Introduction
Sebaceous carcinoma (SGC) of the eyelid is a
malignant tumor arising from the meibomian or Zeiss glands
(1). SGC of the eyelid is highly
prevalent in Asia and represents the most common eyelid malignancy
(accounting for 43.7% of cases in Japan) (2). The tumor is generally resistant to
radiotherapy and chemotherapy, with complete surgical resection
remaining the mainstay of treatment. However, postoperative local
recurrence and lymph node metastasis have been reported in 6-24 and
8-21% of patients, respectively, reflecting a relatively high-grade
malignancy (3-6).
Adjuvant and combination therapies may improve prognosis; however,
the pathogenesis of SGC of the eyelid remains unexplored, and
sufficient research to guide diagnosis and treatment has not been
performed.
Several clinical and pathological factors have been
identified as poor prognostic indicators and risk factors for
postoperative recurrence in SGC of the eyelid. Clinical factors
include increased tumor diameter (6-8),
tumor localization (6,9), and delayed therapeutic intervention
due to misdiagnosis at initial presentation (10). Pathologic factors include diffuse
growth patterns, multicentric tumor origin, non-lobular pattern,
and poor differentiation (11-14).
In addition, pagetoid spread is a critical poor prognostic factor.
It is characterized by diffuse intraepithelial dissemination and a
tendency to invade the cornea and conjunctiva, complicating
complete surgical excision. Furthermore, it is frequently
misdiagnosed as chronic blepharoconjunctivitis, contributing to the
unfavorable prognosis (12,15-21).
Recent advances in genetic analyses have provided
insights into the biological pathways involved in the tumor
microenvironment and pathogenesis of SGC. Whole-genome studies
(22-25)
and microRNA profiling (26-28)
have been performed to elucidate tumor biology. These studies have
revealed recurrent genetic alterations and dysregulated miRNA
expression profiles associated with tumor proliferation, invasion,
and immune-related pathways in SGC.
Increasing evidence suggests that
immune-inflammatory pathways play a critical role in tumor invasion
and progression by shaping the tumor microenvironment (29,30).
In various malignancies, dysregulated immune responses and
immune-related signaling pathways have been implicated in
epithelial-mesenchymal transition, tumor invasiveness, and
metastatic potential (29,30). In sebaceous carcinoma, recent
genomic and transcriptomic studies have identified dysregulated
immune- and inflammation-related pathways associated with
aggressive tumor behavior and invasion (22-28).
These findings suggest that immune-inflammatory mechanisms may
contribute to the invasive behavior of SGC and provide a biological
rationale for investigating immune-related molecular networks in
this disease.
In recent years, integrated bioinformatic approaches
combining miRNA and mRNA expression data have been widely applied
to various malignancies to elucidate regulatory networks underlying
tumor invasiveness and therapeutic resistance, as well as to
identify potential diagnostic and prognostic biomarkers (29,30).
Despite these advances, controlling local recurrence
and lymph node metastasis remains challenging, and clinical
outcomes are often suboptimal. In particular, limited genetic
research has focused on the pathogenesis of pagetoid spread, a
major determinant of poor prognosis in SGC.
Our previous study published in 2021(28) primarily focused on global miRNA
expression profiling in eyelid sebaceous carcinoma. In contrast,
the present study uniquely applies an integrated miRNA-mRNA network
analysis to specifically explore the molecular mechanisms
underlying pagetoid spread, with particular emphasis on
invasion-related and immune-inflammatory pathways.
Therefore, this study aimed to elucidate the genetic
mechanisms involved in SGC of the eyelid with pagetoid spread.
Materials and methods
Patient and tissue samples
Nine patients who underwent surgical treatment for
eyelid tumors at Toyama University Hospital were included in this
study. These patients underwent surgical treatment between January
2020 and October 2024. Cases were selected based on the
availability of sufficient tumor tissue for molecular analyses and
a confirmed pathological diagnosis by histopathological
examination. Based on pathological findings, the patients were
divided into three groups, each comprising three participants: SGC
of the eyelid with pagetoid spread, SGC of the eyelid without
pagetoid spread, and normal controls. The pagetoid spread group
included an 88-year-old woman, a 74-year-old woman, and an
81-year-old man. The maximum tumor diameters in this group were 6,
7, and 8 mm, respectively, and all tumors were classified as T1
according to the American Joint Committee on Cancer (AJCC) staging
system (8th edition). Regarding histopathological growth patterns,
one case in the pagetoid spread group (a 74-year-old woman) showed
a diffuse growth pattern, whereas the remaining two cases showed a
nodular growth pattern. The non-pagetoid spread group consisted of
an 84-year-old woman, a 69-year-old woman, and an 80-year-old man.
The maximum tumor diameters and T classifications in this group
were 10 mm (T1), 6 mm (T1), and 11 mm (T2b), respectively. All
cases in the non-pagetoid spread group exhibited a nodular growth
pattern. No cases in either group showed evidence of multicentric
origin based on clinicopathological evaluation. Normal control
samples were obtained from a 72-year-old woman, a 74-year-old
woman, and a 77-year-old man, from whom normal tarsal plate tissue
was collected during free tarsal plate reconstruction. This study
was approved by the Institutional Review Board of the University of
Toyama (approval no. R2015051), and all procedures were conducted
in accordance with the tenets of the Declaration of Helsinki.
Written informed consent was obtained from all patients prior to
enrollment.
RNA extraction and quality
control
Total RNA was extracted from whole tumor tissue
samples stored at -80˚C after surgical excision, without prior
formalin fixation. Tumor-predominant areas were selected as much as
possible based on pathological evaluation; however, RNA was
extracted from whole tissue sections without manual
macrodissection, and adjacent non-neoplastic tissues may have been
partially included. Total RNA, including miRNA, was extracted from
tissue samples using the NucleoSpin miRNA kit (Macherey-Nagel GmbH
& Co., Düren, Germany) according to the manufacturer's
protocol. RNA quality and quantity were assessed using a
Bioanalyzer 2100 system with the RNA 6000 Nano kit (Agilent
Technologies, Santa Clara, CA, USA) (31).
Library preparation and RNA sequencing
(mRNA and miRNA)
Comprehensive mRNA and miRNA analyses were
subcontracted to OligoAtenta, Inc. (Tokyo, Japan) and performed
using their next-generation sequencing (NGS) service. For library
preparation, Illumina-compatible kits were used, and sequencing was
conducted with either paired-end or single-end reads on the NovaSeq
6000 system (Illumina, San Diego, CA, USA).
For mRNA analysis, mRNA was selectively extracted
and purified from total RNA. Sequencing libraries were subsequently
prepared and analyzed using 150 bp paired-end reads. The resulting
FASTQ data were processed using a standard pipeline provided by
OligoAtenta, which included adapter removal, quality assessment,
read mapping, and expression quantification. Reads were mapped to
the human reference genome (GRCh38), and gene expression levels
were normalized using transcripts per million (TPM) or fragments
per kilobase of transcript per million mapped reads.
In the miRNA analysis, small RNA libraries were
prepared, and reads of 83 nucleotides in length were obtained. Each
read contained a unique molecular identifier (UMI) and a 3' adapter
sequence, which were automatically trimmed during data processing.
Clean reads were mapped to miRbase, and UMI-based deduplication was
performed to obtain expression data normalized by reads per
million. Additional processing steps included adapter removal (N)
using Trim Galore, genome mapping with Bowtie2, file conversion via
Samtools, and annotation analysis using Strand NGS software. The
raw RNA sequencing data generated in the present study have been
deposited in the DNA Data Bank of Japan (DDBJ) Sequence Read
Archive under BioProject accession number PRJDB40390.
Raw read data processing
The first 50 bp sequences of each raw 150 bp
sequence read were extracted using Seqkit. Adaptor sequences were
subsequently trimmed from the 50 bp reads using Cutadapt.
Low-quality (Q score <20) and short reads (<10 bp) were
removed using the FASTX-Toolkit. The filtered reads were then
aligned to hg19, and miRNA annotation was performed using Strand
NGS version 3.3.
Integrated miRNA-mRNA interaction
analyses
To examine the molecular functions and interaction
networks of expressed miRNA and mRNA, combined data from the
present study and previously published datasets generated by our
group were analyzed using Ingenuity Pathways Analysis (IPA)
software (Ingenuity Systems, Redwood City, CA, USA).
Results
Identification of expressed miRNAs and
mRNAs
Before molecular analyses, clinicopathological
characteristics were reviewed to confirm the comparability of the
study groups. All pagetoid spread cases were classified as
pathological T1 according to the AJCC staging system (8th edition);
one case exhibited a diffuse growth pattern, whereas all remaining
cases in both the pagetoid and non-pagetoid groups showed a nodular
growth pattern, and no case in either group showed evidence of
multicentric origin. A total of 280,241,626 raw reads were obtained
for the entire SGC of the eyelid dataset, with each sample yielding
at least 20 million reads. This sequencing depth was sufficient to
ensure reliable miRNA and mRNA expression analyses. Quality
assessment was performed on the raw sequencing data. Low-quality
reads and adapter sequences were removed from the datasets. For
miRNA analysis, the filtered reads were aligned to the human
miRBase using Strand NGS and mapped to known miRNAs. For mRNA
analysis, quality-filtered reads were similarly mapped to the human
reference genome (GRCh38) using the STAR aligner. The transcript
expression levels were quantified as TPM. Expression variation and
functional analyses were subsequently performed on both datasets.
miRNAs with at least a 1.5-fold change and mRNAs with at least a
2.0-fold change in expression relative to control samples were
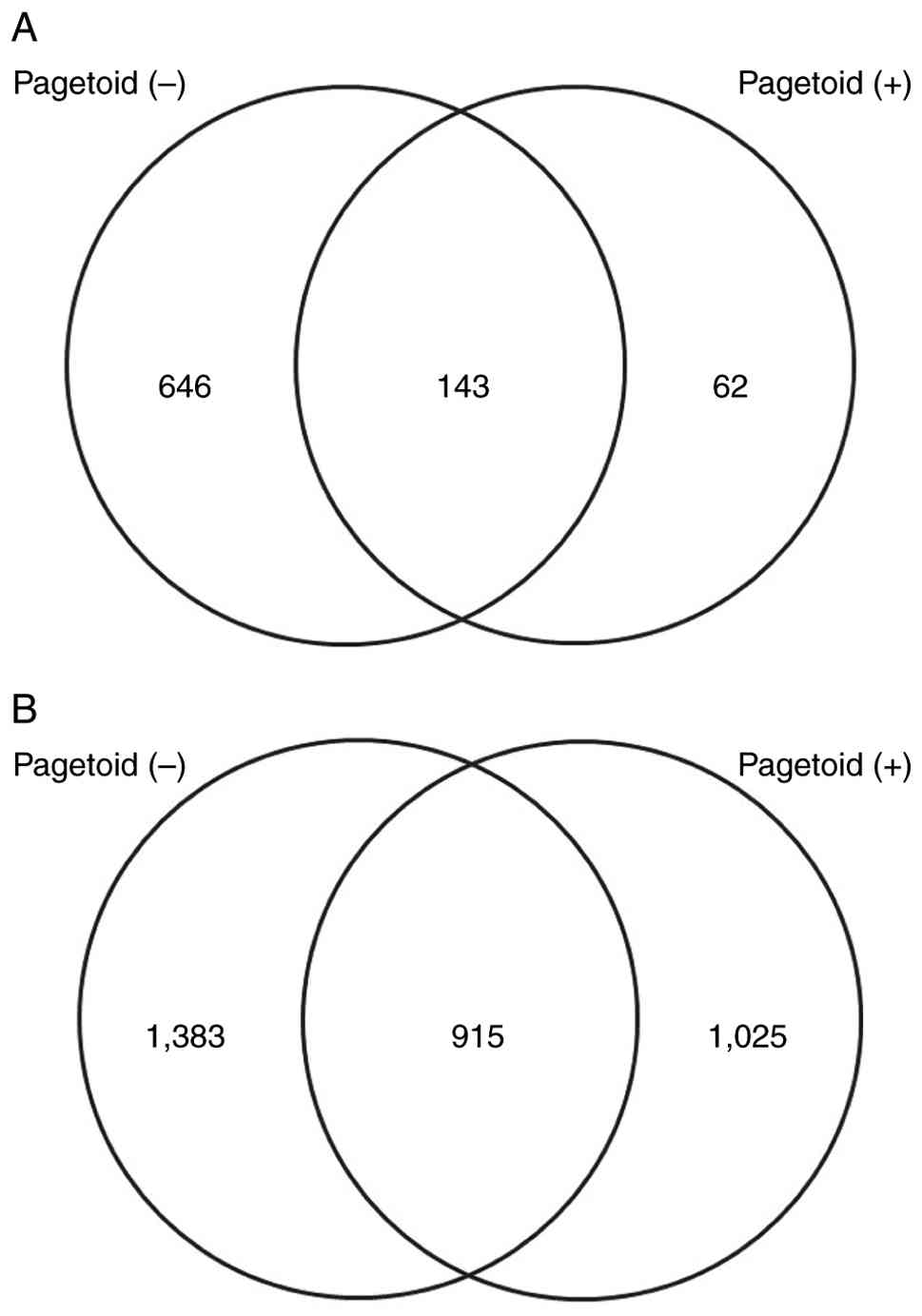
identified. The Venn diagram in Fig.
1 illustrates the number of genes with increased expression.
Fig. 1A presents the number of
upregulated mRNAs, whereas Fig. 1B
depicts the number of downregulated miRNAs known to suppress the
expression of their target genes. Compared with the control group,
789 and 2,205 mRNAs were upregulated in the non-pagetoid and
pagetoid groups, respectively, of which 62 were uniquely
upregulated in the pagetoid group. By contrast, 2,298 and 1,940
miRNAs were downregulated in the non-pagetoid and pagetoid groups,
respectively, of which 1,025 were uniquely downregulated in the
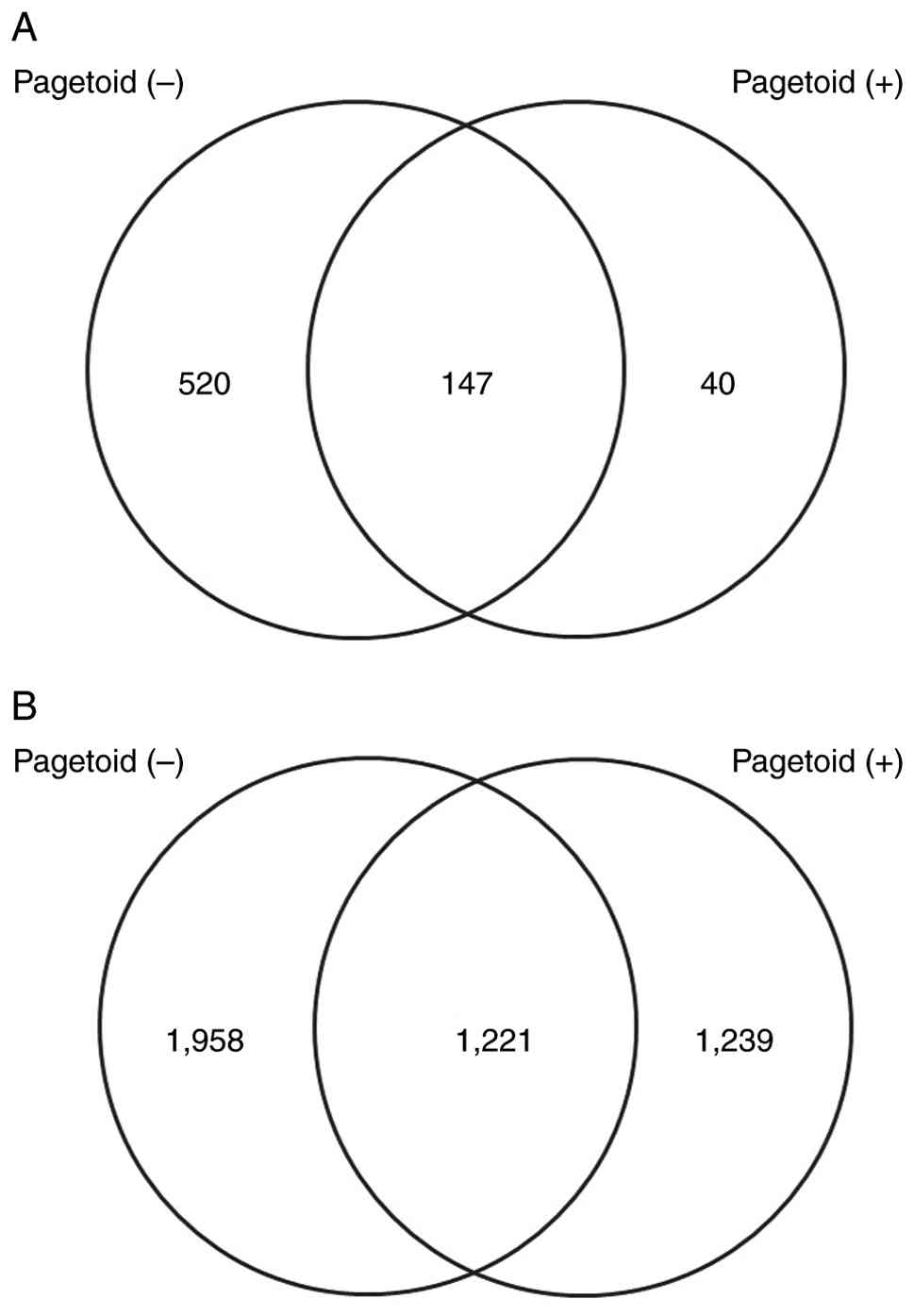
pagetoid group. The Venn diagram in Fig. 2 illustrates the number of genes
with decreased expression. Fig. 2A
and B present the numbers of
downregulated and upregulated mRNAs, respectively. Compared with
the control group, 667 and 187 mRNAs were downregulated in the
non-pagetoid and pagetoid groups, respectively, with 40 uniquely
downregulated in the pagetoid group. Conversely, 3,179 and 2,640
miRNAs were upregulated in the non-pagetoid and pagetoid groups,
respectively, of which 1,239 were uniquely upregulated in the
pagetoid group.
Functional analyses of expressed
miRNAs and mRNAs
To investigate the biological functions and
canonical pathways associated with pagetoid spread, an integrated
miRNA-mRNA dataset comprising 62 upregulated mRNAs and 1,025
downregulated miRNAs specifically identified in the pagetoid group
was analyzed using IPA software. Consequently, biological functions
with positive Z-scores (indicating predicted activation) were
predominantly associated with tumor cell infiltration and migration
(e.g., cell movement, migration, and invasion of tumor cell lines)
and with immune responses (e.g., proliferation and activation of
leukocytes and lymphocytes). The top five biological functions in
each category are presented in Tables
I and II. An integrated
dataset consisting of 40 downregulated mRNAs and 1,239 upregulated
miRNAs specifically observed in the pagetoid group was subsequently
analyzed. IPA analysis predicted the inhibition of biological
functions with negative Z-scores, particularly those related to
lipid metabolism, including lipid synthesis and fatty acid
metabolism. The top five inhibited biological functions are
presented in Table III.
 | Table ITop 5 biological functions (excluding
immune- and inflammation-related functions) associated with
differentially expressed miRNAs (|fold change| ≥1.5) and mRNAs
(|fold change| ≥2.0) in eyelid sebaceous gland carcinoma with
pagetoid spread. |
Table I
Top 5 biological functions (excluding
immune- and inflammation-related functions) associated with
differentially expressed miRNAs (|fold change| ≥1.5) and mRNAs
(|fold change| ≥2.0) in eyelid sebaceous gland carcinoma with
pagetoid spread.
| Functional
annotation | P-value | Predicted
activation state | Activation
z-score | No. of
molecules |
|---|
| Cell movement of
tumor cell lines |
5.03x10-6 | Increased | 5.635 | 54 |
| Migration of tumor
cell lines |
9.67x10-6 | Increased | 5.106 | 50 |
| Invasion of tumor
cell lines |
3.78x10-6 | Increased | 4.761 | 48 |
| Invasion of
cells |
1.93x10-6 | Increased | 4.735 | 53 |
| Cell movement |
1.01x10-11 | Increased | 4.67 | 96 |
 | Table IITop 5 immune- and
inflammation-related biological functions associated with
differentially expressed miRNAs (|fold change| ≥1.5) and mRNAs
(|fold change| ≥2.0) in eyelid sebaceous gland carcinoma with
pagetoid spread. |
Table II
Top 5 immune- and
inflammation-related biological functions associated with
differentially expressed miRNAs (|fold change| ≥1.5) and mRNAs
(|fold change| ≥2.0) in eyelid sebaceous gland carcinoma with
pagetoid spread.
| Functional
annotation | P-value | Predicted
activation state | Activation
z-score | No. of
molecules |
|---|
| Leukopoiesis |
1.58x10-20 | Increased | 4.241 | 58 |
| Cytotoxicity of
leukocytes |
6.6x10-16 | Increased | 4.204 | 23 |
| Activation of
leukocytes |
2.52x10-28 | Increased | 4.19 | 63 |
| Hematopoiesis of
mononuclear leukocytes |
2.42x10-19 | Increased | 4.104 | 51 |
| Cytotoxicity of
lymphocytes |
2.35x10-15 | Increased | 4.088 | 22 |
 | Table IIITop 5 inhibited biological functions,
particularly those related to lipid metabolism, associated with
differentially expressed miRNAs (|fold change| ≥1.5) and mRNAs
(|fold change| ≥2.0) in eyelid SGC with pagetoid spread. |
Table III
Top 5 inhibited biological functions,
particularly those related to lipid metabolism, associated with
differentially expressed miRNAs (|fold change| ≥1.5) and mRNAs
(|fold change| ≥2.0) in eyelid SGC with pagetoid spread.
| Functional
annotation | P-value | Predicted
activation state | Activation
z-score | No. of
molecules |
|---|
| Transport of
molecule | 0.00783 | Decreased | -3.291 | 25 |
| Fatty acid
metabolism | 0.000301 | Decreased | -2.202 | 15 |
| Transport of
lipid | 0.00429 | Decreased | -1.331 | 7 |
| Synthesis of
lipid | 0.00318 | Decreased | -1.15 | 16 |
| Absorption of
lipid | 0.000598 | Decreased | -1.123 | 4 |
Construction of molecular interaction
networks of expressed miRNAs and mRNAs
To further elucidate regulatory interactions between
miRNAs and mRNAs in the pagetoid group, target prediction analysis
was performed using the miRNA Target Filter tool in IPA.
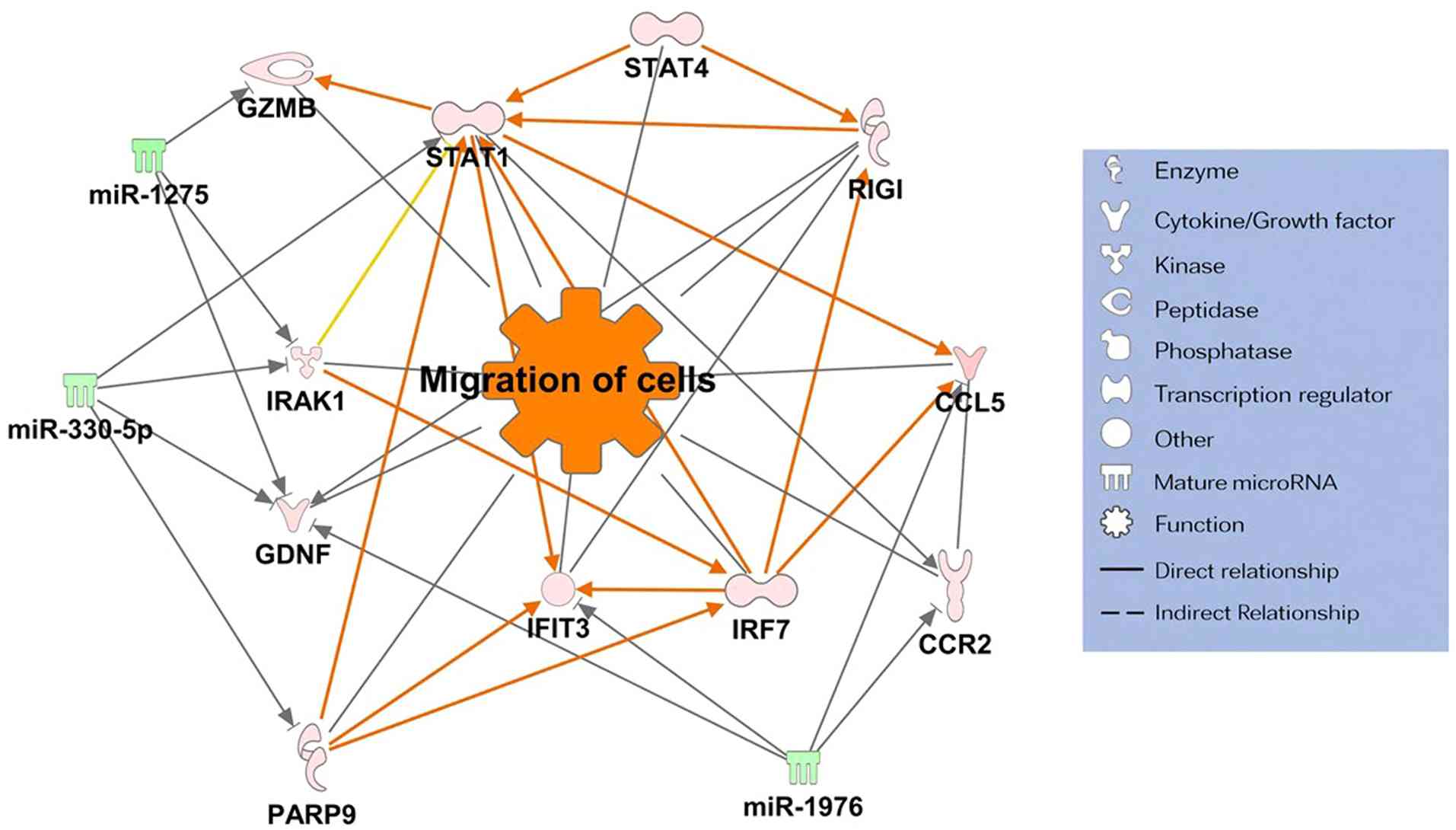
Consequently, two miRNA-mRNA networks associated with increased
gene expression were identified: i) The invasion and migration
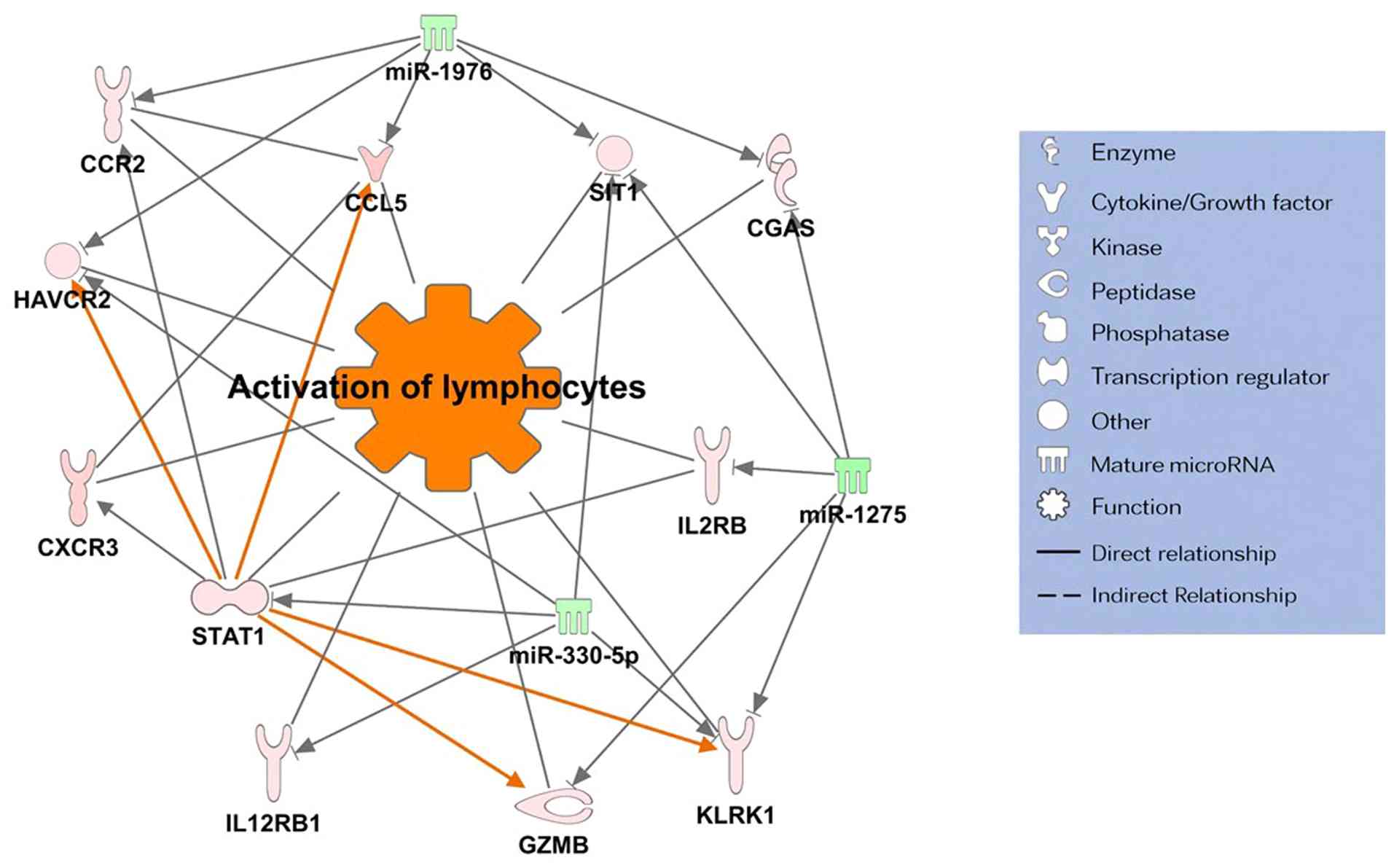
pathway (Fig. 3) and ii) the
immune-inflammatory response pathway (Fig. 4). Notably, both networks shared
three common target miRNAs: miR-330-5p, miR-1275, and miR-1976.
These miRNAs were consistently involved in gene expression
regulation in the pagetoid group and functioned as hub nodes,
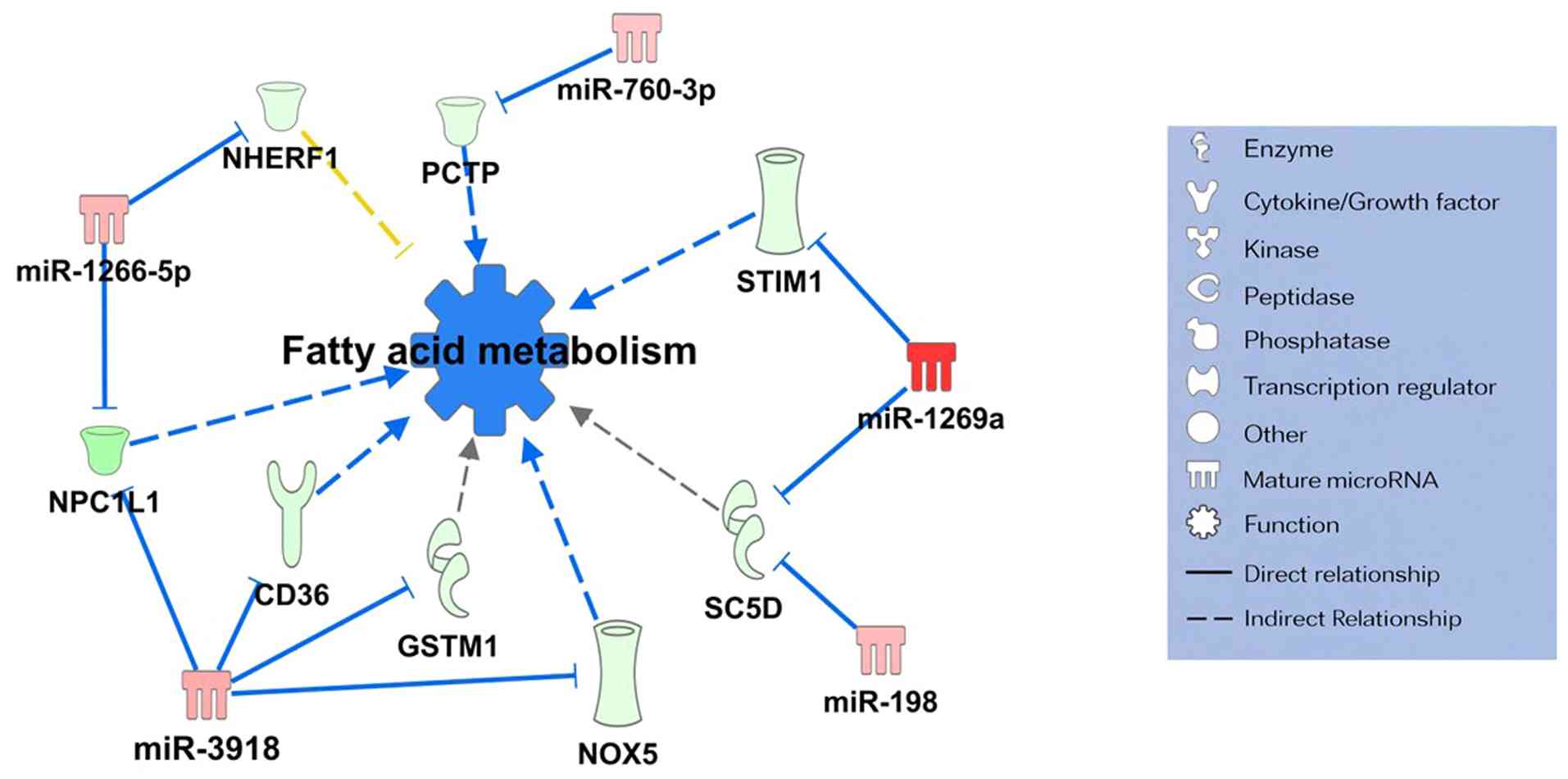
playing central roles within the networks. Moreover, a miRNA-mRNA
network associated with lipid metabolism and gene expression
suppression was identified (Fig.
5). This network analysis suggested that five miRNAs,
miR-760-3p, miR-1266-5p, miR-3918, miR-1269a, and miR-198, were
involved in the regulation of lipid metabolism.
Discussion
Complete surgical resection is the standard
treatment for SGC of the eyelid; however, the risk of recurrence
remains relatively high even when surgery is performed with an
adequate safety margin (3-6).
Intraepithelial invasion through pagetoid spread is a major factor
contributing to local recurrence (15-19).
Therefore, elucidating the pathogenesis of SGC of the eyelid with
pagetoid spread, exploring adjuvant or combination therapies
alongside surgery, and developing diagnostic biomarkers are
essential for improving patient outcomes. In the present study, the
expression profiles of miRNAs and mRNAs specifically associated
with SGC of the eyelid with pagetoid spread were analyzed.
Integrated miRNA-mRNA datasets were analyzed to investigate the
biological functions, canonical pathways, and miRNA-mRNA networks
associated with clinicopathological features. Consequently, SGC of
the eyelid with pagetoid spread exhibited upregulated expression of
genes related to invasion, migration, and immune-inflammatory
responses, organized into interconnected regulatory networks. These
networks appear to be epigenetically regulated by miR-330-5p,
miR-1275, and miR-1976. Furthermore, downregulated expression of
genes involved in lipid metabolism was observed, with related
networks regulated by miR-760-3p, miR-1266-5p, miR-3918, miR-1269a,
and miR-198.
Several studies have examined miRNA expression in
SGC. The downregulation of miR-200c and miR-141, which regulate the
expression of ZEB1, a transcription factor involved in
epithelial-mesenchymal transition, has been associated with SGC
malignancy (32). ZEB2 has been
identified as a target of miR-651-5p, and its expression is
regulated by this miRNA (33).
Furthermore, miR-3907 promotes proliferation and migration in SGC
(34). Reduced expression of
miR-518d and miR-211, both of which suppress cell proliferation,
has also been associated with SGC (27). Additionally, miR-205 and miR-199a
have been implicated in the maintenance of cancer stemness
(26). To date, only one study has
specifically investigated miRNAs involved in SGC of the eyelid with
pagetoid spread. That study demonstrated that, compared with the
nodular type, the pagetoid type exhibited overexpression of miR-205
and downregulation of miR-199a, with corresponding overexpression
of their respective target genes EZH2 (regulated by miR-205) and
CD44 (regulated by miR-199a) (26). CD44 overexpression is recognized as
a marker of cancer stemness and has been reported to promote cell
adhesion to the extracellular matrix and migration through
epithelial-to-mesenchymal transition, thereby activating adhesion,
migration, and proliferative signaling pathways (35). These findings are consistent with
the present study, in which invasion- and migration-related
pathways were upregulated. In our previous work, miR-146a-5p,
miR-149-3p, miR-193a-3p, miR-195-5p, and miR-4671-3p were
identified as miRNAs involved in the proliferation of SGC, whereas
miR-130a-3p and miR-939-5p were associated with the suppression of
lipid metabolism (28). Thus, SGC
is regulated by miRNAs that promote proliferation, enhance invasion
and migration through epithelial-mesenchymal transition, and
suppress lipid metabolism.
Consistent with these previous findings, the present
study further demonstrated that SGC of the eyelid with pagetoid
spread exhibits greater invasive and migratory potential and a more
pronounced reduction in lipid metabolism compared with the
non-pagetoid type.
SGC with pagetoid spread was found to form specific
gene networks involving key molecules that promote cancer cell
metastasis, invasion, and proliferation. These molecules include
signal transducer and activator of transcription 1 (STAT1)
(36,37), glial cell line-derived neurotrophic
factor (38,39), interferon regulatory factor
7(40), retinoic acid-inducible
gene I (40), poly (ADP-ribose)
polymerase family member 9 (41,42),
granzyme B (43), and
interleukin-1 receptor-associated kinase 1(44).
Furthermore, SGC with pagetoid spread forms
immune-inflammatory networks, which contribute to the recruitment
and migration of immune cells, such as lymphocytes and monocytes,
and may be involved in chronic inflammation, immune evasion, and
tumor immunity. The molecules involved in this network include CC
chemokine ligand 5(45), cyclic
GMP-AMP synthase (46),
STAT1(47), interleukin-12
receptor β1 subunit (48),
hepatitis A virus cellular receptor 2(49), C-X-C motif chemokine ligand
3(50), signaling threshold
regulating transmembrane adaptor 1(51), and granzyme B (43). Notably, both the
invasion/proliferation-related and inflammation-related gene
networks were commonly regulated by three downregulated miRNAs:
miR-330-5p, miR-1275, and miR-1976. miR-1275 is downregulated in
head and neck, colorectal, and esophageal cancers, where it
regulates cancer cell migration, invasion, and proliferation
(52-54).
In pancreatic cancer, it also plays a key role in natural killer
cell function and immune evasion (55). miR-1976 is downregulated in breast
and non-small cell lung cancers, and its decreased expression has
been associated with the promotion of epithelial-mesenchymal
transition (56,57). In ovarian cancer, it has been
implicated in the regulation of tumor-infiltrating immune cells
(58). Similarly, miR-330-5p is
downregulated in non-small cell lung and papillary thyroid cancers,
where it promotes cell proliferation, migration, and invasion
(59,60). In ovarian cancer, it regulates
antigen presentation by tumor cells and immune cell infiltration
(61). Thus, the downregulation of
miR-330-5p, miR-1275, and miR-1976 are thought to play key
epigenetic roles in regulating invasion, migration, and
inflammatory immune networks in SGC with pagetoid spread.
These invasion- and migration-related networks may
not be specific to pagetoid spread alone but could also reflect
other established clinicopathological risk factors, such as tumor
size and invasive growth patterns. Likewise, the
immune-inflammatory networks identified may be related to
tumor-infiltrating lymphocytes, which are recognized as important
biological features of SGC.
SGC with pagetoid spread exhibits suppressed lipid
metabolism and forms distinct gene networks. Intracytoplasmic lipid
accumulation in SGC can be detected by immunohistochemical staining
for adipophilin or Oil Red O staining, both of which are recognized
as pathological markers of SGC (62,63).
Abnormal lipid metabolism in sebaceous glands contributes to the
development of SGC and represents a key factor in understanding its
pathogenesis. In a previous study, lipid metabolism was suppressed
in SGC compared with sebaceous adenoma (28). The present findings suggest that
SGC of the eyelid with pagetoid spread is associated with a more
pronounced dysregulation of lipid metabolism-related pathways than
the non-pagetoid type, which may be related to its higher malignant
potential. The gene network related to lipid metabolism includes
sterol-C5-desaturase (64),
cluster of differentiation 36(65), phosphatidylcholine transfer protein
(66), Niemann-Pick C1-like
1(67), glutathione S-transferase
M1(68), NADPH oxidase 5(69), stromal interaction molecule
1(70), and Na+/H+ exchanger
regulatory factor 1(71). The
downregulation of this gene network was regulated by miR-760-3p,
miR-1266-5p, miR-3918, miR-1269a, and miR-198. These miRNAs are
suggested to function as upstream regulators of genes involved in
lipid metabolism in SGC with pagetoid spread.
This study has several limitations. First, the small
sample size precluded stratified or multivariate analyses adjusting
for established high-risk clinicopathological factors, such as
tumor size, pathological T stage, diffuse growth pattern, and
multicentric origin. Although tumor size, T stage, and
multicentricity were carefully documented, the molecular
differences observed in this study may not be exclusively
attributable to pagetoid spread alone and could be influenced by
overlapping pathological features. Second, the miRNA-mRNA
interactions identified in this study were based on integrated
expression analyses and bioinformatic predictions derived from
clinical tumor specimens. Direct experimental validation of miRNA
binding to the 3' untranslated regions of target genes, such as
dual-luciferase reporter assays or functional assays using cell
line models, was not performed. Accordingly, the proposed
regulatory relationships should be considered putative, and further
functional studies using appropriate experimental models will be
required to confirm direct miRNA-mRNA interactions and their
biological effects. Third, although immune-inflammatory pathways
were identified at the transcriptomic level, no histopathological
or immunohistochemical validation of the tumor immune
microenvironment was performed. In particular, the expression of
immune checkpoint molecules such as PD-L1 and the extent of
tumor-infiltrating lymphocytes, including CD8+ T cells,
were not evaluated in this study. Therefore, the immune-related
findings should be interpreted as transcriptional alterations, and
future studies incorporating immunohistochemical analyses will be
required to clarify their pathological and clinical significance.
In addition, this study was not designed as a direct experimental
comparison between pagetoid and nodular subtypes of SGC. Rather, it
aimed to explore molecular features associated with pagetoid spread
using integrated miRNA-mRNA expression analyses; therefore, the
identified molecular networks should be interpreted as
pagetoid-associated signatures rather than definitive
subtype-specific determinants. Taken together, this study should be
regarded as an exploratory, hypothesis-generating analysis
identifying pagetoid-associated miRNA-mRNA regulatory networks in
clinical SGC specimens. These findings provide a conceptual
framework for future mechanistic and translational studies,
including functional validation in experimental models and
comprehensive clinicopathological correlation analyses.
This study provides the first comprehensive
characterization of miRNA-mRNA interaction networks in SGC of the
eyelid with pagetoid spread. Several miRNAs with altered expression
were identified, potentially regulating critical functional
changes, including enhanced invasion, migration, and inflammatory
responses, and suppressed lipid metabolism. These findings advance
the current understanding of the pathophysiological mechanisms
underlying SGC of the eyelid and may aid in the identification of
potential biomarkers and therapeutic targets. Further experimental
validation and detailed investigation are required to elucidate the
precise functional roles of these miRNA-mRNA networks associated
with pagetoid spread. Future studies are expected to clarify how
these networks contribute to improved diagnostic and therapeutic
strategies for SGC.
Acknowledgements
Not applicable.
Funding
Funding: This study was supported in part by a Grant-in-Aid for
Scientific Research (grant no. 22K09786) from the Japan Society for
the Promotion of Science.
Availability of data and materials
The data generated in the present study may be found
in the DNA Data Bank of Japan Sequence Read Archive under
BioProject accession number PRJDB40390 or at the following URL:
https://ddbj.nig.ac.jp/resource/bioproject/PRJDB40390.
The dataset is also accessible via the NCBI BioProject database at
the following URL: https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJDB40390.
Authors' contributions
TY conceived and designed the study, performed the
experiments and drafted the manuscript. TH, YF and YT contributed
to the experimental work and data acquisition. AH contributed to
the conception and design of the study, and to the interpretation
of the data. AH and YT supervised the project and provided critical
revision of the manuscript. AH and YT confirm the authenticity of
all the raw data. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by the Institutional
Review Board of the University of Toyama (approval no. R2015051).
All procedures were conducted in accordance with the ethical
standards of the institutional and/or national research committee
and with the 1964 Declaration of Helsinki and its later amendments.
Written informed consent was obtained from all patients prior to
surgery, including consent for participation in the study and the
use of resected specimens for research purposes.
Patient consent for publication
Written informed consent for publication was
obtained from all patients.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Shields JA, Demirci H, Marr BP, Eagle RC
Jr and Shields CL: Sebaceous carcinoma of the ocular region: A
review. Surv Ophthalmol. 50:103–122. 2005.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Goto H, Yamakawa N, Komatsu H, Asakage M,
Tsubota K, Ueda SI, Nemoto R, Shibata M, Umazume K, Usui Y and Mori
H: Epidemiological characteristics of malignant eyelid tumors at a
referral hospital in Japan. Jpn J Ophthalmol. 66:343–349.
2022.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Goto H, Tsubota K, Nemoto R, Ueda S,
Umazume K, Usui Y and Matsumura H: Clinical features and prognosis
of sebaceous carcinoma arising in the eyelid or conjunctiva. Jpn J
Ophthalmol. 64:549–554. 2020.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Shields JA, Demirci H, Marr BP, Eagle RC
Jr and Shields CL: Sebaceous carcinoma of the eyelids: Personal
experience with 60 cases. Ophthalmology. 111:2151–2157.
2004.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Sa HS, Rubin ML, Xu S, Ning J, Tetzlaff M,
Sagiv O, Kandl TJ and Esmaeli B: Prognostic factors for local
recurrence, metastasis, and survival for sebaceous carcinoma of the
eyelid: observations in 100 patients. Br J Ophthalmol. 103:980–984.
2019.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Kaliki S, Ayyar A, Dave TV, Ali MJ, Mishra
DK and Naik MN: Sebaceous gland carcinoma of the eyelid:
clinicopathological features and outcome in Asian Indians. Eye
(Lond). 29:958–963. 2015.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Watanabe A, Sun MT, Pirbhai A, Ueda K,
Katori N and Selva D: Sebaceous carcinoma in Japanese patients:
Clinical presentation, staging and outcomes. Br J Ophthalmol.
97:1459–1463. 2013.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Choi YJ, Jin HC, Lee MJ, Kim N, Choung HK
and Khwarg SI: Prognostic value of clinical and pathologic T stages
defined by the American Joint Committee on Cancer for eyelid
sebaceous carcinoma in Korea. Jpn J Ophthalmol. 58:327–333.
2014.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Mulay K, Aggarwal E and White VA:
Periocular sebaceous gland carcinoma: A comprehensive review. Saudi
J Ophthalmol. 27:159–165. 2013.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Muqit MM, Foot B, Walters SJ, Mudhar HS,
Roberts F and Rennie IG: Observational prospective cohort study of
patients with newly-diagnosed ocular sebaceous carcinoma. Br J
Ophthalmol. 97:47–51. 2013.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Takahashi Y, Takahashi E, Nakakura S,
Kitaguchi Y, Mupas-Uy J and Kakizaki H: Risk factors for local
recurrence or metastasis of eyelid sebaceous gland carcinoma after
wide excision with paraffin section control. Am J Ophthalmol.
171:67–74. 2016.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Rao NA, Hidayat AA, McLean IW and
Zimmerman LE: Sebaceous carcinomas of the ocular adnexa: A
clinicopathologic study of 104 cases, with five-year follow-up
data. Hum Pathol. 13:113–122. 1982.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Esmaeli B, Nasser QJ, Cruz H, Fellman M,
Warneke CL and Ivan D: American Joint Committee on Cancer T
category for eyelid sebaceous carcinoma correlates with nodal
metastasis and survival. Ophthalmology. 119:1078–1082.
2012.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Muqit MM, Roberts F, Lee WR and Kemp E:
Improved survival rates in sebaceous carcinoma of the eyelid. Eye
(Lond). 18:49–53. 2004.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Sa HS, Tetzlaff MT and Esmaeli B:
Predictors of local recurrence for eyelid sebaceous carcinoma:
questionable value of routine conjunctival map biopsies for
detection of pagetoid spread. Ophthalmic Plast Reconstr Surg.
35:419–425. 2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Shields JA, Saktanasate J, Lally SE,
Carrasco JR and Shields CL: Sebaceous carcinoma of the ocular
region: the 2014 Professor Winifred Mao lecture. Asia Pac J
Ophthalmol (Phila). 4:221–227. 2014.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Chao AN, Shields CL, Krema H and Shields
JA: Outcome of patients with periocular sebaceous gland carcinoma
with and without conjunctival intraepithelial invasion.
Ophthalmology. 108:1877–1883. 2001.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Kaliki S, Vempuluru VS, Tanna V and Luthra
A: Eyelid and periocular sebaceous gland carcinoma: risk factors
for recurrence, exenteration, metastasis, and death in 355
patients. Can J Ophthalmol. 60:e395–e402. 2025.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Gu X, Xie M, Luo Y, Song X, Xu S and Fan
X: Diffuse pattern, orbital invasion, perineural invasion, and
Ki-67 are associated with nodal metastasis in patients with eyelid
sebaceous carcinoma. Br J Ophthalmol. 107:756–762. 2023.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Kaliki S, Morawala A, Dharap RS and
Mohamed A: Pagetoid tumour spread in periocular sebaceous gland
carcinoma: A comparative analysis in 130 patients. Eye (Lond).
35:2864–2870. 2021.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Shields JA, Demirci H, Marr BP, Eagle RC
Jr, Stefanyszyn M and Shields CL: Conjunctival epithelial
involvement by eyelid sebaceous carcinoma. The 2003 J. Howard
Stokes lecture. Ophthalmic Plast Reconstr Surg. 21:92–96.
2005.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Xu S, Moss TJ, Laura Rubin M, Ning J,
Eterovic K, Yu H, Jia R, Fan X, Tetzlaff MT and Esmaeli B:
Whole-exome sequencing for ocular adnexal sebaceous carcinoma
suggests PCDH15 as a novel mutation associated with metastasis. Mod
Pathol. 33:1256–1263. 2020.PubMed/NCBI View Article : Google Scholar
|
|
23
|
North JP, Golovato J, Vaske CJ, Sanborn
JZ, Nguyen A, Wu W, Goode B, Stevers M, McMullen K, Perez White BE,
et al: Cell of origin and mutation pattern define three clinically
distinct classes of sebaceous carcinoma. Nat Commun.
9(1894)2018.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Tetzlaff MT, Singh RR, Seviour EG, Curry
JL, Hudgens CW, Bell D, Wimmer DA, Ning J, Czerniak BA, Zhang L, et
al: Next-generation sequencing identifies high frequency of
mutations in potentially clinically actionable genes in sebaceous
carcinoma. J Pathol. 240:84–95. 2016.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Peterson C, Moore R, Hicks JL, Morsberger
LA, De Marzo AM, Zou Y, Eberhart CG and Campbell AA: NGS analysis
confirms common TP53 and RB1 mutations, and suggests MYC
amplification in ocular adnexal sebaceous carcinomas. Int J Mol
Sci. 22(8454)2021.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Bladen JC, Wang J, Sangaralingam A,
Moosajee M, Fitchett C, Chelala C, Beaconsfield M, O'Toole EA,
Philpott MP and Ezra DG: MicroRNA and transcriptome analysis in
periocular Sebaceous Gland Carcinoma. Sci Rep.
8(7531)2018.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Tetzlaff MT, Curry JL, Yin V,
Pattanaprichakul P, Manonukul J, Uiprasertkul M, Manyam GC, Wani
KM, Aldape K, Zhang L, et al: Distinct pathways in the pathogenesis
of sebaceous carcinomas implicated by differentially expressed
microRNAs. JAMA Ophthalmol. 133:1109–1116. 2015.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Hirano T, Yunoki T, Furusawa Y, Tabuchi Y
and Hayashi A: Bioinformatics analysis of the microRNA-mRNA network
in sebaceous gland carcinoma of the eyelid. Mol Med Rep.
23(44)2021.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Chu M, Zhang Y, Chen J, Cong H, Yin Y and
Chen H: Breast cancer bone metastasis: Novel prognostic biomarkers
identified. Phenomics. 5:404–417. 2025.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Zhong MF, Luo YJ, Guo YY, Xiang S and Lin
WF: Jiedu Fang inhibits hypoxia-induced angiogenesis in
hepatocellular carcinoma by targeting Aurora A/STAT3/IL-8 signaling
pathway. J Integr Med. 23:683–693. 2025.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Furusawa Y, Yunoki T, Hirano T, Minagawa
S, Izumi H, Mori H, Hayashi A and Tabuchi Y: Identification of
genes and genetic networks associated with BAG3dependent cell
proliferation and cell survival in human cervical cancer HeLa
cells. Mol Med Rep. 18:4138–4146. 2018.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Bhardwaj M, Sen S, Chosdol K, Sharma A,
Pushker N, Kashyap S, Bakhshi S and Bajaj MS: miRNA-200c and
miRNA-141 as potential prognostic biomarkers and regulators of
epithelial-mesenchymal transition in eyelid sebaceous gland
carcinoma. Br J Ophthalmol. 101:536–542. 2017.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Zhao H, Yang X, Liu J, Han F, Yang Z, Hu
Z, Liu M and Mei Y: Overexpression of miR-651-5p inhibits
ultraviolet radiation-induced malignant biological behaviors of
sebaceous gland carcinoma cells by targeting ZEB2. Ann Transl Med.
10(517)2022.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Zhang C, Zhu L, Liu X, Jiang M, Tang Q, Xu
F, Lin T, Dong L and He Y: MicroRNA-3907 promotes the proliferation
and migration of sebaceous gland carcinoma of the eyelid by
targeting thrombospondin 1. Oncol Lett. 22(833)2021.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Chen C, Zhao S, Karnad A and Freeman JW:
The biology and role of CD44 in cancer progression: Therapeutic
implications. J Hematol Oncol. 11(64)2018.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Anderson K, Ryan N, Nedungadi D, Lamenza
F, Swingler M, Siddiqui A, Satoskar A, Upadhaya P, Pietrzak M and
Oghumu S: STAT1 is regulated by TRIM24 and promotes
immunosuppression in head and neck squamous carcinoma cells, but
enhances T cell antitumour immunity in the tumour microenvironment.
Br J Cancer. 127:624–636. 2022.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Feng J, Li Y, Zhu L, Zhao Q, Li D, Li Y
and Wu T: STAT1 mediated long non-coding RNA LINC00504 influences
radio-sensitivity of breast cancer via binding to TAF15 and
stabilizing CPEB2 expression. Cancer Biol Ther. 22:630–639.
2021.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Garnis C, Davies JJ, Buys TP, Tsao MS,
MacAulay C, Lam S and Lam WL: Chromosome 5p aberrations are early
events in lung cancer: Implication of glial cell line-derived
neurotrophic factor in disease progression. Oncogene. 24:4806–4812.
2005.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Lu DY, Leung YM, Cheung CW, Chen YR and
Wong KL: Glial cell line-derived neurotrophic factor induces cell
migration and matrix metalloproteinase-13 expression in glioma
cells. Biochem Pharmacol. 80:1201–1209. 2010.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Tang XD, Zhang DD, Jia L, Ji W and Zhao
YS: lncRNA AFAP1-AS1 promotes migration and invasion of non-small
cell lung cancer via up-regulating IRF7 and the RIG-I-like receptor
signaling pathway. Cell Physiol Biochem. 50:179–195.
2018.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Juszczynski P, Kutok JL, Li C, Mitra J,
Aguiar RC and Shipp MA: BAL1 and BBAP are regulated by a gamma
interferon-responsive bidirectional promoter and are overexpressed
in diffuse large B-cell lymphomas with a prominent inflammatory
infiltrate. Mol Cell Biol. 26:5348–5359. 2006.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Tang X, Zhang H, Long Y, Hua H, Jiang Y
and Jing J: PARP9 is overexpressed in human breast cancer and
promotes cancer cell migration. Oncol Lett. 16:4073–4077.
2018.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Lu Z, Huang X, Shen Q, Chen E and Feng Y:
Granzyme B promotes proliferation, migration, and EMT process in
gastric cancer. Biochem Genet. 63:2513–2529. 2025.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Bennett J and Starczynowski DT: IRAK1 and
IRAK4 as emerging therapeutic targets in hematologic malignancies.
Curr Opin Hematol. 29:8–19. 2022.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Liu C, Yao Z, Wang J, Zhang W, Yang Y,
Zhang Y, Qu X, Zhu Y, Zou J, Peng S, et al: Macrophage-derived CCL5
facilitates immune escape of colorectal cancer cells via the
p65/STAT3-CSN5-PD-L1 pathway. Cell Death Differ. 27:1765–1781.
2020.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Motwani M, Pesiridis S and Fitzgerald KA:
DNA sensing by the cGAS-STING pathway in health and disease. Nat
Rev Genet. 20:657–674. 2019.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Verhoeven Y, Tilborghs S, Jacobs J, De
Waele J, Quatannens D, Deben C, Prenen H, Pauwels P, Trinh XB,
Wouters A, et al: The potential and controversy of targeting STAT
family members in cancer. Semin Cancer Biol. 60:41–56.
2020.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Langowski JL, Zhang X, Wu L, Mattson JD,
Chen T, Smith K, Basham B, McClanahan T, Kastelein RA and Oft M:
IL-23 promotes tumour incidence and growth. Nature. 442:461–465.
2006.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Yu M, Lu B, Liu Y, Me Y, Wang L and Zhang
P: Tim-3 is upregulated in human colorectal carcinoma and
associated with tumor progression. Mol Med Rep. 15:689–695.
2017.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Tokunaga R, Zhang W, Naseem M, Puccini A,
Berger MD, Soni S, McSkane M, Baba H and Lenz HJ: CXCL9, CXCL10,
CXCL11/CXCR3 axis for immune activation-a target for novel cancer
therapy. Cancer Treat Rev. 63:40–47. 2018.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Jia M, Liu C, Liu Y, Bao Z, Jiang Y and
Sun X: Discovery and validation of a SIT1-related prognostic
signature associated with immune infiltration in cutaneous
melanoma. J Pers Med. 13(13)2022.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Liu MD, Wu H, Wang S, Pang P, Jin S, Sun
CF and Liu FY: MiR-1275 promotes cell migration, invasion and
proliferation in squamous cell carcinoma of head and neck via
up-regulating IGF-1R and CCR7. Gene. 646:1–7. 2018.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Kahlert C, Klupp F, Brand K, Lasitschka F,
Diederichs S, Kirchberg J, Rahbari N, Dutta S, Bork U, Fritzmann J,
et al: Invasion front-specific expression and prognostic
significance of microRNA in colorectal liver metastases. Cancer
Sci. 102:1799–1807. 2011.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Xie C, Wu Y, Fei Z, Fang Y, Xiao S and Su
H: MicroRNA-1275 induces radiosensitization in oesophageal cancer
by regulating epithelial-to-mesenchymal transition via
Wnt/beta-catenin pathway. J Cell Mol Med. 24:747–759.
2020.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Ou Z, Lu Y, Xu D and Luo Z: Hypoxia
mediates immune escape of pancreatic cancer cells by affecting
miR-1275/AXIN2 in natural killer cells. Front Immunol.
14(1271603)2023.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Wang J, Ma G, Han X, Liang M, Wang X, Xia
T and Wang S: The low expression of miR-1976 in plasma samples
indicating its biological functions in the progression of breast
cancer. Clin Transl Oncol. 22:2111–2120. 2020.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Chen G, Hu J, Huang Z, Yang L and Chen M:
MicroRNA-1976 functions as a tumor suppressor and serves as a
prognostic indicator in non-small cell lung cancer by directly
targeting PLCE1. Biochem Biophys Res Commun. 473:1144–1151.
2016.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Chen L, Gao W, Lin L, Sha C, Li T, Chen Q,
Wei H, Yang M, Xing J, Zhang M, et al: A methylation- and
immune-related lncRNA signature to predict ovarian cancer outcome
and uncover mechanisms of chemoresistance. J Ovarian Res.
16(186)2023.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Cui LH, Xu HR, Yang W and Yu LJ: lncRNA
PCAT6 promotes non-small cell lung cancer cell proliferation,
migration and invasion through regulating miR-330-5p. Onco Targets
Ther. 11:7715–7724. 2018.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Gao Y, Wang F, Zhang L, Kang M, Zhu L, Xu
L, Liang W and Zhang W: LINC00311 promotes cancer stem-like
properties by targeting miR-330-5p/TLR4 pathway in human papillary
thyroid cancer. Cancer Med. 9:1515–1528. 2020.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Liu C, Huang Y, Cui Y, Zhou J, Qin X,
Zhang L, Li X, Li Y, Guo E, Yang B, et al: The immunological role
of CDK4/6 and potential mechanism exploration in ovarian cancer.
Front Immunol. 12(799171)2022.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Jakobiec FA and Mendoza PR: Eyelid
sebaceous carcinoma: clinicopathologic and multiparametric
immunohistochemical analysis that includes adipophilin. Am J
Ophthalmol. 157:186–208.e2. 2014.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Milman T, Schear MJ and Eagle RC Jr:
Diagnostic utility of adipophilin immunostain in periocular
carcinomas. Ophthalmology. 121:964–971. 2014.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Nishi S, Nishino H and Ishibashi T: cDNA
cloning of the mammalian sterol C5-desaturase and the expression in
yeast mutant. Biochim Biophys Acta. 1490:106–108. 2000.PubMed/NCBI View Article : Google Scholar
|
|
65
|
Mundi MS, Velapati S, Patel J, Kellogg TA,
Abu Dayyeh BK and Hurt RT: Evolution of NAFLD and its management.
Nutr Clin Pract. 35:72–84. 2020.PubMed/NCBI View Article : Google Scholar
|
|
66
|
Kang HW, Wei J and Cohen DE: PC-TP/StARD2:
Of membranes and metabolism. Trends Endocrinol Metab. 21:449–456.
2010.PubMed/NCBI View Article : Google Scholar
|
|
67
|
Liu W, Liang B, Zeng J, Meng J, Shi L,
Yang S, Chang J, Wang C, Hu X, Wang X, et al: First discovery of
cholesterol-lowering activity of parthenolide as NPC1L1 inhibitor.
Molecules. 27(6270)2022.PubMed/NCBI View Article : Google Scholar
|
|
68
|
Liu YK, Dong YH, Liang XM, Qiang S, Li ME,
Sun Z, Zhao X, Yan ZH and Zheng J: Application of integrated omics
in aseptic loosening of prostheses after hip replacement. Mol Med
Rep. 31(65)2025.PubMed/NCBI View Article : Google Scholar
|
|
69
|
García JG, Ansorena E, Milagro FI, Zalba G
and de Miguel C: Endothelial Nox5 expression modulates glucose
uptake and lipid accumulation in mice fed a high-fat diet and
3T3-L1 adipocytes treated with glucose and palmitic acid. Int J Mol
Sci. 22(2729)2021.PubMed/NCBI View Article : Google Scholar
|
|
70
|
Maus M, Cuk M, Patel B, Lian J, Ouimet M,
Kaufmann U, Yang J, Horvath R, Hornig-Do HT,
Chrzanowska-Lightowlers ZM, et al: Store-operated Ca2+
entry controls the induction of lipolysis and the transcriptional
reprogramming to lipid metabolism. Cell Metab. 25:698–712.
2017.PubMed/NCBI View Article : Google Scholar
|
|
71
|
Wan Y, Hudson R, Smith J, Forman-Kay JD
and Ditlev JA: Protein interactions, calcium, phosphorylation, and
cholesterol modulate CFTR cluster formation on membranes. Proc Natl
Acad Sci USA. 122(e2424470122)2025.PubMed/NCBI View Article : Google Scholar
|