Introduction
Helicobacter pylori (H. pylori)
infection is a predominant cause of gastristis and peptic ulcer
disease, as well as an early risk factor for gastric cancer. It is
a globally epidemic pathogenic bacteria and more than half of the
world’s population has been infected with this bacterium. H.
pylori infection is common; however, its precise mode of
transmission has not yet been identified. The main reason for this
is that the etiology of H. pylori has not been solved
effectively. Therefore restrictng the formulation of prevention
measures against H. pylori.
A biofilm is a multicellular layer of bacteria,
which attaches to surfaces, interfaces or other cells and is
embedded within a matrix of extracellular polymeric substances
under stress conditions (1).
Growth in a biofilm provides a number of advantages for bacteria,
including enhanced power of resistance to environmental stresses as
well as to host defenses, which leads to serious clinical problems,
particularly chronic and refractory infection. The biofilm of H.
pylori has been observed in the water system, including
drinking water, groundwater and sewage (2,3) and
also on the human gastric mucosal epithelium (4,5). A
number of studies have been performed on H. pylori biofilms
(6,7). Thus, biofilms are hypothesized to be
another adaptive survival mode of H. pylori. In conventional
experiments, H. pylori is cultured in a rich medium
containing fetal bovine serum (FBS). However, this may not
effectively imitate the in vitro and in vivo survival
environment of H. pylori. Nutrient starvation is a common
environmental pressure that H. pylori is faced with. Serum
starvation effectively mimics the microenvironment H. pylori
within healthy human carriers of H. pylori and patients with
chronic atrophic gastritis (8) and
also the in vitro pressure environment pressure that H.
pylori is subjected to. The current study identified that H.
pylori adhered to each other and formed a population structure
similar to that of a bacterial biofilm by removing serum from
medium. Williams et al(9)
observed that serum significantly affects the movement, morphology
and adhesion of H. pylori and may result in agglutination in
1% serum concentration. Campylobacter jejuni also
agglutinate in nutrient-limiting conditions. Reeser et
al(10) observed that the
autoagglutinated structure is a biofilm by analyzing its
extracellular polymer matrix.
The formation of a biofilm is a complex and dynamic
process. Under adverse conditions, the bacteria induce a unique set
of genes, adhere to each other and form structural groups or
subgroups. During biofilm formation, each link may be regarded as a
key point to treat infection and drug design target, particularly
at the early stage. In the present study, proteomic technologies
were used to analyze the proteomic expression profiles of H.
pylori biofilms at an early stage, and aimed to reveal the
proteins and signaling molecules involved in H. pylori
biofilm formation and identify novel pathogenic factors and
antibacterial targets.
Materials and methods
Bacterial strain and culture
conditions
The H. pylori strain 26695 was provided by Dr
Zhang Jianzhong from the Chinese Disease Control and Prevention
Center. It was cultured in Brucella broth with 10% FBS with 120 rpm
agitation under microaerobic conditions (5% O2, 10%
CO2, 85% N2, v/v) at 37°C. The same quantity
of exponentially growing H. pylori was resuspended in
Brucella broth with and without 10% FBS. Bacterial morphological
changes were then observed using Gram staining at intervals of 2 h
until the autoagglutinated structure formed. The initial time of
biofilm formation was recorded. Subsequently, the planktonic H.
pylori and the biofilm were fixed with 2.5% glutaraldehyde
solution for >2 h. The critical-point dried sample was observed
with a Hitachi S-520 scanning electron microscope (Hitachi Limited
Company, Tokyo, Japan).
2-dimentional gel electrophoresis (2-DE)
and image analysis
The planktonic H. pylori and its early
biofilm cultured with and without 10% FBS for 4 h were harvested by
centrifugation at 5000 × g for 10 min at 4°C and washed three times
with ice-cold PBS (pH 7.2). The pellets were solubilized in lysis
buffer containing 8 M urea, 2 M thiourea, 4% CHAPS, 1% DTT, 1%
pharmalyte (pH range, 3–10), 1% protease inhibitor and 1% nuclease
mix (Amersham Pharmacia Biotech, Piscatawway, NJ, USA). Following
sonication, the solution was centrifugated at 20,000 × g for 60 min
at 4°C to remove cell debris. Protein concentrations were
determined using the Bradford method and proteins were stored at
−80°C. The 2-DE analysis was performed as described previously
(11). The SDS gels were
silver-stained and the protein patterns were scanned using an
ImageScannerII (Amersham Pharmacia Biotech). The gels were analyzed
by ImageMaster 2D Elite 5.0 (Amersham Pharmacia Biotech) to
determine differentially expressed protein spots whose vol% ratio
increased >2-fold (P<0.05).
In-gel digestion and matrix assisted
laser desorption ionzation time of flight mass spectrometry
(MALDI-TOF-TOF MS)
Differentially expressed protein spots were excised,
tryptic digested and identified with a 4700 MALDI-TOF-TOF proteomic
system (Applied Biosystems) as described previously (11). The MS and MS/MS spectra were
analyzed with a 50 ppm mass tolerance by GPS Explorer V.2.0.1 and
Mascot V1.9 based on the NCBI and SWISSPROT databases.
Quantitative PCR
Total RNA was isolated from three independent
cultures of planktonic H. pylori and biofilm at early stages
using TRIzol reagent (Invitrogen Life Technologies, Carlsbad, CA,
USA) according to the manufacturer’s instructions. The quantity was
measured by absorbance at 260 nm. Subsequently, 4 μg total template
RNA was reverse transcribed into cDNA using M-MLV reverse
transcriptase and random hexamer primer (MBI). The primers for PCR
are listed in Table I. The 16S
rRNA gene served as the endogenous control. PCR was performed as
described previously (12). The
operating conditions were as follows: One cycle at 50°C for 2 min,
one cycle at 95°C for 10 min, 40 cycles at 95°C for 15 sec and 60°C
for 1 min. Gene expression was quantified by the comparative
threshold cycle (Ct) method and normalized to the quantity of 16S
rRNA cDNA in each sample. The relative quantity of the target was
calculated by the ΔΔCT-method.
 | Table IPrimers used in the study |
Table I
Primers used in the study
| Primers | Sequence
(5′-3′) |
|---|
| cagA Forward |
GCTTACCGCCTGAAGCTAGG |
| cagA Reverse |
CCTTTCTCACCACCTGCTATG |
| 16S rRNA
Forward |
GCTCTTTACGCCCAGTGATTC |
| 16S rRNA
Reverse |
GCGTGGAGGATGAAGGTTTT |
Western blot analysis
Protein samples were separated by sodium dodecyl
sulfate (SDS)-polyacrylamide gel electrophoresis (8% SDS-acrylamide
gels) and transferred to a nitrocellulose membrane. The membranes
were incubated in blocking buffer [TBS containing 0.1% Tween-20
(TBST) and 5% non-fat powdered milk] for 1.5 h and immunoblotted
for 1.5 h with antibodies against β-actin or C-CagA (Santa Cruz
Biotechnology, Inc. Santa Cruz, CA, USA). Membranes were washed
three times with the TBST solution and incubated with the
corresponding antibodies conjugated with horseradish peroxidase for
50 min. The washed membranes were developed by the Chemilucent ECL
Detection system (Millipore, Billerica, MA, USA).
Results and Discussion
In recent years, increasing evidence has indicated
that bacterial biofilms are important in infectious diseases.
Biofilms effectively protect bacteria from external environmental
damage and are an effective strategy to adapt to environmental
pressures in vivo and in vitro. As one of the
predominant pathogens affecting humans, H. pylori was
subjected to various types of adverse conditions, including
nutrient starvation. To produce the survival environment H.
pylori generated when under starvation stress, exponentially
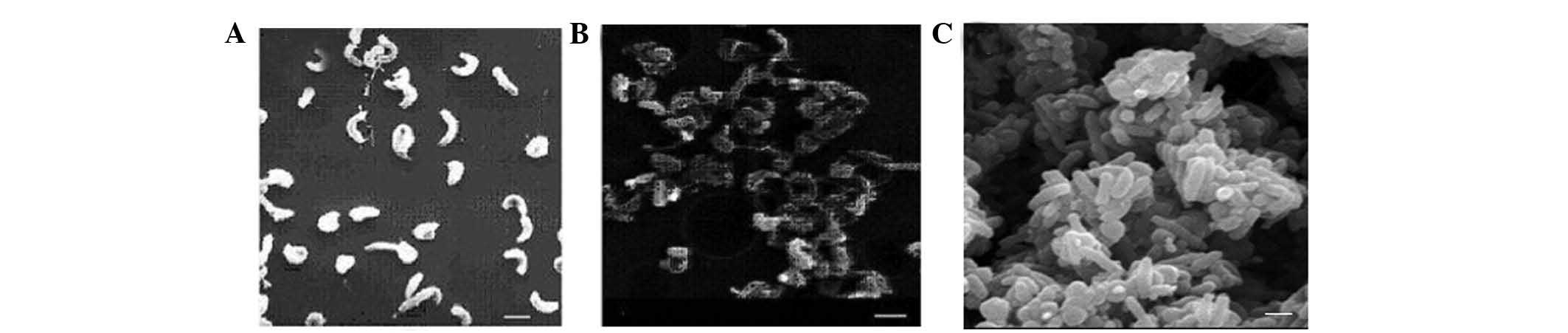
growing H. pylori 26695(Fig.
1A) was inoculated into medium without FBS. Following 4 h, the
bacteria began to adhere to each other and formed a subgroup
structure (Fig. 1B). Typical
community structures were observed following 12 h starvation and a
number of H. pylori changed to a dormant coccoid form
(Fig. 1C). Thus, a biofilm of
H. pylori was successfully induced with serum
starvation.
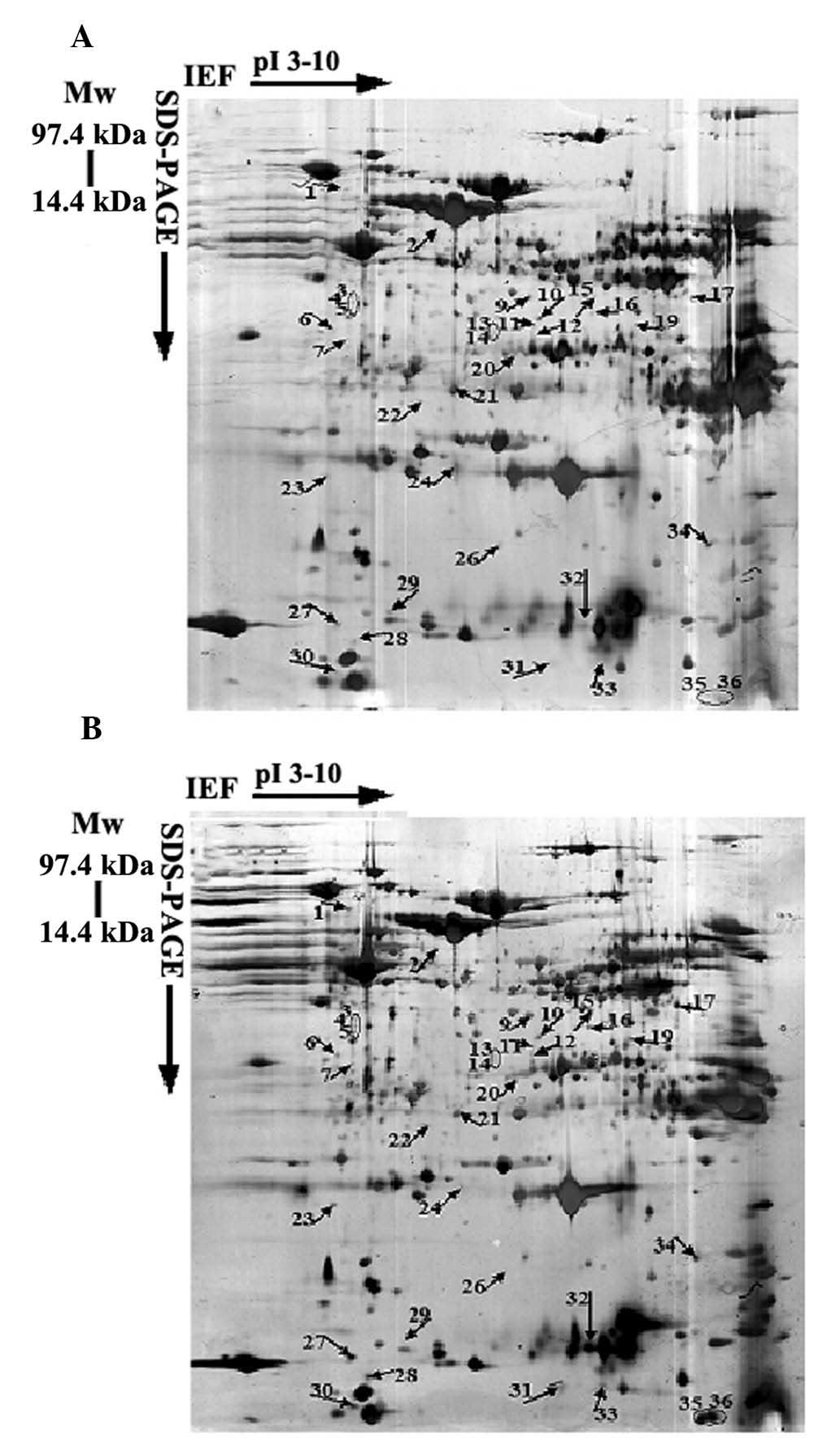
To identify novel proteins associated with H.
pylori biofilm formation, high-resolution 2-DE was performed to
obtain the proteome profiles of spiral H. pylori and its
early biofilm, which was induced by 4 h of starvation (Fig. 2). 2-DE analysis was repeated in
triplicate using independently grown cultures. The comparison of
the protein patterns of these two forms of H. pylori
revealed that 36 spots exhibited different levels of expression
(>2-fold, P<0.05). These spots were subjected to
MALDI-TOF-TOF MS analysis to determine their putative genes and
infer their functions. A total of 35 proteins with different
functions were identified (Table
II); of which 29 were upregulated and six were downregulated in
the early biofilm of H. pylori. These proteins belong to
diverse functional classes, including movement, virulence, energy
metabolism, regulator and chaperone proteins. This suggests that
H. pylori invokes a multi-mechanism response to adapt to
morphological changes under starvation stress.
 | Table IISummary of protein spots showing
altered expression between planktonic H. pylori and its
early biofilm. |
Table II
Summary of protein spots showing
altered expression between planktonic H. pylori and its
early biofilm.
| Spot numbera | Protein (gene) | TIGR ORF
numberb | Gi | Top score | %vol ratioc |
|---|
| 1 | Cag pathogenicity
island protein (cag26) | Hp0547 | 2313664 | 139 | 2.89 |
| 2 | Flagellar capping
protein(fliD) | Hp0752 | 2313869 | 85 | 2.21 |
| 3 | Aconitase B
(acnB) | Hp0779 | 2313908 | 105 | 2.36 |
| 4 | Aconitase B
(acnB) | Hp0779 | 2313908 | 133 | 2.13 |
| 6 | Type I restriction
enzyme R protein (hsdR) | Hp0846 | 2313977 | 46 | 2.05 |
| 6 | Urease β-subunit
(urea amidohydrolase) (ureB) | Hp0072 | 2313153 | 45 | 2.05 |
| 6 | Arginase
(rocF) | Hp1399 | 2314565 | 50 | 2.05 |
| 7 | H. pylori
predicted coding region HP0013 | Hp0013 | 2313087 | 64 | 2.02 |
| 9 | Transaldolase
(tal) | Hp1495 | 2314674 | 175 | 2.04 |
| 10 | Holliday junction
DNA helicase (ruvB) | Hp1059 | 2314203 | 161 | 2.04 |
| 10 | GTP-binding
protein, fusA-homolog (yihK) | Hp0480 | 2313589 | 75 | 2.04 |
| 11 |
UDP-N-acetylenolpyruvoylglucosamine
reductase (murB) | Hp1418 | 2314592 | 69 | 2.15 |
| 12 | Chaperone and heat
shock protein (groEL) | Hp0010 | 2313084 | 106 | 7.45 |
| 13 | Poly E-rich
protein | Hp0322 | 2313421 | 64 | 2.14 |
| 14 | DNA-directed RNA
polymerase, β-subunit (rpoB) | Hp1198 | 2314357 | 64 | 2.26 |
| 15 | GTP-binding protein
(obg) | Hp0303 | 2313401 | 128 | 3.24 |
| 15 |
UDP-N-acetylenolpyruvoylglucosamine
reductase (murB) | Hp1418 | 2314592 | 61 | 3.24 |
| 16 |
Delta-aminolevulinic acid dehydratase
(hemB) | Hp0163 | 2313250 | 111 | 2.13 |
| 17 |
Aspartate-semialdehyde dehydrogenase
(asd) | Hp1189 | 2314350 | 257 | 2.31 |
| 19 | Aspartyl-tRNA
synthetase (aspS) | Hp0617 | 2313739 | 65 | 3.16 |
| 20 | H. pylori
predicted coding region HP0958 | Hp0958 | 2314102 | 90 | 4.76 |
| 21 | Conserved
hypothetical protein | Hp1588 | 2314773 | 73 | 0.32 |
| 21 | 3-oxoadipate
coA-transferase subunit A (yxjD) | Hp0691 | 2313815 | 68 | 0.32 |
| 22 | Response regulator
(ompR) | Hp0166 | 2313252 | 105 | 2.07 |
| 23 | H. pylori
predicted coding region HP0406 | Hp0406 | 2313513 | 131 | 2.01 |
| 24 | Alkyl hydroperoxide
reductase (AhpC) | Hp1563 | 2314747 | 146 | 0.31 |
| 26 | Cag pathogenicity
island protein (cag24) | Hp0504 | 2313660 | 77 | 0.44 |
| 26 | Translation
elongation factor EF-Tu (tufB) | Hp1205 | 2314366 | 45 | 0.44 |
| 27 | Conserved
hypothetical protein | Hp1046 | 2314193 | 76 | 2.57 |
| 27 | Hemolysin secretion
protein precursor (hylB) | Hp0599 | 2313716 | 49 | 2.57 |
| 28 | Riboflavin synthase
beta chain (ribE) | Hp0002 | 2313079 | 121 | 2.66 |
| 29 | Nonheme
iron-containing ferritin (pfr) | Hp0653 | 2313771 | 82 | 0.03 |
| 30 | Hydrogenase
expression/formation protein (hypA) | Hp0869 | 2313996 | 104 | 4.37 |
| 31 | H. pylori
predicted coding region HP1377 | Hp1377 | 2314555 | 111 | 2.55 |
| 32 | Co-chaperone
(groES) | Hp0011 | 2313085 | 82 | 5.08 |
| 34 | H. pylori
predicted coding region HP1029 | Hp1029 | 2314185 | 142 | 2.41 |
| 35 | Thioredoxin | Hp1458 | 2314636 | 97 | 81.74 |
| 36 | Thioredoxin | Hp1458 | 2314636 | 126 | 2.07 |
The adhesion of bacteria is the early stages of
biofilm formation and the flagellar movement are important in this
process. The present study identified that the flagellar filament
capping protein (HAP2) was increased in the early biofilm of H.
pylori. This protein is encoded by the fliD gene, constitutes
the flagellar hook together with HAP1 and HAP3, and is involved in
the flagellar assembly of H. pylori. Martin et al
observed that proteins associated with flagella exhibited higher
expression in the biofilm of Campylobacter and that the
deletion of flagellar genes may seriously affect the biofilm
formation of bacteria (13,10).
In Escherichia coli (E. coli), flagellar
motility may significantly affect the three-dimensional structure
of the biofilm (14). In addition,
aconitase was observed to be upregulated in H. pylori early
biofilms. The structural analysis identified that this protein
contained a domain similar to the HEAT domain, which mediates the
interaction between proteins. In E. coli and Bacillus
subtilis, the HEAT protein has been confirmed to have a dual
function and regulate flagellar movement as a post-transcriptional
regulator under iron and oxygen conditions (15,16).
Under serum starvation conditions, the overexpression of aconitase
in H. pylori is hypothesized to be involved in biofilm
formation by the promotion of flagellar movement. However, this
requires further study to be confirmed.
In present study, the cagA protein encoded by cag
pathogenicity islands was identified to be induced in H.
pylori biofilms. As an important virulence factor of H.
pylori, this protein is injected into gastric mucosa epithelial
cells through the type IV secretion system when the cagA positive
H. pylori infect the individual. CagA is then localized in
the inner surface of the plasma membrane and results in a series of
changes, including excessive proliferation and phenotypic change,
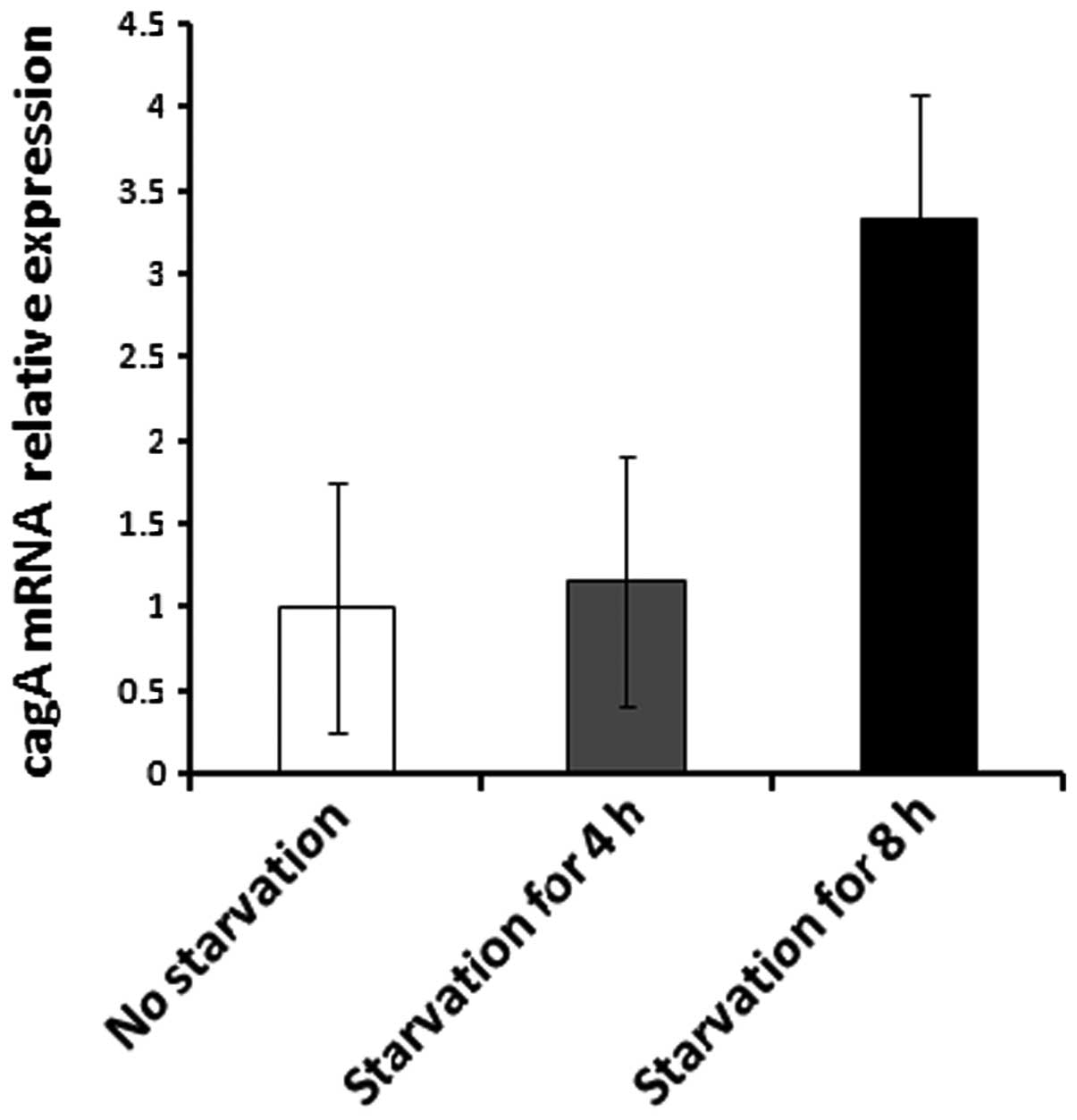
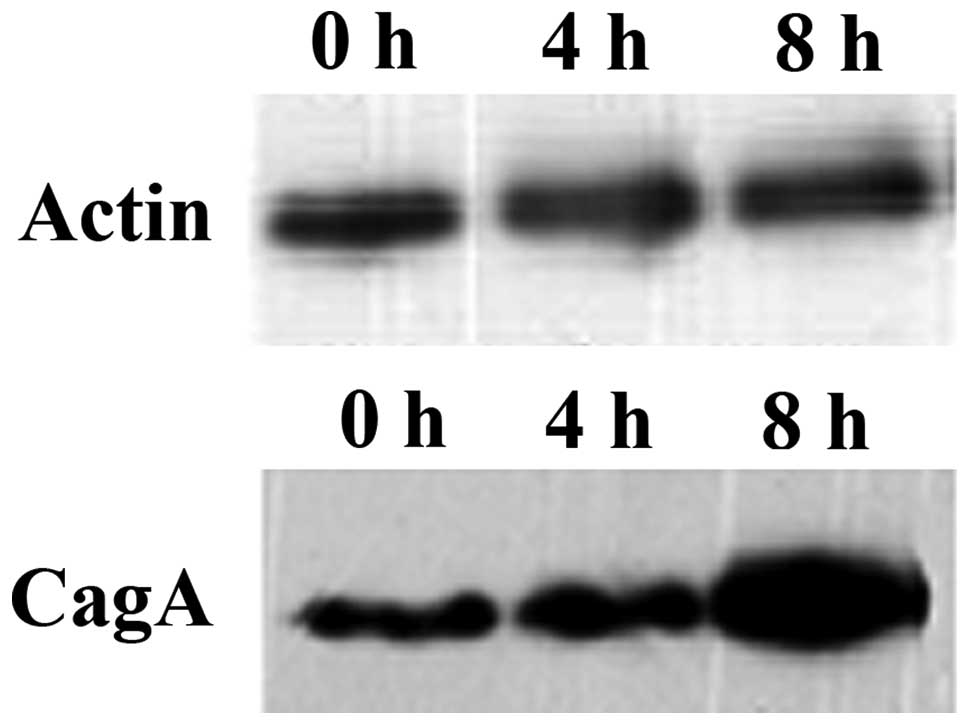
such as extended ‘hummingbird’ morphology (17). To confirm the results, the
expression of cagA was examined at the mRNA and protein levels by
quantitative PCR and western blot analysis, respectively. The
results showed that the expression of cagA was not significantly
changed following 4 h of starvation. This may be due to higher
resolution of the 2-DE compared with 1-D SDS. However, after 8 h
cagA expression was upregulated significantly (Figs. 3 and 4). Thus the upregulation of this protein
was hypothesized to aid in the improvement of the viability of
H. pylori under adverse environments.
UspA is a conservative and integral promoter of a
number of stress-associated proteins in a variety of bacteria. The
β-subunit of RNA polymerase positively regulates the expression of
UspA under a number of environmental pressures, including
starvation. Thus, a variety of stress-related proteins show higher
expression levels and bacterial biofilms have strong resistance to
the outside pressure (18). In the
current study, the β-subunit of RNA polymerase and two chaperone
protein GroES and GroEL of H. pylori were observed to be
overexpressed under serum starvation. In addition, two oxidative
stress-related proteins, alkyl hydroperoxide reductase and
thioredoxin, were identified to have a high level of expression in
the biofilm of H. pylori. This is consistent with the
biofilms of E. coli and Campylobacter(13,19).
The two-component system is a common signal
transduction mechanism in bacterial responses to various stresses.
This system is composed of two types of proteins, a histidine
kinase and a response regulator. These two components exchange
signals by phosphate transfer. Two-component systems are involved
in the biofilm formation of E. coli, Pseudomonas
aeruginosa and other bacteria (20,21).
For example, the EnvZ/OmpR two-component system in E. coli
promotes biofilm formation by regulating the expression of proteins
associated with bacterial fimbriae to enhance its adhesion
(22). In the present study, the
response regulator Hp0166 was observed to be induced in H.
pylori biofilms. A previous study also demonstrated that the
Ars two-component system is involved in H. pylori responses
to acid stress by regulating the expression of a variety of genes
(23).
In addition, the current study also identified that
the expression of other proteins changed in H. pylori
biofilms. The expression of urease and arginase was upregulated.
These two proteins generate ammonia in the process of amino acid
metabolism, which is essential to biofilm formation to maintain the
pH balance. The accessory protein of urease, ureE, is highly
expressed in Staphylococcus aureus biofilms (24). Peptidoglycan is a predominant
component of the cell walls of bacteria, UDP-N acetylene alcohol
acid glucosamine reductase is an enzyme that is required in
peptidoglycan biosynthesis. It catalyzes the reduction of UDPN
GlcNAc to UDPN acetyl muramic acid (25). This protein was overexpressed in
H. pylori biofilms and it is consistent with a previous
study of Staphylococcus aureus(14). The function of peptidoglycan
remains unclear, but it was observed that fracture of the
peptidoglycan may promote the formation of bacterial biofilms
(26). In addition, this study
also showed that the expression of a number of other unknown
proteins, including Hp1377 and Hp1029, changed in H. pylori
biofilm formation. However, this also requires further study.
In the present study, H. pylori biofilm
formation was induced successfully by serum starvation and its
proteome was analyzed using proteomic methods. This study showed
that ≥35 proteins are involved in biofilm formation. These proteins
are associated with a number of types of biological functions,
including flagellar movement, bacterial virulence, signal
transduction and regulation. Such results showed that H.
pylori biofilms are formed through multiple mechanisms
involving numerous signal pathways. These findings may provide
valuable information in understanding the survival mechanism of
this bacterium in animals and humans.
Acknowledgements
This study was supported by grants from the National
Natural Science Foundation of China (grant nos. 81171536 and
30800037) and the Science Foundation of Shandong Province, PR China
(grant nos. ZR2009CZ001 and ZR2009CM002).
References
|
1
|
Donlan RM and Costerton JW: Biofilms:
survival mechanisms of clinically relevant microorganisms. Clin
Microbiol Rev. 15:167–193. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Moreno Y and Ferrús MA: Specific detection
of cultivable Helicobacter pylori cells from wastewater
treatment plants. Helicobacter. 17:327–332. 2012.
|
|
3
|
Braganra SM, Azevedo NF, Simões LC, Keevil
CW and Vieira MJ: Use of fluorescent in situ hybridisation for the
visualisation of Helicobacter pylori in real drinking water
biofilms. Water Sci Technol. 55:387–393. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Cammarota G, Sanguinetti M, Gallo A and
Posteraro B: Review article: biofilm formation by Helicobacter
pylori as a target for eradication of resistant infection.
Aliment Pharmacol Ther. 36:222–230. 2012.PubMed/NCBI
|
|
5
|
Carron MA, Tran VR, Sugawa C and Coticchia
JM: Identification of Helicobacter pylori biofilms in human
gastric mucosa. J Gastrointest Surg. 10:712–717. 2006.
|
|
6
|
Yang FL, Hassanbhai AM, Chen HY, Huang ZY,
Lin TL, Wu SH and Ho B: Proteomannans in biofilm of Helicobacter
pylori ATCC 43504. Helicobacter. 16:89–98. 2011.
|
|
7
|
Grande R, Di Giulio M, Bessa LJ, Di Campli
E, Baffoni M, Guarnieri S and Cellini L: Extracellular DNA in
Helicobacter pylori biofilm: a backstairs rumour. J Appl
Microbiol. 110:490–498. 2011.
|
|
8
|
Testerman TL, Conn PB, Mobley HL and McGee
DJ: Nutritional requirements and antibiotic resistance patterns of
Helicobacter species in chemically defined media. J Clin
Microbiol. 44:1650–1658. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Williams JC, McInnis KA and Testerman TL:
Adherence of Helicobacter pylori to abiotic surfaces is
influenced by serum. Appl Environ Microbiol. 74:1255–1258.
2008.
|
|
10
|
Reeser RJ, Medler RT, Billington SJ, Jost
BH and Joens LA: Characterization of Campylobacter jejuni
biofilms under defined growth conditions. Appl Environ Microbiol.
73:1908–1913. 2007.
|
|
11
|
Shao C, Zhang Q, Sun Y, et al:
Helicobacter pylori protein response to human bile stress. J
Med Microbiol. 57:151–158. 2008. View Article : Google Scholar
|
|
12
|
Shao C, Zhou Y, Sun Y, et al: Analysis of
aztreonam-inducing proteome changes in nondividing filamentous
Helicobacter pylori. Curr Microbiol 2012. 65:108–115. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kalmokoff M, Lanthier P, Tremblay TL, et
al: Proteomic analysis of Campylobacter jejuni 11168
biofilms reveals a role for the motility complex in biofilm
formation. J Bacteriol. 188:4312–4320. 2006.
|
|
14
|
Wood TK, González Barrios AF, Herzberg M
and Lee J: Motility influences biofilm architecture in
Escherichia coli. Appl Microbiol Biotechnol. 72:361–367.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tang Y, Guest JR, Artymiuk PJ and Green J:
Switching aconitase B between catalytic and regulatory modes
involves iron-dependent dimer formation. Mol Microbiol.
56:1149–1158. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tang Y, Guest JR, Artymiuk PJ, Read RC and
Green J: Post-transcriptional regulation of bacterial motility by
aconitase proteins. Mol Microbiol. 51:1817–1826. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Backert S, Moese S, Selbach M, Brinkmann V
and Meyer TF: Phosphorylation of tyrosine 972 of the
Helicobacter pylori CagA protein is essential for induction
of a scattering phenotype in gastric epithelial cells. Mol
Microbiol. 42:631–644. 2001.PubMed/NCBI
|
|
18
|
Kvint K, Hosbond C, Farewell A, Nybroe O
and Nyström T: Emergency derepression: stringency allows RNA
polymerase to override negative control by an active repressor. Mol
Microbiol. 35:435–443. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Schembri MA, Kjaergaard K and Klemm P:
Global gene expression in Escherichia coli biofilms. Mol
Microbiol. 48:253–267. 2003.
|
|
20
|
Li YH, Lau PC, Tang N, Svensäter G, Ellen
RP and Cvitkovitch DG: Novel two-component regulatory system
involved in biofilm formation and acid resistance in
Streptococcus mutans. J Bacteriol. 184:6333–6342. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Otto K and Silhavy TJ: Surface sensing and
adhesion of Escherichia coli controlled by the Cpx-signaling
pathway. Proc Natl Acad Sci USA. 99:2287–2292. 2002.PubMed/NCBI
|
|
22
|
Prigent-Combaret C, Brombacher E, Vidal O,
Ambert A, Lejeune P, Landini P and Dorel C: Complex regulatory
network controls initial adhesion and biofilm formation in
Escherichia coli via regulation of the csgD gene. J
Bacteriol. 183:7213–7223. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wen Y, Feng J, Scott DR, Marcus EA and
Sachs G: Involvement of the HP0165-HP0166 two-component system in
expression of some acidic-pH-upregulated genes of Helicobacter
pylori. J Bacteriol. 188:1750–1761. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Resch A, Leicht S, Saric M, Pásztor L,
Jakob A, Götz F and Nordheim A: Comparative proteome analysis of
Staphylococcus aureus biofilm and planktonic cells and
correlation with transcriptome profiling. Proteomics. 6:1867–1877.
2006.
|
|
25
|
Andres CJ, Bronson JJ, D’Andrea SV, et al:
4-Thiazolidinones: novel inhibitors of the bacterial enzyme MurB.
Bioorg Med Chem Lett. 10:715–717. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Mercier C, Durrieu C, Briandet R, Domakova
E, Tremblay J, Buist G and Kulakauskas S: Positive role of
peptidoglycan breaks in lactococcal biofilm formation. Mol
Microbiol. 46:235–243. 2002. View Article : Google Scholar : PubMed/NCBI
|