Introduction
Esophageal cancer has malignant potential and often
results in early recurrence and a poor prognosis (1). Squamous cell carcinoma antigen
(SCC-Ag), cytokeratin 19 fragment (CYFRA), and carcinoembryonic
antigen (CEA) have been employed as biomarkers for esophageal
cancer. MicroRNA and serological markers have been reported as new
biomarkers, but the results have been unsatisfactory (2,3). The
serum anti-p53 antibody test has recently been developed and has
been employed worldwide to detect superficial esophageal SCC
(4,5). The positive rate of serum anti-p53
was 30% (6); however, there are
few biomarkers that have shown such high sensitivity and
specificity (7-10).
New biomarkers that can estimate the short-term postoperative
prognosis are effective because they can help physicians select the
appropriate postoperative adjuvant therapy during the early
postoperative period.
Serological identification of antigens by
recombinant cDNA expression cloning (SEREX) is an effective
screening method for tumor markers (11). SEREX involves the immune-screening
of cDNA libraries prepared from tumor specimens from patients'
sera. The sequencing of isolated cDNA clones makes SEREX suitable
for the large-scale screening of tumor antigens. SEREX has been
applied to various human tumor types and has identified more than
1000 novel tumor antigens (SEREX antigens) (12).
We conducted a large-scale SEREX screening using
sera from patients with esophageal cancer and identified
TROP2/TACSTD2(13),
SLC2A1/GLUT1(14), tripartite
motif-containing 21(15),
myomegalin (16), makorin
1(17) and esophageal carcinoma
SEREX antigen (ECSA) (18). In the
present study, we further identified striatin 4 (STRN4) as a novel
esophageal SEREX antigen and evaluated clinicopathological
significance of serum anti-STRN4 antibody (s-STRN4-Ab) levels in
patients with solid tumors.
Materials and methods
Collection of serum samples
The study was approved by the Ethics Committee of
Toho University, Graduate School of Medicine (no. A18103) and Chiba
University Graduate School of Medicine (no. 2018-320) (Japan). We
collected sera from patients who had provided written informed
consent.
Serum samples were obtained from 672 patients,
including 192 with esophageal cancer, 96 with gastric cancer, 192
with colorectal cancer, 96 with lung cancer, and 96 with breast
cancer from Toho University Omori Hospital (Japan) between June
2010 and February 2016. Among the 192 patients with esophageal
cancer, 91 underwent radical surgery. Among them, 63 patients
underwent neoadjuvant chemotherapy. The number of patients in each
stage for esophageal cancer (Japanese Classification of Esophageal
Cancer, 11th Edition (19) was as
follows: 9 patients in stage 0, 14 in stage I, 25 in stage II, 34
in stage III, and 9 in stage IVa. Gastric cancer was analyzed in 57
cases and colorectal cancer was analyzed in 113 cases, of which all
cases were underwent radical surgery. Excluded cases were those
with double cancer and pathologically difficult staging due to
neoadjuvant chemotherapy. The number of patients in each stage for
gastric cancer (Japanese classification of gastric carcinoma: 3rd
English edition (20) was as
follows: 28 patients in stage I, 14 in stage II, 8 in stage III,
and 7 in stage IV. And the number of patients for colorectal cancer
(Japanese Classification of Colorectal, Appendiceal, and Anal
Carcinoma: the 3d English Edition [Secondary Publication] (21) was as follows: 5 patients in stage
0, 29 in stage I, 32 in stage II, 31 in stage III, and 16 in stage
IV.
All patients were regularly followed-up until July
2018 or death. Healthy donor sera were obtained from Port Square
Kashiwado Clinic.
SEREX screening
SEREX screening was performed using sera from
patients with esophageal cancer as previously described (13-18).
The isolated STRN4 cDNA was recombined into the
EcoRI/XhoI site of pGEX-4T-3 (GE Healthcare Life
Sciences, Pittsburgh, PA). Expression of STRN4-GST or control GST
was induced by treating pGEX-4T-3-STRN4-transformed or control
pGEX-4T-3-transformed Escherichia coli (E. coli) BL21
with 0.1-mM isopropyl-β-D-thiogalactoside (IPTG) at 37˚C for 3 h.
The cells were subsequently lysed in BugBuster Master Mix (Merck
Millipore, Darmstadt, Germany). STRN4-GST and GST proteins were
purified by glutathione-Sepharose (GE Healthcare Life Sciences)
column chromatography according to the manufacturer's instructions
as previously described (22-26).
Serum sampling and AlphaLISA for serum
markers
Serum samples were collected prior to treatment and
were kept frozen at -80˚C until use. s-STRN4-Ab levels were
examined using amplified luminescence proximity homogeneous
assay-linked immunosorbent assay (AlphaLISA), which was performed
using 384-well microtiter plates (white opaque OptiPlate™; Perkin
Elmer) containing 2.5 µl of 1/100-diluted sera and 2.5 µl of GST or
GST-fusion STRN4 protein (10 µg/ml) in AlphaLISA buffer (25 mM
HEPES, pH 7.4, 0.1% casein, 0.5% Triton X-100, 1-mg/ml dextran-500,
and 0.05% Proclin-300) according to the manufacturer's instructions
(Perkin Elmer, http://www.perkinelmer.com/lab-solutions/resources/docs/GDE_ELISA-to-AlphaLISA.pdf).
The reaction mixture was incubated at room temperature for 6-8 h.
Next, anti-human IgG-conjugated acceptor beads (2.5 µl of 40 µg/ml)
and glutathione-conjugated donor beads (2.5 µl of 40 µg/ml) were
added and incubated further for 14 days at room temperature in the
dark. The chemical emission was measured by an EnSpire Alpha
microplate reader (PerkinElmer) as previously described (27-29).
Specific reactions were calculated by subtracting the Alpha values
of the GST control from the GST-STRN4 values. The serum p-53
antibodies (p53-Abs) (30) and
SCC-Ag (31) were also evaluated
as previously described. Cutoff values for serum p53-Abs and SCC-Ag
were fixed at 1.3 IU/ml and 1.5 ng/ml, respectively.
Statistical analysis
The continuous data are expressed as mean ± standard
deviation. Receiver operating characteristic curve analysis was a
typical method to determine the sensitivity and specificity from
the serum STRN4-Ab (s-STRN4-Ab) level between healthy donor and
cancer patients to set at the values that maximize the sums of the
sensitivity and specificity. Area under the curve (AUC) was defined
by the area of under the curve of the ROC curve. The closer the AUC
is to 1, it is judged that the discrimination performance is high.
Comparisons between unpaired groups were conducted using the
Mann-Whitney U test. Differences in the distribution of two
variables were evaluated using Fisher's exact test or the
chi-squared test. The corresponding differences among three
variables were evaluated with the Kruskal-Wallis test. We analyzed
the clinicopathological data using logistic regression analysis to
evaluate the association with s-STRN4-Ab level. We calculated the
survival curves using the Kaplan-Meier method and compared them
using the log-rank test. All analyses were performed using EZR
software (32), and statistical
significance levels were defined as P<0.05.
Results
Comparison of s-STRN4-Ab levels
between healthy donors and patients with solid tumors
SEREX screening identified striatin 4 (STRN4;
accession number: NM_013403) as an antigen recognized by serum IgG
antibodies in patients with esophageal cancer. We then recombined
the cDNA, purified the STRN4-GST protein and examined the
s-STRN4-Ab levels in patients with esophageal cancer, gastric
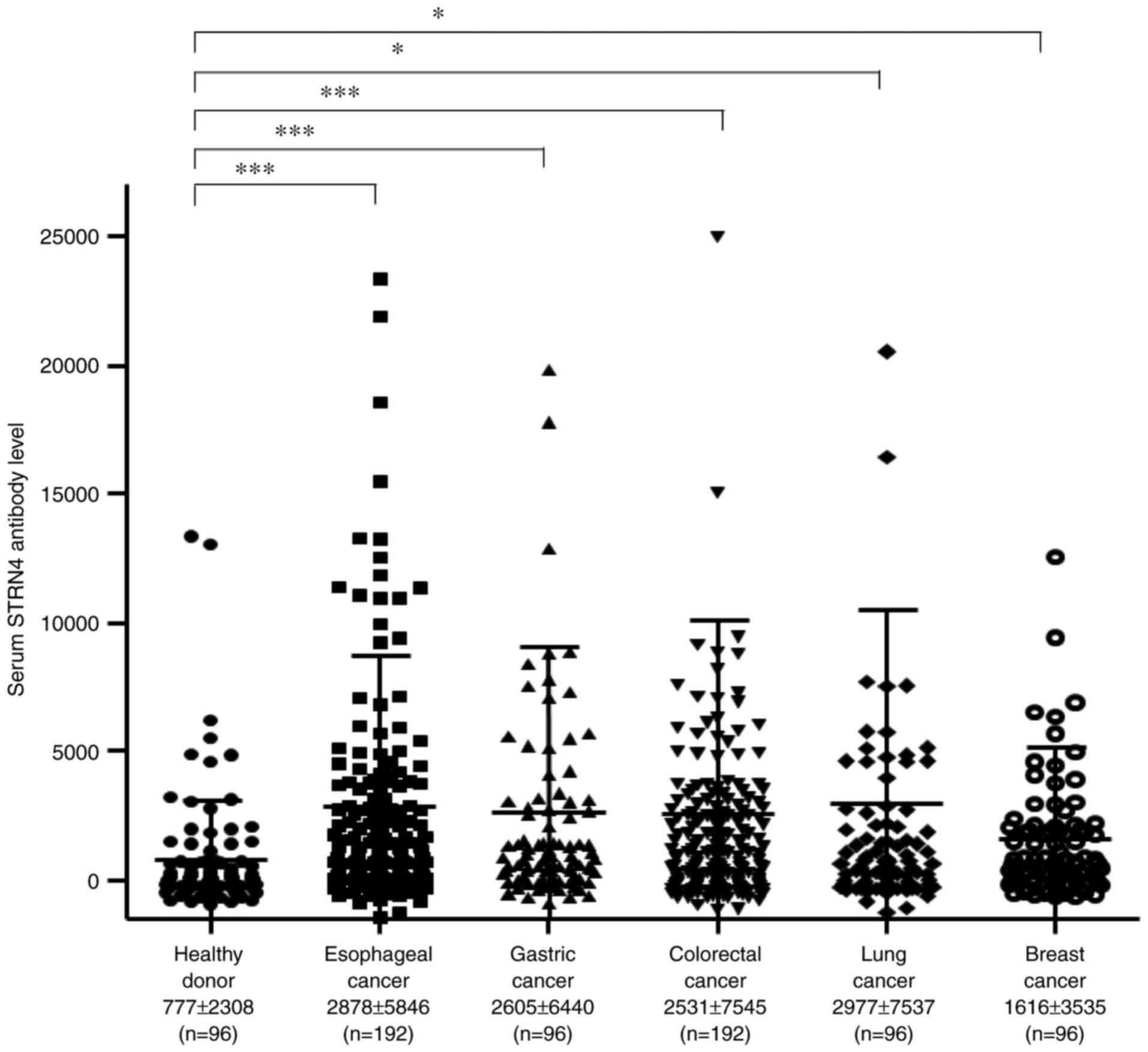
cancer, colorectal cancer, lung cancer, and breast cancer. The mean
s-STRN4-Ab levels (± standard deviation) were as follows: healthy
donors, 777±2308; esophageal cancer, 2878±5846; gastric cancer,
2605±6440; colorectal cancer, 2531±7545; lung cancer, 2977±7537;
and breast cancer, 1616±3535. Compared with the healthy donors, the
patients with any type of cancer showed significantly higher levels
of s-STRN4-Abs in the Kruskal-Wallis test (Fig. 1), suggesting that s-STRN4-Ab is a
common marker for solid cancers. The levels were especially higher
for esophageal cancer and lung cancer but relatively lower for
breast cancer.
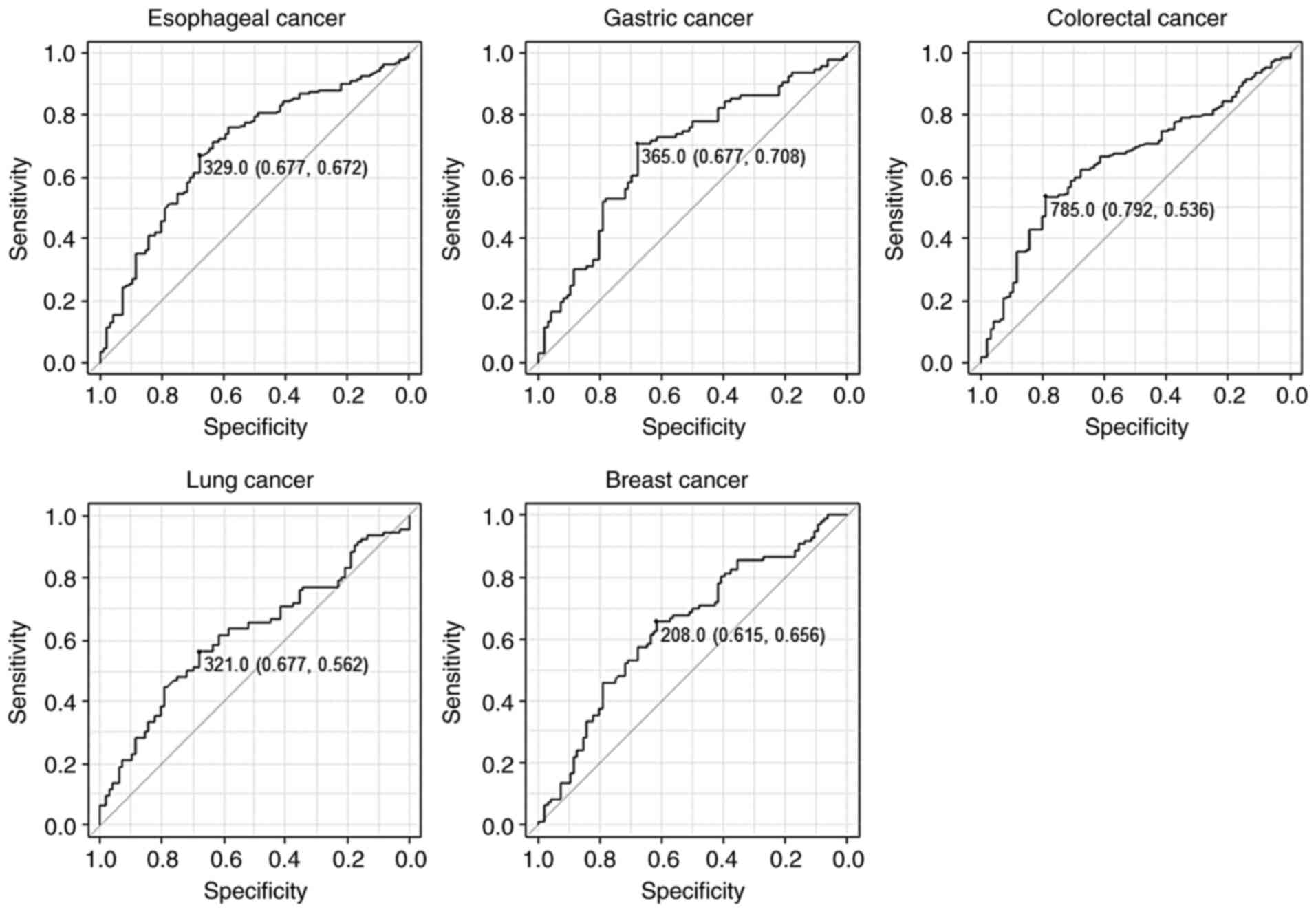
We then performed a receiver operating
characteristic curve analysis to evaluate the sensitivity and
specificity between each cancer and the healthy donors (Fig. 2). The results revealed that the AUC
values were >0.6 for all cancers (Table I). When the cutoff value was set at
the values that maximize the sums of the sensitivity and
specificity, the sensitivity and specificity of s-STRN4-Abs for
esophageal cancer were 67.2 and 67.7%, respectively.
 | Table IComparison of AUC, 95% confidence
intervals, cutoff levels, sensitivity, specificity and P-values
from patients with various cancer types. |
Table I
Comparison of AUC, 95% confidence
intervals, cutoff levels, sensitivity, specificity and P-values
from patients with various cancer types.
| Variable | Esophageal
cancer | Gastric cancer | Colorectal
cancer | Lung cancer | Breast cancer |
|---|
| AUC | 0.694 | 0.682 | 0.653 | 0.616 | 0.639 |
| 95% CI | 0.630-0.758 | 0.606-0.758 | 0.587-0.718 | 0.536-0.696 | 0.560-0.718 |
| Cutoff | >329.0 | >365.0 | >785.0 | >321.0 | >208.0 |
| Sensitivity
(%) | 67.2 | 70.8 | 53.6 | 56.2 | 65.6 |
| Specificity
(%) | 67.7 | 67.7 | 79.2 | 67.7 | 61.5 |
| P-value | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 |
Comparison of s-STRN4-Ab levels
according to various laboratory data
Given that the AUC was the highest for esophageal
cancer among the solid cancers (Table
I), we focused on 91 surgical cases of esophageal cancer and
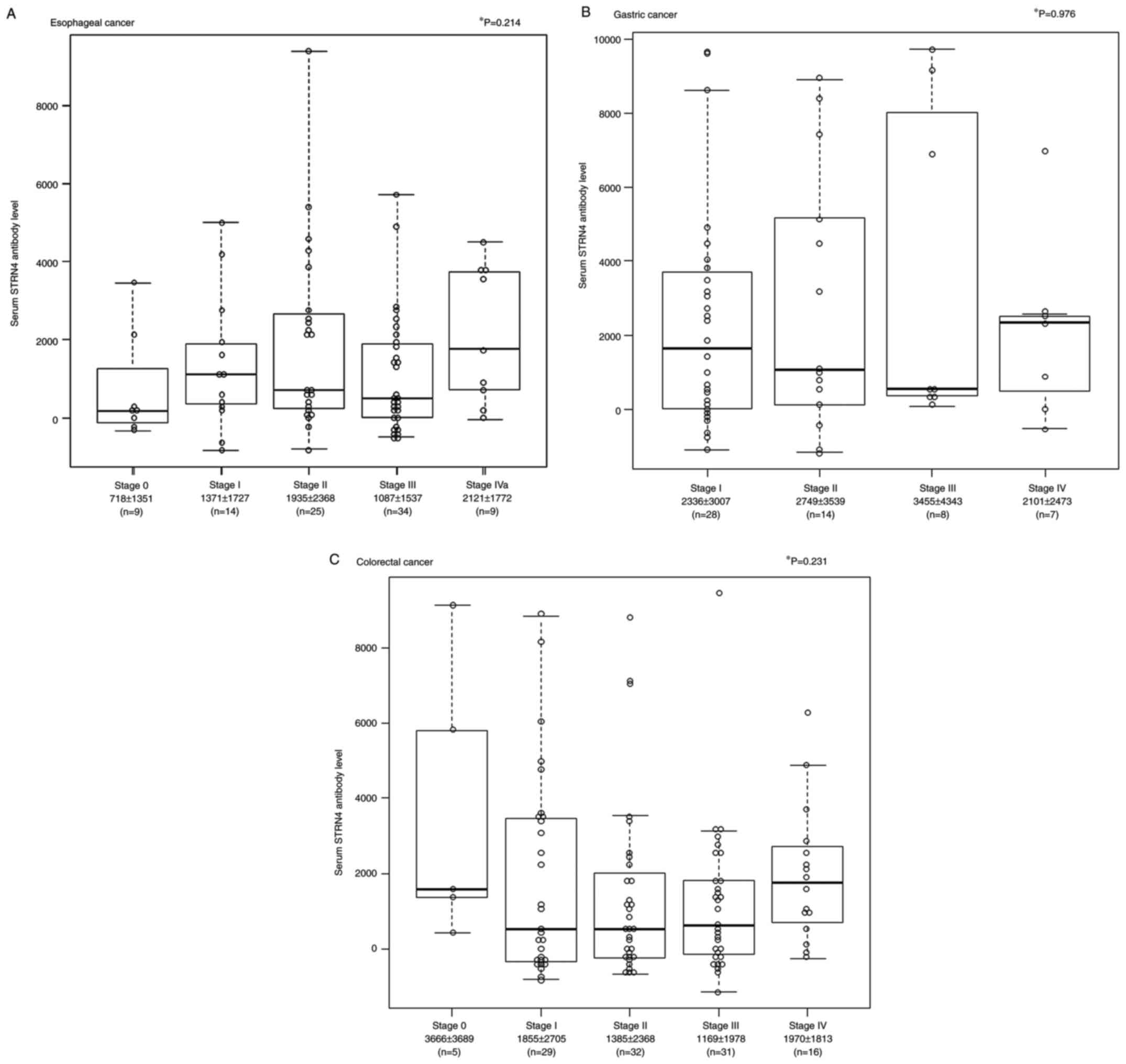
examined their clinicopathological features. The mean (± standard
deviation (SD)) s-STRN4-Ab levels for stage 0, stage I, stage II,
stage III, and stage IVa were 718±1351, 1371±1727, 1935±2368,
1087±1537 and 2121±1772, respectively (Fig. 3A). In the gastric cancer, the mean
± SD of s-STRN4-Ab levels for stage I, stage II, stage III, and
stage IV were 2336±3007, 2749±3539, 3455±4343, and 2101±2473,
respectively (Fig. 3B). In the
colorectal cancer, the mean ± SD s-STRN4-Ab levels for stage 0,
stage I, stage II, stage III, and stage IV were 3666±3689,
1855±2705, 1385±2368, 1169±1978, and 1970±1813, respectively
(Fig. 3C). Although there was no
significant association between each stage and a s-STRN4 antibody
level in the Kruskal-Wallis test in all three cancers, the mean
STRN4 levels for stages I, II, III, and IV were higher than for
stage 0 in terms of esophageal cancer.
Relationship between s-STRN4-Ab levels
and overall survival
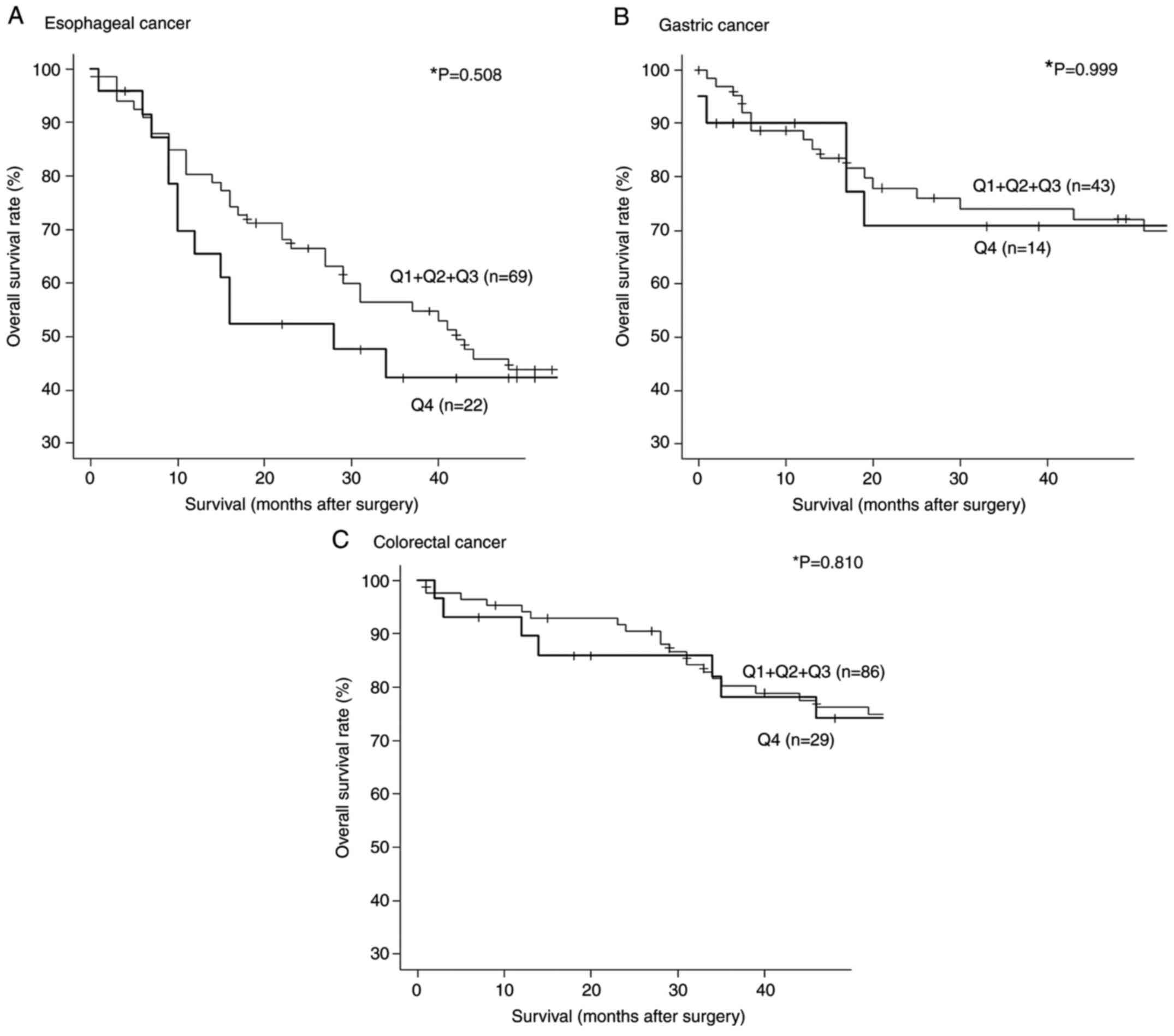
We then divided the s-STRN4-Ab levels for esophageal
cancer into quartiles (Q1, Q2, Q3 and Q4). The s-STRN4-Ab levels
ranged from -827 to 184 for Q1, 190 to 873 for Q2, 1106 to 2746 for
Q3 and 2876 to 39727 for Q4. Although there were no significant
differences in overall survival between the Q1+Q2+Q3 and Q4 groups
(P=0.508), Q4 group showed poor prognosis at early stage (10-36
weeks) after surgery. (Fig. 4A).
In terms of gastric and colorectal cancer, there was no
statistically significant correlation between Q1+Q2+Q3 vs. Q4
groups (Fig. 4B and C).
Relationship between high serum STRN4
antibody levels and clinicopathological factors
The univariate and multivariate analysis showed no
significant association for esophageal cancer between s-STRN4-Ab
levels and sex, age, tumor depth, lymph node status, location,
white blood cell count, neutrophil count, lymphocyte count,
hemoglobin count, platelet count, C-reactive protein level, albumin
level, SCC-Ag level, and p53-Abs (Tables II and III). In terms of gastric and colorectal
cancer, there was no significant association for
clinicopathological factors (data not shown).
 | Table IIFisher's exact comparison of serum
levels according to the clinicopathological characteristics of
patients with esophageal cancer. |
Table II
Fisher's exact comparison of serum
levels according to the clinicopathological characteristics of
patients with esophageal cancer.
| Variables | STRN4 Q1+Q2+Q3 | STRN4 Q4 | P-value |
|---|
| Sex | | | >0.999 |
|
Male | 51 | 19 | |
|
Female | 16 | 5 | |
| Age | | | 0.810 |
|
>65 | 38 | 15 | |
|
≤65 | 29 | 9 | |
| Location | | | 0.753 |
|
Upper | 11 | 3 | |
|
Lower | 56 | 21 | |
| Tumor depth | | | 0.454 |
|
T1 | 23 | 6 | |
|
T2-T4 | 44 | 18 | |
| Lymph node
status | | | 0.636 |
|
N0 | 29 | 12 | |
|
N1 | 38 | 12 | |
| WBC (µl) | | | >0.999 |
|
>8,000 | 8 | 3 | |
|
≤8,000 | 59 | 21 | |
| Neutrophil (%) | 15 | 4 | 0.771 |
|
>70 | 15 | 4 | |
|
≤70 | 52 | 20 | |
| Lymphocyte (%) | | | 0.371 |
|
>35 | 11 | 6 | |
|
≤35 | 56 | 18 | |
| Hemoglobin
(g/dl) | | | 0.450 |
|
>12 | 43 | 18 | |
|
≤12 | 24 | 6 | |
| Platelet | | | 0.375 |
|
>150,000 | 63 | 21 | |
|
≤150,000 | 4 | 3 | |
| CRP
(mg/dl)a | | | 0.450 |
|
>0.3 | 20 | 9 | |
|
≤0.3 | 46 | 14 | |
| Albumin (g/dl) | | | 0.596 |
|
>3.5 | 51 | 17 | |
|
≤3.5 | 16 | 7 | |
| SCC-Ag
(ng/ml)a | | | 0.447 |
|
>1.5 | 21 | 10 | |
|
≤1.5 | 44 | 13 | |
| p53-Abs
(U/ml)a | | | 0.770 |
|
>1.30 | 12 | 5 | |
|
≤1.30 | 53 | 19 | |
 | Table IIILogistic regression analysis of serum
levels according to the clinicopathological characteristics of
patients with esophageal cancer. |
Table III
Logistic regression analysis of serum
levels according to the clinicopathological characteristics of
patients with esophageal cancer.
| Variable | Odds ratio | 95% CI | P-value |
|---|
| Tumor depth | 1.890 | 0.545-6.570 | 0.315 |
| Lymphocyte (%) | 2.260 | 0.643-7.920 | 0.204 |
| Hemoglobin
(g/dl) | 0.417 | 0.130-1.340 | 0.141 |
| Platelet | 2.240 | 0.436-11.50 | 0.333 |
| SCC-Aga | 1.540 | 0.522-4.520 | 0.436 |
Combination assessment of s-STRN4-Ab
and conventional tumor markers
To evaluate the usefulness of s-STRN4-Ab for
esophageal cancer, we compared it with the conventional tumor
markers, SCC-Ag and p53-Abs. When we decided the s-STRN4-Ab cutoff
level at 2746 (the border between Q3 and Q4), the positive rate was
26.4%, which was comparable to 19.1% for p53-Ab and 35.2% for
SCC-Ag (Fig. 5). We then examined
the positive rates of s-STRN4-Ab combined with the conventional
tumor markers. The positive rates for SCC-Ag plus s-STRN4-Ab,
p53-Ab plus s-STRN4-Ab, and SCC-Ag plus p53-Abs were 49.4, 40.4,
and 48.3%, respectively. By combining all of the markers, the
positivity increased to 59.1%. Thus, addition of s-STRN4-Ab marker
improved the sensitivity.
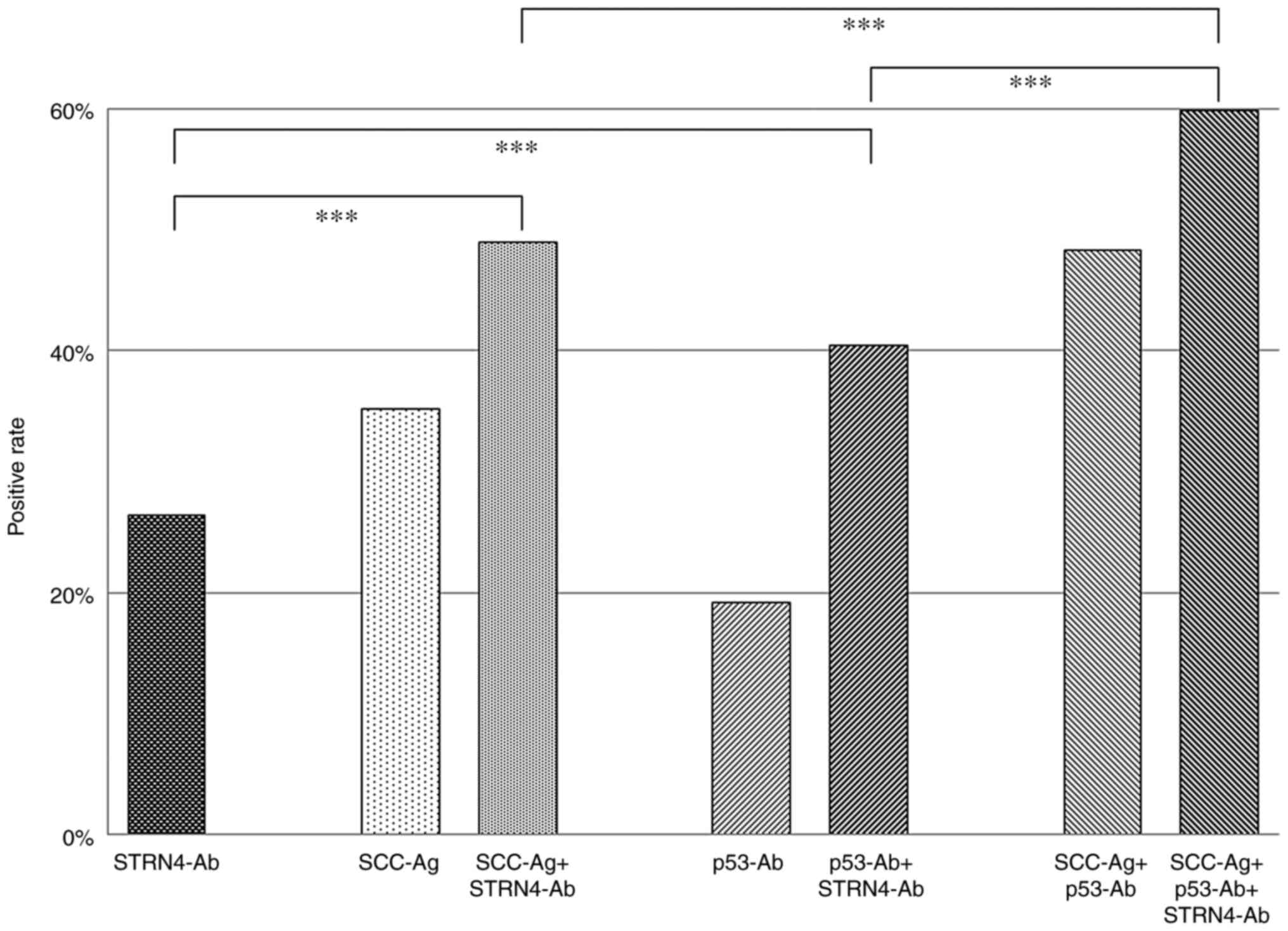 | Figure 5Positive rates of each or combined
tumor markers for patients with esophageal cancer. The positive
percentages of STRN4-Ab, SCC-Ag, SCC-Ag + STRN4-Ab, p53-Ab, p53-Ab
+ STRN4-Ab, SCC-Ag + p53-Ab, and SCC-Ag + p53-Ab + STRN4-Ab are
presented. The cutoff level of STRN4-Ab was set to 2,746 (Q3 upper
limit; 2,746) irrespective of the levels of SCC-Ag and p53-Ab.
P-values were calculated with Fisher's exact test.
***P<0.001. The positive percentages have no
variation. Q represents quartiles of serum STRN4 antibody levels.
Q1 levels ranged from -827 to 184, 190 to 873 for Q2, 1,106 to
2,746 for Q3 and 2,876 to 39,727 for Q4. STRN4, striatin 4; Ab,
antibody; SCC-Ag, squamous cell carcinoma antigen. |
Discussion
We found that s-STRN4-Ab levels were significantly
higher in the solid cancer group (esophageal cancer, gastric
cancer, colorectal cancer, lung cancer, and breast cancer) than the
healthy donor group. P-values of digestive organ cancers vs. HD
were much lower (<0.001) than that of lung cancer (<0.05).
Given that the AUC value for esophageal cancer was highest among
all examined AUC values, we focused on esophageal cancer and
evaluated its clinicopathological character and prognosis; however,
there were no statistically significant correlation between
s-STRN4-Ab and clinicopathological character including TNM. In the
stage-by-stage analysis, however, s-STRN4-Ab levels at stages II,
III and IV were higher than at stage 0. There was no correlation
between STRN4 and other tumor markers in the univariate and
multivariate analysis, indicating that the positive rate increased
by the combined diagnosis of these markers.
STRN, STRN3 and STRN4 are members of the striatin
family of proteins that contain multiple protein-binding domains
including the coiled-coil domain,
Ca2+-calmodulin-binding domain, caveolin-binding domain,
and WD-repeat domain (33-35).
Striatin family proteins are also known to form a complex with
protein phosphatases and protein kinases, which further regulates
the signaling pathways involved in cell proliferation,
differentiation, apoptosis, and transformation (36,37).
These results indicate that STRN4 is involved in the early stage of
carcinogenesis. It has also been reported that silencing of STRN4
suppresses the proliferation, migration, invasion, and
anchorage-independent growth of several cancer cells (38). Wang et al suggested that
miR-6165 inhibits the migration and invasion of gastric cancer
cells by targeting STRN4(39).
Jiang et al indicated that Pokemon, through stimulation of
STRN4 expression, promotes prostate tumor progression via a
Pokemon/STRN4 axis (40). Wong
et al reported that STRN4 was highly expressed in
digestive-organ cancer cell lines and a lung cancer cell line
(38). They also showed that STRN4
knockdown suppressed proliferation and metastasis of these cell
lines. Consequently, STRN4 can facilitate carcinogenesis from the
early stage to the advanced stage, which is consistent with our
results that the s-STRN4-Ab marker was not correlated with the
stage or TNM classification.
High s-STRN4-Ab levels indicated a poor
postoperative prognosis for esophageal cancer, with similar results
reported for liver cancer and ovarian cancer (41). High expression of STRN4 protein in
these cancers indicates a poor prognosis especially in the early
phase. For example, at 20 months after surgery, the survival rates
for the Q1+Q2+Q3 and Q4 groups were 68 and 50%, respectively. The
50% survival periods for the Q1+Q2+Q3 and Q4 groups were 42 and 28
months, respectively. Thus, s-STRN4 levels might reflect a poor
postoperative prognosis. Determining the risk of early
postoperative death through the use of STRN4 antigen and antibody
markers can be useful, because early treatment such as adjuvant
therapy can improve the prognosis.
Antibody markers have differing characteristics from
those of antigen markers. In the early stages of cancer, low-level
tissue destruction and subsequent leakage of intracellular
antigenic proteins lead to the emergence of autoantigens. Further
repeated tissue destruction and leakage result in a remarkable
increase in antibody levels, with antigen levels remaining
constant. In the early stages, the sensitivity of the antibody
markers is therefore much higher than that of antigen markers. In
the advanced stages, tissue destruction becomes evident, and a
large amount of antigen leaks out, which can result in partial
absorption of the autoantibodies (29). Thus, the antigen markers might be
suitable for diagnosing the late stages of cancer.
Although we have identified s-STRN4-Ab as a marker
for digestive solid cancer, this is insufficient for achieving high
sensitivity for diagnosing the early stage of cancers. Therefore we
need to find additional useful biomarkers to improve the prognosis.
s-STRN4-Ab levels were higher in patients with solid cancers than
in healthy donors. The STRN4-Ab marker might reflect the poor
postoperative prognosis of esophageal cancer.
Acknowledgements
The authors would like to thank Ms. Seiko Otsuka,
Ms. Chiho Kusaka and Ms. Satoko Ishibashi (Department of
Gastroenterological Surgery, Toho University School of Medicine)
for preparing patient data.
Funding
This research was supported by the Japan Agency for Medical
Research and Development, AMED and supported by a Grant-in-Aid for
Scientific Research from the Ministry of Education, Culture,
Sports, Science and Technology of Japan (grant nos. 16K10519,
16K10520 and 21K08695).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
MI, TH and HS conceived the project and designed the
experiments. YO, SY, TS, TN, MS, FS, KF, HT and KK performed
acquired the data. YO, SY and NT collected clinicopathological
information. MI, TH, MS and FS performed the experiments. MI and TH
wrote the manuscript. All authors revised the manuscript. All
authors have read and approved the manuscript. MI and TH confirm
the authenticity of all the raw data.
Ethics approval and consent to
participate
The present study was approved by the Institutional
Ethics Committee of Toho University Graduate School of Medicine
(Tokyo, Japan; approval no. A18103) as well as the review board of
Chiba University Hospital. Serum was collected from patients who
had provided written informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Shimada H: Revisiting radiation therapy
for esophageal cancer. Esophagus. 17(99)2020.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Yang B, Liu Y, Li L, Deng H and Xian L:
MicroRNA-200a promotes esophageal squamous cell carcinoma cell
proliferation, migration and invasion through extensive target
genes. Mol Med Rep. 21:2073–2084. 2020.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Fernandes E, Sores J, Cotton S, Peixoto A,
Ferreira D, Freitas R, Reis CA, Santos LL and Ferreira JA:
Esophageal, gastric and colorectal cancers: Looking beyond
classical serological biomarkers towards glycoproteomics-assisted
precision oncology. Theranostics. 31:4903–4928. 2020.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Shimada H, Takeda A, Arima M, Okazumi S,
Matsubara H, Nabeya Y, Funami Y, Hayashi H, Gunji Y, Suzuki T, et
al: Serum p53 antibody is a useful tumor marker in superficial
esophageal squamous cell carcinoma. Cancer. 89:1677–1683.
2000.PubMed/NCBI
|
|
5
|
Kochi R, Yajima S, Nanami T, Suzuki T,
Oshima Y, Tokura N, Takatsuka J, Funahashi K, Tochigi N and Shimada
H: Five-year postsurgical monitoring of serum p53 antibody for
locally advanced esophageal squamous cell carcinoma. Clin J
Gastroenterol. 11:278–281. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Suzuki T, Yajima S, Ishioka N, Nanami T,
Oshima Y, Washizawa N, Funahashi K, Otsuka S, Nemoto T and Shimada
H: Prognostic significance of high serum p53 antibody titers in
patients with esophageal squamous cell carcinoma. Esophagus.
15:294–300. 2018.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Ito M, Yajima S, Suzuki T, Oshima Y,
Nanami T, Sumazaki M, Shiratori F, Funahashi K, Tochigi N and
Shimada H: High serum PD-L1 level is a poor prognostic biomarker in
surgically treated esophageal cancer. Cancer Med. 9:1321–1327.
2020.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Ito M, Oshima Y, Yajima S, Suzuki T,
Nanami T, Shiratori F, Funahashi K, Nemoto T and Shimada H: Is high
serum programmed death ligand 1 level a risk factor for poor
survival in patients with gastric cancer? Ann Gastroenterol Surg.
2:313–318. 2018.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Ito M, Oshima Y, Yajima S, Suzuki T,
Nanami T, Shiratori F, Funahashi K and Shimada H: Diagnostic impact
of high serum midkine level in patients with gastric cancer. Ann
Gastroenterol Surg. 3:195–201. 2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Oshima Y, Shimada H, Yajima S, Nanami T,
Matsushita K, Nomura F, Kainuma O, Takiguchi N, Soda H, Ueda T, et
al: NY-ESO-1 autoantibody as a tumor-specific biomarker for
esophageal cancer: Screening in 1969 patients with various cancers.
J Gastroenterol. 51:30–34. 2016.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Sahin U, Tureci O, Schmitt H, Cochlovius
B, Johannes T, Schmits R, Stenner F, Luo G, Schobert I and
Pfreundschuh M: Human neoplasms elicit multiple specific immune
responses in the autologous host. Proc Natl Acad Sci USA.
92:11810–11813. 1995.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Cancer Immunome Database: Available from:
http://www2.licr.org/CancerImmunomeDB/.
|
|
13
|
Nakashima K, Shimada H, Ochiai T,
Kuboshima M, Kuroiwa N, Okazumi S, Matsubara H, Nomura F, Takiguchi
M and Hiwasa T: Serological identification of TROP2 by recombinant
cDNA expression cloning using sera of patients with esophageal
squamous cell carcinoma. Int J Cancer. 112:1029–1035.
2004.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Kuboshima M, Shimada H, Liu TL, Nakashima
K, Nomura F, Takiguchi M, Hiwasa T and Ochiai T: Identification of
a novel SEREX antigen, SLC2A1/GLUT1, in esophageal squamous cell
carcinoma. Int J Oncol. 28:463–468. 2006.PubMed/NCBI
|
|
15
|
Kuboshima M, Shimada H, Liu TL, Nomura F,
Takiguchi M, Hiwasa T and Ochiai T: Presence of serum tripartite
motif-containing 21 antibodies in patients with esophageal squamous
cell carcinoma. Cancer Sci. 97:380–386. 2006.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Shimada H, Kuboshima M, Shiratori T,
Nabeya Y, Takeuchi A, Takagi H, Nomura F, Takiguchi M, Ochiai T and
Hiwasa T: Serum anti-myomegalin antibodies in patients with
esophageal squamous cell carcinoma. Int J Oncol. 30:97–103.
2007.PubMed/NCBI
|
|
17
|
Shimada H, Shiratori T, Yasuraoka M,
Kagaya A, Kuboshima M, Nomura F, Takiguchi M, Ochiai T, Matsubara H
and Hiwasa T: Identification of Makorin 1 as a novel SEREX antigen
of esophageal squamous cell carcinoma. BMC Cancer.
9(232)2009.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Kagaya A, Shimada H, Shiratori T,
Kuboshima M, Nakashima-Fujita K, Yasuraoka M, Nishimori T, Kurei S,
Hachiya T, Murakami A, et al: Identification of a novel SEREX
antigen family, ECSA, in esophageal squamous cell carcinoma.
Proteome Sci. 9(31)2011.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Japan Esophageal Society. Japanese
Classification of Esophageal Cancer, 11th Edition: Part I.
Esophagus. 14:1–36. 2017.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Japanese Gastric Cancer Association.
Japanese classification of gastric carcinoma: 3rd English edition
Japanese Gastric Cancer Association. Gastric Cancer. 14:101–112.
2011.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Japanese Society for Cancer of the Colon
and Rectum. Japanese Classification of Colorectal, Appendiceal, and
Anal Carcinoma: The 3d English Edition [Secondary Publication]. J
Anus Rectum Colon. 3:175–195. 2019.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Shimada H, Kagaya A, Shiratori T, Nomura
F, Takiguchi M, Matsubara H and Hiwasa T: Detection of anti-CUEC-23
antibodies in serum of patients with esophageal squamous cell
carcinoma: A possible new serum marker for esophageal cancer. J
Gastroenterol. 44:691–696. 2009.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Matsutani T, Hiwasa T, Takiguchi M, Oide
T, Kunimatsu M, Saeki N and Iwadate Y: Autologous antibody to
Src-homology 3-domain GRB2-like 1 specifically increases in the
sera of patients with low-grade gliomas. J Exp Clin Cancer Res.
31(85)2012.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Machida T, Kubota M, Kobayashi E, Iwadate
Y, Saeki N, Yamaura A, Nomura F, Takiguchi M and Hiwasa T:
Identification of stroke-associated-antigens via screening of
recombinant proteins from the human expression cDNA library
(SEREX). J Transl Med. 13(71)2015.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Yoshida Y, Wang H, Hiwasa T, Machida T,
Kobayashi E, Mine S, Tomiyoshi G, Nakamura R, Shinmen N, Kuroda H,
et al: Elevation of autoantibody level against PDCD11 in patients
with transient ischemic attack. Oncotarget. 9:8836–8848.
2017.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Wang H, Zhang XM, Tomiyoshi G, Nakamura R,
Shinmen N, Kuroda H, Kimura R, Mine S, Kamitsukasa I, Wada T, et
al: Association of serum levels of antibodies against MMP1, CBX1,
and CBX5 with transient ischemic attack and cerebral infarction.
Oncotarget. 9:5600–5613. 2017.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Kobayashi S, Hiwasa T, Ishige T,
Rahmutulla B, Kano M, Hoshino T, Minamoto T, Shimada H, Nomura F,
Matsubara H and Matsushita K: Anti-FIRΔexon2, a splicing variant
form of PUF60, autoantibody is detected in the sera of esophageal
squamous cell carcinoma. Cancer Sci. 110:2004–2013. 2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Hiwasa T, Machida T, Zhang XM, Kimura R,
Wang H, Iwase K, Ashino H, Taira A, Arita E, Mine S, et al:
Elevated levels of autoantibodies against ATP2B4 and BMP-1 in sera
of patients with atherosclerosis-related diseases. Immunome Res.
11(97)2015.
|
|
29
|
Hiwasa T, Tomiyoshi G, Nakamura R, Shinmen
N, Kuroda H, Kunimatsu M, Mine S, Machida T, Sato E, Takemoto M, et
al: Serum SH3BP5-specific antibody level is a biomarker of
atherosclerosis. Immunome Res. 13(132)2017.
|
|
30
|
Shimada H, Ochiai T and Nomura F: Japan
p53 Antibody Research Group. Titration of serum p53 antibodies in
1,085 patients with various types of malignant tumors: A
multiinstitutional analysis by the Japan p53 Antibody Research
Group. Cancer. 97:682–689. 2003.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Shimada H, Nabeya Y, Okazumi S, Matsubara
H, Shiratori T, Gunji Y, Kobayashi S, Hayashi H and Ochiai T:
Prediction of survival with squamous cell carcinoma antigen in
patients with resectable esophageal squamous cell carcinoma.
Surgery. 133:486–494. 2003.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Kanda Y: Investigation of the freely
available easy-to-use software ‘EZR’ for medical statistics. Bone
Marrow Transplant. 48:452–458. 2013.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Castets F, Bartoli M, Barnier JV, Baillat
G, Salin P, Moqrich A, Bourgeois JP, Denizot F, Rougon G, Calothy G
and Monneron A: A novel calmodulin-binding protein, belonging to
the WD-repeat family, is localized in dendrites of a subset of CNS
neurons. J Cell Biol. 134:1051–1062. 1996.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Muro Y, Chan EK, Landberg G and Tan EM: A
cell-cycle nuclear autoantigen containing WD-40 motifs expressed
mainly in S and G2 phase cells. Biochem Biophys Res Commun.
207:1029–1037. 1995.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Castets F, Rakitina T, Gaillard S, Moqrich
A, Mattei MG and Monneron A: Zinedin, SG2NA, and striatin are
calmodulin-binding, WD repeat proteins principally expressed in the
brain. J Biol Chem. 275:19970–19977. 2000.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Janssens V and Goris J: Protein
phosphatase 2A: A highly regulated family of serine/threonine
phosphatases implicated in cell growth and signalling. Biochem J.
353(417)2001.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Hoof CV and Goris J: Phosphatases in
apoptosis: To be or not to be, PP2A is in the heart of the
question. Biochim Biophys Acta. 1640:97–104. 2003.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Wong M, Hyodo T, Asano E, Funasaka K,
Miyahara R, Hirooka Y, Goto H, Hamaguchi M and Senga T: Silencing
of STRN4 suppresses the malignant characteristics of cancer cells.
Cancer Sci. 105:1526–1532. 2014.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Wang Z, Li Y, Cao J, Zhang W, Wang Q,
Zhang Z, Gao Z, Ye Y, Jiang K and Wang S: MicroRNA profile
identifies miR-6165 could suppress gastric cancer migration and
invasion by targeting STRN4. Onco Targets Ther. 13:1859–1869.
2020.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Jiang F, Zheng Q, Chang L, Li X, Wang X
and Gu X: Pro-oncogene pokemon promotes prostate cancer progression
by inducing STRN4 expression. J Cancer. 10:1833–1845.
2019.PubMed/NCBI View Article : Google Scholar
|
|
41
|
The Human Protein Atlas:Available from:
https://www.proteinatlas.org/ENSG00000090372-STRN4/pathology.
|