Introduction
High blood pressure is a major risk factor for
cardiovascular disease. The pattern of blood pressure in western
populations is such that the majority of the population is at an
increasing risk for blood pressure-related cardiovascular diseases
(1–5). High blood pressure has been
associated with a variety of nutritional abnormalities and an
increased prevalence of physical inactivity (6–8).
Although some of the early studies argue against the genetic cause
of hypertension (9,10), several more recent ones provide
evidence in favor of the genetic explanation (11,12).
Cloning of leptin gene and characterization of its product leptin
was an important advance in the study of hormonal and metabolic
alterations associated with this gene (13). Leptin is mainly produced by adipose
tissue, and leptin levels are strongly and positively correlated
with body mass index (BMI) (14,15).
However, the role of leptin deficiency in human obesity has also
been considered (16,17).
Previous research has demonstrated that leptin is a
pleiotropic hormone with multiple actions that is potentially
involved in the control of feeding, as well as in cardiovascular
function, insulin secretion, angiogenesis, immune response and
haematopoiesis (18). A direct
effect of leptin on blood pressure has been reported (19–21),
and it has been suggested that this effect is mediated by
sympathetic activation (22).
Moreover, a direct relationship between plasma leptin and heart
rate has been observed in hypertensive patients (20,23).
In normotensive men, serum leptin levels were found to be related
to serum angiotensin II levels (33), which has a well established role in
the regulation of blood pressure. Also, a positive correlation was
found between these two parameters in hypertensive subjects
(34). This relationship appears
to be independent of BMI, plasma insulin and physical activity. The
presence of a microsatellite polymorphism in the 3′UTR of the
leptin gene was found to be significantly correlated with
hypertension, independent of obesity in a Japanese population
(24). A similar polymorphism in
the leptin gene was found in an Italian population, but no
association of this polymorphism with obesity-dependent
hypertension was observed (25).
Another variant, the C538T in the 3′UTR of the
leptin gene, was similarly shown to be associated with essential
hypertension independent of obesity (26). However, in another study on an
African-American group, several genetic markers at the leptin
locus, including the one described by Shintani et al
(24), were not significantly
linked to hypertension (27).
These contradictory data prompted us to investigate the presence of
common tetranucleotide repeat polymorphisms and the rare C538T SNP
in the 3′ flanking region of the leptin gene in: hypertensive
patients independent of obesity, in obese hypertensive patients and
in controls. Since several hormones, such as insulin and steroids
(35,36), affect angiotensin mRNA, this study
was designed to investigate whether leptin upregulates the activity
of the gene in adipocytes, thereby contributing to
hypertension.
Materials and methods
Subjects
In the present study, 362 patients with hypertension
were investigated; 119 subjects were hypertensive with a BMI of
22.10±2.8 and 155 were hypertensive with a BMI of 35.2±5.12.88.
Normotensive subjects with different BMIs were also studied as
controls. All subjects were either first time diagnosed or had not
taken any medication at least 15 days prior to the study. All
subjects underwent a physical examination that included measurement
of the BMI (evaluated by weight/height2;
kg/m2) and levels of serum urea, creatinine, billuribin,
cholesterol, glucose and triglyceride. All subjects had normal
thyroid function. No patient with renal, hepatic or cardiac disease
was included and none of the subjects had diabetes. Therefore, only
a specific group of subjects with essential hypertension was
selected. After determining all the biochemical parameters and
measuring other anthropometric parameters, blood was obtained from
the subjects, collected in EDTA vials and stored at −20°C for
genomic studies. The study was approved by the local institutional
ethics committee, and informed consent was obtained from all of the
subjects.
Genotyping
Genomic DNA was extracted from whole blood using a
commercial kit (Zymo Research Corp., Irvine, CA, USA). Genotyping
of the tetranucleotide polymorphisms in the 3′ flanking region of
the leptin gene was detected by polymerase chain reaction using
forward primer 5′-AGTTCAAATAGAGGTCCAAATCA-3′ and reverse primer
5′-TTCTGAGGTTGTGTCACTGGCA-3′ that flanks the microsatellite in the
3′ flanking region of the leptin gene. Another primer pair,
5′-CGACCTGGAGAACCTCCG-3′ as forward and 5′-GTCCTGGATAAGGGGTGT-3′ as
reverse, was used for amplification of the 316-bp amplicon in the
3′UTR of the leptin gene for the screening of the C538T variant.
PCR contained 100 ng of genomic DNA template, 0.2 μM of each
primer, 2 mM of Mg2+, 0.2 mM of each dNTP, 1.5 units of
Taq polymerase and 1X reaction buffer in a total volume of
25 μl. The PCR was performed for 30 cycles of 30 sec at 94°C, 30
sec at 55°C and 1 min at 72°C, with an initial denaturation of 5
min at 94°C and a final extension of 10 min at 72°C. The reaction
conditions were the same for both pairs of primers. PCR products
were run in 2% agarose gel along with 100-bp ladder as a molecular
weight marker. Amplified products were visualized by staining with
ethidium bromide. In the case of the tetranucleotide repeat
polymorphisms, alleles were distinguished on the basis of amplicon
length, which varies for different alleles due to the variable
number of tetranucleotide repeats. For the screening of the C538T
variant, a 316-bp fragment was digested with the HypCH4IV enzyme.
The digested products were run on 8% PAGE along with pBR322MspI
digest as a molecular weight marker. Ten percent of the digested
samples were sequenced directly as well.
Analysis of serum leptin
concentration
To study the relationship between the leptin gene
polymorphism and the serum leptin concentrations, serum leptin
levels were determined with the human leptin immunoassay kit (DRG
International Inc., Mountainside, NJ, USA). Serum was collected
from each patient in the morning. Frozen samples were kept at −20°C
until analysis.
Analysis of serum angiotensin II
concentration
To study whether leptin upregulates the expression
of angiotensin II levels, which is the main regulator of blood
pressure, the serum angiotensin II concentration was determined
with the angiotensin II ELISA kit (DRG International Inc.).
Statistical analysis
Data are expressed as the means ± SD. The means were
compared by the independent sample t-test. Fisher’s exact test or
χ2 test was used to compare the frequencies. Regression
analysis was used to describe the relationship between serum leptin
as a dependent variable, and the BMI and serum angiotensin II
levels as independent variables. Statistical analysis was carried
out using the statistical package SPSS.
Results
Table I shows the
characteristics of the population that was genotyped for the
microsatellite polymorphisms and the C538T variant in the 3′
flanking region of the leptin gene. The age of the hypertensive
group ranged from 45 to 65 years (mean ± SD, 53±8.35), while the
age of the control group ranged from 35 to 50 years (42.025±8.6).
In the hypertensive group, 168 subjects were female and 106 were
male, and in the normotensive group 30 were female and 58 male.
There was no statistically significant difference in the serum
urea, creatinine, glucose, bilirubin, triglyceride and angiotensin
II, when compared between the hypertensive vs. normotensive class,
or between the obese vs. lean class (p>0.05). A significant
correlation was found between serum leptin when compared between
the hypertensive vs. normotensive class, and also between the obese
and lean class (p≤0.05). Linear regression analysis showed an
independent correlation of leptinemia with BMI (p=0.019), but no
significant correlation was found between the serum leptin
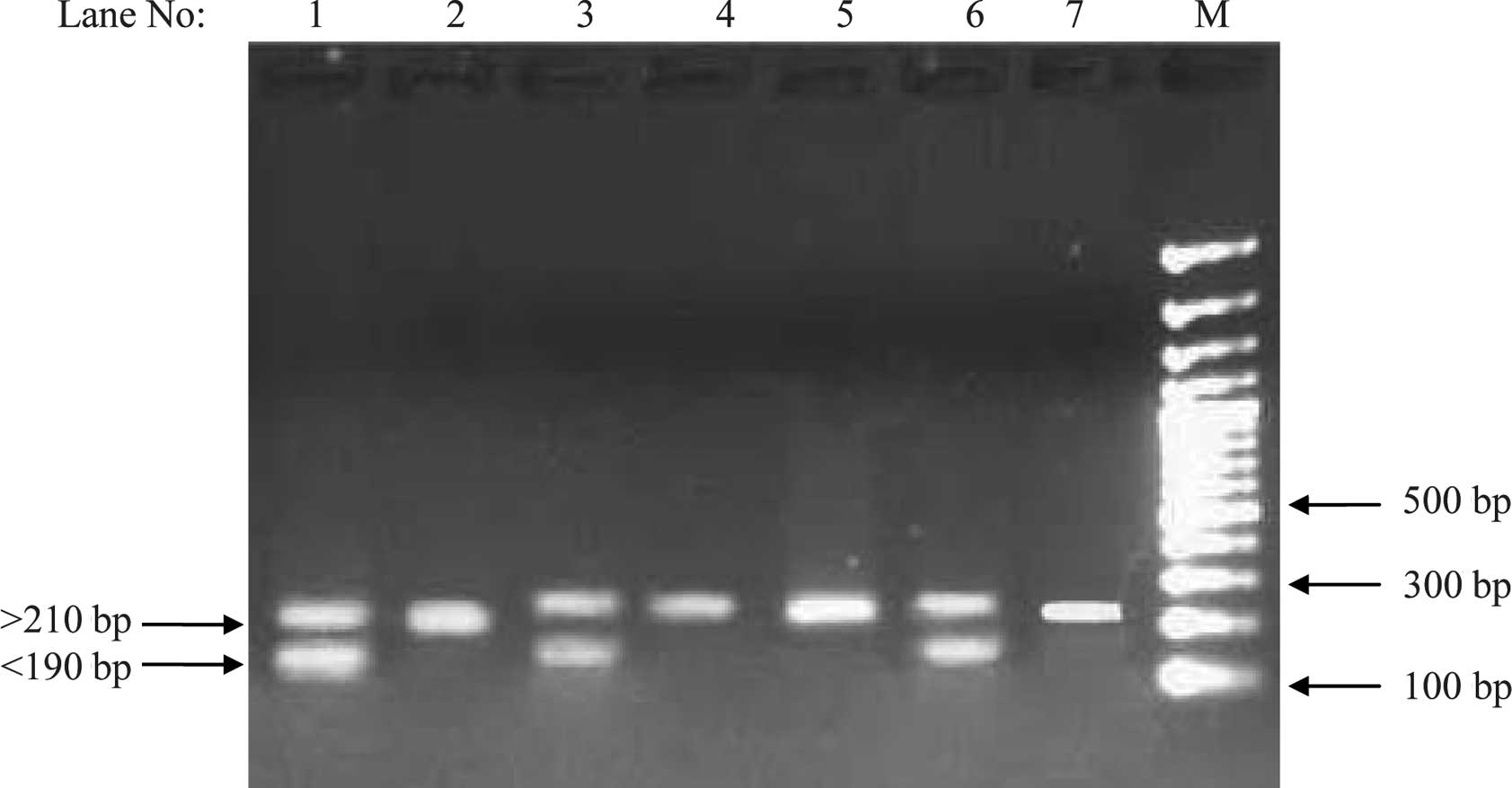
concentration and angiotensin II (Table IV). Alleles of the microsatellite
were composed of two groups with different size distribution; a
shorter one, <190 bp in size (termed as Class I), and a longer
one >210 bp in size (termed as Class II; Fig. 2). The frequency of Class I/Class I
genotype were higher in the hypertensive group than in the
normotensive group (Table II).
Class I/I and Class I/II was found to be significantly correlated
with serum leptin concentration (p≤0.05), but no such correlation
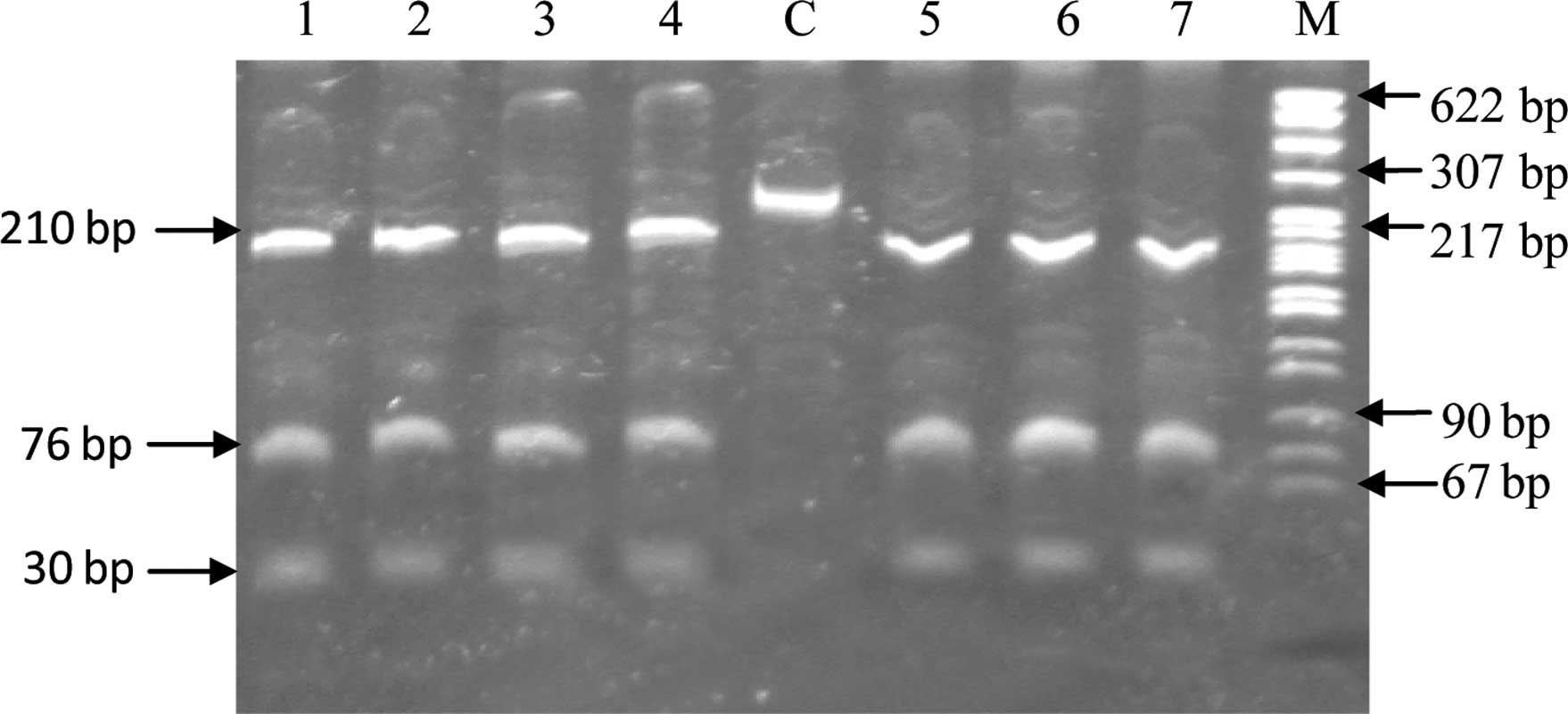
was observed with serum angiotensin II levels (Table III). In the case of the C538T
variant, upon digestion with the HpyCH4IV restriction enzyme, the
wild-type genotype provided three fragments of size 210, 76 and 30
bp, carriers had four fragments of size 210, 76, 30 and 286 bp,
whereas mutant homozygotes gave two bonds of size 286 and 30 bp.
All 362 subjects studied showed the digestion pattern of wild-type
homozygotes (Fig. 1) and direct
sequencing of 10% of the subjects confirmed this.
 | Table IAnthropometric and laboratory data of
the study subjects. |
Table I
Anthropometric and laboratory data of
the study subjects.
| Variables | Hypertensive
lean | Hypertensive
obese | Normotensive
obese | Normotensive
lean | Sign. of
difference |
|---|
| Age (years) | 53.00±8.35 | 50.00±8.35 | 43.20±8.60 | 42.025±8.60 | p=0.0001 |
| BMI
(kg/m2) | 22.10±2.80 | 35.20±5.12 | 29.50±3.20 | 21.850±1.01 | p=0.0001 |
| Systolic blood
pressure (mmHg) | 144.51±16.01 | 145.10±15.21 | 120.90±2.20 | 120.500±5.75 | p=0.0001 |
| Diastolic blood
pressure (mmHg) | 92.25±10.61 | 92.20±12.18 | 80.50±3.20 | 80.050±3.12 | p=0.0001 |
| Serum creatinine
(mg/dl) | 0.90±0.23 | 1.05±0.22 | 0.90±0.11 | 0.920±0.25 | p=0.4230 |
| Urea (mg/dl) | 25.75±9.39 | 30.00±6.70 | 31.00±10.50 | 26.320±8.70 | p=0.7780 |
| Cholesterol
(mg/dl) | 165.00±23.80 | 175.00±23.80 | 172.00±19.80 | 160.770±25.60 | P=0.0250
P2=0.0470 |
| Glucose R
(mg/dl) | 98.40±22.62 | 94.00±7.13 | 102±14 | 95.500±15.70 | p=0.5070 |
| Triglycerides
(mg/dl) | 165.37±21.90 | 182.02±15.80 | 180±23 | 165±15.20 | P=0.3800
P2=0.7170 |
| Na+ | 144.00±5.10 | 141.00±6.20 | 139±5.30 | 142±8.23 | P=0.2020 |
| K+ | 4.50±0.46 | 4.03±0.48 | 4.14±0.44 | 3.900±0.56 | P=0.4100 |
| Leptin (ng/dl) | 11.90±3.77 | 18.71±15.50 | 13.82±12.63 | 9.420±12.20 | P=0.0070
P2=0.0630 |
| Angiotensin II
(ng/dl) | 1.17±1.36 | 1.47±1.60 | 1.34±2.20 | 1.250±1.23 | P=0.9760
P2=0.1280 |
 | Table IVLinear regression analysis for all
subjects (HL, HO, NL and NO), obese subjects (HO and NO), and lean
subjects (HL and NL). |
Table IV
Linear regression analysis for all
subjects (HL, HO, NL and NO), obese subjects (HO and NO), and lean
subjects (HL and NL).
| Independent
variables | Regression
coefficients | 95% confidence
interval of regression coefficient | p-value | R2 |
|---|
| BMI (HO, HL, NO,
NL) | 0.255 | 0.02384 to
0.26150 | 0.019 | 0.065 |
| ANG II (HO, HL, NO,
NL) | 0.171 | −0.01370 to
0.04150 | 0.120 | 0.029 |
| BMI (HO, NO) | 0.482 |
−0.06166±0.01653 | 0.001 | 0.232 |
| ANG II (HO,
NO) | 0.148 |
−0.05043±0.05405 | 0.357 | 0.025 |
| BMI (HL, NL) | 0.167 |
0.02759±0.02397 | 0.256 | 0.028 |
| ANG II (HL,
NL) | 0.203 | −0.01253 to
0.04931 | 0.235 | 0.041 |
 | Table IIFrequency of the genotypes of the
leptin gene polymorphism in hypertensive and normotensive
subjects. |
Table II
Frequency of the genotypes of the
leptin gene polymorphism in hypertensive and normotensive
subjects.
| Variants/Genotypic
distribution | Hypertensive
lean | Hypertensive
obese | Normotensive | Total |
|---|
| C538T |
| C/C | 119 | 155 | 88 | |
| C/T | Nil | Nil | Nil | 362 |
| T/T | Nil | Nil | Nil | |
| Microsatellite |
| I/I | 53 (44.5) | 71 (45.8) | 12 (13.6) | 362 |
| I/II | 41 (34.4) | 60 (38.7) | 42 (47.7) | p≤0.0001 |
| II/II | 25 (21.0) | 24 (15.4) | 34 (38.6) | |
 | Table IIIAssociation of different alleles with
serum leptin and serum angiotensin II levels. |
Table III
Association of different alleles with
serum leptin and serum angiotensin II levels.
| Microsatellite
genotypes | Serum leptin
(ng/dl) | Serum angiotensin
II (ng/dl) | Significance |
|---|
| Genotype I/I
(n=26) | 19.2±6.12 | 1.2±1.35 |
Pa=0.022
Pa1=0.737 |
| Genotype I/II
(n=34) | 18.7±15.2 | 1.40±1.55 |
Pb=0.080
Pa2=0.640 |
| Genotype II/II
(n=24) | 16.2±12.6 | 1.28±1.26 |
Pc=0.298
Pa3=0.336 |
Discussion
Obesity and essential hypertension are considered to
be the result of multiple environmental and genetic determinants.
These two factors are closely linked in epidemiological studies
(28), and essential hypertension
is estimated to be up to three times more prevalent among obese
populations (29). One of the
major mechanisms leading to the development of obesity-induced
hypertension appears to be leptin-mediated sympatho-activation. In
African-Americans, highly polymorphic markers in leptin locus were
not significantly linked to the trait of essential hypertension
(27). Later, several studies on a
Japanese population showed a significant association of the
polymorphism in the 3′UTR region of the leptin gene with
hypertension, while this association was independent of BMI. In
another study involving a British population, another rare variant
in the 3′UTR of the leptin gene was found to impact pulse pressure
considerably, independent of obesity and cortico intima-mediated
thickness (26). The results of
our study confirmed that the 3′ flanking region polymorphism of the
gene previously described by Shintani et al (24) is also found in a pure ethnic
population of Kashmir. A significant association of the
microsatellite polymorphism with hypertension was found. Another
variant, C538T, was observed to be very rare in some populations
(26,32), but seems to be either absent or
even rarer in the Kashmiri population. However, a larger sample
size is required to confirm this possibility. Although the
microsatellite polymorphism examined is located in non-coding
region, similarly located variants in the 3′UTR of other genes are
known to play a key role in their expression (30,31).
There are generally some cis-acting determinants in the 3′UTR to
which proteins bind and either stabilize or destabilize the mRNA.
Cis-acting determinants in the 3′UTR may also interact with other
sequences within that mRNA. Therefore, the variation in these
cis-acting determinants may affect the expression of this
particular gene. This case may be similar to the present study, as
we found that serum leptin is significantly associated with Class
I/Class I and Class I/Class II genotypes. Thus, the Class I/I and
I/II genotypes were shown to be closely associated to hypertensive
and serum leptin concentration. To determine whether this relation
of leptin is mediated through angiotensin II, whose role in the
regulation of blood pressure is well established, the concentration
of angiotensin II was also assessed. However, a negative
correlation was found between leptin and angiotensin II in all
groups. The present data suggest that the common polymorphism in
the 3′ flanking region of the leptin gene is relative to essential
hypertension. Further in vitro and animal model research is
necessary to elucidate how the variation in the size of
microsatellite in the 3′ flanking region of the gene is involved in
the regulation of blood pressure in obesity-dependent and
obesity-independent hypertension.
Acknowledgements
This study was supported by research grants from the
Department of Science and Technology, Government of India New
Delhi, under the Women Scientist Scheme.
References
|
1
|
J ClausenG JensenAre blood pressure levels
increasing in Denmark?J Intern
Med228443450198510.1111/j.1365-2796.1990.tb00261.x
|
|
2
|
FH EpsteinT FrancisNS HaynerPrevalence of
chronic diseases and distribution of selected physiologic variables
in a total community, Tecumseh, MichiganAm J
Epidemiol81307322196514294415
|
|
3
|
E FerranniniSM HaffnerMP SternInsulin
sensitivity and hypertensionJ Hypertension8Suppl II371990
|
|
4
|
WB KannelPW WilsonTJ ZhangThe epidemiology
of impaired glucose tolerance and hypertensionAm Heart
J12112681273199110.1016/0002-8703(91)90432-H2008855
|
|
5
|
LS WebberAW VoorsSR SrinivasanRR
FrerichsGS BerensonOccurrence in children of multiple risk factors
for coronary artery disease: the Bogalusa heart studyPrev
Med8407418197910.1016/0091-7435(79)90018-5471959
|
|
6
|
J TuomilehtoP ZimmetR TaylorP BennettE
WolfA cross-sectional ecological analysis of blood pressure and its
determinants in eleven Pacific populationsJ Am Coll
Nutr8151165198910.1080/07315724.1989.107202902785129
|
|
7
|
PS SeverD GordonWS PeartP
BeightonBlood-pressure and its correlates in urban and tribal
AfricaLancet26064198010.1016/S0140-6736(80)92940-26105245
|
|
8
|
A Costa EdeG RoseCH KleinSalt and blood
pressure in Rio Grande do Sul, BrazilBull Pan Am Health
Organ2415917619902379022
|
|
9
|
A AmeryR FagardC GuoJ StaessenL
ThusIsolated systolic hypertension in elderlyJAMA25070741991
|
|
10
|
WH FrishmanEpidemiology, pathophysiologic,
and management of isolated systolic hypertension in the elderlyAm J
Med9Suppl 3A14201992
|
|
11
|
A BonnardeauxE DaviesX
JeunemaitreAngiotensin II type 1 receptor gene polymorphisms in
human essential
hypertensionHypertension246369199410.1161/01.HYP.24.1.638021009
|
|
12
|
S MarkanM SachdevaDS SehrawatS KumariS
JainM KhullarMTHFR 677 CT/MTHFR 1298 CC genotypes are associated
with increased risk of hypertension in IndiansMol Cell
Biochem302125131200710.1007/s11010-007-9434-517333388
|
|
13
|
Y ZhangR ProencaM MaffeiM BaroneL
LeopoldJM FriedmanPositional cloning of the mouse obese gene and
its human homologueNature372425432199410.1038/372425a07984236
|
|
14
|
TW StephensM BasinskiPK BristowThe role of
neuropeptide Y in the antiobesity action of the obese gene
productNature377530532199510.1038/377530a07566151
|
|
15
|
H MasuzakiY OgawaN IsseHuman obese gene
expression. Adipocyte-specific expression and regional differences
in the adipose
tissueDiabetes44855888199510.2337/diab.44.7.8557789654
|
|
16
|
CT MontagueSI FarooqiJP
WhiteheadCongenital leptin deficiency is associated with severe
early-onset obesity in
humansNature387903908199710.1038/431859202122
|
|
17
|
RV ConsidineMK SinhaML HeimanSerum
immunoreactive-leptin concentrations in normal-weight and obese
humansNew Engl J
Med334292295199610.1056/NEJM1996020133405038532024
|
|
18
|
P StenvinkelF LonnqvistM
SchallingMolecular studies of leptin: implications for renal
diseaseNephrol Dial
Transplant1411031112199910.1093/ndt/14.5.110310344346
|
|
19
|
K NarkiewiczVK SomersL MosM KatoV AccursoP
PalatiniAn independent relationship between plasma leptin and heart
rate in untreated patients with essential hypertensionJ
Hypertens17245249199910.1097/00004872-199917020-0000910067794
|
|
20
|
EW ShekMW BrandsJE HallChronic leptin
infusion increases arterial
pressureHypertension31409414199810.1161/01.HYP.31.1.4099453337
|
|
21
|
JC DunbarY HuH LuIntracerebroventricular
leptin increases lumber and renal sympathetic nerve activity and
blood pressure in normal
ratsDiabetes4620402043199710.2337/diab.46.12.20409392493
|
|
22
|
M Aizawa-AbeY OgawaH
MasuzakiPathophysiological role of leptin in obesity-related
hypertensionJ Clin Invest10512431252200010.1172/JCI834110791999
|
|
23
|
PM SuterR LocherE HaslerW VetterIs there a
role for the ob gene product leptin in essential hypertension?Am J
Hypertens1113051311199810.1016/S0895-7061(98)00162-99832173
|
|
24
|
M ShintaniH IkegamiE YamatoA novel
microsatellite polymorphism in the human OB gene: a highly
polymorphic marker for linkage
analysisDiabetologia3913981401199610.1007/s0012500505898933011
|
|
25
|
S MaestriniM MencarelliB VertiLack of
association between the tetranucleotide repeat polymorphism in the
3′-flanking region of the leptin gene and hypertension in severely
obese patientsJ Endocrinol Invest297767802006
|
|
26
|
N GaukrodgerBM MayosiH ImrieA rare variant
of the leptin gene has large effects on blood pressure and carotid
intima-medial thickness: a study of 1428 individuals in 248
familiesJ Med Genet42474478200510.1136/jmg.2004.02763115937081
|
|
27
|
MP RutkowskiCA KlankeYR SuGenetic markers
at the leptin (OB) locus are not significantly linked to
hypertension in African
AmericansHypertension3112301234199810.1161/01.HYP.31.6.12309622134
|
|
28
|
JS KuafmanRA Durazo-ArvizuCN RotimiDL
McGeeRS CooperObesity and hypertension prevalence in populations of
African origin. The Investigators of the International
Collaborative Study on Hypertension in
BlacksEpidemiology7398405199610.1097/00001648-199607000-000108793366
|
|
29
|
TB Van ItallieHealth implications of
overweight and obesity in the United StatesAnn Intern
Med1039839881985
|
|
30
|
CO JacobNB TashmanDisruption in the AU
motif of the mouse TNF-alpha 3′ UTR correlates with reduced TNF
production by macrophages in vitroNucleic Acids
Res212761276619938332472
|
|
31
|
CM MisquittaVR IyerES WerstiukAK GroverThe
role of 3′-untranslated region (3′-UTR) mediated mRNA stability in
cardiovascular pathophysiologyMol Cell Biochem22453672001
|
|
32
|
MK KarvonenU PesonenP
HeinonenIdentification of new sequence variants in the leptin geneJ
Clin Endocrinol
Metab8332393242199810.1210/jcem.83.9.51359745435
|
|
33
|
U SchorrK BlaschkeS TuranA DistlerAM
SharmaRelationship between angiotensinogen, leptin and blood
pressure levels in young normotensive menJ
Hypertens1614751480199810.1097/00004872-199816100-000119814618
|
|
34
|
G UckayaM OzataA SonmezPlasma leptin
levels strongly correlate with plasma renin activity in patients
with essential hypertensionHorm Metab
Res31435438199910.1055/s-2007-97876910450836
|
|
35
|
J AubertC DarimontI SafonovaG AilhaudR
NegrelRegulation by glucocorticoids of angiotensinogen gene
expression and secretion in adipose cellsBiochem
J32870170619979371734
|
|
36
|
BH JonesMK StandridgeJW TaylorN
MoustaidAngio-tensinogen gene expression in adipose tissue:
analysis of obese models and hormonal and nutritional controlAm J
Physiol273R236R24219979249555
|