Introduction
Gestational choriocarcinoma is a type of gestational
trophoblastic neoplasia (GTN) that originates from trophoblasts and
can develop from a normal pregnancy, miscarriage, or molar
pregnancy. Its estimated incidence in Japan is 1.9-5.5 cases per
100,000 live births (1).
Non-gestational choriocarcinoma shows the same morphological
pattern as that of the gestational form but originates mostly from
germ cells in the ovary and is rarer and associated with worse
outcomes. Short tandem repeat analysis using microsatellite markers
is useful for distinguishing gestational choriocarcinoma from
non-gestational choriocarcinoma (2).
Chemotherapy is effective for treating gestational
choriocarcinoma. However, patients with multiple metastases or
metastases to sites other than the lungs often do not achieve
complete remission (3). When the
cancer develops chemoresistance, more tailored therapies are
required, such as drugs selected based on the specific cancer
genome. The OncoGuide™ NCC Oncopanel System (Sysmex
Corporation) (4) is a
next-generation sequencing (NGS)-based tumor panel that is covered
by health insurance in Japan. This panel facilitates the
identification of variants of 114 cancer-related genes through
matched normal-tumor pair analysis. Here, we report a case of
intractable gestational choriocarcinoma identified using this
system.
Case report
A 51-year-old Japanese woman was diagnosed with
choriocarcinoma with metastases to the lung, spleen, and lymph
nodes. Histopathological examination of the uterine biopsy showed a
two-cell pattern of choriocarcinoma, consisting of
syncytiotrophoblastic cells and cytotrophoblastic cells. She had
experienced six pregnancies, and the last pregnancy ended in
spontaneous abortion approximately 4 years prior. The patient was
treated with etoposide, methotrexate, actinomycin D,
cyclophosphamide, and vincristine (EMA/CO) and had suspected
drug-induced pneumonia after the third course (Fig. 1). Therefore, she could not continue
EMA-CO therapy although it was effective. The regimen was modified,
but the new regimen proved ineffective. She was then referred to
our institution for further treatment. We performed a drug-induced
lymphocyte stimulation test and found that the anticancer drugs
etoposide, etoposide, methotrexate, actinomycin, cyclophosphamide,
and vincristine did not induce the allergy. Thus, we concluded that
her pneumonia was induced by infection and that we could use these
anticancer drugs. After obtaining written informed consent from the
patient and her partner, we performed short tandem repeat analysis
of DNA extracted from the oral mucosal cells of the patient and her
partner and from the paraffin-embedded sections of the
micro-dissected tumor, as previously described (5). This study was approved by the Ethics
Committee of Nagoya University Graduate School of Medicine. Tumor
analysis revealed gestational choriocarcinoma of both maternal and
paternal origins (Table I). The
patient underwent four types of chemotherapy regimens and was then
treated with high-dose ifosfamide, carboplatin, and etoposide
(ICE), along with autologous peripheral blood stem cell
transplantation (2,6,7).
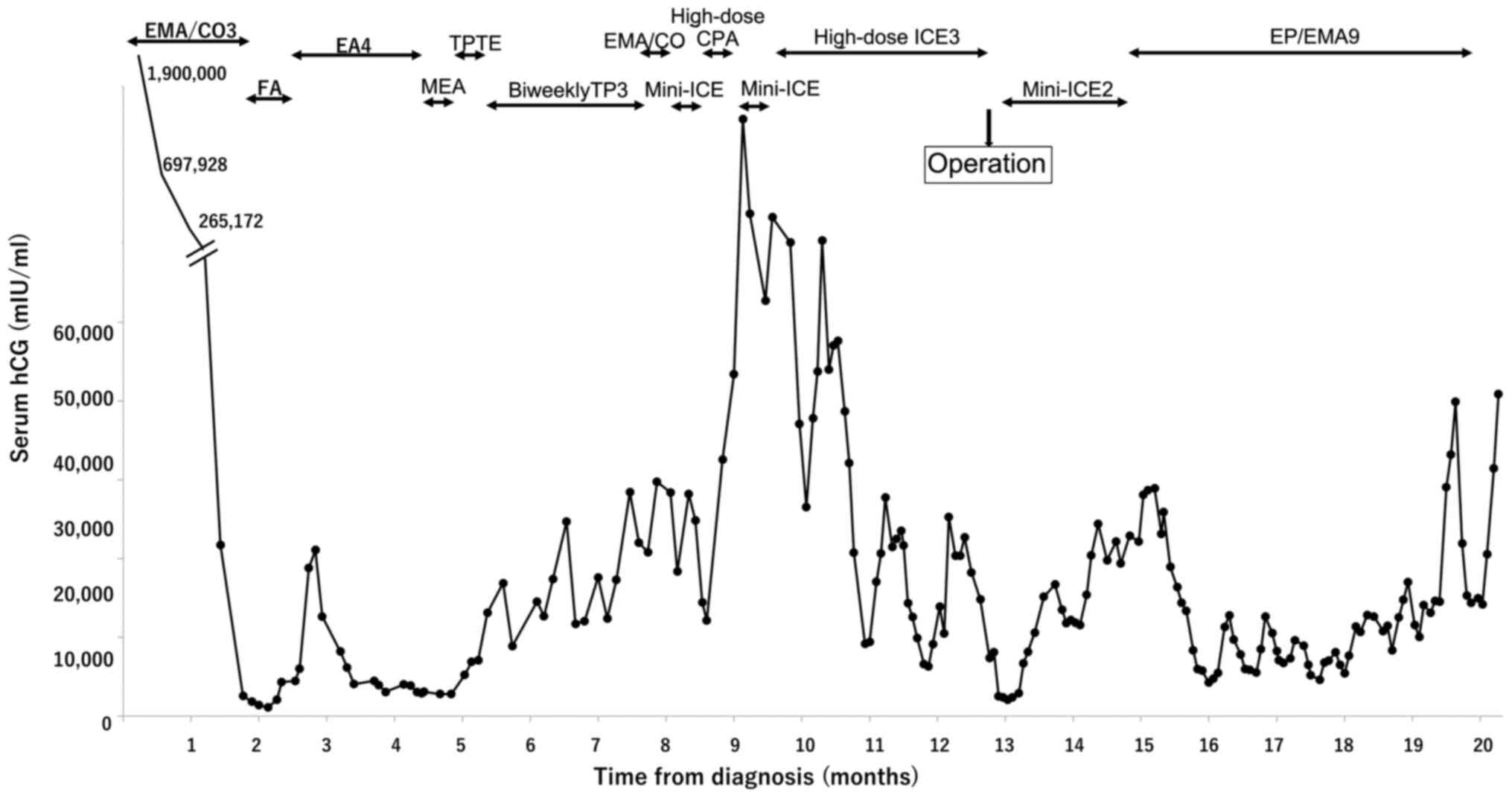 | Figure 1Changes in the serum hCG level of the
patient and the treatment progress of choriocarcinoma. EMA/CO,
etoposide, methotrexate, actinomycin D, cyclophosphamide, and
vincristine; FA, fluorouracil and actinomycin D; EA, etoposide and
actinomycin D; MEA, methotrexate, etoposide, and actinomycin D;
TPTE, paclitaxel, cisplatin, and etoposide; biweekly TP, biweekly
paclitaxel and cisplatin; ICE, ifosfamide, carboplatin, and
etoposide; CPA, cyclophosphamide; EP/EMA, etoposide, cisplatin,
methotrexate, and actinomycin D; hCG, human chorionic
gonadotropin. |
 | Table IShort tandem repeat analysis of DNA
from the tumor, patient and her partner. |
Table I
Short tandem repeat analysis of DNA
from the tumor, patient and her partner.
| Marker | Maternal | Paternal | Tumor |
|---|
| D8S1179 | 10,14 | 13,13 | 10,13 |
| D21S11 | 30,31 | 30,30 | 30 |
| D7S820 | 11,12 | 9,12 | 9,12 |
| CFS1PO | 10,11 | 10,10 | 10 |
| D3S1358 | 16,16 | 16,17 | 16 |
| TH01 | 6,6 | 6,9 | 6 |
| D13S317 | 11,12 | 11,11 | 11,12 |
| D16S539 | 11,11 | 9,9 | 9,11 |
| D2S1338 | 17,20 | 23 | - |
| D19S433 | 13,13 | 13,15.2 | 13 |
| vWA | 17,19 | 16,16 | 16,17,19 |
| TPOX | 8,11 | 8,11 | 11 |
| D18S51 | 14,18 | 14,17 | 17,18 |
| Amerogenin | X,X | X,Y | X,X |
| D5S818 | 10,12 | 9,11 | 10,11,12 |
| FGA | 23,26 | 23,24 | 23,24,26 |
After three courses of high-dose ICE, we performed
total hysterectomy and bilateral adnexectomy to reduce the total
choriocarcinoma volume. The patient was administered two courses of
mini-ICE after the operation, but multiple metastases were found in
the brain. She was thus treated with whole-brain radiotherapy (20
Gy); etoposide, cisplatin, methotrexate, and actinomycin D (EP-EMA)
chemotherapy; and radiotherapy for the bone metastases.
To identify drugs appropriate for treating the
choriocarcinoma in this case, we utilized the NCC Oncopanel System
to compare the uterine choriocarcinoma DNA with the patient's
germline DNA extracted from peripheral blood. Microdissection was
performed to obtain the choriocarcinoma tissue from
formaldehyde-fixed and paraffin-embedded tissue sections (10 µm
thickness). The samples were prepared and analyzed as previously
reported (4). NCC Oncopanel test
revealed 245 single-nucleotide variants (SNVs). Compared with the
usual allele frequency of SNVs in matched normal-tumor pair
analysis of ≤30%, the mean SNV allele frequency of the patient was
more than double, at 63.1%. Initially, experimental errors such as
sample misidentification were suspected; however, we eventually
concluded that part of the gestational choriocarcinoma DNA was
derived from the partner of the patient, whereby the SNV burden was
increased.
The tumor DNA contained 19 variants in 13 of the 114
cancer-related genes (Table II).
The GNAQ p.T96S and TP53 p.R213P variants were
considered to be pathogenic variants; the remaining 17 variants are
frequent in the Japanese population. There are no targeted
therapies for these two pathogenic variants. After nine courses of
EP-EMA, the patient was unable to undergo chemotherapy because of
pancytopenia and febrile neutropenia. She was treated for 20 months
but ultimately died of choriocarcinoma 7 months after the
operation.
 | Table IIGenomic findings for the tumor and
blood of the patient, obtained using the NCC Oncopanel System
Test. |
Table II
Genomic findings for the tumor and
blood of the patient, obtained using the NCC Oncopanel System
Test.
| Gene name | Mutation allele
frequency | Amino acid
change | dbSNP | HGVD allele
frequency |
|---|
| BARD1 | 66.3 | R24S | rs1048108 | 0.350 |
| SETD2 | 68.0 | M1080I | rs76208147 | 0.143 |
| ROS1 | 64.9 | S2229C | rs619203 | 0.145 |
| ROS1 | 67.9 | K2228Q | rs529156 | 0.146 |
| ROS1 | 68.4 | D2213N | rs529038 | 0.151 |
| GNAQ | 12.8 | T96S | rs777679970 | Νot detected |
| TP53 | 61.7 | R213P | rs587778720 | Νot detected |
| BRCA1 | 66.5 | S1613G | rs1799966 | 0.331 |
| BRCA1 | 62.8 | K1183R | rs16942 | 0.329 |
| BRCA1 | 63.0 | E1038G | rs16941 | 0.329 |
| BRCA1 | 70.4 | R871L | rs799917 | 0.331 |
| FGFR4 | 74.9 | G388R | rs351855 | 0.414 |
| NOTCH2 | 69.0 | R1260H | rs75423398 | 0.070 |
| PRKCI | 73.0 | R327R | rs55683301 | 0.061 |
| ESR1 | 78.5 | P146Q | rs17847065 | 0.047 |
| PTCH1 | 70.1 | R893H | rs138154222 | 0.019 |
| BRCA2 | 62.8 | M784V | rs11571653 | 0.095 |
| CREBBP | 64.6 | L551I | rs61753381 | 0.032 |
| BRCA1 | 51.0 | Y856H | rs80356892 | 0.009 |
Discussion
This is the first study using the NCC Oncopanel
System test for gestational choriocarcinoma. The test was performed
to seek appropriate drugs for the intractable choriocarcinoma, but
no drug matched the tumor genome. A hospital-based prospective
study using the NCC Oncopanel System test showed that only 13.4% of
the patients were eligible for targeted drug therapies based on the
sequencing results, and this result is similar to that obtained
using another cancer-gene panel (11%) (8). The relatively low likelihood of
identifying a targeted therapy should be explained to patients
before applying a gene-panel test. Additional genome-matched
clinical trials are required to determine the applications that
these tests would suit the most.
The results of the NCC Oncopanel test in our case
indicate two limitations to using NGS-based tumor-profiling
multiplex gene panels for GTN patients. First, panel tests for
tumor and matched non-tumor samples, such as the NCC Oncopanel
test, show a high SNV burden in the genomic DNA from GTNs, and such
results may be misinterpreted as ‘tumor-derived’ variants. The NCC
Oncopanel test is inappropriate for tumors like GTNs containing the
DNA of other persons. Tumor-profiling gene-testing using only tumor
samples should be used for GTNs. Second, the patient, her partner,
and/or their children might be the source of pathogenic variants or
secondary genetic findings in the tumor DNA of GTNs. A case of
choriocarcinoma in a woman whose partner had a genomic TP53
variant leading to Li-Fraumeni syndrome has been reported, wherein
the TP53 variant was detected in her tumor but not in her
germline DNA (9). Since there are
ethical issues associated with genomic screening in GTN cases,
informed consent should be obtained from patients and their
partners, and specific ethical guidelines should be laid down for
tumor-panel testing of patients with GTN.
In the present case, two pathogenic variants
(GNAQ p.T96S and TP53 p.R213P) were identified in the
tumor DNA. The sequence report from the NCC Oncopanel revealed a
12.8% variant allele frequency for GNAQ p.T96S, classifying
this allele as a somatic variant. The variant allele frequency of
TP53 p.R213P was 61.7%, indicating that this allele might
have been a germline variant, but the DNA of the patient's blood
did not have it. We realized that this allele might have been a
true somatic variant in her tumor or a germline variant in the
partner, one of the children of the patient, or the lost pregnancy,
because the tumor was a gestational choriocarcinoma. Genetic
counseling sessions were conducted with the partner of the patient
to discuss our findings for this variant and its association with
Li-Fraumeni syndrome. Upon the request of the partner, we checked
the existence of TP53 p.R213P variants with only his blood
but not her children's blood. We found that he did not have this
variant.
We here report a case of intractable gestational
choriocarcinoma resistant to numerous chemotherapies, including
high-dose ICE with peripheral stem cell rescue. It is suggested
that it is difficult for choriocarcinoma patients to achieve
complete remission when the second chemotherapy regimen fails and
multiple metastases exist (3).
High-dose chemotherapy with stem cell rescue and anti-programmed
cell death-1 (PD-1) antibody therapy might be an option for
intractable choriocarcinoma (2,7,10). The
effectiveness of anti-PD-1 antibody therapy for intractable GTN
patients has recently been reported (10). Our patient was not eligible for this
treatment because her choriocarcinoma did not show a high
microsatellite instability status, which is required for
health-insurance coverage in Japan. Clinical trials of anti-PD-1
antibody therapy for intractable GTN are needed, as this therapy
has been shown to be effective for patients with GTN with unknown
microsatellite instability statuses (11).
In conclusion, our experience of an intractable
choriocarcinoma case screened with the NCC Oncopanel System
suggests that matched normal-tumor pair analysis is not appropriate
for GTN. When using an NGS-based tumor panel to assess
choriocarcinoma, researchers must consider whether the patient's
and partner's genomic DNA is involved in the GTN.
Acknowledgements
Not applicable.
Funding
This work was financially supported by grants-in-aid numbers
17K16845 and 20K09639 (to KN) by the Ministry of Education,
Culture, Sports, Science, and Technology of Japan. The funding
bodies had no role in study design or data collection, analysis or
interpretation.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
KNii, EY, SM, HH, MMorik, HK and FK designed the
study. KNii wrote the final manuscript. SM, HH, MMorik, MH and
MMorit analyzed and interpreted the data, and wrote the outline of
the manuscript regarding genetic analysis. MMorik, MH and MMorit
were responsible for genetic counseling. KNis, YO, EW and TY
analyzed and interpreted the data of STR analysis. KNii, KNis, YO,
EW, HK and FK were involved in the treatment of patients as
attending physicians and provided important advice for decision
making. EY, TY, HK and FK critically revised the paper for
important intellectual content. KNii and EY confirmed the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
Written informed consent was obtained from the study
subjects prior to the collection of all the biological samples
according to the regulations set out by the Ethics Committee at
Nagoya University.
Patient consent for publication
Written informed consent was obtained from the
partner of the patient for the publication of these data.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Albrecht C, Chamley L, Charnock-Jones DS,
Collins S, Fujiwara H, Golos T, Grayo S, Hannan N, Harris L,
Ichizuka K, et al: IFPA meeting 2018 workshop report II: Abnormally
invasive placenta; inflammation and infection; preeclampsia;
gestational trophoblastic disease and drug delivery. Placenta.
84:9–13. 2019.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Yamamoto E, Niimi K, Fujikake K, Nishida
T, Murata M, Mitsuma A, Ando Y and Kikkawa F: High-dose
chemotherapy with autologous peripheral blood stem cell
transplantation for choriocarcinoma: A case report and literature
review. Mol Clin Oncol. 5:660–664. 2016.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Powles T, Savage PM, Stebbing J, Short D,
Young A, Bower M, Pappin C, Schmid P and Seckl MJ: A comparison of
patients with relapsed and chemo-refractory gestational
trophoblastic neoplasia. Br J Cancer. 96:732–737. 2007.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Sunami K, Ichikawa H, Kubo T, Kato M,
Fujiwara Y, Shimomura A, Koyama T, Kakishima H, Kitami M,
Matsushima H, et al: Feasibility and utility of a panel testing for
114 cancer-associated genes in a clinical setting: A hospital-based
study. Cancer Sci. 110:1480–1490. 2019.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Yamamoto E, Niimi K, Shinjo K, Yamamoto T,
Fukunaga M and Kikkawa F: Identification of causative pregnancy of
gestational trophoblastic neoplasia diagnosed during pregnancy by
short tandem repeat analysis. Gynecol Oncol Case Rep. 9:3–6.
2014.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Piamsomboon S, Kudelka AP,
Termrungruanglert W, Van Besien K, Edwards CL, Lifshitz S, Schomer
DF, Champlin R, Mante RP, Kavanagh JJ and Verschraegen CF:
Remission of refractory gestational trophoblastic disease in the
brain with ifosfamide, carboplatin, and etoposide (ICE): First
report and review of literature. Eur J Gynaecol Oncol. 18:453–456.
1997.PubMed/NCBI
|
|
7
|
van Besien K, Verschraegen C, Mehra R,
Giralt S, Kudelka AP, Edwards CL, Piamsonboom S, Termrungruanglert
W, Champlin R and Kavanagh JJ: Complete remission of refractory
gestational trophoblastic disease with brain metastases treated
with multicycle ifosfamide, carboplatin, and etoposide (ICE) and
stem cell rescue. Gynecol Oncol. 65:366–369. 1997.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Zehir A, Benayed R, Shah RH, Syed A,
Middha S, Kim HR, Srinivasan P, Gao J, Chakravarty D, Devlin SM, et
al: Mutational landscape of metastatic cancer revealed from
prospective clinical sequencing of 10,000 patients. Nat Med.
23:703–713. 2017.PubMed/NCBI View
Article : Google Scholar
|
|
9
|
Cotter JA, Szymanski L, Karimov C,
Boghossian L, Margol A, Dhall G, Tamrazi B, Varaprasathan GI,
Parham DM, Judkins AR and Biegel JA: Transmission of a TP53
germline mutation from unaffected male carrier associated with
pediatric glioblastoma in his child and gestational choriocarcinoma
in his female partner. Cold Spring Harb Mol Case Stud.
4(a002576)2018.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Choi MC, Oh J and Lee C: Effective
anti-programmed cell death 1 treatment for chemoresistant
gestational trophoblastic neoplasia. Eur J Cancer. 121:94–97.
2019.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Ghorani E, Kaur B, Fisher RA, Short D,
Joneborg U, Carlson JW, Akarca A, Marafioti T, Quezada SA, Sarwar N
and Seckl MJ: Pembrolizumab is effective for drug-resistant
gestational trophoblastic neoplasia. Lancet. 390:2343–2345.
2017.PubMed/NCBI View Article : Google Scholar
|















