Introduction
The latest data indicates that liver cancer is the
sixth most prevalent malignant tumor worldwide and the fourth most
prevalent in China, and primary liver cancer caused ~757,948
mortalities worldwide in 2022 (1,2).
Hepatocellular carcinoma (HCC) is the main histopathological type
of primary liver cancer, accounting for 75-85% of cases, with
chronic hepatitis B or C viral infections being the primary risk
factors (1,2). HCC has an insidious onset and rapid
progression, with the majority of patients diagnosed at a late
stage. In addition, HCC tumors are heterogeneous (3). At present, the HCC treatment options
are diverse and include surgical resection, liver transplantation,
chemotherapy, radiotherapy, targeted therapy and interventional
therapy. However, recurrence and metastasis following treatment are
the main contributors to the poor prognosis of patients with HCC
(4-7).
Therefore, investigating the mechanisms of HCC recurrence and
metastasis, identifying the specific metastatic pathways and
developing novel treatment strategies are key to improving patient
prognosis.
The epithelial-to-mesenchymal transition (EMT) is a
biological process in which epithelial cells lose their polarity
and tight connections, transforming into mesenchymal cells. During
this process, the expression of epithelial cell markers (such as
E-cadherin) decrease, while mesenchymal markers (such as
N-cadherin, vimentin and fibronectin) increase. Concurrently, the
cells lose their polarity and acquire a motile phenotype (8-10).
Based on this characteristic, the EMT is a key process in tumor
invasion and metastasis. Long non-coding RNAs (lncRNAs) are
functionally defined as transcripts of >200 nucleotides in
length that lack protein-coding potential. lncRNAs exhibit aberrant
expression in various types of tumors (such as endometrial and
ovarian cancer) and are notable regulators of the tumor metastasis
process (11-13).
Increasing evidence demonstrates that lncRNAs participate in the
regulation of the EMT, affecting tumor invasion and metastasis and
are hypothesized to be potential targets for tumor diagnosis and
treatment (14-16).
For example, a study by Yuan et al (17) reports that lncRNA UCID enhances
Snail mRNA stability, promotes the EMT and accelerates the
metastasis of HCC cells by interacting with Snail mRNA.
Additionally, a study by Zhao et al (18) demonstrates that lncRNA PNUTS
facilitates metastasis and invasion of HCC cells by targeting zinc
finger E-box binding homeobox 1 and activating the EMT pathway.
The functional role of lncRNAs in metastasis is
often associated with the activity of microRNA (miRNA/miR). A
predominant mechanism involves the lncRNA acting as a competing
endogenous RNA or molecular sponge, sequestering specific miRNAs
and preventing them from binding and repressing their target mRNAs
(19). For example, in HCC,
lncRNAs such as H19, MALAT1 and HULC facilitate metastasis by
sponging tumor-suppressive miRNAs (such as miR-200a-3p and
miR-15b), which derepresses oncogenic mRNAs that promote the EMT
and metastatic progression (20,21).
This lncRNA-miRNA-mRNA axis represents a key layer of epigenetic
regulation in cancer cell dissemination. Therefore, comprehensively
profiling lncRNA and mRNA expression changes during the EMT process
is a key step towards identifying novel nodes within this
regulatory network.
In the present study, RNA sequencing was used to
identify differentially expressed (DE) lncRNAs and mRNAs in a
transforming growth factor-β (TGF-β)-induced EMT model of Huh7
cells. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and
Genomes (KEGG) enrichment analyses were carried out on the
identified DE lncRNAs and mRNAs in order to highlight potential
lncRNAs and signaling pathways that may be involved in the
regulation of the EMT and HCC migration.
Materials and methods
Cell culture
Huh7 cells (cat. no. SCSP-526) were obtained from
The Cell Bank of Type Culture Collection of the Chinese Academy of
Sciences. Huh7 cells were cultured in Dulbecco's Modified Eagle's
Medium (DMEM; Gibco; Thermo Fisher Scientific, Inc.) enriched with
10% fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc.)
and 1% penicillin-streptomycin at 37˚C and under 5% CO2.
Routine medium replacement was carried out according to cell
proliferation status, as assessed by microscopic confluence and
medium color change, which is indicative of metabolic activity.
In vitro HCC EMT cell model
To induce the in vitro EMT cell model, Huh7
cells were cultured at 37˚C in DMEM with 2.5% FBS and 10 ng/ml
TGF-β (cat. no. HY-P7118; MedChemExpress) for 4 days. This
treatment was used to construct an in vitro EMT model (n=3).
Cells in the control group were cultured in DMEM containing 2.5%
FBS without TGF-β (n=3). The successful induction of the EMT was
confirmed by observing a transition from an epithelial
cobblestone-like to a mesenchymal spindle-shaped morphology under a
phase-contrast microscope. Successful induction of the EMT was
further confirmed using reverse transcription-quantitative PCR
(RT-qPCR) analysis after a downregulation of E-cadherin and an
upregulation of N-cadherin, Slug and Snail were revealed.
Cell migration assay
After the 4-day EMT induction, both TGF-β-treated
and control Huh7 cells were detached using trypsin and resuspended
in serum-free medium with or without 10 ng/ml TGF-β, respectively.
Subsequently, the cells were seeded into the upper chamber of
Transwell inserts (8 µm pore size with a polycarbonate membrane;
cat. no. 3422; Corning Life Sciences) without Matrigel at a density
of 1x105 cells/well. The lower chamber was filled with
600 µl medium containing 10% FBS. The plate was then cultured at
37˚C with 5% CO2 for 24 h. The cells were then fixed
with 4% paraformaldehyde at room temperature for 30 min and stained
with 1% crystal violet for 5 min at 37˚C. The migrated cells were
imaged in five randomly selected fields of view using an inverted
light microscope (Olympus IX73) at a x200 magnification and then
manually counted.
RT-qPCR
According to the manufacturer's protocol, total RNA
was extracted from the aforementioned TGF-β-treated and control
Huh7 cells using the Total RNA Miniprep kit (cat. no.
UE-MN-MS-RNA-50; Suzhou UELandy Biotechnology Co., Ltd.), which
included an RNA extraction buffer. The concentration of RNA was
quantified using a UV spectrophotometer. Subsequently, cDNA
synthesis was carried out following the protocol provided with the
RevertAidTM First Strand cDNA Synthesis kit (cat. no. K1622; Thermo
Fisher Scientific, Inc.). qPCR was carried out using the
synthesized cDNA as a template, following the manufacturers
protocol of the TB Green® Premix Ex Taq™ II (Tli RNase H
Plus) kit (cat. no. RR820A; Takara Biotechnology Co., Ltd.). The
thermocycling conditions used were as follows: Initial denaturation
at 95˚C for 30 sec, followed by 40 cycles of 95˚C for 5 sec and
then 60˚C for 34 sec; and subsequently, a melt curve analysis was
carried out at 95˚C for 15 sec, 60˚C for 1 min and finally 95˚C for
15 sec. The relative expression level of the target gene was
calculated using the 2-ΔΔCq method
(22), with GAPDH as the internal
reference. Primers were synthesized by Sangon Biotech Co., Ltd.,
(Table I).
 | Table IPrimers for reverse
transcription-quantitative PCR. |
Table I
Primers for reverse
transcription-quantitative PCR.
| Gene | Primer
direction | Primer sequences
(5'-3') |
|---|
| E-cadherin | F |
GAATGACAACAAGCCCGAAT |
| | R |
GACCTCCATCACAGAGGTTCC |
| N-cadherin | F |
GGTGGAGGAGAAGAAGACCAG |
| | R |
GCATCAGGCTCCACAGT |
| Slug | F |
TGGTTGCTTCAAGGACACAT |
| | R |
GTTGCAGTGAGGGCAAGAA |
| Snail | F |
GCTGCAGGACTCAATCCAGA |
| | R |
ATCTCCGGAGGTGGGATG |
| NNMT-205 | F |
CAGGAGCTGGAGAAGTGGCTG |
| | R |
TGCTTGACCGCCTGTCTCAAC |
| CASC15-204 | F |
GCACTGACCTCCTTCATTCTGC |
| | R |
AAGCAACTCCAGATGAATCCAGG |
| UBASH3B-202 | F |
TGTGTCGGCATGGTGAGAGGAT |
| | R |
GGCATGTTCAGGTTGGTGCGTA |
| CAPN2-206 | F |
AGGACATGCACACCATCGGCTT |
| | R |
CGGAGGTTGATGAAGGTGTCTG |
| CAV2-214 | F |
ATCGCAGAGCCGGTGACTAC |
| | R |
AAGCATCGTCCTACGCTCGT |
| COL1A1 | F |
GAGGGCCAAGACGAAGACATC |
| | R |
CAGATCACGTCATCGCACAAC |
| BMP6 | F |
TTCCTCAACGACGCGGACATG |
| | R |
CACCCTCAGGAATCTGGGATAAG |
| TUBA1A | F |
TGCCCCTGTCATCTCTGCTG |
| | R |
CGCTTGGTCTTGATGGTGGC |
| ATP2B2 | F |
CATCCTCAACGAACTCACCTGC |
| | R |
GATATTGTCGCCAGTGACCATGC |
| F2 | F |
ATGGGCTGGATGAGGACTCAGA |
| | R |
CGGTTTTGTCCTCCAGCGACTT |
| GAPDH | F |
GAGAAGGCTGGGGCTCAT |
| | R |
TGCTGATGATCTTGAGGCTG |
RNA sequencing
RNA sequencing was carried out by Guangzhou
Genedenovo Biotechnology Co., Ltd (http://www.genedenovo.com/). Total RNA was isolated
using the TRIzol® reagent kit (cat. no.15596026,
Invitrogen; Thermo Fisher Scientific, Inc.) following the
manufacturer's protocol. The integrity and quality of the RNA were
evaluated using an Agilent 2100 Bioanalyzer (Agilent Technologies,
Inc.) and further confirmed via RNase-free agarose gel
electrophoresis. Following the extraction of total RNA, ribosomal
RNAs were eliminated to enrich mRNAs and ncRNAs. Sequencing
libraries were prepared using the NEBNext® UltraTM II
Directional RNA Library Prep kit (cat. no. E7760S; New England
Biolabs). The resultant mRNAs and ncRNAs were fragmented using the
fragmentation buffer from the aforementioned kit. These fragments
were reverse-transcribed into cDNA using random primers. The
synthesis of the second-strand cDNA was facilitated by DNA
polymerase I, RNase H and dNTPs (with dUTP replacing dTTP) in DNA
polymerase reaction buffer from the same aforementioned kit. The
cDNA fragments were then purified using the QIAquick PCR
Purification kit (cat. no. 28104; Qiagen Benelux B.V.), followed by
end repair, the addition of poly(A) tails and ligation to Illumina
sequencing adapters. Subsequently, uracil-N-glycosylase was used to
digest the second-strand cDNA. The resulting digested products
underwent size selection via agarose gel electrophoresis, followed
by PCR amplification. Sequencing was then carried out by Guangzhou
Genedenovo Biotechnology Co. Ltd. using the Illumina HiSeq™ 4000
platform with paired-end 150 bp strand-specific reads. For cluster
generation and sequencing, the HiSeq 3000/4000 PE Cluster kit (cat.
no. PE-410-1001; Illumina, Inc.) and SBS kit (cat. no. FC-410-1003;
Illumina, Inc.) were used according to the manufacturer's
protocols. The final library was loaded at a concentration of 20 nM
as measured using a Qubit fluorometer. For bioinformatic analysis,
raw sequencing reads were subjected to quality control using FastQC
software (version 0.12.1; https://www.bioinformatics.babraham.ac.uk/projects/fastqc/),
followed by adapter trimming and quality filtering. The cleaned
reads were then aligned to the reference genome using HISAT2
software (version 2.2.1; https://daehwankimlab.github.io/hisat2/download/).
Gene expression levels were then quantified using featureCounts
software (version 2.0.6; http://subread.sourceforge.net). Subsequently, the
sequencing data were analyzed using edgeR software (version
4.0.16;https://bioconductor.org/packages/edgeR). The
screening threshold used to identify the significance of DE lncRNAs
and mRNAs was false-discovery rate (FDR)<0.05 and [log2
fold-change (FC)]>1.
GO and KEGG analyses
GO and KEGG analyses were carried out using the
‘clusterProfiler’ R package (version 4.0.5; R Development Core
Team) to elucidate the potential molecular mechanisms through which
DE lncRNAs and mRNAs influence HCC migration by regulating the EMT
process. GO analysis was used to classify and annotate the
functions of the DE genes, as well as to perform enrichment
analysis. KEGG analysis was used to identify significantly enriched
signaling pathways or metabolic pathways associated with the DE
genes. Q<0.05 was set as the screening criterion.
Construction of the lncRNA-mRNA
interaction network
To investigate the potential roles of DE lncRNAs in
HCC migration and the EMT, a regulatory network was constructed
based on the lncRNA-mRNA pairs predicted by trans-action analysis.
The interaction pairs were formatted as a two-column source-target
list and imported into Cytoscape software (version 3.10.3;
https://cytoscape.org/download.html)
using default settings to generate the network visualization.
Statistical analysis
All data were statistically analyzed using SPSS
(version 16.0; SPSS, Inc.). Quantitative data are expressed as the
mean ± SD. Differences between two groups were compared using an
unpaired t-test. All of the experiments were carried out in three
independent biological replicates. P<0.05 was considered to
indicate a statistically significant difference.
Results
Validation of the HCC cell EMT
model
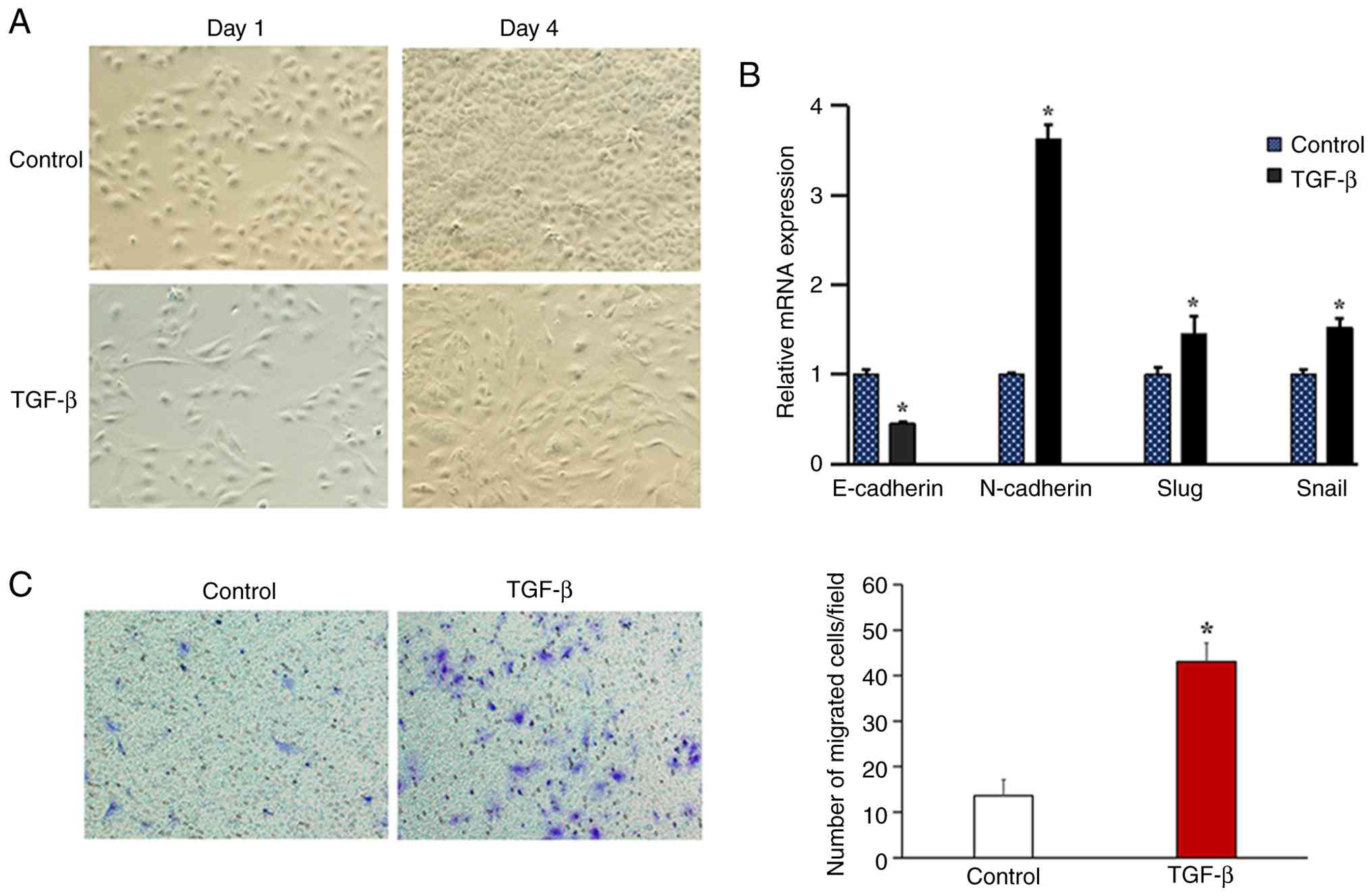
To construct an in vitro EMT model, Huh7
cells were stimulated with 10 ng/ml TGF-β for 4 days, with
untreated cells serving as the control group. Compared with this
control, the TGF-β-treated Huh7 cells lost their original polarity
and transformed into a spindle or typical fibroblast-like phenotype
(Fig. 1A). At the molecular level,
TGF-β treatment significantly downregulated the mRNA expression of
the epithelial marker E-cadherin and upregulated the expression of
mesenchymal markers (N-cadherin, Snail and Slug) compared with the
control group (Fig. 1B).
Functionally, the induction of the EMT led to a significant
enhancement in the cell migration ability in the TGF-β treatment
group compared with the control (Fig.
1C).
Analysis of the DE lncRNAs and mRNAs
in TGF-β-treated Huh7 cells
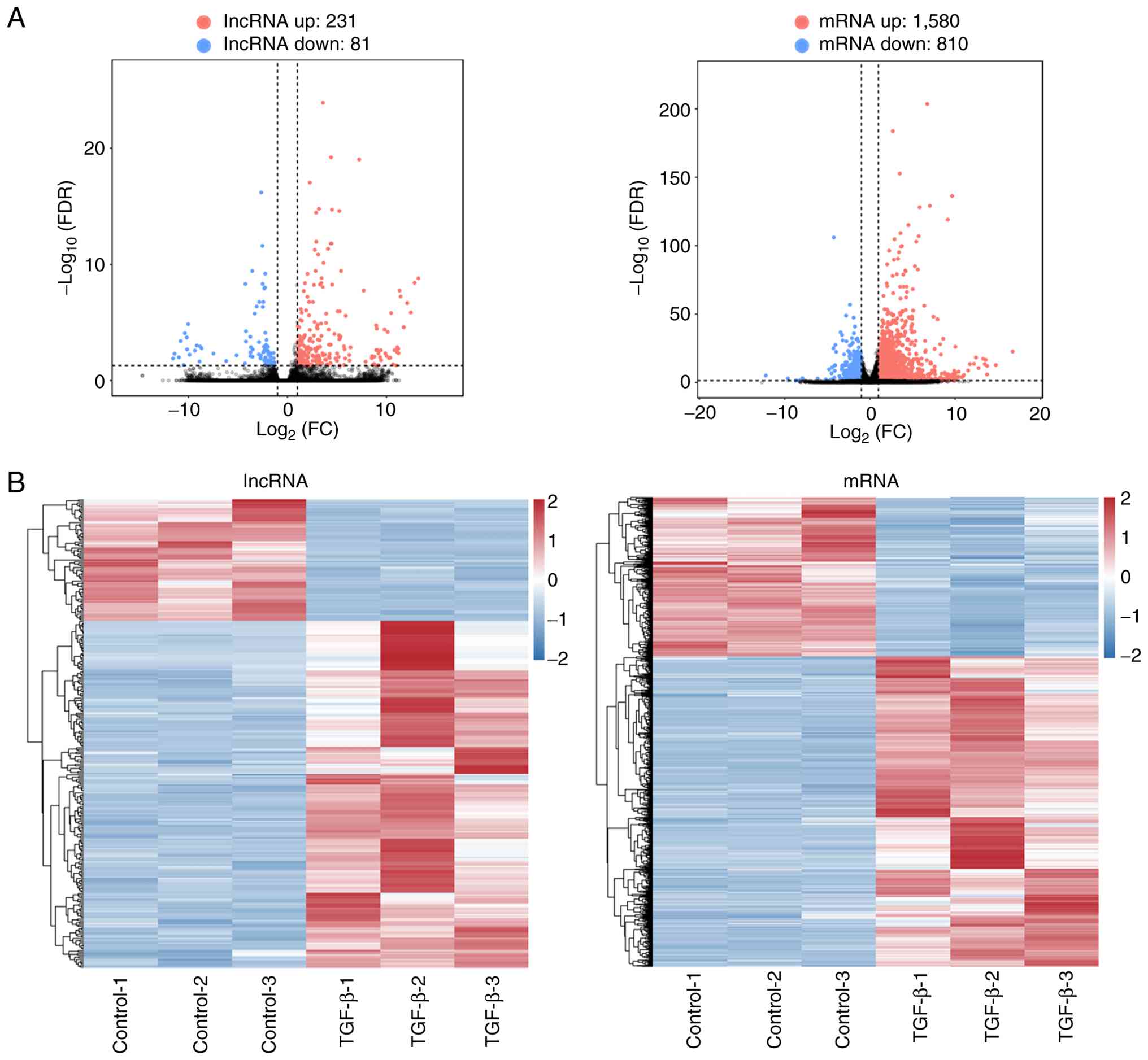
RNA sequencing was carried out to detect the DE
mRNAs and lncRNAs in Huh7 cells treated with or without TGF-β. The
volcano plots revealed 231 upregulated and 81 downregulated DE
lncRNAs (Tables SI and SII) as well as 1,580 upregulated and 810
downregulated DE mRNAs (Tables
SIII and SIV) (Fig. 2A). In addition, hierarchical
clustering of the DE lncRNAs and mRNAs was carried out and notable
differences between the Huh7 cells with or without TGF-β treatment
were identified (Fig. 2B). These
findings potentially identify key DE lncRNAs (for example, CAV2-214
and TACC1-226) and DE mRNAs (for example, TUBA1A and H4C11) that
regulate the EMT process and promote HCC migration.
GO and KEGG enrichment analyses
The function of lncRNAs is associated with their
co-expressed protein-coding genes. Trans-action analysis was
conducted to identify associations between DE lncRNAs and mRNAs.
The prediction results indicated that 312 lncRNAs may act on 2,367
target genes, forming a total of 101,780 lncRNA-gene connections
(Table SV). To further
investigate the potential function of these identified DE lncRNAs,
both GO and KEGG functional enrichment analyses were carried out.
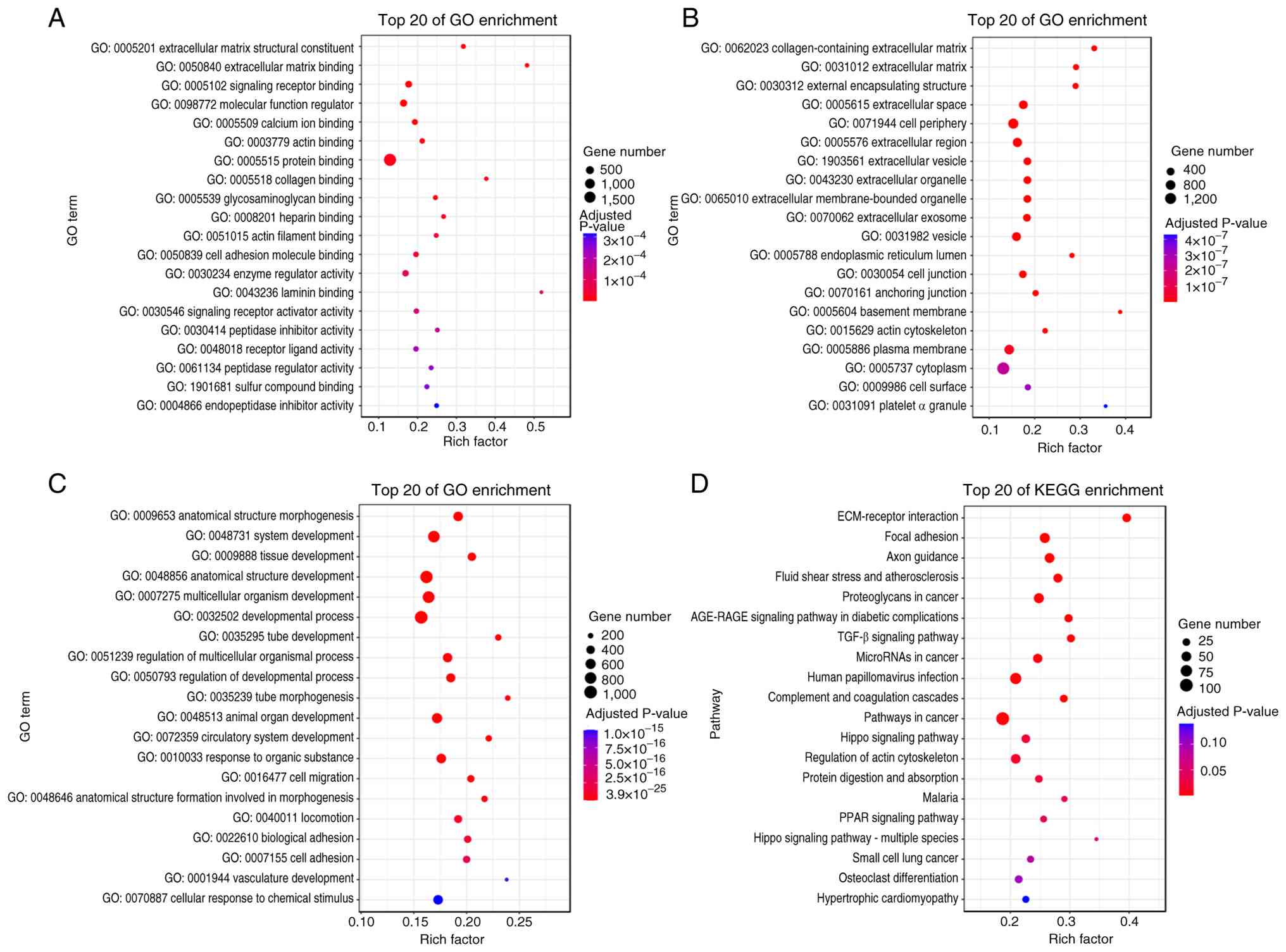
Within the GO molecular function classification, several key GO
terms were enriched, including ‘extracellular matrix structural
constituent’, ‘extracellular matrix binding’ and ‘signaling
receptor binding’ (Fig. 3A).
Within the GO cellular component classification, several key GO
terms were enriched, including ‘collagen-containing extracellular
matrix’, ‘extracellular matrix’ and ‘external encapsulating
structure’ (Fig. 3B). Within the
GO biological process classification, several key GO terms were
enriched, including ‘anatomical structure morphogenesis’, ‘system
development’ and ‘tissue development’ (Fig. 3C). A number of these GO terms have
been reported to be associated with EMT processes, such as ‘actin
binding’, the ‘extracellular region’ and ‘cell migration’ (23-25).
The results of the KEGG enrichment analysis revealed that the
associated signaling pathways included ‘ECM-receptor interaction’,
‘focal adhesion’ and ‘axon guidance’ (Fig. 3D). These pathways have also been
reported to be associated with the EMT process (26-28).
These results were used to select the lncRNAs that are potentially
involved in the regulation of the EMT in the progression of
HCC.
Establishment of the lncRNA-mRNA
interaction network
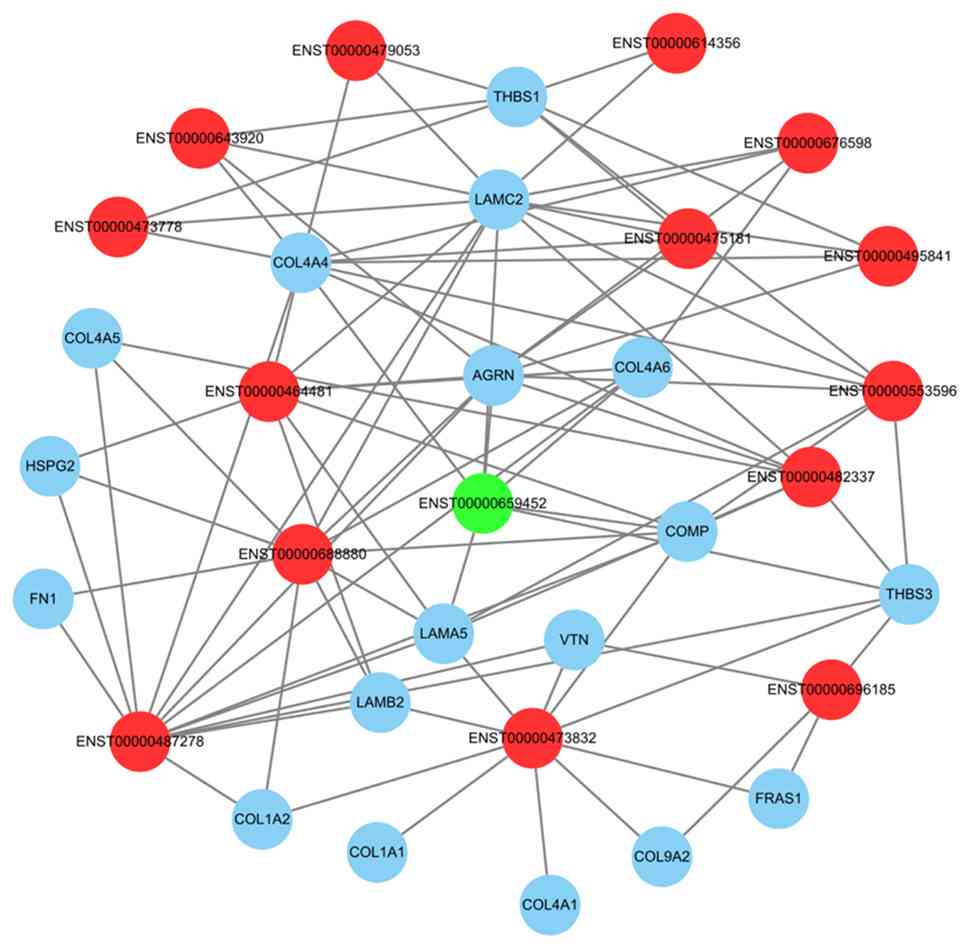
To further investigate how DE lncRNAs potentially
affect the migration and development of HCC by regulating the EMT,
a network map was constructed based on 15 DE lncRNAs and 18 DE
mRNAs. The lncRNA-mRNA interactions were predicted by the previous
trans-action analysis (Fig.
4).
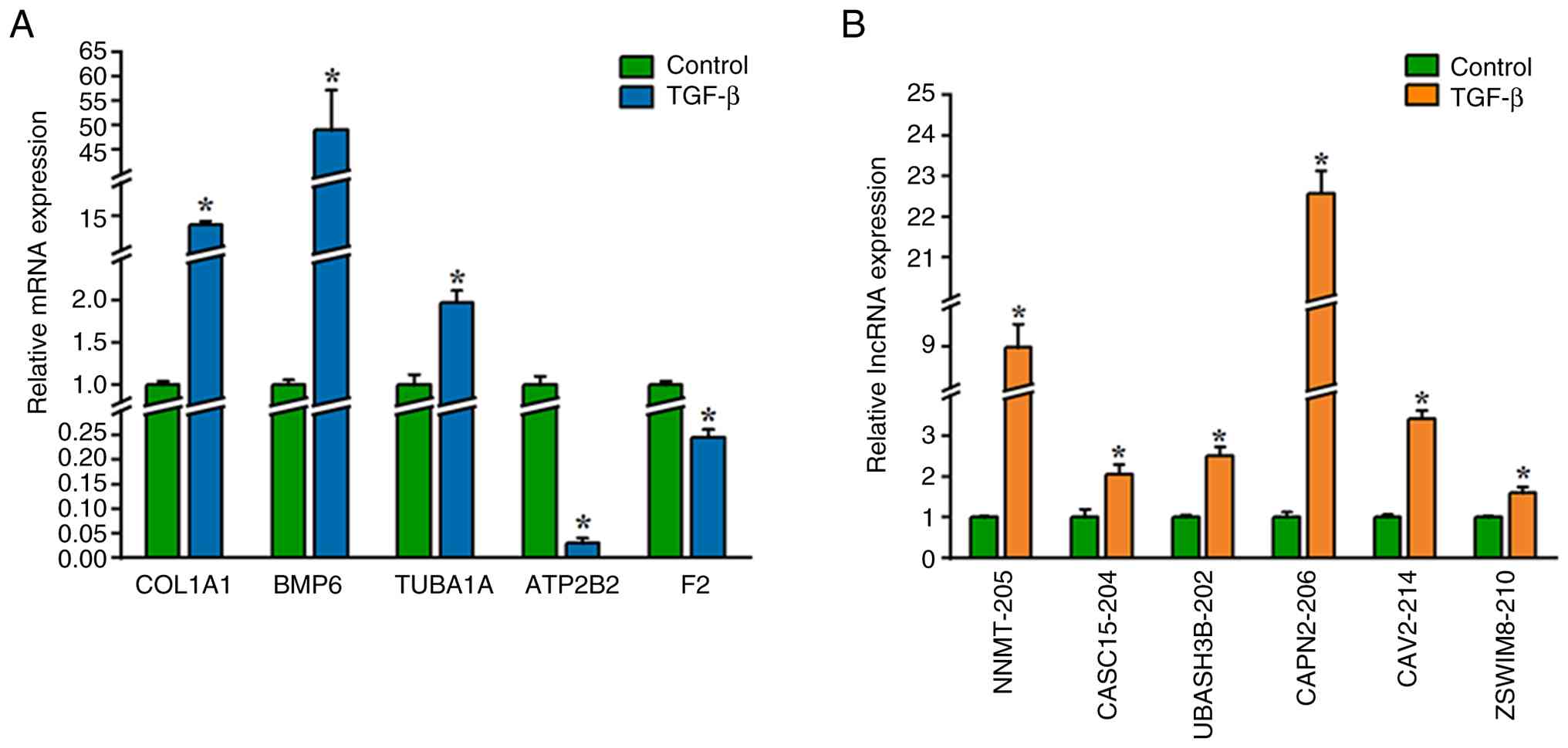
Validation of sequencing data using
RT-qPCR
To validate the RNA sequencing results, 5 DE mRNAs
(COL1A1, BMP6, TUBA1A, ATP2B2 and F2) and 6 DE lncRNAs (NNMT-205,
CASC15-204, UBASH3B-202, CAPN2-206, CAV2-214 and ZSWIM8-210) were
selected for RT-qPCR analysis based on their statistical
significance and putative association with the EMT. The RT-qPCR
results were consistent with the sequencing data. Compared with the
control, TGF-β treatment significantly upregulated the expression
of COL1A1, BMP6, TUBA1A, NNMT-205, CASC15-204, UBASH3B-202,
CAPN2-206, CAV2-214 and ZSWIM8-210. Furthermore, TGF-β treatment
significantly downregulated the expression of ATP2B2 and F2
compared with the control (Fig. 5A
and B).
Discussion
The EMT is involved in the metastasis of various
types of tumors (such as colorectal and breast cancer) and is a key
mechanism for the recurrence and metastasis of malignant tumors,
including HCC (29-31).
Despite the role of the EMT in HCC progression, therapeutic
strategies targeting regulators of the EMT, such as TGF-β/Smad
signaling or Snail family transcription factors, have not yet been
clinically successful. This highlights that the complexity and
compensatory mechanisms in the EMT regulatory network are
intricate.
Increasing evidence suggests that lncRNAs can affect
the migration and progression of HCC by regulating the EMT process
(32,33). For example, a study by Chen et
al (34) reports that lncRNA
MALAT1 is highly expressed in HCC cells and promotes the EMT
through the miRNA22-Snail family transcriptional repressor 1 axis,
which accelerates the progression of HCC. Furthermore, a study by
Zhang et al (35)
identifies that lncRNA SNHG 3 is aberrantly activated in highly
metastatic HCC and can accelerate the EMT process and invasion of
HCC cells. However, despite a small number of well-characterized
molecules (such as MALAT1 and SNHG 3), the majority of lncRNAs that
are dynamically expressed during the EMT are still functionally
unannotated, suggesting that there may be a number of potential
therapeutic targets that still require investigation. Therefore,
the present study aimed to generate a catalog of potential
candidate lncRNAs to suggest novel, uncharacterized regulators
beyond these established molecules, in order to potentially provide
insights into the mechanisms of HCC tumorigenesis and potential
therapeutic targets.
The present study investigated the molecular
mechanisms by which lncRNAs affect HCC migration and development by
regulating the EMT. An in vitro EMT model was successfully
constructed by treating Huh7 cells with 10 ng/ml TGF-β for 4 days.
RNA sequencing analysis of the lncRNAs and mRNAs in the model and
control groups was then carried out. The results revealed 312
significantly DE lncRNAs and 2,390 significantly DE mRNAs
(FDR<0.05 and |log2FC|>1). The expression profiles
of the lncRNAs and mRNAs in the model and control groups were then
systematically investigated through trans-action analysis, which
predicted 101,780 potential lncRNA-mRNA target gene pairs by
analyzing 312 significantly DE lncRNAs and 2,367 significantly DE
mRNAs. To further reveal the key lncRNAs, mRNAs and their regulated
signaling pathways, GO and KEGG enrichment analyses of the
significantly DE lncRNAs were carried out. The GO analysis results
revealed that DE lncRNAs were significantly enriched in cellular
components such as ‘collagen-containing extracellular matrix’,
‘extracellular matrix’ and ‘external encapsulation structure’. KEGG
pathway analysis revealed several key signaling pathways associated
with disease or biological processes, such as ‘ECM-receptor
interaction’, ‘focal adhesion’ and ‘axon guidance’. These findings
are consistent with previous reports and confirm the relevance of
the ECM-receptor interaction, focal adhesion and
collagen-containing extracellular matrix components in the
regulation of the EMT (36,26).
Furthermore, a lncRNA-mRNA network was constructed
in the present study, which further elucidated that lncRNAs affect
cellular molecular functions or signaling pathways by regulating
mRNAs, thereby influencing the EMT process. The lncRNA-mRNA network
results demonstrated that lncRNAs such as ZSWIM8-210
(ENST00000487278), BMS1P23-222 (ENST00000688880) and DGCR2-204
(ENST00000473832) were core nodes that interacted with multiple
mRNAs; suggesting that these lncRNAs may serve a key role in the
regulatory network. Upon reviewing the literature, it was
identified that lncRNAs ZSWIM8-210, BMS1P23-222 and DGCR2-204 are
yet to be elucidated in the field of HCC research. However, the
expression level of ZSWIM8 is associated with drug resistance in
breast cancer cells and epithelial ovarian cancer cells (37,38).
In the present study, the role of ZSWIM8-210 as a core node in the
lncRNA-mRNA network highlights its potential importance in the
regulatory landscape of cancer biology, thus, further investigation
into its functional roles and therapeutic relevance is
warranted.
The present study has several limitations that
should be considered. Firstly, the primary findings are based on
sequencing and bioinformatic analyses of in vitro
experiments. A key limitation of this approach is that it fails to
recapitulate the key influence of the in vivo
microenvironment. Secondly, although the co-expression network
analysis provided valuable predictive insights and has been
partially validated by the results of RT-qPCR, the functional roles
of the identified lncRNAs (including the validated candidates such
as ZSWIM8-210 and CAV2-214) in EMT regulation remain to be
experimentally verified. Future studies should include a functional
characterization through in vitro gain-of-function and
loss-of-function experiments to establish causative relationships
between the identified lncRNAs and the EMT phenotypes. Thirdly, the
precise molecular mechanisms through which these lncRNAs operate,
such as whether through chromatin remodeling, transcriptional
regulation or post-transcriptional modulation, requires further
elucidation. Investigating these mechanisms through subcellular
localization studies, RNA-protein interaction assays and validation
of downstream pathways are suggested experiments for future
studies.
Despite these limitations, the results of the
present study provided an understanding of the lncRNA regulatory
networks in TGF-β-induced EMT. The aforementioned experiments that
were suggested for future studies may help to translate these
bioinformatic predictions into mechanistic insights with potential
therapeutic implications for HCC metastasis.
In conclusion, the present study identified
significantly DE lncRNAs in Huh7 cells treated with TGF-β. GO
enrichment analysis revealed that these significantly DE lncRNAs
were involved in ‘cell migration’, ‘cell adhesion’, ‘tube
morphogenesis’ and ‘regulation of multicellular organismal
process’. KEGG pathway analysis suggested that these significantly
DE lncRNAs were involved in pathways associated with ‘ECM-receptor
interaction’, ‘focal adhesion’ and ‘axon guidance’. Lastly, by
constructing a lncRNA-mRNA network, potential DE lncRNAs and mRNAs
that may affect the EMT process were highlighted. These results
indicated that these DE lncRNAs may contribute to tumor development
and progression by participating in the EMT process. The present
study established a framework for the lncRNA regulatory network,
with the lncRNAs highlighted in the present study serving as
potential candidates for future research. To translate these
suggested bioinformatic predictions into mechanistic insights,
future work should address current knowledge gaps through in
vivo validation, functional assays, mechanistic studies and
clinical correlation. Such efforts are required in order to advance
therapeutic and prognostic strategies for HCC.
Supplementary Material
Upregulated differentially expressed
long non-coding RNAs upon treatment with transforming growth
factor-β.
Downregulated differentially expressed
long non-coding RNAs upon treatment with transforming growth
factor-β.
Upregulated differentially expressed
mRNAs upon treatment with transforming growth factor-β.
Downregulated differentially expressed
mRNAs upon treatment with transforming growth factor-β.
lncRNA targeting results.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the Guangxi Natural
Science Foundation (grant no. 2024GXNSFBA010071) and Research Team
Incubation Project of Minzu Hospital of Guangxi Zhuang Autonomous
Region (grant no. MKFY202104).
Availability of data and materials
The data generated in the present study may be found
in the Sequence Read Archive database under accession number
(PRJNA1017537) or at the following URL: https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA1017537.
Authors' contributions
QD and LM searched the literature, designed the
present study, interpreted the findings and revised the manuscript.
QD, JZ, YZ, XL, TG and QM carried out data management, statistical
analysis and performed the experiments. QD, JZ, YZ, XL and TG
confirm the authenticity of all the raw data. All authors read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates ofincidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Zheng RS, Chen R, Han BF, Wang SM, Li L,
Sun KX, Zeng HM, Wei WW and He J: Cancer incidence and mortality in
China, 2022. Zhonghua Zhong Liu Za Zhi (Chinese). 46:221–231.
2022.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Villanueva A: Hepatocellular carcinoma. N
Engl J Med. 380:1450–1462. 2019.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Luo Y, Lin J, Zhang J, Song Z, Zheng D,
Chen F, Zhuang X, Li A and Liu X: LncRNA SNHG17 contributes to
proliferation, migration, and poor prognosis of hepatocellular
carcinoma. Can J Gastroenterol Hepatol.
2021(9990338)2021.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Wu J, Lan Z, Li X, He J, Zhang D and Jin
T: A novel recombinant adenovirus expressing apoptin and melittin
genes kills hepatocellular carcinoma cells and inhibits the growth
of ectopic tumor. Invest New drugs. 42:428–441. 2024.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Choi JH and Thung SN: Advances in
histological and molecular classification of hepatocellular
carcinoma. Biomedicines. 11(2582)2023.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Ma PC, Wang GR, Men K, Li CJ, Gao N and Li
NJ: Advances in clinical application of nanoparticle-based therapy
for cancer treatment: A systematic review. Nano TransMed.
3(100036)2024.
|
|
8
|
Bure IV, Nemtsova MV and Zaletaev DV:
Roles of Ecadherin and noncoding RNAs in the epithelial-mesenchymal
transition and progression in gastric cancer. Int J Mol Sci.
20(2870)2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Gong J, Wang Y and Shu C: LncRNA CHRF
promotes cell invasion and migration via EMT in gastric cancer. Eur
Rev Med Pharmacol Sci. 24:1168–1176. 2020.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Saitoh M: Involvement of partial EMT in
cancer progression. J Biochem. 164:257–264. 2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Wang J, Shi P, Teng H, Lu L, Guo H and
Wang X: Corrigendum: lncRNA DCST1-AS1 promotes endometrial cancer
progression by modulating the miR-665/HOXB5 and miR-873-5p/CADM1
pathways. Front Oncol. 12(819908)2022.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Tang K, Lu D, Miao L, Mao Y and Yu X:
LncRNA TUG1 functions as a ceRNA for miR-1-3p to promote cell
proliferation in hepatic carcinogenesis. J Clin Lab Anal.
36(e24415)2022.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Zhao Y, Yuan D, Zhu D, Huang A, Jiang L,
Liu C, Qian H and Bu X: LncRNA-MSC-AS1 inhibits the ovarian cancer
progression by targeting miR-425-5p. J Ovarian Res.
14(109)2021.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Li WJ, Li G, Liu ZW, Chen ZY and Pu R:
LncRNA LINC00355 promotes EMT and metastasis of bladder cancer
cells through the miR424-5p/HMGA2 axis. Neoplasma. 68:1225–1135.
2021.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Gugnoni M and Ciarrocchi A: Long noncoding
RNA and epithelial mesenchymal transition in cancer. Int J Mol Sci.
20(1924)2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Heery R, Finn SP, Cuffe S and Gray SG:
Long non-coding RNAs: Key regulators of epithelial-mesenchymal
transition, tumour drug resistance and cancer stem cells. Cancers
(Basel). 9(38)2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Yuan S, Si W, Zhuang K, Li Y, Zhang Y, Liu
J, Yang L and Zhang X: LncRNA UCID promotes hepatocellular
carcinoma metastasis via stabilization of Snail. Onco Targets Ther.
14:725–736. 2021.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Zhao H, Liu C, Zhao C, Che C, Liu W and
Mei Y, Liu W and Mei Y: Alternatively-spliced lncRNA-PNUTS promotes
HCC cell EMT via regulating ZEB1 expression. Tumori. 109:28–37.
2023.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Wu Y and Qian Z: Long non-coding RNAs
(lncRNAs) and microRNAs regulatory pathways in the tumorigenesis
and pathogenesis of glioma. Discov Med. 28:129–138. 2019.PubMed/NCBI
|
|
20
|
Zhou Y, Fan RG, Qin CL, Jia J, Wu XD and
Zha WZ: LncRNA-H19 activates CDC42/PAK1 pathway to promote cell
proliferation, migration and invasion by targeting miR-15b in
hepatocellular carcinoma. Genomics. 111:1862–1872. 2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Li SP, Xu HX, Yu Y, He JD, Wang Z, Xu YJ,
Wang CY, Zhang HM, Zhang RX, Zhang JJ, et al: LncRNA HULC enhances
epithelial-mesenchymal transition to promote tumorigenesis and
metastasis of hepatocellular carcinoma via the miR-200a-3p/ZEB1
signaling pathway. Oncotarget. 7:42431–42446. 2016.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Zou W, Huang C, Chen Y, Tang J, Li Q, Fang
Q, Ma Y, Wu W and Feng S: Role of HDAC3 in the
epithelial-mesenchymal transition of retinal pigment epithelium
cells: Implications for proliferative vitreoretinopathy. Heliyon.
10(e39333)2024.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Mathieu M, Isomursu A and Ivaska J:
Positive and negative durotaxis - Mechanisms and emerging concepts.
J Cell Sci. 137(jcs261919)2024.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Liu X, Liu M, Ji M, Ma B, Hou YC, Yao XY,
Cheng QC and Chen L: Bone morphogenetic protein-6 suppresses
TGF-β2-induced epithelial-mesenchymal transition in retinal pigment
epithelium. Int J Ophthalmol. 17:646–652. 2024.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Horta CA, Doan K and Yang J:
Mechanotransduction pathways in regulating epithelial-mesenchymal
plasticity. Curr Opin Cell Biol. 85(102245)2023.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Capasso G, Mouawad N, Castronuovo M,
Ruggeri E, Visentin A, Trentin L and Frezzato F: Focal adhesion
kinase as a new player in the biology of onco-hematological
diseases: The starting evidence. Front Oncol.
14(1446723)2024.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Xiao GY, Tan X, Rodriguez BL, Gibbons DL,
Wang S, Wu C, Liu X, Yu J, Vasquez ME, Tran HT, et al: EMT
activates exocytotic Rabs to coordinate invasion and
immunosuppression in lung cancer. Proc Natl Acad Sci USA.
120(e2220276120)2023.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Dou R, Liu K, Yang C, Zheng J, Shi D, Lin
X, Wei C, Zhang C, Fang Y, Huang S, et al: EMT-cancer cells-derived
exosomal miR-27b-3p promotes circulating tumour cells-mediated
metastasis by modulating vascular permeability in colorectal
cancer. Clin Transl Med. 11(e595)2021.PubMed/NCBI View
Article : Google Scholar
|
|
30
|
Brown MS, Abdollahi B, Wilkins OM, Lu H,
Chakraborty P, Ognjenovic NB, Muller KE, Jolly MK, Christensen BC,
Hassanpour S and Pattabiraman DR: Phenotypic heterogeneity driven
by plasticity of the intermediate EMT state governs disease
progression and metastasis in breast cancer. Sci Adv.
8(eabj8002)2022.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Wang J, Peng J and Chen Y: The role of
stromal cells in epithelial-mesenchymal plasticity and its
therapeutic potential. Discov Oncol. 15(13)2024.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Yang X, Xu C, Liu C, Wu X, Chen X, Hou J
and Wang L: TGF-β1-Induced LINC01094 promotes
epithelial-mesenchymal transition in hepatocellular carcinoma
through the miR-122-5p/ TGFBR2-SAMD2-SMAD3 Axis. Funct Integr
Genomics. 24(123)2024.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Shah M and Sarkar D: HCC-Related lncRNAs:
Roles and Mechanisms. Int J Mol Sci. 25(597)2024.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Chen S, Wang G, Tao K, Cai K, Wu K, Ye L,
Bai J, Yin Y, Wang J, Shuai X, et al: Long noncoding RNA
metastasis-associated lung adenocarcinoma transcript 1 cooperates
with enhancer of zeste homolog 2 to promote hepatocellular
carcinoma development by modulating the microRNA-22/Snail family
transcriptional repressor 1 axis. Cancer Sci. 111:1582–1595.
2020.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Zhang PF, Wang F, Wu J, Wu Y, Huang W, Liu
D, Huang XY, Zhang XM and Ke AW: LncRNA SNHG3 induces EMT and
sorafenib resistance by modulating the miR-128/CD151 path way in
hepatocellular carcinoma. J Cell Physiol. 234:2788–2794.
2019.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Liu ZH, Zhang XF, Ben TR, Li M, Jin Y,
Wang TL and Song YQ: Focal adhesion in the tumour metastasis: from
molecular mechanisms to therapeutic targets. Biomark Res.
13(38)2025.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Gong K, Song K, Zhu Z, Xiang Q, Wang K and
Shi J: SWIM domain protein ZSWIM4 is required for JAK2 inhibition
resistance in breast cancer. Life Sci. 279(119696)2021.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Gong K, Huang Y, Zheng Y, Hao W and Shi K:
ZSWIM4 inhibition improves chemosensitivity in epithelial ovarian
cancer cells by suppressing intracellular glycine biosynthesis. J
Transl Med. 22(192)2024.PubMed/NCBI View Article : Google Scholar
|