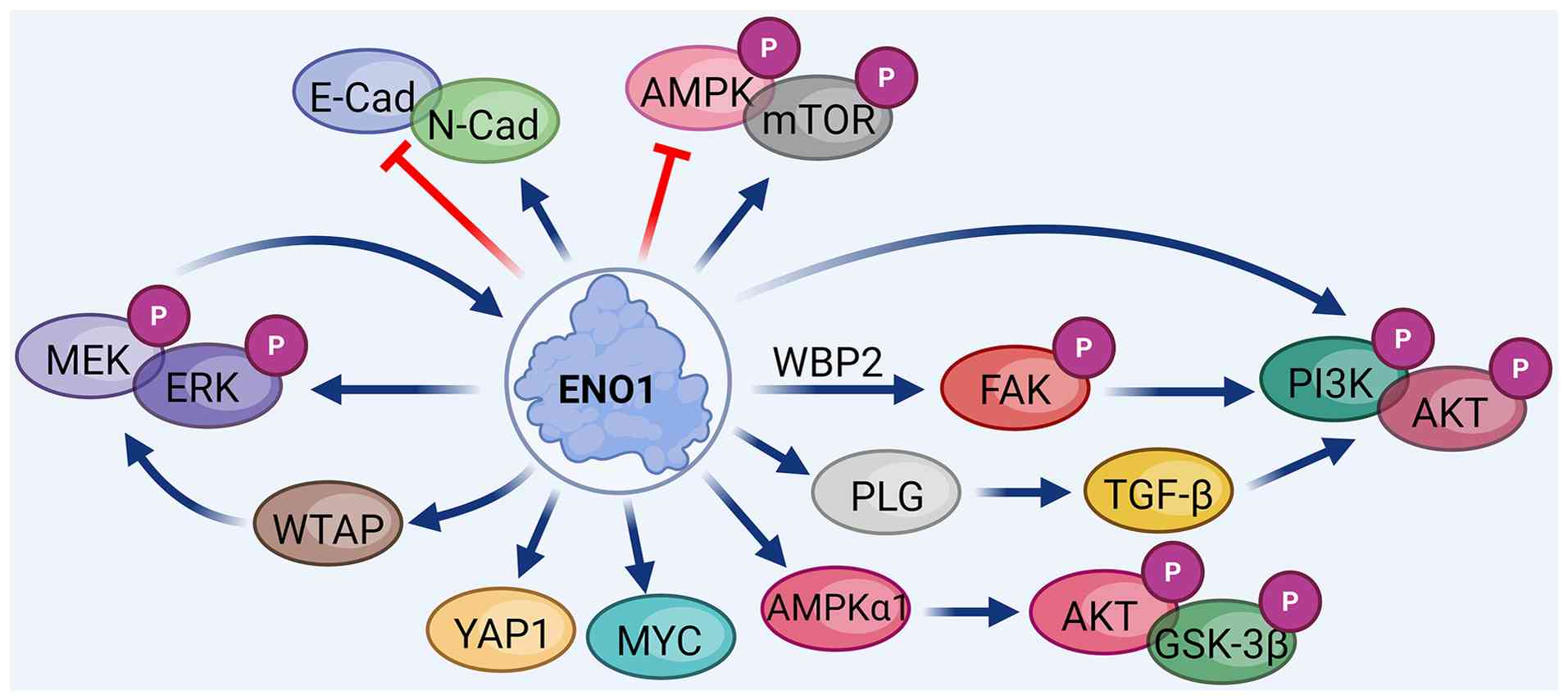
In tumorigenesis and progression, ENO1 not only
fuels abnormal metabolic reorganization in tumor cells but also
extensively contributes to malignant phenotypes such as
proliferation, invasion, metastasis and drug resistance by
regulating multiple signaling pathways. It activates the PI3K/AKT
pathway to promote survival signaling, suppresses the AMPK/mTOR
pathway to enhance anabolic metabolism, forms a positive feedback
loop with the extracellular signal-regulated kinase (ERK) signaling
pathway and regulates resistance mechanisms against various
chemotherapeutic agents, including gemcitabine, cisplatin and
5-fluorouracil. Thus, ENO1 serves as a central hub within tumor
signaling networks (Fig. 3).
In tumors, the PI3K/AKT pathway is often
constitutively activated due to genetic or epigenetic alterations
(122). This pathway drives
tumor cells toward aerobic glycolysis rather than mitochondrial
oxidation, providing metabolic advantages and promoting malignant
phenotypes (123-126). ENO1 facilitates the
phosphorylation of FAK at Tyr397, leading to increased levels of
phosphorylated (p-)PI3K (Tyr458) and p-AKT (Ser473), thereby
activating the PI3K/AKT signaling pathway (127). Additionally, ENO1 can be
phosphorylated at Y44, directly promoting the activation of PI3K
and AKT at the aforementioned sites (128). ENO1 also activates TGF-β1
through PLG recruitment and plasmin (PL) generation, thereby
activating the PI3K/AKT pathway (129). These mechanisms collectively
support ENO1's role in enhancing this signaling pathway.
Theoretically, any pathway regulating ENO1 protein levels and
stability could influence its downstream signaling pathways.
Reports confirm that circRPN2 and CCDC65 inhibit PI3K/AKT signaling
by binding to ENO1 and accelerating its degradation (130,131). Conversely, fibroblast growth
factor receptor-like 1, WW domain-binding protein 2, family with
sequence similarity 126 member A and transient receptor potential
cation channel subfamily C member 5 opposite strand positively
regulate ENO1 function and downstream signaling by directly binding
to the ENO1 protein, thereby promoting tumor proliferation and
chemotherapy resistance (123,132-134).
As a key glycolytic enzyme, ENO1 drives high
glycolytic flux and ATP production in cancer cells, suppressing
AMPK activation and subsequently enhancing mTOR signaling (135). Mechanistically, ENO1 inhibits
AMPK phosphorylation at Thr172 while promoting phosphorylation of
mTOR at Ser2447 and Akt at T308/S473, facilitating oncogenic growth
and metastasis in cancers such as CRC (29,30,136-138). Notably, ENO1 can also activate
AMPKα1 under certain conditions, contributing to cell proliferation
and apoptosis resistance through AKT/GSK3β phosphorylation
(136). Moreover, ENO1 can act
through the PI3K/Akt/mTOR signaling, a pathway independent of AMPK,
highlighting its multifaceted role in metabolic signaling (139,140). Silencing ENO1 induces
autophagy-dependent ferroptosis in breast cancer cells, closely
linked to AMPK/mTOR signaling, as mTOR is a well-known autophagy
inducer (141). Emerging
evidence suggests that the AMPK/mTOR pathway may reciprocally
regulate ENO1 expression and activity through transcriptional or
post-translational mechanisms, indicating bidirectional crosstalk
between ENO1 and this central metabolic hub (142). Further studies are needed to
fully elucidate the complex interplay between ENO1 and AMPK/mTOR
signaling in tumor metabolic rewiring.
The ERK signaling pathway, a critical member of the
mitogen-activated protein kinase (MAPK) signaling pathway, plays a
vital role in cellular processes such as proliferation,
differentiation, migration and apoptosis (32). Substantial evidence indicates that
ENO1 forms a positive feedback loop with the ERK signaling pathway.
In CRC, high ENO1 expression promotes ERK phosphorylation,
enhancing glycolysis and tumor growth (85). ENO1 may regulate ERK partly
through its PLG-activating function, as its knockdown disrupts
integrin-mediated cell-matrix adhesion, a process linked to ERK
activation (12). Activated
ERK1/2 signaling phosphorylates WTAP and enhances its stability,
promoting m6A modification of ENO1 mRNA, thereby
increasing its stability and translation efficiency (64) and supporting tumor cell
proliferation and survival (143). This positive feedback loop
sustains the tumor's proliferative signal. Targeting ENO1
(ENOblock) and targeting ERK (SCH772984) both demonstrate antitumor
effects, particularly in ENO1/ERK-hyperactive tumor subtypes,
indicating that the ERK signaling pathway and ENO1 jointly drive
tumor progression through multidimensional bidirectional
interactions.
ENO1 overexpression drives resistance to gemcitabine
in cancers such as prostate, pancreatic and cholangiocarcinoma
through multiple interconnected mechanisms. A central pathway
involves the post-transcriptional upregulation of YAP1. ENO1 binds
to C-U-G-rich elements in YAP1 mRNA, enhancing its translation.
Elevated YAP1 activates the Hippo signaling pathway and promotes
protective autophagy, thus shielding tumor cells from GTP-binding
protein overexpressed in skeletal muscle-induced apoptosis
(21). The oncogenic effects of
YAP1 are further amplified through the YAP1/phospholipase C beta
1/15-hydroxyprostaglandin dehydrogenase axis, stimulating
arachidonic acid metabolism and leading to prostaglandin E2
accumulation, a key driver of ENO1-mediated progression that can be
pharmacologically inhibited by aspirin (56). Additionally, ENO1 contributes to
GEM resistance by modulating cellular redox balance, reducing
intracellular ROS via glycolysis-enhanced 'Warburg’ metabolism
(144). ROS can reciprocally
regulate YAP1 stability through CBP-mediated phosphorylation at
Ser127, forming a regulatory circuit that sustains the resistant
phenotype (145). Targeting this
resistance pathway, modulation of ENO1 protein degradation has been
shown to reverse gemcitabine resistance in preclinical models
(80).
ENO1 enhances resistance to platinum-based
DNA-targeted drugs in multiple tumor types. The formation of ENO1
dimers markedly contributes to lactate levels, thereby mediating
cisplatin resistance (79,108).
Lactic acid accumulation promotes DNA homologous recombination
repair, a mechanism that is crucial for resistance to DNA-targeting
drugs, including cisplatin, temozolomide and doxorubicin (146,147). Lactate further activates
pro-survival signaling pathways such as Wnt and PI3K/Akt,
contributing to cisplatin tolerance (132,148,149). These observations highlight
ENO1-mediated glycolysis as a central metabolic determinant of
broad chemoresistance. Increased glycolytic activity driven by ENO1
also underlies resistance to nucleoside analogs such as
gemcitabine, often associated with MYC pathway activation and
upregulation of ribonucleotide reductase M1 (RRM1) (150). Separately, in CRC, ENO1 directly
induces epithelial-mesenchymal transition (EMT) and confers
resistance to 5-fluorouracil (5-FU) (151). Inhibition of ENO1 at Thr205
elevates CDH1 expression, reverses EMT and restores
chemosensitivity (148). This
strategy is particularly effective in TP53-mutated CRC; combined
targeting of the JAK2-STAT3-UCHL3-ENO1 axis with pacritinib
synergizes strongly with 5-FU, achieving >90% tumor reduction in
preclinical models (152).
ENO1 also contributes to therapy resistance through
additional pathways. In prostate cancer, cell surface-localized
ENO1 interacts with extracellular matrix protein 1, inducing its
phosphorylation at Y189. This modification facilitates the
recruitment of adaptor proteins growth factor receptor-bound
protein 2 and son of sevenless homolog 1, leading to downstream
MAPK signaling activation and promoting resistance to hormonal
therapies such as enzalutamide (153). Moreover, ENO1 plays a key role
in castration-resistant prostate cancer. The histone demethylase
lysine-specific demethylase 4B cooperates with c-Myc to bind the
c-Myc response element within the ENO1 promoter, driving ENO1
transcription and positioning it as a promising therapeutic target
in castration-resistant prostate cancer (154). These findings illustrate the
diverse mechanisms through which ENO1 fosters drug resistance.
Targeting ENO1 or its associated signaling networks represents a
compelling strategy for overcoming treatment resistance and
improving therapeutic outcomes.
Overexpression of ENO1 is strongly linked to the
upregulation of immune checkpoint molecules and resistance to
immunotherapy, suppressing antitumor immune responses across
diverse types of cancer through multiple mechanisms. It drives T
cell exhaustion and interacts with key immune molecules, ultimately
fostering an immunosuppressive microenvironment.
ENO1 is crucial in shaping an immunosuppressive
TME, mainly by inducing T cell dysfunction and exhaustion (155). Its overexpression is associated
with increased expression of inhibitory immune checkpoints such as
programmed cell death protein 1, cytotoxic T-lymphocyte-associated
protein 4 (CTLA-4) and T-cell immunoreceptor with Ig and ITIM
domains and contributes to resistance against anti-programmed
death-ligand 1 (PD-L1) therapy (156). The mechanisms of ENO1-mediated
immunosuppression differ among types of cancer. In pancreatic
cancer (PC), ENO1 knockout reduces regulatory T cells (Tregs) and
boosts IFN-γ and TNF-α production, transforming immunologically
'cold’ tumors into 'hot’ ones (157). It can also directly upregulate
PD-L1 to hinder CD8+ T cell infiltration (158). In intrahepatic
cholangiocarcinoma, ENO1 activates AKT signaling to elevate
fibrinogen-like protein 1, which binds to lymphocyte-activation
gene 3 on T cells and suppresses their function (159). In esophageal squamous cell
carcinoma, PLCE1-driven ENO1 upregulation enhances aerobic
glycolysis and lactate production, impairing CD8+ T cell
activation and cytokine secretion (160). Moreover, hydrogen sulfide
(H2S)-induced disulfide modification at ENO1 Cys119
inhibits Treg activation. Reducing H2S levels through a
sulfur-restricted diet or endogenous depletion synergizes with
anti-PD-L1/CTLA-4 therapy in CRC (161). In glioblastoma, ENO1 promotes
M2-type microglial polarization (162), while in multiple myeloma, it
dampens plasmacytoid dendritic cell (pDC) activity. This effect can
be reversed by ENO1 inhibitors, which restore pDC-mediated
activation of CD8+ T and NK cells (163). Collectively, these findings
underscore ENO1 as a master regulator of immune evasion and support
its dual targeting for direct antitumor effects and immunotherapy
enhancement.
ENO1 participates in immune regulation by
interacting with various immune-related molecules through direct
and indirect mechanisms, with context-dependent effects across
types of cancer. In CRC, O-GlcNAcylation of ENO1 at Ser249 disrupts
its interaction with PD-L1, reducing PD-L1's association with the
E3 ligase STIP1 homology and U-box containing protein 1. This
inhibits PD-L1's ubiquitin-mediated degradation, increases its
stability and ultimately facilitates immune evasion by suppressing
T-cell activity (112).
Conversely, in lung cancer, ENO1 interacts with PD-L1 and promotes
its degradation via the ubiquitin-proteasome pathway, enhancing T
cell-mediated antitumor immunity (27). In breast cancer, ENO1 contributes
to an immunosuppressive microenvironment by directly binding to and
stabilizing SPP1 mRNA, leading to elevated SPP1 expression.
Subsequently, SPP1 activates the ITGB1 pathway, impairing
CD8+ T cell function and promoting tumor-associated
macrophage polarization, thereby reducing the efficacy of
anti-PD-L1 therapy (164).
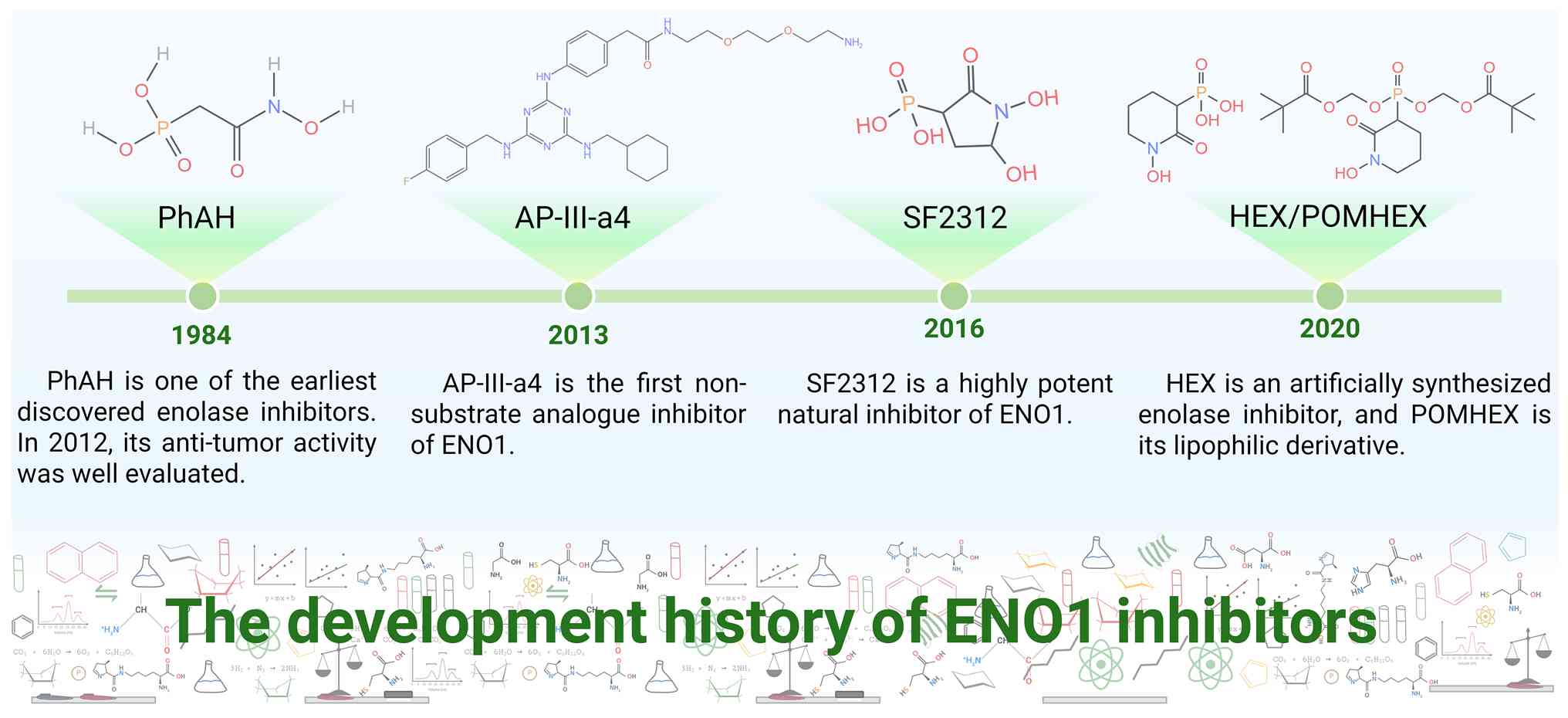
As a multifunctional oncogene, ENO1 has driven the
development of various targeted therapeutic strategies. Research on
ENO1 inhibitors has evolved from early natural products to a range
of novel compounds identified through high-throughput virtual
screening and rational drug design. Additionally, the exploration
of therapeutic vaccines has broadened the therapeutic options for
ENO1-related cancers (Fig.
4).
As early as the 1980s, numerous ENO1 inhibitors
were discovered. Most of these inhibitors either directly target
ENO1 to reduce its enzymatic activity or mimic metabolic
intermediates to inhibit binding (165). Fluoride, identified as an ENO1
inhibitor during this period, markedly inhibits ENO activity in
Streptococcus mutans at concentrations between 50 and 300
μM (Table I). It acts as a
competitive inhibitor of enolase (166). Another ENO1 inhibitor discovered
around the same period was phosphonoacetohydroxamate (PhAH)
(165), which was not thoroughly
evaluated until 2012. PhAH efficiently inhibits human enolases,
showing similar inhibitory effects on both ENO1 and ENO2 and
effectively suppresses glioblastoma proliferation and progression
(30,167).
2-Fluoro-2-phosphonoacetohydroxamate (FPAH), a fluorinated
derivative of the known inhibitor PhAH, has a lower pKa value for
its phosphate group due to fluorination, bringing it closer to that
of the natural substrate PEP. However, upon binding to ENO1, the
fluorine atom of FPAH forms a tight non-covalent interaction with
Gln164, forcing the nearby His156 to flip outward, resulting in a
relatively weak binding affinity (167). AP-III-a4 is the first
non-substrate analog inhibitor targeting ENO1 and also functions as
a non-enzymatic site inhibitor (30,168). It inhibits tumor survival and
migration, enhances the sensitivity of gastric cancer to cisplatin
and increases the sensitivity of breast cancer to radiotherapy
(169). In multiple myeloma,
AP-III-a4 restores T cell and NK cell-mediated tumor killing by
activating pDCs and synergistically enhances T cell-mediated cancer
cell lysis with anti-PD-L1 antibodies (163). Additionally, AP-III-a4 induces
the translocation of enolase into the cell nucleus, where it acts
as a transcriptional repressor (168,170). HEX, a synthetic enolase
inhibitor, selectively kills glioblastoma cells at low nanomolar
concentrations, eradicates intracranial orthotopic tumors in mice
and is well-tolerated. The carbonyl and hydroxylamine groups of HEX
form chelates with magnesium ions (Mg2+), while the
anionic phosphonate forms a salt bridge with the R373 residue,
thereby inhibiting ENO enzyme activity. Notably, HEX exhibits
higher selectivity toward ENO2, with Ki values of 232 and 64 nM for
ENO1 and ENO2, respectively. This makes HEX particularly suitable
for tumors lacking ENO1, enabling maximal inhibition of tumor cell
glycolysis while minimizing effects on normal cells (171). Due to its highly polar
phosphoric acid structure, HEX may have impaired plasma membrane
penetration. Consequently, the lipophilic POMHEX was developed,
which undergoes enzymatic hydrolysis inside cells to release the
active HEX, markedly improving its pharmacokinetic properties.
HuL227 is a multi-target monoclonal antibody that binds to ENO1 on
the tumor surface. It blocks ENO1's function as a PLG receptor,
reducing plasmin activation, inhibiting cancer cell degradation of
the extracellular matrix and impeding tumor invasion and
metastasis. Simultaneously, HuL227 inhibits VEGF-A-induced PLG
activation via ENO1 and markedly reduces vascular endothelial cell
luminal formation. Additionally, HuL227 suppresses the migration
and chemotaxis of androgen-independent prostate cancer cells (PC-3,
DU145) induced by inflammatory mediators (TNFα, CCL2, TGFβ)
(172). In recent years, Lung
et al (173) screened 22
million chemical structures from the ZINC database that comply with
Lipsky's five rules and identified compounds that bind to ENO1
through virtual screening. ZINC1304634, ZINC16124623, ZINC1702762
and ZINC72415103 are four ENO1 inhibitors identified through
comprehensive computer-aided screening. These inhibitors are
classified as non-mutagenic, non-carcinogenic and suitable for oral
administration, indicating development potential (174).
Several natural products and their derivatives have
demonstrated promising ENO1-targeting activities through distinct
mechanisms. The phosphonate antibiotic SF2312 binds to the ENO1
active site by coordinating two Mg2+ ions via its
phosphate and carbonyl groups, while its 5-hydroxyl group forms
hydrogen bonds with the catalytic residues Glu166 and His370. This
interaction locks ENO1 in a closed conformation, blocking substrate
entry and halting the catalytic cycle (29). Ciwujianoside E, a natural product,
specifically inhibits the interaction between ENO1 and PLG, thereby
preventing plasmin generation and TGF-β1 activation. In both in
vitro and in vivo studies on Burkitt's lymphoma,
Ciwujianoside E exhibited potent antitumor effects, suppressing
cell proliferation and invasion (129). Subsequent studies have revealed
that ginsenosides and their derivatives also exhibit inhibitory
effects on ENO1. Rh2E2 (175)
and 20 (S)-Rh2E2 (176), two
synthetic ginsenoside derivatives, inhibit tumor growth and
metastasis. Rh2E2 exhibits no toxic reactions at the maximum oral
dose of 5.000 mg/kg. It specifically downregulates tumor
glycolysis, fatty acid β-oxidation and the tricarboxylic acid
cycle, thereby inhibiting ATP production and targeting tumor cell
metabolism. 20 (S)-Rh2E2 reduces ENO1 protein levels, suppressing
lactate and ATP production in lung cancer cells without affecting
normal cells.
Tumor therapeutic vaccines are a form of
immunotherapy that activates the patient's immune system to
recognize and attack cancer cells. Their core principle involves
presenting tumor-specific antigens expressed by the tumor to the
immune system (177,178). The earliest ENO1-targeted tumor
therapeutic vaccine was a DNA vaccine. This ENO1 DNA vaccine
induced both antibody and cellular immune responses, extending the
average survival time of PC mice by 138 days. Vaccinated mice
showed a significant increase in serum levels of anti-ENO1
immunoglobulin G. This antibody binds to cancer cell surfaces,
inducing complement-dependent cell-mediated cytotoxicity. The ENO1
DNA vaccine reduced the number of myeloid-derived suppressor cells
(MDSCs) and Tregs while enhancing multiple responses of T helper
cells (179). Combining this
vaccine with a PI3Kγ inhibitor produced synergistic effects
(180). By inhibiting PI3Kγ to
target MDSCs, this approach increased CD8+ T cell and M1
macrophage infiltration in tumors while reducing Treg cells.
Specific IgG and IFNγ targeting ENO1 also increased in the mouse
circulation. In the meantime, pretreating PC mice with gemcitabine
before inoculation with the ENO1 DNA vaccine unleashed the
antitumor activity of CD4+ T cells, resulting in superior tumor
suppression (181). These
findings highlight the potential of ENO1-directed vaccines,
especially when combined with immunomodulatory or chemotherapeutic
agents, to overcome immunosuppression and enhance antitumor
immunity.
Designing antigens targeting post-translational
modification epitopes represents a novel strategy for activating
tumor immune responses (182).
Citrullinated ENO1 peptides, such as ENO1 11-25cit (15) in melanoma and ENO1 241-260cit
(253) in HLA-DR4 transgenic mice, elicit potent Th1 responses and
demonstrate antitumor activity not observed with their unmodified
counterparts (183). Building on
this, a refined citrullinated ENO1 (citENO1) vaccine peptide was
shown to enhance CD8+ T cell activation, inhibit tumor
growth and synergize with PD-1 blockade (184). Another vaccine design
incorporating the tumor-associated antigen mENO1 (Ag85B-ENO1 46-82)
effectively boosted CD8+ T cell infiltration and
cytokine production (IFN-γ, TNF-α), promoted M1-like macrophage
polarization and suppressed tumor progression in a lung cancer
model (185). Collectively,
these advances underscore the potential of ENO1-directed
therapeutic vaccines to reprogram the immunosuppressive TME and
enhance antitumor immunity.
ENO1 is notably overexpressed in various types of
cancer, such as HCC, PC, CRC and breast cancer. Its expression
levels are closely associated with tumor stage, invasiveness and a
poor prognosis. The diagnostic accuracy is substantially enhanced
when ENO1 is used alone or in combination with traditional
biomarkers such as CA19-9 and CEA.
The diagnostic value of ENO1 mainly stems from its
combined analysis with established biomarkers, which markedly
improves diagnostic sensitivity and specificity. In lung cancer
diagnosis, co-analyzing ENO1 with CEA, SCC, NSE and CYFRA21-1
effectively boosts detection sensitivity (186). In gastric cancer, elevated serum
anti-ENO1 autoantibody titers (AUC=0.656) have diagnostic utility
and their combination with CEA levels can further inform patient
prognosis (187). For PC
diagnosis, ENO1 alone shows a sensitivity of 75.8% and a
specificity of 88.2%. When combined with CA19-9, diagnostic
sensitivity reaches 94.5% with an AUC of 0.935, outperforming any
single currently available biomarker (188). This approach is particularly
valuable in Lewis-negative patients with normal CA19-9 levels.
Moreover, in oral submucosal fibrosis with atypical hyperplasia,
ENO1 expression levels can predict the malignant progression of
precancerous lesions (189). In
summary, ENO1 holds substantial diagnostic value.
ENO1 expression is markedly prognostic across
various types of cancer. In HCC, elevated ENO1 mRNA and protein
levels in tumor tissues are linked to poorer overall survival and
disease-free survival (3,190). Similarly, in CRC, ENO1
overexpression is strongly associated with advanced
clinicopathological features, including deeper tumor invasion,
lymph node metastasis, perineural invasion and a higher TNM stage,
serving as an indicator of an unfavorable prognosis (2,138). Elevated ENO1 protein levels are
detectable in the plasma of PC patients, with its expression
markedly associated with lymph node metastasis, clinical staging
and a poor prognosis (1,188). In triple-negative breast cancer,
ENO1 overexpression is markedly associated with high-grade tumors
and a poor prognosis (5).
Additionally, combined detection of CD47 and ENO1 provides a
reliable prognostic biomarker for CRC patients (85). Collectively, these findings
establish ENO1 as a valuable prognostic marker in multiple types of
cancer.
ENO1, as a multifunctional glycolytic enzyme, has
roles that extend far beyond its traditional function in energy
metabolism. The present review systematically clarified the
multifaceted functions of ENO1 in tumorigenesis and progression,
covering its diverse side roles as a metabolic enzyme, PLG
receptor, nucleic acid-binding protein and signaling scaffold
protein. The execution of these functions heavily relies on its
extensive PTMs, which dynamically regulate its enzymatic activity,
stability, subcellular localization and functional transitions.
ENO1 promotes tumor proliferation, invasion, metastasis and drug
resistance through mechanisms such as activating the PI3K/AKT
pathway, inhibiting the AMPK/mTOR pathway and forming positive
feedback loops with ERK. Furthermore, the regulatory role of ENO1
in the tumor immune microenvironment is gradually coming to light.
Therapeutically, small-molecule inhibitors, natural product
derivatives and therapeutic vaccines targeting ENO1 have
demonstrated significant antitumor potential in preclinical models,
especially when combined with immune checkpoint inhibitors or
chemotherapy, displaying favorable synergistic effects. Regarding
diagnosis and prognosis, ENO1 is highly expressed in multiple types
of cancer and is associated with tumor malignancy and poor patient
outcomes, making it a clinically valuable biomarker, either
independently or in combination with others.
Despite significant progress in understanding
ENO1's role in tumors, numerous unknowns remain to be explored.
First, although the present review summarized the extensive PTM
sites on ENO1, whether crosstalk exists between different
modifications is still unknown. Elucidating how PTM combinations
dynamically regulate ENO1's functional transitions and subcellular
localization is of great value, particularly its response
mechanisms within the dynamically changing TME. ENO1 plays critical
functions in normal cells and physiological states, yet its
functional regulation regarding tissue or tumor type specificity
remains poorly characterized. Simultaneously, clarifying its unique
mechanisms across diverse cancer contexts could facilitate the
development of more targeted therapeutic strategies. Existing ENO1
inhibitors mainly target enzymatic activity, with limited
approaches available for its non-enzymatic functions. Future
approaches may involve designing multifunctional inhibitors or
combination therapies that simultaneously block its metabolic and
signaling scaffold functions. As a novel topic, the specific
mechanisms by which ENO1 regulates immune cell function within the
TME require further elucidation. Investigating the metabolic-immune
crosstalk it mediates could offer new targets for overcoming
immunotherapy resistance. Future interdisciplinary, multi-level
investigations will advance ENO1 from mechanistic research to
clinical application, ultimately driving breakthroughs in cancer
treatment.
Not applicable.
XN completed the initial manuscript draft. MZ was
responsible for translation and language editing. KZ handled the
graphical visualizations. CW, JG and WF were responsible for
literature collection, organization and screening. LZ, TJ and GZ
provided financial support and ultimately reviewed the authenticity
of the article content and references. GZ was responsible for
writing guidance and topic selection. Data authentication is not
applicable. All authors read and approved the final manuscript.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
The authors utilized DeepL, an AI-powered
translation website, to assist in completing this article. While
the website provided support during the translation process, the
author bears full responsibility for the final content. Retrieved
from: https://www.deepl.com/zh/translator.
Not applicable.
National Natural Science Foundation of China Regional Innovation
Development Joint Fund Key Support Project (grant no. U23A20499),
National Natural Science Foundation of China Key Project (grant no.
82030119), Chunyuan Traditional Chinese Medicine Development
Special Fund Research Project for Achievement Transformation (grant
no. CY202302), National Natural Science Foundation of China Youth
Fund Project (grant no. 82204950), Zhejiang Provincial Natural
Science Foundation Youth Project (grant no. LQ23H270013), Zhejiang
Province Traditional Chinese Medicine Science and Technology
Project (grant no. 2023ZL292) and Zhejiang Province Medical and
Health Science and Technology Project (grant no. 2023KY617).
|
1
|
Almaguel FA, Sanchez TW, Ortiz-Hernandez
GL and Casiano CA: Alpha-enolase: Emerging tumor-associated
antigen, cancer biomarker, and oncotherapeutic target. Front Genet.
11:6147262021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cheng Z, Shao X, Xu M, Zhou C and Wang J:
ENO1 acts as a prognostic biomarker candidate and promotes tumor
growth and migration ability through the regulation of Rab1A in
colorectal cancer. Cancer Manag Res. 11:9969–9978. 2019. View Article : Google Scholar
|
|
3
|
Zhu W, Li H, Yu Y, Chen J, Chen X, Ren F,
Ren Z and Cui G: Enolase-1 serves as a biomarker of diagnosis and
prognosis in hepatocellular carcinoma patients. Cancer Manag Res.
10:5735–5745. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yang T, Shu X, Zhang HW, Sun LX, Yu L, Liu
J, Sun LC, Yang ZH and Ran YL: Enolase 1 regulates stem cell-like
properties in gastric cancer cells by stimulating glycolysis. Cell
Death Dis. 11:8702020. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Giannoudis A, Heath A and Sharma V: ENO1
as a biomarker of breast cancer progression and metastasis: A
bioinformatic approach using available databases. Breast Cancer
(Auckl). 18:117822342412856482024. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Song Y, Luo Q, Long H, Hu Z, Que T, Zhang
X, Li Z, Wang G, Yi L, Liu Z, et al: Alpha-enolase as a potential
cancer prognostic marker promotes cell growth, migration, and
invasion in glioma. Mol Cancer. 13:652014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhu M, Xiao Q, Cai X, Chen Z, Shi Q, Sun
X, Xie X and Sun M: Predicting lymphoma prognosis using machine
learning-based genes associated with lactylation. Transl Oncol.
49:1021022024. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhu X, Miao X, Wu Y, Li C, Guo Y, Liu Y,
Chen Y, Lu X, Wang Y and He S: ENO1 promotes tumor proliferation
and cell adhesion mediated drug resistance (CAM-DR) in
non-Hodgkin's lymphomas. Exp Cell Res. 335:216–223. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Su Z, You L, He Y, Chen J, Zhang G and Liu
Z: Multi-omics reveals the role of ENO1 in bladder cancer and
constructs an epithelial-related prognostic model to predict
prognosis and efficacy. Sci Rep. 14:21892024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Tsai ST, Chien IH, Shen WH, Kuo YZ, Jin
YT, Wong TY, Hsiao JR, Wang HP, Shih NY and Wu LW: ENO1, a
potential prognostic head and neck cancer marker, promotes
transformation partly via chemokine CCL20 induction. Eur J Cancer.
46:1712–1723. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li HJ, Ke FY, Lin CC, Lu MY, Kuo YH, Wang
YP, Liang KH, Lin SC, Chang YH, Chen HY, et al: ENO1 promotes lung
cancer metastasis via HGFR and WNT signaling-driven
epithelial-to-mesenchymal transition. Cancer Res. 81:4094–4109.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Principe M, Borgoni S, Cascione M,
Chattaragada MS, Ferri-Borgogno S, Capello M, Bulfamante S,
Chapelle J, Di Modugno F, Defilippi P, et al: Alpha-enolase (ENO1)
controls alpha v/beta 3 integrin expression and regulates
pancreatic cancer adhesion, invasion, and metastasis. J Hematol
Oncol. 10:162017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhang K, Tian R, Zhang W, Li Y, Zeng N,
Liang Y and Tang S: α-Enolase inhibits apoptosis and promotes cell
invasion and proliferation of skin cutaneous melanoma. Mol Biol
Rep. 49:8241–8250. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Crunkhorn S: Inhibiting enolase eradicates
tumours. Nat Rev Drug Discov. 20(20)2021.
|
|
15
|
Zhong X, He X, Wang Y, Hu Z, Huang H, Zhao
S, Wei P and Li D: Warburg effect in colorectal cancer: The
emerging roles in tumor microenvironment and therapeutic
implications. J Hematol Oncol. 15:1602022. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chu Z, Zhu M, Luo Y, Hu Y, Feng X, Wang H,
Sunagawa M and Liu Y: PTBP1 plays an important role in the
development of gastric cancer. Cancer Cell Int. 23:1952023.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Jiang H and Ye J: The Warburg effect: The
hacked mitochondrial-nuclear communication in cancer. Semin Cancer
Biol. 112:93–111. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hicks D and O'Regan RM: Improving outcomes
for high-risk hormone receptor-positive breast cancer with CDK
inhibition. J Clin Oncol. 40:1142–1146. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hsiao KC, Shih NY, Fang HL, Huang TS, Kuo
CC, Chu PY, Hung YM, Chou SW, Yang YY, Chang GC and Liu KJ: Surface
α-enolase promotes extracellular matrix degradation and tumor
metastasis and represents a new therapeutic target. PLoS One.
8:e693542013. View Article : Google Scholar
|
|
20
|
Li P and Hammes-Schiffer S:
Substrate-to-product conversion facilitates active site loop
opening in yeast enolase: A molecular dynamics study. ACS Catal.
9:8985–8990. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ma H, Kong L, Liu L, Du Y, Zhu X, Wang J
and Zhao W: ENO1 contributes to the gemcitabine resistance of
pancreatic cancer through the YAP1 signaling pathway. Mol Carcinog.
63:1221–1234. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Steinberg GR and Hardie DG: New insights
into activation and function of the AMPK. Nat Rev Mol Cell Bio.
24:255–272. 2023. View Article : Google Scholar
|
|
23
|
Nakayasu ES, Burnet MC, Walukiewicz HE,
Wilkins CS, Shukla AK, Brooks S, Plutz MJ, Lee BD, Schilling B,
Wolfe AJ, et al: Ancient regulatory role of lysine acetylation in
central metabolism. mBio. 8:e01894–17. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Xiao S, Jiang S, Wen C, Wang H, Nie W,
Zhao J and Zhang B: EMC2 promotes breast cancer progression and
enhances sensitivity to PDK1/AKT inhibition by deubiquitinating
ENO1. Int J Biol Sci. 21:2629–2646. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Xie F, Zhang H, Zhu K, Jiang CS, Zhang X,
Chang H, Qiao Y, Sun M, Wang J, Wang M, et al: PRMT5 promotes
ovarian cancer growth through enhancing Warburg effect by
methylating ENO1. MedComm (2020). 4:e2452023. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zeng K and Yin H: KAT2A changes the
function of endometrial stromal cells via regulating the
succinylation of ENO1. Open Life Sci. 19:202207852024. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhang C, Zhang K, Gu J and Ge D: ENO1
promotes antitumor immunity by destabilizing PD-L1 in NSCLC. Cell
Mol Immunol. 18:2045–2047. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Tian Y, Guo J, Mao L, Chen Z, Zhang X, Li
Y, Zhang Y, Zha X and Luo OJ: Single-cell dissection reveals
promotive role of ENO1 in leukemia stem cell self-renewal and
chemoresistance in acute myeloid leukemia. Stem Cell Res Ther.
15:3472024. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Leonard PG, Satani N, Maxwell D, Lin YH,
Hammoudi N, Peng Z, Pisaneschi F, Link TM, Lee GR IV, Sun D, et al:
SF2312 is a natural phosphonate inhibitor of enolase. Nat Chem
Biol. 12:1053–1058. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Muller FL, Colla S, Aquilanti E, Manzo VE,
Genovese G, Lee J, Eisenson D, Narurkar R, Deng P, Nezi L, et al:
Passenger deletions generate therapeutic vulnerabilities in cancer.
Nature. 488:337–342. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Song Q, Zhang K, Sun T, Xu C, Zhao W and
Zhang Z: Knockout of ENO1 leads to metabolism reprogramming and
tumor retardation in pancreatic cancer. Front Oncol.
13:11198862023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Yuan J, Dong X, Yap J and Hu J: The MAPK
and AMPK signalings: Interplay and implication in targeted cancer
therapy. J Hematol Oncol. 13:1132020. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang T, Sun L, Hao Y, Suo C, Shen S, Wei
H, Ma W, Zhang P, Wang T, Gu X, et al: ENO1 suppresses cancer cell
ferroptosis by degrading the mRNA of iron regulatory protein 1. Nat
Cancer. 3:75–89. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Giallongo A, Oliva D, Calì L, Barba G,
Barbieri G and Feo S: Structure of the human gene for
alpha-enolase. Eur J Biochem. 190:567–573. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Semenza GL, Jiang BH, Leung SW, Passantino
R, Concordet JP, Maire P and Giallongo A: Hypoxia response elements
in the aldolase A, enolase 1, and lactate dehydrogenase A gene
promoters contain essential binding sites for hypoxia-inducible
factor 1. J Biol Chem. 271:32529–32537. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Oliva D, Calì L, Feo S and Giallongo A:
Complete structure of the human gene encoding neuron-specific
enolase. Genomics. 10:157–165. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Maranto C, Perconti G, Contino F, Rubino
P, Feo S and Giallongo A: Cellular stress induces cap-independent
alpha-enolase/MBP-1 translation. FEBS Lett. 589:2110–2116. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Kang HJ, Jung SK, Kim SJ and Chung SJ:
Structure of human alpha-enolase (hENO1), a multifunctional
glycolytic enzyme. Acta Crystallogr D Biol Crystallogr. 64:651–657.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Schulz EC, Tietzel M, Tovy A, Ankri S and
Ficner R: Structure analysis of Entamoeba histolytica enolase. Acta
Crystallogr D Biol Crystallogr. 67:619–627. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Liu H, Zhang Y and Yang W: How is the
active site of enolase organized to catalyze two different reaction
steps? J Am Chem Soc. 122:6560–6570. 2000. View Article : Google Scholar
|
|
41
|
Nagatani RA, Gonzalez A, Shoichet BK,
Brinen LS and Babbitt PC: Stability for function trade-offs in the
enolase superfamily 'catalytic module’. Biochemistry. 46:6688–6695.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Nagatani RJA and Babbitt PC: Investigating
the properties of catalytic module residues in the enolase
superfamily: Effects on thermostability. FASEB J. 20:A9012006.
View Article : Google Scholar
|
|
43
|
Morrow JR, Amyes TL and Richard JP:
Phosphate binding energy and catalysis by small and large
molecules. Accounts Chem Res. 41:539–548. 2008. View Article : Google Scholar
|
|
44
|
Poyner RR, Cleland WW and Reed GH: Role of
metal ions in catalysis by enolase: An ordered kinetic mechanism
for a single substrate enzyme. Biochemistry. 40:8009–8017. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang E, Hatada M, Brewer JM and Lebioda
L: Catalytic metal ion binding in enolase: The crystal structure of
an enolase-Mn2+-phosphonoacetohydroxamate complex at 2.4-A
resolution. Biochemistry. 33:6295–6300. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Lebioda L and Stec B: Mechanism of
enolase: The crystal structure of
enolase-Mg2(+)-2-phosphoglycerate/phosphoenolpyruvate complex at
2.2-A resolution. Biochemistry. 30:2817–2822. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Reed GH, Poyner RR, Larsen TM, Wedekind JE
and Rayment I: Structural and mechanistic studies of enolase. Curr
Opin Struc Biol. 6:736–743. 1996. View Article : Google Scholar
|
|
48
|
Sims PA, Menefee AL, Larsen TM,
Mansoorabadi SO and Reed GH: Structure and catalytic properties of
an engineered heterodimer of enolase composed of one active and one
inactive subunit. J Mol Biol. 355:422–431. 2006. View Article : Google Scholar
|
|
49
|
Schreier B and Höcker B: Engineering the
enolase magnesium II binding site: Implications for its evolution.
Biochemistry. 49:7582–7589. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Krucinska J, Falcone E, Erlandsen H,
Hazeen A, Lombardo MN, Estrada A, Robinson VL, Anderson AC and
Wright DL: Structural and functional studies of bacterial enolase,
a potential target against gram-negative pathogens. Biochemistry.
58:1188–1197. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Principe M, Ceruti P, Shih NY,
Chattaragada MS, Rolla S, Conti L, Bestagno M, Zentilin L, Yang SH,
Migliorini P, et al: Targeting of surface alpha-enolase inhibits
the invasiveness of pancreatic cancer cells. Oncotarget.
6:11098–11113. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Andronicos NM, Ranson M, Bognacki J and
Baker MS: The human ENO1 gene product (recombinant human
alpha-enolase) displays characteristics required for a plasminogen
binding protein. Biochim Biophys Acta. 1337:27–39. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
El Otmani H, Frunt R, Smits S, Barendrecht
AD, de Maat S, Fijnheer R, Lenting PJ and Tersteeg C:
Plasmin-cleaved von Willebrand factor as a biomarker for
microvascular thrombosis. Blood. 143:2089–2098. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Bergmann S, Wild D, Diekmann O, Frank R,
Bracht D, Chhatwal GS and Hammerschmidt S: Identification of a
novel plasmin(ogen)-binding motif in surface displayed
alpha-enolase of Streptococcus pneumoniae. Mol Microbiol.
49:411–423. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Ji H, Wang J, Guo J, Li Y, Lian S, Guo W,
Yang H, Kong F, Zhen L, Guo L and Liu Y: Progress in the biological
function of alpha-enolase. Anim Nutr. 2:12–17. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Sun L, Suo C, Zhang T, Shen S, Gu X, Qiu
S, Zhang P, Wei H, Ma W, Yan R, et al: ENO1 promotes liver
carcinogenesis through YAP1-dependent arachidonic acid metabolism.
Nat Chem Biol. 19:1492–1503. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zhao Y, He X, Yang X, Hong Z, Xu Y, Xu J,
Zheng H, Zhang L, Zuo Z and Hu X: CircFndc3b mediates
exercise-induced neuroprotection by mitigating
microglial/macrophage pyroptosis via the ENO1/KLF2 axis in stroke
mice. Adv Sci. 12:e24038182025. View Article : Google Scholar
|
|
58
|
Miller DM, Thomas SD, Islam A, Muench D
and Sedoris K: c-Myc and cancer metabolism. Clin Cancer Res.
18:5546–5553. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Chen B, Deng Y, Hong Y, Fan L, Zhai X, Hu
H, Yin S, Chen Q, Xie X, Ren X, et al: Metabolic recoding of
NSUN2-Mediated m5C modification promotes the progression
of colorectal cancer via the NSUN2/YBX1/m5C-ENO1
positive feedback loop. Adv Sci (Weinh). 11:e23098402024.
View Article : Google Scholar
|
|
60
|
Zhang H, Zhai X, Liu Y, Xia Z, Xia T, Du
G, Zhou H, Franziska Strohmer D, Bazhin AV, Li Z, et al:
NOP2-mediated m5C modification of c-Myc in an EIF3A-dependent
manner to reprogram glucose metabolism and promote hepatocellular
carcinoma progression. Research (Wash D C). 6:01842023.PubMed/NCBI
|
|
61
|
Zhang Z, Xu P, Hu Z, Fu Z, Deng T, Deng X,
Peng L, Xie Y, Long L, Zheng D, et al: CCDC65, a gene knockout that
leads to early death of mice, acts as a potentially novel tumor
suppressor in lung adenocarcinoma. Int J Biol Sci. 18:4171–4186.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Ma L, Xue X, Zhang X, Yu K, Xu X, Tian X,
Miao Y, Meng F, Liu X, Guo S, et al: The essential roles of
m6A RNA modification to stimulate ENO1-dependent
glycolysis and tumorigenesis in lung adenocarcinoma. J Exp Clin
Canc Res. 41:362022. View Article : Google Scholar
|
|
63
|
Shen C, Liu J, Xie F, Yu Y, Ma X, Hu D,
Liu C and Wang Y: N6-methyladenosine enhances the translation of
ENO1 to promote the progression of bladder cancer by inhibiting
PCNA ubiquitination. Cancer Lett. 595:2170022024. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Ou B, Liu Y, Yang X, Xu X, Yan Y and Zhang
J: C5aR1-positive neutrophils promote breast cancer glycolysis
through WTAP-dependent m6A methylation of ENO1. Cell Death Dis.
12:7372021. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Liu XS, Zhou LM, Yuan LL, Gao Y, Kui XY,
Liu XY and Pei ZJ: NPM1 is a prognostic biomarker involved in
immune infiltration of lung adenocarcinoma and associated with m6A
modification and glycolysis. Front Immunol. 12:7247412021.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Lu L, Li X, Zhong Z, Zhou W, Zhou D, Zhu M
and Miao C: KMT5A downregulation participated in high
glucose-mediated EndMT via upregulation of ENO1 expression in
diabetic nephropathy. Int J Biol Sci. 17:4093–4107. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Liu Y, Liao L, An C, Wang X, Li Z, Xu Z,
Liu J and Liu S: α-Enolase lies downstream of mTOR/HIF1α and
promotes thyroid carcinoma progression by regulating CST1. Front
Cell Dev Biol. 9:6700192021. View Article : Google Scholar
|
|
68
|
Sun H, Mo J, Cheng R, Li F, Li Y, Guo Y,
Li Y, Zhang Y, Bai X, Wang Y, et al: ENO1 expression and Erk
phosphorylation in PDAC and their effects on tumor cell apoptosis
in a hypoxic microenvironment. Cancer Biol Med. 19:1598–1616. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Beckner ME, Pollack IF and Hamilton RL:
Abstract 3940: Transformation by ENO1 highlights the positive
relationship between HIF1A's and VEGFA's RNA expression levels,
putatively by counteracting heterogeneity in glioblastomas. Cancer
Res. 77(13 Suppl): S39402017. View Article : Google Scholar
|
|
70
|
Zheng F, Jang WC, Fung FKC, Lo ACY and
Wong IYH: Up-regulation of ENO1 by HIF-1α in retinal pigment
epithelial cells after hypoxic challenge is not involved in the
regulation of VEGF secretion. PLoS One. 11:e01479612016. View Article : Google Scholar
|
|
71
|
Sedoris KC, Thomas SD and Miller DM:
Hypoxia induces differential translation of enolase/MBP-1. BMC
Cancer. 10:1572010. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Zhang S, Wang YS, Li Y, To KI, Zhang ET
and Jin YH: Annexin A2 binds the 3'-UTR of H2AX mRNA and regulates
histone-H2AX-derived hypoxia-inducible factor 1-alpha activation.
Cell Signal. 132:1117812025. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Chaika NV, Gebregiworgis T, Lewallen ME,
Purohit V, Radhakrishnan P, Liu X, Zhang B, Mehla K, Brown RB,
Caffrey T, et al: MUC1 mucin stabilizes and activates
hypoxia-inducible factor 1 alpha to regulate metabolism in
pancreatic cancer. Proc Natl Acad Sci USA. 109:13787–13792. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Gong W, Ekmu B, Wang X, Lu Y and Wan L:
AGR2-induced glucose metabolism facilitated the progression of
endometrial carcinoma via enhancing the MUC1/HIF-1α pathway. Hum
Cell. 33:790–800. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Si Y, Ou H, Jin X, Gu M, Sheng S, Peng W,
Yang D, Zhan X, Zhang L, Yu Q, et al: G protein pathway suppressor
2 suppresses aerobic glycolysis through RACK1-mediated HIF-1α
degradation in breast cancer. Free Radical Bio Med. 222:478–492.
2024. View Article : Google Scholar
|
|
76
|
Li L, Liang Y, Kang L, Liu Y, Gao S, Chen
S, Li Y, You W, Dong Q, Hong T, et al: Transcriptional regulation
of the Warburg effect in cancer by SIX1. Cancer Cell. 33368–385.
(e7)2018. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Arkinson C, Dong KC, Gee CL and Martin A:
Mechanisms and regulation of substrate degradation by the 26S
proteasome. Nat Rev Mol Cell Bio. 26:104–122. 2025. View Article : Google Scholar
|
|
78
|
Zhan P, Wang Y, Zhao S, Liu C, Wang Y, Wen
M, Mao JH, Wei G and Zhang P: FBXW7 negatively regulates ENO1
expression and function in colorectal cancer. Lab Invest.
95:995–1004. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Wei X, Feng J, Chen L, Zhang C, Liu Y,
Zhang Y, Xu Y, Zhang J, Wang J, Yang H, et al: METTL3-mediated m6A
modification of LINC00520 confers glycolysis and chemoresistance in
osteosarcoma via suppressing ubiquitination of ENO1. Cancer Lett.
2171942024.Epub ahead of print. PubMed/NCBI
|
|
80
|
Xu X, Chen Y, Shao S, Wang J, Shan J, Wang
Y, Wang Y, Chang J, Zhou T, Chen R, et al: USP21 deubiquitinates
and stabilizes HSP90 and ENO1 to promote aerobic glycolysis and
proliferation in cholangiocarcinoma. Int J Biol Sci. 20:1492–1508.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Zhang G, Zhao X and Liu W: NEDD4L inhibits
glycolysis and proliferation of cancer cells in oral squamous cell
carcinoma by inducing ENO1 ubiquitination and degradation. Cancer
Biol Ther. 23:243–253. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Zhang M, Zhang Z, Tian X, Zhang E, Wang Y,
Tang J and Zhao J: NEDD4L in human tumors: Regulatory mechanisms
and dual effects on anti-tumor and pro-tumor. Front Pharmacol.
14:12917732023. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Ma J, Zhu J, Li J, Liu J, Kang X and Yu J:
Enhanced E6AP-mediated ubiquitination of ENO1 via LINC00663
contributes to radiosensitivity of breast cancer by regulating
mitochondrial homeostasis. Cancer Lett. 560:2161182023. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Zhang QW, Lin XL, Dai ZH, Zhao R, Hou YC,
Liang Q, Zhang Y and Ge ZZ: Hypoxia and low-glucose environments
co-induced HGDILnc1 promote glycolysis and angiogenesis. Cell Death
Discov. 10:1322024. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Hu T, Liu H, Liang Z, Wang F, Zhou C,
Zheng X, Zhang Y, Song Y, Hu J, He X, et al: Tumor-intrinsic CD47
signal regulates glycolysis and promotes colorectal cancer cell
growth and metastasis. Theranostics. 10:4056–4072. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Tharuka MDN, Courelli AS and Chen Y:
Immune regulation by the SUMO family. Nat Rev Immunol. 25:608–620.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Pienkowski T, Kowalczyk T, Cysewski D,
Kretowski A and Ciborowski M: Glioma and post-translational
modifications: A complex relationship. Biochim Biophys Acta Rev
Cancer. 1878:1890092023. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Li TY, Sun Y, Liang Y, Liu Q, Shi Y, Zhang
CS, Zhang C, Song L, Zhang P, Zhang X, et al: ULK1/2 constitute a
bifurcate node controlling glucose metabolic fluxes in addition to
autophagy. Mol Cell. 62:359–370. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Lee YH, Lim H, Kim G, Jang G, Kuk MU, Park
JH, Yoon JH, Lee YJ, Kim D, So B, et al: Elucidating the role and
mechanism of alpha-enolase in senescent amelioration via metabolic
reprogramming. Cell Prolif. 58:e700492025. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Bian Y, Song C, Cheng K, Dong M, Wang F,
Huang J, Sun D, Wang L, Ye M and Zou H: An enzyme assisted RP-RPLC
approach for in-depth analysis of human liver phosphoproteome. J
Proteomics. 96:253–262. 2014. View Article : Google Scholar
|
|
91
|
Dephoure N, Zhou C, Villén J, Beausoleil
SA, Bakalarski CE, Elledge SJ and Gygi SP: A quantitative atlas of
mitotic phosphorylation. Proc Natl Acad Sci USA. 105:10762–10767.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Qiao G, Wu A, Chen X, Tian Y and Lin X:
Enolase 1, a moonlighting protein, as a potential target for cancer
treatment. Int J Biol Sci. 17:3981–3992. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Ross SH, Rollings C, Anderson KE, Hawkins
PT, Stephens LR and Cantrell DA: Phosphoproteomic analyses of
interleukin 2 signaling reveal integrated JAK kinase-dependent and
-independent networks in CD8(+) T cells. Immunity. 45:685–700.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Zhou H, Di Palma S, Preisinger C, Peng M,
Polat AN, Heck AJ and Mohammed S: Toward a comprehensive
characterization of a human cancer cell phosphoproteome. J Proteome
Res. 12:260–271. 2013. View Article : Google Scholar
|
|
95
|
Rush J, Moritz A, Lee KA, Guo A, Goss VL,
Spek EJ, Zhang H, Zha XM, Polakiewicz RD and Comb MJ:
Immunoaffinity profiling of tyrosine phosphorylation in cancer
cells. Nat Biotechnol. 23:94–101. 2005. View Article : Google Scholar
|
|
96
|
Zhou MM and Cole PA: Targeting lysine
acetylation readers and writers. Nat Rev Drug Discov. 24:112–133.
2025. View Article : Google Scholar
|
|
97
|
Zhu D, Hou L, Hu B, Zhao H, Sun J, Wang J
and Meng X: Crosstalk among proteome, acetylome and succinylome in
colon cancer HCT116 cell treated with sodium dichloroacetate. Sci
Rep. 6:374782016. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Sharma V, Christodoulidou A, Yue L,
Alontaga A, Goodheart W, Hesterberg R, Zheng X, Zheng MW, Lee JY,
Burnette P and Wright KL: Abstract LB-249: HDAC11 regulates lysine
acetylation of enolase 1. Cancer Res. 78(13 Suppl): LB–249. 2018.
View Article : Google Scholar
|
|
99
|
Arito M, Nagai K, Ooka S, Sato T, Takakuwa
Y, Kurokawa MS, Sase T, Okamoto K, Suematsu N and Kato T: Altered
acetylation of proteins in patients with rheumatoid arthritis
revealed by acetyl-proteomics. Clin Exp Rheumatol. 33:877–886.
2015.PubMed/NCBI
|
|
100
|
Zhang S, Wu G, Shi L, Ding P, Liu G, Chang
W, Dai Y, Han X and Ma X: ARID1A deficiency promotes malignant
proliferation of hepatocellular carcinoma by activating HDAC7/ENO1
signaling pathway. Hepatol Commun. 9:e07382025. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Lv M, Yang X, Xu C, Song Q, Zhao H, Sun T,
Liu J, Zhang Y, Sun G, Xue Y and Zhang Z: SIRT4 promotes pancreatic
cancer stemness by enhancing histone lactylation and epigenetic
reprogramming stimulated by calcium signaling. Adv Sci (Weinh).
12:e24125532025. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Huppertz I, Perez-Perri JI, Mantas P,
Sekaran T, Schwarzl T, Russo F, Ferring-Appel D, Koskova Z,
Dimitrova-Paternoga L, Kafkia E, et al: Riboregulation of Enolase 1
activity controls glycolysis and embryonic stem cell
differentiation. Mol Cell. 82:2666–2680. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Yao W, Hu X and Wang X: Crossing
epigenetic frontiers: the intersection of novel histone
modifications and diseases. Signal Transduct Target Ther.
9:2322024. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Du J, Zhou Y, Su X, Yu JJ, Khan S, Jiang
H, Kim J, Woo J, Kim JH, Choi BH, et al: Sirt5 is a NAD-dependent
protein lysine demalonylase and desuccinylase. Science.
334:806–809. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Kurmi K, Hitosugi S, Wiese EK,
Boakye-Agyeman F, Gonsalves WI, Lou Z, Karnitz LM, Goetz MP and
Hitosugi T: Carnitine palmitoyltransferase 1A has a lysine
succinyltransferase activity. Cell Rep. 22:1365–1373. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Guo Z, Hui Y, Sun S and Kong F: KAT3B
promotes the glycolysis and malignant progression of lung cancer by
mediating the succinylation modification of PKM2. J Biochem Mol
Toxicol. 39:e702592025. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Zakrzewicz D, Didiasova M, Krüger M,
Giaimo BD, Borggrefe T, Mieth M, Hocke AC, Zakrzewicz A, Schaefer
L, Preissner KT and Wygrecka M: Protein arginine methyltransferase
5 mediates enolase-1 cell surface trafficking in human lung
adenocarcinoma cells. Biochim Biophys Acta Mol Basis Dis.
1864:1816–1827. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Sun M, Li L, Niu Y, Wang Y, Yan Q, Xie F,
Qiao Y, Song J, Sun H, Li Z, et al: PRMT6 promotes tumorigenicity
and cisplatin response of lung cancer through triggering 6PGD/ENO1
mediated cell metabolism. Acta Pharm Sin B. 13:157–173. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Shi Q, Shen Q, Liu Y, Shi Y, Huang W, Wang
X, Li Z, Chai Y, Wang H, Hu X, et al: Increased glucose metabolism
in TAMs fuels O-GlcNAcylation of lysosomal Cathepsin B to promote
cancer metastasis and chemoresistance. Cancer Cell.
40:1207–1222.e10. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Xu X, Peng Q, Jiang X, Tan S, Yang W, Han
Y, Oyang L, Lin J, Shen M, Wang J, et al: Altered glycosylation in
cancer: Molecular functions and therapeutic potential. Cancer
Commun (Lond). 44:1316–1336. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Chung IC, Huang WC, Huang YT, Chen ML,
Tsai AW, Wu PY and Yuan TT: Unrevealed roles of extracellular
enolase-1 (ENO1) in promoting glycolysis and pro-cancer activities
in multiple myeloma via hypoxia-inducible factor 1α. Oncol Rep.
50:2052023. View Article : Google Scholar
|
|
112
|
Zhu Q, Li J, Sun H, Fan Z, Hu J, Chai S,
Lin B, Wu L, Qin W, Wang Y, et al: O-GlcNAcylation of enolase 1
serves as a dual regulator of aerobic glycolysis and immune evasion
in colorectal cancer. Proc Natl Acad Sci USA. 121:e24083541212024.
View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Huang H, Tang S, Ji M, Tang Z, Shimada M,
Liu X, Qi S, Locasale JW, Roeder RG, Zhao Y and Li X: p300-mediated
lysine 2-hydroxyisobutyrylation regulates glycolysis. Mol Cell.
70663–678. (e6)2018. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Dong H, Zhai G, Chen C, Bai X, Tian S, Hu
D, Fan E and Zhang K: Protein lysine de-2-hydroxyisobutyrylation by
CobB in prokaryotes. Sci Adv. 5:eaaw67032019. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Moellering RE and Cravatt BF: Functional
lysine modification by an intrinsically reactive primary glycolytic
metabolite. Science. 341:549–553. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Baker SA and Rutter J: Metabolites as
signalling molecules. Nat Rev Mol Cell Bio. 24:355–374. 2023.
View Article : Google Scholar
|
|
117
|
Giannakopoulos NV, Luo JK, Papov V, Zou W,
Lenschow DJ, Jacobs BS, Borden EC, Li J, Virgin HW and Zhang DE:
Proteomic identification of proteins conjugated to ISG15 in mouse
and human cells. Biochem Biophys Res Commun. 336:496–506. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Hou JY, Cao J, Gao LJ, Zhang FP, Shen J,
Zhou L, Shi JY, Feng YL, Yan Z, Wang DP and Cao JM: Upregulation of
α enolase (ENO1) crotonylation in colorectal cancer and its
promoting effect on cancer cell metastasis. Biochem Biophys Res
Commun. 578:77–83. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Su W, Kang K, Li X and Huang H: A
prognostic lysine crotonylation signature shapes the immune
microenvironment in hepatocellular carcinoma. Curr Med Chem.
Jun;13:2025.Epub ahead of print.
|
|
120
|
Zhu D, Zhang Y and Wang S: Histone
citrullination: A new target for tumors. Mol Cancer. 20:902021.
View Article : Google Scholar : PubMed/NCBI
|
|
121
|
He Y, Sareila O, Johansson L, Agelii ML,
Cheng L, Lundquist A, Lönnblom E, Gröndal G, Gudbjornsson B,
Hørslev-Petersen K, et al: Epitopes targeted by autoantibodies in
presymptomatic individuals predict early rheumatoid arthritis. Ann
Rheum Dis. 84:1090–1103. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Hao C, Wei Y, Meng W, Zhang J and Yang X:
PI3K/AKT/mTOR inhibitors for hormone receptor-positive advanced
breast cancer. Cancer Treat Rev. 132:1028612025. View Article : Google Scholar
|
|
123
|
Li Y, Li Y, Luo J, Fu X, Liu P, Liu S and
Pan Y: FAM126A interacted with ENO1 mediates proliferation and
metastasis in pancreatic cancer via PI3K/AKT signaling pathway.
Cell Death Discov. 8:2482022. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Luo Y, Guo C, Ling C, Yu W, Chen Y, Jiang
L, Luo Q, Wang C and Xu W: Pine pollen reverses the function of
hepatocellular carcinoma by inhibiting α-Enolase mediated PI3K/AKT
signaling pathway. PLoS One. 19:e03124342024. View Article : Google Scholar
|
|
125
|
Ma J, Zhang M, Mulati S, Nabi X and Zhang
W: Esculetin inhibits the PI3K/Akt/mTOR pathway and enhances
anti-colorectal cancer activity via binding to ENO1. Front
Pharmacol. 16:16271142025. View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Wu X, Ding C, Liu Y, Dong K and Zhang H:
B7-H3 promotes proliferation and migration of lung cancer cells by
modulating PI3K/AKT pathway via ENO1 activity. Transl Cancer Res.
13:833–846. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Fu QF, Liu Y, Fan Y, Hua SN, Qu HY, Dong
SW, Li RL, Zhao MY, Zhen Y, Yu XL, et al: Alpha-enolase promotes
cell glycolysis, growth, migration, and invasion in non-small cell
lung cancer through FAK-mediated PI3K/AKT pathway. J Hematol Oncol.
8:222015. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Feng T, Feng N, Zhu T, Li Q, Zhang Q, Wang
Y, Gao M, Zhou B, Yu H, Zheng M and Qian B: A SNP-mediated lncRNA
(LOC146880) and microRNA (miR-539-5p) interaction and its potential
impact on the NSCLC risk. J Exp Clin Canc Res. 39:1572020.
View Article : Google Scholar
|
|
129
|
Wang H, Zhang S, Kui X, Ren J, Zhang X,
Gao W, Zhang Y, Liu H, Yan J, Sun M, et al: Ciwujianoside E
inhibits Burkitt lymphoma cell proliferation and invasion by
blocking ENO1-plasminogen interaction and TGF-β1 activation. Biomed
Pharmacother. 177:1169702024. View Article : Google Scholar
|
|
130
|
Deng T, Shen P, Li A, Zhang Z, Yang H,
Deng X, Peng X, Hu Z, Tang Z, Liu J, et al: CCDC65 as a new
potential tumor suppressor induced by metformin inhibits activation
of AKT1 via ubiquitination of ENO1 in gastric cancer. Theranostics.
11:8112–8128. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Li J, Hu ZQ, Yu SY, Mao L, Zhou ZJ, Wang
PC, Gong Y, Su S, Zhou J, Fan J, et al: CircRPN2 inhibits aerobic
glycolysis and metastasis in hepatocellular carcinoma. Cancer Res.
82:1055–1069. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Chen R, Li D, Zheng M, Chen B, Wei T, Wang
Y, Li M, Huang W, Tong Q, Wang Q, et al: FGFRL1 affects
chemoresistance of small-cell lung cancer by modulating the
PI3K/Akt pathway via ENO1. J Cell Mol Med. 24:2123–2134. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Chen S, Zhang Y, Wang H, Zeng YY, Li Z, Li
ML, Li FF, You J, Zhang ZM and Tzeng CM: WW domain-binding protein
2 acts as an oncogene by modulating the activity of the glycolytic
enzyme ENO1 in glioma. Cell Death Dis. 9:3472018. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Cui Y, Peng J, Zheng M, Ge H, Wu X, Xia Y,
Huang Y, Wang S, Yin Y, Fu Z and Xie H: TRPC5OS induces
tumorigenesis by increasing ENO1-mediated glucose uptake in breast
cancer. Transl Oncol. 22:1014472022. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Shu X, Cao KY, Liu HQ, Yu L, Sun LX, Yang
ZH, Wu CA and Ran YL: Alpha-enolase (ENO1), identified as an
antigen to monoclonal antibody 12C7, promotes the self-renewal and
malignant phenotype of lung cancer stem cells by AMPK/mTOR pathway.
Stem Cell Res Ther. 12:1192021. View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Dai J, Zhou Q, Chen J, Rexius-Hall ML,
Rehman J and Zhou G: Alpha-enolase regulates the malignant
phenotype of pulmonary artery smooth muscle cells via the AMPK-Akt
pathway. Nat Commun. 9:38502018. View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Wang G, Yu Y, Wang YZ, Yin PH, Xu K and
Zhang H: The effects and mechanisms of isoliquiritigenin loaded
nanoliposomes regulated AMPK/mTOR mediated glycolysis in colorectal
cancer. Artif Cells Nanomed Biotechnol. 48:1231–1249. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Zhan P, Zhao S, Yan H, Yin C, Xiao Y, Wang
Y, Ni R, Chen W, Wei G and Zhang P: α-enolase promotes
tumorigenesis and metastasis via regulating AMPK/mTOR pathway in
colorectal cancer. Mol Carcinog. 56:1427–1437. 2017. View Article : Google Scholar
|
|
139
|
Shi Y, Liu J, Zhang R, Zhang M, Cui H,
Wang L, Cui Y, Wang W, Sun Y and Wang C: Targeting endothelial ENO1
(alpha-enolase)-PI3K-Akt-mTOR axis alleviates hypoxic pulmonary
hypertension. Hypertension. 80:1035–1047. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
140
|
Udawant S, Litif C, Lopez A, Gunn B,
Schuenzel E and Keniry M: PI3K pathway inhibition with NVP-BEZ235
hinders glycolytic metabolism in glioblastoma multiforme cells.
Cells. 10:30652021. View Article : Google Scholar : PubMed/NCBI
|
|
141
|
Huang G, Lu L, You Y, Li J and Zhang K:
Knockdown of ENO1 promotes autophagy dependent-ferroptosis and
suppresses glycolysis in breast cancer cells via the regulation of
CST1. Drug Develop Res. 85:e700042024. View Article : Google Scholar
|
|
142
|
Sukumaran A, Choi K and Dasgupta B:
Insight on transcriptional regulation of the energy sensing AMPK
and biosynthetic mTOR pathway genes. Front Cell Dev Biol.
8:6712020. View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Chen S, Duan G, Zhang R and Fan Q:
Helicobacter pylori cytotoxin-associated gene A protein upregulates
α-enolase expression via Src/MEK/ERK pathway: Implication for
progression of gastric cancer. Int J Oncol. 45:764–770. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
144
|
Wang L, Bi R, Yin H, Liu H and Li L: ENO1
silencing impaires hypoxia-induced gemcitabine chemoresistance
associated with redox modulation in pancreatic cancer cells. Am J
Transl Res. 11:4470–4480. 2019.PubMed/NCBI
|
|
145
|
Jin J, Zhang L, Li X, Xu W, Yang S, Song
J, Zhang W, Zhan J, Luo J and Zhang H: Oxidative stress-CBP axis
modulates MOB1 acetylation and activates the Hippo signaling
pathway. Nucleic Acids Res. 50:3817–3834. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
146
|
Chen H, Li Y, Li H, Chen X, Fu H, Mao D,
Chen W, Lan L, Wang C, Hu K, et al: NBS1 lactylation is required
for efficient DNA repair and chemotherapy resistance. Nature.
631:663–669. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
147
|
Chen Y, Wu J, Zhai L, Zhang T, Yin H, Gao
H, Zhao F, Wang Z, Yang X, Jin M, et al: Metabolic regulation of
homologous recombination repair by MRE11 lactylation. Cell.
187:294–311.e21. 2024. View Article : Google Scholar
|
|
148
|
Chen T, Liu G, Chen S, Zhang F, Ma S, Bai
Y, Zhang Q and Ding Y: Natural product mediated
mesenchymal-epithelial remodeling by covalently binding ENO1 to
degrade m6A modified β-catenin mRNA. Acta Pharm Sin B. 15:467–483.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
149
|
Mohapatra P, Shriwas O, Mohanty S, Ghosh
A, Smita S, Kaushik SR, Arya R, Rath R, Das Majumdar SK, Muduly DK,
et al: CMTM6 drives cisplatin resistance by regulating Wnt
signaling through the ENO-1/AKT/GSK3β axis. JCI Insight.
6:e1436432021.
|
|
150
|
Lu Y, Ma H, Xiong X, Du Y, Liu L, Wang J
and Zhao W: Deletion of ENO1 sensitizes pancreatic cancer cells to
gemcitabine via MYC/RRM1-mediated glycolysis. Sci Rep. 15:99412025.
View Article : Google Scholar : PubMed/NCBI
|
|
151
|
Gu J, Zhong K, Wang L, Ni H, Zhao Y, Wang
X, Yao Y, Jiang L, Wang B and Zhu X: ENO1 contributes to
5-fluorouracil resistance in colorectal cancer cells via EMT
pathway. Front Oncol. 12:10130352022. View Article : Google Scholar
|
|
152
|
Xin H, Zhao Z, Guo S, Tian R, Ma L, Yang
Y, Zhao L, Wang G, Li B, Hu X, et al: Targeting the
JAK2-STAT3-UCHL3-ENO1 axis suppresses glycolysis and enhances the
sensitivity to 5-FU chemotherapy in TP53-mutant colorectal cancer.
Acta Pharm Sin B. 15:2529–2544. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
153
|
Wang X, Wang M, Lin Q, He L, Zhang B, Chen
X, Chen G, Du H, Lang C, Peng X and Dai Y: Osteoblast-derived ECM1
promotes anti-androgen resistance in bone metastatic prostate
cancer. Adv Sci (Weinh). 12:e24076622025. View Article : Google Scholar :
|
|
154
|
Wu MJ, Chen CJ, Lin TY, Liu YY, Tseng LL,
Cheng ML, Chuu CP, Tsai HK, Kuo WL, Kung HJ and Wang WC: Targeting
KDM4B that coactivates c-Myc-regulated metabolism to suppress tumor
growth in castration-resistant prostate cancer. Theranostics.
11:7779–7796. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
155
|
Dai L, Fan G, Xie T, Li L, Tang L, Chen H,
Shi Y and Han X: Single-cell and spatial transcriptomics reveal a
high glycolysis B cell and tumor-associated macrophages cluster
correlated with poor prognosis and exhausted immune
microenvironment in diffuse large B-cell lymphoma. Biomark Res.
12:582024. View Article : Google Scholar : PubMed/NCBI
|
|
156
|
Santiago Torres KR, Duran A, Whitley K,
Roy S, Almaguel F and Casiano CA: Abstract 4301: ENO1 as a
potential target for anti-cancer therapy: Insights into its role in
neuroendocrine prostate cancer cell growth and survival. Cancer
Res. 85(8 Suppl 1): S43012025. View Article : Google Scholar
|
|
157
|
Huang CK, Lv L, Chen H, Sun Y and Ping Y:
ENO1 promotes immunosuppression and tumor growth in pancreatic
cancer. Clin Transl Oncol. 25:2250–2264. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
158
|
Tulamaiti A, Xiao SY, Yang Y, Mutailifu M,
Li XQ, Yin SQ, Ma HT, Yao HF, Yao LL, Hu LP, et al: ENO1 promotes
PDAC progression by inhibiting CD8+ T cell infiltration
through upregulating PD-L1 expression via HIF-1α signaling. Transl
Oncol. 52:1022612025. View Article : Google Scholar
|
|
159
|
Wan W, Li Y, Sun W, Cheng Z, Ma F, Shen S,
Liu H and Zhang J: The DCDC2/ENO1 axis promotes tumor progression
and immune evasion in intrahepatic cholangiocarcinoma via
activating FGL1-LAG3 checkpoint. J Exp Clin Canc Res. 44:1772025.
View Article : Google Scholar
|
|
160
|
Cui X, Yang J, Sun M, He S, Zhou Z and
Chen Y: Effect of PLCE1-driven aerobic glycolysis through ENO1
modulation on immune evasion in esophageal squamous cell carcinoma
(ESCC). J Clin Oncol. 42(16 Suppl): e160162024. View Article : Google Scholar
|
|
161
|
Yue T, Li J, Zhu J, Zuo S, Wang X, Liu Y,
Liu J, Liu X, Wang P and Chen S: Hydrogen sulfide creates a
favorable immune microenvironment for colon cancer. Cancer Res.
83:595–612. 2023. View Article : Google Scholar
|
|
162
|
Liang X, Wang Z, Dai Z, Zhang H, Zhang J,
Luo P, Liu Z, Liu Z, Yang K, Cheng Q and Zhang M: Glioblastoma
glycolytic signature predicts unfavorable prognosis, immunological
heterogeneity, and ENO1 promotes microglia M2 polarization and
cancer cell malignancy. Cancer Gene Ther. 30:481–496. 2023.
|
|
163
|
Ray A, Song Y, Du T, Chauhan D and
Anderson KC: Preclinical validation of Alpha-Enolase (ENO1) as a
novel immunometabolic target in multiple myeloma. Oncogene.
39:2786–2796. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
164
|
Shen C, Liu J, Hu D, Liu C, Xie F and Wang
Y: Tumor-intrinsic ENO1 inhibition promotes antitumor immune
response and facilitates the efficacy of anti-PD-L1 immunotherapy
in bladder cancer. J Exp Clin Canc Res. 44:2072025. View Article : Google Scholar
|
|
165
|
Anderson VE, Weiss PM and Cleland WW:
Reaction intermediate analogues for enolase. Biochemistry.
23:2779–2786. 1984. View Article : Google Scholar : PubMed/NCBI
|
|
166
|
Guha-Chowdhury N, Clark AG and Sissons CH:
Inhibition of purified enolases from oral bacteria by fluoride.
Oral Microbiol Immunol. 12:91–97. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
167
|
de A S Navarro MV, Gomes Dias SM, Mello
LV, da Silva Giotto MT, Gavalda S, Blonski C, Garratt RC and Rigden
DJ: Structural flexibility in Trypanosoma brucei enolase revealed
by X-ray crystallography and molecular dynamics. FEBS J.
274:5077–5089. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
168
|
Jung DW, Kim WH, Park SH, Lee J, Kim J, Su
D, Ha HH, Chang YT and Williams DR: A unique small molecule
inhibitor of enolase clarifies its role in fundamental biological
processes. Acs Chem Biol. 8:1271–1282. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
169
|
Ni J, Huang Y, Li C, Yin Q and Ying J:
Beyond ENO1, emerging roles and targeting strategies of other
enolases in cancers. Mol Ther-Oncolytics. 31:1007502023. View Article : Google Scholar : PubMed/NCBI
|
|
170
|
Cho H, Um J, Lee JH, Kim WH, Kang WS, Kim
SH, Ha HH, Kim YC, Ahn YK, Jung DW and Williams DR: ENOblock, a
unique small molecule inhibitor of the non-glycolytic functions of
enolase, alleviates the symptoms of type 2 diabetes. Sci Rep.
7:441862017. View Article : Google Scholar : PubMed/NCBI
|
|
171
|
Lin YH, Satani N, Hammoudi N, Yan VC,
Barekatain Y, Khadka S, Ackroyd JJ, Georgiou DK, Pham CD, Arthur K,
et al: An enolase inhibitor for the targeted treatment of
ENO1-deleted cancers. Nat Metab. 2:1413–1426. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
172
|
Chen ML, Yuan TT, Chuang CF, Huang YT,
Chung IC and Huang WC: A novel enolase-1 antibody targets multiple
interacting players in the tumor microenvironment of advanced
prostate cancer. Mol Cancer Ther. 21:1337–1347. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
173
|
Lung J, Chen KL, Hung CH, Chen CC, Hung
MS, Lin YC, Wu CY, Lee KD, Shih NY and Tsai YH: In silico-based
identification of human α-enolase inhibitors to block cancer cell
growth metabolically. Drug Des Devel Ther. 11:3281–3290. 2017.
View Article : Google Scholar
|
|
174
|
Möbitz H: Design principles for balancing
lipophilicity and permeability in beyond rule of 5 space.
ChemMedChem. 19:e2023003952024. View Article : Google Scholar
|
|
175
|
Wong VK, Dong H, Liang X, Bai LP, Jiang
ZH, Guo Y, Kong AN, Wang R, Kam RK, Law BY, et al: Rh2E2, a novel
metabolic suppressor, specifically inhibits energy-based metabolism
of tumor cells. Oncotarget. 7:9907–9924. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
176
|
Huang Q, Zhang H, Bai LP, Law BYK, Xiong
H, Zhou X, Xiao R, Qu YQ, Mok SWF, Liu L and Wong VKW: Novel
ginsenoside derivative 20(S)-Rh2E2 suppresses tumor growth and
metastasis in vivo and in vitro via intervention of cancer cell
energy metabolism. Cell Death Dis. 11:6212020. View Article : Google Scholar : PubMed/NCBI
|
|
177
|
Pail O, Lin MJ, Anagnostou T, Brown BD and
Brody JD: Cancer vaccines and the future of immunotherapy. Lancet.
406:189–202. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
178
|
Rappaport AR, Kyi C, Lane M, Hart MG,
Johnson ML, Henick BS, Liao CY, Mahipal A, Shergill A, Spira AI, et
al: A shared neoantigen vaccine combined with immune checkpoint
blockade for advanced metastatic solid tumors: Phase 1 trial
interim results. Nat Med. 30:1013–1022. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
179
|
Cappello P, Rolla S, Chiarle R, Principe
M, Cavallo F, Perconti G, Feo S, Giovarelli M and Novelli F:
Vaccination with ENO1 DNA prolongs survival of genetically
engineered mice with pancreatic cancer. Gastroenterology.
144:1098–1106. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
180
|
Curcio C, Mucciolo G, Roux C, Brugiapaglia
S, Scagliotti A, Guadagnin G, Conti L, Longo D, Grosso D, Papotti
MG, et al: PI3Kγ inhibition combined with DNA vaccination unleashes
a B-cell-dependent antitumor immunity that hampers pancreatic
cancer. J Exp Clin Canc Res. 43:1572024. View Article : Google Scholar
|
|
181
|
Mandili G, Curcio C, Bulfamante S, Follia
L, Ferrero G, Mazza E, Principe M, Cordero F, Satolli MA, Spadi R,
et al: In pancreatic cancer, chemotherapy increases antitumor
responses to tumor-associated antigens and potentiates DNA
vaccination. J Immunother Cancer. 8:e0010712020. View Article : Google Scholar : PubMed/NCBI
|
|
182
|
Hu Q, Shi Y, Wang H, Bing L and Xu Z:
Post-translational modifications of immune checkpoints: Unlocking
new potentials in cancer immunotherapy. Exp Hematol Oncol.
14:372025. View Article : Google Scholar : PubMed/NCBI
|
|
183
|
Cook K, Daniels I, Symonds P, Pitt T,
Gijon M, Xue W, Metheringham R, Durrant L and Brentville V:
Citrullinated α-enolase is an effective target for anti-cancer
immunity. Oncoimmunology. 7:e13906422017. View Article : Google Scholar
|
|
184
|
León-Letelier RA, Sevillano-Mantas AM,
Chen Y, Park S, Vykoukal J, Fahrmann JF, Ostrin EJ, Garrett C, Dou
R, Cai Y, et al: Citrullinated ENO1 vaccine enhances PD-1 blockade
in mice implanted with murine triple-negative breast cancer cells.
Vaccines (Basel). 13:6292025. View Article : Google Scholar : PubMed/NCBI
|
|
185
|
Liu F, Huang H, Yang X, Jiang S, Xu A, Yu
Z, Li J, Yu M, Wang Y and Wang B: Ag85B-ENO1-46-82 therapeutic
vaccines enhance anti-tumor immunity by inducing CD8+ T
cells and remodeling tumor microenvironment. Int Immunopharmacol.
130:1117072024. View Article : Google Scholar
|
|
186
|
Wang L: Screening and biosensor-based
approaches for lung cancer detection. Sensors (Basel). 17:24202017.
View Article : Google Scholar : PubMed/NCBI
|
|
187
|
Yajima S, Ito M, Suzuki T, Oshima Y,
Sumazaki M, Shiratori F, Takizawa H, Li SY, Zhang BS, Yoshida Y, et
al: Application of serum anti-ENO1 and anti-SSNA1 antibody
biomarkers in predicting the prognosis of gastric cancer. Oncol
Lett. 30:3602025. View Article : Google Scholar : PubMed/NCBI
|
|
188
|
Yin H, Wang L and Liu HL: ENO1
overexpression in pancreatic cancer patients and its clinical and
diagnostic significance. Gastroenterol Res Pract. 2018:38421982018.
View Article : Google Scholar : PubMed/NCBI
|
|
189
|
Bag S, Dutta D, Chaudhary A, Sing BC, Pal
M, Ray AK, Banerjee R, Paul RR, Basak A, Das AK and Chatterjee J:
Identification of α-enolase as a prognostic and diagnostic
precancer biomarker in oral submucous fibrosis. J Clin Pathol.
71:228–238. 2018. View Article : Google Scholar
|
|
190
|
Li G, Li Y, Zhou J, Tang S, Guo H and Lin
J: Prognostic evaluation of glycolysis markers in hepatocellular
carcinoma: insights from meta-analysis and multi-omics approaches.
BMC Med Genomics. 18:1792025. View Article : Google Scholar : PubMed/NCBI
|
|
191
|
Wang L, Qu M, Huang S, Fu Y, Yang L, He S,
Li L, Zhang Z, Lin Q, Zhang L, et al: A novel α-enolase-targeted
drug delivery system for high efficacy prostate cancer therapy.
Nanoscale. 10:13673–13683. 2018. View Article : Google Scholar : PubMed/NCBI
|