Introduction
Renal cell carcinoma (RCC) is a prevalent and highly
lethal cancer affecting the genitourinary system. It is
particularly prevalent in men, with a mortality rate ranging from
30-40%. Risk factors for RCC include sex, obesity, hypertension,
smoking and chronic kidney disease (1,2).
Treatment typically involves surgical intervention, namely either
total or partial nephrectomy, but such radical procedures can
significantly impact the quality of life of patients. Complementary
to surgery, targeted pharmacotherapies are pivotal in managing RCC
(3). Nevertheless, the high cost
associated with these advanced therapies limits their widespread
adoption amongst patients with RCC, underscoring the critical need
for cost-effective treatment alternatives (4,5).
Traditional Chinese medicines have been garnering
increasing global attention due to their extensive historical use
and notable efficacy. With the advancement of modern pharmacology
and systems biology approaches, increasing efforts have been
devoted to elucidating the molecular mechanisms underlying their
therapeutic effects. Understanding of their mechanisms of action
has advanced as a result of progress in modern pharmacological
techniques (6).
Plants of the genus Trigonostemon, belonging
to the Euphorbiaceae family, are prevalent in tropical and
temperate regions. They have a long history of medicinal use
(7,8). In particular, Trigonostemon
xyphophyllorides (Croizat) L. K. Dai and T. L. Wu (TX) root is
a local herb that is traditionally used in Thailand for treating
food poisoning and snake bites. In China, the bark of this herb has
been utilized to alleviate asthma symptoms (9). Modern pharmacological studies have
identified TX to possess antiviral, antitumor, antibacterial,
anti-inflammatory and insecticidal properties (10-13).
Previous in vitro experiments have demonstrated that the
coumarin compound psoralen (PSO, PubChem CID 6199), which is found
in TX, exhibits potent antitumor effects, suggesting its potential
as a natural anticancer agent (14,15).
Despite accumulating evidence supporting the antitumor activity of
PSO, its therapeutic efficacy and more importantly its underlying
molecular mechanisms in RCC regulation remain poorly understood.
The signaling pathways and key molecular targets through which PSO
exerts anti-RCC effects remain to be fully elucidated. Therefore,
the aim of the present study was to systematically investigate the
anti-RCC effects of PSO using an integrated multi-omics strategy,
combining network pharmacology, molecular docking, molecular
dynamics simulation, transcriptomic analysis and experimental
validation. The present study seeks to offer both experimental and
mechanistic evidence supporting the clinical utilization of TX in
treating RCC, thereby advancing its research and development.
Materials and methods
Plant sources
The TX branches utilized in the present were
obtained from Haikou (China) and authenticated as TX by Professor
Guangtong Chen from the Department of Pharmacy, of Nantong
University. A voucher specimen (no. 2023026) has been archived at
the Herbarium Center, School of Pharmacy of Nantong University.
PSO extraction and structure
determination
The branches of TX were subjected to three rounds of
extraction with 95% ethanol under reflux conditions for 2 h each.
The herbal extract underwent purification through sequential
passage through silica gel column chromatography, gel column
chromatography and high-performance liquid chromatography [JAI
LC9103 Recycling preparative HPLC (Japan Analytical Industries),
JAIGEL-ODS-AP-P column and JAIGEL-GS310 column using a JAI
refractive index detector and a JAI UV-3702 detector with MultiChro
2000 workstation, 50 µl, MeOH-H2O (55:45), 10 ml/min] to
isolate pure PSO (11.2 mg). The identity of PSO was confirmed by
comparison of the monomer with literature data using 1H
nuclear magnetic resonance (NMR) and 13C NMR analyses
(16,17).
Target gene prediction
Potential gene targets of PSO were predicted using
the Swiss Target Prediction database (https://www.swisstargetprediction.ch/). To identify
target genes associated with RCC, databases, including OMIM
(https://www.omim.org/), DrugBank (https://go.drugbank.com/), DisGeNet (https://disgenet.com/), Genecards (https://www.genecards.org/) and Therapeutic Target
Database (TTD; https://ttd.idrblab.cn/), were queried. Compound and
disease-related gene sets were compiled by merging and removing
duplicates. The overlapping genes were analyzed to propose
potential compound target genes for RCC. These databases were
selected due to their complementary strengths in covering genetic
associations, curated therapeutic targets and disease relevance,
which are widely used in network pharmacology studies to improve
target prediction reliability.
Protein-protein interaction (PPI)
network construction
The potential target genes were inputted into the
STRING database https://string-db.org/ at a confidence level of 0.7
and subsequently imported into Cytoscape 3.10.2 software
(https://cytoscape.org/). Utilizing the metrics of
Degree, Betweenness Centrality (BC) and Closeness Centrality (CC),
the PPIs were visualized. A positive correlation between these
metrics and the therapeutic efficacy of target genes on RCC was
observed, indicating a stronger influence of genes with higher
values on the therapeutic potential.
Kyoto Encyclopedia of Genes and
Genomes (KEGG) and Gene Ontology (GO) analysis
The high-impact target genes were inputted into the
Sandbox database (http://www.sangerbox.com/home.html), with human
subjects selected, a minimum overlap value of 3, a significance
threshold of P<0.05, and a minimum enrichment threshold of 1.5.
Subsequently, KEGG pathway enrichment and GO function analyses were
conducted, where the top-ranked KEGG and GO data were summarized
and visualized through histogram analysis (18).
Molecular docking
The PDB file of the high-impact target gene was
downloaded from the database (https://www.rcsb.org/) and Discovery Studio (v 1.21)
was used to visualize its ligand and water molecule configuration.
Subsequently, ChemDraw 3D software (2020) was used to open the
compound structure and execute energy minimization to produce the
MOL2 file format. AutoDock software (1.5.7) was then used for
docking the compound and high-impact target genes to assess the
binding free energy, where a greater absolute value signifies a
stronger interaction (18,19).
Molecular dynamics simulations
Following molecular docking, the top-performing
complexes were chosen for kinetic simulation and visualization.
Protein FASTA sequences were retrieved from the RCSB PDB and
modeled using Swiss-Model (https://swissmodel.expasy.org/). The protein structure
was refined by filling in missing data and adding hydrogen atoms
via Protein Plus (https://g6altair.sci.hokudai.ac.jp/g6/service/pocasa/).
Topology files were generated using ACPYPE (2022.1.3) for molecular
dynamics simulations. GROMACS software (2024.2) was employed with
the AMBER 14SB force field and TIP3P water model, ensuring a
minimum distance of 1.2 nm from the protein edges to the box
boundaries.
Cell viability assay
The cell lines 769-P, 786-0, ACHN and HK-2 were
procured from the National Collection of Authenticated Cell
Cultures. Cells underwent STR certification and were stored in a
humidified environment at 37˚C with 5% CO2. The cells
were treated with varying concentrations of PSO (0, 2, 4, 8, 16 µM)
to induce stabilization, followed by assessment of cell viability
using the Cell Counting Kit-8 (CCK-8) assay post drug
administration. Cells in the logarithmic growth phase
(5x104 cells/ml) were seeded into a 96-well plate at 100
µl per well. Subsequently, 10 µl of test substances at varying
concentrations were added to each well for a 24-h treatment period.
Following this, 10 µl of CCK-8 solution was introduced to each
well, and the plate was returned to the incubator for an additional
2 h. Absorbance (OD value) was then measured at 450 nm using a
microplate reader, and cell survival rates at different
concentrations, along with the IC50 value, were
calculated.
Transcriptomic analysis
Tumor tissues underwent standard preprocessing
procedures to prepare them for transcriptome sequencing in
subsequent investigations. RNA quality in skin samples was assessed
using an Agilent 2100 bioanalyzer. Ribosomal RNA (rRNA) removal,
library construction, and Illumina sequencing were conducted by
Beijing Novogene Bioinformatics Technology Co., Ltd. Bowtie
software (v2.5.0) was utilized to eliminate rRNA sequences from the
raw sequencing reads, using the mouse-derived rRNA reference
sequence as the index. The mem algorithm of BWA software
(v0.7.17-r1188) aligned the filtered clean reads with the mouse
reference genome (Mus Musculus (GRCm38/mm10)). SAM format files
were converted to BAM format using Samtools software (v1.8),
followed by sorting and indexing. FeatureCounts software (v2.0.1)
counted the uniquely aligned reads to determine the raw expression
levels of each gene. Differentially expressed genes were then
identified using R language, following the standard differential
expression analysis workflow. For small sample sizes, DESeq2
(v1.34.0) was employed, while the Wilcoxon rank-sum test was used
for larger sample sizes. Genes were considered differentially
expressed if they had a corrected P-value <1.0. Initially, reads
exhibiting low-quality base ratios exceeding 40%, comprising over
1% unknown bases, and showing contamination from adapter sequences
were eliminated using SOAPnuke (2024.2). DESeq2 was employed to
pinpoint differentially expressed genes with Fold Change (FC) >2
or 1.0 and P<0.05.
Statistical analysis
Data analysis was conducted using SPSS 22.0 software
(IBM Corp.). Mean values with standard deviations were utilized to
represent data variability. Group comparisons were performed using
one-way paired t-tests. P<0.05 was considered to indicate a
statistically significant difference.
Results
Structural determination
The structure of the compound was elucidated using
various chromatographic techniques, where its NMR data are
presented as follows: 1H NMR (400 MHz, DMSO-d6)
δH 8.17 (d, J=9.6 Hz, 1H, H-4), 8.11 (d,
J=2.3 Hz, 1H, H-2'), 8.00 (s, 1H, H-6), 7.72 (s, 1H, H-8),
7.10 (dd, J=2.3, 0.9 Hz, 1H, H-3'), 6.43 (d, J=9.6
Hz, 1H, H-3) (Fig. S1).
13C NMR (100 MHz, DMSO-d6) δC
160.2(C-2), 155.7(C-7), 151.5(C-9), 147.9(C-2'), 145.0(C-4),
124.6(C-6), 120.7(C-5), 115.4(C-10), 114.1(C-3), 106.8(C-3'),
99.3(C-8) (Fig. S2). A literature
review confirmed the compound as PSO (20), a natural furanocoumarin initially
isolated from Psoralea corylifolia.
Target gene prediction
Prediction of PSO using the Swiss Target Prediction
database identified 162 potentially relevant genes. Searching for
genes related to RCC yielded 3,687 genes from GeneCards, 179 genes
from DisGeNET, 200 genes from OMIM, 30 genes from DrugBank and 54
genes from the TTD database. After removing duplicates, a total of
3,923 genes were predicted. The intersection of these datasets
revealed 78 target genes associated with RCC (Fig. S3).
PPI network analysis
A PPI network diagram was constructed based on the
78 predicted target genes, where the top 20 genes were ranked
according to Degree, BC and CC metrics. The results, presented in
Table I, revealed that the top
five genes were ESR1 (Degree=41, BC=0.1679 and CC=0.6788), HSP90AB1
(Degree=38, BC=0.118611 and CC=0.6666), NFKB1 (Degree=38, BC=0.1186
and CC=0.6666), NFKB1 (Degree=38, BC=0.0552 and CC=0.6491) and
STAT1 (Degree=31, BC=0.0338 and CC=0.5967).
 | Table IGene names, degree value, betweenness
centrality and closeness centrality of key targets. |
Table I
Gene names, degree value, betweenness
centrality and closeness centrality of key targets.
| Gene name | Betweenness
centrality | Closeness
centrality | Degree |
|---|
| ESR1 | 0.167942 | 0.678899 | 41 |
| HSP90AB1 | 0.118611 | 0.666667 | 38 |
| NFKB1 | 0.055249 | 0.649123 | 38 |
| STAT1 | 0.033857 | 0.596774 | 31 |
| GSK3B | 0.052042 | 0.611570 | 31 |
| TLR4 | 0.055006 | 0.606557 | 31 |
| PARP1 | 0.043182 | 0.587302 | 28 |
| PIK3CA | 0.046844 | 0.573643 | 27 |
| RELA | 0.015787 | 0.578125 | 26 |
| ICAM1 | 0.028730 | 0.564885 | 24 |
| JAK2 | 0.028848 | 0.544118 | 23 |
| NFE2L2 | 0.015432 | 0.564885 | 23 |
| NFKBIA | 0.018060 | 0.560606 | 22 |
| MAPK14 | 0.043738 | 0.544118 | 22 |
| PGR | 0.013787 | 0.540146 | 21 |
| RPS6KB1 | 0.006633 | 0.536232 | 20 |
| AR | 0.007943 | 0.540146 | 20 |
| PPARA | 0.041333 | 0.536232 | 19 |
| ESR2 | 0.034383 | 0.532374 | 17 |
| HDAC2 | 0.018560 | 0.503401 | 16 |
KEGG and GO analysis
After analyzing 78 potential target genes using KEGG
pathway enrichment in the database, a total of 131 signaling
pathways were identified. The top 10 enriched KEGG pathways
included ‘Pathways in cancer’, ‘Metabolic pathways’, ‘Kaposi
sarcoma-associated herpesvirus infection’, ‘Hepatitis B’,
‘Chemokine signaling pathway’, ‘PI3K-Akt signaling pathway’, ‘Human
cytomegalovirus infection’, ‘Prolactin signaling pathway’,
‘Epstein-Barr virus infection’ and ‘cAMP signaling pathway’
(Fig. S4).
The 78 potential target genes underwent analysis
using the GO Functional Enrichment Database, yielding 267 molecular
function (MF) results, 98 cellular component (CC) results, and
2,134 biological process (BP) results. The top 5 MF results
included ‘enzyme binding’, ‘catalytic activity’, ‘protein kinase
activity’, ‘phosphotransferase activity’ and ‘alcohol group as
acceptor’. The top 5 CC results consisted of ‘cytosol’, ‘cell
surface’, ‘whole membrane’, ‘nucleoplasm’ and ‘plasma membrane
part’. The top 5 BP results were ‘response to organic substance’,
‘response to chemical’, ‘response to oxygen-containing compound’,
‘cellular response to chemical stimulus’ and ‘regulation of
biological quality’ (Fig.
S5).
Molecular docking
The protein genes with high degree values within the
PI3K/Akt/mTOR signaling pathway, crucial in the therapeutic process
of RCC, were chosen for molecular docking simulations with PSO to
assess their binding capabilities. Selected proteins included heat
shock protein (HSP)90AB1 (PDB ID: 5UC4), NFKB1 (PDB ID: 1U3Y),
glycogen synthase kinase (GSK)3B (PDB ID: 6AE3), toll-like receptor
(TLR)4 (PDB ID: 5UCA), PIK3CA (PDB ID: 9B4T), RELA (PDB ID: 1VJ7),
Janus kinase (JAK)1 (PDB ID: 6GGH) and JAK2 (PDB ID: 7F7W). The
docking results revealed binding energies of -6.14 kcal/mol with
5UC4, -5.226 kcal/mol with 1U3Y, -7.33 kcal/mol with 6AE3, -0.198
kcal/mol with 5UCA, -7.47 kcal/mol with 9B4T, -7.07 kcal/mol with
1VJ7, -6.514 kcal/mol with 6GGH and -6.18 kcal/mol with 7F7W. The
complex with the highest absolute docking binding energies was
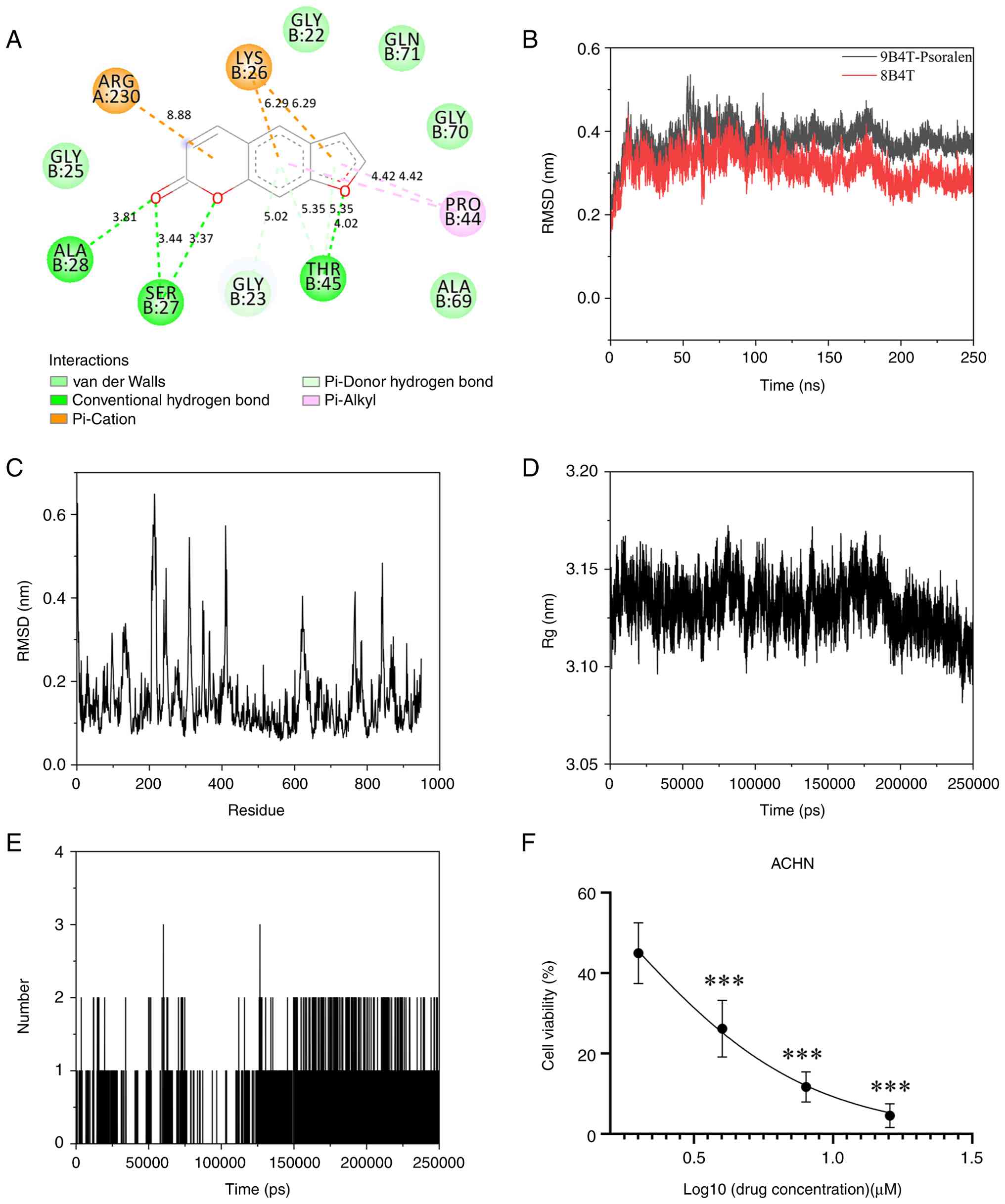
chosen for in-depth molecular docking analysis (Fig. 1A).
Molecular dynamics simulations
PSO with 9B4T kinetic simulation preprocessing was
employed, modeling the system with dimensions of
13.711x13.711x13.711 nm, each angle at 90.00˚ and a volume of
2,577.47 nm³. The final configuration consisted of 254,810 atoms
distributed over 80,730 residues, with a solvent density of
1,001.99 g/l and 79,781 solvent molecules. Sodium ions were
introduced to maintain charge balance, with a salt solution
concentration of 0.15 mol/l to mimic physiological conditions.
Electrostatic interactions were computed using Particle-mesh Ewald
for energy minimization over 50,000 steps. The system was simulated
in canonical ensemble and isothermal-isobaric ensemble states at
310 K and 1 bar pressure. Molecular dynamics simulations were
conducted for 250 ns under these conditions, with a non-bonded
interaction cutoff of 10 Å. The simulation results were evaluated
based on root mean square deviation (RMSD), root mean square
fluctuation (RMSF), radius of gyration (RG) and hydrogen bond
energy.
The RMSD of the PSO/9B4T complex was comparable to
but slightly lower compared with that of the unbound protein,
indicating reduced structural deviation and enhanced stability
(Fig. 1B). PSO/9B4T maintains
lower and less fluctuating RMSF values, indicating higher
structural stability and suggesting a more stable structure for the
PSO/9B4T complex (Fig. 1C). The RG
of the PSO/9B4T complex fluctuated within a narrow range (3.11-3.16
nm), with an average value of ~3.13 nm, indicating a compact and
stable complex throughout the simulation (Fig. 1D). In the PSO/9B4T complex, the
majority of hydrogen bonds were concentrated between 1 and 2,
indicating a relatively stable state of low-frequency hydrogen
bonding throughout most time points (Fig. 1E). Quantitative analysis showed
that the average RMSD of the PSO-PIK3CA complex was lower compared
with that of the unbound protein during the equilibrium phase,
indicating enhanced structural stability. The average RG of the
complex remained stable at ~3.13 nm, with smaller fluctuations
compared with the apo protein, further supporting the formation of
a stable complex.
Cell viability test results
Initially, the toxic impact of PSO was assessed on
normal cells. Various doses of PSO were administered to normal
renal epithelial cells (HK-2), where cell viability was assessed
using the CCK-8 assay. The results indicated that PSO exhibited no
significant toxicity towards HK-2 cells within the 0-8 µM range,
instead demonstrating slight cytotoxicity at a higher concentration
of 16 µM. PSO treatment resulted in the significantly decreased
cellular viability of 769-P and 786-0 cells (IC50 values
of 6.02±0.88 and 6.26±0.96 µM, respectively). Furthermore, PSO
significantly decreased the cellular viability of ACHN cells, with
an IC50 of 1.74±0.90 µM (Fig. 1F).
Transcriptomics results
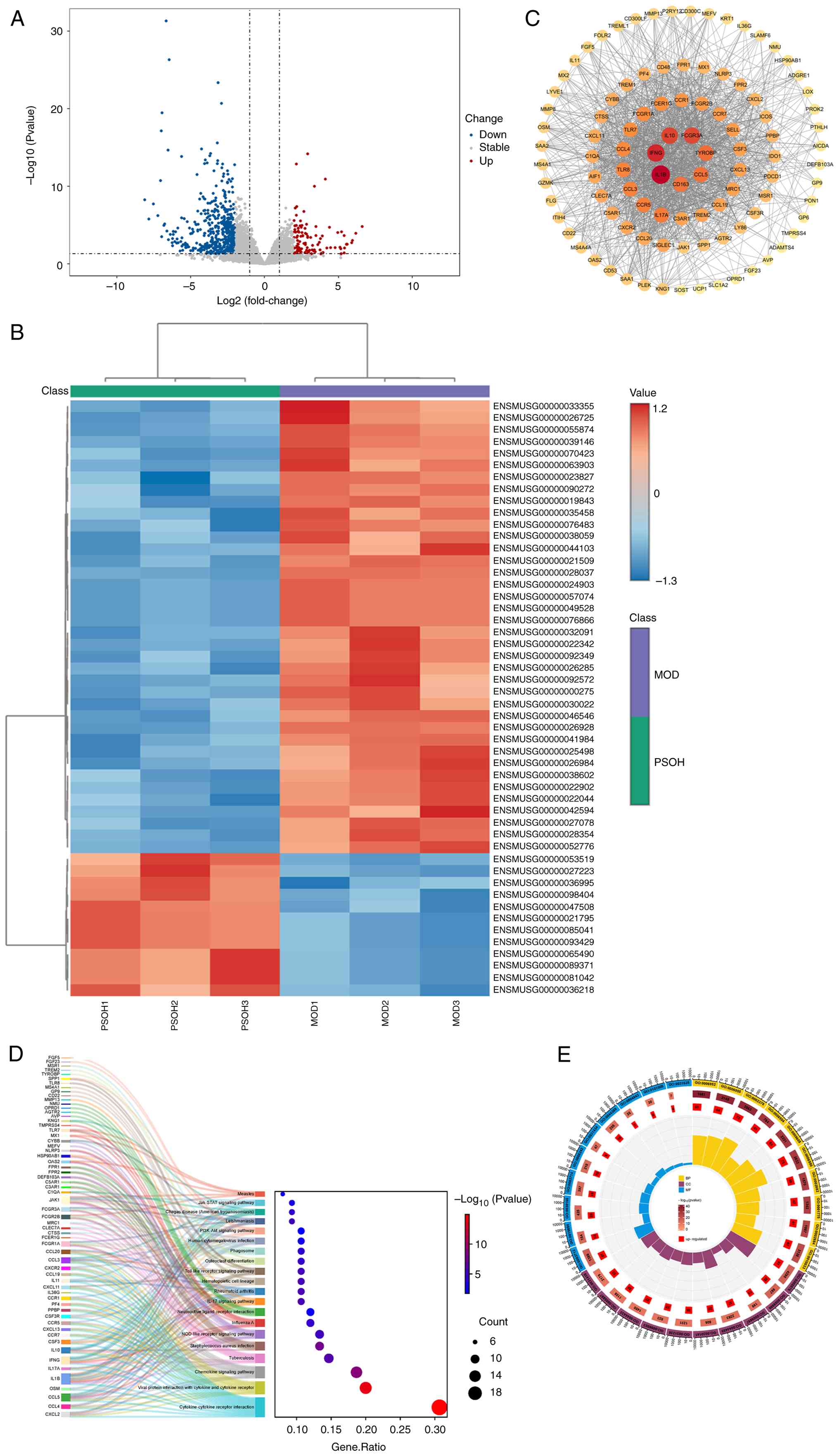
Transcriptomics analysis was utilized to
comprehensively assess genetic changes in tumor cells before and
after drug administration. Comparison of gene expression
differences between MOD and PSO groups through volcano and heat map
analyses (Fig. 2A and B) revealed
112 upregulated genes and 481 downregulated genes. Due to the
substantial number of differential genes, PPI analysis was
conducted using the STRING database to identify potential target
genes. Selection criteria included Degree ≥7, BC ≥0.0019 and CC
≥0.2976. Ultimately, 100 significant target genes were identified
and visualized based on their degree values (Fig. 2C). KEGG pathway enrichment analysis
of these target genes identified 48 highly enriched pathways, with
the top 20 pathways including ‘cytokine-cytokine receptor
interaction’, ‘chemokine signaling pathway’, ‘tuberculosis’ and
others (Fig. 2D). GO functional
analysis visualized the top 10 results for MF, CC and BP (Fig. 2E). Transcriptomics pathway
enrichment results indicated significant downregulation of OSM,
FGF23, FGF5 and other genes in the PI3K/Akt pathway (P<0.001),
suggesting differential changes at the transcriptome level and
supporting the pathway's role in the therapeutic process.
Discussion
PSO is a bioactive furanocoumarin that can be
isolated from TX and has been reported to exhibit broad antitumor
activities in multiple cancer types. Previous studies have
demonstrated that PSO can suppress tumor cell proliferation by
inducing apoptosis in hepatocellular carcinoma, breast cancer,
gastric cancer and osteosarcoma through various mechanisms,
including DNA damage, inhibition of DNA polymerase and
topoisomerase II, endoplasmic reticulum stress and regulation of
microRNA-associated signaling pathways (17-19).
However, these studies primarily focused on tumor types outside the
renal system, where the molecular mechanisms underlying PSO
activity in RCC remained largely unexplored.
In the present study, the anti-RCC effects of PSO
were systematically investigated using an integrated multi-omics
strategy. Compared with previous studies in other cancer models,
the present findings highlight a distinct but convergent mechanism
in RCC, characterized by suppression of the PI3K/AKT signaling
pathway (20). Network
pharmacology and transcriptomic analyses consistently identified
PI3K/AKT-related genes as central nodes in PSO-mediated regulation,
whilst molecular docking and molecular dynamics simulations further
supported a stable interaction between PSO and PIK3CA.
The PI3K/AKT signaling pathway is a well-established
driver of RCC progression, regulating cell proliferation, survival,
metabolism and therapeutic resistance. Natural products targeting
this pathway have shown promise in RCC treatment. Hemsleya
amabilis Diels extract has been shown to induce G2/M cell cycle
arrest and apoptosis in RCC models through the PI3K/AKT pathway
inhibition, effects that were reversed by PI3K inhibition (21). Similarly, gypenosides from
Gynostemma pentaphyllum was observed to suppress RCC cell
proliferation and promoted apoptosis by downregulating PI3K, AKT
and mTOR expression (22).
Consistent with these studies, the results of the present study
demonstrated that PSO significantly downregulates PI3K/AKT
pathway-related genes and inhibits RCC cell proliferation,
positioning PSO as a natural PI3K/AKT-modulating agent in RCC.
Based on these findings, a schematic mechanism was
proposed in which PSO interacts with PIK3CA, leading to the
suppression of PI3K/AKT signaling, disruption of downstream
survival and proliferative signals, ultimately inhibiting RCC tumor
growth. This integrated mechanism bridges computational predictions
with experimental validation and provides a coherent framework for
understanding anti-RCC activity of PSO.
In summary, the present study systematically
elucidated the anti-RCC effects of PSO derived from TX using an
integrated multi-omics approach. The present findings demonstrated
that PSO exerts significant antiproliferative effects on RCC cells
primarily through suppression of the PI3K/AKT signaling pathway,
supported by network pharmacology, molecular docking, molecular
dynamics simulations, transcriptomics and in vitro
validation.
These results not only extend the antitumor spectrum
of PSO to RCC but also highlight its potential as a natural
PI3K/AKT pathway modulator. Given the clinical importance of
PI3K/AKT signaling in RCC progression and therapeutic resistance,
PSO may represent a promising lead compound for the development of
cost-effective and mechanism-based therapeutic strategies for RCC.
Further preclinical and clinical investigations are warranted to
explore its translational potential.
Supplementary Material
1H nuclear magnetic
resonance spectra of psoralen.
13C nuclear magnetic
resonance spectra of psoralen.
Intersection plot, blue area
represents the number of genes implicated by psoralen and the red
area represents the number of genes associated with RCC. RCC, renal
cell carcinoma.
Mulberry plot of top ten pathways
identified in Kyoto Encyclopedia of Genes and Genomes enrichment
analysis.
Top ten results from Gene Ontology
functional analysis graph.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the National Natural
Science Foundation of China (grant no. 32200314), the Ningbo Health
Technology Project (grant no. 2022Y18) and the Research Project of
Zhejiang Chinese Medical University (grant no. 2022FSYYZQ19) and
the Ningbo Natural Science Foundation (grant no. 2024J261).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
DC and TC made substantial contributions to
conception and design; acquisition, analysis and interpretation of
data and were involved in drafting or revising the manuscript
critically for important intellectual content. TC and AW confirm
the authenticity of all the raw data and provided final approval
for publication. DC, TC, YY and AW participated sufficiently in the
work to take public responsibility for appropriate portions of the
content and agreed to be accountable for all aspects of the work in
ensuring that questions related to the accuracy or integrity of any
part of the work are appropriately investigated and resolved. All
authors read and approved the final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Chen YW, Wang L, Panian J, Dhanji S,
Derweesh I, Rose B, Bagrodia A and McKay RR: Treatment landscape of
renal cell carcinoma. Curr Treat Options Oncol. 24:1889–1916.
2023.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Nezami BG and MacLennan GT: Clear cell
renal cell carcinoma: A comprehensive review of its histopathology,
genetics, and differential diagnosis. Int J Surg Pathol.
33:265–280. 2025.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Wolf MM, Kimryn Rathmell W and Beckermann
KE: Modeling clear cell renal cell carcinoma and therapeutic
implications. Oncogene. 39:3413–3426. 2020.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Siva S, Bressel M, Sidhom M, Sridharan S,
Vanneste BGL, Davey R, Montgomery R, Ruben J, Foroudi F, Higgs B,
et al: Stereotactic ablative body radiotherapy for primary kidney
cancer (TROG 15.03 FASTRACK II): A non-randomised phase 2 trial.
Lancet Oncol. 25:308–316. 2024.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Wang L, Zong L, Cao D and Guan M:
Unraveling the molecular mechanisms of PFOA in clear cell renal
cell carcinoma through network toxicology and molecular docking
strategies. Int J Surg. 111:4842–4853. 2025.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Jin J, Xie Y, Zhang JS, Wang JQ, Dai SJ,
He WF, Li SY, Ashby CR Jr, Chen ZS and He Q: Sunitinib resistance
in renal cell carcinoma: From molecular mechanisms to predictive
biomarkers. Drug Resist Updat. 67(100929)2023.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Song K, Li M, Yang Y, Zhang Z, Zhang J,
Zhu Q, Liu J and Wang A: Trigonostemon species in south China:
Insights on its chemical constituents towards pharmacological
applications. J Ethnopharmacol. 281(114504)2021.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Wang A, Yang Y, Chen T, He J, Ling B and
Cheng X: Antiproliferative effects of Trigonostemon
xyphophyllorides on renal cell carcinoma via the PI3K/AKT
pathway. Front Pharmacol. 16(1594461)2025.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Ban NK, Linh TM, Mai NC, Tai BH, Nhiem NX,
Hoang NH and Kiem PV: New 3,4-seco-diterpene and coumarin
derivative from the leaves of Trigonostemon flavidus Gagnep.
Nat Prod Res. 36:3247–3254. 2022.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Ban NK, Truong LH, Tiep TV, Yen DTH, Doan
VV, Nhiem NX, Seo Y, Namkung W, Kim SH, Tai BH and Kiem PV: Four
new sucrose diesters of substituted truxinic acids from
trigonostemon honbaensis with their anoctamin-1 inhibitory
activity. Bioorg Chem. 102(104058)2020.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Cho YR, Park K, Kang JS, Byun HW, Oh JS,
Seo DW and Ahn EK: Trigonostemon reidioides modulates
endothelial cell proliferation, migration and tube formation via
downregulation of the Akt signaling pathway. Oncol Lett.
14:4677–4683. 2017.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Liu YP, Hu S, Wen Q, Ma YL, Jiang ZH, Tang
JY and Fu YH: Novel γ-lactone derivatives from trigonostemon
heterophyllus with their potential antiproliferative activities.
Bioorg Chem. 79:107–110. 2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Liu YP, Wen Q, Hu S, Ma YL, Jiang ZH, Tang
JY and Fu YH: Structurally diverse diterpenoids from trigonostemon
howii. Nat Prod Res. 33:1169–1174. 2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Sun B, Shen K, Zhao R, Li Y and Lin J:
Clarithromycin attenuates airway epithelial-mesenchymal transition
in ovalbumin-induced asthmatic mice through modulation of Kv1.3
channels and PI3K/Akt signaling. Int Immunopharmacol.
139(112624)2024.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Chen S, Guo W, Liu H, Zheng J, Lu D, Sun
J, Li C, Liu C, Wang Y, Huang Y, et al: Mechanistic study of
cytochrome P450 enzyme-mediated cytotoxicity of psoralen and
isopsoralen. Food Chem Toxicol. 180(114011)2023.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Takeda J, Iwao Y, Karashima M, Yamamoto K
and Ikeda Y: Structural evaluation of the choline and geranic
acid/water complex by SAXS and NMR analyses. ACS Biomater Sci Eng.
7:595–604. 2021.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Wang Y, Zhang J, Xu S, Li J, Liu W, Jiang
M and Bai G: Psoralen alleviates acute lung injury by covalently
targeting Cys106 of HMGB1 in macrophages to inhibit inflammatory
responses. Phytomedicine. 142(156807)2025.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Li X, Miao F, Xin R, Tai Z, Pan H, Huang
H, Yu J, Chen Z and Zhu Q: Combining network pharmacology,
molecular docking, molecular dynamics simulation, and experimental
verification to examine the efficacy and immunoregulation mechanism
of FHB granules on vitiligo. Front Immunol.
14(1194823)2023.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Lu S, Sun X, Zhou Z, Tang H, Xiao R, Lv Q,
Wang B, Qu J, Yu J, Sun F, et al: Mechanism of bazhen decoction in
the treatment of colorectal cancer based on network pharmacology,
molecular docking, and experimental validation. Front Immunol.
14(1235575)2023.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Chang J, Hu P, Zhang B, Liu Y, Cheng Y and
Li L and Li L: Perilla seed oil alleviates high-fat-diet-induced
hyperlipidemia by regulating fatty acid metabolism via the
PI3K/Akt/NOS3 pathway. Foods. 14(4125)2025.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Chen LM, Qian ST, Li ZQ, He MF and Li HJ:
Psoralen and isopsoralen, two estrogen-like natural products from
psoraleae fructus, induced cholestasis via activation of ERK1/2.
Chem Res Toxicol. 37:804–813. 2024.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Shen Z, Gao X, Huang D, Xu X and Shen J:
The potential of gynostemma pentaphyllum in the treatment of
hyperlipidemia and its interaction with the LOX1-PI3K-AKT-eNOS
pathway. Food Sci Nutr. 12:8000–8012. 2024.PubMed/NCBI View Article : Google Scholar
|