Introduction
Cytological examination of the thyroid gland by
fine-needle aspiration biopsy (FNAB) of thyroid nodules is
important for the verification of a pathology and for differential
diagnosis between benign nodular neoplasms and malignant tumors
(1-3).
Differential diagnosis of Hürthle cell (oncocytic)
tumors of the thyroid gland represents a special problem. Hürthle
cells are normally associated with lymphocytic thyroiditis
(Hashimoto's disease) and multinodular goiters; however, these
cells can also give rise to such tumors as Hürthle cell adenoma
(HCA) and Hürthle cell carcinoma (HCC). The WHO Classification of
Tumors has singled out HCA and HCC as nosological entities other
than follicular tumors, because of morphological and genetic
differences between these tumors (4). According to the Bethesda System for
Reporting Thyroid Cytopathology 2018, HCA and HCC are in diagnostic
category IV (suspicious for a follicular neoplasm), with molecular
testing or surgical lobectomy recommended (1).
A typical feature of tumors that grow from Hürthle
cells is that cellular atypia does not necessarily mean malignancy
(1,5,6). FNAB
material appears to consist exclusively of Hürthle cells with
abundant finely granular cytoplasm, an enlarged central or
eccentric round nucleus, and prominent nucleolus. Some cells are
stand-alone entities, and others are in syncytial-like sheets. The
oncocytes vary in size from giant cells to small monomorphic cells
with a high nuclear-to-cytoplasmic ratio. Some cells feature marked
anisonucleosis: A large hyperchromic nucleus with irregular
contours of the nuclear membrane, sometimes with nuclear grooves
and pseudo-inclusions.
As mentioned above, nuclear atypia, which is
indicative of Hürthle cells, is by itself not a criterion for
malignancy of a tumor. Hürthle cell patterns should be
distinguished between nontumors (Hashimoto's disease and
multinodular goiters) and tumors, including papillary thyroid
carcinoma, follicular thyroid carcinoma, and medullary thyroid
carcinoma. Cytologically, HCA and HCC are identical; accordingly,
the diagnosis can be made only after histological examination
following nodule resection. Consequently, due to additional
molecular oncomarkers in FNAB, clinical decision making is expected
to become easier, and prognoses may become more accurate.
Over the past decade, it has been demonstrated that
microRNAs (miRNAs, miRs)-short (18- to 24-nucleotide-long) RNA
molecules with an important role in post-transcriptional regulation
of a large number of genes-can be used as biological oncomarkers
(7-9).
Levels of many miRNAs are substantially altered in diverse tumors,
including thyroid neoplasms (10-12).
Isolation and profiling of miRNA in the material used for
cytological examination of thyroid nodules can improve diagnostic
accuracy when combined with the cytological method of
diagnosis.
In a recent study we described our original
diagnostic algorithm to identify and type malignant thyroid tumors
(including HCC) via analysis of a small number of molecular markers
in FNAC preparations (levels of HMGA2 mRNA and miR-375, -221, and
-146b in combination with the mitochondrial-to-nuclear DNA ratio)
(13). It has already been
validated on 122 samples (14), but
at this validation, we did not focus on Hürthle cell tumors.
The aim of the present study was to assess the
effectiveness of the same algorithm at detecting Hürthle cell
thyroid tumors in three well-characterized samples obtained by FNAB
and used for cytological examination.
Materials and methods
Clinical material
A cytological examination of the FNAB material and
intraoperative imprint cytological analysis of the thyroid lesions
were performed for three patients with nodular neoplasms. The
cytological smears and intraoperative imprints was air-dried and
stained by a standard procedure [May-Grünwald-Giemsa (MGG)
staining]. Histological analysis of surgically removed thyroid
tissue was conducted by staining tissue sections (5-µm thick) with
hematoxylin and eosin. Briefly, after resection of the thyroid
lobe, the tissue is fixed in formalin for 24 h at room temperature
then histological processing of the tissue was carried out using
vacuum infiltration processor Tissue-Tek® VIP™ 6 (Sakura
Finetek Europe B.V., The Netherlands) with the routine overnight
run program (processing time: 12.7 h, temp=40-58˚C). After
sectioning of the paraffin-embedded tissue, the sections were
manually stained with hematoxylin (NPF ABRIS+) and eosin
(ErgoProduction), following a basic protocol: Dewaxing (with
xylene), dehydration (ethanol then water), hematoxylin (10 min),
differentiation (acid alcohol), bluing (aqueous ammonium
hydroxide), eosin (5 min), dehydration (ethanol then water),
clearing (xylene), cover-slipping. The preparations were examined
under a light microscope with a magnification of 100 and 400. The
preparations were examined under a light microscope with a
magnification of 100 and 400. The histological and cytological
material was obtained in accordance with Russian laws and
regulations; all data were depersonalized.
Total-nucleic-acid isolation
Nucleic acid was isolated from the cytological
preparations (reviewed in ref. 13). The dried cytological
preparation was washed into a microcentrifuge tube with three 200
µl portions of guanidine lysis buffer, then the sample was
vigorously mixed and incubated in a thermal shaker for 15 min at
65˚C. Next, an equal volume of isopropanol was added. The reaction
solution was thoroughly mixed and kept at room temperature for 5
min. After centrifugation for 10 min at 14,00 x g, the supernatant
was discarded, and the pellet was washed with 500 µl of 70% ethanol
and 300 µl of acetone. Finally, the RNA was dissolved in 200 µl of
deionized water. If not analyzed immediately, RNA samples were
stored at -20˚C until further use.
Molecular analysis
Assessment of relative expression levels of the
HMGA2 gene (normalized to housekeeping gene PGK1),
miR-146b, -221, and -375 (normalized to the geometric mean of
miR-29b, -23a, and -197 levels) and calculation of the ratio of
mitochondrial DNA (mtDNA) to nuclear DNA (nDNA) were done as
described previously (13). The
details are presented below.
MiRNA detection and quantitation
The detection of the 6 miRNAs was performed on three
samples of oncocytic tumors. To quantify miRNA, we followed the
protocol published by Chen et al in 2005, which includes
reverse transcription of mature miRNA using a long stem-loop primer
followed by the detection of cDNA via real-time PCR (15). Reverse-transcription reactions were
set up individually for each miRNA to be quantified. The obtained
cDNA was subjected to further PCR analysis immediately. The
reverse-transcription reaction and real-time PCR were carried out
(reviewed in ref. 16). The miRNA content was normalized to the
geometric mean of the amounts of the three reference miRNAs by the
2-ΔCq method (17).
Quantification of HMGA2 mRNA
The relative concentration of HMGA2 mRNA was
estimated by real-time RT-PCR, where PGK1 (phosphoglycerate
kinase 1) mRNA served for normalization. The following RT-PCR
program was employed: Incubation for 30 min at 45˚C, preheating for
2 min at 95˚C, and then 50 cycles of denaturation for 10 sec at
94˚C with annealing and elongation for 20 sec at 60˚C (13). The relative expression level was
calculated by the 2-ΔCq method.
Determination of the mtDNA/nDNA
ratio
Detection of specific sites in mtDNA and nDNA was
performed independently by real-time PCR. The thermal cycling
conditions were as follows: Preheating for 2 min at 95˚C, then 50
cycles of denaturation for 10 sec at 94˚C with annealing and
elongation for 20 sec at 60˚C (13). The ratio was determined with the
2-ΔCq method.
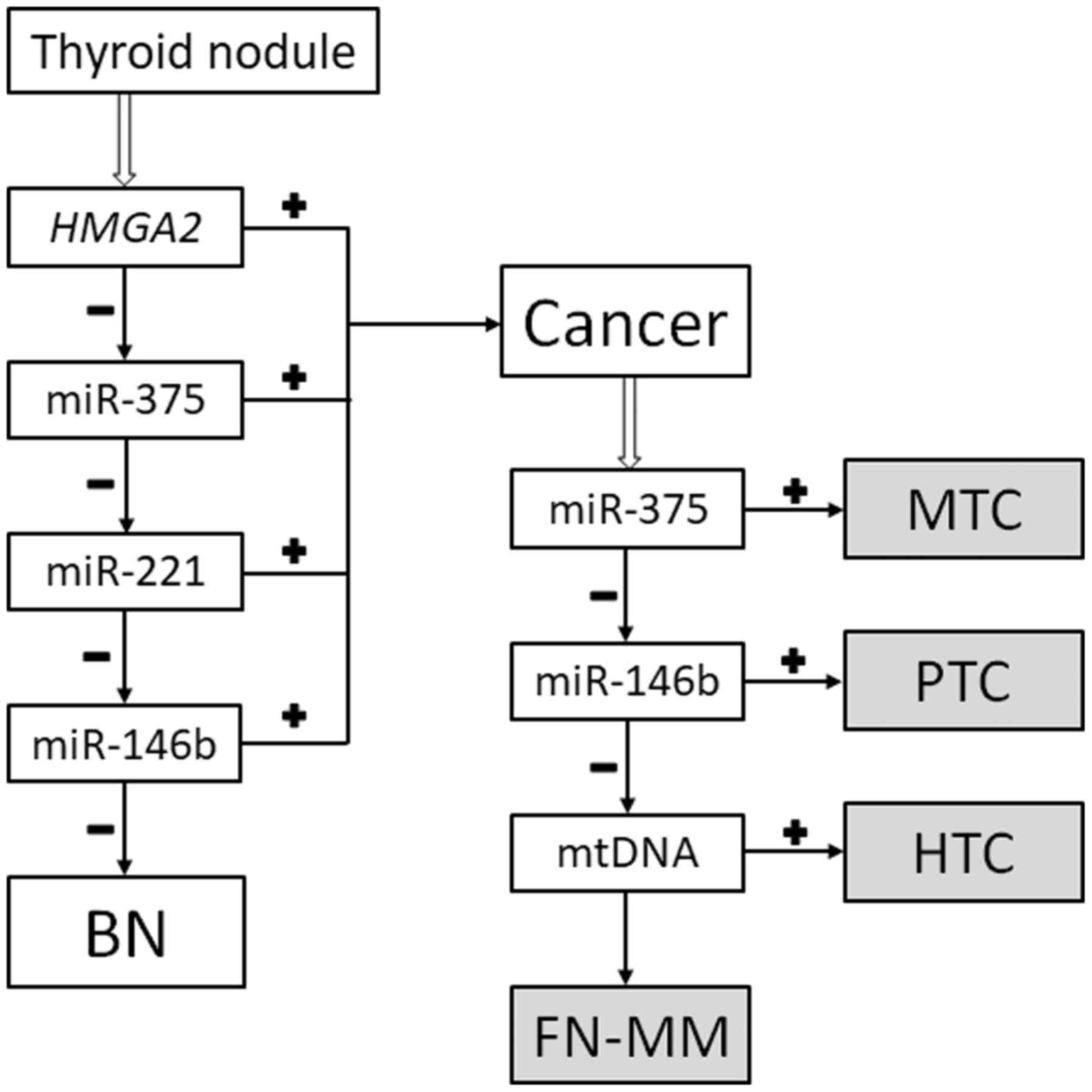
The classifier
The classification method (reviewed in ref. 13) was
used (Fig. 1). Thus, the specimens
are classified as either benign (goiters and follicular neoplasms
without markers of malignancy) or malignant (papillary thyroid
carcinoma, medullary thyroid carcinoma, HCC, and a follicular
neoplasm with markers of malignancy). Additionally, the ratio of
mtDNA to nDNA allows to determine whether a given case of papillary
thyroid carcinoma can be categorized as the Hürthle cell type.
Results
Case 1
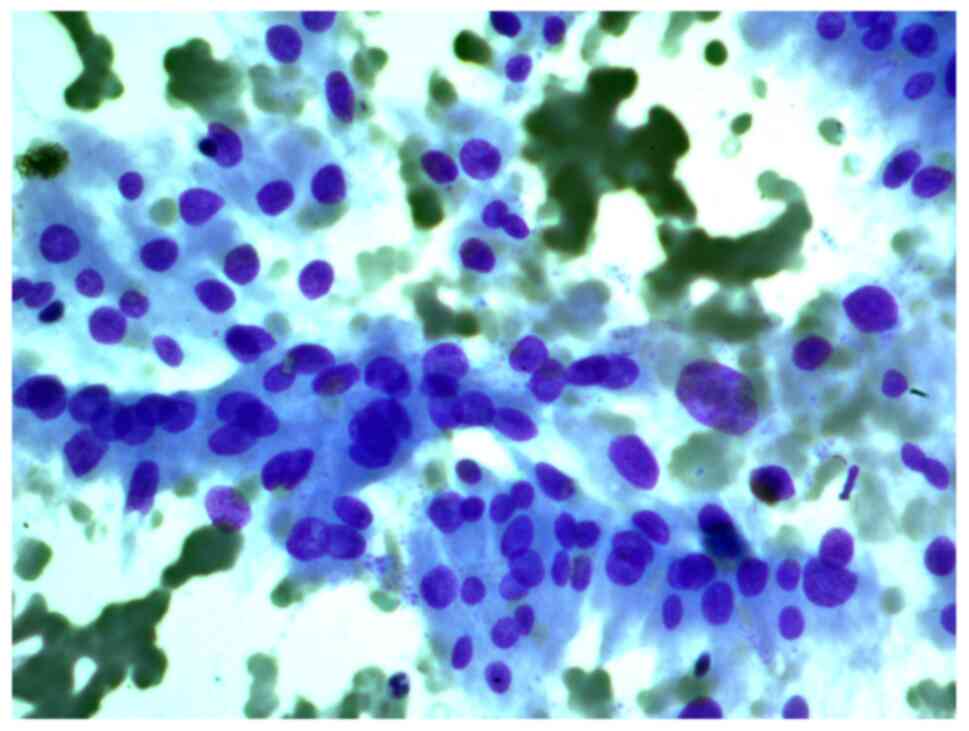
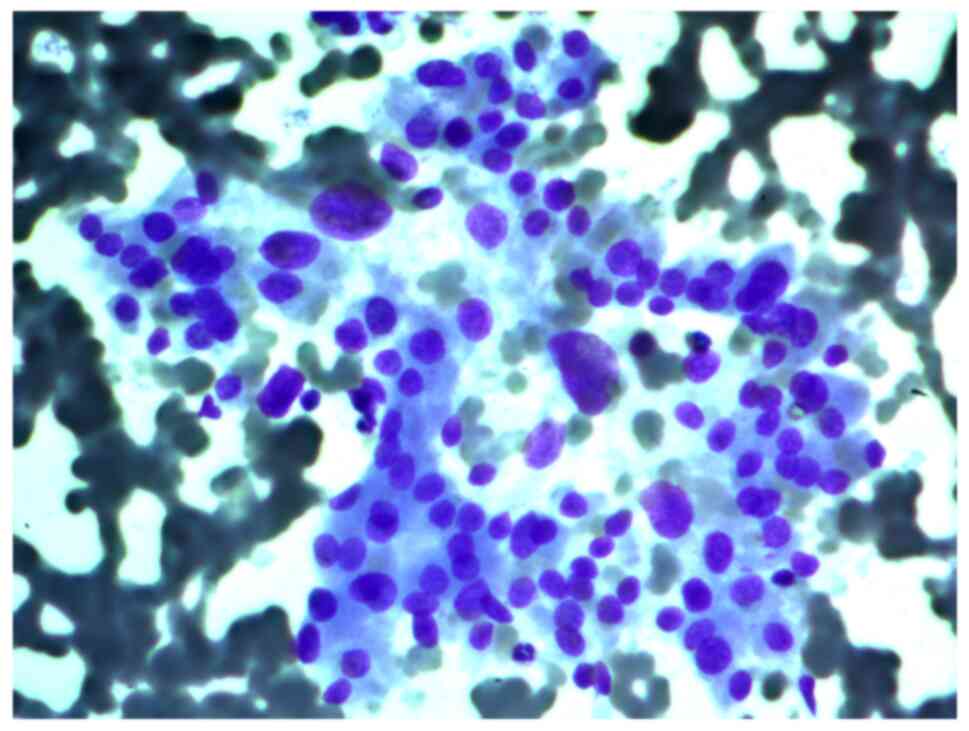
Aspirates of FNAB from a 34-year-old female patient
with a 2 cm nodule in the left thyroid lobe were investigated. The
cytological report stated that this was a Hürthle cell (oncocytic)
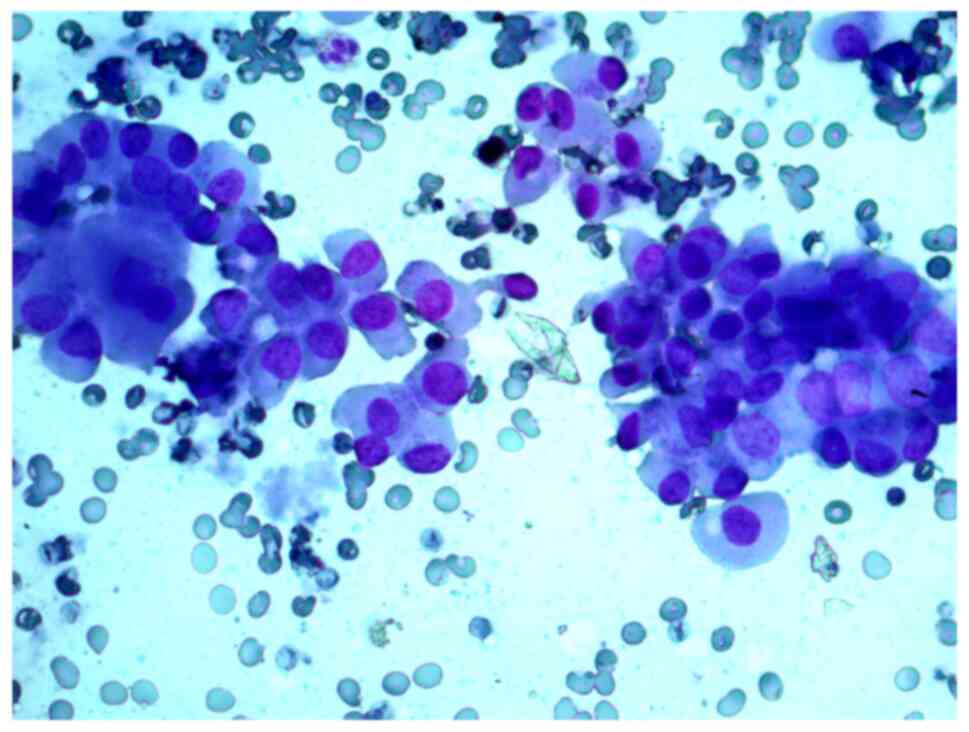
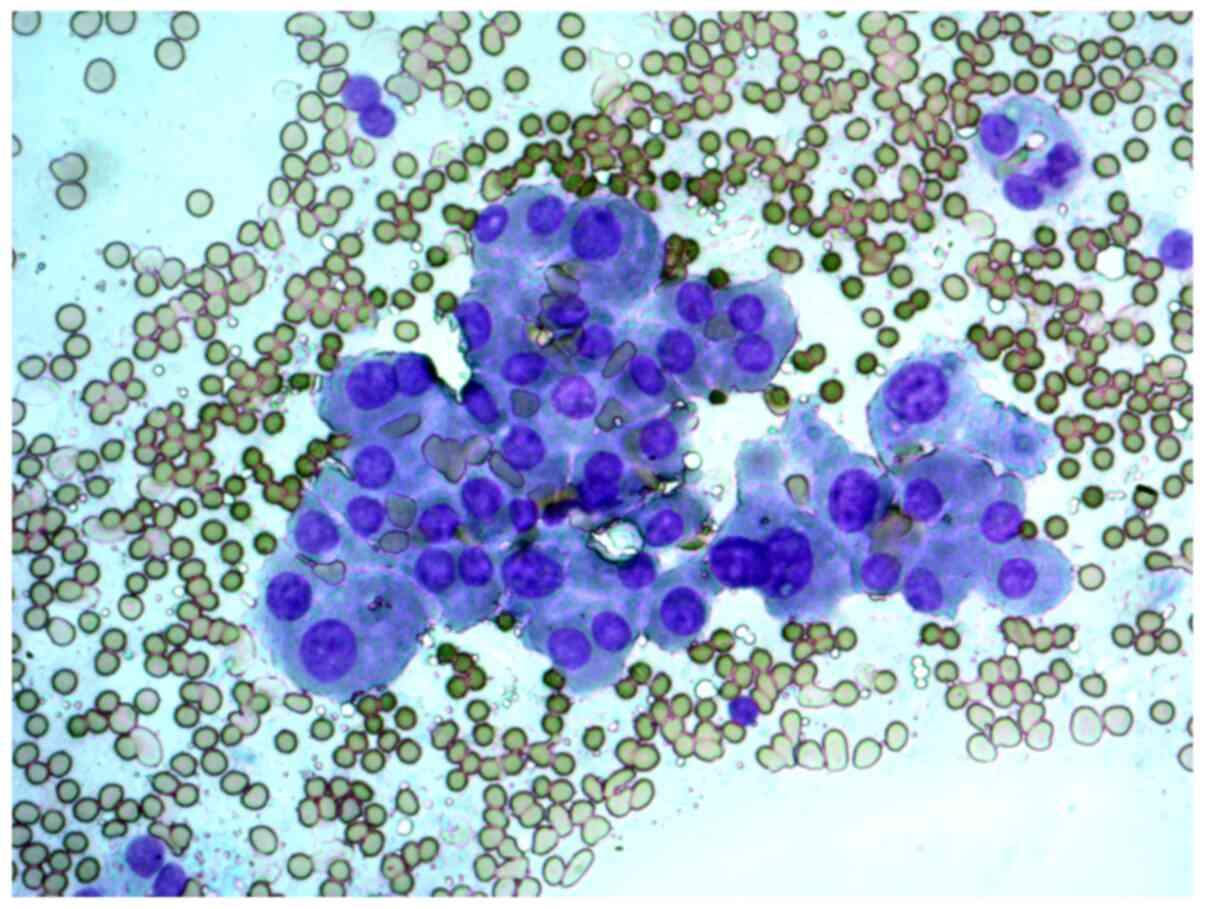
neoplasm, and that in the studied samples, there were groups of
Hürthle cells of various sizes, with subpopulations showing
pronounced polymorphism, abundant fine-grained cytoplasm, an
eccentrically or centrally located nucleus, and enlarged nucleolus
(Figs. 2 and 3).
Three slides were examined, and the results are
given in Table I; our final opinion
was HCC. The histological report on the surgical material indicated
that this was Hürthle cell (oncocytic) thyroid carcinoma, and that
the trabecular structure prevailed in the tumor tissue, with
minimal invasive growth into the capsule. The report also stated
that the Hürthle cells that formed the tumor featured pronounced
cellular and nuclear polymorphism, eosinophilic granular cytoplasm,
and enlarged nucleoli.
 | Table IMolecular analysis of the material on
slides from case 1. |
Table I
Molecular analysis of the material on
slides from case 1.
| Variable | Slide 1 | Slide 2 | Slide 3 | Specimen
classification cut-off values | Cancer
characterization cut-off values |
|---|
| HMGA2 | 0.0139 | 0.0033 | 0.0130 | 0.092 | - |
| miR-146b | -1.48 | -1.69 | -1.03 | 1.53 | 0.17 |
| miR-221 | -1.74 | 1.15a | 1.75a | 0.01 | - |
| miR-375 | -131.56 | -54.00 | -192.44 | -12.12 | 5.25 |
| mtDNA | 10,391a | 19,756a | 30,055a | - | 5,716 |
| Opinion | Benign Hürthle cell
tumor | HCC | HCC | - | - |
Case 2
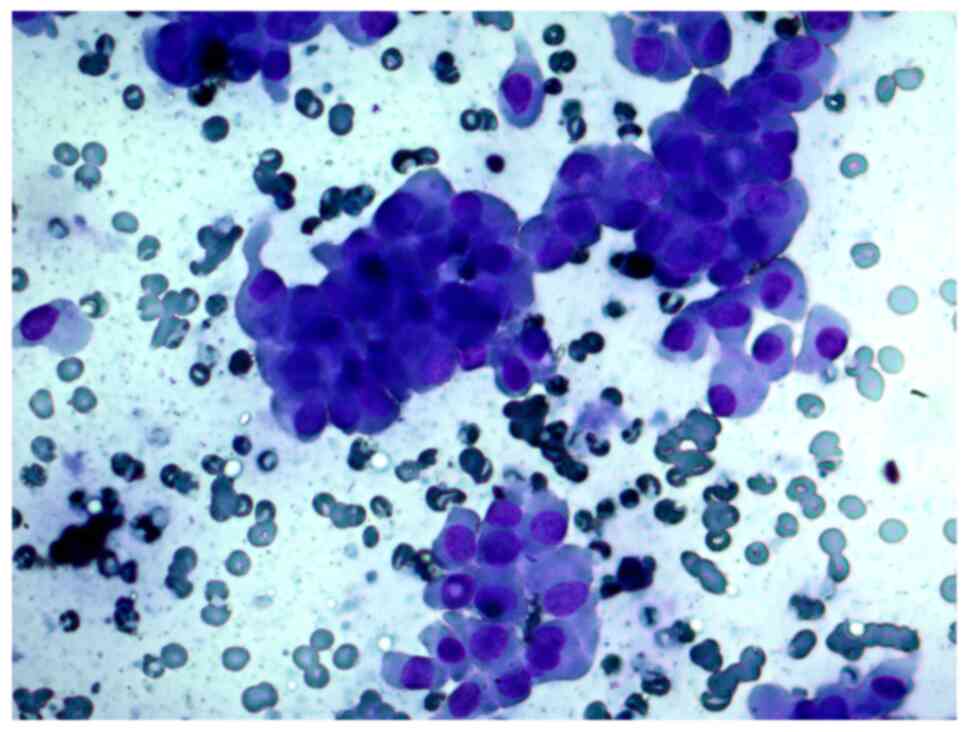
FNAB was performed on a 56-year-old female patient
with a thyroid nodule. The cytological report indicated a suspected
Hürthle cell (oncocytic) neoplasm and that the cellular aspirate
consisted of cells with oncocytic signs, forming sporadic papillary
structures; some cells had features characteristic of papillary
carcinoma: Uneven nuclear contours and nuclear grooves. According
to the cytological report, papillary thyroid carcinoma could not be
ruled out (Figs. 4 and 5).
One slide was examined; the results are listed in
Table II; our conclusion is
Hürthle cell papillary thyroid carcinoma.
 | Table IIMolecular analysis of the material on
a slide from case 2. |
Table II
Molecular analysis of the material on
a slide from case 2.
| Variable | Slide 1 | Specimen
classification cut-off values | Cancer
characterization cut-off values |
|---|
| HMGA2 | 0.0018 | 0.0920 | - |
| miR-146b | 1.61a | 1.53 | 0.17 |
| miR-221 | -7.25 | 0.01 | - |
| miR-375 | -153.75 | -12.12 | 5.25 |
| mtDNA | 8,257a | - | 5,716 |
| Opinion | Hürthle cell
papillary thyroid carcinoma | - | - |
Total thyroidectomy was performed. The histological
report stated that this was papillary thyroid carcinoma, and that
the tumor tissue was predominantly papillar. Tumor cells belonged
to the oncocytic type, with abundant eosinophilic granular
cytoplasm and a moderately polymorphic nucleus with typical signs
of papillary cancer. According to the report, the tumor showed
invasive capsular growth, and there were regional lymph node
metastases.
Case 3
A 29-year-old woman with a solitary thyroid nodule
in the right lobe. The cytological report on the FNAB samples
indicated that this was a suspected Hürthle cell (oncocytic)
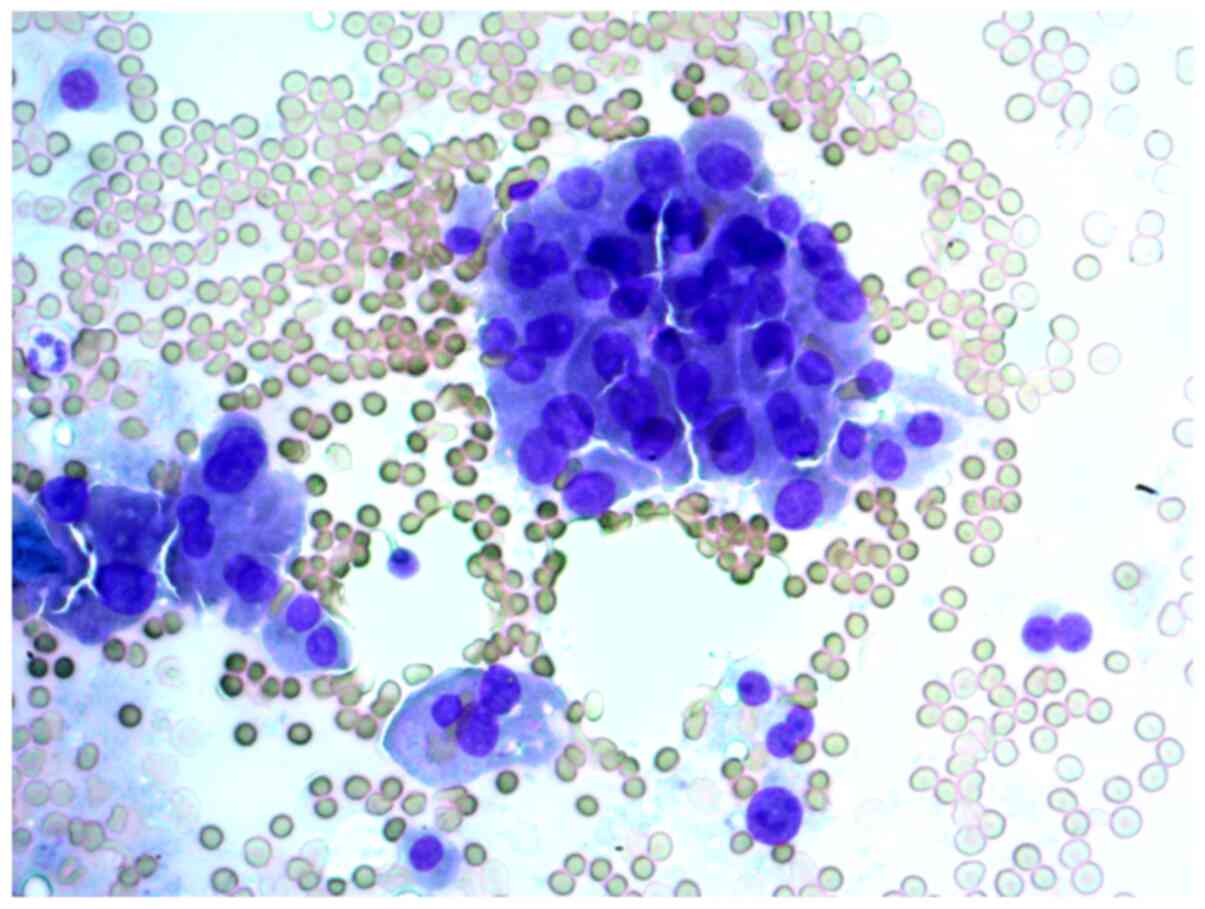
neoplasm, and that in cytological samples, there were
syncytial-like sheets and clusters of Hürthle cells with moderate
cellular and nuclear polymorphism (Figs. 6 and 7).
One slide was examined; the results are presented in
Table III; we concluded that this
is a benign Hürthle cell tumor.
 | Table IIIMolecular analysis of the material on
a slide from case 3. |
Table III
Molecular analysis of the material on
a slide from case 3.
| Variable | Slide 1 | Specimen
classification cut-off values | Cancer
characterization cut-off values |
|---|
| HMGA2 | 0.0025 | 0.0920 | - |
| miR-146b | -10.89 | 1.53 | 0.17 |
| miR-221 | -10.53 | 0.01 | - |
| miR-375 | -157.04 | -12.12 | 5.25 |
| mtDNA | 12,868a | - | 5,716 |
| Opinion | Benign Hürthle cell
tumor | - | - |
Surgical lobectomy was performed. The histological
report stated that this was oncocytic adenoma, i.e., an
encapsulated tumor with a solid and trabecular structure,
consisting of cells of the oncocytic type, with abundant
eosinophilic granular cytoplasm, a large nucleus, and enlarged
nucleolus. According to this report, no capsular or vascular
invasion was found.
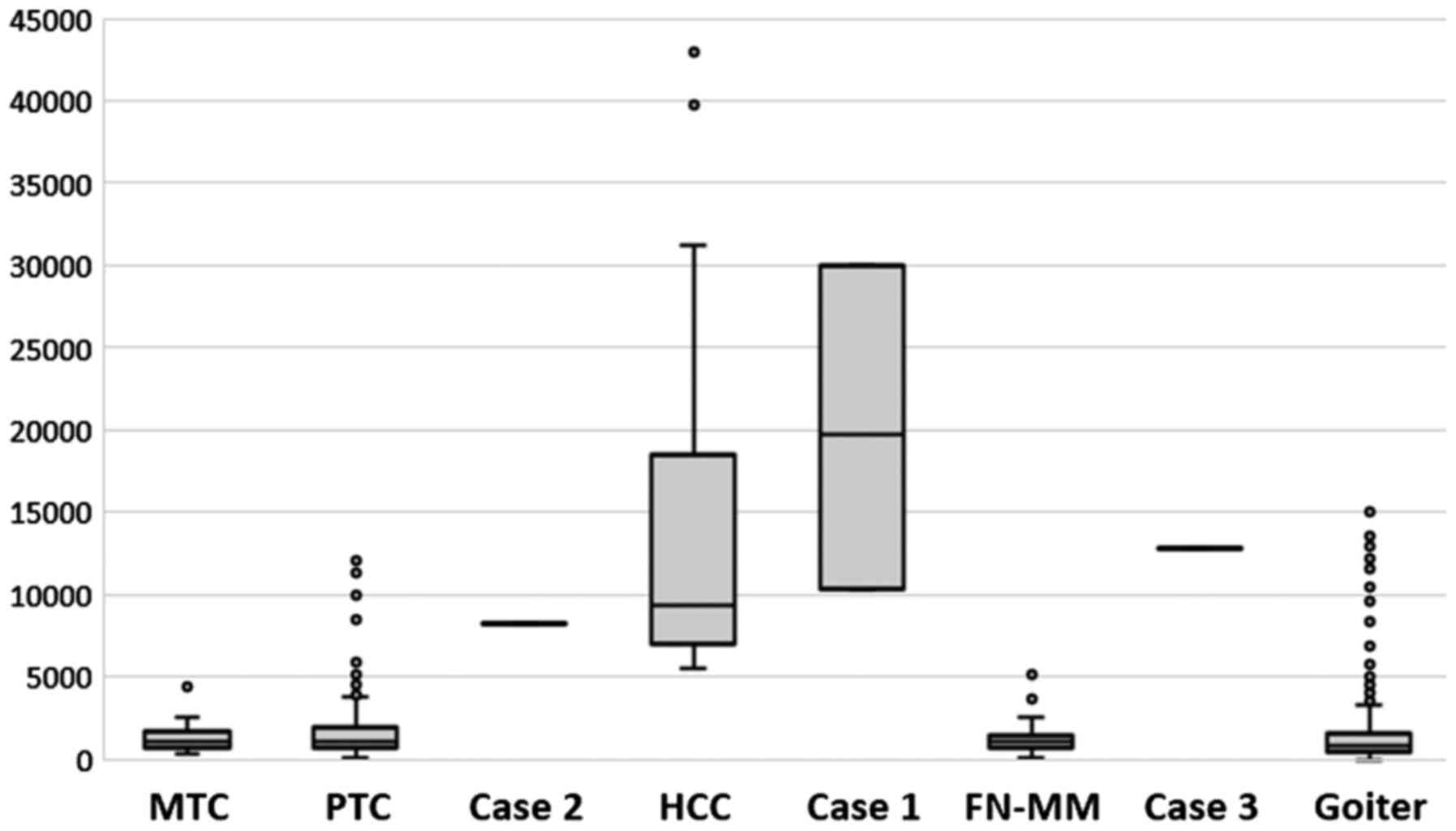
We verified the validity of all the above reports.
In the cases presented, the ratio of mtDNA to nDNA, which is a
marker of Hürthle cells, was within previously described ranges
(13) and helped to make the
diagnoses (Fig. 8).
Discussion
In our recent study, we proposed an original version
of the diagnostic algorithm that identifies and types malignant
thyroid tumors via analysis of molecular markers in cytological
preparations. This algorithm involves the ratio of mtDNA to nDNA as
a criterion for the presence of Hürthle cells because these cells,
according to the literature (18)
and our original data, contain many more mitochondria than
follicular cells do. This is apparently because at least in the
thyroid gland, Hürthle cells exhibit decreased cellular performance
(19). These data suggest a defect
in the energy production machinery of the cells and indicate that
the increased mitochondrial content may be compensatory.
It is now generally accepted that almost all
histological types of benign and malignant thyroid tumors have a
Hürthle cell counterpart (20). In
these tumors, the Hürthle cell appearance is thought to represent a
phenotype that is superimposed on the genotypic and conventional
histopathologic features of the tumors. After the recognition of
the clinicopathologic and genetic differences between PTC and FTC
composed of Hürthle cells, it has been generally accepted that the
Hürthle cell variant of PTC should be referred to as such, whereas
the Hürthle cell variant of FTC may be designated simply as HCC
(21) (the same implicitly follows
from Fig. 7, if we compare the
mtDNA content between FN-MM and HCC). It should be emphasized that
the preoperative detection of FTC is a complex diagnostic problem,
but our proposed algorithm allows this to be done reasonably
accurately at least for the Hürthle cell variant of FTC (i.e.,
HCC).
Therefore, elevated mtDNA content of a tissue sample
is indicative of Hürthle cells but not of malignancy. The hallmark
of HCC is overexpression of miR-221. Although this upregulation
alone is not unique to HCC, its combination with elevated mtDNA
content is. This diagnostic algorithm can be applied to any
cytological slides but primarily to Bethesda categories III and IV.
The limitations of this method are the same as those for
cytological analysis: An insufficient cell number in a sample and
contamination with blood. These limitations may distort the result
or prevent it from being obtained, however, this method is supposed
to be used for samples that have already been analyzed by a
cytologist, so that unsuitable samples will not be tested by our
diagnostic algorithm.
At the moment, there is another diagnostic test that
enables clinicians to identify benign and malignant Hürthle cell
tumors: The Afirma Genomic Sequencing Classifier (GSC), which
involves next-generation RNA sequencing. To identify Hürthle cell
tumors, the GSC system includes two components: Hürthle cell
index-mRNA expression plus mitochondrial transcripts as well as
Hürthle neoplasm index-mRNA expression plus chromosomal level of
loss of heterozygosity (22). Thus,
just as in our algorithm, in the GSC, an estimate of the number of
mitochondria is used to identify Hürthle cells (via determination
of expression levels of mitochondrial genes). In contrast,
malignancy is determined differently, through the assessment of
genetic instability, whereas in our version, this is done via
evaluation of the expression of miR-221, which is a simpler
approach.
The main limitation of this study is the small
number of samples involved (three). This is due to the relative
rarity of Hürthle cell tumors, but this number is planned to
increase over time. As far as the analyzed cases are concerned, the
use of molecular markers in the analysis of cytological material
helped to make a correct diagnosis preoperatively, demonstrating
potential usefulness of this approach for addressing and solving
cytologically unclear or ambivalent cases.
Acknowledgements
Not applicable.
Funding
The present study was supported by the Russian Science
Foundation (grant no. 20-14-00074).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on request.
Authors' contributions
ST and TLP conceived the current study. TLP and VVA
collected patient samples. ST, YAV, and TLP conducted the
experiments. ST and TLP wrote the manuscript. VVA and YAV revised
the manuscript. VVA interpreted the data and gave final approval
for the manuscript to be published. All the authors read and
approved the final manuscript.
Ethics approval and consent to
participate
The Ethical Committee of the Clinical Hospital
RZD-Medicine approved the study protocol. The procedures in the
present study adhere to the tenets of the Declaration of Helsinki.
Written informed consent was obtained from all the participating
patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ali SZ and Cibas ES (eds.): The Bethesda
System for Reporting Thyroid Cytopathology. Springer, New York, NY,
2018.
|
|
2
|
Baloch ZW, LiVolshi VA, Asa SL, Rosai J,
Merino MJ, Randolph G, Vielh P, DeMay RM, Sidawy MK and Frable WJ:
Diagnostic terminology and morphologic criteria for cytologic
diagnosis of thyroid lesions: A synopsis of the national cancer
institute thyroid fine-needle aspiration state of the science
conference. Diagn Cytopathol. 36:425–437. 2008.PubMed/NCBI View
Article : Google Scholar
|
|
3
|
Nguyen GK, Lee MW, Ginsberg J, Wragg T and
Bilodeau D: Fine-needle aspiration of the thyroid: An overview.
Cytojournal. 2(12)2005.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Lloyd RV, Osamura RY, Klöppel G and Rosai
J (eds): WHO Classification of Tumours of Endocrine Organs. IARC,
Lyon, 2017.
|
|
5
|
Montone KT, Baloch ZW and LiVolsi VA: The
thyroid Hürthle (oncocytic) cell and its associated pathologic
conditions: A surgical pathology and cytopathology review. Arch
Pathol Lab Med. 132:1241–1250. 2008.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Nikiforov YE, Biddinger PW, Thompson LDR
and Nikiforova MN (eds): Diagnostic Pathology and Molecular
Genetics of the Thyroid. Lippincott Williams & Wilkins,
Piladelphia, PA, 2009.
|
|
7
|
Lagos-Quintana M, Rauhut R, Lendeckel W
and Tuschl T: Identification of novel genes coding for small
expressed RNAs. Science. 294:853–858. 2001.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Wang J, Chen J and Sen S: MicroRNA as
biomarkers and diagnostics. J Cell Physiol. 231:25–30.
2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Nikiforova MN, Tseng GC, Steward D, Diorio
D and Nikiforov YE: MicroRNA expression profiling of thyroid
tumors: Biological significance and diagnostic utility. J Clin
Endocrinol Metab. 93:1600–1608. 2008.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Cancer Genome Atlas Research Network.
Integrated genomic characterization of papillary thyroid carcinoma.
Cell. 159:676–690. 2014.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Pallante P, Battista S, Pierantoni GM and
Fusco A: Deregulation of microRNA expression in thyroid neoplasias.
Nat Rev Endocrinol. 10:88–101. 2014.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Swierniak M, Wojcicka A, Czetwertynska M,
Stachlewska E, Maciag M, Wiechno W, Gornicka B, Bogdanska M,
Koperski L, de la Chapelle A and Jazdzewski K: In-depth
characterization of the microRNA transcriptome in normal thyroid
and papillary thyroid carcinoma. J Clin Endocrinol Metab.
98:E1401–E1409. 2013.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Titov SE, Ivanov MK, Demenkov PS, Katanyan
GA, Kozorezova ES, Malek AV, Veryaskina YA and Zhimulev IF:
Combined quantitation of HMGA2 mRNA, microRNAs, and
mitochondrial-DNA content enables the identification and typing of
thyroid tumors in fine-needle aspiration smears. BMC Cancer.
19(1010)2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Titov S, Demenkov PS, Lukyanov SA,
Sergiyko SV, Katanyan GA, Veryaskina YA and Ivanov MK: Preoperative
detection of malignancy in fine-needle aspiration cytology (FNAC)
smears with indeterminate cytology (Bethesda III, IV) by a combined
molecular classifier. J Clin Pathol. 73:722–727. 2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee
DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, et al:
Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic
Acids Res. 33(e179)2005.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Titov SE, Ivanov MK, Karpinskaya EV,
Tsivlikova EV, Shevchenko SP, Veryaskina YA, Akhmerova LG, Poloz
TL, Klimova OA, Gulyaeva LF, et al: miRNA profiling, detection of
BRAF V600E mutation and RET-PTC1 translocation in patients from
Novosibirsk oblast (Russia) with different types of thyroid tumors.
BMC Cancer. 16(201)2016.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Cannon J: The Significance of Hürthle
Cells in Thyroid Disease. Oncologist. 16:1380–1387. 2011.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Valenta LJ, Michel-Bechet M, Warshaw JB
and Maloof F: Human thyroid tumors composed of mitochondrion-rich
cells: Electron microscopic and biochemical findings. J Clin
Endocrinol Metab. 39:719–733. 1974.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Sobrinho-Simoes M, Asa SL, Kroll TG,
Nikiforov YE, DeLellis RA, Farid P, Kitamura Y, Noguchi S, Eng C,
Harach HR, et al: Follicular carcinoma. In: World Health
Organization Classification of Tumours. Pathology and Genetics of
Tumours of Endocrine Organs. deLellis RA, Lloyd RV, Heitz PU and
Eng C (eds). IARC Press, Lyon, pp 67-72, 2004.
|
|
21
|
Máximo V, Lima J, Prazeres H, Soares P and
Sobrinho-Simões M: The biology and the genetics of Hurthle cell
tumors of the thyroid. Endocr Relat Cancer. 19:R131–R147.
2012.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Patel KN, Angell TE, Babiarz J, Barth NM,
Blevins T, Duh QY, Ghossein RA, Harrell RM, Huang J, Kennedy GC, et
al: Performance of a genomic sequencing classifier for the
preoperative diagnosis of cytologically indeterminate thyroid
nodules. JAMA Surg. 153:817–824. 2018.PubMed/NCBI View Article : Google Scholar
|