Introduction
Gastric cancer (GC) is one of the deadliest cancers
worldwide. It is the fifth most common malignancy worldwide and
fourth leading cause of cancer-related deaths (1). The incidence rate of GC in men is
twice that in women and there has been a notable increase in the
number of cases among individuals under 50 years of age. Despite
advances in surgery, chemotherapy and targeted therapy, the overall
survival rate of patients with advanced GC remains <40%.
Therefore, it is imperative to thoroughly investigate its
pathogenesis in order to develop novel diagnostic and therapeutic
approaches (2).
Research suggests that aberrant epigenetic
regulation is closely related to GC and plays a significant role in
the course of this illness (3).
Post-translational modifications (PTMs) of histones, the core
epigenetic mechanism, affect cell function by dynamically
regulating DNA and gene transcriptional activity. PTMs are closely
related to the emergence of GC resistance and are involved in
pathological processes, such as malignant proliferation (4). Ubiquitination, acetylation and the
newly discovered lactylation are three histone changes that have
gained much interest because of their crucial function in GC
(5,6). An imbalance in ubiquitination can
impair the stability of critical tumor suppressor genes, which is
similar to a 'double-edged switch' in the course of tumorigenesis.
This switch has the ability to stimulate and suppress tumor
development (7). Histone
deacetylases (HDACs) are overexpressed in GC, resulting in
chromatin defects and promoting tumor growth. Hence, epigenetic
treatment of GC has increasingly focused on HDACs (8). Histone lactylation alterations
deserve special consideration in the study of GC. The emergence of
a unique connection between epigenetic regulation and metabolic
reprogramming is the cause of this phenomenon. This change has the
potential to directly affect histones, which in turn causes the
expression of cancer-causing genes. It has the potential to alter
the metabolic trait of high glycolysis into tumorigenic epigenetic
alterations. Furthermore, there is growing evidence that
lactylation plays a role in controlling the tumor immune
microenvironment (9).
The present review aimed to systematically examine
the role of lactylation in GC and clarify the molecular processes
underlying major histone modifications during the initiation,
development, migration and multidrug tolerance of GC. Additionally,
it examined the therapeutic promise of these alteration pathways in
order to improve our knowledge of the epigenetic disruption
mechanisms in GC and offer a basis for creating new, targeted
treatment techniques.
Broader epigenetic landscape in GC
A number of studies demonstrate that there is a
communication system between them known as 'cross-talk' (10-12). DNA methyltransferases (DNMTs,
including DNMT1, DNMT3A and DNMT3B) are the enzymes that catalyze
DNA methylation, which involves the selective addition of methyl
groups (-CH3) to specific bases in DNA molecules. In GC,
it can result in the transcriptional silencing of important tumor
suppressor genes, such as CDH1, MLH1, CDH11 and IGFBP7. This is
frequently the case in highly methylated local areas of the genome.
Low-methylation areas often lead to genomic instability, whereas
the clustering of histone modifications or methylation-binding
proteins further strengthens gene silencing. The position,
composition and stability of nucleosomes are dynamically regulated
by chromatin remodeling complexes, including the switching
defective/sucrose nonfermenting (SWI/SNF) family, the imitation SWI
(ISWI) family, the nucleosome remodeling and
deacetylation/Menin-MLL Inhibitor (Mi-2)/chromodomain helicase
DNA-binding (CHD) family and the inositol requiring 80 family,
which, via ATP hydrolysis, allows transcription factors to approach
DNA and regulate transcription, replication, repair, development
and genome stability. The SWI/SNF complex consists of components
such as AT-rich interaction domain 1A (ARID1A) (13) and the chromatin remodeling factors
CHD4 (14) and CHD5 (15), all of which have the potential to
cause GC metastasis. Based on their length, non-coding RNAs
(ncRNAs) are primarily classified as circular RNAs (circRNAs),
PIWI-interacting RNAs (piRNAs), long non-coding RNAs (lncRNAs) and
small non-coding RNAs (sncRNAs). Although they do not encode
proteins, they are involved in GC angiogenesis, metabolism, immune
escape and chemotherapy resistance. CircPOFUT1 (16), circVAPA (17) and other similar compounds increase
cisplatin resistance in GC and facilitate cell proliferation,
migration and invasion. hsa_circ_0073453 (18) and circ_0136666 (19) influence CD8+ cells, eventually
resulting in immune evasion (20).
Histone modification
Direct protein modification regulates
transcriptional activity by altering the histone structure or
charge, which in turn affects chromatin compaction and gene
accessibility. It regulates gene 'activation' or 'inactivation'
without affecting DNA sequences, making it a fundamental component
of epigenetic regulation (21,22).
Histones are fundamental proteins that reside in the
nucleus of eukaryotic cells. They make up the core elements of
nucleosome subunits, which are made up of an octamer of four core
histones (H3, H4, H2A and H2B) surrounded by a 147-base-pair DNA
segment (23). Each histone has a
unique side chain or tail, which is highly enriched in lysine and
arginine residues (24). These
changes alter the charge density between histones and DNA, allowing
the precise regulation of chromatin transcription (Fig. 1). They are one of the core
mechanisms behind the epigenetic changes that regulate gene
activation or suppression without altering the underlying DNA
sequence. Histone modifications take numerous forms, including
methylation, acetylation, ubiquitination and lactylation (25).
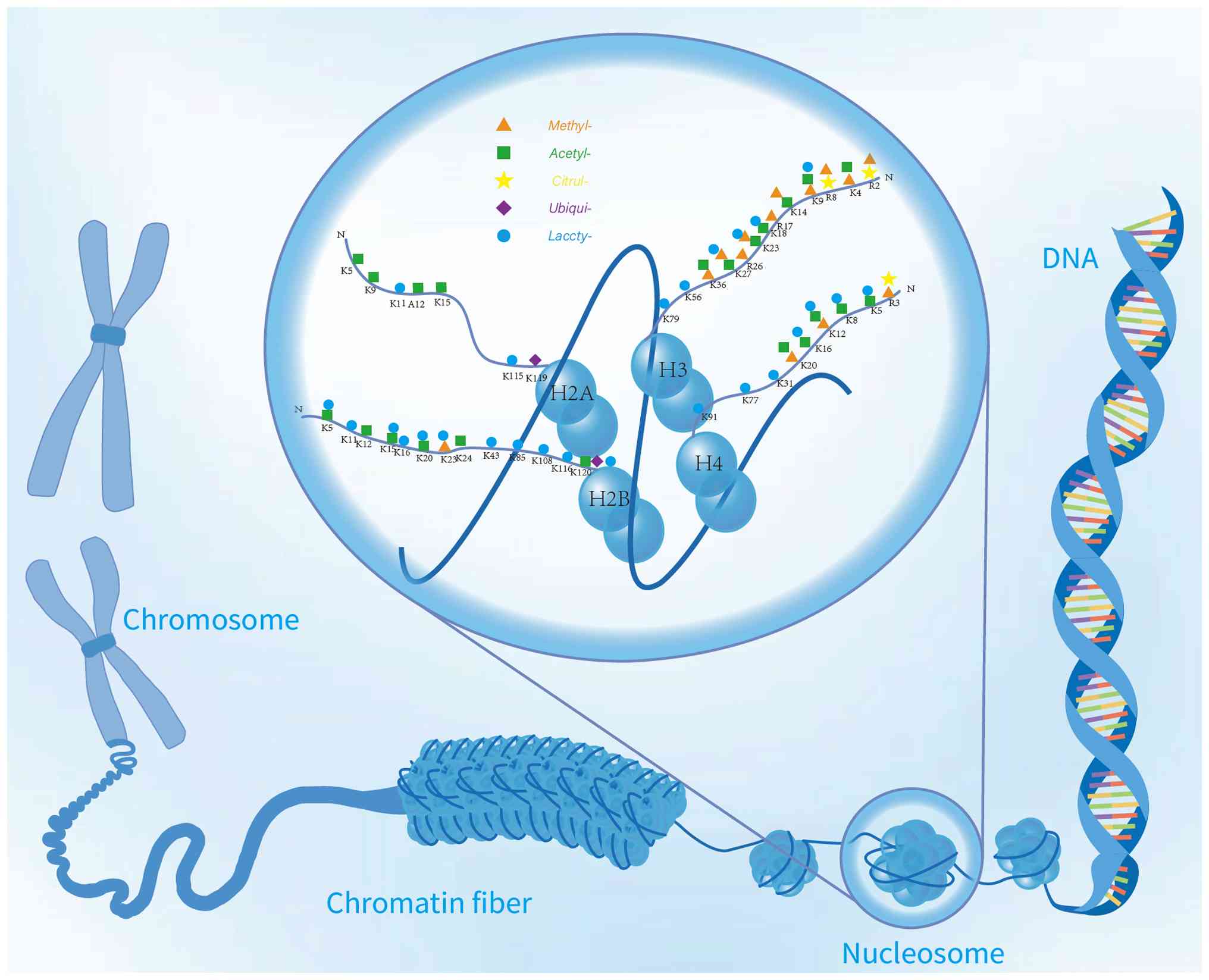 | Figure 1Distribution of multi-type
modifications of histone in different residual base sites and
structure in chromatin. The various types of histone modification
and its distribution in the chromatin structure from the advanced
structure of chromosomes to nucleosome, chromatin fiber and DNA
double helix structure. The figure marks the modification types of
some different histones, including methylation, acetylation,
citrullination, ubiquitination and lactylation and the specific
distribution of these modifications in different amino acid residue
bases of histone H2A, H2B, H3 and H4 (such as K14, K27, K36, etc.),
which reflects the effect of histone modification on the structure
and function of chromatin in epigenetic regulation. |
Acetylation modification
Histone acetyltransferase (HAT) mediates the
chemical process of acetylation. In this procedure, acetyl groups
are transferred from an acetyl coenzyme A (acetyl-CoA) molecule to
another molecule. This procedure activates the protein function by
attaching an acetyl group supplied by acetyl-CoA to the lysine
residues of histones. Thus, acetyl-CoA, a crucial intermediate, has
a significant influence on acetylation activity. The main HAT
families include the MYST proteins, p300/CBP and the Gcn5-related
N-acetyltransferase (GNAT) superfamily (26). The GNAT superfamily includes the
members Gcn5, PCAF, Elp3, Hpa2 and HAT1 (27), which can interact with particular
substrates and play roles in chromatin remodeling and gene
transcription. In addition to their HAT activity, p300/CBP proteins
act as transcriptional coactivators and interact with other
transcription factors to promote gene transcription. Moz,
Ybf2/Sas3, Sas2 and Tip60 are examples of MYST proteins (28). Additionally, acetylated lysine
residues in proteins have been identified by dual PHD finger (DPF)
domain proteins (e.g., MOZ and DPF2), YEATS domain proteins (e.g.,
MLLT3 and Taf14) and bromodomain proteins (e.g., BRD4 and BRDT)
(29). They function as lysine
acetylation readers, receiving signals from acetylated lysine
residues and binding to specific acetylating complexes. BRD4, which
has been found to be a histone acetylation reader and an upcoming
anti-cancer treatment target, is one of these that encourages GC
progression and metastasis (30).
The chemical process of removing acetyl groups from lysine
residues, which is opposite to acetylation, is catalyzed by HDACs.
HDACs are primary deacetylases that remove acetyl groups (31). The main categories of HDACs are
NAD+- and Zn2+-dependent HDACs (32). By eliminating acetyl groups, the
positive charge of histones is restored, which increases the
affinity between histones and the negatively charged DNA backbone,
resulting in chromatin compaction. As a result, the mixing of
transmission factors and RNA polymerase with DNA is prevented,
inhibiting gene expression (33).
The dysregulation of HDAC activity is closely associated with
cancer, fibrotic illnesses, cardiovascular and renal diseases,
neurodegenerative diseases and mental disorders.
Methylation modification
Histone methylation, which was first discovered in
the 1960s, is a reversible process in all living organisms. It
involves alkylation reactions in which methyl groups replace
hydrogen atoms (34). Histone
methyltransferases (HMTs) and histone demethylases (HDMs) work
together to maintain histone methylation balance. Using the
cofactor S-5'-adenosyl-L-methionine as a methyl donor, histone
methylation is catalyzed by HMTs. Methylation of a lysine or
arginine residue at a particular site on the tail of histones
(primarily H3 and H4) is a part of this process (35). Arginine methylation can occur as
symmetric mono-methylated (ME), symmetric di-methylated (ME2S), or
asymmetric di-methylated (ME2A), whereas histone lysine residues
can be methylated at the ε-amino group as mono-methylated (Me),
di-methylated (Me2), or tri-methylated (Me3) (36). The effects of methylation are what
make its function so complicated. These results demonstrate
site-specific and degree-specific methylation. Some of the histone
methylation sites are H3K4, H3K9, H3K27, H3K36, H3K79 and H4K20.
The arginine methylation sites include H3R2, H3R8, H3R17, H3R26 and
H4R3. Methylation is controlled by HMTs. There are three families
of enzymes: N-methyltransferases, lysine-acting Dot1-like proteins
and SE domain-containing enzymes. These enzymes act on both free
and non-histone proteins. Thus, the functions of these proteins are
triggered (37). These enzymes
are involved in various biological processes, including
transcription factor recruitment, binding and chromosome packing.
They also control RNA processing/editing, DNA binding and
initiation/elongation and are involved in several other connected
procedures (38). Additionally,
HMTs govern genomic alterations that contribute to cancer
development. Indeed, previous studies have shown that they are
essential for the development and formation of tumors, while also
aiding in preserving stem cell populations in tumors. HMTs are
subject to rigorous regulations governing essential cellular
activities under typical conditions (39).
HDMs are involved in the process of using specific
enzymatic catalysts to eliminate methyl groups from lysine or
arginine residues on histone tails (37). The two main enzyme categories in
this catalytic process include flavin-dependent amine oxidases,
which are the enzyme family to which the first class of HDMs (such
as LSD1) belongs and the JmjC domain family, which are the enzyme
family to which the second class of HDMs belongs and which
catalyzes ferrous ion oxidation and employs ketoglutarate as a
cofactor to demethylate histone lysine residues. Other proteins
involved in histone demethylation were also found in the histone
lysine demethylase (KDMs) cohort (40) within the KDM1 to KDM9 subfamilies
(41,42). HDMs have a dynamic effect on
chromatin regions rather than altering the DNA sequences. In
addition to preserving genome integrity and stability, they play
essential regulatory roles in embryonic development, genetic
control, cellular reprogramming and other biological processes.
They play an important role in cancer because they are closely
linked to disease pathogenesis; this includes boosting mutation
rates, changing the copy number of gene segments, enriching
transcription factors and remethylating genes that control cell
fate. Targeting particular HDMs has opened up an intriguing new
area in cancer treatment (Fig. 2)
(39).
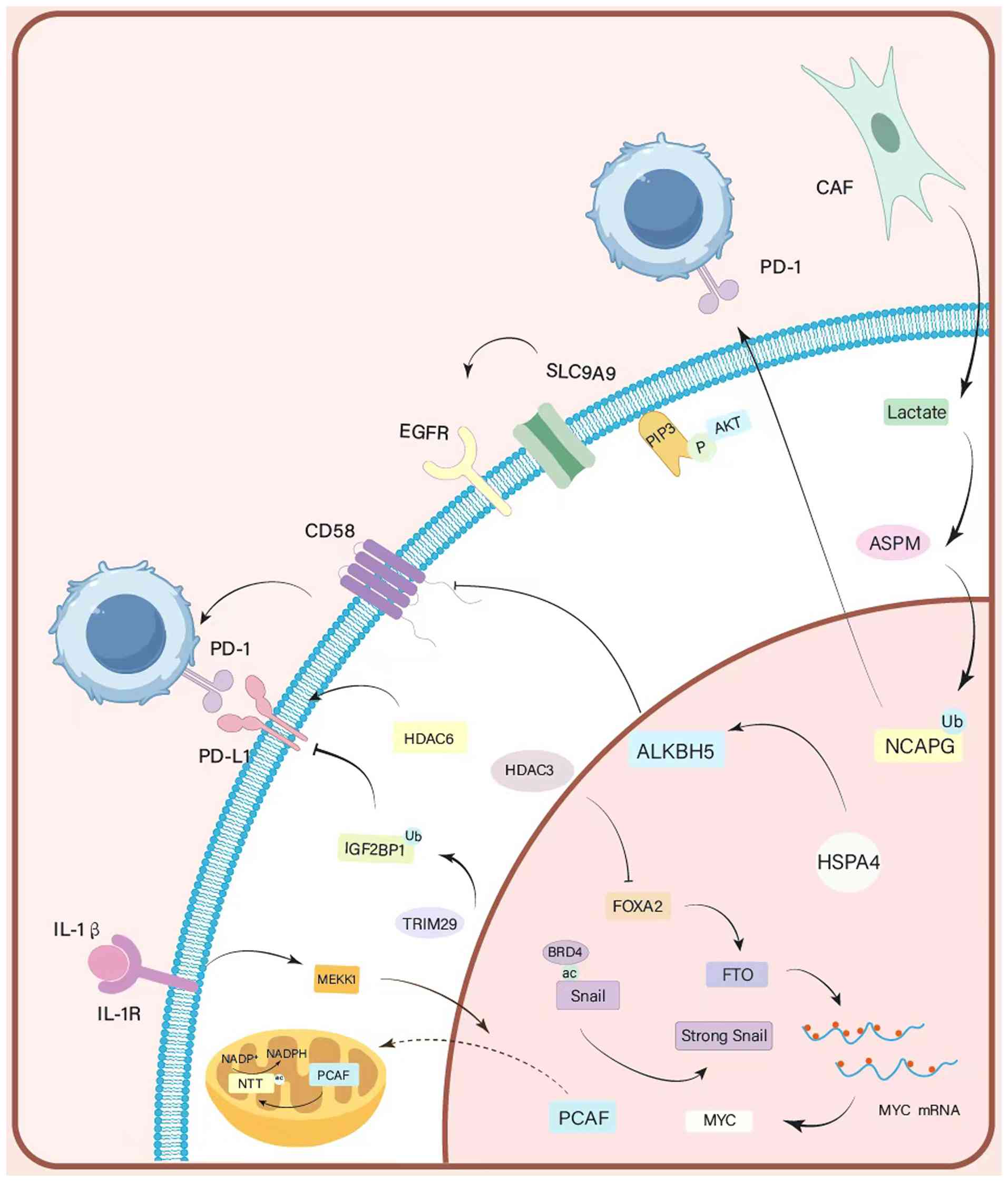 | Figure 2Mechanisms of histone acetylation and
ubiquitination. These two major histone modifications regulate the
expression of key effector molecules (such as MYC, Snail, NCAPG)
and activate downstream signaling pathways (such as EGFR/AKT,
IL-1β/MEKK1), thereby directly promoting malignant proliferation
and survival of GC cells. MYC, myelocytomatosis oncogene; NCAPG,
non-SMC condensin I complex subunit G; EGFR, epidermal growth
factor receptor; AKT, protein kinase B; IL-1β, interleukin-1 beta;
MEKK1, mitogen-activated protein kinase kinase kinase 1; GC,
gastric cancer. |
Lactylation modification
In the 1920s, Otto Warburg discovered that tumor
tissue uses considerably more glucose than healthy tissue. The
Warburg effect is a counterintuitive observation in that tumor
cells continue to produce lactate from glucose, even in an
oxygen-rich environment (13).
Lactate was previously thought to be both an energy source and a
metabolic byproduct of the Warburg effect (43,44). The important functions of lactate
in the body have been discovered in an increasing number of studies
over the past few years. It is now appreciated beyond a superficial
metabolic byproduct role. Professor Yingming Zhao at the University
of Chicago initially demonstrated the existence of histone lysine
lactylation (Kla) in 2019 and proved that it was a novel epigenetic
change (45). Through specific
receptors, lactate can activate downstream signaling pathways and
serve as a substrate to support Kla activity. Understanding how Kla
functions may offer insights into tumor treatment because it has a
unique impact on tumor growth (46). Kla is a covalent modification in
which the lactate moiety (-CO-CH3OH) is linked to lysine
residues on proteins via an amide bond. It consists of three
stereochemically distinct isomers that are structurally similar:
L-lactylation (KL-La), D-lactylation (KD-La) and
N-ε-(carboxyethyl)-lysine (Kce). At present, 9,275 Kla locations
have been identified, of which 9,256 are non-histones (47). Most Kla is present on the lysines
of H2A, H2B, H3 and H4, especially H3K18 (48). Using liquid chromatography-mass
spectrometry to analyze animal cells, Varner et al (49) found that the levels of lactoyl-CoA
were 20-350 times lower than those of other key lactoyl-CoA
compounds. Kla's dynamic equilibrium uses lactoyl-CoA as its
substrate and is jointly controlled by lactyltransferase and
delactylase enzymes, resulting in covalent bonding under certain
enzymatic or non-enzymatic conditions (50). The source of lactoyl-CoA
production may be glycolysis, lactate buildup, or metabolic
bypasses that use coenzyme A. P300/CBP was the first possible
lactyltransferase discovered (45,51); however, indirect participation
cannot be excluded. Lactyltransferases, including alanine tRNA
synthetase (AARS1) (52), KAT2A,
HBO1 (53), KAT8 (54), GNAT13, YiaC and CobB (55,56), have been reported to increase in
recent years. Known deacetylases include Class I histone
deacetylases (HDAC1-3) (56) and
SIRT1-3 (57). Research suggests
that repair genes, pluripotency genes, transcription factors
(58), pro-inflammatory factors
(59), m6A-associated regulatory
proteins (60), fibrosis-related
genes and essential signaling pathway proteins are the downstream
target genes of H3K18la-mediated transcriptional activation
(61). Additionally, H3K18la and
H4K5la may function together to regulate PD-L1 expression (Fig. 3) (62).
Other modifications
Histone modifications vary and include histone
ubiquitination and citrullination, as well as the previously
mentioned changes. Citrullination of histones is a unique
modification procedure, which converts positively charged or
methylated arginine residues into neutral citrulline residues. The
interaction between proteins as well as between proteins and
nucleic acids is impacted by this shift (63). Enzymes, such as peptide arginine
deaminase (PAD), which transforms arginine residues into citrulline
residues, play a major role in this process. When arginine is
highly alkaline, positively charged side chains are hydrolyzed and
neutral urea is produced (64).
This alteration affects the fundamental processes of cells in a
significant way, such as via apoptosis, differentiation and immune
response. The fact that this change can directly stimulate the
growth and spread of cancer is even more significant (65,66). PAD can worsen rheumatoid arthritis
in the pathological state by causing autoantibodies and setting off
the body's inflammatory reaction (67,68). Studies have demonstrated that PAD
can be used as both a therapeutic target and a diagnostic
indicator. Non-small cell lung cancer, GC, hepatitis B
virus-related hepatocellular carcinoma and various malignant blood
tumors are all associated diseases (69). PAD inhibitors can also be used in
cancer therapy; PAD4 inhibitors are particularly noteworthy because
they can stop tumor metastasis in patients with cancer and prevent
accompanying thrombosis (70,71). The ATP-dependent process of
ubiquitination links ubiquitin molecules to substrate proteins,
with cell survival, proliferation and differentiation all being
affected by this process (72).
Ubiquitination can be categorized into four types: Mono-, poly-,
homogeneous and heterogeneous polyubiquitination. E1-Ub, E2-Ub and
E3-Ub ligases are three enzymes that mediate ubiquitination, which
is a histone modification (73).
Ubiquitinated proteins can be removed by deubiquitination enzymes
(DUBs), which reverse the ubiquitination process. Early research
has demonstrated that several DUBs, including USP4 (UNP), USP6
(Tre-2) and USP8 (UBPY), are essential for cancer development.
programmed death-1/programmed cell death protein 1(PD-1/PD-L1)
inhibitors can be used in conjunction with USP8/USP7 inhibitors to
greatly improve antitumor effectiveness (74). USP9X deubiquitinase is capable of
stabilizing oncogenic genes such as β-catenin and MCL1, both of
which are involved in tumor development and spread. In addition,
USP9X governs epithelial-mesenchymal transition, which raises the
chance of tumor metastasis. USP9X also eliminates ubiquitin from
PD-L1 cells. Due to this activity, cancer cells can avoid the
immune system. MCL1 is destroyed by USP9X inhibitors and cancer
cell apoptosis is initiated by this breakdown, suggesting that
USP9X may be a therapeutic target. Indeed, USP9X has attracted
considerable interest for its potential therapeutic use in
neurodegenerative disorders (75). The complex interplay between
ubiquitination and deubiquitination is closely associated with
several cellular processes. Disruption of this mechanism can result
in neurodegenerative illnesses, cancer and other illnesses. New
approaches for treating these disorders may arise from this
association (Table I) (76).
 | Table ILandscape of major histone
modifications and their regulators in GC. |
Table I
Landscape of major histone
modifications and their regulators in GC.
| Modification | Enzymatic
regulators | Relevance in
GC | Key associated
molecules/targets in GC |
|---|
| Acetylation | Writers:
HATs
• p300/CBP
• GNAT family
• MYST family
Erasers: HDACs
• Class I/II (Zn2+-dependent)
• Class III (NAD+-dependent, SIRTs) | HDAC overexpression
correlates with poor prognosis; HDACs inhibitors are promising
therapeutic targets. HDAC6 upregulates PD-1/PD-L1. BRD4 promotes GC
progression and metastasis. | HDAC1, HDAC3,
HDAC6; BRD4; HSPA4; SLC9A9/pomiferin; NNT K1042ac. |
| Methylation | Writers:
HMTs
• EZH2
•SUV39H1/2
• SETD7
• PRMTs
Erasers: KMTs/HDMs
• LSD1 (KDM1A)
• KDM5A/B/C
• KDM6B | EZH2 overexpression
correlates with progression and poor prognosis. SUV39H1/2
participate in GC initiationand migration. LSD1 and KDM5A/B/C are
overexpressed and associated with proliferation and
metastasis. | EZH2; SUV39H1,
SUV39H2; LSD1 (KDM1A); KDM5A, KDM5B, KDM5C; KDM6B; SETD7/YAP1 |
| Lactylation | Writers:
Lactyltransferases
• p300/CBP
• AARS1
• KAT2A, HBO1
Erasers: Delactylases
• SIRT1-3
• HDAC1-3 | H3K18la level is
markedly increased in GC and closely related to poor prognosis. Kla
drives metabolic reprogramming, proliferation, migration, immune
escape and therapy resistance. | H3K18la; AARS1;
LDHA; GLUT3; PFKM; HKDC1; NLRP12; PD-L1. |
| Citrullination | Writers: PADs,
especially PAD4
Erasers: (Not well characterized) | PAD4 has genetic
susceptibility to GC and its overexpression promotes tumor
angiogenesis, proliferation and migration. PAD4 regulates H3R26
citrullination in GC cell lines. | PAD4; H3R26; CXCR2,
KRT14, TNF-α. |
| Ubiquitination | Writers: E3
ubiquitin ligases (e.g., RNF112, TRIM29)
Erasers: DUBs (e.g., USP9X, OTUD5) | Ubiquitination
regulates glycolytic reprogramming and promotes GC metastasis.
TRIM29/IGF2BP1/PD-L1 axis is a potential therapeutic target. |
TRIM29/IGF2BP1/PD-L1; |
Application of histone modifications in
gastric cancer
Acetylation modifications and gastric
cancer
Several targets for histone acetylation have been
identified during GC treatment. Based on acetylation patterns, four
lncRNAs (AC114730.1, AL445250.1, LINC01778 and AL163953.1) have
been identified as potential chemotherapeutic drugs for adjuvant
treatment in various risk categories and to create a prognostic
model for the GC response to immunotherapy (77). The responsiveness of patients with
GC to immune checkpoint inhibitor treatment can be predicted by
histone acetylation-induced upregulation of HSPA4 in GC tumor
tissues (78). In cancer cells,
the acetylation of nicotinamide nucleotide transferase (NNT) at
lysine (K) 1042 (NNT K1042ac) under interleukin-1β (IL-1β)
stimulation markedly reduces tumor immune escape brought about by
IL-1β. By blocking NNT acetylation, therapeutic benefits are
attained by interfering with the IL-1β-tumor cell axis (79). By blocking EGFR/PI3K/AKT signaling
and preventing GC development, the natural flavonoid pomiferin
binds to SLC9A9, a marker of aberrant histone acetylation, in GC
cells (2). Owing to their role as
epigenetic regulators of chromatin condensation and decondensation
(80), HDACs are potential
therapeutic candidates for cancer treatment. The hydroxyacetamide
derivatives vorinostat and panobinostat are examples of HDAC
inhibitors licensed for the treatment of hematologic malignancies
(81) and their use in solid
tumors is currently under research (82). By contrast, excessive expression
of HDAC1 and HDAC3 (83) in GC is
associated with poor prognosis, whereas HDAC6 contributes to an
increase in important elements of cancer immunotherapy targets,
such as PD-1 and PD-L1 receptors (84). Additionally, HDAC3, HDAC4, HDAC5,
HDAC6, HDAC7, SIRT3, SIRT5, SIRT6 and SIRT7 seem to be possible
prognostic indicators, while HDAC1, HDAC2 (85) and HDAC4 may also be used as
predictive biomarkers (86).
These results provide a plethora of possible targets and biomarkers
for targeted therapy, prognostic evaluation and treatment
optimization in GC.
Methylation modifications and gastric
cancer
Several malignancies and developmental problems are
linked to dysregulation of histone lysine methylation. Thus,
histone lysine methylation is a potential therapeutic target
(87). Since cancer frequently
exhibits aberrant histone methylation caused by gene mutations,
translocations, or dysregulation, a number of HMTs and HDMs are
possible therapeutic targets (88). The histone methyltransferase EZH2
has been thoroughly investigated in the context of GC pathogenesis
(89). Its overexpression in GC
is linked to progression, malignancy and poor prognosis and
promotes H3K27 trimethylation. Other HMTs involved in GC include
SUV39H1 and SUV39H2, which promote H3K9 methylation and play roles
in the initiation, progression and spread of GC (90,91). Methylation of the ribosomal
protein L40 (rpL40) has been linked to the ribosomal export process
in GC by the lysyl methyltransferase SMYD5. One possible component
of combination treatment for this cancer is targeting SMYD5
(92).
Although some HDMs exhibit tumor-suppressive
properties, their potential functions in cancer are mainly
oncogenic. By demethylating H3K4, which silences tumor suppressor
genes and promotes tumorigenesis, LSD1, a member of the LSD family
of HDMs, functions as a transcriptional co-repressor (93,94). Additionally, numerous KDMs are
overexpressed in GC tissues. These KDMs include KDM1A, KDM5A, KDM5B
and KDM5C, which are involved in cell proliferation and metastasis
(95,96) and are members of the H3K4
demethylase family; KDM2A and KDM2B, which drive cell growth and
migration and KDM4B and KDM4C, which are members of the H3K9
demethylase family and are involved in tumor growth and invasion
(97). Additionally, GC
development is linked to excessive expression of the H3K27
demethylase KDM6B. There have been numerous reports of aberrant
expression of enzymes controlling histone methylation (91,98). Therefore, there are possible
treatment targets for the dysregulation of histone lysine
methylation, which is closely associated with developmental
problems and cancer. Genetic mutations or other mechanisms
frequently cause such dysregulation in GC, with aberrant expression
or function of HMTs (e.g., EZH2 and SUV39H2) and HDMs/KDMs (e.g.,
LSD1 and KDM1A) being pertinent to the initiation, progression and
prognosis of GC, respectively, thereby providing possible
therapeutic targets.
Other modifications and gastric
cancer
Histone citrullination is an increasingly important
therapeutic target, which aids in addressing GC (99). PAD4 is an essential enzyme that
controls histone citrullination and is crucial in initiating and
promoting GC. The PAD4 monomer is composed of two domains, one of
which is the N-terminal domain (residues Met1-Pro300) and the other
is the C-terminal catalytic domain, which consists of residues
Asn301-Pro663. Two immunoglobulin-like subdomains make up the
N-terminal domain. Residues Met1-Cys118 are included in subdomain
I, while the residues between Ala119 and Pro300 are included in
Subdomain II (100). The PAD4
gene is genetically predisposed to GC. GC is strongly associated
with tag SNPs at the PAD4 locus. Overexpression of PAD4 converts
arginine residues to citrulline residues. By increasing the
expression of TNF-α, KRT14 and CXCR2, this effect might encourage
tumor angiogenesis, cell proliferation and cell movement.
Collectively, these modifications create an immune microenvironment
that supports GC development (101). PAD4 is essential for GC cell
lines and may increase the citrulline modification of H3R26,
supporting the hypothesis that PAD4 contributes to GC development
by regulating histone citrullination (102).
In conclusion, the histone citrullination alteration
mediated by PAD4 is a potential therapeutic target in GC. By
suppressing the activity or expression of PAD4, it is possible to
lower the level of histone citrullination and consequently, its
regulatory influence on downstream cancer-causing genes. This may
open up a novel avenue for GC therapy based on the precise control
of histone citrullination and may prevent the spread, movement and
angiogenesis of GC cells, ultimately improving patient prognosis.
Protein ubiquitination plays a critical role in GC progression by
affecting the reprogramming of glycolytic metabolism and
advancement of GC metastasis (103). Tripartite motif-containing
protein 29 (TRIM29) expression is lower in GC tumor tissues than in
normal tissues. The TRIM29/IGF2BP1/PD-L1 axis includes TRIM29 and
represents a potential therapeutic target (104). By contrast, the promotion of
ubiquitination prevents GC growth and enhances the effectiveness of
anticancer medications. Tumor tissues produce more lactate in
cancer-associated fibroblasts (CAFs). In GC cells, this lactate
encourages the lactylation of H3K18. Deubiquitination of the
non-SMC condensin I complex subunit G (NCAPG) is indirectly
enhanced by the activation of spindle microtubule assembly factor
(ASPM). PD-L1 expression is enhanced by NCAPG. The effectiveness of
anti-PD-1 treatment for GC may be enhanced by daturilin, a
low-molecular-weight NCAPG antagonist (105).
The intricate regulatory system of histone
ubiquitination has great potential for reversing the aberrant
proliferation and metastatic behavior of GC cells. This strategy
creates new opportunities for precisely targeted therapies based on
the control of histone ubiquitination and aids in the creation of
more successful GC treatment approaches.
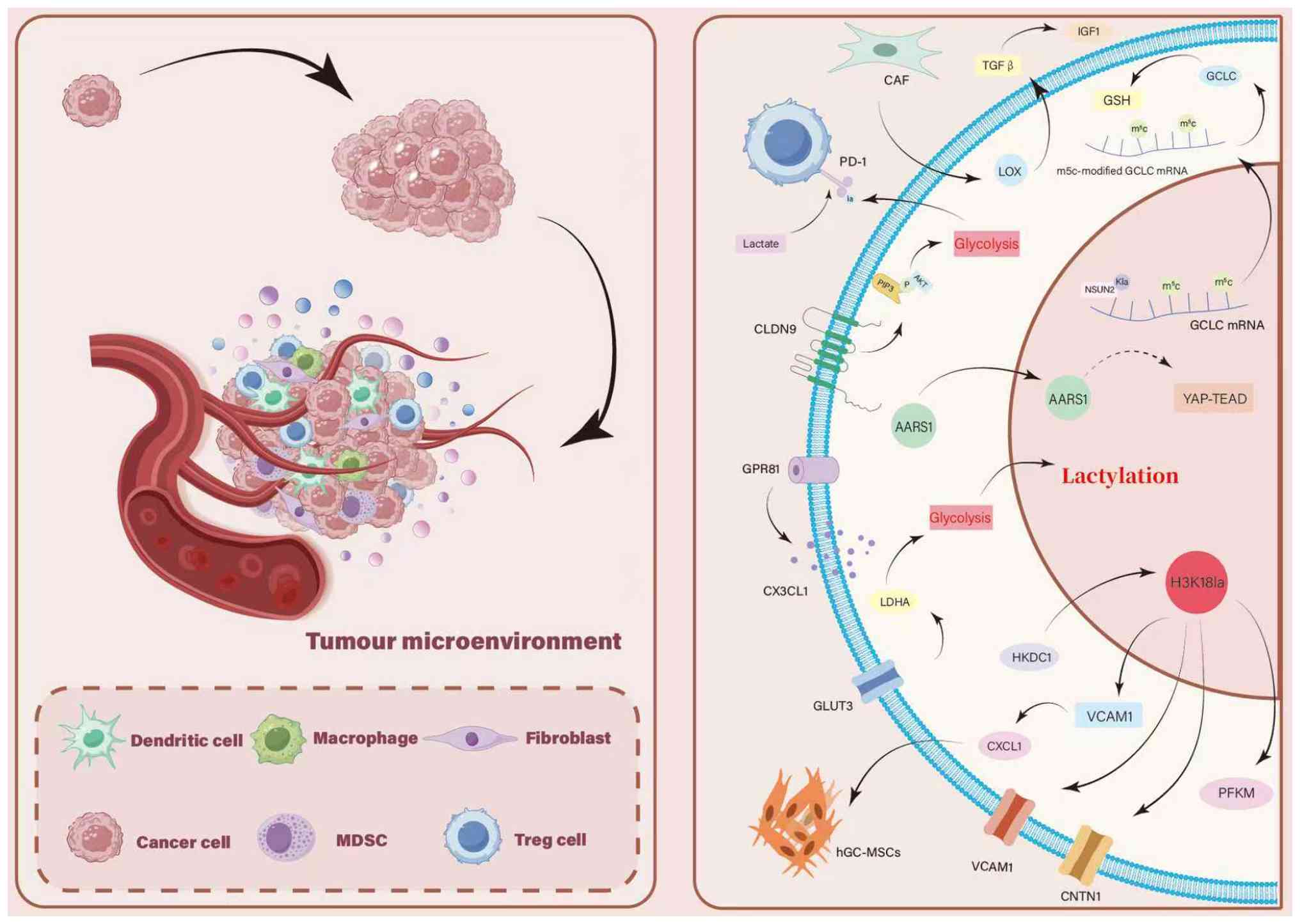
Function of histone lactylation in gastric
cancer
Kla uses a variety of processes to markedly affect
the development and course of GC, which has a profound effect on
the course of the illness (106). Kla controls metabolic
reprogramming in GC, markedly alters a wide range of proteins in GC
cells, affects major metabolic pathways, such as fatty acid
metabolism and glycolysis, and simultaneously promotes metabolic
abnormalities and malignant progression. Its mode of action
involves the aberrant expression of molecules such as AARS1 and
NLRP12, as well as lactation-induced changes to histones such as
H3K18la. Conversely, Kla modulates histone modification and
controls the activity of molecules such Glucose transporter 3
(GLUT3), Phosphofructokinase muscle isoform (PFKM) and HKDC1. By
activating the appropriate signaling pathways, Kla promotes the
growth, migration and invasiveness of GC cells. Furthermore, GC
cells can evade immune system surveillance by influencing PD-L1
expression and regulating the function of tumor microenvironmental
components, including cancer-related fibroblasts, macrophages and
regulatory T cells. As a result, Kla is a major catalyst for the
advancement of GC and management of drug resistance. The method by
which these factors are lactylated will then be examined in
depth.
Lactylation and metabolic
reprogramming
Kla is essential for GC development because it
controls metabolic reprogramming. According to previous research,
the protein Kla varies greatly among the liver cancer, pancreatic
cancer, colorectal cancer and GC. A variety of functional routes
are enriched in proteins with specific Kla sites, particularly
those involved in metabolic processes, such as fatty acid
metabolism, glycolysis, glucuronidation and amino acid metabolism
(107). Thus, Kla might affect
the course of GC by changing the fundamental metabolic pathways
(108). A total of 2,375 Kla
sites were identified among 1,014 GC cell proteins (109). The fact that the amount of
H3K18la is much higher in GC and is strongly associated with poor
prognosis is noteworthy, reinforcing the importance of Kla in GC
metabolic reprogramming and disease progression, particularly
through regulation of metabolic gene expression mediated by
histones (57,108). Lactate dehydrogenase A (LDHA)
and AARS1 are upregulated in GC and their increased expression is
associated with negative patient outcomes. Abnormal expression of
AARS1, a crucial enzyme in lactate metabolism, may indirectly
control metabolic reprogramming in GC by upsetting the dynamic
equilibrium of Kla, making it a potential marker of metabolic
abnormalities and malignancy in GC (52). By contributing to the metabolic
reprogramming of GC cells, improving the expression of H3K18la and
stabilizing the essential glycolytic enzyme hexokinase 2 (HK2), the
NLR family pyrin domain, which includes 12 members (NLRP12),
primarily advances the progression of GC (110). Kla, similar to H3K18la, plays a
significant role in the metabolic reprogramming of GC by regulating
the expression of essential metabolic pathways and associated
proteins (e.g., AARS1, NLRP12 and HK2) in GC cells. The
significance of Kla in the pathogenesis of GC and its enormous
potential in scientific studies and clinical practice have been
emphasized.
Promotion of gastric cancer proliferation
and migration
Lactate is an important metabolic intermediate,
which serves as a mediator for Kla and promotes the development and
progression of GC. In this process, several molecules either
control Kla or are directly involved in the pathways linked to
lactylation. Previous studies have demonstrated that lactate
directly promotes tumor cell progression in GC cells (111-113). This mechanism corresponds to the
function of lactate as a metabolic intermediate as it facilitates
histone modification and promotes the transcription of Vascular
Cell Adhesion Molecule 1 (VCAM1) by inducing H3K18la. The AKT-mTOR
signaling pathway increases C-X-C Motif Chemokine Ligand 1 (CXCL1)
expression via VCAM1. This encourages the recruitment of human
GC-derived mesenchymal stem cells, increases immunosuppression and
hastens the course of cancer (114). Primary and metastatic GC tumors
both exhibit high levels of GLUT3 expression. Its expression is
positively associated with lactate-associated pathways and LDHA. By
controlling H3 Kla, which facilitates cell proliferation,
metastasis and invasion, GLUT3 modulates GC cell activity (115). PFKM is one of the primary
rate-limiting enzymes in glycolysis. Its expression is controlled
by lactate and is overexpressed in GC. By increasing Contactin-1
(CNTN1) via H3K18la, PFKM promotes GC progression; by decreasing
PFKM, GC cell proliferation and migration are inhibited (116,117). Intracellular lactate levels are
detected by AARS1, which then translocates to the nucleus. Lactate
activates the YAP-TEAD transcription factor complex, which
initiates a positive feedback loop that encourages the growth of GC
cells. Poor patient prognosis is associated with high AARS1
expression in GC (52,118). In GC, hexokinase
domain-containing 1 (HKDC1) protein stimulates the expression of
H3K18la. Controlling VCAM1 expression accelerates tumor development
(119). Kla changes, such as
H3K18la changes, are mediated by lactate via its own activity and
by important molecules such as GLUT3, PFKM, AARS1 and HKDC1. This
either controls signaling pathways (such as AKT-mTOR and YAP-TEAD)
or activates the expression of target genes (such as VCAM1 and
CNTN1). Ultimately, it promotes GC development through a variety of
methods, including boosting tumor cell activity, attracting
immunosuppressive cells and triggering pro-oncogenic signals.
Lactylation and immune
microenvironment
Kla is a vital mechanism that facilitates the
progression of GC, influencing tumor-immune interactions in a
variety of ways during its development (120). According to previous research,
there is a clear link between lactate and tumor immune evasion; the
amount of lactate released by tumor tissues is markedly higher than
that released by healthy tissues. Increased lactate concentrations
in GC tissues lead to much higher PD-L1 expression than that in the
normal control group, demonstrating that lactate aids cancer cells
in evading the immune system (121). Lactic-acid-mediated immune
escape also involves cellular elements in the tumor
microenvironment (TME) (122).
Expression of the closely linked protein-9 (CLDN9) gene and
interleukin IL-4 both cause CD8+ T cell dysfunction in GC, which
contributes to immune escape. Through the PI3K/AKT/HIF1α signaling
pathway, CLDN9 promotes the glycolysis pathway while also fostering
the production and stability of PD-L1 lactate (123). By boosting ATP synthesis and
increasing sugar fermentation, inducing lactate production,
upregulating the expression of FcγRIIB on the surface of
macrophages and inducing PD-1 antibody to treat drug resistance,
IL-4 stimulates macrophage metabolic changes, activates the
PI3K/AKT/mTOR pathway and facilitates immune escape (124,125). By lowering the inhibitory effect
on CD8+ T cell activity, deletion of G protein-coupled receptor 81
(GPR81) reduces the entry of regulatory T cells into the TME and
slows the development of GC (113). By controlling the release of
lysyl oxidase (LOX) and CLDN9 by CAFs, activating the
PI3K/AKT/HIF1α pathway and inducing macrophage metabolic changes
caused by IL-4, Kla directly encourages the expression of PD-L1 in
tumor cells and mediates immunity throughout the development of GC.
Kla also promotes medication resistance in GC therapy by
influencing Treg cell infiltration through GPR81. In conclusion,
lactylation is an essential primary immune-related process in GC
development.
Lactylation and therapy resistance
Lactate plays an essential role in the mechanism of
tumor drug resistance by controlling the immune microenvironment,
DNA repair, gene expression and metabolic pathways (126) and contributes to drug resistance
in various tumor types (127).
In the context of immunological drug resistance, lactate induces
the expression of chemokine CX3CL1 by activating the GPR81
signaling pathway, which promotes the entry of regulatory T cells
into the TME. By reducing regulatory T cell infiltration and
mitigating its inhibitory effect on CD8+ T cell function, GPR81
deletion can prevent the central lymph node from progressing
(117). Kla is widely involved
in various drug resistance mechanisms, providing key theoretical
support and potential ways to develop new Kla targeting strategies
to overcome tumor drug resistance.
Clinical significance of histone lactylation
modifications
Biomarkers for gastric cancer diagnosis
and prognosis
The spread and invasion of GC cells is facilitated
by GLUT3. In GC cells, LDH activity, L-lactic acid concentration
and Kla levels can be markedly reduced by knocking out GLUT3,
indicating that it may be used as a possible diagnostic marker of
GC (128). The gene associated
with hypoxic-glycolysis-lactic acidation mirrors the intrinsic
metabolic condition, drug reactivity and immunological
microenvironment makeup of GC, which aids in improving prognostic
assessment (129). The SIRT
family is a subset of class III HDACs. The combination of oxamate
and a low dose of the SIRT1 activator SRT2104 may markedly limit
the proliferation of GC cells (57). The efficacy of these treatment
strategies, which are regulated by lactylation, supports the use of
lactylation-related chemicals as prognostic indicators.
Models that incorporate numerous genes related to
lactylation reflect disease features more accurately than models
that focus on a single molecule. The lactate production scoring
model is a promising diagnostic biomarker for GC because database
analysis has revealed four basic lactylation-related pathways in GC
tissues, which led to the selection of six lactylation-related
genes for model development (130). Based on the Cancer Genome Atlas
study, a risk model for lactate production-related genes
efficiently differentiates molecular and immunological features in
patients with GC, laying the groundwork for prognostic evaluation
(121).
Therapeutic targets
Extensive studies have shown that epigenetic drugs
can suppress oncogenes or activate tumor suppressor genes to treat
specific cancers. In research on lactate-related therapeutic
targets for GC, molecules involved in lactate metabolism and
transport are important research directions (131). As a key enzyme in lactate
production, LDHA has shown potential in preclinical studies on
targeted drugs (132). Zhao
et al (128) conducted
experiments using samples from patients who underwent gastrectomy
and found that oxamate, a widely used LDHA inhibitor, suppressed GC
cell proliferation when used alone. However, this has not been
clinically validated. Lactate transporters are closely associated
with LDHA. A previous study suggested that inhibiting
monocarboxylate transporters targeting glycolytic and oxidative
tumor cells disrupts the lactate shuttle, thereby overcoming immune
evasion, enhancing immunity and strengthening antitumor effects
(126). Among them, the MCT1
inhibitor AZD3956 has entered clinical trials and is expected to
further inhibit GC progression by blocking lactate transport.
However, it should be noted that studies have identified cases of
malignant hyperlactemic acidosis occurring in patients with
metastatic melanoma immediately after their first exposure to
AZD3965. Therefore, the efficacy and safety of AZD3956 in GC still
require confirmation in clinical trials (127). β-Aminopropionitrile, a LOX
inhibitor compound, has been shown to be effective in treating
breast cancer by inhibiting LOX. In GC, subsequent studies have
clarified that CAFs can drive tumor progression through the
LOX/transforming growth factor β(TGFβ)/IGF1 pathway. Earlier
experiments revealed that dextran sulfate can simultaneously reduce
the expression of LOX and TGF-β under hypoxic conditions, while
inhibiting the invasion and migration of GC cells. Although this
early study did not explicitly link these molecules to a single
pathway, its findings align closely with the subsequently
elucidated pathway inhibition effects. Therefore,
β-aminopropionitrile likely exerts its anti-GC activity by
inhibiting the same LOX/TGFβ/IGF1 pathway, though this remains to
be confirmed through targeted experimental validation (133-135). Monocarboxylate transporter 4 is
highly overexpressed in malignant GC cells with peritoneal
carcinomatosis and silencing this protein reduces tumor cell
proliferation and lactate uptake in malignant ascites, making it a
clinically specific target for GC (136). In addition, there are multiple
notable targets for lactylation-related epigenetic regulation and
cell death mechanisms. Tumor stemness refers to tumor cells with
stem cell behavior and characteristics that play key roles in
tumorigenesis (137). METTL14
expression is upregulated by histone H3Lys18 lactylation, which
inhibits GC stemness by mediating m6A modification of ATF5 mRNA;
knocking down METTL14 activates the WDR74/β-catenin axis to promote
stemness and reduced METTL14 expression levels in patients with GC
correlate with poor prognosis, making it a potential therapeutic
target (138). In GC therapeutic
research, these targets have shown potential for inhibiting disease
progression and improving prognosis, providing multidimensional
approaches for precision treatment.
Current limitations and unresolved
questions in histone lactylation research in GC
Currently, it is quite difficult to identify Kla,
especially at particular histone locations such as H3K18la, owing
to the lack of antibody specificity. According to recent research,
most PTMs have not yet been isolated. Instead, it has been
suggested that two or more separate PTMs could interact with each
other. Due to the potential for cross-reactivity with other
acylation changes, such as acetylation, the validity of existing
antibodies may be jeopardized (49). Second, mass spectrometry analysis
struggles with signal interference because lactyl-lysine and
N6-carboxyethyl lysine are isomers with the same molecular weight,
making it difficult to distinguish them. Additionally, lactoyl-CoA,
the major substrate, is found at very low concentrations in cells
and traditional mass spectrometry techniques are not sufficiently
sensitive to identify it. This constraint prevents precise
measurements and dynamic tracking investigations (50). Most importantly, most of the
possible targets mentioned in the present review, such as GLUT3,
LDHA and MCT1, are still in the early stages of research,
underscoring the universal difficulties in translating histone Kla
research from basic mechanisms to therapeutic uses. For example,
CLDN9 is essential for GC immune evasion. Systemic suppression of
this protein, which regulates several cellular processes, may have
significant systemic consequences. Therefore, it is essential to
develop therapies that target CLDN9, particularly for the treatment
of GC, without causing harm to the body. However, modern
technologies are still unable to satisfy this need.
Outlook
GC is one of the most prevalent types of cancers
worldwide. The pathogenesis is complex and exhibits significant
heterogeneity. Clinical diagnosis and treatment still face numerous
challenges, including difficulties in early detection, drug
resistance and poor prognosis. Overall, the pivotal role of
epigenetic regulation in tumor development has become increasingly
prominent.
Epigenetic regulation constitutes a multilevel
network for the occurrence and development of GC, in which classic
mechanisms such as DNA methylation, chromatin remodeling complexes
and non-coding RNA work together with histone modifications to
regulate gene expression and cell fate. As important epigenetic
regulatory machinery, histone modifications dynamically regulate
chromatin structure and gene expression through acetylation,
ubiquitination and Kla. These modifications are involved in
numerous biological processes such as proliferation, migration,
metabolic reprogramming and immune escape of tumor cells,
representing hot spots in GC research.
The present review focused on histone modification
in GC and systematically reviewed its molecular mechanism,
functional role and clinical transformation potential. It outlined
how different types of modifications regulate the development of GC
and explain how they affect the process of metabolism,
proliferation, migration and immunity to drive the progression of
tumors. It also focused on the clinical value of histone
modification (especially Kla modification) in GC as a diagnostic
biomarker and treatment target, as well as the prospect of
application in analyzing the mechanism of drug resistance.
Ultimately, the present review combined existing research results
to further transform these findings into clinical practice.
Availability of data and materials
Not applicable.
Authors' contributions
Writing and original draft preparation was by YZ and
SW. Writing, reviewing and editing was by YZ, SW and ML.
Visualization was by YZ, ML, WL, ZL and WL. Data curation and
investigation, was by LZ, XM and YG. SW supervised the review,
critically evaluated and revised the manuscript. Funding
acquisition was by SW and LM. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
This study was funded by the Science and Technology Project
Founded by the Education Department of Jiangxi Province (GJJ2201928
to LLW).
References
|
1
|
Smyth EC, Nilsson M, Grabsch HI, van
Grieken NC and Lordick F: Gastric cancer. Lancet. 396:635–648.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Guang D, Xiaofei Z, Yu M, Hui N, Min S and
Xiaonan S: Pomiferin targeting SLC9A9 based on histone acetylation
modification pattern is a potential therapeutical option for
gastric cancer with high malignancy. Biochem Pharmacol.
226:1163332024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Christodoulidis G, Koumarelas KE, Kouliou
MN, Thodou E and Samara M: Gastric cancer in the era of
epigenetics. Int J Mol Sci. 25:33812024. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yang Q, Chen Y, Guo R, Dai Y, Tang L, Zhao
Y, Wu X, Li M, Du F, Shen J, et al: Interaction of ncRNA and
epigenetic modifications in gastric cancer: Focus on histone
modification. Front Oncol. 11:8227452022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wang Y, Liu H, Zhang M, Xu J, Zheng L, Liu
P, Chen J, Liu H and Chen C: Epigenetic reprogramming in
gastrointestinal cancer: Biology and translational perspectives.
MedComm (2020). 5:e6702024. View
Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zong Z, Zhang L and Zhou F: Lactylation in
cancer: Advances, challenges, and future perspectives. Cancer Res.
85:3192–3195. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Han S, Wang R, Zhang Y, Li X, Gan Y, Gao
F, Rong P, Wang W and Li W: The role of ubiquitination and
deubiquitination in tumor invasion and metastasis. Int J Biol Sci.
18:2292–2303. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Regel I, Merkl L, Friedrich T,
Burgermeister E, Zimmermann W, Einwächter H, Herrmann K, Langer R,
Röcken C, Hofheinz R, et al: Pan-histone deacetylase inhibitor
panobinostat sensitizes gastric cancer cells to anthracyclines via
induction of CITED2. Gastroenterology. 143:99–109.e10. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yu X, Yang J, Xu J, Pan H, Wang W, Yu X
and Shi S: Histone lactylation: From tumor lactate metabolism to
epigenetic regulation. Int J Biol Sci. 20:1833–1854. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yang J, Xu J, Wang W, Zhang B, Yu X and
Shi S: Epigenetic regulation in the tumor microenvironment:
Molecular mechanisms and therapeutic targets. Signal Transduct
Target Ther. 8:2102023. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Schmitz KM, Mayer C, Postepska A and
Grummt I: Interaction of noncoding RNA with the rDNA promoter
mediates recruitment of DNMT3b and silencing of rRNA genes. Genes
Dev. 24:2264–2269. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Tsai MC, Manor O, Wan Y, Mosammaparast N,
Wang JK, Lan F, Shi Y, Segal E and Chang HY: Long noncoding RNA as
modular scaffold of histone modification complexes. Science.
329:689–693. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Arai J, Hayakawa Y, Suzuki N, Kinoshita H,
Hata M, Kurokawa K, Matsushita Y, Abe S, Oya Y, Tsuboi M, et al:
ARID1A mutation drives gastric tumorigenesis via activating type 2
immune dominant microenvironment. iScience. 28:1131172025.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Shi Y, Zhao Z, Zhou S, Zhou Z, Huang Z,
Zhou Z and Zhang C: CHD4 drives gastric cancer metastasis via
MYH9/GSK3β/β-catenin axis and WNT/EMT pathway activation. Cancer
Lett. 628:2178132025. View Article : Google Scholar
|
|
15
|
Hashimoto T, Kurokawa Y, Wada N, Takahashi
T, Miyazaki Y, Tanaka K, Makino T, Yamasaki M, Nakajima K, Mori M
and Doki Y: Clinical significance of chromatin remodeling factor
CHD5 expression in gastric cancer. Oncol Lett. 19:1066–1073.
2020.PubMed/NCBI
|
|
16
|
Luo M, Deng X, Chen Z and Hu Y: Circular
RNA circPOFUT1 enhances malignant phenotypes and
autophagy-associated chemoresistance via sequestrating miR-488-3p
to activate the PLAG1-ATG12 axis in gastric cancer. Cell Death Dis.
14:102023. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Deng P, Sun M, Zhao WY, Hou B, Li K, Zhang
T and Gu F: Circular RNA circVAPA promotes chemotherapy drug
resistance in gastric cancer progression by regulating
miR-125b-5p/STAT3 axis. World J Gastroenterol. 27:487–500. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wu C, Cao X, Xu J, Wang L, Huang J, Wen J,
Wang X, Sang X, Zhu W, Yao Y, et al: Hsa_circ_0073453 modulates
IL-8 secretion by GC-MSCs to promote gastric cancer progression by
sponging miR-146a-5p. Int Immunopharmacol. 119:1101212023.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Miao Z, Li J, Wang Y, Shi M, Gu X, Zhang
X, Wei F, Tang X, Zheng L and Xing Y: Hsa_circ_0136666 stimulates
gastric cancer progression and tumor immune escape by regulating
the miR-375/PRKDC Axis and PD-L1 phosphorylation. Mol Cancer.
22:2052023. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Han Z, Liu W, Zhu Y, Sun Y, Sun D, Jia R,
Yang Y, Qi H, Zhang L, Huo Y, et al: Non-coding RNAs in gastric
cancer: Mechanisms and therapeutic prospects. Mol Cancer.
24:2442025. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Dai E, Zhu Z, Wahed S, Qu Z, Storkus WJ
and Guo ZS: Epigenetic modulation of antitumor immunity for
improved cancer immunotherapy. Mol Cancer. 20:1712021. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dawson MA and Kouzarides T: Cancer
epigenetics: From mechanism to therapy. Cell. 150:12–27. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Audia JE and Campbell RM: Histone
modifications and cancer. Cold Spring Harb Perspect Biol.
8:a0195212016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Yun M, Wu J, Workman JL and Li B: Readers
of histone modifications. Cell Res. 21:564–578. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Cao J and Yan Q: Cancer epigenetics, tumor
immunity, and immunotherapy. Trends Cancer. 6:580–592. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Carrozza MJ, Utley RT, Workman JL and Côté
J: The diverse functions of histone acetyltransferase complexes.
Trends Genet. 19:321–329. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Salah Ud-Din AIM, Tikhomirova A and
Roujeinikova A: Structure and functional diversity of GCN5-related
N-acetyltransferases (GNAT). Int J Mol Sci. 17:10182016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang Z, Zang C, Cui K, Schones DE, Barski
A, Peng W and Zhao K: Genome-wide mapping of HATs and HDACs reveals
distinct functions in active and inactive genes. Cell.
138:1019–1031. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Fujisawa T and Filippakopoulos P:
Functions of bromodomain-containing proteins and their roles in
homeostasis and cancer. Nat Rev Mol Cell Biol. 18:246–262. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Qin ZY, Wang T, Su S, Shen LT, Zhu GX, Liu
Q, Zhang L, Liu KW, Zhang Y, Zhou ZH, et al: BRD4 promotes gastric
cancer progression and metastasis through acetylation-dependent
stabilization of snail. Cancer Res. 79:4869–4881. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Sun L, Zhang H and Gao P: Metabolic
reprogramming and epigenetic modifications on the path to cancer.
Protein Cell. 13:877–919. 2022. View Article : Google Scholar :
|
|
32
|
Shvedunova M and Akhtar A: Modulation of
cellular processes by histone and non-histone protein acetylation.
Nat Rev Mol Cell Biol. 23:329–349. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Seto E and Yoshida M: Erasers of histone
acetylation: The histone deacetylase enzymes. Cold Spring Harb
Perspect Biol. 6:a0187132014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Michalak EM, Burr ML, Bannister AJ and
Dawson MA: The roles of DNA, RNA and histone methylation in ageing
and cancer. Nat Rev Mol Cell Biol. 20:573–589. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Suraweera A, O'Byrne KJ and Richard DJ:
Epigenetic drugs in cancer therapy. Cancer Metastasis Rev.
44:372025. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Yang C, Zhang J, Ma Y, Wu C, Cui W and
Wang L: Histone methyltransferase and drug resistance in cancers. J
Exp Clin Cancer Res. 39:1732020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gong F and Miller KM: Histone methylation
and the DNA damage response. Mutat Res Rev Mutat Res. 780:37–47.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Marzochi LL, Cuzziol CI, Nascimento Filho
CHVD, Dos Santos JA, Castanhole-Nunes MMU, Pavarino ÉC, Guerra ENS
and Goloni-Bertollo EM: Use of histone methyltransferase inhibitors
in cancer treatment: A systematic review. Eur J Pharmacol.
944:1755902023. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhang Y, Chen J, Liu H, Mi R, Huang R, Li
X, Fan F, Xie X and Ding J: The role of histone methylase and
demethylase in antitumor immunity: A new direction for
immunotherapy. Front Immunol. 13:10998922023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Gray ZH, Honer MA, Ghatalia P, Shi Y and
Whetstine JR: 20 Years of histone lysine demethylases: From
discovery to the clinic and beyond. Cell. 188:1747–1783. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Srivastava R, Singh R, Jauhari S, Lodhi N
and Srivastava R: Histone demethylase modulation: Epigenetic
strategy to combat cancer progression. Epigenomes. 7:102023.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Sterling J, Menezes SV, Abbassi RH and
Munoz L: Histone lysine demethylases and their functions in cancer.
Int J Cancer. 148:2375–2388. 2021. View Article : Google Scholar
|
|
43
|
Koppenol WH, Bounds PL and Dang CV: Otto
Warburg's contributions to current concepts of cancer metabolism.
Nat Rev Cancer. 11:325–337. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Li X, Yang Y, Zhang B, Lin X, Fu X, An Y,
Zou Y, Wang JX, Wang Z and Yu T: Lactate metabolism in human health
and disease. Signal Transduct Target Ther. 7:3052022. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang D, Tang Z, Huang H, Zhou G, Cui C,
Weng Y, Liu W, Kim S, Lee S, Perez-Neut M, et al: Metabolic
regulation of gene expression by histone lactylation. Nature.
574:575–580. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wan L, Zhang H, Liu J, He Q, Zhao J, Pan
C, Zheng K and Tang Y: Lactylation and human disease. Expert Rev
Mol Med. 27:e102025. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Yang Z, Zheng Y and Gao Q: Lysine
lactylation in the regulation of tumor biology. Trends Endocrinol
Metab. 35:720–731. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zhao L, Qi H, Lv H, Liu W, Zhang R and
Yang A: Lactylation in health and disease: Physiological or
pathological? Theranostics. 15:1787–1821. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Varner EL, Trefely S, Bartee D, von
Krusenstiern E, Izzo L, Bekeova C, O'Connor RS, Seifert EL, Wellen
KE, Meier JL and Snyder NW: Quantification of lactoyl-CoA
(lactyl-CoA) by liquid chromatography mass spectrometry in
mammalian cells and tissues. Open Biol. 10:2001872020. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Dai X, Lv X, Thompson EW and Ostrikov KK:
Histone lactylation: Epigenetic mark of glycolytic switch. Trends
Genet. 38:124–127. 2022. View Article : Google Scholar
|
|
51
|
Li F, Si W, Xia L, Yin D, Wei T, Tao M,
Cui X, Yang J, Hong T and Wei R: Positive feedback regulation
between glycolysis and histone lactylation drives oncogenesis in
pancreatic ductal adenocarcinoma. Mol Cancer. 23:902024. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Ju J, Zhang H, Lin M, Yan Z, An L, Cao Z,
Geng D, Yue J, Tang Y, Tian L, et al: The alanyl-tRNA synthetase
AARS1 moonlights as a lactyltransferase to promote YAP signaling in
gastric cancer. J Clin Invest. 134:e1745872024. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Niu Z, Chen C, Wang S, Lu C, Wu Z, Wang A,
Mo J, Zhang J, Han Y, Yuan Y, et al: HBO1 catalyzes lysine
lactylation and mediates histone H3K9la to regulate gene
transcription. Nat Commun. 15:35612024. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Xie B, Zhang M, Li J, Cui J, Zhang P, Liu
F, Wu Y, Deng W, Ma J, Li X, et al: KAT8-catalyzed lactylation
promotes eEF1A2-mediated protein synthesis and colorectal
carcinogenesis. Proc Natl Acad Sci USA. 121:e23141281212024.
View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Dong H, Zhang J, Zhang H, Han Y, Lu C,
Chen C, Tan X, Wang S, Bai X, Zhai G, et al: YiaC and CobB regulate
lysine lactylation in Escherichia coli. Nat Commun. 13:66282022.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Moreno-Yruela C, Zhang D, Wei W, Bæk M,
Liu W, Gao J, Danková D, Nielsen AL, Bolding JE, Yang L, et al:
Class I histone deacetylases (HDAC1-3) are histone lysine
delactylases. Sci Adv. 8:eabi66962022. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Tsukihara S, Akiyama Y, Shimada S, Hatano
M, Igarashi Y, Taniai T, Tanji Y, Kodera K, Yasukawa K, Umeura K,
et al: Delactylase effects of SIRT1 on a positive feedback loop
involving the H19-glycolysis-histone lactylation in gastric cancer.
Oncogene. 44:724–738. 2025. View Article : Google Scholar
|
|
58
|
Fu C, Jiang W, Wang C, Song SJ, Tao H,
Zhang XG, Li WT, Jin X, Yu BB, Hao JJ, et al: AP001885.4 promotes
the proliferation of esophageal squamous cell carcinoma cells by
histone lactylation- and NF-κB (p65)-dependent transcription
activation and METTL3-mediated mRNA stability of c-myc. Anim Cells
Syst (Seoul). 28:536–550. 2024. View Article : Google Scholar
|
|
59
|
Yao X and Li C: Lactate dehydrogenase A
mediated histone lactylation induced the pyroptosis through
targeting HMGB1. Metab Brain Dis. 38:1543–1553. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Wu D, Spencer CB, Ortoga L, Zhang H and
Miao C: Histone lactylation-regulated METTL3 promotes ferroptosis
via m6A-modification on ACSL4 in sepsis-associated lung injury.
Redox Biol. 74:1031942024. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Lin X, Lei Y, Pan M, Hu C, Xie B, Wu W, Su
J, Li Y, Tan Y, Wei X, et al: Augmentation of scleral glycolysis
promotes myopia through histone lactylation. Cell Metab.
36:511–525.e7. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Hu XT, Wu XF, Xu JY and Xu X:
Lactate-mediated lactylation in human health and diseases: Progress
and remaining challenges. J Adv Res. 75:229–248. 2025. View Article : Google Scholar :
|
|
63
|
Song YH, Wang ZJ, Kang L, He ZX, Zhao SB,
Fang X, Li ZS, Wang SL and Bai Y: PADs and NETs in digestive
system: From physiology to pathology. Front Immunol.
14:10770412023. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Zhu D, Zhang Y and Wang S: Histone
citrullination: A new target for tumors. Mol Cancer. 20:902021.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Yuzhalin AE: Citrullination in cancer.
Cancer Res. 79:1274–1284. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Pitter MR, Kryczek I, Zhang H, Nagarsheth
N, Xia H, Wu Z, Tian Y, Okla K, Liao P, Wang W, et al: PAD4
controls tumor immunity via restraining the MHC class II machinery
in macrophages. Cell Rep. 43:1139422024. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Myers LK, Ouyang YX, Patel JR, Odens HH,
Woo-Rasberry V, Park J, Yi AK, Rosloniec EF, Brand DD, Stuart JM
and Kang AH: Role of citrullinated collagen in autoimmune
arthritis. Int J Mol Sci. 23:98332022. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Yu K and Proost P: Insights into
peptidylarginine deiminase expression and citrullination pathways.
Trends Cell Biol. 32:746–761. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Yao W, Hu X and Wang X: Crossing
epigenetic frontiers: The intersection of novel histone
modifications and diseases. Signal Transduct Target Ther.
9:2322024. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Wang Y, Lyu Y, Tu K, Xu Q, Yang Y, Salman
S, Le N, Lu H, Chen C, Zhu Y, et al: Histone citrullination by
PADI4 is required for HIF-dependent transcriptional responses to
hypoxia and tumor vascularization. Sci Adv. 7:eabe37712021.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Mondal S and Thompson PR: Protein arginine
deiminases (PADs): Biochemistry and chemical biology of protein
citrullination. Acc Chem Res. 52:818–832. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Popovic D, Vucic D and Dikic I:
Ubiquitination in disease pathogenesis and treatment. Nat Med.
20:1242–1253. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Zheng N and Shabek N: Ubiquitin ligases:
Structure, function, and regulation. Annu Rev Biochem. 86:129–157.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Gao H, Xi Z, Dai J, Xue J, Guan X, Zhao L,
Chen Z and Xing F: Drug resistance mechanisms and treatment
strategies mediated by Ubiquitin-Specific Proteases (USPs) in
cancers: New directions and therapeutic options. Mol Cancer.
23:882024. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Gao H, Chen Z, Zhao L, Ji C and Xing F:
Cellular functions, molecular signalings and therapeutic
applications: Translational potential of deubiquitylating enzyme
USP9X as a drug target in cancer treatment. Biochim Biophys Acta
Rev Cancer. 1879:1890992024. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Sun T, Liu Z and Yang Q: The role of
ubiquitination and deubiquitination in cancer metabolism. Mol
Cancer. 19:1462020. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Wu Z, Yang X, Yuan Z, Guo Y, Wang X and Qu
L: Identification of a novel histone acetylation-related long
non-coding RNA model combined with qRT-PCR experiments for
prognosis and therapy in gastric cancer. Heliyon. 10:e366152024.
View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Suo D, Gao X, Chen Q, Zeng T, Zhan J, Li
G, Zheng Y, Zhu S, Yun J, Guan XY and Li Y: HSPA4 upregulation
induces immune evasion via ALKBH5/CD58 axis in gastric cancer. J
Exp Clin Cancer Res. 43:1062024. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Han Y, Zhang YY, Pan YQ, Zheng XJ, Liao K,
Mo HY, Sheng H, Wu QN, Liu ZX, Zeng ZL, et al: IL-1β-associated NNT
acetylation orchestrates iron-sulfur cluster maintenance and cancer
immunotherapy resistance. Mol Cell. 83:1887–902.e8. 2023.
View Article : Google Scholar
|
|
80
|
Falkenberg KJ and Johnstone RW: Histone
deacetylases and their inhibitors in cancer, neurological diseases
and immune disorders. Nat Rev Drug Discov. 13:673–691. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Jenke R, Oliinyk D, Zenz T, Körfer J,
Schäker-Hübner L, Hansen FK, Lordick F, Meier-Rosar F, Aigner A and
Büch T: HDAC inhibitors activate lipid peroxidation and ferroptosis
in gastric cancer. Biochem Pharmacol. 225:1162572024. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Biersack B, Polat S and Höpfner M:
Anticancer properties of chimeric HDAC and kinase inhibitors. Semin
Cancer Biol. 83:472–486. 2022. View Article : Google Scholar
|
|
83
|
Yang Z, Jiang X, Zhang Z, Zhao Z, Xing W,
Liu Y, Jiang X and Zhao H: HDAC3-dependent transcriptional
repression of FOXA2 regulates FTO/m6A/MYC signaling to contribute
to the development of gastric cancer. Cancer Gene Ther. 28:141–155.
2021. View Article : Google Scholar
|
|
84
|
Lin Y, Jing X, Chen Z, Pan X, Xu D, Yu X,
Zhong F, Zhao L, Yang C, Wang B, et al: Histone
deacetylase-mediated tumor microenvironment characteristics and
synergistic immunotherapy in gastric cancer. Theranostics.
13:4574–4600. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Shetty MG, Pai P, Deaver RE, Satyamoorthy
K and Babitha KS: Histone deacetylase 2 selective inhibitors: A
versatile therapeutic strategy as next generation drug target in
cancer therapy. Pharmacol Res. 170:1056952021. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Badie A, Gaiddon C and Mellitzer G:
Histone deacetylase functions in gastric cancer: Therapeutic
target? Cancers (Basel). 14:54722022. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Hyun K, Jeon J, Park K and Kim J: Writing,
erasing and reading histone lysine methylations. Exp Mol Med.
49:e3242017. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Song Y, Wu F and Wu J: Targeting histone
methylation for cancer therapy: Enzymes, inhibitors, biological
activity and perspectives. J Hematol Oncol. 9:492016. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Morera L, Lübbert M and Jung M: Targeting
histone methyltransferases and demethylases in clinical trials for
cancer therapy. Clin Epigenetics. 8:572016. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Popovic R and Licht JD: Emerging
epigenetic targets and therapies in cancer medicine. Cancer Discov.
2:405–413. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
De Marco K, Sanese P, Simone C and Grossi
V: Histone and DNA methylation as epigenetic regulators of DNA
damage repair in gastric cancer and emerging therapeutic
opportunities. Cancers (Basel). 15:49762023. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Park J, Wu J, Szkop KJ, Jeong J, Jovanovic
P, Husmann D, Flores NM, Francis JW, Chen YC, Benitez AM, et al:
SMYD5 methylation of rpL40 links ribosomal output to gastric
cancer. Nature. 632:656–663. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Dong L, Zhu J, Deng A, Wei J, Li J, Mao X
and Jia Z: Relationship between histone demethylase LSD family and
development and prognosis of gastric cancer. Front Immunol.
14:11707732023. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Metzger E, Wissmann M, Yin N, Müller JM,
Schneider R, Peters AHFM, Günther T, Buettner R and Schüle R: LSD1
demethylates repressive histone marks to promote
androgen-receptor-dependent transcription. Nature. 437:436–439.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Wang F, Hao P, Pu Y, He X, He Q, Cui H and
Jiang S: KDM1A-driven RNF81 downregulation promotes gastric cancer
progression via KLF4 destabilization. Oncogene. 44:3864–3878. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Zheng L, Xu H, Di Y, Chen L, Liu J, Kang L
and Gao L: ELK4 promotes the development of gastric cancer by
inducing M2 polarization of macrophages through regulation of the
KDM5A-PJA2-KSR1 axis. J Transl Med. 19:3422021. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Wang Z, Cai H, Zhao E and Cui H: The
diverse roles of histone demethylase KDM4B in normal and cancer
development and progression. Front Cell Dev Biol. 9:7901292022.
View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Liu F, Wang Y, Yang Z, Cui X, Zheng L, Fu
Y, Shao W, Zhang L, Yang Q and Jia J: KDM6B promotes gastric
carcinogenesis and metastasis via upregulation of CXCR4 expression.
Cell Death Dis. 13:10682022. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Pitter MR and Zou W: Citrullination in
tumor immunity and therapy. J Clin Invest. 135:e1963482025.
View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Liu X, Arfman T, Wichapong K,
Reutelingsperger CPM, Voorberg J and Nicolaes GAF: PAD4 takes
charge during neutrophil activation: Impact of PAD4 mediated NET
formation on immune-mediated disease. J Thromb Haemost.
19:1607–1617. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Zheng Y, Zhao G, Xu B, Liu C, Li C, Zhang
X and Chang X: PADI4 has genetic susceptibility to gastric
carcinoma and upregulates CXCR2, KRT14 and TNF-α expression levels.
Oncotarget. 7:62159–62176. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Song S, Xiang Z, Li J, Ji J, Yan R, Zhu Z
and Yu Y: A novel citrullinated modification of histone 3 and its
regulatory mechanisms related to IPO-38 antibody-labeled protein.
Front Oncol. 9:3042019. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Zhang L, Li Q, Yang J, Xu P, Xuan Z, Xu J
and Xu Z: Cytosolic TGM2 promotes malignant progression in gastric
cancer by suppressing the TRIM21-mediated
ubiquitination/degradation of STAT1 in a GTP binding-dependent
modality. Cancer Commun (Lond). 43:123–149. 2023. View Article : Google Scholar :
|
|
104
|
Jiang T, Xia Y, Li Y, Lu C, Lin J, Shen Y,
Lv J, Xie L, Gu C, Xu Z and Wang L: TRIM29 promotes antitumor
immunity through enhancing IGF2BP1 ubiquitination and subsequent
PD-L1 downregulation in gastric cancer. Cancer Lett.
581:2165102024. View Article : Google Scholar
|
|
105
|
Zhou S, Xiao L, Hu L, Zuo F, Wang Y, Fei
B, Dai J and Zhou X: CAFs promote immune evasion in gastric cancer
through histone lactylation-mediated suppression of NCAPG
ubiquitination. J Transl Med. 23:9892025. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Lv M, Huang Y, Chen Y and Ding K:
Lactylation modification in cancer: Mechanisms, functions, and
therapeutic strategies. Exp Hematol Oncol. 14:322025. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Shan Z and Liu Y: Harnessing glycolysis in
gastric cancer: molecular targets, therapeutic strategies, and
clinical horizons. Front Immunol. 16:16289372025. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Duan Y, Zhan H, Wang Q, Li B, Gao H, Liu
D, Xu Q, Gao X, Liu Z, Gao P, et al: Integrated lactylome
characterization reveals the molecular dynamics of protein
regulation in gastrointestinal cancers. Adv Sci (Weinh).
11:e24002272024. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Yang D, Yin J, Shan L, Yi X, Zhang W and
Ding Y: Identification of lysine-lactylated substrates in gastric
cancer cells. iScience. 25:1046302022. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Zhou L, Wang Z, Huang Y, Zhang X, Jiang H,
Guo Z, Zhou G and Liu H: NLRP12 decreases TRIM25-mediated HK2
degradation to promote glycolysis and H3K18la in gastric cancer.
Cell Death Dis. 16:6152025. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Hui S, Ghergurovich JM, Morscher RJ, Jang
C, Teng X, Lu W, Esparza LA, Reya T, Le Zhan, Yanxiang Guo J, et
al: Glucose feeds the TCA cycle via circulating lactate. Nature.
551:115–118. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Lin K, Lin X and Luo F: IGF2BP3 boosts
lactate generation to accelerate gastric cancer immune evasion.
Apoptosis. 29:2147–2160. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Su J, Mao X, Wang L, Chen Z, Wang W, Zhao
C, Li G, Guo W and Hu Y: Lactate/GPR81 recruits regulatory T cells
by modulating CX3CL1 to promote immune resistance in a highly
glycolytic gastric cancer. Oncoimmunology. 13:23209512024.
View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Zhao Y, Jiang J, Zhou P, Deng K, Liu Z,
Yang M, Yang X, Li J, Li R and Xia J: H3K18 lactylation-mediated
VCAM1 expression promotes gastric cancer progression and metastasis
via AKT-mTOR-CXCL1 axis. Biochem Pharmacol. 222:1161202024.
View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Yang H, Yang S, He J, Li W, Zhang A, Li N,
Zhou G and Sun B: Glucose transporter 3 (GLUT3) promotes
lactylation modifications by regulating lactate dehydrogenase A
(LDHA) in gastric cancer. Cancer Cell Int. 23:3032023. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Ishaq AR, Younis T, Lin S, Usman M, Wang T
and Chen ZS: Phosphofructokinase-1 in cancer: A promising target
for diagnosis and therapy. Recent Pat Anticancer Drug Discov.
20:667–680. 2025. View Article : Google Scholar
|
|
117
|
Shen X, Shi H, Liu L, Chong S, Han J and
Gu Y: PFKM promotes the progression of gastric cancer by
up-regulating CNTN1 expression through H3K18la modification. Appl
Biochem Biotechnol. 197:5885–5901. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Zong Z, Xie F, Wang S, Wu X, Zhang Z, Yang
B and Zhou F: Alanyl-tRNA synthetase, AARS1, is a lactate sensor
and lactyltransferase that lactylates p53 and contributes to
tumorigenesis. Cell. 187:2375–92.e33. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Jiang G, Zhao Y, Zhang Y, Liu C, Wu J, Sun
Y, Xu Z and Shi X: Unraveling the prognostic significance and
molecular mechanisms of HKDC1 in cancer via integrated multi-omics
and experimental validation. Pathol Res Pract. 272:1561012025.
View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Huang M: Histone lactylation in
gastrointestinal cancers: Developing immunotherapeutic drugs
targeting epigenetics. Arch Pharm Res. 48:831–842. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Sun X, Dong H, Su R, Chen J, Li W, Yin S
and Zhang C: Lactylation-related gene signature accurately predicts
prognosis and immunotherapy response in gastric cancer. Front
Oncol. 14:14855802024. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Che X, Zhang Y, Chen X, Xie G, Li J, Xu C,
Zhang C, Zhu Y and Yang X: The lactylation-macrophage interplay:
Implications for gastrointestinal disease therapeutics. Front
Immunol. 16:16081152025. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Hu X, Ouyang W, Chen H, Liu Z, Lai Z and
Yao H: Claudin-9 (CLDN9) promotes gastric cancer progression by
enhancing the glycolysis pathway and facilitating PD-L1 lactylation
to suppress CD8+ T cell anti-tumor immunity. Cancer
Pathog Ther. 3:253–266. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Zhang G, Gao Z, Guo X, Ma R, Wang X, Zhou
P, Li C, Tang Z, Zhao R and Gao P: CAP2 promotes gastric cancer
metastasis by mediating the interaction between tumor cells and
tumor-associated macrophages. J Clin Invest. 133:e1662242023.
View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Zhang J, Dong Y, Yu S, Hu K, Zhang L,
Xiong M, Liu M, Sun X, Li S, Yuan Y, et al: IL-4/IL-4R axis
signaling drives resistance to immunotherapy by inducing the
upregulation of Fcγ receptor IIB in M2 macrophages. Cell Death Dis.
15:5002024. View Article : Google Scholar
|
|
126
|
Dong Z, Yuan Z, Jin T, Gao C, Wang X and
Xu F: Lactate at the crossroads of tumor metabolism and immune
escape: A new frontier in cancer therapy. J Transl Med.
23:12392025. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Faghihkhorasani F, Moosavi M, Rasool
Riyadh Abdulwahid AH, Kavei M, Karimi S, Seyed Karimi M, Vezvaei P,
Manafi Varkiani M, Aref AR and Ebrahimi N: Role of monocarboxylate
transporters in cancer immunology and their therapeutic potential.
Br J Pharmacol. 182:4421–4457. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Zhao Z, Han F, Yang S, Wu J and Zhan W:
Oxamate-mediated inhibition of lactate dehydrogenase induces
protective autophagy in gastric cancer cells: Involvement of the
Akt-mTOR signaling pathway. Cancer Lett. 358:17–26. 2015.
View Article : Google Scholar
|
|
129
|
Zhang X, Li Y and Chen Y: Development of a
comprehensive gene signature linking hypoxia, glycolysis,
lactylation, and metabolomic insights in gastric cancer through the
integration of bulk and single-Cell RNA-Seq data. Biomedicines.
11:29482023. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Yang H, Zou X, Yang S, Zhang A, Li N and
Ma Z: Identification of lactylation related model to predict
prognostic, tumor infiltrating immunocytes and response of
immunotherapy in gastric cancer. Front Immunol. 14:11499892023.
View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Ramezankhani R, Solhi R, Es HA, Vosough M
and Hassan M: Novel molecular targets in gastric adenocarcinoma.
Pharmacol Ther. 220:1077142021. View Article : Google Scholar
|
|
132
|
Chen AN, Luo Y, Yang YH, Fu JT, Geng XM,
Shi JP and Yang J: Lactylation, a novel metabolic reprogramming
code: Current status and prospects. Front Immunol. 12:6889102021.
View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Ferreira S, Saraiva N, Rijo P and
Fernandes AS: LOXL2 inhibitors and breast cancer progression.
Antioxidants (Basel). 10:3122021. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Xu Y, Wang X, Huang Y, Ma Y, Jin X, Wang H
and Wang J: Inhibition of lysyl oxidase expression by dextran
sulfate affects invasion and migration of gastric cancer cells. Int
J Mol Med. 42:2737–2749. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Li Z, Liang P, Chen Z, Chen Z, Jin T, He
F, Chen X and Yang K: CAF-secreted LOX promotes PD-L1 expression
via histone Lactylation and regulates tumor EMT through TGFβ/IGF1
signaling in gastric cancer. Cell Signal. 124:1114622024.
View Article : Google Scholar
|
|
136
|
Lee JY, Lee I, Chang WJ, Ahn SM, Lim SH,
Kim HS, Yoo KH, Jung KS, Song HN, Cho JH, et al: MCT4 as a
potential therapeutic target for metastatic gastric cancer with
peritoneal carcinomatosis. Oncotarget. 7:43492–43503. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Feng Q, Liu Z, Yu X, Huang T, Chen J, Wang
J, Wilhelm J, Li S, Song J, Li W, et al: Lactate increases stemness
of CD8 + T cells to augment anti-tumor immunity. Nat Commun.
13:49812022. View Article : Google Scholar
|
|
138
|
Zhang P, Xiang H, Peng Q, Ma L, Weng C,
Liu G and Lu L: METTL14 attenuates cancer stemness by suppressing
ATF5/WDR74/β-catenin axis in gastric cancer. Cancer Sci.
116:112–127. 2025. View Article : Google Scholar :
|