Introduction
Cervical cancer is one of the most common
gynecological carcinomas in the world with 569,847 cases and
311,365 mortalities annually (1).
It ranks the second most prevalent lethal tumors in developing
countries and the tenth in developed countries; in 2020, ~600,000
women were diagnosed with invasive cervical cancer, mostly in
advanced stages (2). The World
Health Organization (WHO) announced cervical cancer as a public
health problem in May 2018. Cervical cancer is classified into two
histological subtypes including squamous cell carcinoma (SCC) and
adenocarcinoma (ADC) accounting for 68 and 21%, respectively. These
two subtypes have different features in accordance with etiology,
screening effectiveness, and prognosis (3). Accumulated studies have suggested
that ADC exhibits worse prognosis compared with SCC (4). The major risk factors related to
cervical cancer involve persistent human papilloma virus (HPV)
infection, smoking, presence of sexual transmitted disease, such as
human immunodeficiency virus or chlamydia infection (5). Despite early screening and
vaccination, it remains the second leading cause of mortality in
women aged 20-39 (6). There are
critical individual differences in the prognosis of patients
because of tumor heterogeneity. Thus, it is essential to
investigate molecular heterogeneity and establish a complete
prognostic assessment system, improving the precision treatment for
cervical cancer.
Cancer testis antigens (CTAs) are specifically
expressed in but abnormally expressed in some types of cancer
including melanoma, lung cancer, liver cancer even in female cancer
such as ovarian cancer and breast cancer (7-11).
CTAs are expressed through embryonic development but abnormally
re-expressed during tumorigenesis. Studies suggest that
overexpression of oncogenic CTAs improves cancer cell stemness,
tumorigenicity, metastasis, and drug resistance (12-15). The oncogenic CTAs provoke the
initiation and development of malignant tumors (16). Physiological expression of CTAs in
normal tissues is mostly limited to testis, and few are detected in
normal adult somatic cells and tissues such as placental tissue
(17). Until now, more than 200
CTAs have been identified and ~100 gene families are extremely
expressed in malignant tumors. Due to unique expression patterns,
these CTAs are considered as immunogenic tumor-associated antigens
promising targets for cancer treatment (18).
The first discovered gene among CTAs was the
Melanoma Antigen gene (MAGE) family and is highly conserved in all
eukaryotes. The MAGE family can be categorized in accordance with
the pattern of tissue expression: Type I MAGEs are categorized as
CTAs, including MAGE-A, -B, and -C subfamilies, whose genes are
specifically located on the X chromosome. Type II MAGEs include
MAGE-D to -L are expressed around various tissues in human body
(19,20). Among the MAGE family, the role of
MAGE-A groups is mostly unknown, exerting different functions in
terms of subcellular localizations. Nonetheless, MAGEA1 is known to
be located in the cytosol of melanoma cells and has also been
detected in both the cytoplasm and nucleus of spermatogonia
(21). MAGEA1 belongs to a
cluster of 12 MAGE-A genes in X chromosome q28 region (22). Previous studies showed that
promoter methylation and histone acetylation improve the expression
of MAGEA1 in human cancer cells including lung cancer, breast
cancer and colorectal cancer (23,24). However, detail functions are
mainly unrevealed. Thus, this study aims to investigate the role of
MAGEA1 in cervical cancer and its potential as a therapeutic target
and biomarker.
Materials and methods
Survival prognosis analysis
Kaplan-Meier survival analysis was performed using
the KM Plotter online database (https://kmplot.com/analysis/) to evaluate the
prognostic significance of MAGEA1 expression levels in cancer
patients. This publicly available platform contains comprehensive
survival data, including overall survival information for patients
with ovarian, lung, gastric, colon cancers, and cervical squamous
cell carcinomas. Kaplan-Meier plots were generated to assess the
correlation between MAGEA1 expression levels and patient survival
outcomes across these cancer types.
Cells and cell culture
HeLa (CCL-2) and Panc1 (CRL-1469) were purchased
from American Type Culture Collection. SK-OV-3 (cat. no. 30077),
T24 (cat. no. 30004), SiHa (cat. no. 30035) were purchased from
Korean Cell Line Bank. The cells were cultured in Dulbecco's
modified Eagle's medium (Biowest) with 10% fetal bovine serum
(Gibco; Thermo Fisher Scientific, Inc.) and 1%
antibiotic-antimycotic (Gibco; Thermo Fisher Scientific, Inc.) in a
humidified atmosphere of 5% CO2 at 37°C.
Reverse transcription-quantitative (RT-q)
PCR
The total RNA was extracted from 1×106
cells with TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.). The concentration and quality of the total RNA
were measured by Nanodrop (NanoDrop Technologies; Thermo Fisher
Scientific, Inc.). For cDNA synthesis, the reverse transcription
was performed using reverse transcription system (Promega
Corporation). Primers for each target gene were designed by
extracting the coding sequence from the corresponding NCBI
reference sequence and generating forward and reverse primers
accordingly. The primer sequences used in this study were as
follows: MAGEA1 forward: 5'-AGGTGGCTGATTTGGTTGGT-3', reverse:
5'-GACCAGCTGCAAGGACTCAG-3'; β-actin forward:
5'-GCACTCTTCCAGCCTTCCTT-3', reverse: 5'-GTTGGCGTACAGGTCTTTGC-3'.
The mRNA level of MAGEA1 was determined by qPCR according to
triplicate with SYBR green mix (Bio-Rad Laboratories, Inc.) and
analyzed on a CFX connect Real-Time System (Bio-Rad Laboratories,
Inc.). 95°C for 5 min, followed by 40 cycles of 90°C for 10 sec and
60°C for 30 sec, with a final melt-curve analysis. The relative
expression level was calculated based on the 2−ΔΔCq
method (25).
Western blot analysis
Total proteins were extracted from cells using RIPA
buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.5% sodium
deoxycholate, 1% Triton X-100, 0.1% SDS, 2 mM EDTA) supplemented
with protease inhibitor cocktail (Thermo Fisher Scientific, Inc.).
Protein concentration was determined using a BCA assay (Thermo
Fisher Scientific, Inc.). Equal amounts of protein (20 μg
per lane) were resolved by 10% SDS-PAGE, transferred to PVDF
membranes (MilliporeSigma) and blocked with 5% skimmed milk at room
temperature for 1 h. Membranes were incubated with primary
antibodies against MAGEA1 (1:100; cat. no. sc-20033; Santa Cruz
Biotechnology, Inc.) and GAPDH (1:4,000; cat. no. sc-32233; Santa
Cruz Biotechnology, Inc.), followed by HRP-conjugated secondary
antibody (1:5,000; cat. no. 1706516; Bio-Rad Laboratories, Inc.).
Bands were visualized using an enhanced chemiluminescence system
(Intron Biotechnology, Inc.). Densitometric analysis was performed
using ImageJ software, version 1.53k (National Institutes of
Health).
Lentiviral transduction for stable
knockdown cell construction
Lentiviral transduction was performed using MAGEA1
short hairpin RNAs (shRNAs) lentiviral plasmids (#1 and #2, cat.
no. 4100; OriGene Technologies, Inc.) in pGFP-C-shLenti vector. The
non-targeting scrambled shRNA, expressing GFP was purchased from
OriGene Technologies, Inc.; cat. no. TR30021V). The shRNA sequences
used were as follows: shMAGEA1 #1:
5'-TGGAGTCCTTGTTCCGAGCAGTAATCACT-3', shMAGEA1 #2:
5'-CAGTCACAAAGGCAGAAATGCTGGAGAGT-3'. 293T cells (OriGene
Technologies, Inc.) were transfected with 5 μg of shRNA
plasmids and 3rd-generation lentiviral components (REV, RRE and
MD2.G vector) at a weight ratio of 4:2:1:1 using transfection
reagents (Promega Corporation). Additional information is provided
in Table SI. Media was replaced with antibiotic-free DMEM/10% FBS
overnight post-transfection at 37°C. Viral supernatant was
harvested 72 h later and centrifuged at 500 × g at 4°C for 5 min.
HeLa cells were infected with the viral supernatant at a
multiplicity of infection (MOI) of 10 for 24 h and selected with 1
μg/ml puromycin for four days after 72 h post-infection
(26). Subsequent experiments
were conducted 48 h after the selection process. Following initial
selection, cells were maintained in 0.8 μg/ml puromycin to
preserve stable integration.
Cell proliferation assay
HeLa cells were seeded into 6-well plate at a
density of 1.5×105 cells/well for 48 h and
1×105 cells/well 37°C for 72 h. Each sample was set in
triplicate. Cells were detached with EDTA-trypsin and the collected
cells were counted with hemocytometer C-Chip (Incyto) using a light
microscope (CKX3-SLP; Olympus Corporation) at ×40
magnification.
Wound scratch assay
HeLa cells were cultured in 6 cm plates until they
formed a confluent layer. The scratch was inflicted with a sterile
10 μl pipette tip across the center and side of cell
monolayer. The medium was removed and washed PBS. Then, 5 ml DMEM
without FBS (serum-free) per plate was replaced and incubated as
previously described (27). Each
image were acquired with light microscope (Ts2R-FL; Nikon
Corporation) with a camera for 24, 48, 72 and 96 h following
scratch. To determine the wound closure rate, phase contrast images
were estimated through the image processing program ImageJ,
software, version 1.53k (National Institutes of Health).
Transwell migration and invasion
assays
Transwell migration and invasion assays were
performed using Transwell plates (SPLInsert hanging; SPL Life
Sciences Co., Ltd.). For migration assays, upper chambers with 8
μm pores were coated with 0.2% gelatin. Cells
(5×104 cells/well) were seeded in serum-free DMEM in the
upper chamber, with 10% FBS-containing medium in the lower chamber.
After 6 h, migrated cells were fixed with methanol at room
temperature for 15 min and stained with 1% crystal violet. For
invasion assays, upper chambers were coated with 0.2% gelatin and
Basement Membrane Extract (BME; equivalent to Matrigel). The
pre-coating was performed at 37°C for 1 h to allow the BME to
solidify. Cells (5×104 cells/well) were seeded similarly
and incubated for 24 h. Invaded cells were fixed, stained, and
counted in five fields per well.
Cell viability assay
Cell viability was measured by estimating the
reduction of 3-(4,5-dimethylthiazol-2) 2,5-diphenyltetrazolium
bromide (MTT) to formazan by mitochondrial dehydrogenase.
5×103 cells/well were seeded on 96 well plates a diluted
MTT/PBS solution (5 mg/ml) with media was then added to the wells
and incubated at 37°C for 2 h. Living cells reduced MTT to formazan
which was quantified by evaluating absorbance at 570 nm wavelength
using a microplate reader (GloMax Explorer; Promega Corporation).
The cell viability was calculated as CV(%)=(Absorbance of test
sample/Absorbance of control) ×100.
Cell cycle analysis
For flow cytometric analysis of cell cycle, cells
were harvested and fixed with cold 70% ethanol for 48 h at −20°C.
The cells were washed and resuspended in PBS. RNase A (50
μl; 100 μg/ml; Thermo Fisher Scientific, Inc.) was
added and incubated at 37°C water bath for 20 min. Then, the cells
were treated with 5 μl propidium iodide (10 mg/ml,
Invitrogen; Thermo Fisher Scientific, Inc.) seeded in 96 well
plates with density at 1×106 cells/well and incubated at
4°C in dark until flow cytometry readings. Cell cycle distribution
was measured by flow cytometry (CytoFLEX V0-B3-R2 Flow Cytometer;
Beckman Coulter, Inc.). The data was analyzed using the CytoFLEX
platform (v2.4, Beckman Coulter, Inc.).
Construction of biomimetic 3D culture
systems; gelatin methacryloyl (GelMA) colony formation
Lyophilized GelMA was dissolved to 5% GelMA in cell
culture media (DMEM) with 0.1% Lithium
phenyl-2,4,6-trimethylbenzoylphosphinate. Dissolved GelMA hydrogels
were fabricated by dispensing the hydrogel precursors into 24-well
culture plate and exposing GelMA with cells at a density of
1.5×104 cells/well to 405 nm UV light for 30 sec. Then,
1 ml of DMEM was added and incubated at 37°C for seven days. The
culture media were replaced every three days to maintain optimal
cell growth conditions. Following incubation, the cells were
stained with DAPI (MilliporeSigma) at room temperature for 10 min.
Each image was captured with a light microscope (Ts2R-FL; Nikon
Corporation) with an attached camera.
RNA sequencing (RNA-seq)
Cellular RNA was isolated from shNC and shMAGEA1 #1
and #2 using TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) according to manufacturer's instructions. TruSeq
Stranded mRNA Library Prep Kit (Illumina, Inc.) was used to isolate
poly(A)+ mRNA and synthesize cDNA for sequencing library
construction. The prepared libraries were sequenced in triplicate
on the Illumina NovaSeq 6000 platform (Illumina, Inc.) with a
paired-end 150 bp configuration.
Raw sequencing reads were preprocessed using
Trimmomatic v0.39 (28), where
low-quality bases (Q-score <20) and adapter sequences were
removed. The trimmed reads were aligned to the Homo sapiens
GRCh38 reference genome available on ENSEMBL (release 110;
http://ftp.ensembl.org/pub/release-110/fasta/homo_sapiens/dna/)
using STAR aligner v2.7.11b (29), and the alignment results were
converted to BAM format and sorted using samtools v1.7 (30). The read counts were obtained using
FeatureCounts v2.0.8 (31).
Normalization of raw count data was conducted using edgeR v4.2.1
(https://bioconductor.org/packages/edgeR/based on the
TMM method, followed by GLM fitting and likelihood ratio test for
differential expression analysis (32).
Principal Component Analysis (PCA) was performed
using the plotPCA function in R to visualize the variance and
separation between the control and MAGEA1 knockdown groups based on
the normalized gene expression data. Gene expression patterns and
enrichment analyses were conducted across three groups (one control
and two experimental groups) using the pooled count data to assess
gene expression patterns. To visualize this result, a volcano plot
was generated using ggrepel v0.9.6 (33) and ggplot2 v3.5.1 (34), where differentially expressed
genes (DEGs) were identified based on P-value <0.05.
Additionally, the top 10 most statistically significant genes were
labeled for emphasis. Gene set enrichment analysis (GSEA) was
performed using GSEA software v4.3.3 (35,36) with the MSigDB Hallmark gene sets
(h.all.v2024.1.Hs.symbols.gmt) database (37). Heatmaps were created using
pheatmap v1.0.12 (38) and
RColorBrewer v1.1.3 (39) to
compare the expression patterns of enriched gene sets.
Functional annotation of DEGs was conducted using
the DAVID web server (40,41),
where gene ontology pathway analysis was performed (42,43). DEGs were selected based on
overlapping genes between two experimental groups compared with the
control [false discovery rate (FDR) <0.01] and classified as
upregulated or downregulated according to absolute log2 fold change
>0. Furthermore, functional interactions among DEGs were
explored using ClueGO v2.5.8 (44,45), with pathway network visualization
filtered at term P<0.05.
Tumor xenograft model in nude mice
All animal experimental procedures were performed
and approved by accordance with the Catholic University of Korea
(approval no. CUK-IACUC-2022-009). Female nude mice (nu/nu;
4-week-old; initial weight, 18-22 g; n=15) were purchased from
Orient bio (https://www.orient.co.kr/). The mice
were housed in a specific pathogen-free (SPF) facility maintained
at 22±2°C with 55±5% humidity and a 12-h light/dark cycle. To
induce tumor genesis, right flank of nude mice (n=5 per group) were
subcutaneously inoculated with 1×107 cells transfected
with scrambled and shMAGEA1 #1 and #2 resuspended in 150 μl
PBS. The mice were randomized after one week and observed for 28
days. Tumor volumes and body weights were calculated twice per week
after injection; Tumor volume (mm3)=length (mm) ×
width2 (mm2) × 1/2. All treated mice were
observed and weighed until they met one of the criterion for
sacrifice as specified in the IACUC-approved protocol, including
tumor volume and body weight thresholds, or until the experimental
endpoint was reached. Mice were sacrificed using a gradual-fill
CO2 method at a displacement rate of 30-70% of the
chamber volume per min.
Immunohistochemistry (IHC) staining
The tumor tissue samples were fixed with 4%
paraformaldehyde at 4°C for 24 h. Following fixation, the tumor
tissues were dehydrated through a graded series of ethanol (70, 80,
90, 95, and 100%), cleared in xylene, and then embedded in paraffin
wax at 60°C. Sections (4 μm thick) were deparaffinized,
rehydrated, and permeabilized with 0.1% Triton X-100. IHC was
conducted with IHC kit (Abcam; cat. no. ab64264) according to the
manufacturer's instructions. Briefly, nonspecific background was
blocked using the provided Protein Block for 30 min at room
temperature. The sections were incubated with primary antibodies
including Ki-67 (1:500; cat. no. HGS-S239; Acro Biosystems Ltd.)
and MAGEA1 (1:100; cat. no. sc-20033; Santa Cruz Biotechnology,
Inc.) at 4°C overnight. Then, the sections were incubated with the
kit's HRP-conjugated secondary antibody (1:1,000) for 1 h at room
temperature. Signal was visualized using DAB solution for 5 min at
room temperature, followed by counterstaining with Mayer's
Hematoxylin for 2 min at room temperature. Each image was captured
with microscope slide scanner (Aperio CS2; Leica Biosystems).
Statistical analysis
All experimental data were performed in triplicate.
Statistical significance was determined using GraphPad Prism (v9;
Dotmatics) with one-way ANOVA for normally distributed datasets,
followed by a suitable post hoc multiple comparisons test (Tukey's
multiple comparisons test) to compare the control and MAGEA1
knockdown groups. Error bars in replicate datasets are presented as
the standard error of the mean (SEM). P<0.05 was considered to
indicate a statistically significant difference.
Results
Effects of MAGEA1 on cervical cancer cell
proliferation and motility
The association between MAGEA1 expression levels and
overall survival across various cancer types was evaluated using
Kaplan-Meier survival analysis. This analysis demonstrated that
lower MAGEA1 expression in patients with cancer, particularly those
with ovarian, lung, gastric, colon, and cervical squamous cell
cancers, is markedly associated with improved overall survival,
suggesting that high MAGEA1 expression may negatively affect
patient prognosis (Fig. S1A-E). Subsequently, the present study
analyzed MAGEA1 protein expression levels in various cancer cell
lines. Western blot analysis revealed that MAGEA1 expression was
markedly elevated in cervical cancer cells (Fig. 1A). To investigate the role of
MAGEA1 in cervical cancer, stable MAGEA1 knockdown cell lines (Hela
and SiHa) were established by infecting cells with lentiviruses
carrying green fluorescence protein (GFP)-tagged constructs
encoding either a scrambled shRNA or MAGEA1-specific shRNAs
designed to effectively silence MAGEA1 expression. In addition, GFP
fluorescence was consistently observed in all groups treated with
scrambled shRNA, shMAGEA1 #1 and shMAGEA1 #2, indicating successful
and stable transduction using GFP-tagged vectors (Fig. S1F). The
cells transduced with shMAGEA1 #1 and shMAGEA1 #2 effectively
suppressed both MAGEA1 mRNA and protein expression levels (Figs. 1B and C and S2A and B).
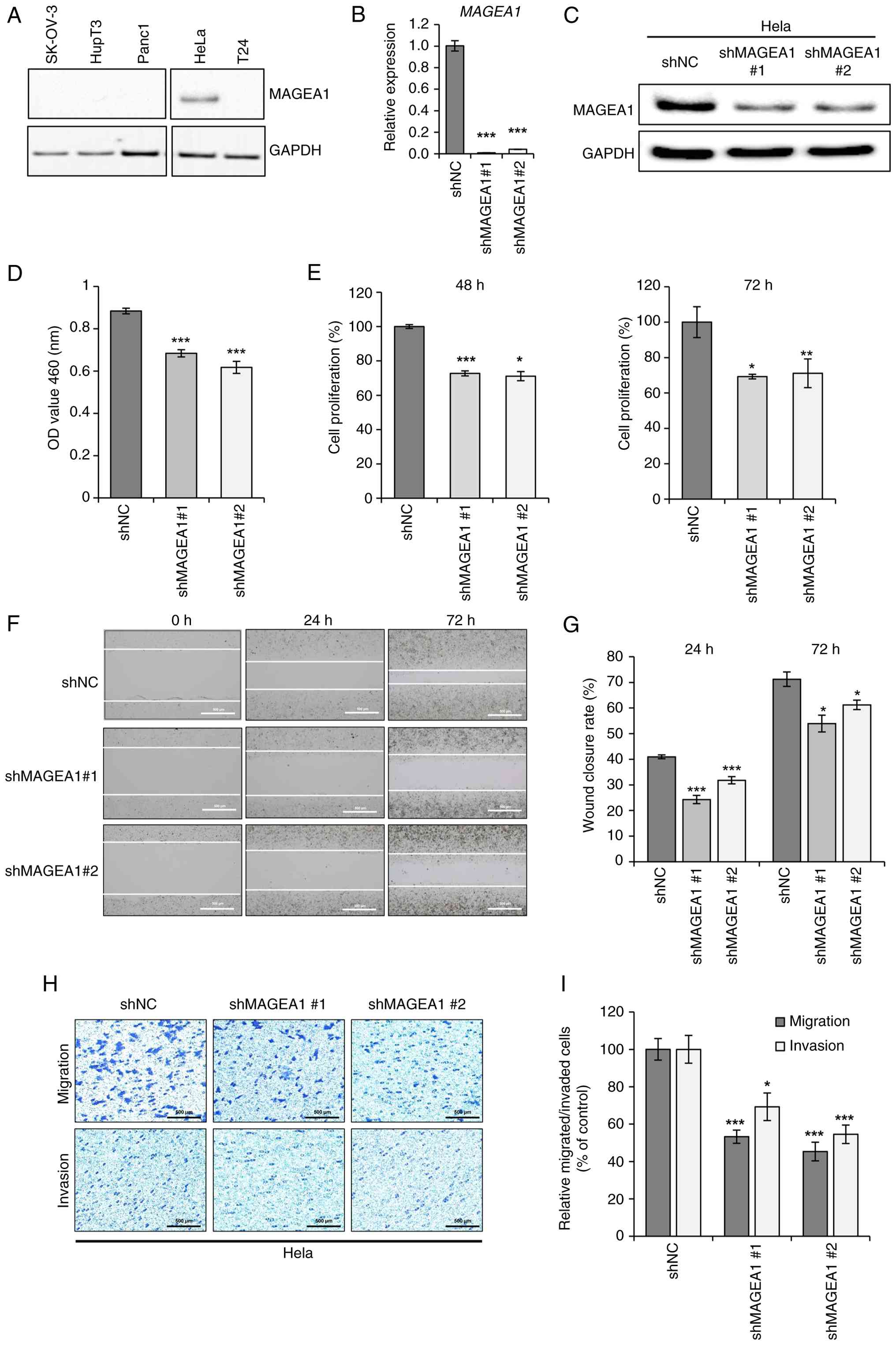 | Figure 1Effects of MAGEA1 knockdown on
mRNA/protein expression and cellular functions including
proliferation, viability, migration, and invasion. (A) Western
blotting detected MAGEA1 expression in SK-OV-3 (ovarian cancer),
Panc1 (pancreatic cancer), HupT3 (pancreatic cancer), HeLa
(cervical cancer), and T24 (urinary bladder carcinoma). GAPDH was
used as a loading control. HeLa was transfected with shMAGEA1 #1
and #2 and shNC was established as a control. (B) Reverse
transcription-quantitative PCR and (C) western blot analysis were
used to estimate the efficiency of knockdown. (D) Cell viability
was determined by MTT assay after 24 h of cell seeding. (E) A
significant decrease in cell proliferation was observed in MAGEA1
knockdown compared with control. The number of proliferated cells
is presented as a percentage of the control. (F) Representative
images from wound scratch at different time points (magnification,
×40). (G) Percentages of wound closure at 24 and 72 h are shown as
a bar graph. The scratched area and lines were quantified by ImageJ
software with the MRI tool. (H) Cell migration and invasion were
confirmed by Transwell migration and invasion assays.
Representative images of cells are illustrated below. Scale bar,
500 μm. (I) Quantification of migrated and invaded cells in
distinct groups. The number of migrated and invaded cells is
presented as a percentage of the control. Error bars represent the
mean ± SEM. *P<0.05, **P<0.01,
***P<0.005. MAGEA1, MAGE family member A1; sh, short
hairpin; NC, negative control. |
Next, to elucidate the functional role of MAGEA1 in
cervical cancer, a series of in vitro assays was performed
using stable MAGEA1-knockdown cells. The MTT assay revealed that
MAGEA1 knockdown markedly reduced cell viability to 77 and 69%
compared with the control group (Fig.
1D). Cell proliferation assays also showed that MAGEA1
knockdown markedly reduced cell proliferation at both 48 and 72 h
(Figs. 1E and S2C). In wound healing assays performed
to assess cellular migration, MAGEA1 depletion inhibited the
migratory ability of cervical cancer cells (Fig. 1F and G). In addition, Transwell
chamber assays revealed a significant reduction in both the
migratory and invasive abilities of MAGEA1-knockdown cells
(Figs. 1HI and S2D), suggesting that MAGEA1 contributes
not only to the proliferative capacity of cervical cancer cells but
also to their motility.
Effects of MAGEA1 on
anchorage-independent cell growth and drug sensitivity
While 2D cell culture systems are widely used in
cancer research, they have limitations in accurately replicating
the complex 3D architecture and cell-cell interactions of the TME.
3D cell culture using hydrogels is commonly employed for an
improved modeling of the native extracellular matrix and
intercellular communication. In the field of tissue engineering,
hydrogels are regarded as promising scaffolds due to their
biocompatibility, tunable mechanical properties, and ability to
encapsulate cells and bioactive molecules (46). GelMA hydrogel, a bioink commonly
used for 3D bioprinting, is a synthetically modified form of
gelatin, in which methacrylate-based groups are introduced into the
gelatin side chains through substitution reactions targeting amino
(-NH2) and hydroxyl (-OH) groups. The incorporation of
methacrylamide groups enhances GelMA's functionality by enabling
photo-initiated cross-linking, which facilitates the formation of
hydrogels with tunable stiffness and density. These adjustable
properties make GelMA an ideal material for constructing biomimetic
3D culture systems. Using GelMA-based hydrogels, the present study
established a 3D culture system to evaluate the colony-forming
ability of cervical cancer cells under more physiologically
relevant conditions (Fig. 2A). In
the colony formation assay, most cells in the control group formed
colonies consisting of 6-8 cells, whereas the MAGEA1 knockdown
groups predominantly formed smaller colonies composed of 2-4 cells
(Fig. 2B). Notably, the number of
colonies containing eight or more cells was notably reduced in the
MAGEA1 knockdown groups, accounting for only 29 and 32% relative to
the control group (Fig. 2C),
suggesting that MAGEA1 is closely associated with
anchorage-independent cell growth in 3D environments.
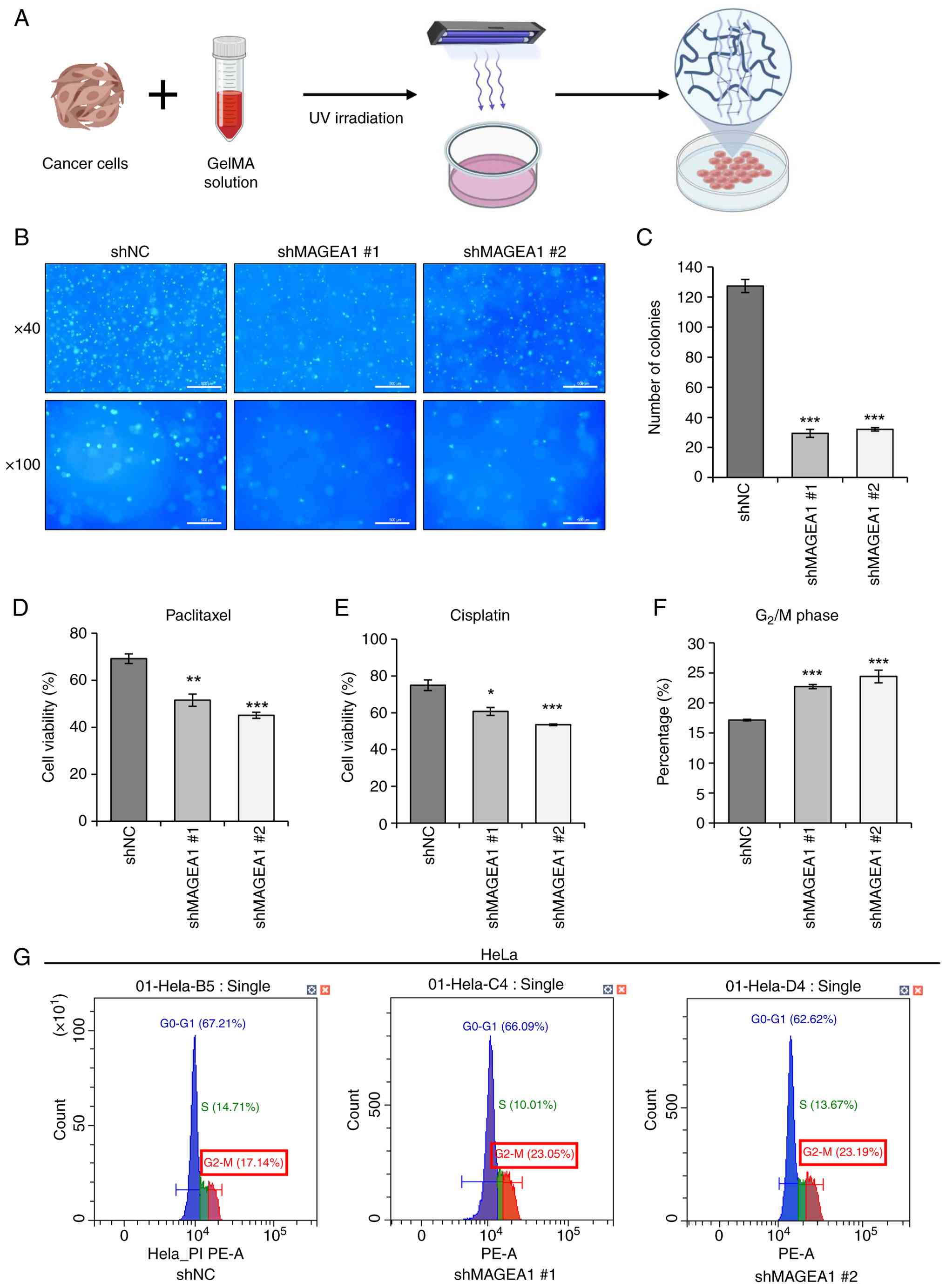 | Figure 2Effect of MAGEA1 knockdown on colony
formation, chemosensitivity, and cell cycle progression. (A) A
Schematic diagram of 3D experimental design. (B) Fluorescent
microscope images of DAPI staining (magnification ×40, ×100). Scale
bar, 500 μm. (C) Quantification of colony formation with 8
or more colonies on GelMA. Cells were treated with (D) 1 μM
of paclitaxel and (E) 20 μM of cisplatin for 48 h and cell
viability was measured by MTT assays. (F) Cells were stained with
PI and analyzed using Cytexpert software (v2.4). (G) Representative
DNA content histograms showing distributions across G0-G1, S, and
G2-M phases in each group samples. Error bars represent
the mean ± SEM. *P<0.05, **P<0.01,
***P<0.005. MAGEA1, MAGE family member A1; GelMA,
gelatin methacryloyl. |
Paclitaxel is a widely used chemotherapeutic
diterpenoid alkaloid that exerts its anti-cancer effects by
promoting tubulin polymerization and inhibiting mitosis (47). Cisplatin, a platinum-based
chemotherapeutic agent, acts as a DNA-damaging compound that
inhibits tumor growth. Through preliminary dose-response
experiments, we determined the concentrations of paclitaxel (1
μM) and cisplatin (20 μM) that inhibit ~30% of cell
growth in HeLa cells. Treatment with these concentrations of
paclitaxel and cisplatin resulted in a significant increase in drug
sensitivity in MAGEA1 knockdown cells compared with the control
group (Fig. 2D and E), suggesting
that MAGEA1 may be involved in drug resistance in cervical cancer
cells.
Based on the observation that MAGEA1 is closely
associated with cancer cell growth, the present study analyzed the
cell cycle distribution of cervical cancer cells using flow
cytometry. As a result, the proportion of cells in the
G2/M phase was markedly increased in the MAGEA1
knockdown group compared with the control group (Fig. 2F and G). This accumulation of
cells in the G2/M phase suggested that MAGEA1 depletion
impaired the ability of cervical cancer cells to progress through
mitosis, potentially contributing to the reduced proliferation
observed in knockdown cells.
MAGEA1 alters the expression of genes
related to the cell cycle, hypoxia, and apoptosis
To investigate MAGEA1-mediated regulation of gene
expression, RNA-seq was performed on both MAGEA1 knockdown cells
and control cells. Principal component analysis revealed a clear
separation between control and MAGEA1 knockdown groups, indicating
distinct transcriptomic differences (Fig. S3A). A total of 2,289
upregulated and 2,016 downregulated DEGs were identified between
the control and MAGEA1 knockdown groups (FDR <0.01; |log2 fold
change|>0). A heatmap illustrated the clustering and expression
patterns of these DEGs between shNC- and shMAGEA1-transfected
cells, offering a comprehensive overview of transcriptomic
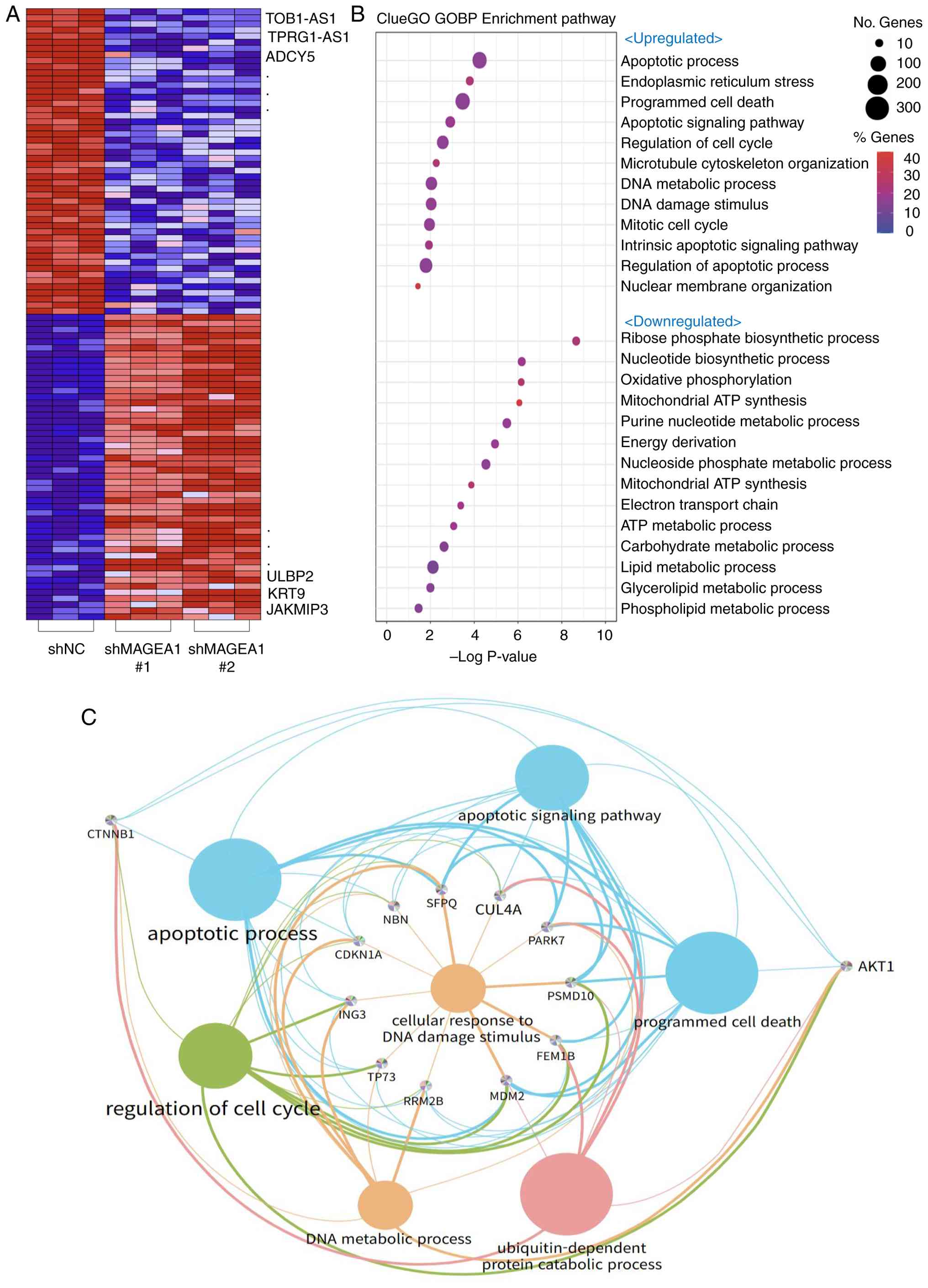
alterations (Fig. 3A). In
addition, a volcano plot depicted the distribution of fold changes,
highlighting the most markedly dysregulated genes (Fig. S3B).
ClueGO-based Gene Ontology Biological Process analysis revealed
significant upregulation of pathways related to apoptotic
processes, cell cycle regulation and DNA damage response, while
pathways associated with metabolic functions, including ATP
synthesis, nucleotide biosynthesis, oxidative phosphorylation, and
lipid metabolism, were substantially downregulated (Fig. 3B). Functional enrichment network
analysis revealed crosstalk among upregulated pathways, with
apoptotic processes, DNA damage response, and cell cycle regulation
forming interconnected clusters (Fig.
3C). Conversely, the downregulated gene network was
predominantly enriched in metabolic pathways, including oxidative
phosphorylation, the electron transport chain, and phospholipid
metabolism (Fig. S3C).
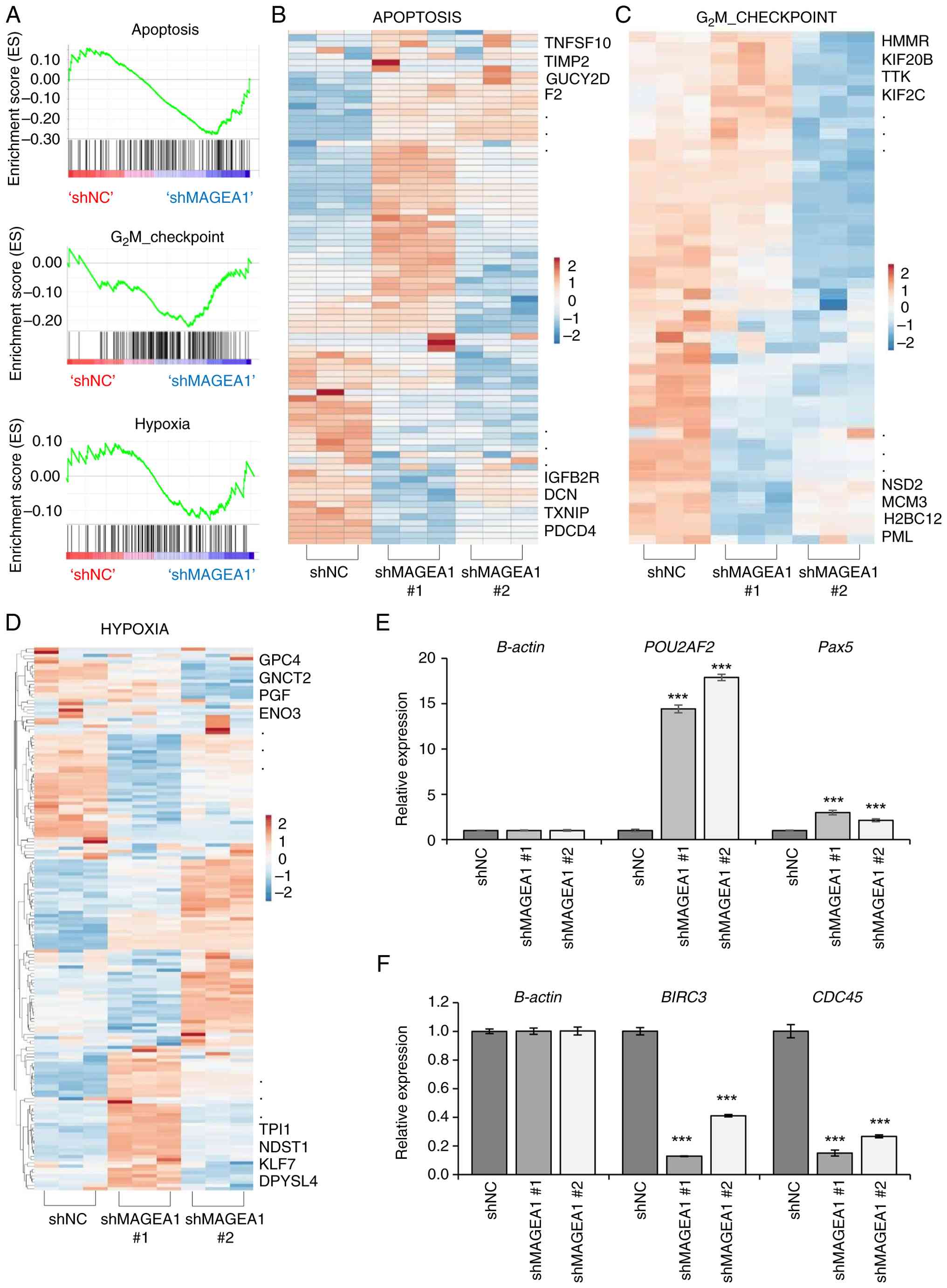
GSEA and expression heatmaps confirmed significant
positive enrichment of apoptosis and G2/M checkpoint
gene sets in MAGEA1 knockdown cells, whereas hypoxia-related
pathways exhibited negative enrichment (Fig. 4A-D). To validate the
RNA-sequencing results, several key DEGs were selected for
confirmation via RT-qPCR. The data showed elevated mRNA expression
levels of POU2AF2 and Pax5 (Fig. 4E; Tables SII and SIII). POU2AF2, a
regulator of tuft cell differentiation, may be inversely associated
with the increased risk of cancer development (48). Pax5 is primarily known as a B
cell-specific transcription factor and also exhibits
tumor-suppressive activity by inducing cell cycle arrest in primary
effusion lymphoma (49). In
addition, RT-qPCR analysis confirmed the downregulation of
BIRC3, and CDC45 (Fig.
4F; Tables SII and SIII). These genes are associated with
cervical cancer cell survival and the inhibition of apoptosis
(50). In particular, CDC45, a
gene implicated in cervical cancer progression, may contribute to
G2/M phase cell cycle arrest and suppressed cell
proliferation (51). POU2AF2,
Pax5, BIRC3 and CDC45 were prioritized because they exhibit
biological relevance to MAGEA1-associated pathways, as previous
evidence indicates functional interactions or co-regulation with
MAGEA1 in tumor progression.
Collectively, these transcriptomic analyses
demonstrated that MAGEA1 knockdown induced profound alterations in
the gene expression profile of cervical cancer cells, particularly
impacting pathways related to cell survival, DNA damage response
and cellular metabolism. These molecular changes identify MAGEA1 as
a key regulator of cervical cancer progression and emphasize its
potential as a promising therapeutic target.
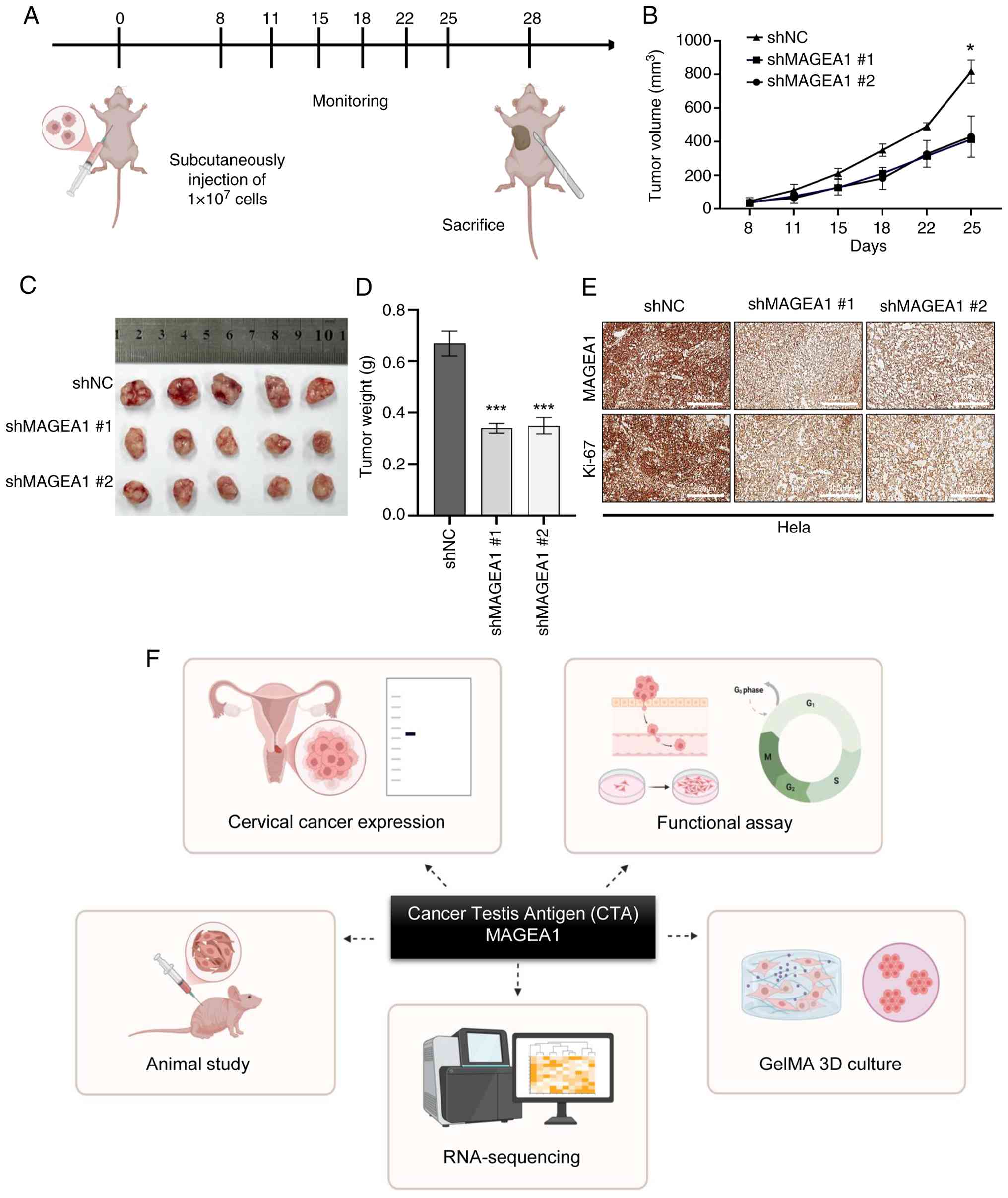
MAGEA1 depletion suppresses in vivo tumor
growth
To evaluate the effect of MAGEA1 on in vivo
tumor growth, HeLa cells stably expressing shNC, shMAGEA1 #1, or
shMAGEA1 #2 were subcutaneously injected into BALB/c nude mice.
Tumor size and mouse body weight were measured twice weekly, and at
the end of the experiment, tumors were excised for imaging and
weighed (Fig. 5A). Monitoring of
mouse body weights throughout the experiment revealed no
significant differences between the control group and the shMAGEA1
groups (Fig. S4A). During the three-week observation period, tumors
in the control group grew progressively, while those in the
shMAGEA1 #1 and #2 groups showed markedly slower growth rates. On
day 25, analysis of individual tumor volumes showed that the mean
tumor volumes in the shMAGEA1 #1 group (413.97±45.35
mm3) and shMAGEA1 #2 group (430.24±122.87
mm3) were reduced by 49.3 and 48.3%, respectively,
compared with the control group (816.84±156.23 mm3;
Fig. 5B). Tumor weight
measurements demonstrated that tumors in the shMAGEA1 #1 group
(0.34 g) and shMAGEA1 #2 group (0.35 g) were markedly reduced
compared with the control group (0.67 g; Fig. 5C and D). IHC analysis of tumor
tissues confirmed the sustained suppression of MAGEA1 protein
expression in MAGEA1 knockdown tumors (Fig. 5E, top). Furthermore,
immunostaining for the proliferation marker Ki-67 showed that the
number of Ki-67-positive cells was markedly reduced in shMAGEA1
tumors (Fig. 5E, bottom).
Collectively, these results clearly demonstrated that MAGEA1
depletion effectively suppresses in vivo tumor growth in
cervical cancer.
Discussion
The present study revealed through a series of
functional assays that MAGEA1 was highly upregulated in cervical
cancer and functioned as a key regulator of tumor progression
(Fig. 5F). Knockdown of MAGEA1
markedly inhibited cell proliferation, migration, invasion and
tumor growth, while enhancing chemosensitivity. The present study
further explained the molecular mechanisms underlying the role of
MAGEA1 in cervical cancer through comprehensive RNA-seq analysis.
The bioinformatics evaluation disclosed significant alterations in
key cellular pathways following MAGEA1 knockdown, notably the
upregulation of apoptosis, DNA repair and cell cycle checkpoint
pathways. Additionally, the downregulation of metabolic pathways,
including ATP metabolism and lipid metabolism, suggested that
MAGEA1 plays a critical role in regulating cancer cell metabolism,
a hallmark of cancer progression. To validate the in vitro
findings in a more physiologically relevant context, a xenograft
model was established using BALB/c nude mice. Especially, both
shMAGEA1 groups exhibited markedly reduced tumor growth and volume
compared with the control group, strongly supporting the in
vivo tumor-suppressive effect of MAGEA1 knockdown. These in
vivo findings were consistent with the in vitro data and
further validated MAGEA1 as a key regulator of cervical cancer
progression.
MAGEA1 is a member of the CTA family and is normally
restricted to immune-privileged tissues such as the testis.
However, it is aberrantly re-expressed in a wide range of
malignancies, including melanoma, lung, gastric, colorectal and
breast cancers, as well as hematologic cancers (22,52-56). High MAGEA1 expression has been
associated with aggressive clinicopathological features and poor
patient survival, particularly in breast, lung and gastric cancer
(57). In gastric and colorectal
cancers, MAGEA1 genes have been implicated in carcinogenesis and
proposed as diagnostic markers of malignant transformation
(53,58). Functional studies in lung
adenocarcinoma further indicate that MAGEA1 promotes tumor growth
and may represent a promising target for antigen-specific
immunotherapy (59). However,
compared with other CTAs, MAGEA1 remains relatively poorly
understood. Notably, to the best of the authors' knowledge, no
previous studies have established a relationship between MAGEA1 and
cervical cancer. In the present study, the RNA-seq analysis
provided the first evidence that MAGEA1 may serve as a novel
biomarker not only for cervical cancer but also potentially for a
broad spectrum of types of cancer. To evaluate the potential of
MAGEA1 as a therapeutic target in cervical cancer, the present
study investigated the effects of shRNA-mediated MAGEA1 knockdown
on cell proliferation, migration, invasion and colony formation.
Consistent upregulation of MAGEA1 was observed in cervical cancer
cells and the present study demonstrated that MAGEA1 played a key
role in promoting cell migration, invasion, wound healing and
anchorage-independent growth, consistent with previous findings in
melanoma and breast cancer.
Notably, MAGEA1 knockdown markedly increased
sensitivity to chemotherapeutic agents, suggesting a potential
involvement in drug resistance mechanisms, implying that targeting
MAGEA1 could enhance the therapeutic efficacy of conventional
chemotherapies in cervical cancer treatment. Despite favorable
5-year survival rates >80% for early-stage cervical cancer,
clinical data indicates that in young women under 30 years,
high-risk HPV infections can rapidly progress to high-grade lesions
within two years (60).
Therefore, the identification and functional characterization of
host proteins interacting with HPV oncoproteins remains a
significant challenge in advancing cervical cancer therapy.
Understanding these interactions could provide critical insights
into cervical tumorgenesis and contribute to the development of
novel therapeutic strategies targeting these magnalities.
Nevertheless, translating these findings into
clinical applications will require further investigation to
optimize the delivery of MAGEA1-targeted agents and assess
potential off-target effects in complex physiological environments.
Future research should focus on clarifying the molecular
interactions between MAGEA1 and HPV oncoproteins, evaluating
synergistic effects with conventional therapies, and investigating
the role of MAGEA1 in immune evasion mechanisms. Such studies could
provide the basis for personalized immunotherapeutic strategies,
utilizing the immunogenic properties of MAGEA1 as a cancer-testis
antigen.
Supplementary Data
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author. All RNA-seq data generated
in this may be found in the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject)
under accession number PRJNA1362940 or at the following URL:
https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1362940.
Authors' contributions
Conceptualization was by AK, JK, WK, JWY and HY. AK,
JK, WK, KM, SC, MJ, JL, JWY and HY wrote the original draft. WK,
JWY and HY were responsible for writing, reviewing and editing. JWY
and HY confirm the authenticity of all the raw data. All authors
read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare they have no competing
interests.
Acknowledgements
The schematics of Figs. 2 and 5 were created by the authors using
images provided by BioRender (https://biorender.com/).
Funding
The present study was funded by the Brain Korea 21, grant no.
M2022B002600003, Korea Health Industry Development Institute, grant
no. RS-2025-02214206 and the Ministry of Food and Drug Safety in
Korea, grant nos. 22213MFDS421 and RS-2025-02213409.
References
|
1
|
Causin RL, Freitas AJA, Hidalgo Filho CMT,
Reis RD, Reis RM and Marques MMC: A systematic review of MicroRNAs
involved in cervical cancer progression. Cells. 10:6682021.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Plante M, Kwon JS, Ferguson S, Samouëlian
V, Ferron G, Maulard A, de Kroon C, Van Driel W, Tidy J, Williamson
K, et al: Simple versus radical hysterectomy in women with low-risk
cervical cancer. N Engl J Med. 390:819–829. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wang M, Huang K, Wong MCS, Huang J, Jin Y
and Zheng ZJ: Global cervical cancer incidence by histological
subtype and implications for screening methods. J Epidemiol Glob
Health. 14:94–101. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Cohen CM, Wentzensen N, Castle PE,
Schiffman M, Zuna R, Arend RC and Clarke MA: Racial and ethnic
disparities in cervical cancer incidence, survival, and mortality
by histologic subtype. J Clin Oncol. 41:1059–1068. 2023. View Article : Google Scholar
|
|
5
|
Yang ST, Wang PH, Liu HH, Chang CW, Chang
WH and Lee WL: Cervical cancer: Part II the landscape of treatment
for persistent, recurrent and metastatic diseases (I). Taiwan J
Obstet Gynecol. 63:637–650. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lai W, Liao J, Li X, Liang P, He L, Huang
K, Liang X and Wang Y: Characterization of the microenvironment in
different immune-metabolism subtypes of cervical cancer with
prognostic significance. Front Genet. 14:10676662023. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Craig AJ, Garcia-Lezana T, Ruiz de
Galarreta M, Villacorta-Martin C, Kozlova EG, Martins-Filho SN, von
Felden J, Ahsen ME, Bresnahan E, Hernandez-Meza G, et al:
Transcriptomic characterization of cancer-testis antigens
identifies MAGEA3 as a driver of tumor progression in
hepatocellular carcinoma. PLoS Genet. 17:e10095892021. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Vlasenkova R, Konysheva D, Nurgalieva A
and Kiyamova R: Characterization of cancer/testis antigens as
prognostic markers of ovarian cancer. Diagnostics (Basel). 13. pp.
30922023, View Article : Google Scholar
|
|
9
|
Chen L, Wu Q, Xu X, Yang C, You J, Chen F
and Zeng Y: Cancer/testis antigen LDHC promotes proliferation and
metastasis by activating the PI3K/Akt/GSK-3β-signaling pathway and
the in lung adenocarcinoma. Exp Cell Res. 398:1124142021.
View Article : Google Scholar
|
|
10
|
Danilova A, Misyurin V, Novik A, Girdyuk
D, Avdonkina N, Nekhaeva T, Emelyanova N, Pipia N, Misyurin A and
Baldueva I: Cancer/testis antigens expression during cultivation of
melanoma and soft tissue sarcoma cells. Clin Sarcoma Res. 10:32020.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Cui Z and Chen Y, Hu M, Lin Y, Zhang S,
Kong L and Chen Y: Diagnostic and prognostic value of the
cancer-testis antigen lactate dehydrogenase C4 in breast cancer.
Clin Chim Acta. 503:203–209. 2020. View Article : Google Scholar
|
|
12
|
Yu QY, Wang ZW, Zhou MY, Li SF and Liao
XH: MAGE-A3 regulates tumor stemness in gastric cancer through the
PI3K/AKT pathway. Aging (Albany NY). 14:9579–9598. 2022.PubMed/NCBI
|
|
13
|
Bai R and Yuan C: Kita-kyushu lung cancer
antigen-1 (KK-LC-1): A promising cancer testis antigen. Aging Dis.
13:1267–1277. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wei R, Dean DC, Thanindratarn P, Hornicek
FJ, Guo W and Duan Z: Cancer testis antigens in sarcoma:
Expression, function and immunotherapeutic application. Cancer
Lett. 479:54–60. 2020. View Article : Google Scholar
|
|
15
|
Wang Z, Liu Z and Wu S: Long non-coding
RNA CTA sensitizes osteosarcoma cells to doxorubicin through
inhibition of autophagy. Oncotarget. 8:31465–31477. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yang P, Meng M and Zhou Q: Oncogenic
cancer/testis antigens are a hallmarker of cancer and a sensible
target for cancer immunotherapy. Biochim Biophys Acta Rev Cancer.
1876:1885582021. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hikmet F, Rassy M, Backman M, Méar L,
Mattsson JSM, Djureinovic D, Botling J, Brunnström H, Micke P and
Lindskog C: Expression of cancer-testis antigens in the immune
microenvironment of non-small cell lung cancer. Mol Oncol.
17:2603–2617. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ren S, Zhang Z, Li M, Wang D, Guo R, Fang
X and Chen F: Cancer testis antigen subfamilies: Attractive targets
for therapeutic vaccine (review). Int J Oncol. 62:712023.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Weon JL and Potts PR: The MAGE protein
family and cancer. Curr Opin Cell Biol. 37:1–8. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yang SW, Huang X, Lin W, Min J, Miller DJ,
Mayasundari A, Rodrigues P, Griffith EC, Gee CT, Li L, et al:
Structural basis for substrate recognition and chemical inhibition
of oncogenic MAGE ubiquitin ligases. Nat Commun. 11:49312020.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Laduron S, Deplus R, Zhou S, Kholmanskikh
O, Godelaine D, De Smet C, Hayward SD, Fuks F, Boon T and De Plaen
E: MAGE-A1 interacts with adaptor SKIP and the deacetylase HDAC1 to
repress transcription. Nucleic Acids Res. 32:4340–4350. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang D, Wang J, Ding N, Li Y, Yang Y, Fang
X and Zhao H: MAGE-A1 promotes melanoma proliferation and migration
through C-JUN activation. Biochem Biophys Res Commun. 473:959–965.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Lian Y, Meng L, Ding P and Sang M:
Epigenetic regulation of MAGE family in human cancer
progression-DNA methylation, histone modification, and non-coding
RNAs. Clin Epigenetics. 10:1152018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Oh C, Kim HR, Oh S, Ko JY, Kim Y, Kang K,
Yang Y, Kim J, Park JH, Roe JS and Yoo KH: Epigenetic upregulation
of MAGE-A isoforms promotes breast cancer cell aggressiveness.
Cancers (Basel). 13. pp. 31762021, View Article : Google Scholar
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
26
|
Matentzoglu K and Scheffner M:
Ubiquitin-fusion protein system: A powerful tool for ectopic
protein expression in mammalian cells. Biotechniques. 46:212224–26.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tsai HC, Li YC, Hsu SH, Young TH and Chen
MH: Inhibition of growth and migration of oral and cervical cancer
cells by citrus polyphenol. J Formos Med Assoc. 115:171–185. 2016.
View Article : Google Scholar
|
|
28
|
Bolger AM, Lohse M and Usadel B:
Trimmomatic: A flexible trimmer for Illumina sequence data.
Bioinformatics. 30:2114–2120. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Dobin A, Davis CA, Schlesinger F, Drenkow
J, Zaleski C, Jha S, Batut P, Chaisson M and Gingeras TR: STAR:
Ultrafast universal RNA-seq aligner. Bioinformatics. 29:15–21.
2013. View Article : Google Scholar
|
|
30
|
Li H, Handsaker B, Wysoker A, Fennell T,
Ruan J, Homer N, Marth G, Abecasis G and Durbin R; 1000 Genome
Project Data Processing Subgroup: The sequence alignment/map format
and SAMtools. Bioinformatics. 25:2078–2079. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Liao Y, Smyth GK and Shi W: featureCounts:
An efficient general purpose program for assigning sequence reads
to genomic features. Bioinformatics. 30:923–930. 2014. View Article : Google Scholar
|
|
32
|
Chen Y, Chen L, Lun ATL, Baldoni PL and
Smyth GK: edgeR v4: Powerful differential analysis of sequencing
data with expanded functionality and improved support for small
counts and larger datasets. Nucleic Acids Res. 53:gkaf0182025.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wickham H: Data analysis In: ggplot2:
Elegant graphics for data analysis. Springer; pp. 189–201. 2016
|
|
34
|
Wickham H: ggplot2: Elegant graphics for
data analysis. 2nd edition. Springer; New York: 2009, View Article : Google Scholar
|
|
35
|
Subramanian A, Tamayo P, Mootha VK,
Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub
TR, Lander ES and Mesirov JP: Gene set enrichment analysis: A
knowledge-based approach for interpreting genome-wide expression
profiles. Proc Natl Acad Sci USA. 102:15545–15550. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Mootha VK, Lindgren CM, Eriksson KF,
Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E,
Ridderstråle M, Laurila E, et al: PGC-1alpha-responsive genes
involved in oxidative phosphorylation are coordinately
downregulated in human diabetes. Nat Genet. 34:267–273. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Liberzon A, Birger C, Thorvaldsdóttir H,
Ghandi M, Mesirov JP and Tamayo P: The molecular signatures
database (MSigDB) hallmark gene set collection. Cell Syst.
1:417–425. 2015. View Article : Google Scholar
|
|
38
|
Kolde R and Kolde MR: Package 'pheatmap’.
R Package. 1:7902015.
|
|
39
|
Neuwirth E: RColorBrewer: Colorbrewer
palettes. R Package Version. 1:1–3. 2014.
|
|
40
|
Sherman BT, Hao M, Qiu J, Jiao X, Baseler
MW, Lane HC, Imamichi T and Chang W: DAVID: A web server for
functional enrichment analysis and functional annotation of gene
lists (2021 update). Nucleic Acids Res. 50(W1): W216–W221. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Huang DW, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: Gene ontology: Tool for the unification of biology. The gene
ontology consortium Nat Genet. 25:25–29. 2000.
|
|
43
|
Gene Ontology Consortium; Aleksander SA,
Balhoff J, Carbon S, Cherry JM, Drabkin HJ, Ebert D, Feuermann M,
Gaudet P, Harris NL, et al: The gene ontology knowledgebase in
2023. Genetics. 224:iyad0312023. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Bindea G, Mlecnik B, Hackl H, Charoentong
P, Tosolini M, Kirilovsky A, Fridman WH, Pagès F, Trajanoski Z and
Galon J: ClueGO: A Cytoscape plug-in to decipher functionally
grouped gene ontology and pathway annotation networks.
Bioinformatics. 25:1091–1093. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Bindea G, Galon J and Mlecnik B: CluePedia
Cytoscape plugin: Pathway insights using integrated experimental
and in silico data. Bioinformatics. 29:661–663. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Imere A, Foster NC, Hajiali H, Okur KE,
Wright AL, Barroso IA and Haj AJE: Enhanced chondrogenic potential
in GelMA-based 3D cartilage model via Wnt3a surface immobilization.
Sci Rep. 14:150222024. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Qiao JX, Guo DY, Tian H, Wang ZP, Fan QQ,
Tian Y, Sun J, Zhang XF, Zou JB, Cheng JX, et al: Research progress
of paclitaxel nanodrug delivery system in the treatment of
triple-negative breast cancer. Mater Today Bio. 29:1013582024.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Rajasekaran V, Harris BT, Osborn RT,
Smillie C, Donnelly K, Bacou M, Esiri-Bloom E, Ooi LY, Allan M,
Walker M, et al: Genetic variation at 11q23.1 confers colorectal
cancer risk by dysregulation of colonic tuft cell transcriptional
activator POU2AF2. Gut. 74:787–803. 2025. View Article : Google Scholar :
|
|
49
|
Goto H, Kariya R, Kudo E, Katano H and
Okada S: PAX5 functions as a tumor suppressor by RB-E2F-mediated
cell cycle arrest in Kaposi sarcoma-associated herpesvirus-infected
primary effusion lymphoma. Neoplasia. 56:1010352024. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Bloomstein JD, von Eyben R, Chan A, Rankin
EB, Fregoso DR, Wang-Chiang J, Lee L, Xie LX, David SM, Stehr H, et
al: Validated limited gene predictor for cervical cancer lymph node
metastases. Oncotarget. 11:2302–2309. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
He Z, Wang X, Yang Z, Jiang Y, Li L, Wang
X, Song Z, Wang X, Wan J, Jiang S, et al: Expression and prognosis
of CDC45 in cervical cancer based on the GEO database. PeerJ.
9:e121142021. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Yi E, Chang JE, Leem C, Jeon CH and Jheon
S: Association of MAGE A1-6 expression with lung cancer
progression. J Cancer. 8:1324–1329. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Lian Y, Sang M, Gu L, Liu F, Yin D, Liu S,
Huang W, Wu Y and Shan B: MAGE-A family is involved in gastric
cancer progression and indicates poor prognosis of gastric cancer
patients. Pathol Res Pract. 213:943–948. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Almutairi MH, Alotaibi MM, Alonaizan R,
Alrefaei AF and Almutairi BO: Identification of MAGE-A family genes
in colon cancer patients and their expression mechanism. J King
Saud Univ Sci. 34:1022512022. View Article : Google Scholar
|
|
55
|
Mi Y, Liu F, Liang X, Liu S, Huang X, Sang
M and Geng C: Tumor suppressor let-7a inhibits breast cancer cell
proliferation, migration and invasion by targeting MAGE-A1.
Neoplasma. 66:54–62. 2019. View Article : Google Scholar
|
|
56
|
Suyama T, Ohashi H, Nagai H, Hatano S,
Asano H, Murate T, Saito H and Kinoshita T: The MAGE-A1 gene
expression is not determined solely by methylation status of the
promoter region in hematological malignancies. Leuk Res.
26:1113–1118. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Jin S, Cao S, Li J, Meng Q, Wang C, Yao L,
Lang Y, Cao J, Shen J, Pan B, et al: Cancer/testis antigens (CTAs)
expression in resected lung cancer. Onco Targets Ther.
11:4491–4499. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Tarnowski M, Czerewaty M, Deskur A,
Safranow K, Marlicz W, Urasińska E, Ratajczak MZ and Starzyńska T:
Expression of cancer testis antigens in colorectal cancer: New
prognostic and therapeutic implications. Dis Markers.
2016:19875052016. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Fanipakdel A, Seilanian Toussi M,
Rezazadeh F, Mohamadian Roshan N and Javadinia SA: Overexpression
of cancer-testis antigen melanoma-associated antigen A1 in lung
cancer: A novel biomarker for prognosis, and a possible target for
immunotherapy. J Cell Physiol. 234:12080–12086. 2019. View Article : Google Scholar
|
|
60
|
Lei J, Ploner A, Elfström KM, Wang J, Roth
A, Fang F, Sundström K, Dillner J and Sparén P: HPV vaccination and
the risk of invasive cervical cancer. N Engl J Med. 383:1340–1348.
2020. View Article : Google Scholar : PubMed/NCBI
|