Introduction
MicroRNAs (miRNAs or miRs) constitute a class of
small non-coding RNAs, typically ~22 nucleotides in length, that
are essential for gene regulation. They primarily modulate gene
expression by base-pairing with target messenger RNAs (mRNAs),
resulting in either translational repression or mRNA degradation
(1). A single miRNA can regulate
multiple target genes, thereby exerting pleiotropic effects.
Consequently, miRNAs are pivotal in various physiological and
pathological processes, such as morphogenesis, disease progression
and tumorigenesis (2). Among
them, miR-124 is a highly conserved miRNA found in various species
and tissues. It is often downregulated in numerous cancers, where
it acts as a tumor suppressor by inhibiting multiple oncogenic
pathways (3). These pathways
include cell proliferation, metastasis, stemness, drug resistance,
and modulation of the tumor microenvironment (TME) and immune
response (4). Moreover, recent
research suggests that miR-124 plays a pivotal role in regulating
genes involved in metabolism and contributes to the metabolic
reprogramming observed in tumors (5).
This discovery highlights its functional complexity
in diverse physiological and pathological contexts. In cancer, the
upstream regulatory mechanisms of miR-124 constitute a complex,
multi-layered network that is regulated by transcription factors,
epigenetic modifications, competing endogenous RNA (ceRNA)
networks, and various signaling pathways (6,7).
These mechanisms collectively and precisely modulate miR-124
expression, thereby affecting numerous downstream target genes and
signal transduction pathways. miR-124 has emerged as a promising
therapeutic target due to its fundamental tumor-suppressive
function. Research has implicated miR-124 in cancer diagnosis,
therapeutic response, and prognosis assessment, underscoring its
significant potential as both a biomarker and a therapeutic target
(3). A comprehensive
investigation into the functional mechanisms of miR-124 in tumor
progression is poised to significantly advance the development of
personalized medicine and targeted therapies, thereby paving the
way for novel clinical interventions. Despite the significant
challenges associated with miR-124 translation into clinical
practice, a systematic examination of miR-124 and its regulatory
network is anticipated to significantly enrich our understanding of
cancer biology.
Multifaceted roles of miR-124 in tumors
miR-124, a broad-acting tumor suppressor, plays a
crucial role in tumorigenesis and cancer progression, as evident by
its consistent downregulation in various human cancers (3,8).
miR-124 is involved in numerous biological processes during
tumorigenesis and progression, attributed to its multifaceted
regulatory capabilities. It directly inhibits the proliferation,
migration and invasion of tumor cells. Furthermore, miR-124 exerts
tumor-suppressive effects across multiple fronts by modulating
cancer stemness, drug sensitivity, TME remodeling and immune
responses. This chapter systematically elucidates the roles and
mechanisms underlying miR-124 in tumor growth, metastasis, stemness
maintenance, drug resistance, microenvironment regulation and
immune modulation. It also elaborates on its complex regulatory
network, thereby providing a theoretical foundation for developing
miR-124-based diagnostic and therapeutic strategies.
Suppression of tumor growth and
metastasis by miR-124
miR-124 is an important tumor suppressor frequently
downregulated in various cancers. Its primary role is to inhibit
the progression of malignant tumors, particularly by targeting
tumor growth and metastasis (9).
This inhibitory effect is mediated by a sophisticated regulatory
network that concurrently targets multiple critical nodes within
complementary and synergistic signaling pathways (3,10,11). The tumor-suppressive activity of
miR-124 is characterized by three hallmark features. First, the
systematic, cancer-type-specific nature of miR-124 target selection
facilitates its simultaneous inhibition of essential proliferative
processes and tissue-specific invasive behaviors. Particularly,
miR-124 suppresses key pathways that regulate cell cycle
progression and proliferation by targeting widely acting
oncoproteins, such as signal transducer and activator of
transcription 3 (STAT3) (3),
enhancer of zeste homolog 2 (EZH2) (12), cyclin-dependent kinase 6 (CDK6)
(13), and SET and MYND domain
containing 3 (14). Furthermore,
it affects tissue-specific factors, including androgen receptor in
prostate cancer (PC) (15,16),
Forkhead box Q1 in nasopharyngeal carcinoma and breast cancer (BC)
(17-19), integrin subunit beta 1 in
osteosarcoma (20), and receptor
tyrosine kinase like orphan receptor 2 in medulloblastoma (21,22) thereby disrupting
cancer-type-specific migration, invasion, and survival signaling
pathways.
This coordinated regulation of both universal and
context-dependent targets enables miR-124 to effectively suppress
tumor progression across various cancer types. Second, the wide
spectrum of miR-124 activity underlies its tumor-suppressive
effects, extending beyond cancer cells themselves by directly
targeting and remodeling the metastatic microenvironment. For
instance, miR-124 in oral cancer inhibits the tumor-promoting
functions of cancer-associated fibroblasts (CAFs) by downregulating
C-C motif chemokine ligand 2 and interleukin-8 (23). In BC, miR-124 disrupts the
formation of the bone metastatic niche by targeting interleukin-11
(24). Furthermore, in non-small
cell lung cancer, exosome-mediated intercellular transmission of
miR-124 facilitates the spatiotemporal extension of its
tumor-suppressive effects (25,26). Through these multifaceted
mechanisms, miR-124 effectively inhibits metastasis at both local
and systemic levels. Third, the significant therapeutic potential
of miR-124 is underscored by the intricate nature of upstream
regulatory mechanisms and the extensive scope of its functional
network. miR-124 expression is modulated by ceRNA molecules,
including long non-coding RNAs (lncRNAs) SND1-IT1 (27) and LINC00240 (28), indicating the complex layers of
its regulatory control. The concurrent targeting of multiple
critical nodes by miR-124 that promote metastasis, such as IQ Motif
Containing GTPase activating protein 1 in colorectal cancer (CRC)
(29), arrestin domain containing
1 in hepatocellular carcinoma (HCC) (30), and histidine rich carboxyl
terminus 1 in gastric cancer (GC) (31), suggests that restoring or
simulating miR-124 function could be a promising multi-target
therapeutic strategy. This approach can inhibit tumor growth and
metastasis, thereby improving patient prognosis. Collectively,
these findings underscore the consistent role of miR-124 in tumor
suppression across diverse cancer types, underscoring its broad
therapeutic applicability.
Modulation of cancer stem cell (CSC)
properties by miR-124
CSCs are a minor yet critical subpopulation within
tumors, distinguished by their ability to self-renew and to
differentiate into multiple lineages. These cells play a crucial
role in tumor initiation, metastasis and therapeutic resistance
(32,33). miR-124 significantly affects the
self-renewal and differentiation capabilities of CSCs by regulating
key target genes (34). Notably,
the dysregulation of miR-124-mediated pathways is strongly
associated with the emergence of resistance in CSCs to standard
chemotherapy and radiotherapy (35). Studies have demonstrated that the
natural compound sulforaphane (SFN), in addition to its
wide-ranging antitumor effects, can particularly improve the
transcription of miR-124. This upregulation leads to the
downregulation of essential stemness genes, including β-catenin,
SRY-box transcription factor 2 and octamer-binding transcription
factor 4 (Oct4). Consequently, SFN significantly reduces stemness
and the self-renewal capacity of CSCs (36). miR-124 exerts a significant
inhibitory effect on CSCs, as observed across various tumors. For
instance, miR-124 directly downregulates the expression of the
junctional adhesion molecule A in nasopharyngeal carcinoma, thereby
reducing stem cell-like characteristics of tumor cells and
increasing their sensitivity to radiotherapy (37). In HCC, miR-124 inhibits CSC
proliferation and reverses sorafenib resistance (38).
The regulatory mechanisms of miR-124 demonstrate
significant heterogeneity across various cancer types. In
glioblastoma (GBM), miR-124 directly targets the oncogene EPH
receptor A2 by disrupting AlkB homolog 5-mediated
N6-methyladenosine RNA modification, thereby inhibiting tumor cell
stemness and tumorigenic potential (39). This discovery highlights a novel
epitranscriptomic mechanism of miR-124 regulation. Conversely,
circular RNA circ-trichorhinophalangeal syndrome type 1 functions
as a molecular sponge in PC, sequestering miR-124, thereby
indirectly upregulating EZH2 expression and enhancing stem
cell-like characteristics. However, the exogenous reintroduction of
miR-124 counteracts the EZH2-mediated epigenetic silencing pathway,
thereby reducing tumor stemness and improving patient prognosis
(40). Collectively, these
findings clarify the critical role of the circRNA-miRNA-epigenetic
axis in regulating stemness from a ceRNA perspective. The
significant metastatic potential of CSCs is closely linked to their
tumorigenic properties, a highly aggressive characteristic
primarily attributed to their ability to undergo
epithelial-mesenchymal transition (EMT). It is a process deeply
embedded in their intrinsic cellular plasticity. miR-124 partially
mitigates the metastatic potential of CSCs by inhibiting EMT. EMT
endows cells with migratory and invasive capabilities and is a
fundamental mechanism by which CSCs facilitate metastasis.
Mechanistically, miR-124 has been demonstrated to directly target
key transcription factors that induce EMT, such as Zinc Finger
E-Box Binding Homeobox 1 and Snail Family Transcriptional Repressor
1 (SNAI1) (20,41). For instance, miR-124 reduces SNAI1
expression in osteosarcoma, thereby attenuating transforming growth
factor-beta-induced EMT and reducing metastatic potential (20). miR-124 disrupts the CSC-induced
metastatic cascade by simultaneously suppressing both stemness and
EMT, underscoring its critical role as a tumor suppressor.
In conclusion, miR-124 modulates numerous signaling
pathways to play a critical inhibitory role in maintaining stemness
in various CSCs. These findings provide new insights into the
heterogeneity and plasticity of CSCs and establish a theoretical
basis for the development of miR-124-based precision therapies.
Despite the cancer-type specificity of miR-124 mechanisms,
targeting miR-124 or its associated pathways offers a promising
universal approach to overcoming resistance to tumor therapies.
Future research should focus on elucidating the specific targets of
miR-124 and the interactions between the pathways it regulates in
different TMEs. These endeavors will be indispensable for advancing
the clinical translation of miR-124-based therapeutic
strategies.
Role of miR-124 in reversing tumor drug
resistance
Tumor drug resistance is characterized by the
ability of cancer cells to endure chemotherapy, targeted therapy,
and other anticancer agents, resulting in reduced therapeutic
efficacy and unfavorable patient outcomes (42,43). Recent advances in cancer
therapeutics have highlighted the importance of miR-124 in
modulating drug resistance (44).
miR-124 exhibits multifaceted regulatory functions in counteracting
tumor drug resistance. miR-124 improves drug sensitivity by
targeting pro-survival signaling pathways at the cellular level.
Notably, miR-124, a key negative regulator of the Janus
kinase-signal transducer and activator of transcription pathway,
suppresses the expression of the glycosyltransferase
beta-1,4-galactosyltransferase 1, thereby reversing chemotherapy
resistance in tumor cells (45).
In liver cancer, miR-124 enhances the sensitivity of HCC cells to
sorafenib by inhibiting the AKT serine/threonine kinase 2/sirtuin 1
pathway, thereby activating Forkhead Box O3a-mediated apoptosis.
Furthermore, miR-124 directly targets TNF receptor-associated
factor 6, negatively regulating the nuclear factor kappa B (NF-κB)
pathway and thereby counteracting pro-survival signaling-induced
drug resistance (46,47). Regarding the DNA damage response,
miR-124 improves the efficacy of genotoxic agents, such as
temozolomide, by impairing the DNA repair mechanisms of tumor
cells, a phenomenon particularly well-documented in glioma
(48,49). Moreover, miR-124 suppresses CSC
characteristics. For instance, in BC, it diminishes stemness and
doxorubicin resistance by disrupting the STAT3/hypoxia inducible
factor 1 alpha (HIF-1α) signaling axis (50,51). Given its influence on TME and cell
death regulation, miR-124 modulates immune microenvironment
dynamics and pyroptosis-related processes, which, in turn, affect
the response to gemcitabine chemotherapy in bladder cancer
(52,53). In conclusion, miR-124
significantly enhances the efficacy of various anticancer agents by
orchestrating multiple mechanisms, including apoptosis, DNA repair,
stemness, the TME, and epigenetic modifications. These diverse
functions highlight miR-124 as a promising therapeutic target for
combating drug resistance.
Although miR-124 significantly reverses drug
resistance in preclinical studies, its translation from
experimental findings to clinical application is hindered by
numerous obstacles. As elaborated in subsequent sections, these
challenges are predominantly related to delivery systems, stability
and targeting strategies. The systematic resolution of these issues
could establish miR-124 as an innovative and effective therapeutic
approach to overcome tumor drug resistance.
Modulation of the TME by miR-124
TME is a complex ecosystem comprising tumor cells,
immune cells, stromal cells, the extracellular matrix (ECM),
soluble factors, and physicochemical conditions such as hypoxia and
acidic pH. This intricate assembly facilitates tumor proliferation,
invasion, metastasis and drug resistance (54-56). miRNAs are crucial regulators
within the TME, and their expression and activity are tightly
controlled by various TME components while simultaneously
contributing to TME reprogramming. Among these, miR-124 assumes a
central regulatory role within this intricate network (57). Within the immune microenvironment,
miR-124 downregulates the expression of pro-inflammatory factors,
including tumor necrosis factor-alpha and interleukin-1 beta. This
reduction in tumor-associated inflammation subsequently reduces the
risk of tumor initiation and progression (58,59). Furthermore, miR-124 directly
modulates macrophage polarization by inhibiting M1 markers and
promoting the M2 phenotype, thereby reprogramming the immune
microenvironment and affecting tumor progression (60,61). miR-124 regulates T-cells by
inhibiting TH17 cell differentiation. Notably, miR-124 demonstrates
both anti-inflammatory and antitumor properties in models of
colitis-associated CRC (62,63). In the metabolic microenvironment,
miR-124 is an important link between metabolic reprogramming and
inflammatory responses. Particularly, in CRC, cell energy
utilization and signal secretion through lipid metabolic
reprogramming are affected by a regulatory network involving the
RNA-binding protein HuR, miR-124, and its target gene, the vitamin
D receptor (VDR). Subsequently, this process affects immune cell
infiltration and function, including VDR signaling-induced
macrophage polarization (64,65). Regarding the ECM and
physicochemical microenvironment, emerging evidence indicates that
miR-124 is involved in cell-matrix interactions and stress
responses. lncRNA MALAT1 activates the phosphatidylinositol
3-kinase/protein kinase B (PI3K/AKT) pathway through the
miR-124/laminin subunit gamma 1 axis. This mechanism, initially
identified in tissue repair, suggests that MALAT1 may contribute to
tumor ECM remodeling and regulation of the metastatic
microenvironment (66,67).
In conclusion, miR-124 is crucial to prevent the
formation of a pro-TME by regulating various mechanisms, including
immune responses, metabolic reprogramming and physicochemical
properties. miR-124 precisely modulates inflammatory factors,
metabolic pathways, and immune differentiation, through its
post-transcriptional regulatory functions, underscoring its
potential as a viable target for therapies targeting the TME.
Notably, the regulatory effects of miR-124 on immune cells are
highly context-dependent, and the specific outcomes vary based on
tumor type and microenvironmental signals. Therefore, future
research should focus on elucidating the role of miR-124 within
specific components of the TME, such as CAFs, ECM and hypoxic
niches, to elucidate its functional diversity across various
contexts, thereby facilitating its precise translation in tumor
immunology and microenvironment-targeted therapeutic
strategies.
Modulation of tumor immunity by
miR-124
miR-124 regulates tumor immunity by coordinating
multicellular and multipathway synergistic effects. It enhances
antitumor immune responses by simultaneously targeting both immune
and tumor cells (68). In the
adaptive immune system, miR-124 enhances the CD8+
T-cell-mediated clearance of tumor cells and facilitates the
production and function of effector CD4+ T-cells by
inhibiting the STAT3 signaling pathway (69-71). Furthermore, miR-124 increases the
cytotoxic activity of natural killer (NK) cells by modulating the
LINC00240/miR-124/STAT3/MHC Class I polypeptide-related sequence A
axis (28). In neuro-immunity,
miR-124 regulates microglial activation via the MALAT1/miR-124/SGK
axis in myeloid immune cells (72). Similarly, miR-124 may reverse the
immunosuppressive microenvironment by affecting the polarization
state of tumor-associated macrophages through this mechanism.
Moreover, miR-124 directly influences tumor cells by downregulating
immune checkpoint molecules, including programmed death-ligand 1
(PD-L1), while simultaneously upregulating the expression of
pro-inflammatory factors. This dual action impedes immune evasion
and facilitates the immune cell infiltration (61,73). Furthermore, miR-124-regulated
JAK/STAT and NF-κB signaling pathways act as 'bridge pathways' that
connect inflammation and cancer in autoimmune diseases, thereby
reinforcing the critical role of miR-124 in tumor immunology
(74-76). In summary, miR-124 emerges as a
critical therapeutic target that can reverse the immunosuppressive
microenvironment and modulate antitumor immune responses through
its integrated multicellular and multipathway regulatory
mechanisms. The development of effective regulatory strategies,
including nanoparticle-based or natural compound-derived delivery
systems, has the potential to advance the field of cancer
immunotherapy significantly.
miR-124 demonstrates significant functional
pleiotropy and context-dependent effects in regulating the immune
system. miR-124 can manifest diverse and occasionally contradictory
biological functions in distinct TME niches and among various
immune cell subsets. As aforementioned, miR-124 facilitates
macrophage polarization towards the M2 phenotype, primarily
indicating its regulatory role in mitigating inflammatory damage
and preserving tissue homeostasis within specific
microenvironmental contexts. Conversely, this section highlights
its capacity to counteract tumor immune suppression by targeting
key immune checkpoint molecules, such as STAT3 and PD-L1, thereby
augmenting the effector functions of CD8+ T-cells and NK
cells. This functional plasticity demonstrates the complex role of
miR-124 as a microenvironment-responsive regulator. Its ultimate
effect is not an intrinsic property of the molecule itself.
However, it is influenced by the spatiotemporal dynamics of the
tumor type, immune cell subset status, the repertoire of
microenvironmental signals, and intercellular interactions.
Consequently, it is imperative to analyze the immunomodulatory
mechanisms of miR-124 across distinct pathophysiological contexts,
avoiding a reductionist binary perspective on its function.
In summary, miR-124 acts as an important tumor
suppressor by regulating tumor growth, metastasis, stemness, drug
resistance and immune responses. Its multifaceted functions are
systematically cataloged in Table
I and schematically summarized in Fig. 1, highlighting its central role in
cancer biology.
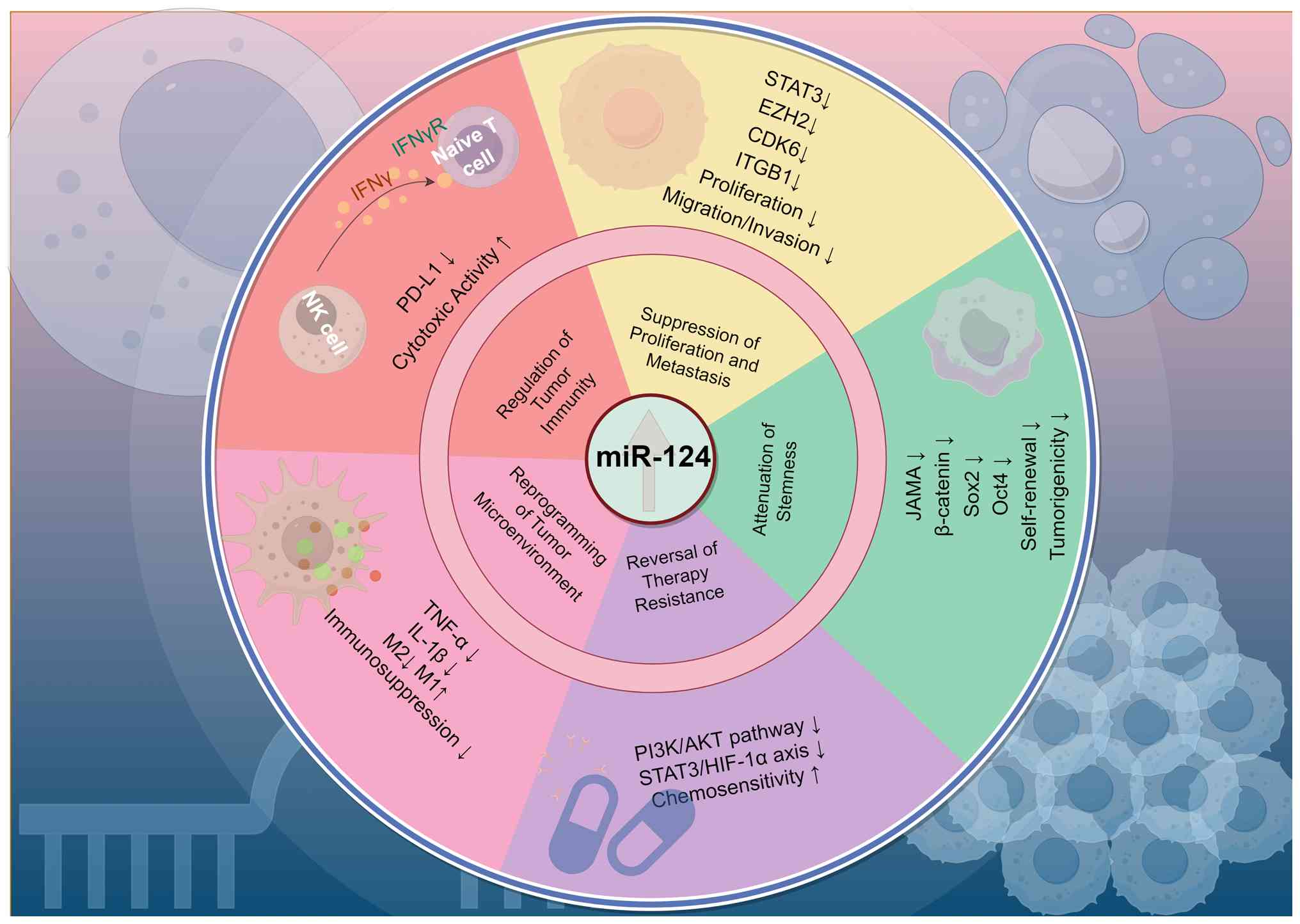 | Figure 1Comprehensive summary of the
tumor-suppressive functions of miR-124. This figure summarizes the
key functional roles of miR-124 discussed throughout this section,
including inhibition of tumor growth and metastasis, modulation of
cancer stem cell properties, reversal of drug resistance, and
regulation of the tumor microenvironment and antitumor immunity.
This figure was created using FigDraw. miR, microRNA; PD-L1,
programmed death-ligand 1; EZH2, enhancer of zeste homolog 2; JAMA,
junctional adhesion molecule A; ITGB1, integrin subunit beta 1;
OCT4, octamer-binding transcription factor 4; SOX2, SRY-box
transcription factor 2; HIF-1α, hypoxia inducible factor 1 alpha;
CDK6, cyclin-dependent kinase 6. |
 | Table IMultifunctional roles of miR-124 in
cancer. |
Table I
Multifunctional roles of miR-124 in
cancer.
| Functional
category | Key mechanisms of
action | Critical
targets/regulatory axes | Associated
cancers |
|---|
| Inhibition of tumor
growth and metastasis | Suppresses tumor
cell proliferation and invasion; impairs the pre-metastatic
niche | STAT3 (13), EZH2 (14), CDK6 (15), SMYD3 (16), AR (17,18), Foxq1 (19-21), ITGB1 (22), ROR2 (23,24); CCL2/IL-8 in CAFs (25); IL-11 in bone metastasis (26); Exosome-mediated transfer (27,28); lncRNA SND1-IT1/LINC00240 (ceRNA)
(29,30); IQGAP1 (31), ARRDC1 (32), HRCT1 (33) | Prostate cancer,
nasopharyngeal carcinoma, breast cancer, osteosarcoma,
medulloblastoma, oral squamous cell carcinoma, non-small cell lung
cancer |
| Reduction of cancer
stem cell stemness | Inhibits
self-renewal and multi-lineage differentiation potential | β-catenin, Sox-2,
Oct4 (via SFN induction) (38);
JAMA (39); EPHA2 (via
ALKBH5/m6A) (41); EZH2 (via
circ-TRPS1 sponge) (20) | Nasopharyngeal
carcinoma, liver cancer, glioblastoma, prostate cancer |
| Reversal of tumor
drug resistance | Enhances
chemosensitivity by promoting drug-induced apoptosis and impairing
DNA damage repair | B4GALT1/JAK-STAT
(45); AKT2/SIRT1/FOXO3a and
TRAF6/NF-κB axes (46,47); DNA repair mechanisms in glioma
(48,49); STAT3/HIF-1α axis (50,51); TME/Pyroptosis in bladder cancer
(52,53) | Liver cancer,
glioma, breast cancer, bladder cancer |
| Modulation of
TME | Reprograms the TME
by regulating immune cell polarization, inflammatory factors,
metabolic reprogramming, and ECM remodeling | TNF-α/IL-1β
(58,69); Macrophage polarization (M1/M2)
(60,61); TH17 cell differentiation (62,63); HuR/miR-124/VDR axis (metabolic
reprogramming) (64,65); lncRNA-Malat1/miR-124/Lamc1 axis
(ECM remodeling) (66,67) | Colorectal cancer,
oral cancer, breast cancer |
| Regulation of tumor
immunity | Potentiates
antitumor immunity by enhancing T and NK cell activity and
suppressing immune checkpoint expression | STAT3 in T cells
(69-71); LINC00240/miR-124/STAT3/MICA axis
in NK cells (28);
MALAT1/miR-124/SGK axis in microglia/TAMs (72); PD-L1 downregulation (61,73); JAK/STAT & NF-κB bridge
pathways (74-76) | Various solid
tumors and hematological malignancies |
Upstream regulatory mechanisms of miR-124 in
tumors
The upstream regulatory mechanisms of miR-124 in
tumors comprise a complex network, including transcription factors,
signaling pathways, epigenetic modifications, and competitive
interactions with non-coding RNAs. Understanding this intricate
network is a primary focus of contemporary research efforts. The
transcriptional regulation of miR-124 involves a coordinated
interplay of various transcription factors and oncogenic signaling
pathways (Fig. 2).
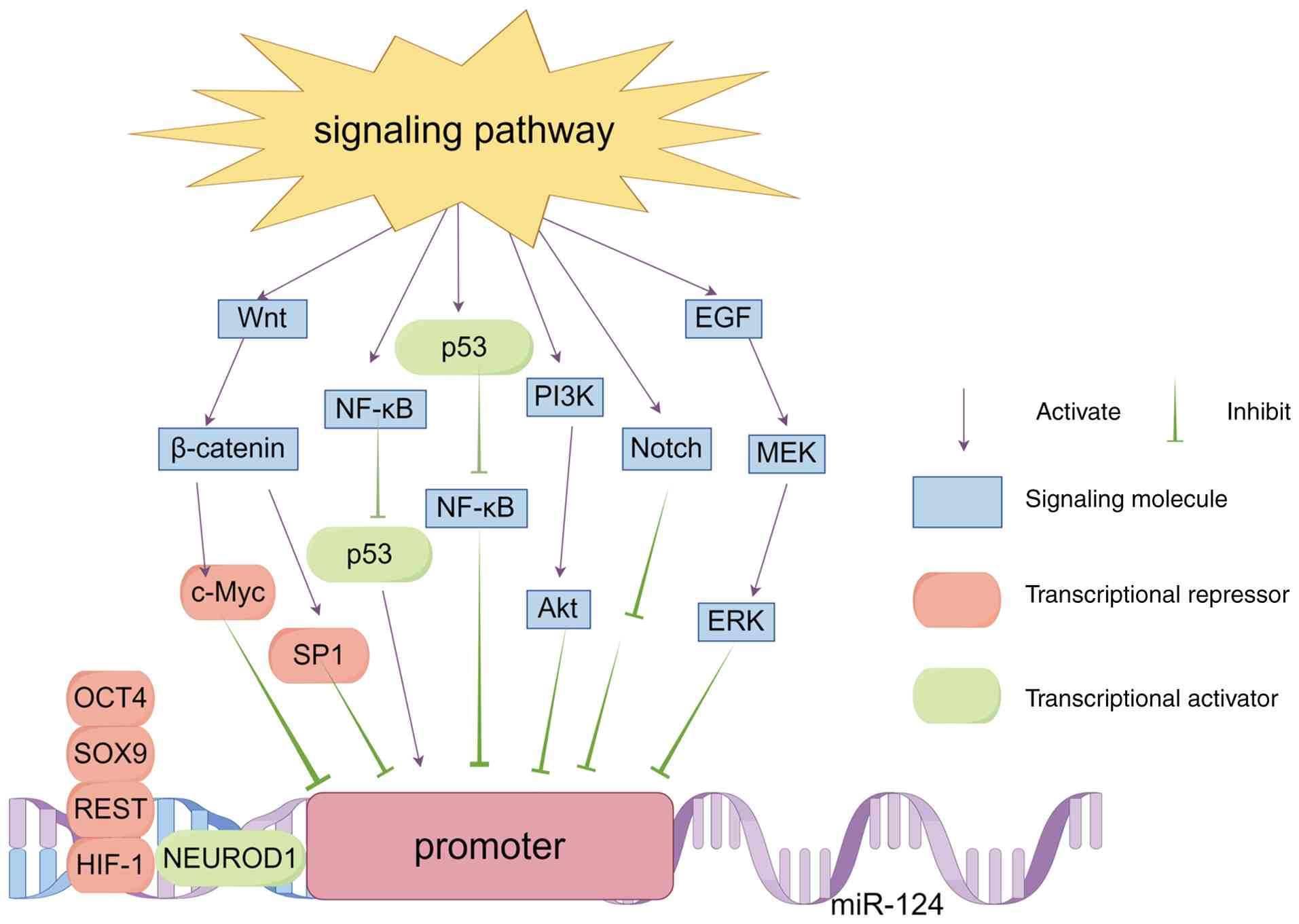 | Figure 2Regulatory network of transcription
factors and upstream signaling pathways controlling miR-124
transcription. This schematic diagram illustrates the sophisticated
network orchestrated by key transcription factors (for instance,
c-Myc, p53 and HIF-1α) and core signaling pathways (for instance,
Wnt/β-catenin, PI3K/Akt and NF-κB), which fine-tunes miR-124
transcription through multiple layers of crosstalk. Dysregulation
of this network and the consequent disruption of miR-124 expression
are important pathogenic events in cancer. This figure was created
using FigDraw. miR, microRNA; HIF-1α, hypoxia inducible factor 1
alpha; NF-κB, nuclear factor kappa B; OCT4, octamer-binding
transcription factor 4; SOX9, SRY-box transcription factor 9;
NEUROD1, neural differentiation factor neuronal differentiation 1;
SP1, specificity protein 1; REST, RE1-silencing transcription
factor. |
Transcriptional and signaling
pathway-mediated upstream regulation of miR-124 in tumors
The expression of miR-124 in tumors is regulated by
a complex network of transcription factors and signaling pathways.
miR-124 regulation can be categorized into repressive and
activating mechanisms at the transcriptional level. Several
cancer-associated transcription factors, such as HIF-1α,
RE1-silencing transcription factor (77) and c-Myc (78,79), directly inhibit miR-124 expression
by binding to cis-regulatory elements, particularly the miR-124
promoter. Similarly, stemness factors, such as SRY-Box
transcription factor 9 (80),
specificity protein 1 (SP1) (81,82) and OCT4 (83), repress transcription by directly
interacting with promoters. Collectively, these mechanisms maintain
cellular stemness, inhibit differentiation, and promote malignant
progression. Conversely, tumor suppressor factors, such as p53
(84) and the neural
differentiation factor neuronal differentiation 1 (85), significantly induce miR-124
expression. The role of p53 is particularly significant, as it
directly activates miR-124 transcription and participates in a
signaling crosstalk network involving the p53-miR-124 axis, NF-κB
and iASPP, thereby affecting therapeutic response (86,87).
In cancer, miR-124 expression is intricately
regulated through various key signaling pathways, forming a complex
repressive network. Evidence suggests that the transcription of
miR-124 is cooperatively suppressed by several core oncogenic
pathways. This upstream regulatory network integrates critical
signaling cascades, including Wnt/β-catenin (88), PI3K/Akt (22,26), Notch (89) and EGF/MEK/ERK (90,91). miR-124 targeting through multiple
pathways highlights its crucial role in maintaining cellular
homeostasis. Consequently, miR-124 inactivation is a critical
mechanism by which cancer cells maintain their malignant phenotype.
Furthermore, NF-κB, a transcription factor and a canonical
inflammatory pathway, negatively regulates miR-124 upon activation.
This regulatory mechanism is particularly important in
tumorigenesis and the establishment of the TME (92-94). By coordinated suppression of their
expression, evolutionarily conserved signaling pathways contribute
to the prevalent silencing of miR-124 in tumors. This regulatory
framework provides a basis for therapeutic targeting of upstream
regulatory nodes of miR-124.
In summary, miR-124 expression in tumors is
meticulously regulated by a complex, finely tuned multi-layered
regulatory network. The frequent silencing of this molecule results
from direct suppression by oncogenic transcription factors, such as
c-Myc and HIF-1α, the functional loss of tumor suppressors, such as
p53, and collaborative suppression through numerous signaling
pathways, including Wnt/β-catenin, PI3K/Akt and NF-κB. There is
extensive crosstalk between these regulatory elements. For
instance, the p53-miR-124 axis regulates NF-κB while concurrently
inhibiting miR-124 expression. Furthermore, signaling pathways such
as Wnt and PI3K/Akt can activate transcription factors, including
c-Myc and SP1, which subsequently increase the transcriptional
repression of miR-124, thereby establishing a multi-layered
regulatory network. A comprehensive analysis of the composition and
dynamic regulation of this network will clarify the crucial role of
miR-124 in tumorigenesis and provide a theoretical basis for
developing restorative therapeutic strategies targeting this
regulatory system.
Epigenetic silencing mechanisms of
miR-124 expression
Epigenetic modification is a crucial mechanism
regulating miR-124 expression (95). At the epigenetic level, DNA
methylation, such as CpG island hypermethylation in promoter
regions, and histone modifications, including repressive
methylation or deacetylation, can silence the miR-124 gene locus.
Furthermore, certain non-coding RNAs, such as lncRNAs, may
indirectly contribute to the silencing or functional inhibition of
miR-124 through mechanisms, such as ceRNA networks (96,97).
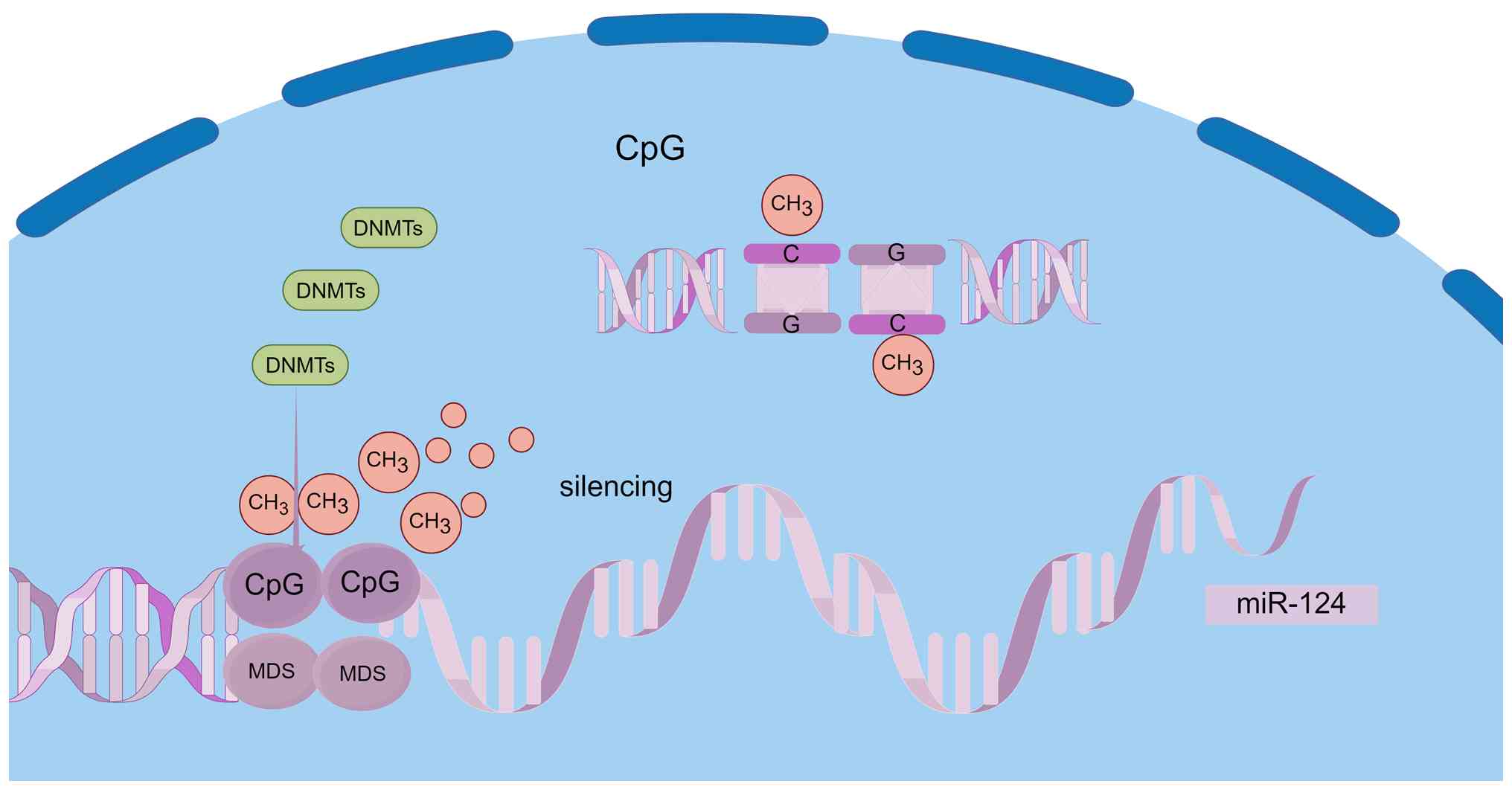
Hypermethylation of CpG islands in the promoter
region of the miR-124 gene is primarily responsible for its
transcriptional silencing. DNA methyltransferases facilitate the
formation of 5-methylcytosine (5-mC), which directly obstructs the
binding of transcription factors to promoter regions. Furthermore,
5-mC recruits repressive complexes, including methyl-CpG-binding
domain proteins, leading to chromatin condensation and subsequent
gene silencing. The molecular mechanism underlying this epigenetic
silencing is illustrated in Fig.
3. Research has demonstrated that miR-124 expression is
silenced through promoter hypermethylation, a critical epigenetic
mechanism, in various cancers, including pancreatic adenocarcinoma,
cutaneous T-cell lymphoma and oral squamous cell carcinoma. This
silencing promotes malignant progression and underscores its
potential as a diagnostic biomarker (98-100).
Histone modifications play a direct, crucial role in
the transcriptional silencing of miR-124. In acute lymphoblastic
leukemia, the miR-124 locus, including miR-124-1, miR-124-2 and
miR-124-3, exhibits a characteristic repressive histone
modification profile. This profile is characterized by enrichment
of repressive histone marks and depletion of activating marks.
Particularly, elevated levels of repressive histone modifications,
including H3K9me2, H3K9me3 and H3K27me3, facilitate the recruitment
of complexes, such as heterochromatin protein 1 and Polycomb
repressive complex 2, thereby promoting heterochromatin formation.
Simultaneously, a significant reduction in activating marks,
including H3K4me3 and acetylated histone H3 (AcH3), maintains a
transcriptionally repressed state. Functionally, the histone
deacetylase inhibitor trichostatin A can reverse the loss of AcH3
and partially restore miR-124 expression. This mechanism of histone
modification-mediated regulation is illustrated in Fig. 4. These histone modifications
synergize with DNA methylation mechanisms, establishing a feedback
loop that stably silences miR-124. Consequently, this silencing
leads to aberrant expression of its target gene, CDK6, contributing
to leukemia progression (101).
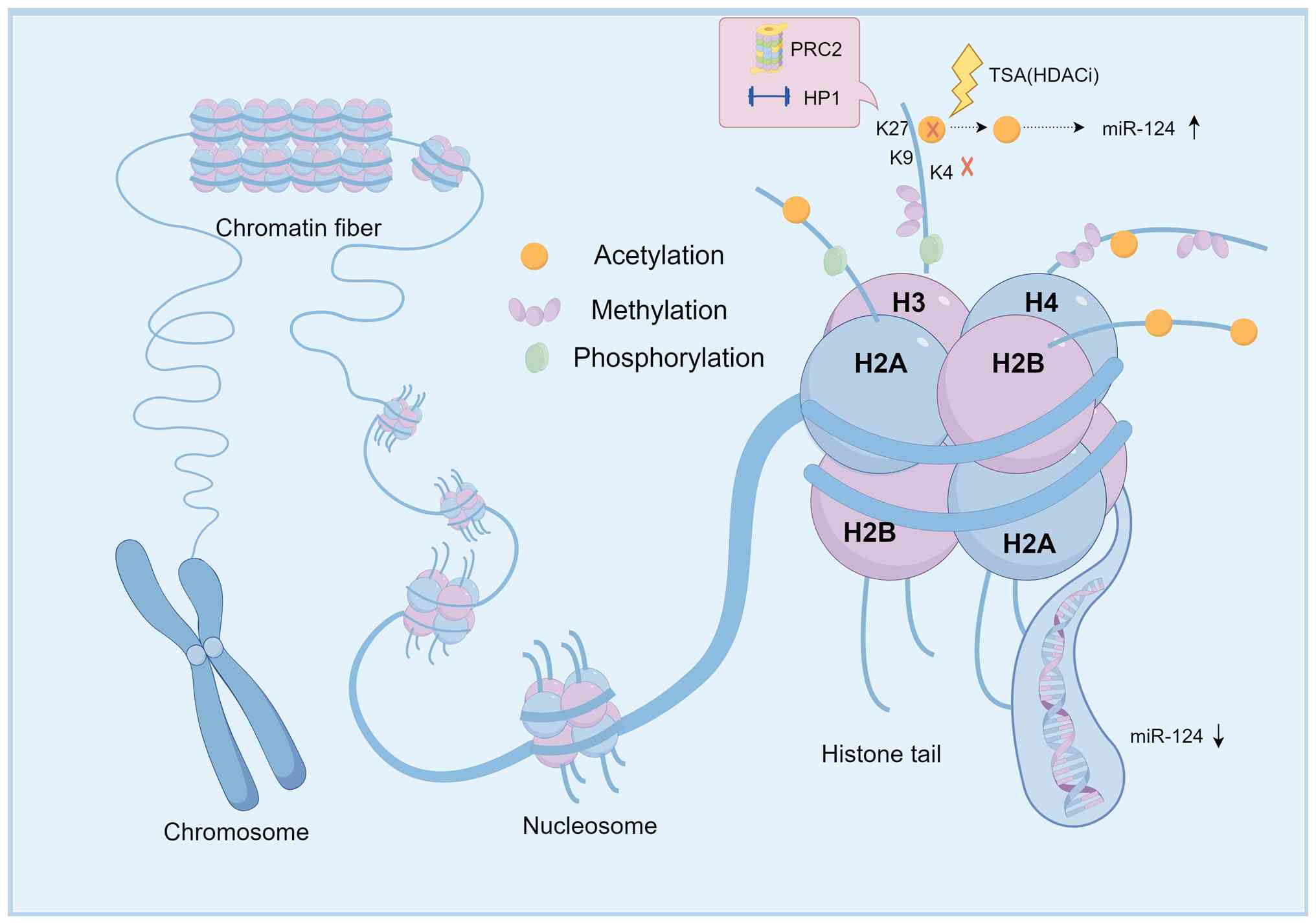 | Figure 4Histone modification-mediated
epigenetic regulation of miR-124 in cancer. This model illustrates
the dynamic balance between activating and repressive histone marks
at the miR-124 locus. Enrichment of repressive marks, including
H3K9me3 and H3K27me3, recruits protein complexes such as HP1 and
PRC2, leading to chromatin condensation and gene silencing.
Conversely, HDAC inhibitors (for instance, TSA) can promote an open
chromatin state by increasing histone acetylation, thereby
reactivating miR-124 expression. This figure was created using
FigDraw. miR, microRNA; HP1, heterochromatin protein 1; PRC2,
polycomb repressive complex 2; HDAC, histone deacetylase; TSA,
trichostatin A. |
Research has demonstrated that the functional
activity of miR-124 is influenced by post-transcriptional
regulation of lncRNAs via a ceRNA mechanism (102). Numerous lncRNAs, including H19
imprinted maternally expressed transcript (103), nuclear paraspeckle assembly
transcript 1 (104), OIP5
antisense RNA 1 (105) and
MALAT1 (72), have been
identified as 'molecular sponges' that sequester miR-124. This
sequestration alleviates miR-124-mediated repression of downstream
target genes and regulates the related signaling pathways. A
schematic representation of this critical regulatory axis of
lncRNA-miR-124 interaction in tumorigenesis is illustrated in
Fig. 5. These findings highlight
the key role of the ceRNA network in the precise regulation of
miR-124 and enhance our understanding of non-coding RNA
interactions, thereby providing novel insights into disease
modulation. Future research should elucidate the context-dependent
dynamics of ceRNA regulation and its modifications in pathological
conditions, and explore the translational potential of ceRNA-based
targeted therapies.
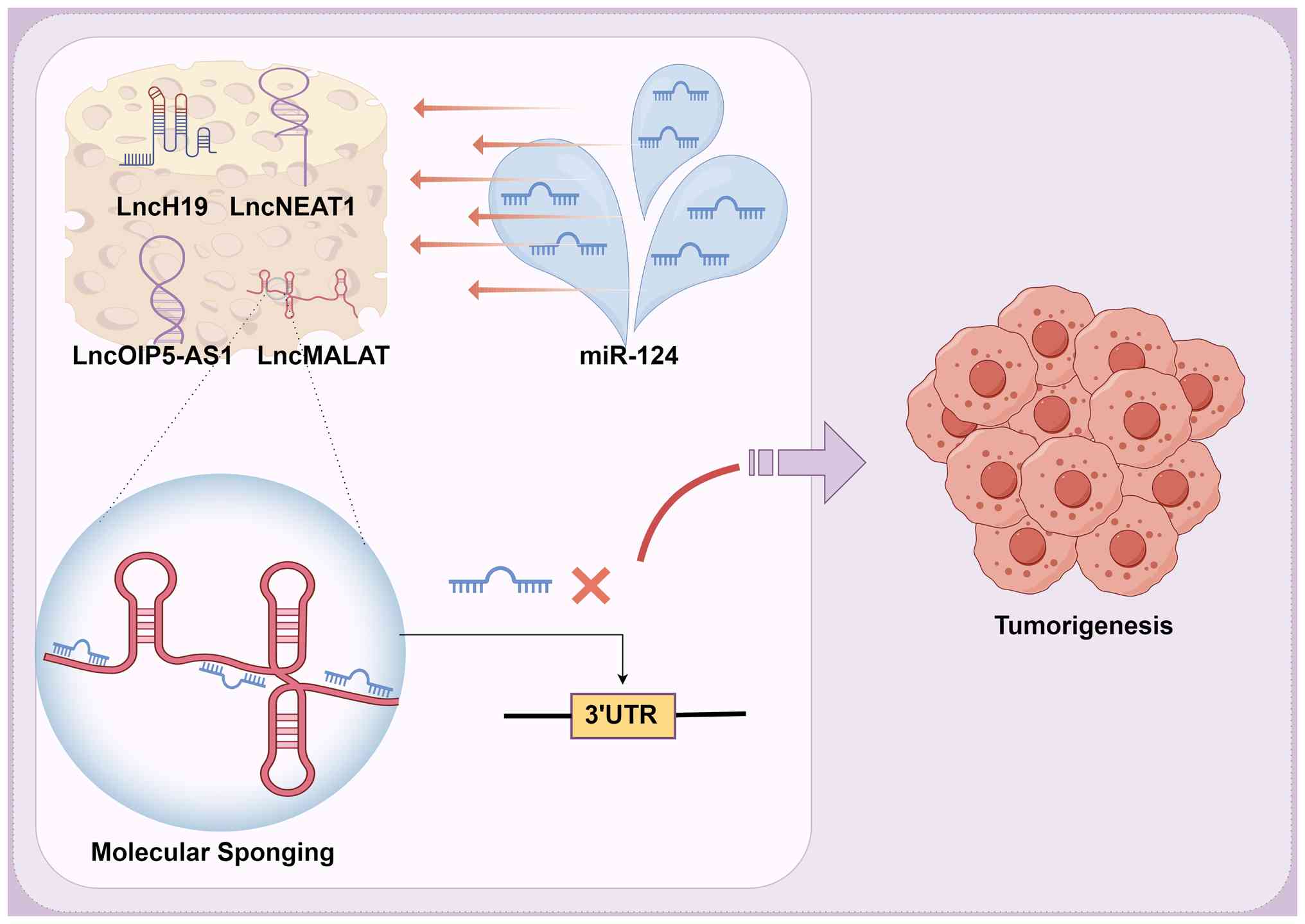 | Figure 5Epigenetic regulation of miR-124 by
lncRNAs as molecular sponges in tumorigenesis. This schematic
illustrates the competing endogenous RNA mechanism, in which
specific lncRNAs, including H19, NEAT1 and MALAT1, act as molecular
sponges to sequester miR-124. This interaction prevents miR-124
from binding its target mRNAs, thereby derepressing oncogenic
signaling pathways and contributing to tumor development. This
figure was created using FigDraw. miR, microRNA; lncRNA, long
non-coding RNA; H19, H19 imprinted maternally expressed transcript;
NEAT1, nuclear paraspeckle assembly transcript 1; UTR, untranslated
region. |
A complex multi-layered network of upstream
regulators that regulate miR-124 expression and activity is
comprehensively summarized in Table
II.
 | Table IIKey upstream regulators of miR-124
expression and activity. |
Table II
Key upstream regulators of miR-124
expression and activity.
| Regulatory
layer | Regulatory
type | Representative
factors | Effect on
miR-124 | Associated
cancers/context |
|---|
| Transcriptional
regulation | Transcription
factors (repressive) | c-Myc 78,79),
HIF-1α (77), REST (77), SOX9 (80), SP1 (81,82), OCT4 (83) | Repression | Ubiquitous in
various cancers |
| Transcription
factors (activatory) | p53 (84), NEUROD1 (85) | Activation | Inactivation
contributes to miR-124 downregulation |
| Signaling
pathways | Wnt/β-catenin
(88), PI3K/Akt 22,26), Notch
(89), EGF/MEK/ERK (90,91), NF-κB (92-94) | Repression | Ubiquitous in
various cancers |
| Epigenetic
regulation | DNA
methylation | Promoter
hypermethylation (by DNA methyltransferases) (98-100) | Repression | Pancreatic cancer,
cutaneous T-cell lymphoma, oral squamous cell carcinoma |
| Histone
modifications | H3K9me3, H3K27me3;
H3K4me3, AcH3 (101) | Repression | Acute lymphoblastic
leukemia |
|
Post-transcriptional regulation (ceRNA
network) | ceRNAs | lncRNAs: H19
(103), NEAT1 (104), OIP5-AS1 (105), MALAT1 (72) | Functional
inhibition (sponging) | Gastric cancer,
glioma, prostate cancer |
Potential clinical applications of miR-124
in cancer therapy
Dual role of miR-124 as a predictive
biomarker and therapeutic target
The significantly reduced expression of miR-124
across various cancers underscores its potential use as a biomarker
for cancer diagnosis and prognostic evaluation (106). The quantification of miR-124
levels in serum or tumor tissue enables early tumor detection and
has the potential to predict patient survival rates and recurrence
risk. Wang et al (107)
reported a significant downregulation of serum miR-124 expression
in patients with bladder cancer. Increased miR-124 levels were
associated with early-stage disease, smaller tumor size, absence of
lymph node metastasis and improved survival outcomes. Moreover,
miR-124 was identified as an independent prognostic factor,
underscoring its potential as a diagnostic and prognostic biomarker
for bladder cancer (107).
Numerous studies in translational research have investigated the
diagnostic potential of miR-124 when used in conjunction with other
miRNAs. For instance, a systematic analysis of 14 miRNAs in the
serum of patients with BC identified a specific panel of miRNAs,
including miR-124, miR-96, miR-183, miR-195, miR-15a and miR-16,
which has significant diagnostic value for differentiating
early-stage BC patients from healthy individuals. This finding was
corroborated by an independent external cohort (n=115), producing
an area under the curve of 0.889 and an accuracy rate of 86.14%.
These results highlight the significant potential of integrating
miR-124 with other miRNAs to improve early detection rates of BC
(108). The clinical relevance
of miR-124 is substantiated by its essential biological functions,
primarily through the modulation of critical signaling pathways. In
addition to its diagnostic applications, miR-124 exhibits
significant promise in modulating chemosensitivity and mitigating
drug resistance. In advanced GC, studies have demonstrated that low
serum miR-124 expression is significantly associated with poor
tumor differentiation, advanced TNM stage and chemotherapy
resistance. Mechanistically, miR-124 modulates chemosensitivity by
downregulating the PI3K/AKT/mTOR signaling pathway. These findings
indicate that miR-124 is a predictive biomarker and a promising
therapeutic target for overcoming drug resistance (109).
In conclusion, miR-124 is a multifunctional molecule
with integrated diagnostic, prognostic and therapeutic guidance
capabilities, offering significant promise for the clinical
management of cancer. However, the therapeutic efficacy of most
related research is currently restricted to the preclinical stage
due to several significant challenges, including suboptimal in
vivo delivery efficiency, insufficient tissue targeting
specificity, and inadequate molecular stability. Future research
should prioritize the development of efficient and safe targeted
delivery systems, such as lipid nanoparticles (LNPs) or exosomal
vectors. Furthermore, it is imperative to elucidate the critical
role of miR-124 within tumor drug resistance networks and its
synergistic interactions with current chemotherapeutic and targeted
agents. Moreover, research should focus on developing precise
therapeutic strategies based on molecular subtyping to identify
patient subgroups that are most likely to benefit from
miR-124-based interventions. Investigating miR-124-based gene
therapies or oligonucleotide drugs will be essential for
translating these findings from the bench to the bedside.
Carrier system-based therapeutic
strategies for miR-124 in tumor treatment
Carrier-based delivery strategies for miR-124 are a
promising development in tumor gene therapy. Current delivery
systems include LNPs, polymeric carriers, inorganic nanoparticles,
and viral vectors, such as adeno-associated virus and lentiviral
vectors, and exosomes. These carriers exhibit structural and
functional diversity, providing a versatile toolkit for delivering
miR-124. Collectively, they facilitate the transition of this
strategy from basic research to clinical applications.
Among synthetic nanocarriers, LNPs have attracted
significant attention owing to their adjustable physicochemical
properties and favorable biocompatibility. For instance, an LNP
system developed with the novel ionizable lipid S-Ac7-DOG
effectively encapsulates and delivers miR-124, significantly
improving its stability and intracellular delivery efficiency
(110). Moreover, liposomal
nanocarriers facilitate co-delivery strategies, such as concurrent
encapsulation of miR-124 and the chemotherapeutic agent SN38,
resulting in synergistic therapeutic effects in the treatment of
HCC (111). Polymeric carriers,
such as cationic polymeric nanoparticles, have been demonstrated to
effectively deliver miR-124 into PC cells, leading to a significant
reduction in tumorigenicity and demonstrating significant
therapeutic potential (112).
Inorganic nanomaterials also have significant potential for miR-124
delivery. Although studies particularly targeting miR-124 are
currently sparse in the literature, various inorganic nanoplatforms
have been successfully used to deliver other functional miRNAs,
such as miR-206. The design principles and technical expertise
derived from these studies offer valuable insights into the
development of miR-124-specific delivery systems. Gold
nanoparticles (AuNPs) exhibit excellent biocompatibility, ease of
functionalization, and distinctive optical properties, which
facilitate efficient miRNA loading and targeted delivery via
surface modifications. Research indicates that PEGylated AuNPs can
efficiently deliver miR-206 mimics, thereby inhibiting BC
progression (113). This
suggests that analogous strategies could be used to improve the
stability and tumor-targeting properties of miR-124. Mesoporous
silica nanoparticles, known for their high stability, adjustable
pore size, and flexible surface chemistry, present opportunities
for high-capacity encapsulation and controlled release of miR-124
(114). Furthermore,
carbon-based materials, including graphene oxide and carbon
nanotubes, have attracted significant interest as innovative
platforms for miRNA delivery owing to their exceptional electrical
properties and easily functionalizable surfaces (115). The combined advantages of these
inorganic carriers, including stability, loading capacity, and
functional versatility, provide a robust foundation for developing
efficient and targeted miR-124 delivery systems.
Compared with synthetic carriers, viral vectors are
distinguished by their superior transfection efficiency. Lentiviral
vectors facilitate stable overexpression of miR-124 in human lung
fibroblasts, which effectively suppress downstream target genes
(85). Conversely, AAV vectors
enable tissue-specific delivery of miR-124, resulting in
significant antitumor effects by inhibiting oncogenes, such as
sphingosine kinase 1, and consequently remodeling the TME (116). Exosomes, endogenous
nanovesicles, offer distinct advantages for targeted delivery. For
instance, exosomes engineered with rabies virus glycoprotein can
cross the blood-brain barrier and deliver miR-124 to regions
affected by cerebral ischemia. This targeted delivery enhances
neural repair and mitigates injury (117), providing proof of concept for
the clinical translation of this strategy.
In summary, critical strategies to enhance the
therapeutic efficacy of miR-124 focus on improving its stability
and bioavailability, as well as on developing efficient and safe
delivery systems. To improve stability, chemical modifications,
such as 2'-O-methylation or phosphorothioate linkages, can be used.
Alternatively, encapsulation within nuclease-resistant carriers can
prolong its half-life. Incorporating functional components, such as
ionizable lipids or cell-penetrating peptides, can improve
bioavailability by increasing cellular uptake and endosomal escape
efficiency. The development of delivery systems requires a balance
between targeting specificity and safety. Surface modification of
carriers with targeting ligands, such as peptides, antibodies, or
aptamers, can increase tumor accumulation while minimizing
off-target effects. Furthermore, stimuli-responsive carriers (for
instance, pH-, reactive oxygen species-, or enzyme-sensitive
materials) facilitate the controlled release of miR-124 within the
TME. It is anticipated that the clinical translation of miR-124 in
precision oncology will be significantly advanced by integrating
personalized delivery strategies with multimodal combination
therapies. The core characteristics, advantages and challenges of
the major delivery systems developed for miR-124 are provided in
Table III.
 | Table IIIDelivery systems for microRNA-124 in
cancer therapy with clinical challenges. |
Table III
Delivery systems for microRNA-124 in
cancer therapy with clinical challenges.
| Delivery
system | Representative
carrier | Core
advantages | Key challenges |
|---|
| LNPs | Ionizable LNP | Highest industrial
maturity, suitable for systemic administration, facile surface
modification | Limited target
specificity, potential acute toxicity |
| Polymeric
nanoparticles | PEI, PLGA | High design
flexibility, capable of stimulus-responsive release | Cytotoxicity
concerns, complex in vivo behavior |
| Inorganic
nanoparticles | AuNPs, AAV | High stability,
potential for theranostic applications | Poor
biodegradability and long-term toxicity risks |
| Viral vectors | AAV,
lentivirus | Sustained and
high-efficiency expression, Inherent tissue tropism | Immunogenicity,
limited cargo capacity, Insertional mutagenesis concerns |
| Exosomes | MSC-derived
exosomes | Innately low
immunogenicity, superior biodistribution and barrier-penetrating
capacity | Major hurdles in
scalable production and quality control, low drug loading
efficiency |
Current status and future challenges of
miR-124 in clinical translation
Although numerous miRNA-based therapeutics have
progressed to phase I/II clinical trials, research particularly
targeting miR-124 remains relatively underdeveloped. Regulatory
bodies, such as the U.S. Food and Drug Administration or the
National Medical Products Administration of China, have not yet
granted clinical approval for any pharmaceutical agent that
directly targets miR-124 or incorporates it as an active
ingredient. Most current studies are limited to the preclinical
stage, with a primary focus on validating miR-124-based diagnostic
markers and therapeutic strategies. These efforts include the
development of nanoparticle delivery systems for conditions such as
HCC and GBM, using both cellular and animal models. Although these
studies provide preliminary evidence suggesting the potential for
clinical translation, numerous critical challenges must be overcome
to realize this potential. The most important challenge is the
delivery system. Serum nucleases can degrade unmodified miRNA
molecules, limiting their ability to selectively access target
cells. Therefore, the development of safe, stable and targeted
delivery carriers, such as LNPs, exosomes, or viral vectors, is a
primary challenge. Furthermore, there is a significant risk of
off-target effects and adverse side effects, as miR-124 can
regulate numerous downstream target genes. Moreover, exogenous RNA
molecules may elicit immunogenic responses, and the in vivo
pharmacokinetic properties, including distribution, metabolism and
stability, require comprehensive evaluation. Furthermore,
establishing large-scale production and quality control systems
that comply with clinical standards remains an unmet requirement.
Consequently, it is anticipated that the clinical translation of
miR-124-based therapies will focus on three principal directions.
First, the integration into precision medicine frameworks will
enable the identification of patient subgroups characterized by low
miR-124 expression through molecular subtyping, thereby enabling
targeted interventions. Second, combination with standard
chemotherapy or targeted agents may improve therapeutic efficacy by
functioning as chemosensitizers or by reversing resistance. Third,
it is expected that advancements in delivery technologies, such as
nanomaterials and engineered exosomes, will accelerate clinical
translation. Despite these challenges, miR-124 remains a highly
promising therapeutic strategy, particularly considering the recent
advancements in RNA therapeutics. The overarching journey from
bench to bedside for miR-124, including its dual roles as a
biomarker and therapeutic target, as well as current delivery
strategies and translational challenges, is comprehensively
summarized in Fig. 6.
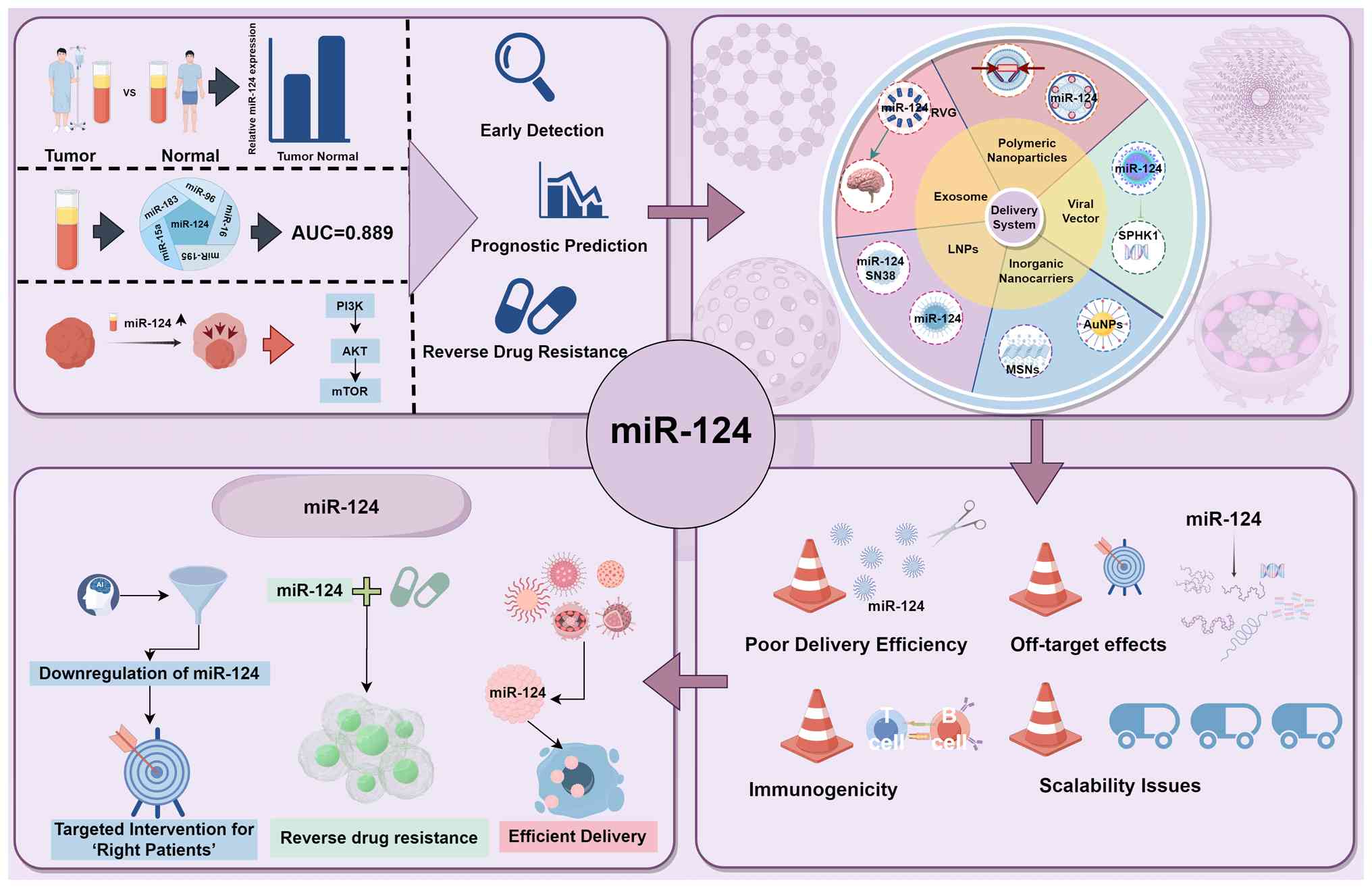 | Figure 6Translational landscape of miR-124 in
cancer therapy: From biological mechanisms to clinical
applications. This schematic integrates the core concepts discussed
in this section, revealing the potential of miR-124 as a
diagnostic/prognostic biomarker and therapeutic agent, currently
developed delivery systems, including LNPs, viral vectors, and
exosomes, key challenges in clinical translation, and future
development prospects. This figure was created using FigDraw. LNPs,
lipid nanoparticles; miR, microRNA; AuNPs, gold nanoparticles;
MSNs, mesoporous silica nanoparticles; AUC, area under the curve;
SPHK1, sphingosine kinase 1; RVG, rabies virus glycoprotein. |
Future perspectives
Prospective studies of miR-124 in oncology present
significant potential, particularly in precision medicine, targeted
therapy, elucidation of underlying mechanisms, and biomarker
development. First, it is anticipated that miR-124-level-based
personalized therapeutic strategies will be developed as
understanding of miR-124 expression profiles across various cancer
types advances. This precision medicine approach could improve
patient response rates to therapy and reduce unnecessary side
effects, thereby improving the overall quality of life. Second, it
is imperative to optimize the targeted delivery systems for miR-124
to ensure its efficient release and high bioavailability within the
TME. Current research primarily focuses on various nanocarriers,
including liposomes, polymeric nanoparticles and viral vectors, to
enhance the specificity and accumulation of miR-124 in tumor
tissues. Furthermore, the development of functionalized
nanoparticles presents innovative strategies to improve miR-124
delivery, potentially enabling controlled release through external
stimuli, such as light or magnetic fields, thereby increasing
treatment precision. In mechanistic studies, it is essential to
elucidate the precise molecular mechanisms by which miR-124
influences tumorigenesis and metastasis. This involves a
comprehensive understanding of its interactions with key signaling
pathways, such as PI3K/Akt, Wnt/β-catenin and NF-κB, as well as the
array of target genes it regulates. These studies will provide
novel insights into tumor biology and may guide the development of
novel targeted therapeutic strategies. Moreover, integrating
miR-124 with conventional chemotherapeutic agents or
immunotherapies represents a promising avenue for research.
Empirical evidence indicates that miR-124 can improve the
sensitivity of tumor cells to specific chemotherapeutic agents,
thereby overcoming drug resistance and improving overall
therapeutic efficacy. Future clinical trials should prioritize the
assessment of the safety and efficacy of such combination therapies
to provide more targeted treatment options for patients with
cancer. Furthermore, miR-124, as a potential biomarker, offers
significant application prospects for early cancer diagnosis,
prognostic evaluation, and therapeutic monitoring. A systematic
analysis of the correlation between miR-124 expression and the
clinical characteristics of patients will provide significant
support for the diagnosis and treatment of cancer. In summary, the
future of miR-124 in cancer research is highly promising. However,
advancing this field will require more in-depth fundamental
research and multidisciplinary collaborations to facilitate its
effective translation from bench to bedside. With the progression
of technologies and ongoing research, miR-124 is poised to emerge
as a novel biomarker and therapeutic target in cancer therapy.
Conclusion
miR-124 is a critical miRNA that has a multifaceted
role in cancer research. Its significant functions include
suppression of tumor growth, regulation of metastasis and cancer
stemness, modulation of drug resistance, remodeling of the TME, and
modulation of immune responses. These diverse functions offer novel
insights into the complex mechanisms underlying tumorigenesis and
cancer progression. Recent studies suggest that miR-124 activity is
modulated by multiple upstream signaling pathways. Moreover, the
broad spectrum of its downstream target genes highlights its
potential as a therapeutic target. Despite these advancements, the
specific roles of miR-124 in different cancer types and its
clinical applications warrant further comprehensive studies. Future
studies should aim to elucidate the precise mechanisms by which
miR-124 affects tumor biology and explore its practical
applications in cancer therapy, including its potential as a
biomarker and in the development of therapeutic strategies. miR-124
is expected to improve the prognosis and quality of life of
patients with cancer by providing novel therapeutic avenues through
the integration of fundamental research and clinical practice. In
conclusion, miR-124 is poised to remain a focus of future research
and clinical practice due to its significant clinical potential and
multifaceted roles in oncology.
Availability of data and materials
Not applicable.
Authors' contributions
JG conceptualized the study, developed methodology,
conducted investigation, performed formal analysis, wrote the
original draft and visualized data. YXL conceptualized and
supervised the study, provided resources, wrote, reviewed and
edited the manuscript, conducted project administration, and
acquired funding. XPY validated data, wrote, reviewed and edited
the manuscript, and supervised the study. YYG performed software
analysis, validated and curated data. PYC conducted investigation
and data curation. WYX performed formal analysis and data
visualization. YXT wrote, reviewed and edited the manuscript. ZYW
provided resources, contributed to the study design, and reviewed
the manuscript. All authors read and approved the final version of
the manuscript. Data authentication is not applicable.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Abbreviations:
|
miR or miRNA
|
microRNA
|
|
TME
|
tumor microenvironment
|
|
CSCs
|
cancer stem cells
|
|
ceRNA
|
competing endogenous RNA
|
|
lncRNA
|
long non-coding RNA
|
|
OCT4
|
octamer-binding transcription factor
4
|
|
EMT
|
epithelial-mesenchymal transition
|
|
SNAI1
|
Snail family transcriptional repressor
1
|
|
PI3K/AKT
|
phosphatidylinositol 3-kinase/protein
kinase b
|
|
PD-L1
|
programmed death-ligand 1
|
|
SP1
|
specificity protein 1
|
|
STAT3
|
signal transducer and activator of
transcription 3
|
|
CDK6
|
cyclin-dependent kinase 6
|
|
CAFs
|
cancer-associated fibroblasts
|
|
NF-κB
|
nuclear factor kappa B
|
|
HIF-1α
|
hypoxia inducible factor 1 alpha
|
|
5-mC
|
5-methylcytosine
|
|
LNPs
|
lipid nanoparticles
|
|
AAV
|
adeno-associated virus
|
|
AuNPs
|
gold nanoparticles
|
Acknowledgements
Not applicable.
Funding
The present study was supported by The National Natural Science
Foundation of China (grant no. 82360609), the Program of Science
and Technology Department of GuiZhou [grant no. Qian Ke He Ji
Chu-ZK(2022)Yiban 619), the Program for High level Innovative
Talents in Guizhou [grant no. QKHRC-CXTD (2025) 046], the Key
Construction Discipline of Immunology and Pathogen biology in
Zhuhai Campus of Zunyi Medical University (grant no.
ZHGF2024-1).
References
|
1
|
Gourishetti K, Balaji Easwaran V, Mostakim
Y, Ranganath Pai KS and Bhere D: MicroRNA (miR)-124: A promising
therapeutic gateway for oncology. Biology (Basel).
12:9222023.PubMed/NCBI
|
|
2
|
Kim H, Lee YY and Kim VN: The biogenesis
and regulation of animal microRNAs. Nat Rev Mol Cell Biol.
26:276–296. 2025. View Article : Google Scholar
|
|
3
|
Sanuki R and Yamamura T: Tumor suppressive
effects of miR-124 and its function in neuronal development. Int J
Mol Sci. 22:59192021. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Liu Y, Yang Y, Wang X, Yin S, Liang B,
Zhang Y, Fan M, Fu Z, Shen C, Han Y, et al: Function of miR-124 in
the pathogenesis of cancer (Review). Int J Oncol. 64:62024.
View Article : Google Scholar
|
|
5
|
Zhou Z, Lv J, Wang J, Yu H, Lu H, Yuan B,
Han J, Zhou R, Zhang X, Yang X, et al: Role of MiR-124 as a
prognostic factor in multiple neoplasms: A meta-analysis. Dis
Markers. 2019:16547802019. View Article : Google Scholar
|
|
6
|
Braga EA, Fridman MV, Burdennyy AM,
Filippova EA, Loginov VI, Pronina IV, Dmitriev AA and Kushlinskii
NE: Regulation of the key epithelial cancer suppressor miR-124
function by competing endogenous RNAs. Int J Mol Sci. 23:136202022.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
López-Urrutia E, Bustamante Montes LP,
Ladrón de Guevara Cervantes D, Pérez-Plasencia C and Campos-Parra
AD: Crosstalk between long Non-coding RNAs, Micro-RNAs and mRNAs:
Deciphering molecular mechanisms of master regulators in cancer.
Front Oncol. 9:6692019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lujambio A, Ropero S, Ballestar E, Fraga
MF, Cerrato C, Setién F, Casado S, Suarez-Gauthier A,
Sanchez-Cespedes M, Git A, et al: Genetic unmasking of an
epigenetically silenced microRNA in human cancer cells. Cancer Res.
67:1424–1429. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Baranwal S and Alahari SK: MicroRNA
control of tumor cell invasion and metastasis. Int J Cancer.
126:1283–1290. 2010. View Article : Google Scholar
|
|
10
|
Liang T, Lu T, Jia W, Li R, Jiang M, Jiao
Y, Wang Y, Cong S, Jiang X, Dong L, et al: Knockdown of lncRNA
MALAT1 induces pyroptosis by regulating the miR-124/SIRT1 axis in
cervical cancer cells. Int J Oncol. 63:1382023. View Article : Google Scholar
|
|
11
|
Chen ZY, Wang XY, Yang YM, Wu MH, Yang L,
Jiang DT, Cai H and Peng Y: LncRNA SNHG16 promotes colorectal
cancer cell proliferation, migration, and epithelial-mesenchymal
transition through miR-124/MCP-1. Gene Ther. 29:193–205. 2022.
View Article : Google Scholar
|
|
12
|
Salehi-Mazandarani S, Mahmoudian-Hamedani
S, Farajzadegan Z and Nikpour P: EZH2: A crucial competing
endogenous RNA in cancer research-A scoping review. Adv Biomed Res.
14:532025. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yueh PF, Chiang IT, Weng YS, Liu YC, Wong
RCB, Chen CY, Hsu JB, Jeng LB, Shyu WC and Hsu FT: Innovative
dual-gene delivery platform using miR-124 and PD-1 via umbilical
cord mesenchymal stem cells and exosome for glioblastoma therapy. J
Exp Clin Cancer Res. 44:1072025. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhao WW, Gao Y, Zhu YT, Zhong FL and Luo
XG: SMYD3 plays a pivotal role in mediating the
epithelial-mesenchymal transition process in breast cancer. Biochem
Biophys Res Commun. 749:1513632025. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Aoyama S, Izumi K, Hiratsuka K, Inaba T,
Koketsu Y, Nakagawa R, Toriumi R, Kamijima T, Kano H, Makino T, et
al: Androgen receptor-controlled miR-124.2 suppresses prostate
cancer progression via CCL2 inhibition. Cancer Sci. 116:2797–2807.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Shi XB, Xue L, Ma AH, Tepper CG,
Gandour-Edwards R, Kung HJ and deVere White RW: Tumor suppressive
miR-124 targets androgen receptor and inhibits proliferation of
prostate cancer cells. Oncogene. 32:4130–4138. 2013. View Article : Google Scholar
|
|
17
|
Peng XH, Huang HR, Lu J, Liu X, Zhao FP,
Zhang B, Lin SX, Wang L, Chen HH, Xu X, et al: MiR-124 suppresses
tumor growth and metastasis by targeting Foxq1 in nasopharyngeal
carcinoma. Mol Cancer. 13:1862014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Luo Y, Wang J, Wang F, Liu X, Lu J, Yu X,
Ma X, Peng X and Li X: Foxq1 promotes metastasis of nasopharyngeal
carcinoma by inducing vasculogenic mimicry via the EGFR signaling
pathway. Cell Death Dis. 12:4112021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kim SH and Singh SV: The role of MiR-124
in breast cancer stem cell inhibition by benzyl isothiocyanate.
Pharm Res. 41:1921–1932. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yu B, Jiang K and Zhang J: MiR-124
suppresses growth and aggressiveness of osteosarcoma and inhibits
TGF-β-mediated AKT/GSK-3β/SNAIL-1 signaling. Mol Med Rep.
17:6736–6744. 2018.PubMed/NCBI
|
|
21
|
Wang T, Xu Y, Li S, Du R, Shi J, Jiang C,
Wang R and Zhu Y: Novel small-molecule miR-124 inducer acts as 'a
Physiological Brake' of inflammation in ulcerative colitis by
targeting the PIK3R2/PI3K/Akt axis. J Med Chem. 68:14114–14126.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang C, Fu R, Wang Y, Wei J, Yu Y, Hu L
and Zhang C: miR-124 and miR-194-5p regulation of the PI3K/AKT
pathway via ROR2 in medulloblastoma progression. Cancer Gene Ther.
31:941–954. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Li X, Fan Q, Li J, Song J and Gu Y:
MiR-124 down-regulation is critical for cancer associated
fibroblasts-enhanced tumor growth of oral carcinoma. Exp Cell Res.
351:100–108. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Cai WL, Huang WD, Li B, Chen TR, Li ZX,
Zhao CL, Li HY, Wu YM, Yan WJ and Xiao JR: miR-124 inhibits bone
metastasis of breast cancer by repressing Interleukin-11. Mol
Cancer. 17:92018. View Article : Google Scholar
|
|
25
|
Zhang S, Liang Y, Xia M, Tian X, Chen Z,
Lin L, Liang J and Liu Y: Serum exosomal microRNA profiling reveals
a down-regulation of hsa-miR-124 in patients with severe acne.
Front Immunol. 16:15548112025. View Article : Google Scholar
|
|
26
|
Zhu Q, Zhang Y, Li M, Zhang Y, Zhang H,
Chen J, Liu Z, Yuan P, Yang Z and Wang X: MiR-124 impedes the
metastasis of non-small cell lung cancer via extracellular exosome
transport and intracellular PI3K/AKT signaling. Biomark Res.
11:12023. View Article : Google Scholar
|
|
27
|
Hu YZ, Hu ZL, Liao TY, Li Y and Pan YL:
LncRNA SND1-IT1 facilitates TGF-β1-induced
epithelial-to-mesenchymal transition via miR-124/COL4A1 axis in
gastric cancer. Cell Death Discov. 8:732022. View Article : Google Scholar
|
|
28
|
Zhang Y, Li X, Zhang J and Liang H:
Natural killer T cell cytotoxic activity in cervical cancer is
facilitated by the LINC00240/microRNA-124-3p/STAT3/MICA axis.
Cancer Lett. 474:63–73. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhou W, Van Sinderen M, Rainczuk K,
Menkhorst E, Sorby K, Osianlis T, Pangestu M, Santos L, Rombauts L,
Rosello-Diez A and Dimitriadis E: Dysregulated miR-124 in
endometrial epithelial cells reduces endometrial receptivity by
altering polarity and adhesion. Proc Natl Acad Sci USA.
121:e24010711212024. View Article : Google Scholar
|
|
30
|
Zhao Q, Jiang F, Zhuang H, Chu Y, Zhang F
and Wang C: MicroRNA miR-124 suppresses proliferation and
epithelial-mesenchymal transition of hepatocellular carcinoma via
ARRDC1 (arrestin domain containing 1). Bioengineered. 13:8255–8265.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Hou F, Shi DB, Guo XY, Zhao RN, Zhang H,
Ma RR, He JY and Gao P: HRCT1, negatively regulated by miR-124,
promotes tumor metastasis and the growth of gastric cancer by
activating the ERBB2-MAPK pathway. Gastric Cancer. 26:250–263.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Deng Q, Hua A, Zhao Q, Zhang Z, Yang T,
Wang Q, Yang X and Li Z: Modulating tumor acidity with hydroxyethyl
starch-based nanoparticles by targeting CA9 to eliminate cancer
stem cells and overcome immunosuppression. Biomaterials.
324:1235012026. View Article : Google Scholar
|
|
33
|
Wang L, Zhu Y, Huang C, Pan Q, Wang J, Li
H, Huang Y, Yi G, Li Z, Qi S, et al: Targeting ferroptosis in
cancer stem cells: A novel strategy to improve cancer treatment.
Genes Dis. 12:1016782025. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Mao X, Peng S, Lu Y and Song L: Regulatory
functions of microRNAs in cancer stem cells: Mechanism, facts, and
perspectives. Cells. 14:10732025. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Shukla D, Mishra S, Mandal T, Charan M,
Verma AK, Khan MMA, Chatterjee N, Dixit AK, Ganesan SK, Ganju RK
and Srivastava AK: MicroRNA-379-5p attenuates cancer stem cells and
reduces cisplatin resistance in ovarian cancer by regulating
RAD18/Polη axis. Cell Death Dis. 16:1402025. View Article : Google Scholar
|
|
36
|
Li Y, Zhang T, Korkaya H, Liu S, Lee HF,
Newman B, Yu Y, Clouthier SG, Schwartz SJ, Wicha MS and Sun D:
Correction: Sulforaphane, a dietary component of broccoli/broccoli
sprouts, inhibits breast cancer stem cells. Clin Cancer Res.
31:20622025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Tian Y, Tian Y, Tu Y, Zhang G, Zeng X, Lin
J, Ai M, Mao Z, Zheng R and Yuan Y: miR-124 inhibits stem-like
properties and enhances radiosensitivity in nasopharyngeal
carcinoma cells via direct repression of expression of JAMA. J Cell
Mol Med 2020. Sep;24:9533–9544. 2020. View Article : Google Scholar
|
|
38
|
Feng Y, Jiang W, Zhao W, Lu Z, Gu Y and
Dong Y: miR-124 regulates liver cancer stem cells expansion and
sorafenib resistance. Exp Cell Res. 394:1121622020. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Tuoheti M, Li J, Zhang C, Gao F, Wang J
and Wu Y: MiR-124-3p inhibits cell stemness in glioblastoma via
targeting EPHA2 through ALKBH5-mediated m6A modification. Hum Cell.
38:102024. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Sha J, Xia L, Han Q, Chi C, Zhu Y, Pan J,
Huang Y, Xia W, Dong B, Xue W and Yang C: Erratum: Downregulation
of circ-TRPS1 suppressed prostatic cancer prognoses by regulating
miR-124-3p/EZH2 axis-mediated stemness. Am J Cancer Res.
12:5694–5695. PubMed/NCBI
Erratum. Am J Cancer Res. 10:4372–4385.
2020.
|
|
41
|
Li Z, Wang X, Li W, Wu L, Chang L and Chen
H: miRNA-124 modulates lung carcinoma cell migration and invasion.
Int J Clin Pharmacol Ther. 54:603–612. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Park MN, Kim M, Lee S, Kang S, Ahn CH,
Tallei TE, Kim W and Kim B: Targeting redox signaling through
exosomal MicroRNA: Insights into tumor microenvironment and
precision oncology. Antioxidants (Basel). 14:5012025. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wei W, Li J, Huang J, Jiang Q, Lin C, Hu
R, Wei J, Li Q, Xu G and Chang Z: Exosomal miR-3681-3p from
M2-polarized macrophages confers cisplatin resistance to gastric
cancer cells by targeting MLH1. Mol Med Rep. 31:942025. View Article : Google Scholar
|
|
44
|
Çerçi Alkaç B, Soyöz M, Kılıçaslan Ayna T,
Pehlivan M and Pirim İ: Antiproliferative and apoptotic effects of
mono/combined treatment of abemaciclib and regulation of
neuroblastoma-related miRNAs. FASEB J. 39:e707602025. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Liu YX, Wang L, Liu WJ, Zhang HT, Xue JH,
Zhang ZW and Gao CJ: MiR-124/B4GALT1 axis plays an important role
in SOCS3-regulated growth and chemo-sensitivity of CML. J Hematol
Oncol. 9:692016. View Article : Google Scholar
|
|
46
|
Dong ZB, Wu HM, He YC, Huang ZT, Weng YH,
Li H, Liang C, Yu WM and Chen W: MiR-124.1 sensitizes
hepatocellular carcinoma cells to sorafenib by regulating FOXO3a by
targeting AKT2 and SIRT1. Cell Death Dis. 13:352022. View Article : Google Scholar
|
|
47
|
Xie C, Zhang LZ, Chen ZL, Zhong WJ, Fang
JH, Zhu Y, Xiao MH, Guo ZW, Zhao N, He X and Zhuang SM: A
hMTR4-PDIA3P1-miR-125/124-TRAF6 regulatory axis and its function in
NF kappa B signaling and chemoresistance. Hepatology. 71:1660–1677.
2020. View Article : Google Scholar :
|
|
48
|
Shi Z, Chen Q, Li C, Wang L, Qian X, Jiang
C, Liu X, Wang X, Li H, Kang C, et al: MiR-124 governs glioma
growth and angiogenesis and enhances chemosensitivity by targeting
R-Ras and N-Ras. Neuro Oncol. 16:1341–1353. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Wei Y, Wang P, Zhao J, Fan X, Jiang J, Mu
X, Wang Y, Yang A, Zhang R, Hu S and Guo Z: Overexpression of
miR-124 enhances the therapeutic benefit of TMZ treatment in the
orthotopic GBM mice model by inhibition of DNA damage repair. Cell
Death Dis. 16:472025. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Smith MA, Chiacchia S, Boehme J, Datar SA,
Morell E, Keller RL, Romer A, Colglazier E, Parker C, Becerra J and
Fineman JR: MicroRNA in pediatric pulmonary hypertension microRNA
profiling to inform disease classification, severity, and treatment
response in pediatric pulmonary hypertension. Am J Physiol Heart
Circ Physiol. 328:H47–H57. 2025. View Article : Google Scholar :
|
|
51
|
Liu C, Xing H, Guo C, Yang Z and Wang Y
and Wang Y: MiR-124 reversed the doxorubicin resistance of breast
cancer stem cells through STAT3/HIF-1 signaling pathways. Cell
Cycle. 18:2215–2227. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Li D, Dai Y, Li Z, Bi H, Li H, Wang Y, Liu
Y, Tian X and Chen L: Resveratrol upregulates miR-124-3p expression
to target DAPK1, regulating the NLRP3/Caspase-1/GSDMD pathway to
inhibit pyroptosis and alleviate spinal cord injury. J Cell Mol
Med. 29:e703382025. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Song Y, Du Y, Qin C, Liang H, Yang W, Lin
J, Ding M, Han J and Xu T: Gemcitabine-resistant biomarkers in
bladder cancer are associated with tumor-immune microenvironment.
Front Cell Dev Biol. 9:8096202022. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Ren X, Zhang L, Zhang Y, Li Z, Siemers N
and Zhang Z: Insights gained from single-cell analysis of immune
cells in the tumor microenvironment. Annu Rev Immunol. 39:583–609.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Yuan S, Almagro J and Fuchs E: Beyond
genetics: Driving cancer with the tumour microenvironment behind
the wheel. Nat Rev Cancer. 24:274–286. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Liu H, Tang L, Li Y, Xie W, Zhang L, Tang
H, Xiao T, Yang H, Gu W, Wang H and Chen P: Nasopharyngeal
carcinoma: current views on the tumor microenvironment's impact on
drug resistance and clinical outcomes. Mol Cancer. 23:202024.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Li H, Jiang L, Zhang S, Miao X and Jiang
SH: Pan-cancer analysis reveals multifaceted roles of
nonmyelinating Schwann cells in gastrointestinal cancers. Cancer
Lett. 628:2178502025. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Rasul KH, Hameed EO, Karim KJ, Moshari S
and Nahali A: Divergent glucose stress responses in TM4 Sertoli
cells: A miRNA-driven regulatory landscape of inflammation and
cytokine expression. Mol Biol Rep. 52:8172025. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Dasgupta D, Ghosh S, Dey I, Majumdar S,
Chowdhury S, Das S, Banerjee S, Saha M, Ghosh A, Roy N, et al:
Influence of polymorphisms in TNF-α and IL1β on susceptibility to
alcohol induced liver diseases and therapeutic potential of miR-124
impeding TNF-α/IL1β mediated multi-cellular signaling in liver
microenvironment. Front Immunol. 14:12417552023. View Article : Google Scholar
|
|
60
|
Asl ER, Hosseini SE, Tahmasebi F, Bolandi
N and Barati S: MiR-124 and MiR-155 as therapeutic targets in
microglia-mediated inflammation in multiple sclerosis. Cell Mol
Neurobiol. 45:632025. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Li Z, Cheng Q, Lin L, Fu X and Wang Y:
Plasma membrane-derived biomimetic apoptotic nanovesicles targeting
inflammation and cartilage degeneration for osteoarthritis. Small
Methods. 9:e24006602025. View Article : Google Scholar
|
|
62
|
Xu Q, Shi MF, Han YF, Liu MY, Liu XB, Ma
XN, Feng W, Lin CS and Liu QP: Kunduan Yimu Decoction affected
Th17/Treg balance through microRNA-124 to improve rheumatoid
arthritis pathology. Phytomedicine. 135:1561292024. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Shi P, Liu Y, Yang H and Hu B: Breast
cancer derived exosomes promoted angiogenesis of endothelial cells
in microenvironment via circHIPK3/miR-124/MTDH axis. Cell Signal.
95:1103382022. View Article : Google Scholar
|
|
64
|
Huang F, Bu L, Li M, Wang Y, Zhang R, Shao
Y, Lin K, Yang H, Zhao Q and Liu L: HuR/miR-124/VDR complex bridges
lipid metabolism and tumor development in colorectal cancer.
Carcinogenesis. 46:bgae0612025. View Article : Google Scholar
|
|
65
|
Dhar P, Moodithaya S, Patil P and Adithi
K: A hypothesis: MiRNA-124 mediated regulation of sirtuin 1 and
vitamin D receptor gene expression accelerates aging. Aging Med
(Milton). 7:320–327. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Zhu Y, Zou W, Sun B, Shen K, Xia F, Wang
H, Jiang F and Lu Z: Ginsenoside Rg1 regulates the activation of
astrocytes through lncRNA-Malat1/miR-124/Lamc1 axis driving
PI3K/AKT signaling pathway, promoting the repair of spinal cord
injury. CNS Neurosci Ther. 30:e701032024. View Article : Google Scholar
|
|
67
|
De Jong MME, Fokkema C, Papazian N, Czeti
Á, Appelman MK, Vermeulen M, van Heusden T, Hoogenboezem RM, van
Beek G, Tahri S, et al: An IL-1β-driven neutrophil-stromal cell
axis fosters a BAFF-rich protumor microenvironment in individuals
with multiple myeloma. Nat Immunol. 25:820–833. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Raza ML, Fatima M, Rawalia MA and Raza R:
A narrative review on exosomes therapeutics in stroke: Advancing
neuroprotection and regeneration. Neuroscience. 584:311–322. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Zhou WJ, Zhang J, Xie F, Wu JN, Ye JF,
Wang J, Wu K and Li MQ: CD45RO-CD8+ T cell-derived exosomes
restrict estrogen-driven endometrial cancer development via the
ERβ/miR-765/PLP2/Notch axis. Theranostics. 11:5330–5345. 2021.
View Article : Google Scholar
|
|
70
|
Wei J, Wang F, Kong LY, Xu S, Doucette T,
Ferguson SD, Yang Y, McEnery K, Jethwa K, Gjyshi O, et al: miR-124
inhibits STAT3 signaling to enhance T cell-mediated immune
clearance of glioma. Cancer Res. 73:3913–3926. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Zhang DH, Wang R, Liu L, Zhang P, Zhao JH
and Liang B: Comprehensive analysis of pyroptosis-related gene
signatures in renal fibrosis. Clin Exp Nephrol. 29:1764–1782. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Liu Z, Lin X, Tu Y, Zhou Y, Huang M and
Fang C: MALAT1 promotes microglia activation and neuronal apoptosis
through via the miR-124/SGK1 axis mediating experimental autoimmune
encephalomyelitis disease progression in mice. Int Immunopharmacol.
152:1144172025. View Article : Google Scholar
|
|
73
|
Sharen G, Meng H, Zhang L, Liu K, Wang Y
and Zhao D: LncRNA XIST enhances gastric cancer cell function by
regulating STAT3/PD-L1 axis as a competing endogenous RNA for
miR-124. Am J Cancer Res. 15:1597–1613. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Qiu B, Xu X, Yi P and Hao Y: Curcumin
reinforces MSC-derived exosomes in attenuating osteoarthritis via
modulating the miR-124/NF-kB and miR-143/ROCK1/TLR9 signalling
pathways. J Cell Mol Med. 24:10855–10865. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Gong X, Jia L, Zhou L and Hu T: USP14
predicts poorer survival outcomes and promotes tumor progression in
endometrial carcinoma by activating NF-κB signaling. Aging (Albany
NY). 15:12120–12135. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Meng Q, Pan B and Sheng P: Histone
deacetylase 1 is increased in rheumatoid arthritis synovium and
promotes synovial cell hyperplasia and synovial inflammation in the
collagen-induced arthritis mouse model via the miR-124-dependent
MARCKS-JAK/STAT axis. Clin Exp Rheumatol. 39:970–981. 2021.
View Article : Google Scholar
|
|
77
|
Rasheed M, Asghar R, Firdoos S, Ahmad N,
Nazir A, Ullah KM, Li N, Zhuang F, Chen Z and Deng Y: A systematic
review of circulatory microRNAs in major depressive disorder:
Potential biomarkers for disease prognosis. Int J Mol Sci.
23:12942022. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Seviour EG, Sehgal V, Lu Y, Luo Z, Moss T,
Zhang F, Hill SM, Liu W, Maiti SN, Cooper L, et al: Functional
proteomics identifies MicroRNAs to target a p27/Myc/phospho-Rb
signature in breast and ovarian cancer. Oncogene. 35:691–701. 2016.
View Article : Google Scholar :
|
|
79
|
Liu K, Lei S, Kuang Y, Jin Q, Long D, Liu
C, Jiang Y, Zhao H and Yao H: A novel mechanism of the c-Myc/NEAT1
axis mediating colorectal cancer cell response to photodynamic
therapy treatment. Front Oncol. 11:6528312021. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Li S, Liu J, Liu S, Jiao W and Wang X:
Mesenchymal stem cell-derived extracellular vesicles prevent the
development of osteoarthritis via the circHIPK3/miR-124-3p/MYH9
axis. J Nanobiotechnology. 19:1942021. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Nguyen HD: Effects of a mixture of ambient
air pollution and its metabolites on depression: From epidemiology
to molecular mechanisms. J Environ Sci (China). 158:405–419. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Wu M, Huang Z, Huang W, Lin M, Liu W, Liu
K and Li C: miR-124 attenuates myocardial injury in sepsis via
modulating SP1/HDAC4/HIF-1α axis. Cell Death Discov. 8:402022.
View Article : Google Scholar
|
|
83
|
Prajapati KS, Shuaib M, Gupta S and Kumar
S: Withaferin A mediated changes of MicroRNA expression in breast
cancer-derived mammospheres. Mol Carcinog. 61:876–889. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Cates K, Yuan L, Yang Y and Yoo AS: Fate
erasure logic of gene networks underlying direct neuronal
conversion of somatic cells by microRNAs. Cell Rep. 44:1151532025.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Mseis-Jackson N, Jiang M, Sharma M,
Ranchod A, Williams C, Chen X and Li H: Dynamic regulation of
NeuroD1 expression level by a novel viral construct during
astrocyte-to-neuron reprogramming. Mol Ther Nucleic Acids.
36:1027042025. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Jeong D, Kim J, Nam J, Sun H, Lee YH, Lee
TJ, Aguiar RC and Kim SW: MiR-124 links p53 to the NF-κB pathway in
B-cell lymphomas. Leukemia. 29:1868–1874. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Liu K, Chen W, Lei S, Xiong L, Zhao H,
Liang D, Lei Z, Zhou N, Yao H and Liang Y: Wild-type and mutant p53
differentially modulate miR-124/iASPP feedback following
pohotodynamic therapy in human colon cancer cell line. Cell Death
Dis. 8:e30962017. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
He L, Zhu C, Jia J, Hao XY, Yu XY, Liu XY
and Shu MG: ADSC-Exos containing MALAT1 promotes wound healing by
targeting miR-124 through activating Wnt/β-catenin pathway. Biosci
Rep. 40:BSR201925492020. View Article : Google Scholar
|
|
89
|
Burbach KF and Yoo AS: Notch inhibition
enhances morphological reprogramming of microRNA-Induced human
neurons. bioRxiv (Preprint). 2024.01.12.575384. 2024.
|
|
90
|
Korkmaz Bayram K, Bayram A, Yilmaz
Sukranli Z, Mehmetbeyoglu Duman E, Aybuga F, Tufan Benli E, Taheri
S, Ozkul Y and Rassoulzadegan M: Dynamic interplay between
miR-124-3p and EGF in the regulation of overgrowth via RNA
signaling. Biomolecules. 15:11862025. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Yang L, Zhu Y, Kong D, Gong J, Yu W, Liang
Y, Nie Y and Teng CB: EGF suppresses the expression of miR-124a in
pancreatic β cell lines via ETS2 activation through the MEK and
PI3K signaling pathways. Int J Biol Sci. 15:2561–2575. 2019.
View Article : Google Scholar
|
|
92
|
Ramar V, Guo S, Wang G and Liu M: The
pivotal role of NF-kappaB in glioblastoma: mechanisms of activation
and therapeutic implications. Int J Mol Sci. 26:78832025.
View Article : Google Scholar
|
|
93
|
Mao H, Zhao X and Sun SC: NF-κB in
inflammation and cancer. Cell Mol Immunol. 22:811–839. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Ge X, Guo M, Hu T, Li W, Huang S, Yin Z,
Li Y, Chen F, Zhu L, Kang C, et al: Increased microglial exosomal
miR-124 alleviates neurodegeneration and improves cognitive outcome
after rmTBI. Mol Ther. 28:503–522. 2020. View Article : Google Scholar
|
|
95
|
Pulliero A, Cassatella G, Astuni P, Khalid
Z, Fiordoro S and Izzotti A: The role of microRNA expression and
DNA Methylation in HPV-Related cervical cancer: A systematic
review. Int J Mol Sci. 25:127142024. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Buttà M, Serra N, Sucato A, Cabibi D,
Campisi G, Panzarella V, Alfedi G, Pistoia D and Capra G: The role
of methylation as an epigenetic marker in HPV-related oral lesions.
J Med Virol. 97:e704592025. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Aschner M, Skalny AV, Santamaria A, Rocha
JBT, Mansouri B, Tizabi Y, Madeddu R, Lu R, Lee E and Tinkov AA:
Epigenetic mechanisms of aluminum-induced neurotoxicity and
Alzheimer's disease: A focus on Non-Coding RNAs. Neurochem Res.
49:2988–3005. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Du X, Yi X, Zou X, Chen Y, Tai Y, Ren X
and He X: PCDH1, a poor prognostic biomarker and potential target
for pancreatic adenocarcinoma metastatic therapy. BMC Cancer.
23:11022023. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
García-Colmenero L, González J, Sandoval
J, Guillén Y, Diaz-Lagares A, Andrades E, Iglesias A, Nonell L,
Pujol RM, Bigas A, et al: Epigenetic silencing of tumor suppressor
miR-124 directly supports STAT3 activation in cutaneous T-Cell
Lymphoma. Cells. 9:26922020. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Liao SW, Liao XH, Wu SH, Li YF, Chen PY,
Wang YL, Lu YC and Tai CK: Methylation-mediated silencing of
miR-124-3 regulates LRRC1 expression and promotes oral cancer
progression. Cancers (Basel). 17:11362025. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Agirre X, Vilas-Zornoza A, Jiménez-Velasco
A, Martin-Subero JI, Cordeu L, Gárate L, San José-Eneriz E,
Abizanda G, Rodríguez-Otero P, Fortes P, et al: Epigenetic
silencing of the tumor suppressor microRNA Hsa-miR-124a regulates
CDK6 expression and confers a poor prognosis in acute lymphoblastic
leukemia. Cancer Res. 69:4443–4453. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Deng L, Tang Q, Lv H and Liu Z: LncRNA
HNF1A-AS1 inhibits ferroptosis in oral squamous cell carcinoma
cells through the miR-124/PKM2 axis. J Stomatol Oral Maxillofac
Surg. 126(6S): 1025222025. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Zhang X, Cai Y, Sit BHM, Jian RX, Malki Y,
Zhang Y, Ong CCY, Li Q, Lam RPK and Rainer TH: Cell-free nucleic
acids for early diagnosis of acute ischemic stroke: A systematic
review and meta-analysis. Int J Mol Sci. 26:15302025. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Chen G, Zhang L, Wang Y, Wang J, Yang K,
Wang X and Chen X: The NEAT1/miR-124-3p/CCL2 axis in chronic kidney
disease progression: Integrated bioinformatics analysis and
experimental validation. Epigenomics. 17:935–952. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Li L, Ma Y, Maerkeya K, Reyanguly D and
Han L: LncRNA OIP5-AS1 regulates the warburg effect through
miR-124-5p/IDH2/HIF-1α pathway in cervical cancer. Front Cell Dev
Biol. 9:6550182021. View Article : Google Scholar
|
|
106
|
Bellon M, Moles R, Chaib-Mezrag H,
Pancewicz J and Nicot C: JAG1 overexpression contributes to Notch1
signaling and the migration of HTLV-1-transformed ATL cells. J
Hematol Oncol. 11:1192018. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Wang D, Zhang QJ and Wu J: The
relationship between serum miR-124 and miR-320a levels and
clinicopathological characteristics as well as survival prognosis
in patients with bladder cancer. International Journal of
Laboratory Medicine. 45:608–613. 2024. View Article : Google Scholar
|
|
108
|
Borsos BN, Páhi ZG and Pankotai T:
Exploring potent miRNA combinations for detecting early-stage
breast cancer. Mol Ther Nucleic Acids. 36:1026212025. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Liu HQ, Zhang YF, Zhang F, et al: The
relationship between serum miR-124 and miR-361-5p levels and
clinicopathological characteristics, chemotherapy sensitivity, and
the PI3K/AKT/mTOR signaling pathway in patients with advanced
gastric cancer. Chinese Journal of Difficult and Complicated Cases.
22:42–48. 2023. View Article : Google Scholar
|
|
110
|
Ma Z, Dang HA, Yang J, Rodella G, Mwema A,
De Lombaerde E, Chen Y, De Geest BG, van Pesch V, Muccioli GG and
des Rieux A: Lipid nanoparticle-mediated delivery of microRNA-124
reduces neuroinflammation. Biomaterials. 325:1235892026. View Article : Google Scholar
|
|
111
|
Wu H: Study on Co-delivery of SN38 and
miR-124 Based on a Liposomal Nanodrug Delivery System for Combined
Therapy of Hepatocellular Carcinoma (D). Zhejiang Province:
Zhejiang University; 2021, View Article : Google Scholar
|
|
112
|
Conte R, Valentino A, Di Cristo F, Peluso
G, Cerruti P, Di Salle A and Calarco A: Cationic polymer
nanoparticles-mediated delivery of miR-124 impairs tumorigenicity
of prostate cancer cells. Int J Mol Sci. 21:8692020. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Ji Z, Cheng S, Li W, Xing Y, Tang Z, Zhu
X, Wang D, Hao C, Wang B and Shi M: Ultrasensitive detection of
miR-31 using a signal-on electrochemiluminescence biosensor based
on CRISPR/Cas12a and MXene nanocomposites. Bioelectrochemistry.
167:1090592026. View Article : Google Scholar
|
|
114
|
Chaudhari R, Nasra S, Meghani N and Kumar
A: MiR-206 conjugated gold nanoparticle based targeted therapy in
breast cancer cells. Sci Rep. 12:47132022. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Gao X, Hang Y, Boryczka J, Tan W, Zheng P,
Yang F, Engler-Chiurazzi E, Simpkins JW and Wu N: Comparative
visible-light and near-infrared fluorescence detection of microRNA
in blood plasma with paper lateral flow test strips. Biosens
Bioelectron. 289:1179032025. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Mansour RM, Shaker AAS, Abulsoud AI,
Mageed SSA, Ashraf A, Elsakka EGE, Dahab MI, Sadek MM, Awad FA,
Lutfy RH, et al: The role of MicroRNAs in neurodegeneration:
Insights from huntington's disease. Mol Neurobiol. 62:8502–8517.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Yang J, Zhang X, Chen X, Wang L and Yang
G: Exosome mediated delivery of miR-124 promotes neurogenesis after
ischemia. Mol Ther Nucleic Acids. 7:278–287. 2017. View Article : Google Scholar : PubMed/NCBI
|