Introduction
Breast cancer (BC) is one of the most prevalent
malignancies and a leading cause of cancer-related death worldwide,
with incidence rates continuing to rise (1,2).
Current treatment strategies, including surgery, chemotherapy and
radiotherapy (3), have markedly
improved patient outcomes. However, individuals with more
aggressive or advanced disease often develop resistance to these
conventional therapies (4). This
underscores an urgent need to identify novel therapeutic targets to
overcome radio-resistance in BC.
Tumor-associated macrophages, known for their
functional diversity and plasticity, are key components of the BC
tumor microenvironment and play crucial roles in disease
progression and therapeutic response (5). Under different conditions,
macrophages can polarize into either the pro-inflammatory,
anti-tumorigenic M1 phenotype or the anti-inflammatory,
pro-tumorigenic M2 phenotype. M1 macrophages mediate anti-tumor
immunity, whereas M2 macrophages facilitate tissue remodeling and
tumor progression (6). Notably,
the suppression of M2 macrophage polarization has been shown to
mitigate radio-resistance in inflammatory BC (7).
Calmodulin-binding transcriptional activators
(CAMTAs) are an evolutionarily conserved family of transcription
regulatory genes (8). Among them,
CAMTA1 is predominantly expressed in neuronal tissues (9). Reduced CAMTA1 expression is
associated with unfavorable tumor biomarkers and poor prognosis in
several cancers (10). Emerging
evidence suggests that CAMTA1 plays an important role in tumor
progression. For instance, CAMTA1 expression is reduced in
colorectal cancer (CRC) tissues and the silence of CAMTA1 renders
CRC cells resistant to oxaliplatin treatment (11). Notably, CAMTA1 expression appears
to be elevated in BC tissues, yet its functional role in BC
progression remains poorly defined.
Neuregulin 1 (NRG1), a member of the neuregulin
family, is widely recognized as an oncogene in multiple cancers and
plays critical roles in tumor progression (12,13). For example, NRG1 elevation is
associated with aggressive clinical manifestations and acts as a
prognostic biomarker in gastric cancer (14). Importantly, increased NRG1
expression has also been reported in BC tissues and cells (15). Bioinformatics analyses reveal a
positive correlation between CAMTA1 and NRG1 expression in BC
tissues. According to the analysis on GEPIA database (http://gepia2.cancer-pku.cn/), NRG1 expression was
positively associated with the M2 macrophage marker CD163. These
findings led the present study to propose a novel regulatory
mechanism whereby CAMTA1 promotes radio-resistance by facilitating
M2 macrophage polarization through NRG1 signaling. This
CAMTA1-NRG1-M2 axis, to the best of the authors' knowledge, has not
been previously reported and represents a key novelty of the
present study.
Accordingly, the present study aimed to investigate
the role of CAMTA1 in BC radio-resistance and to explore whether
CAMTA1 regulates M2 macrophage polarization via NRG1, thereby
providing a potential therapeutic target for improving BC treatment
outcomes.
Material and methods
Cell culture and hypoxia treatment
The human BC cell line MDA-MB-231 (cat. no.
iCell-h133), the mouse BC 4T1 cells (cat. no. iCell-m067), mouse
RAW264.7 macrophages (cat. no. iCell-m047) and human monocytic
leukemia THP-1 cells (cat. no. iCell-h213) were obtained from iCell
Bioscience Inc. All cells were cultured in RPMI-1640 medium
supplemented with 10% FBS (Gibco; Thermo Fisher Scientific, Inc.;
cat. no. 10099-14) and 1% penicillin-streptomycin at 37°C with 5%
CO2. THP-1 cells were differentiated into macrophages
through the treatment with 100 ng/ml PMA (16). To induce M2 macrophage
polarization, THP-1-derived macrophages were stimulated with 20
ng/ml IL-4 for 36 h (17).
For co-culture experiments, RAW264.7 macrophages and
4T1 cells were seeded into a Transwell chamber with 0.4 μm
pore. For next-generation RNA sequencing analysis, RAW264.7
macrophages were subjected to high-energy X-ray irradiation.
For hypoxic treatment and exosome isolation,
MDA-MB-231 cells were incubated in a tri-gas incubator of 1%
O2, 5% CO2 and 94% N2 with
contained exosome-depleted FBS medium for 12 or 24 h, followed by
co-culture with THP-1-derived macrophages for 24 h (18). To inhibit exosome biogenesis,
MDA-MB-231 cells were pretreated with 10 μM GW4869 prior to
further experimentation (19). To
further confirm the role of NRG1 in modulating M2 polarization, an
anti-NRG1 blocking antibody (10 μg/ml; Ab-2, LabVision) was
used for cell treatment (20).
Cell transfection
The pc-DNA3.1 vectors containing the complete
sequence of NRG1 (Ov-NRG1), an empty vector (Ov-NC), the small
interfering RNA targeting NRG1 (si-NRG1) and the corresponding
negative control (si-NC) were purchased from Shanghai GenePharma
Co., Ltd. and transfected into THP-1-derived macrophages. The
sequences were: si-NRG1#1: 5'-CCCGATTGAAAGAGATGAAAAGC-3';
si-NRG1#2: 5'-TGGGAATGAATTGAATCGAAAAA-3'; si-NRG1#3:
5'-TGGCTGATTCTGGAGAGTATATG-3'; si-NC:
5'-AAGACAUUGUGUGUCCGCCTT-3'.
The pc-DNA3.1 vectors containing the complete
sequence of CAMTA1 (Ov-CAMTA1), the empty vector (Ov-NC), the short
hairpin RNA targeting CAMTA1 (sh-CAMTA1) and the corresponding
negative control (sh-NC) were obtained from Shanghai GenePharma
Co., Ltd. and transfected into MDA-MB-231 cells. The target
sequences were: sh-CAMTA1#1: 5'-TGAGGAAATTGCGGCTTATTT-3';
sh-CAMTA1#2: 5'-CCCGACTGTTTCCTCAATAAT-3'; sh-CAMTA1#3:
5'-TCGGTCTGAACCCTCTAATTA-3'; sh-NC:
5'-GGAATCTCATTCGATGCATAC-3'.
Cell transfection was performed using
Lipofectamine® 2000 reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's instructions.
Exosome isolation and fluorescent
labeling
Exosomes were isolated using a standard differential
centrifugation protocol. Initially, the conditioned medium was
centrifuged at 300 × g for 10 min at 4°C to remove cell debris,
2,000 × g for 20 min at 4°C to remove the dead cells and 30 min at
10,000 × g at 4°C to further remove cell debris (21). The pellet of exosomes was
collected through the ultracentrifugation at 100,000 × g for 70 min
at 4°C (22). The exosome
particle size was determined by nanoparticle tracking analysis
using ZetaView PMX 110 (Particle Metrix).
For fluorescent labeling, purified exosomes were
stained with PKH26 red fluorescent labeling kits (MilliporeSigma)
according to the manufacturer's instructions. Briefly,
ultracentrifuged exosomes were re-suspended in diluent C (100
μl) after ultracentrifugation at 100,000 × g for 70 min at
4°C and then incubated with PKH26 dye solution (100 μl) at
room temperature for 5 min, followed by the addition of 200
μl serum. Subsequently, the labeled exosomes were washed by
PBS at 4°C and then incubated with MDA-MB-231 cells. Fluorescence
microscopy was used to visualize exosome uptake.
Transmission electron microscopy
Exosomes were fixed with 2.5% glutaraldehyde at 4°C,
placed onto the copper grids and negatively stained with 2%
phosphotungstic acid for 2 min at room temperature. After
air-drying, the samples were observed using a transmission electron
microscope. Images were analyzed using ImageJ software (version
1.53, National Institutes of Health).
Reverse transcription-quantitative (RT-q)
PCR
Total RNA was extracted from sample cells seeded at
a density of 1×106 cells per well using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.) strictly according to the manufacturer's protocol. 1
μg of RNA was reverse-transcribed into cDNA using a
commercial RevertAid cDNA Synthesis kit (Thermo Fisher Scientific,
Inc.) according to the manufacturer's instructions. The reverse
transcription was performed at 42°C for 60 min, followed by 70°C
for 5 min to inactivate the reverse transcriptase. qPCR was
conducted using SYBR Green PCR Master Mix on the 7500 Fast
Real-time PCR system according to the manufacturer's instructions.
Thermocycling conditions were as follows: Initial denaturation at
95°C for 2 min. Denaturation at 94°C for 15 sec, annealing and
extension at 60°C for 30 sec, 40 cycles. The relative gene
expression was determined using 2−ΔΔCq method (23). GAPDH was used as the internal
reference gene. All RT-qPCR experiments were performed with three
independent biological replicates. The following were the primer
sequences: NRG1 (human) forward, 5'-GATTCCTACCGAGACTCTCCTC-3' and
reverse, 5'-TGGAAGGCATGGACACCGTCAT-3'; NRG1 (mouse) forward,
5'-GCTCATCACTCCACGACTGTCA-3' and reverse,
5'-TGCCTGCTGTTCTCTACCGATG-3'; CAMTA1 (human) forward,
5'-AGTGCAGAAAATGAAGAATGCG-3' and reverse
5'-CAAAATTCTCCTGCTTGATTCG-3'; CAMTA1 (mouse) forward,
5'-CGGTGGTGTTTGAGTACAAGGC-3' and reverse
5'-CCTCCTTTCCATCTGCTCCAGA-3'; GAPDH (human) forward,
5'-GAAGGTGAAGGTCGGAGTC-3' and reverse, 5'-GAAGATGGTGATGGGATTTC-3',
or GAPDH (mouse) forward, 5'-AGGTCGGTGTGAACGGATTTG-3' and reverse,
5'-TGTAGACCATGTAGTTGAGGTCA-3'.
Immunoblotting analysis
The total proteins were extracted from sample cells
using RIPA lysis buffer (Beyotime Biotechnology, China) and the
protein concentration was quantified using BCA assay kits (Thermo
Fisher Scientific, Inc.) according to the manufacturer's
instructions. Separated by 10% SDS-PAGE, equal amounts of protein
(30 μg per lane) were transferred to PVDF membranes
(MilliporeSigma). Blocked by 5% BSA (MilliporeSigma) for 1 h at
room temperature, the membranes were incubated with primary
antibodies against NRG1 (cat. no. 10527-1-AP; 1:1,000; Proteintech
Group, Inc.), hypoxia-inducible factor-1α (HIF-1α; cat. no.
20960-1-AP; 1:2,000; Proteintech Group, Inc.), CD63 (cat. no.
25682-1-AP; 1:2,000; Proteintech Group, Inc.), CD81 (cat. no.
27855-1-AP; 1:1,000; Proteintech Group, Inc.), CD9 (cat. no.
20597-1-AP; 1:2,000; Proteintech Group, Inc.), ALIX (cat. no.
12422-1-AP; 1:5,000; Proteintech Group, Inc.) and GAPDH (cat. no.
10494-1-AP; 1:5,000; Proteintech Group, Inc.) overnight at 4°C. On
the next day, the membranes were incubated with goat anti-rabbit
horseradish peroxidase (HRP)-conjugated secondary antibodies (cat.
no. SA00001-2; Proteintech Group, Inc.) at room temperature for 3
h. The protein bands were visualized using ECL regent (Thermo
Fisher Scientific, Inc.) and protein density was semi-quantified
via ImageJ software (version 1.53; National Institutes of Health,
USA).
Migration assay
For the Transwell migration assay, MDA-MB-231 cells
were suspended in serum-free medium (200 μl) and seeded into
the upper chamber, while THP-1-derived macrophages were cultured in
medium (800 μl) containing 10% FBS in the lower chamber.
After 24 h of co-culture, the non-migrated cells were gently
removed while the migrated cells were stained by 0.1% crystal
violet for 30 min. The migrated cells in five randomly selected
non-overlapping fields were observed under a light microscope.
Detection of M2 macrophage marker
The THP-1 macrophages were suspended in pre-cooled
PBS and then centrifuged at 200 × g for 5 min at 4°C (24). After discarding the supernatant,
cells were fixed with 0.2 ml of fixation buffer for 5 min at room
temperature and then 0.5 ml of the permeabilization wash buffer was
added for 5 min at room temperature. Subsequently, the cells were
incubated with CD163 antibody (cat. no. 16627; 1:50; Cell Signaling
Technology, Inc.) at room temperature for 30 min. A BD FACSCanto II
flow cytometer (BD Biosciences) was used to assess CD163 expression
and the data analysis was processed using FlowJo software (version
10.8.1; FlowJo LLC; BD Biosciences).
Flow cytometry analysis of the cell cycle
and apoptosis
The MDA-MB-231 cells were washed by PBS and
centrifuged to remove the supernatant (25). For cell cycle analysis, MDA-MB-231
cells were treated with 50 μl RNase A and 200 μl PI
and incubated in the dark at room temperature for 20 min, followed
by flow cytometry analysis. For apoptosis analysis, MDA-MB-231
cells were suspended in 500 μl PBS and then incubated with 5
μl Annexin-V-FITC and 5 μl PI in the dark at room
temperature for 15 min, followed by flow cytometry analysis. All
samples were analyzed using a BD FACSCanto II flow cytometer (BD
Biosciences). Data acquisition and analysis were performed using
FlowJo software (version 10.8.1; FlowJo LLC; BD Biosciences). The
apoptotic rate was calculated as follows: Apoptotic rate (%)=early
apoptotic cells (Annexin V+/PI−) + late
apoptotic cells (Annexin V+/PI+).
Colony formation assay and MTT assay
Cell viability was detected using MTT assay and
colony formation assay. For MTT detection, cells were incubated
with MTT solution, followed by the aspiration of supernatant and
the addition of DMSO (100 μl). The optical density was
measured at 490 nm using a microplate reader.
For colony formation assays, the colonies were fixed
with methanol at room temperature for 15 min and stained by 0.1%
crystal violet at room temperature for 20 min. The number of
colonies was counted using under a light microscope (magnification,
×100). Five randomly selected, non-overlapping fields per well were
examined.
ELISA
The supernatant was collected by centrifugation at
2,000 × g for 5 min at 4°C. The concentrations of IL-10 and IL-12
were detected using ELISA-related IL-10 assay kits and IL-12 assay
kits according to the manufacturer's instructions. The optical
density was measured at 450 nm using a microplate reader.
Xenograft tumor model
All animal experiments approved by the Animal Ethics
Committee of Jiangsu Cancer Hospital and Jiangsu Institute of
Cancer Research and The Affiliated Cancer Hospital of Nanjing
Medical University and conducted in compliance with the National
Institutes of Health Guide for the Care and Use of Laboratory
Animals (approval no. IACUC-20210216-01). A total of 50 female
BALB/c nude mice (4-5 weeks old, weighing 18-22 g) were obtained
from Beijing Vital River Laboratory Animal Technology Co., Ltd..
Mice were housed under SPF conditions at a controlled temperature
of 22±2°C, relative humidity of 50-60%, and a 12-h light/dark
cycle, with free access to food and water. The mice were
subcutaneously injected with 1×106 MDA-MB-231 cells
transfected with sh-CAMTA1 or Ov-CAMTA1 and then randomly divided
into five groups (n=10): PBS, Normoxic exo, Hypoxic exo, Hypoxic
exo + Ov-CAMTA1 and Hypoxic exo + sh-CAMTA1 groups. Mice in PBS
group received 30 μl PBS, while mice in remaining groups
were treated with 30 μg exosomes. Local radiotherapy in the
tumors was performed for consecutive 5 days (2 Gy/d). The tumor
growth was monitored every week. After five weeks, the mice were
sacrificed by intraperitoneal injection of pentobarbital sodium
(150 mg/kg). Death was confirmed based on the disappearance of pain
response, no response when pressing the toes with hands or forceps
and observation of cardiac and respiratory arrest. Tumor tissues
were removed, images captured and collected for histological and
immunohistochemistry analysis.
Histological analysis
The collected tumor issues were fixed with 10%
formalin at room temperature for 24 h. After fixation, tissues were
dehydrated through a graded ethanol series, cleared in xylene, and
subsequently embedded in paraffin according to standard protocol.
Paraffin-embedded tissues were then sectioned into 7
μm-thick slices. Subsequently, the slices were stained with
H&E at room temperature, including hematoxylin staining for 5
min followed by eosin staining for 2 min. The stained sections were
observed under a light microscope.
Immunohistochemistry analysis
Immunohistochemistry was performed to assess the
expression of NRG1, CD163, and cleaved caspase-3 in tumor tissues.
Briefly, antigen retrieval was performed by heating the tissue
samples in 1X sodium citrate buffer using a microwave, followed by
the incubation with 3% hydrogen peroxide for 10 min and subsequent
inhibition with 3% normal goat serum (Solarbio, China) for 1 h at
room temperature (26). The
slices were then incubated with primary antibody against NRG1 (cat.
no. 83323-6-RR; 1:500; Proteintech Group, Inc.), CD163 (cat. no.
83285-4-RR; 1:2,000; Proteintech Group, Inc.) and Caspase-3 (cat.
no. 19677-1-AP; 1:500; Proteintech Group, Inc.) overnight at 4°C.
On the next day, the slices were incubated with HRP-labeled
secondary antibody (cat. no. ab6721; 1:1,000; Abcam) for 1 h at
room temperature. Following the treatment with DAB solution, the
counterstaining to hematoxylin was performed for 1 min at room
temperature and a light microscope was used for observation.
Public database validation
To explore the expressions of CAMTA1 and NRG1 in BC
and validate their association with macrophage polarization,
several public databases were used. Gene expression levels of
CAMTA1 and NRG1 in The Cancer Genome Atlas-Breast Invasive
Carcinoma (TCGA-BRCA) samples were obtained from the UCSC Xena
platform (https://xenabrowser.net/). Spearman's
correlation between CAMTA1 and NRG1 was analyzed using R software
(v4.2.2; https://www.r-project.org/). The
correlation between NRG1 expression and M2 macrophage marker CD163
was assessed via the Gene Expression Profiling Interactive Analysis
(GEPIA)2 platform (http://gepia2.cancer-pku.cn/) based on TCGA-BRCA tumor
data. Kaplan-Meier survival analysis was performed using the KMplot
database (https://kmplot.com) with the overall
survival in Affymetrix breast cancer cohorts stratified by NRG1
expression.
Statistical analysis
The collected data was processed using GraphPad
Prism 8 software (Dotmatics) and presented as mean ± SD.
Statistical comparisons between two groups were performed using
unpaired Student's t-test (for two groups) while comparisons among
three or more groups were conducted using one-way ANOVA with
Tukey's test. P<0.05 was considered to indicate a statistically
significant difference.
Results
NRG1 expression is elevated in RAW264.7
macrophages co-cultured with 4T1 cells
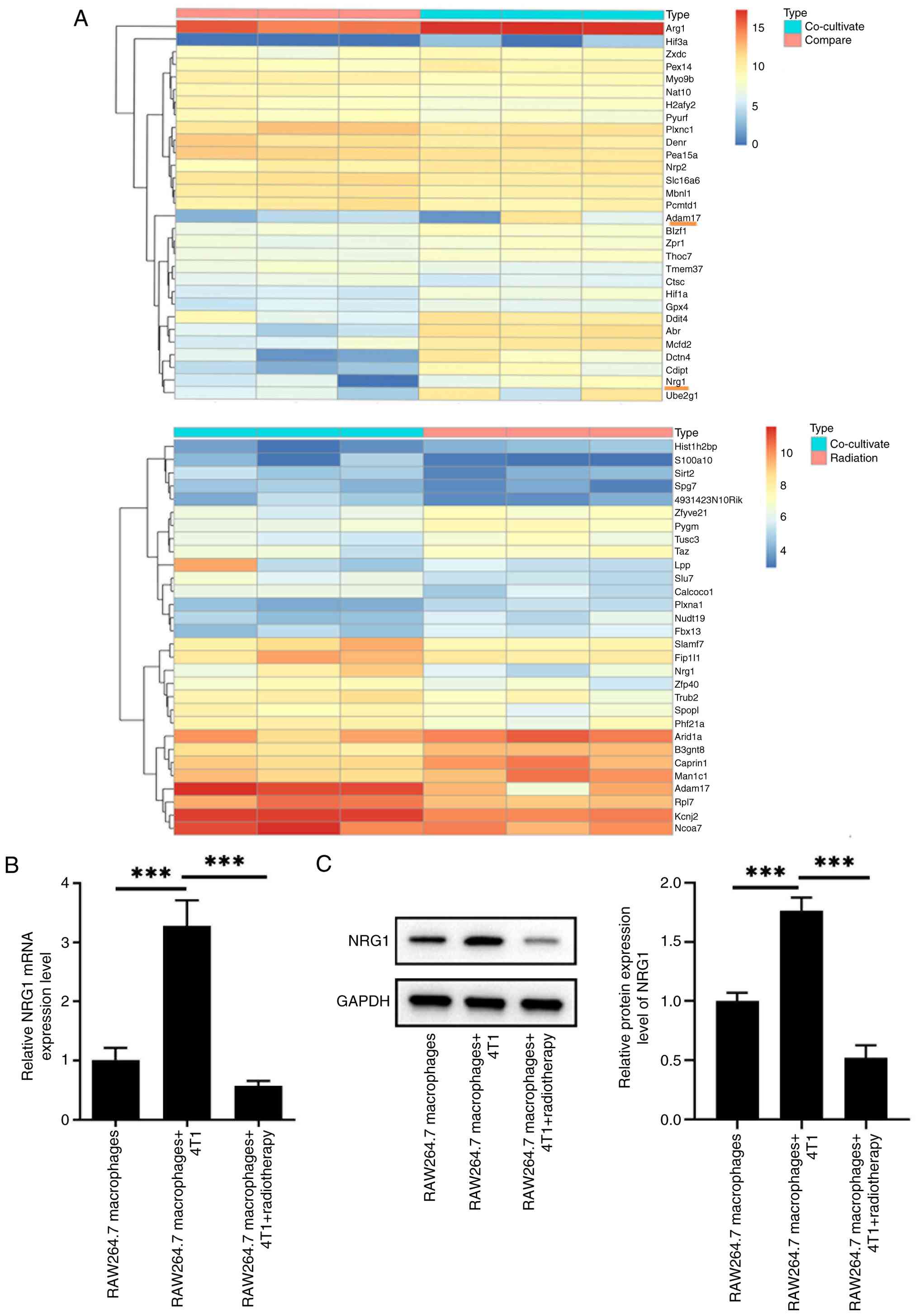
Through next-generation RNA sequencing analysis,
NRG1 expression was found to be increased in RAW264.7 macrophages
co-cultured with 4T1 cells. By contrast, radiotherapy treatment
markedly suppressed NRG1 expression (Fig. 1A). These findings were further
confirmed by RT-qPCR and immunoblotting analysis. Compared with the
RAW264.7 macrophages group, both the mRNA and protein expressions
of NRG1 in RAW264.7 macrophages were markedly increased following
the co-culture with 4T1 cells, whereas radiotherapy treatment
effectively reversed these changes (Fig. 1B and C).
Hypoxic MDA-MB-231 cells induce the M2
polarization of THP-1 macrophages
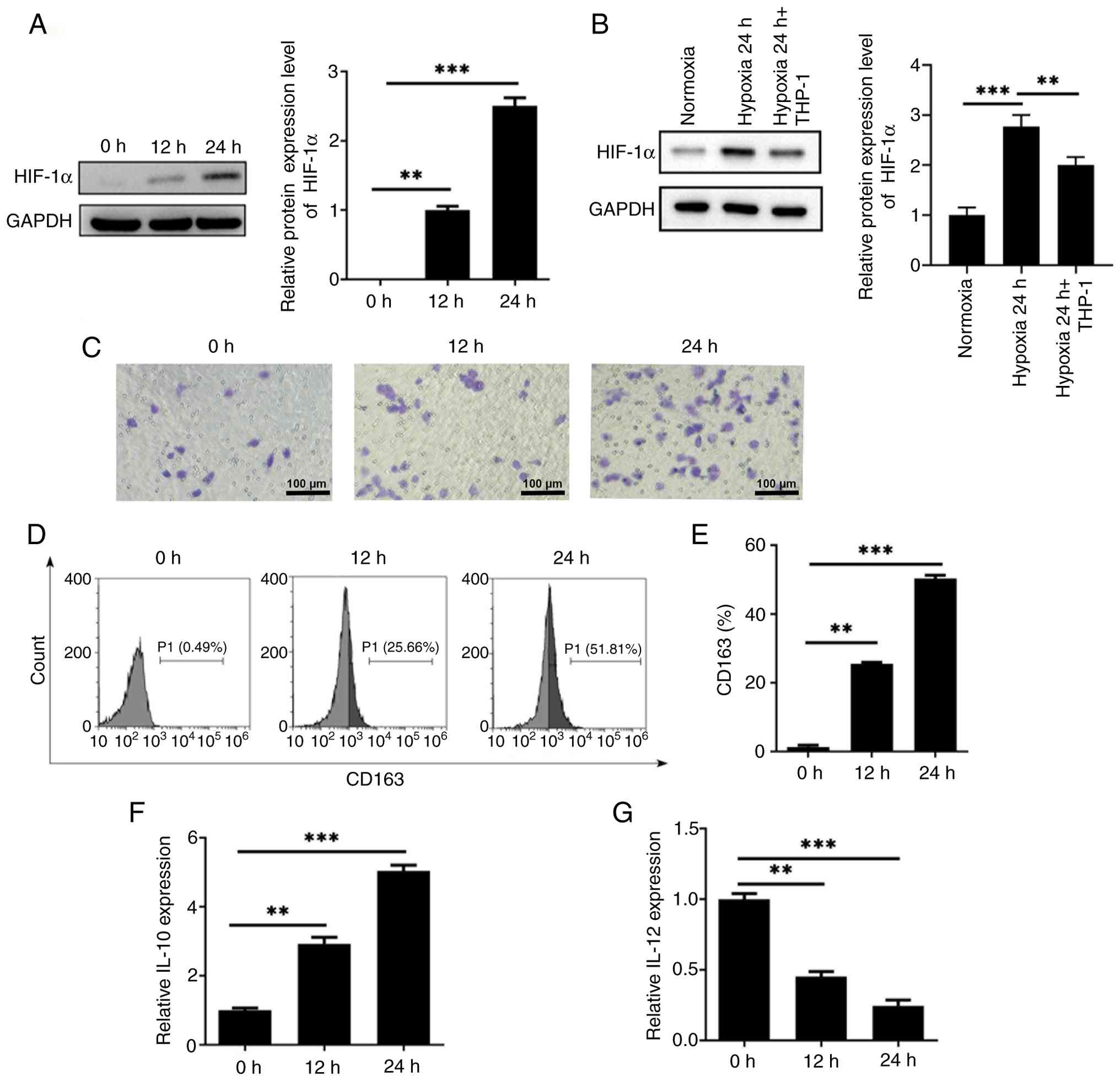
Under hypoxic conditions, the protein expression of
HIF-1α in MDA-MB-231 cells was assessed using immunoblotting
analysis. As shown in Fig. 2A,
HIF-1α protein level progressively increased with prolonged hypoxia
exposure (0, 12 and 24 h). Based on this observation, MDA-MB-231
cells pre-treated with hypoxia for 24 h were subsequently
co-cultured with THP-1-derived macrophages for another 24 h.
Following co-culture, the increased HIF-1α expression induced by
hypoxia was markedly decreased, yet remained above normoxic control
(Fig. 2B). Transwell assays
demonstrated that the migration of THP-1 macrophages was markedly
increased after co-culture with hypoxia-pretreated MDA-MB-231 cells
(Fig. 2C). Flow cytometry
analysis further showed that hypoxia stimulation increased the
expression of CD163, a well-established M2 macrophage marker, in a
time-dependent manner (Fig. 2D and
E). IL-12 is a cytokine associated with the M1 phenotype, while
IL-10 is indicative of M2 polarization. In addition, ELISA results
revealed that hypoxia promoted IL-10 secretion while concomitantly
reducing IL-12 production in THP-1 macrophages (Fig. 2F and G). Taken together, these
findings suggested that hypoxic MDA-MB-231 cells could facilitate
the M2 polarization of THP-1-derived macrophages.
M2 polarization enhances the resistance
of MDA-MB-231 cells to radiotherapy
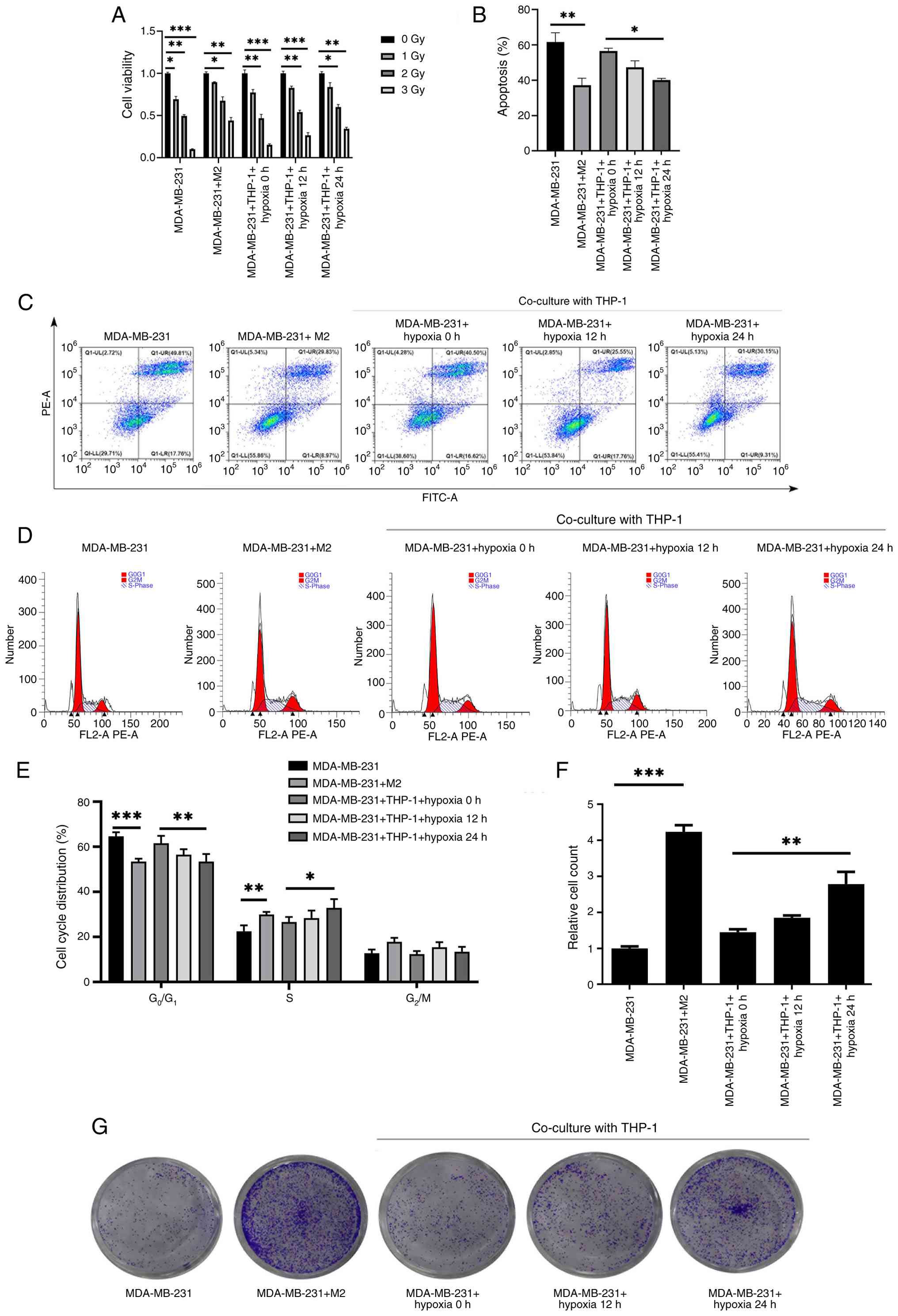
To investigate the radio-resistance of MDA-MB-231
cells, all cells were exposed to X-ray irradiation at doses of 1, 2
and 3 Gy. Given that hypoxic conditions can induce the
differentiation of M0 macrophages into M2 macrophages, MDA-MB-231
cells were co-cultured with THP-1 macrophages under hypoxia for
varying durations. The cells were divided into MDA-MB-231,
MDA-MB-231 + M2, MDA-MB-231 + THP-1 + hypoxia 0 h, MDA-MB-231 +
THP-1 + hypoxia 12 h and MDA-MB-231 + THP-1 + hypoxia 24 h groups.
MDA-MB-231 cell viability was first assessed using the MTT assay.
Compared with the 0 Gy group, MDA-MB-231 cell viability was reduced
with increasing radiation dose, whereas co-culture with M2
macrophages conferred the strongest radiation resistance (Fig. 3A). Based on these findings,
subsequent experiments were conducted using 2 Gy radiation. Flow
cytometry revealed that M2 polarization markedly inhibited the
apoptosis of MDA-MB-231 cells. Moreover, hypoxic co-culture with
THP-1 macrophages suppressed apoptosis in a time-dependent manner
compared with the MDA-MB-231 + THP-1 + hypoxia 0 h group (Fig. 3B and C). To assess whether the
reduced apoptosis was associated with altered cell proliferation,
cell-cycle distribution was analyzed by flow cytometry. The results
showed that M2 polarization markedly increased the proportion of
cells in S phase while decreasing the proportion in
G0/G1 phase, suggesting accelerated DNA
synthesis and proliferation potential. Similarly, hypoxic
co-culture with THP-1 macrophages increased the S-phase fraction in
a time-dependent manner (Fig. 3D and
E). In addition, colony formation assays demonstrated that M2
polarization greatly increased the number of colonies compared with
the MDA-MB-231 group and hypoxic co-culture with THP-1 macrophages
further promoted clonogenic survival (Fig. 3F-G).
NRG1 expression is associated with M2
polarization and poor prognosis
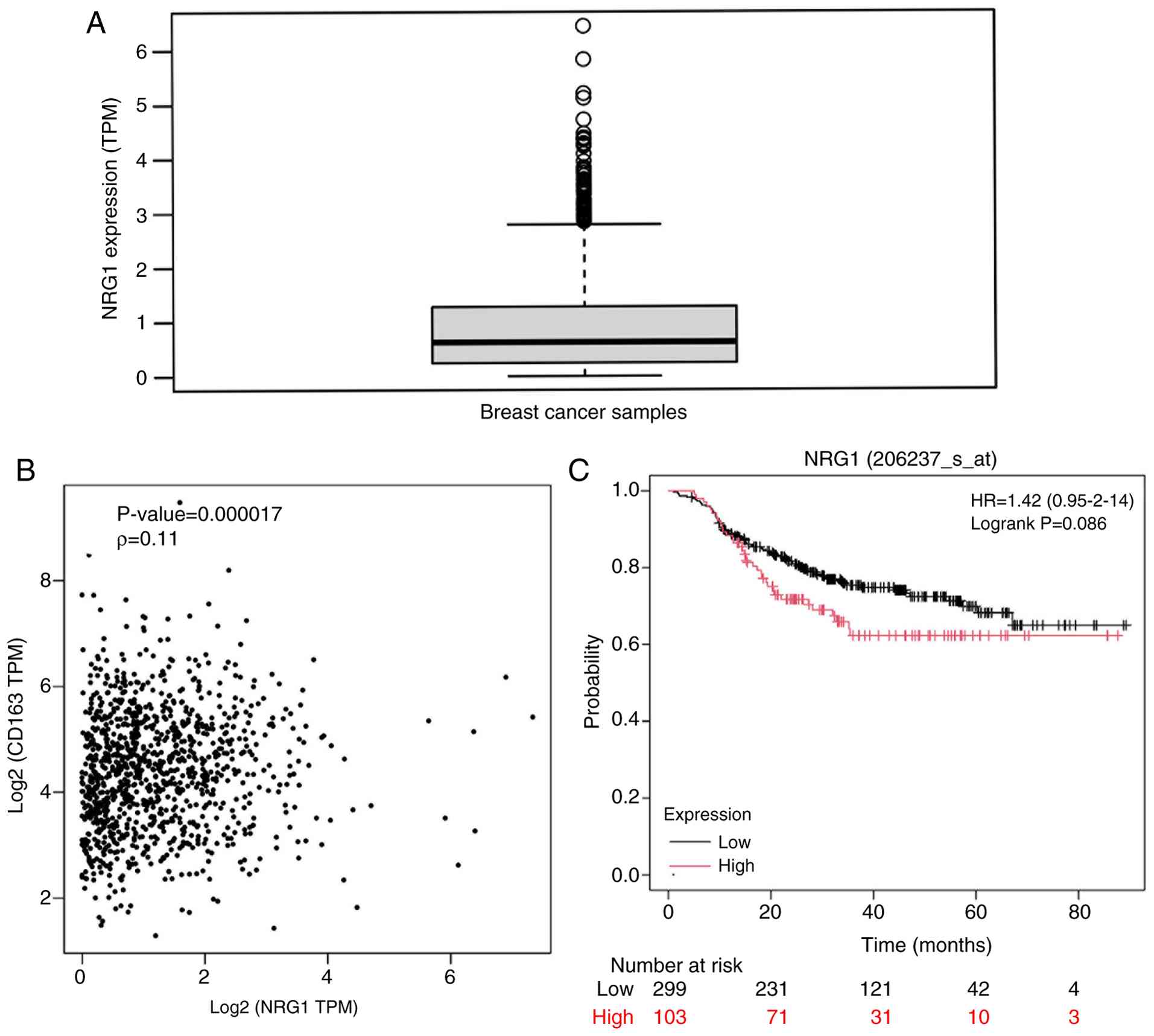
Analysis of the TCGA database revealed that NRG1
expression was markedly upregulated in BC samples (Fig. 4A). GEPIA2 database analysis
further demonstrated a positive correlation between NRG1 expression
and the M2 macrophage marker CD163 (Fig. 4B). Moreover, Kaplan-Meier survival
analysis showed that high NRG1 expression was associated with low
overall survival in BC patients (Fig.
4C).
Hypoxic MDA-MB-231 cells induces the M2
polarization of THP-1 macrophages via NRG1
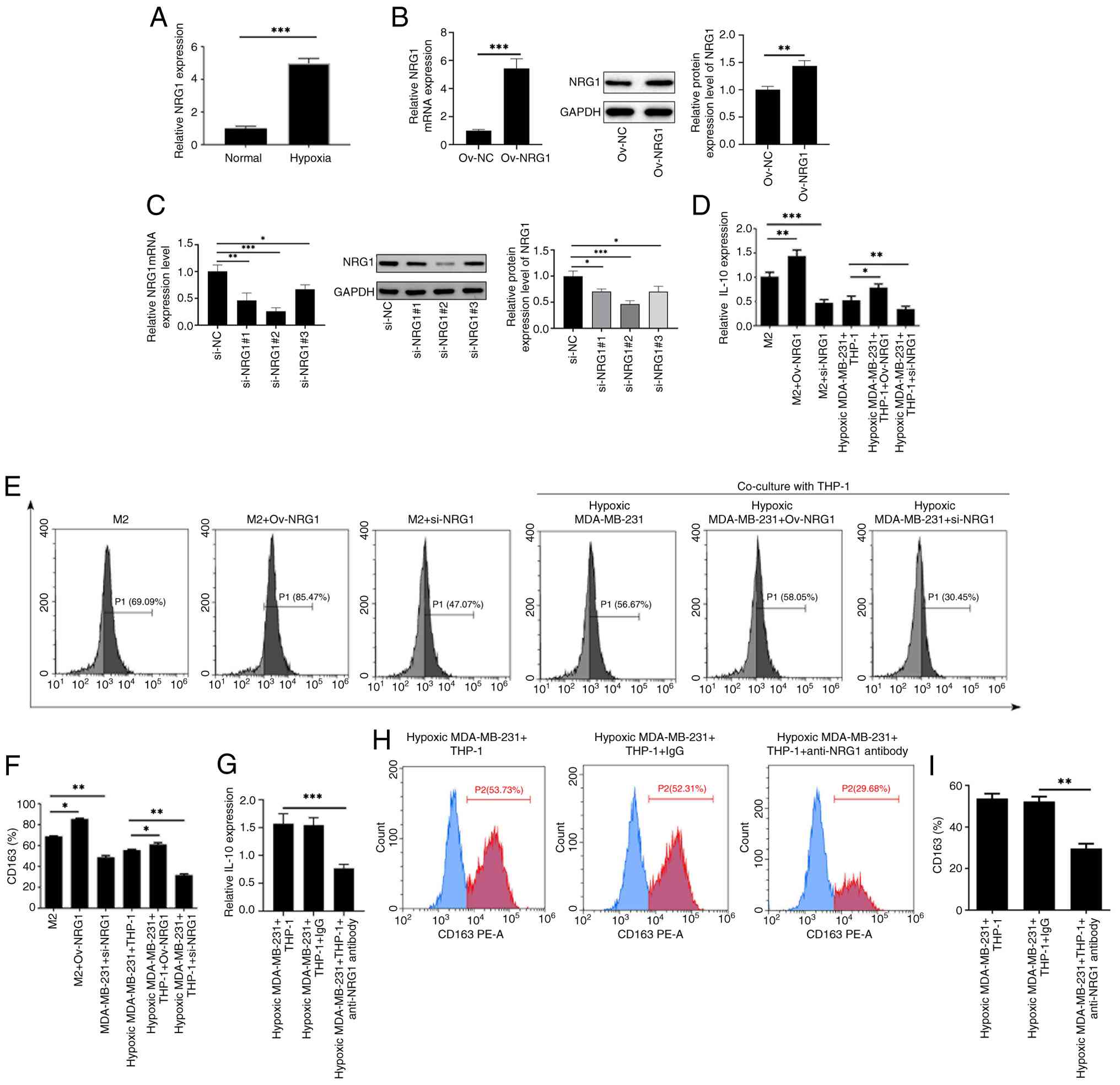
To assess the effects of hypoxia on NRG1 expression,
MDA-MB-231 cells were exposed to hypoxia for 24 h and co-cultured
with THP-1 macrophages. RT-qPCR analysis revealed that hypoxia
markedly upregulated NRG1expression compared with the Normoxia
group (Fig. 5A). To upregulate or
downregulate NRG1 expression, Ov-NRG1 or si-NRG1 was transfected
into cells and the transfection efficacy was examined using RT-qPCR
and immunoblotting analysis (Fig. 5B
and C). si-NRG1#2 (hereinafter referred as si-NRG1) was
selected for subsequent experiments due to its more prominent
transfection efficacy. Then, the cells were assigned into M2, M2 +
Ov-NRG1, M2 + si-NRG1, Hypoxic MDA-MB-231 + THP-1, Hypoxic
MDA-MB-231 + THP-1 + Ov-NRG1 and Hypoxic MDA-MB-231 + THP-1 +
si-NRG1 groups. ELISA results showed that Ov-NRG1 elevated IL-10
secretion, whereas si-NRG1 reduced IL-10 level compared with the M2
or Hypoxic MDA-MB-231 + THP-1 groups (Fig. 5D). Consistently, flow cytometry
analysis demonstrated that CD163 expression was increased by
Ov-NRG1 and decreased by si-NRG1 (Fig. 5E and F). To further verify whether
the reduction in IL-10 was specifically attributable to impaired
NRG1 signaling, hypoxic MDA-MB-231/THP-1 co-cultures were treated
with an anti-NRG1 blocking antibody. Cells were divided into
Hypoxic MDA-MB-231 + THP-1, Hypoxic MDA-MB-231 + THP-1 + IgG and
Hypoxic MDA-MB-231 + THP-1 + anti-NRG1 antibody groups. Blocking
NRG1 markedly decreased IL-10 secretion (Fig. 5G) and consistently reduced CD163
expression (Fig. 5H and I). These
findings indicated that hypoxia-induced NRG1 upregulation is a key
driver of M2 macrophage polarization, at least in part by enhancing
IL-10 secretion and CD163 expression.
Hypoxic MDA-MB-231 cells mediate the M2
polarization of THP-1 macrophages via exosomes
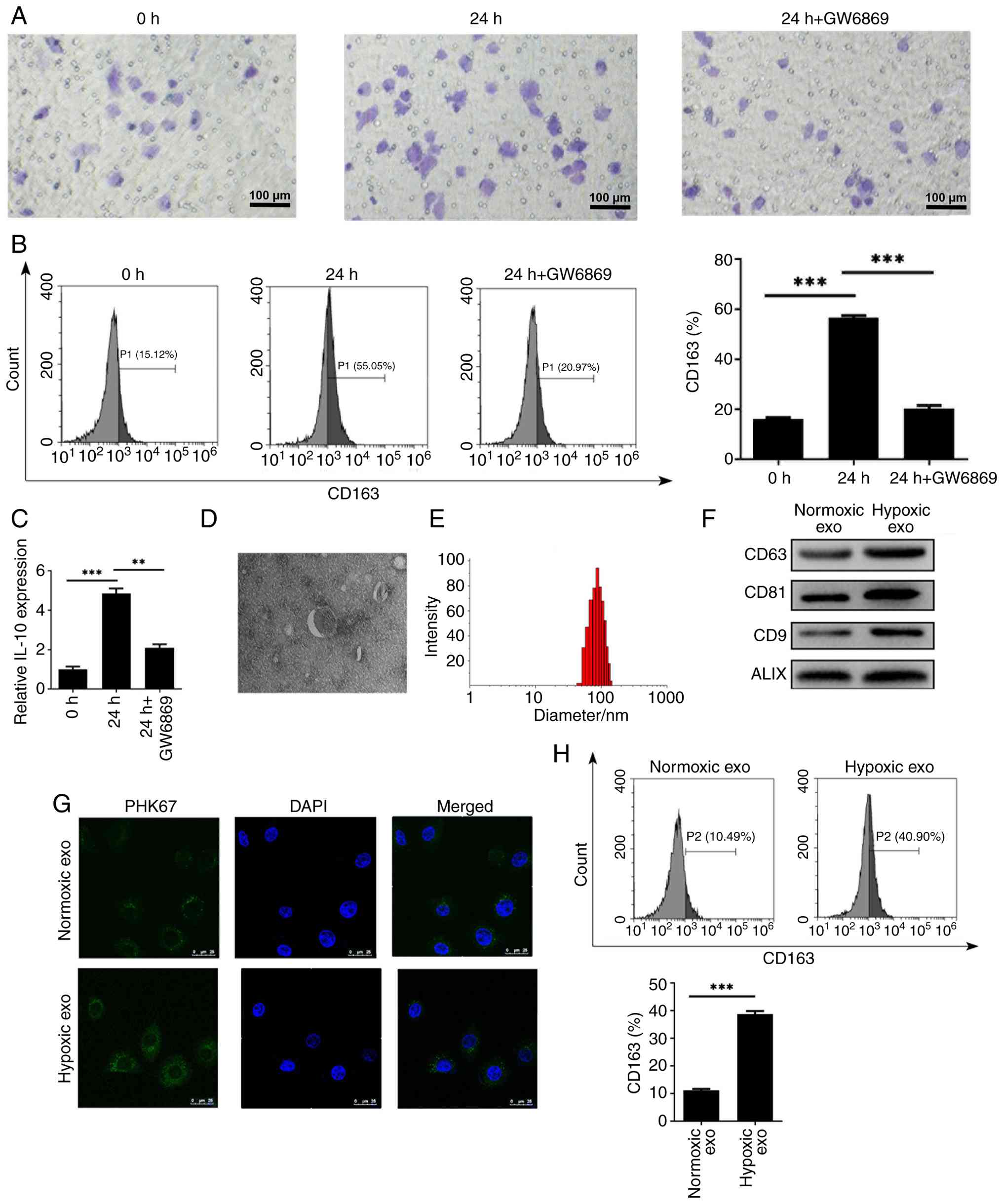
In the tumor microenvironment, macrophages play
crucial roles in tumor initiation and progression. To investigate
whether exosomes contribute to M2 polarization, MDA-MB-231 cells
were treated with the exosome biogenesis inhibitor GW4869 and then
co-cultured with THP-1 macrophages. Transwell assays showed that
hypoxic MDA-MB-231 cells markedly enhanced the migration of THP-1
macrophages, whereas this effect was notably attenuated by GW4869
treatment (Fig. 6A). Moreover,
GW4869 treatment markedly reduced CD163 expression and IL-10
secretion, suggesting that exosomes derived from hypoxic MDA-MB-231
cells are pivotal in promoting M2 polarization (Fig. 6B and C). Transmission electron
microscopy further revealed abundant exosomes with typical bilayer
membrane structures in the supernatant of hypoxic MDA-MB-231 cells.
Nanoparticle size analysis showed that the average exosome diameter
was ~100 nm (Fig. 6D and E).
Immunoblotting analysis confirmed the presence of exosomal marker
proteins CD9, ALIX, CD63 and CD81 in exosome extracts (Fig. 6F).
Exosomes extracted from hypoxic or normoxic
MDA-MB-231 cells were subsequently labeled with PKH67 and incubated
with THP-1 macrophages. Fluorescence imaging confirmed the
successful intake of MDA-MB-231-derived exosomes by THP-1
macrophages (Fig. 6G).
Consistently, flow cytometry analysis demonstrated that
hypoxia-derived exosomes markedly increased CD163 expression in
THP-1 macrophages (Fig. 6H). The
above results indicated that hypoxic MDA-MB-231 cells promoted the
M2 polarization of THP-1 macrophages via exosomes.
Exosomal CAMTA1 promotes the M2
polarization of THP-1 macrophages
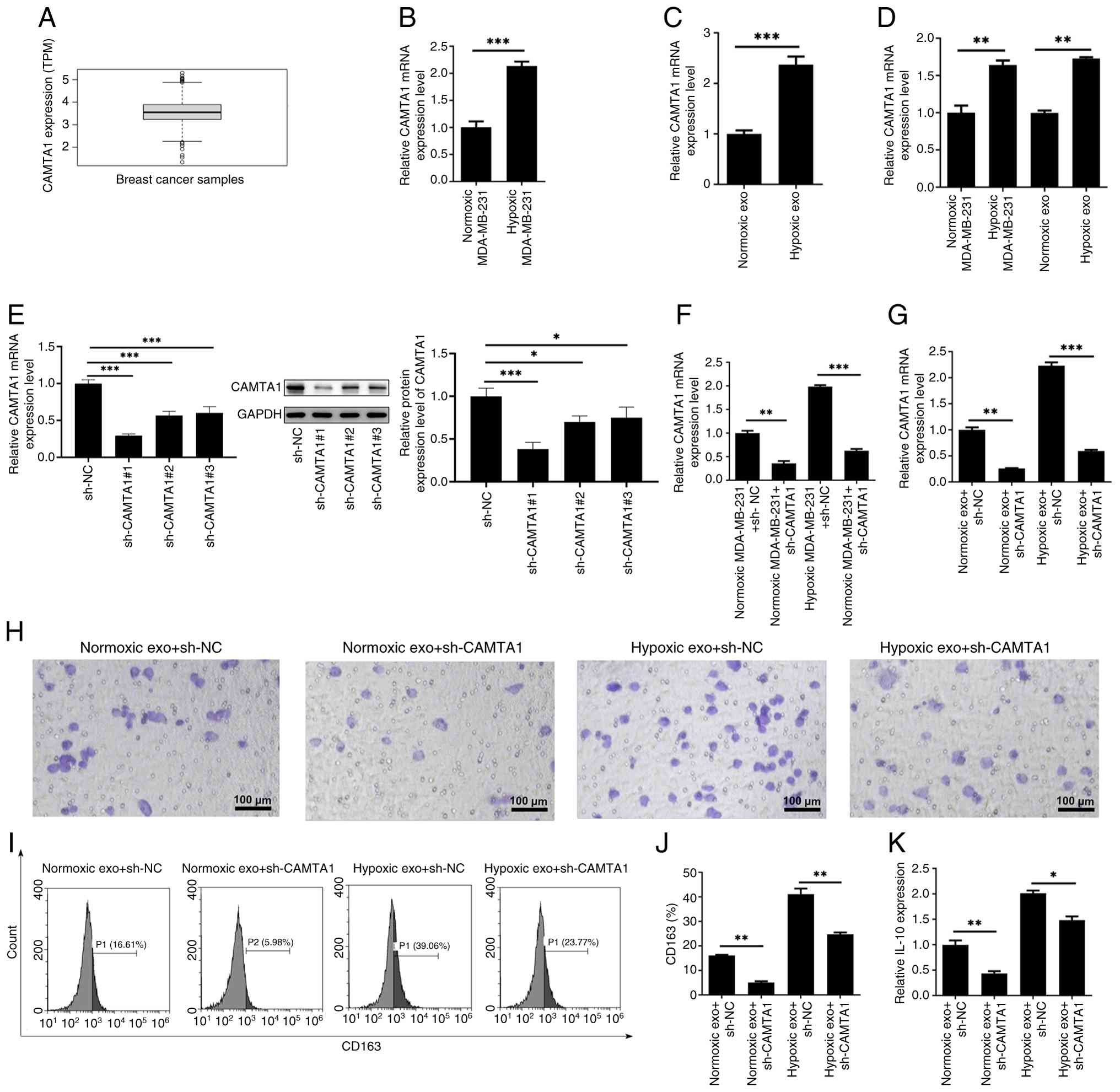
Analysis of the TCGA database revealed that CAMTA1
expression was markedly elevated in BC samples (Fig. 7A). Consistently, RT-qPCR results
showed that hypoxic stimulation markedly increased CAMTA1
expression in both MDA-MB-231 cells and their secreted exosomes
(Fig. 7B and C). Furthermore,
co-culture experiments demonstrated a substantial increase in
CAMTA1 expression with recipient THP-1 macrophages after exposure
to hypoxia-derived exosomes (Fig.
7D). To reduce CAMTA1 expression, sh-CAMTA1 was transfected
into MDA-MB-231 cells and the transfection efficacy was examined
using RT-qPCR and immunoblotting analysis (Fig. 7E). sh-CAMTA1#1 (hereinafter
referred as sh-CAMTA1) was selected for subsequent experiments due
to its improved transfection efficacy. As Fig. 7F and G shows, CAMTA1 expression
was markedly reduced in both MDA-MB-231 cells and exosomes
following the transfection of sh-CAMTA1. Exosomes from
CAMTA1-silenced MDA-MB-231 cells were then incubated with THP-1
macrophages and then divided into Normoxic MDA-MB-231 + exo +
sh-NC, Normoxic MDA-MB-231 + exo + sh-CAMTA1, Hypoxic MDA-MB-231 +
exo + sh-NC and Hypoxic MDA-MB-231 + exo + sh-CAMTA1 groups.
Functional assays revealed that CAMTA1 knockdown markedly impaired
the migration of THP-1 macrophages (Fig. 7H). Moreover, CAMTA1 depletion also
led to a notable reduction in the expressions of CD163 and IL-10
(Fig. 7I-K).
Exosomal CAMTA1 promotes tumor growth in
vivo
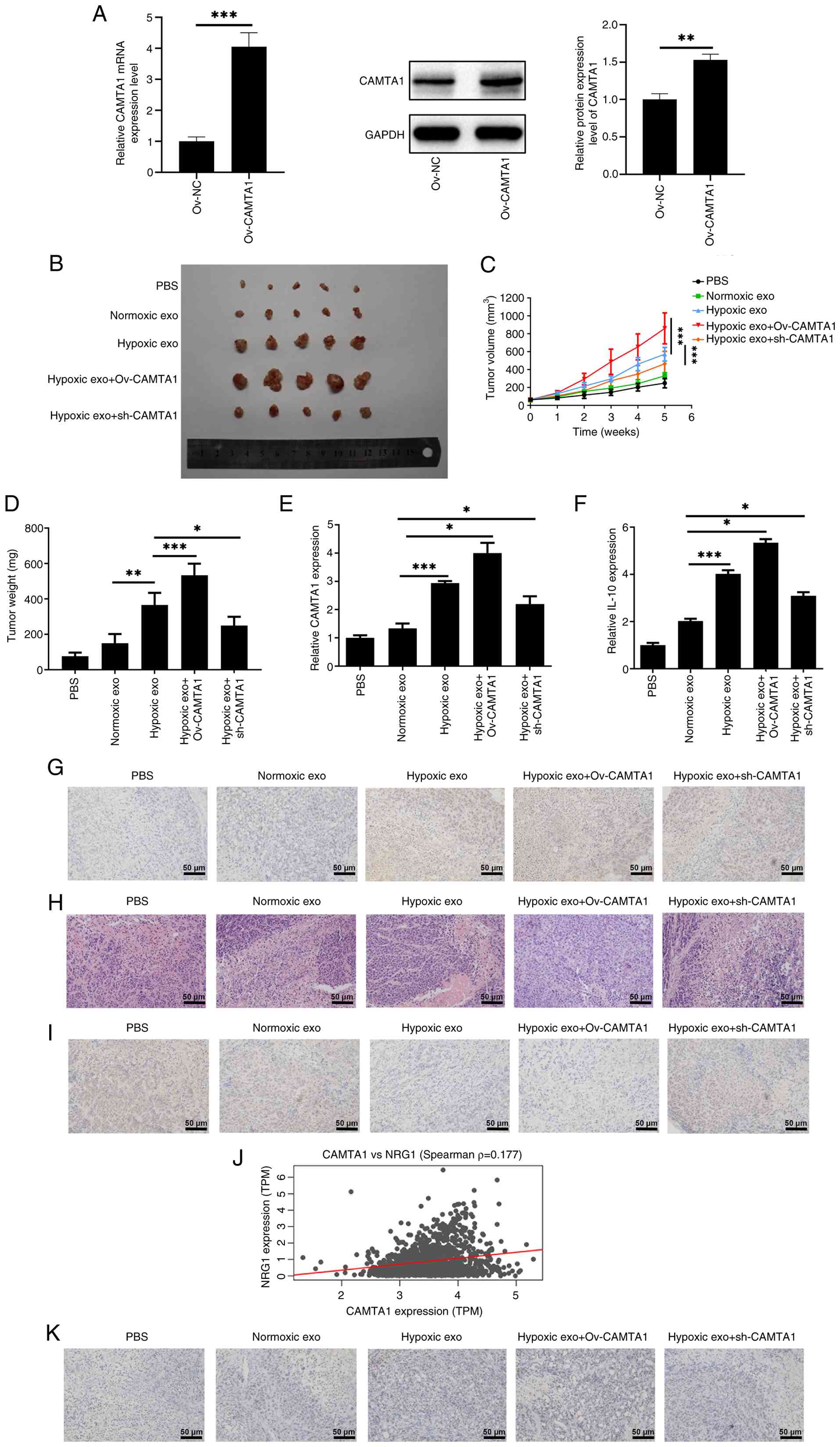
To upregulate CAMTA1 expression, Ov-CAMTA1 was
transfected into MDA-MB-231 cells and the transfection efficacy was
examined (Fig. 8A). To
investigate the role of CAMTA1 in vivo, MDA-MB-231 cells
transfected with sh-CAMTA1 or Ov-CAMTA1 were subcutaneously
injected into mice for the establishment of MDA-MB-231
tumor-bearing nude mice model. The appearance of tumor was shown in
Fig. 8B. As illustrated in
Fig. 8C and D, CAMTA1
overexpression increased the tumor volume and weight, whereas
CAMTA1 knockdown produced the opposite effects. Throughout the
present study, the maximum tumor volume measured was ~850
mm3, with a corresponding maximum diameter of ~10 mm.
RT-qPCR analysis confirmed that CAMTA1 expression in tumor tissues
was elevated by CAMTA1 overexpression and reduced by CAMTA1
silencing compared with the Hypoxia exo group (Fig. 8E). Furthermore, the expression
levels of IL-10 and CD163, which were elevated in the Hypoxia exo
group, were further increased upon CAMTA1 overexpression,
suggesting that CAMTA1 promoted M2 polarization in vivo
(Fig. 8F and G). Additionally,
CAMTA1 overexpression increased tumor cell viability and suppressed
Caspase 3 expression, while CAMTA1 silencing exerted the opposite
effects (Fig. 8H and I).
Correlation analysis revealed a positive association
between CAMTA1 and NRG1 expression (Fig. 8J). Immunohistochemistry further
confirmed that CAMTA1 overexpression markedly upregulated NRG1
expression in tumor tissues, suggesting that CAMTA1 may positively
regulate NRG1 in BC (Fig.
8K).
Discussion
Hypoxia, a hallmark of the tumor microenvironment,
is strongly associated with tumor progression and therapeutic
resistance (27). A previous
study supported that hypoxia stimulates the release of exosomes
from cancer cells, which in turn promote tumor growth and survival
(28). The present study isolated
exosomes from hypoxia-treated MDA-MB-231 cells and investigated
their role in modulating macrophage polarization and
radio-resistance. The findings identified exosomal CAMTA1 as a key
mediator that enhanced the radio-resistance in MDA-MB-231 cells,
possible through promoting M2 macrophage polarization via NRG1. The
present study provided the first evidence, to the best of the
authors' knowledge, linking hypoxia-driven exosomal CAMTA1 to
therapeutic resistance, highlighting it as a novel therapeutic
target for sensitizing BC to radiotherapy.
Exosomes are well recognized as key mediators of
intercellular communication, particularly within the tumor
microenvironment. It is well established that tumor-derived
exosomes contribute to immunosuppression and tumor progression
(21). Among immune cells, M2
macrophages play important roles in driving tumor growth and
migration (29,30). For instance, bladder
cancer-derived exosomes have been reported to accelerate tumor
progression by stimulating M2 macrophage polarization (31). Similarly, Wang et al
(32) demonstrated that liver
cancer cells-derived exosomes under hypoxic conditions promote M2
polarization and increase tumor cell migration. Moreover, a recent
study has showed that BC cell-derived exosomes can be internalized
by macrophages during co-culture, thereby inducing M2 macrophage
polarization and facilitating cell migration (33). Consistent with the aforementioned
findings, the present results showed that hypoxic-derived exosomes
from BC cells facilitated M2 macrophage polarization, which in turn
enhanced the proliferation and suppressed the apoptosis of BC cells
expose to radiotherapy in vitro.
To clarify the molecular contributors to
exosome-mediated polarization, the present study focused on CAMTA1,
a transcriptional regulator previously implicated in tumor
progression (11,34). Analysis of TCGA database revealed
that CAMTA1 expression is markedly upregulated in BC samples.
Consistently, the present study observed increased CAMTA1
expression in both hypoxic MDA-MB-231 cells and their secreted
exosomes. Importantly, CAMTA1 levels were markedly increased in
macrophages after co-culture with hypoxia-derived exosomes,
indicating effective exosomal transfer. To determine its functional
relevance, CAMTA1 was silenced in MDA-MB-231 cells, resulting in a
notable reduction in both cellular and exosomal CAMTA1 levels.
Furthermore, the silencing of CAMTA1 failed to induce typical
M2-like phenotypes in macrophages, as evidenced by decreased
migratory ability and lower CD163 expression. To our knowledge,
this is the first report to identify CAMTA1 as an exosomal cargo
that promotes M2 polarization and contributes to radio-resistance
in BC.
Given the apparent involvement of CAMTA1 in
regulating macrophage polarization, the present study next
investigated the role of NRG1, an oncogene implicated in various
malignancies. A previous study demonstrated elevated NRG1
expression in metastatic hepatocellular carcinoma cells (35) and in thyroid cancer sample tissues
(36). In the present study, NRG1
expression was markedly increased in THP-1 macrophages co-cultured
with hypoxia-treated MDA-MB-231 cells and RNA-seq data from
RAW264.7 macrophages co-cultured with 4T1 cells further confirmed
this upregulation, which was suppressed following irradiation.
These findings implied that NRG1 may mediate immune-modulatory
processes related to radiotherapy. Consistent with this notion,
NRG1 has been shown to enhance M2 macrophage populations (37) and to promote microglia M2
polarization in neuropathic pain following spinal cord injury
(38). Supporting this, GEPIA2
analysis revealed a positive correlation between NRG1 expression
and the M2 marker CD163. Moreover, as IL-10 is a key cytokine
characteristic of reparative M2 macrophages (39), the present study further
corroborated these observations: NRG1 overexpression elevated both
CD163 expression and IL-10 secretion in macrophages co-cultured
with hypoxic MDA-MB-231 cells, confirming its role in promoting M2
macrophage polarization.
To substantiate these findings in vivo,
xenograft tumor experiments were performed in mice, demonstrating
that hypoxic exosomes enriched with CAMTA1 markedly promoted tumor
growth of xenograft tumor mice. Functional analyses revealed that
CAMTA1 overexpression elevated IL-10 and CD163 levels, enhanced
tumor cell viability and inhibited cell apoptosis in xenograft
tumor mice with radiation treatment, suggesting that CAMTA1 could
facilitate M2 polarization and promote the radio-resistance of BC
cells in vivo. Moreover, CAMTA1 overexpression elevated NRG1
expression in tumor tissues, indicating that CAMTA1 may act
upstream of NRG1 in this regulatory axis. This observation was
consistent with Spearman's correlation analysis, which revealed a
positive association between CAMTA1 and NRG1 expression. These
results collectively proposed a novel CAMTA1-NRG1-M2 polarization
pathway that contributed to hypoxia-induced radio-resistance in
BC.
In summary, the present study identifies CAMTA1 as a
key regulator in hypoxia-induced exosomal communication between BC
cells and macrophages. By promoting M2 macrophage polarization and
upregulating NRG1 expression, CAMTA1 contributes to the development
of radio-resistance in MDA-MB-231 cells. Notably, the present study
was the first, to the best of the authors' knowledge, to propose
and provide supporting evidence for a novel CAMTA1-NRG1-M2
polarization axis, uncovering a previously unrecognized mechanism
by which exosomal CAMTA1 shapes the tumor immune microenvironment
and facilitates therapeutic resistance. These findings highlighted
the pivotal role of exosomal CAMTA1 in modulating macrophage
behavior and suggested that targeting this pathway may represent a
novel and clinically relevant strategy to enhance the efficacy of
radiotherapy and improve outcomes for patients with BC.
Despite these novel insights, several limitations
should be acknowledged. First, although the data suggested that
CAMTA1 modulates NRG1 expression, the underlying molecular
mechanism remains undefined. The use of immunohistochemistry
analysis alone is insufficient to confirm a direct regulatory
relationship. Further studies are needed to determine whether
CAMTA1 directly binds to the NRG1 promoter or exerts its effects
through indirect transcriptional regulation. Second, the clinical
relevance of targeting exosomal CAMTA1 in BC has not yet been fully
validated. Comprehensive mechanistic studies and
preclinical/clinical evaluations, are required to clarify the
translational potential of CAMTA1 as a therapeutic target. Third,
although HIF-1α expression was shown to remain elevated after 24 h
of co-culture with macrophages under hypoxic conditions, time
points beyond 24 h was not assessed. As prolonged co-culture could
potentially alter HIF-1α dynamics, this represents another
limitation of the current study.
Availability of data and materials
The raw sequencing data generated in the present
study have been deposited in the NCBI Sequence Read Archive under
BioProject accession PRJNA1372921 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1372921).
Individual SRA accessions include SRR36284160 and SRR36284161:
https://www.ncbi.nlm.nih.gov/sra/SRR36284160,
https://www.ncbi.nlm.nih.gov/sra/SRR36284161. These
datasets are currently under controlled access and will be made
publicly available upon acceptance.
Authors' contributions
QL conceived the experiments. QL and MJ performed
the experiments. QL, BZ, WW and LX analyzed the data. JH, HG and MD
confirmed the authenticity of all the raw data. All authors read
and approved the final manuscript.
Ethics approval and consent to
participate
All animal experiments were approved by the Animal
Ethics Committee of Jiangsu Cancer Hospital and Jiangsu Institute
of Cancer Research and The Affiliated Cancer Hospital of Nanjing
Medical University and conducted in compliance with the National
Institutes of Health Guide for the Care and Use of Laboratory
Animals (approval no. IACUC-20210216-01).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
No funding was received.
References
|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249.
2021.PubMed/NCBI
|
|
2
|
Li T, Mello-Thoms C and Brennan PC:
Descriptive epidemiology of breast cancer in China: Incidence,
mortality, survival and prevalence. Breast Cancer Res Treat.
159:395–406. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Liyanage PY, Hettiarachchi SD, Zhou Y,
Ouhtit A, Seven ES, Oztan CY, Celik E and Leblanc RM:
Nanoparticle-mediated targeted drug delivery for breast cancer
treatment. Biochim Biophys Acta Rev Cancer. 1871:419–433. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Tan C, Hu W, He Y, Zhang Y, Zhang G, Xu Y
and Tang J: Cytokine-mediated therapeutic resistance in breast
cancer. Cytokine. 108:151–159. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Nalio Ramos R, Missolo-Koussou Y,
Gerber-Ferder Y, Bromley CP, Bugatti M, Núñez NG, Tosello Boari J,
Richer W, Menger L, Denizeau J, et al: Tissue-resident
FOLR2+ macrophages associate with CD8+ T cell
infiltration in human breast cancer. Cell. 185:1189–1207.e25. 2022.
View Article : Google Scholar
|
|
6
|
Pan Y, Yu Y, Wang X and Zhang T:
Tumor-associated macrophages in tumor immunity. Front Immunol.
11:5830842020. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Rahal OM, Wolfe AR, Mandal PK, Larson R,
Tin S, Jimenez C, Zhang D, Horton J, Reuben JM, McMurray JS and
Woodward WA: Blocking Interleukin (IL)4- and IL13-mediated
phosphorylation of STAT6 (Tyr641) decreases M2 polarization of
macrophages and protects against macrophage-mediated
radioresistance of inflammatory breast cancer. Int J Radiat Oncol
Biol Phys. 100:1034–1043. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Finkler A, Ashery-Padan R and Fromm H:
CAMTAs: Calmodulin-binding transcription activators from plants to
human. FEBS Lett. 581:3893–3898. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Nagase T, Ishikawa K, Suyama M, Kikuno R,
Hirosawa M, Miyajima N, Tanaka A, Kotani H, Nomura N and Ohara O:
Prediction of the coding sequences of unidentified human genes.
XIII. The complete sequences of 100 new cDNA clones from brain
which code for large proteins in vitro. DNA Res. 6:63–70. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Henrich KO, Claas A, Praml C, Benner A,
Mollenhauer J, Poustka A, Schwab M and Westermann F: Allelic
variants of CAMTA1 and FLJ10737 within a commonly deleted region at
1p36 in neuroblastoma. Eur J Cancer. 43:607–616. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Pan R, Zhang Z, Jia H, Ma J, Wu C, Xue P,
Cai W, Zhang X and Sun J: CAMTA1-PPP3CA-NFATc4 multi-protein
complex mediates the resistance of colorectal cancer to
oxaliplatin. Cell Death Discov. 8:1292022. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Drilon A, Somwar R, Mangatt BP, Edgren H,
Desmeules P, Ruusulehto A, Smith RS, Delasos L, Vojnic M,
Plodkowski AJ, et al: Response to ERBB3-directed targeted therapy
in NRG1-rearranged cancers. Cancer Discov. 8:686–695. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Jonna S, Feldman RA, Swensen J, Gatalica
Z, Korn WM, Borghaei H, Ma PC, Nieva JJ, Spira AI, Vanderwalde AM,
et al: Detection of NRG1 gene fusions in solid tumors. Clin Cancer
Res. 25:4966–6972. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yun S, Koh J, Nam SK, Park JO, Lee SM, Lee
K, Lee KS, Ahn SH, Park DJ, Kim HH, et al: Clinical significance of
overexpression of NRG1 and its receptors, HER3 and HER4, in gastric
cancer patients. Gastric Cancer. 21:225–236. 2018. View Article : Google Scholar
|
|
15
|
Shu L, Chen A, Li L, Yao L, He Y, Xu J, Gu
W, Li Q, Wang K, Zhang T and Liu G: NRG1 regulates Fra-1
transcription and metastasis of triple-negative breast cancer cells
via the c-Myc ubiquitination as manipulated by ERK1/2-mediated
Fbxw7 phosphorylation. Oncogene. 41:907–919. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yi C, Wu S, Duan Q, Liu L, Li L, Luo Y and
Wang A: Ferroptosis-dependent breast cancer cell-derived exosomes
inhibit migration and invasion of breast cancer cells by
suppressing M2 macrophage polarization. PeerJ. 11:e150602023.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Liu C, Huang X, Li S, Ji W, Luo T, Liang J
and Lv Y: M2 macrophage-derived exosomes reverse TGF-β1-induced
epithelial mesenchymal transformation in BEAS-2B cells via the
TGF-βRI/Smad2/3 signaling pathway. Eur J Med Res. 30:2712025.
View Article : Google Scholar
|
|
18
|
Lu F, Ye M, Shen Y, Xu Y, Hu C, Chen J, Yu
P, Xue B, Gu D, Xu L, et al: Hypoxic tumor-derived exosomal
miR-4488 induces macrophage M2 polarization to promote liver
metastasis of pancreatic neuroendocrine neoplasm through RTN3/FABP5
mediated fatty acid oxidation. Int J Biol Sci. 20:3201–3218. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yang Y, Li CW, Chan LC, Wei Y, Hsu JM, Xia
W, Cha JH, Hou J, Hsu JL, Sun L and Hung MC: Exosomal PD-L1 harbors
active defense function to suppress T cell killing of breast cancer
cells and promote tumor growth. Cell Res. 28:862–864. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Hansen MR, Roehm PC, Chatterjee P and
Green SH: Constitutive neuregulin-1/ErbB signaling contributes to
human vestibular schwannoma proliferation. Glia. 53:593–600. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yang Y, Wu T, Wang Y, Luo D, Zhao Z, Sun
H, Zhang M, Zhang B and Han B: Hypoxic tumour-derived exosomal
miR-1290 exacerbates the suppression of CD8+ T cells by promoting
M2 macrophage polarization. Immunology. 173:672–688. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang Y, Zhang J, Shi H, Wang M, Yu D, Fu
M, Qian Y, Zhang X, Ji R, Wang S, et al: M2 Tumor-associated
macrophages-derived exosomal MALAT1 promotes glycolysis and gastric
cancer progression. Adv Sci (Weinh). 11:e23092982024. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
24
|
Ye M, Lu F, Gu D, Xue B, Xu L, Hu C, Chen
J, Yu P, Zheng H, Gao Y, et al: Hypoxia exosome derived CEACAM5
promotes tumor-associated macrophages M2 polarization to accelerate
pancreatic neuroendocrine tumors metastasis via MMP9. FASEB J.
38:e237622024. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Guo M, Wang R, Nie M, Zhang H, Wang C,
Song C and Niu S: H3K27ac-induced RHOXF2 activates Wnt2/β-catenin
pathway by binding to HOXC13 to aggravate the malignant progression
of triple negative breast cancer. Cell Signal. 120:1111962024.
View Article : Google Scholar
|
|
26
|
Jiang D, Gao X, Tan R, Liu X, Zhu Y and
Zhang L: Euphorbia factor L1 suppresses breast cancer liver
metastasis via DDR1-mediated immune infiltration. Aging.
15:9217–9229. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Schito L and Semenza GL: Hypoxia-inducible
factors: Master regulators of cancer progression. Trends Cancer.
2:758–770. 2016. View Article : Google Scholar
|
|
28
|
King HW, Michael MZ and Gleadle JM:
Hypoxic enhancement of exosome release by breast cancer cells. BMC
Cancer. 12:4212012. View Article : Google Scholar :
|
|
29
|
Yang Y, Jin X, Xie Y, Ning C, Ai Y, Wei H,
Xu X, Ge X, Yi T, Huang Q, et al: The CEBPB+
glioblastoma subcluster specifically drives the formation of M2
tumor-associated macrophages to promote malignancy growth.
Theranostics. 14:4107–4126. 2024. View Article : Google Scholar
|
|
30
|
Sun Y, Lian Y, Mei X, Xia J, Feng L, Gao
J, Xu H, Zhang X, Yang H, Hao X and Feng Y: Cinobufagin inhibits
M2-like tumor-associated macrophage polarization to attenuate the
invasion and migration of lung cancer cells. Int J Oncol.
65:1022024. View Article : Google Scholar
|
|
31
|
Jiang Z, Zhang Y, Zhang Y, Jia Z, Zhang Z
and Yang J: Cancer derived exosomes induce macrophages
immunosuppressive polarization to promote bladder cancer
progression. Cell Commun Signal. 19:932021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wang X, Zhou Y, Dong K, Zhang H, Gong J
and Wang S: Exosomal lncRNA HMMR-AS1 mediates macrophage
polarization through miR-147a/ARID3A axis under hypoxia and affects
the progression of hepatocellular carcinoma. Environ Toxicol.
37:1357–1372. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hao C, Sheng Z, Wang W, Feng R, Zheng Y,
Xiao Q and Zhang B: Tumor-derived exosomal miR-148b-3p mediates M2
macrophage polarization via TSC2/mTORC1 to promote breast cancer
migration and invasion. Thorac Cancer. 14:1477–1491. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Schraivogel D, Weinmann L, Beier D,
Tabatabai G, Eichner A, Zhu JY, Anton M, Sixt M, Weller M, Beier CP
and Meister G: CAMTA1 is a novel tumour suppressor regulated by
miR-9/9* in glioblastoma stem cells. EMBO J. 30:4309–4322. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Shi DM, Li LX, Bian XY, Shi XJ, Lu LL,
Zhou HX, Pan TJ, Zhou J, Fan J and Wu WZ: miR-296-5p suppresses EMT
of hepatocellular carcinoma via attenuating NRG1/ERBB2/ERBB3
signaling. J Exp Clin Cancer Res. 37:2942018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhang TT, Qu N, Sun GH, Zhang L, Wang YJ,
Mu XM, Wei WJ, Wang YL, Wang Y, Ji QH, et al: NRG1 regulates redox
homeostasis via NRF2 in papillary thyroid cancer. Int J Oncol.
53:685–693. 2018.PubMed/NCBI
|
|
37
|
Alizadeh A, Santhosh KT, Kataria H, Gounni
AS and Karimi-Abdolrezaee S: Neuregulin-1 elicits a regulatory
immune response following traumatic spinal cord injury. J
Neuroinflammation. 15:532018. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ma Y, Fan P, Zhao R, Zhang Y, Wang X and
Cui W: Neuregulin-1 regulates the conversion of M1/M2 microglia
phenotype via ErbB4-dependent inhibition of the NF-κB pathway. Mol
Biol Rep. 49:3975–3986. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Chen X, Wan Z, Yang L, Song S, Fu Z, Tang
K, Chen L and Song Y: Exosomes derived from reparative M2-like
macrophages prevent bone loss in murine periodontitis models via
IL-10 mRNA. J Nanobiotechnol. 20:1102022. View Article : Google Scholar
|