Introduction
Multiple sclerosis (MS) is an inflammatory
demyelinating disease (IDD) caused by damage to the myelin sheaths
of axons in the central nervous system, leading to demyelination
and scarring (1). A patient with
MS may suffer from a broad spectrum of signs or symptoms,
including, but not limited to, changes in sensation, muscle
weakness, ataxia, difficulties with speech, swallowing and vision,
fatigue and chronic pain. Psychological symptoms such as depression
and unstable mood are also commonly observed. The onset of MS
usually occurs in young adults, and more often in females than in
males. The cause of the disease is not fully understood, but
researchers agree that a combination of genetic and environmental
factors are responsible for the development of the disease
(2). Advances in the field of
genetics research have led to the identification of various genes
related to MS. One of the most well-known genetic regions that
affects MS is the human leukocyte antigen (HLA)
region in chromosome 6. However, this region explains only a
fraction of MS genetic etiology (2). In order to investigate the genetic
factors of MS, a number of studies have searched for risk genes
using the latest technology. As a result, several genes, such as
interleukin-7 receptor (IL7R), interleukin-2
receptor α (IL2RA), glypican-5 (GPC5),
cluster of differentiation 6 (CD6) and tumor necrosis
factor receptor superfamily member 1 α (TNFRSF1A) have
been found to be associated with MS risk (3–6).
Neuromyelitis optica (NMO) is another type of IDD
that particularly affects the optic nerve and spinal cord, leading
to optic neuritis and demyelination. Although its signs and
symptoms overlap with MS in certain ways, evidence from
neuroimaging and laboratory findings indicate that NMO etiology is
different from that of MS (7,8). In
addition, while MS is an uncommon disease in Asian populations, NMO
is more prevalent in Asians when compared with MS (9–11).
Although numerous studies have been conducted on the association
between MS and genetic polymorphisms, studies on correlations
between NMO and polymorphisms are less common. We previously
conducted a genome-wide association study (GWAS) for NMO and MS,
which showed that the risk polymorphisms for NMO were different
from those of MS (12).
IL7R, located on the surface of immune cells, is a
heterodimer known to be important in the development of lymphocytes
(13). This protein also controls
the accessibility of the T-cell receptor γ gene (14). Thus, several studies have
investigated a possible association between genetic polymorphisms
of IL7R and MS. A GWAS revealed that polymorphisms of
IL7R were associated with MS (6), and several follow-up studies have
confirmed this association in different ethnic populations,
including European, Australian, American and Japanese populations
(6,15–17).
In this study, we examined the association analysis
between IL7R polymorphisms and IDD, including MS and NMO, in
a Korean population. Additionally, we conducted a meta-analysis of
MS studies carried out in various populations to compare and
contrast the effects of IL7R polymorphisms.
Materials and methods
Subjects
For the genotyping of IL7R polymorphisms, a
total of 415 patients were recruited, including 98 NMO patients, 80
MS patients (178 IDD patients in total) and 237 normal control
patients. Individuals with each disease were evaluated and invited
to participate in the study at the MS centers of the Asian Medical
Center, Ewha Woman’s University Medical Center and National Cancer
Center of Korea from July 2006 to September 2007. Thorough
attention was given to age, gender, disease duration, age at
disease onset and assessment of disease severity using the Expanded
Disability Status Scale (18). In
addition, 237 healthy and elderly controls of Korean ethnicity were
included who had not suffered from IDDs, including NMO, classical
MS or idiopathic recurrent transverse myelitis. The study protocol
was approved by the Institutional Review Board of the National
Cancer Center of Korea. Written informed consent was obtained from
each subject prior to initiation of the study.
Single-nucleotide polymorphism (SNP)
selection and genotyping
Thirteen SNPs of IL7R were selected based on
linkage disequilibrium (LD), minor allele frequency (MAF)
(>0.05), locations (SNPs in exons were preferred) and amino acid
changes (non-synonymous SNPs were preferred) from the Asian
(Chinese and Japanese) population database of the International
HapMap Project (http://hapmap.ncbi.nlm.nih.gov/). The selected SNPs
were then genotyped in 178 IDD cases and 237 normal control
subjects using a TaqMan assay from the ABI prism 7900HT sequence
detection system (Applied Biosystems, Foster City, CA, USA).
Genotyping quality control was performed in 10% of the samples by
duplicate checking (rate of concordance in duplicates
>99.5%).
Statistical analysis
The LD was obtained using the Haploview v4.2
software from the Broad Institute (http://www.broadinstitute.org/mpg/haploview), with
examination of Lewontin’s D′ (|D′|) and the LD coefficient
r2 between all the pairs of bi-allelic
loci (19). Haplotypes were first
estimated using PHASE software (20), and then computed using a
Statistical Analysis System (SAS). Associations for IDD, MS and NMO
in a logistic model were adjusted for age (continuous value) and
gender (male was 0, female was 1) as covariates, using SAS. In
order to correct for the multiple testing error, the Single
Nucleotide Polymorphism Spectral Decomposition program (http://gump.qimr.edu.au/general/daleN/SNPSpD/) was
used, with the correction number of 9.4353. The meta-analysis was
conducted with the R program package ‘meta’. Comparisons between
ethnic groups were conducted with an SAS using a Chi-square test,
and an LD plot of the ethnic groups was obtained using the
Haploview software. P<0.05 was considered to indicate a
statistically significant difference.
Results
Subjects and IL7R characteristics
A total of 415 subjects were enrolled in the present
study: 178 IDD patients, which included 80 MS patients and 98 NMO
patients, and 237 normal controls. Information about the subjects,
including age, gender, age of disease onset and duration of disease
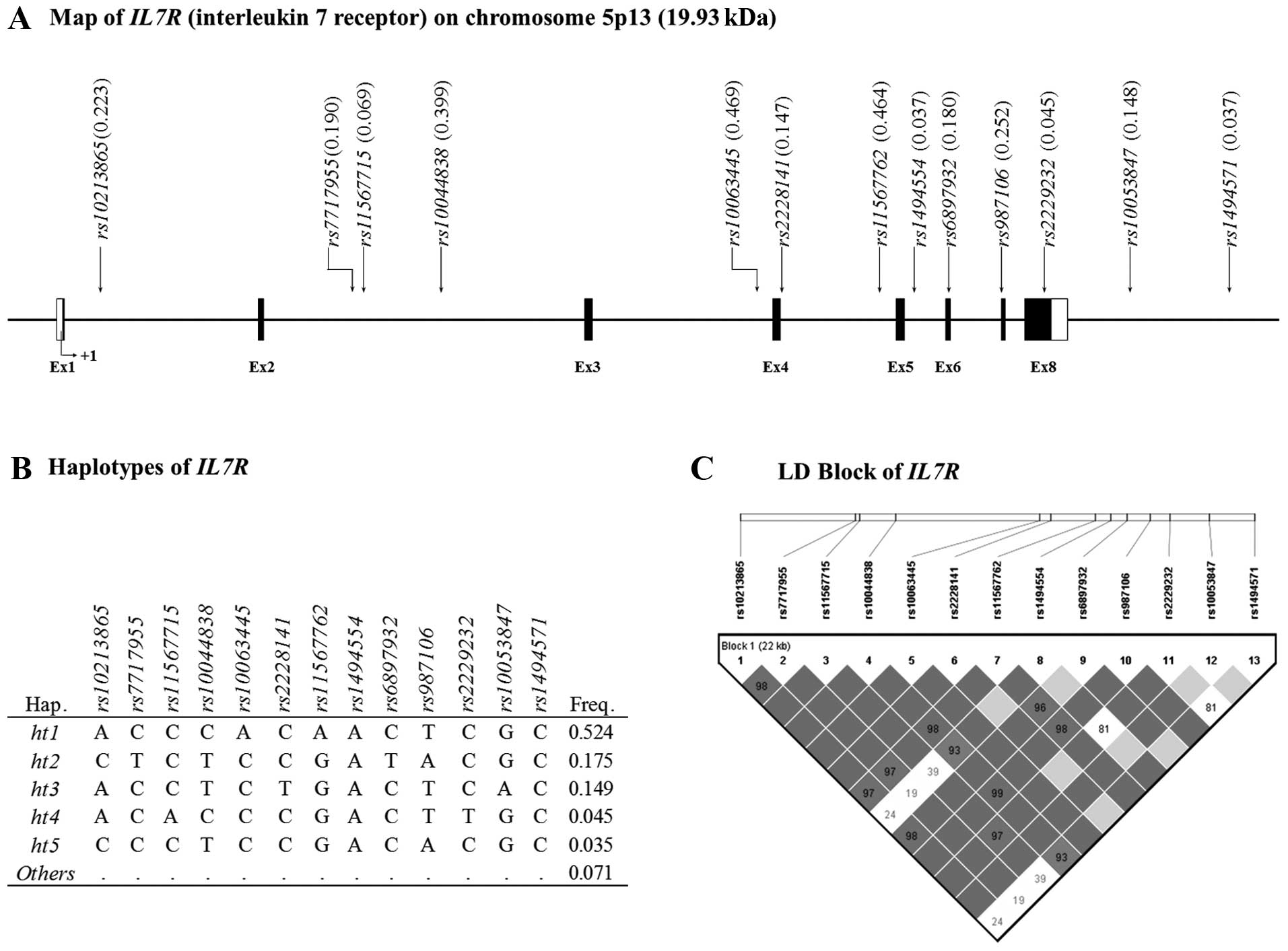
is listed in Table I. A physical
map of IL7R and the location of the SNPs are shown in
Fig. 1A. In addition, information
on five common haplotypes and the LD block of IL7R are shown
in Fig. 1B and C, respectively.
The LD block of IL7R showed that the SNPs formed one tight
block. Detailed information on the 13 SNPs selected from
IL7R is listed in Table
II. All the SNPs had an MAF >0.05 and none of the SNPs broke
the Hardy-Weinberg equilibrium in the case, control or total
populations.
 | Table IClinical characteristics of
subjects. |
Table I
Clinical characteristics of
subjects.
| Characteristics | NMO | MS | IDD (NMO+MS) | NC |
|---|
| No. of subjects | 98 | 80 | 178 | 237 |
| Age [mean
(min-max)] | 39.9 (11–67) | 34.3 (14–57) | 37.0 (11–67) | 47.3 (38–60) |
| Gender
(male/female) | 10/88 | 29/51 | 39/139 | 81/156 |
| Onset age (age, mean
± SD) | 33.4±12.38 | 30.08±10.23 | 31.99±11.50 | - |
| Duration (year, mean
± SD) | 7.0±4.40 | 4.45±3.59 | 5.86±4.26 | - |
 | Table IIGenotype distributions and allele
frequencies of IL7R SNPs. |
Table II
Genotype distributions and allele
frequencies of IL7R SNPs.
| SNP | Position | Genotypes
(n=415) | MAF | Heterozygosity | HWE |
|---|
|
rs10213865 | Intron 1 | AA (247) | AC (142) | CC (20) | 0.222 | 0.346 | 0.944 |
|
rs7717955 | Intron 2 | CC (273) | CT (125) | TT (16) | 0.19 | 0.307 | 0.721 |
|
rs11567715 | Intron 2 | CC (359) | AC (53) | AA (3) | 0.071 | 0.132 | 0.502 |
|
rs10044838 | Intron 2 | CC (158) | CT (183) | TT (72) | 0.396 | 0.478 | 0.135 |
|
rs10063445 | Intron 3 | AA (118) | AC (205) | CC (92) | 0.469 | 0.498 | 0.868 |
|
rs2228141 | Exon 4 | CC (302) | CT (99) | TT (12) | 0.149 | 0.253 | 0.270 |
|
rs11567762 | Intron 4 | AA (119) | AG (199) | GG (90) | 0.464 | 0.497 | 0.693 |
|
rs1494554 | Intron 5 | AA (387) | AC (25) | CC (1) | 0.033 | 0.063 | 0.385 |
|
rs6897932 | Exon 6 | CC (279) | CT (121) | TT (14) | 0.180 | 0.295 | 0.843 |
|
rs987106 | Intron 6 | TT (238) | AT (150) | AA (27) | 0.246 | 0.371 | 0.609 |
|
rs2229232 | Exon 8 | CC (375) | CT (36) | TT (2) | 0.048 | 0.092 | 0.271 |
|
rs10053847 | 3′-UTR | GG (301) | AG (100) | AA (12) | 0.150 | 0.255 | 0.299 |
|
rs1494571 | 3′-UTR | CC (389) | CG (25) | GG (1) | 0.033 | 0.063 | 0.382 |
Association between rs6897932 and
IDD
We analyzed the 13 selected SNPs for the risk of
IDD, MS and NMO (Table III). The
analyses showed rs6897932 and haplotype 2
(ht2) to be significantly associated with IDD, even after
multiple-testing correction (Pcor.=0.03 and 0.04,
respectively). In order to compare the role of rs6897932 in
different populations, we listed studies carried out with
rs6897932 and conducted a meta-analysis (Table IV). There were certain exceptions
in European and Australian populations (21–23);
however, the majority of studies reported a significant correlation
between rs6897932 and MS (6,15–17,24–28).
This association was also detected in Asian populations, in which
the magnitude of risk was greater than in Caucasian populations
[odds ratio (OR)=0.47 and 0.54 in the two studies with Asian
populations and 0.82 in Caucasian populations]. We also
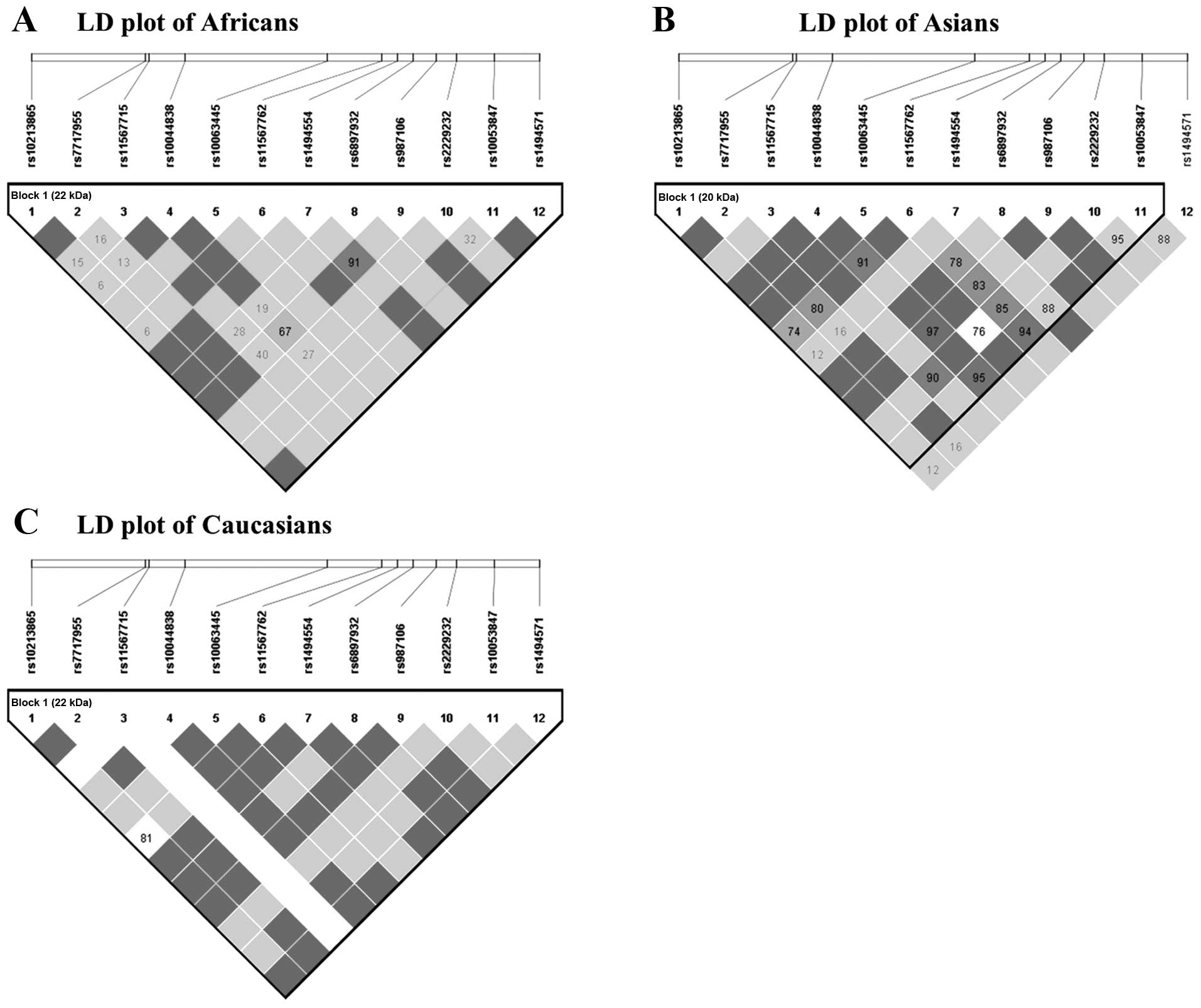
investigated the genetic makeup of three different ethnic groups:
Africans, Asians and Caucasians (Table
V and Fig. 2). As expected,
there were notable differences between the three populations.
 | Table IIILogistic analyses of IL7R
polymorphisms with the risk of inflammatory demyelinating diseases
in a Korean population (n=415). |
Table III
Logistic analyses of IL7R
polymorphisms with the risk of inflammatory demyelinating diseases
in a Korean population (n=415).
| SNP and
haplotypes | Allele change | IDD vs. NC | MS vs. NC | NMO vs. NC |
|---|
|
|
|
|---|
| MAF | Co-dominant | Dominant | Recessive | MAF | Co-dominant | Dominant | Recessive | MAF | Co-dominant | Dominant | Recessive |
|---|
|
|
|
|
|
|
|
|
|
|
|
|
|---|
| Case | Control | P |
Pcor. | P |
Pcor. | P |
Pcor. | Case | Control | P |
Pcor. | P |
Pcor. | P |
Pcor. | Case | Control | P |
Pcor. | P |
Pcor. | P |
Pcor. |
|---|
|
rs10213865 | A>C | 0.220 | 0.225 | 0.90 | NS | 0.57 | NS | 0.34 | NS | 0.191 | 0.225 | 0.29 | NS | 0.35 | NS | 0.43 | NS | 0.242 | 0.225 | 0.41 | NS | 0.19 | NS | 0.46 | NS |
|
rs7717955 | C>T | 0.183 | 0.195 | 0.75 | NS | 0.82 | NS | 0.12 | NS | 0.158 | 0.195 | 0.15 | NS | 0.18 | NS | 0.36 | NS | 0.202 | 0.195 | 0.67 | NS | 0.30 | NS | 0.13 | NS |
|
rs11567715 | C>A | 0.079 | 0.065 | 0.78 | NS | 0.96 | NS | 0.25 | NS | 0.089 | 0.065 | 1.00 | NS | 0.99 | NS | NA | NA | 0.071 | 0.065 | 0.76 | NS | 0.98 | NS | 0.20 | NS |
|
rs10044838 | C>T | 0.381 | 0.407 | 0.51 | NS | 0.96 | NS | 0.23 | NS | 0.380 | 0.407 | 0.68 | NS | 0.75 | NS | 0.70 | NS | 0.383 | 0.407 | 0.71 | NS | 0.85 | NS | 0.34 | NS |
|
rs10063445 | A>C | 0.466 | 0.470 | 0.70 | NS | 0.97 | NS | 0.48 | NS | 0.475 | 0.470 | 0.78 | NS | 0.99 | NS | 0.64 | NS | 0.460 | 0.470 | 0.88 | NS | 1.00 | NS | 0.80 | NS |
|
rs2228141 | C>T | 0.138 | 0.157 | 0.41 | NS | 0.25 | NS | 0.60 | NS | 0.171 | 0.157 | 0.57 | NS | 0.79 | NS | 0.27 | NS | 0.112 | 0.157 | 0.17 | NS | 0.11 | NS | 0.91 | NS |
|
rs11567762 | A>G | 0.466 | 0.463 | 0.83 | NS | 0.83 | NS | 0.55 | NS | 0.475 | 0.463 | 0.85 | NS | 0.93 | NS | 0.66 | NS | 0.459 | 0.463 | 1.00 | NS | 0.91 | NS | 0.90 | NS |
|
rs1494554 | A>C | 0.028 | 0.036 | 0.68 | NS | 0.63 | NS | NA | NA | 0.013 | 0.036 | 0.77 | NS | 0.77 | NS | NA | NA | 0.041 | 0.036 | 0.92 | NS | 0.99 | NS | NA | NA |
|
rs6897932 | C>T | 0.171 | 0.186 | 0.50 | NS | 0.84 | NS | 0.003 | 0.03 | 0.152 | 0.186 | 0.10 | NS | 0.20 | NS | 0.06 | NS | 0.187 | 0.186 | 0.93 | NS | 0.36 | NS | NA | NA |
|
rs987106 | T>A | 0.242 | 0.249 | 0.97 | NS | 0.71 | NS | 0.41 | NS | 0.209 | 0.249 | 0.36 | NS | 0.44 | NS | 0.46 | NS | 0.268 | 0.249 | 0.50 | NS | 0.30 | NS | 0.74 | NS |
|
rs2229232 | C>T | 0.059 | 0.040 | 0.27 | NS | 0.25 | NS | 0.99 | NS | 0.071 | 0.040 | 0.31 | NS | 0.30 | NS | NA | NA | 0.051 | 0.040 | 0.40 | NS | 0.38 | NS | 0.88 | NS |
|
rs10053847 | G>A | 0.138 | 0.159 | 0.36 | NS | 0.22 | NS | 0.60 | NS | 0.171 | 0.159 | 0.60 | NS | 0.82 | NS | 0.27 | NS | 0.112 | 0.159 | 0.15 | NS | 0.09 | NS | 0.91 | NS |
|
rs1494571 | C>G | 0.028 | 0.036 | 0.68 | NS | 0.64 | NS | NA | NA | 0.013 | 0.036 | 0.77 | NS | 0.77 | NS | NA | NA | 0.040 | 0.036 | 0.91 | NS | 0.99 | NS | NA | NA |
|
IL7R_ht1 | NA | 0.466 | 0.470 | 0.70 | NS | 0.97 | NS | 0.48 | NS | 0.475 | 0.470 | 0.78 | NS | 0.99 | NS | 0.64 | NS | 0.46 | 0.470 | 0.88 | NS | 1.00 | NS | 0.80 | NS |
|
IL7R_ht2 | NA | 0.169 | 0.181 | 0.59 | NS | 0.76 | NS | 0.005 | 0.04 | 0.146 | 0.181 | 0.12 | NS | 0.24 | NS | 0.07 | NS | 0.187 | 0.181 | 0.84 | NS | 0.32 | NS | NA | NA |
|
IL7R_ht3 | NA | 0.143 | 0.158 | 0.38 | NS | 0.22 | NS | 0.58 | NS | 0.171 | 0.158 | 0.59 | NS | 0.82 | NS | 0.27 | NS | 0.121 | 0.158 | 0.17 | NS | 0.10 | NS | 0.86 | NS |
|
IL7R_ht4 | NA | 0.059 | 0.040 | 0.33 | NS | 0.30 | NS | 0.99 | NS | 0.070 | 0.040 | 0.37 | NS | 0.36 | NS | NA | NA | 0.051 | 0.040 | 0.40 | NS | 0.38 | NS | 0.88 | NS |
|
IL7R_ht5 | NA | 0.042 | 0.027 | 0.11 | NS | 0.11 | NS | NA | NA | 0.044 | 0.027 | 0.11 | NS | 0.11 | NS | NA | NA | 0.040 | 0.027 | 0.21 | NS | 0.21 | NS | NA | NA |
 | Table IVComparison and meta-analysis of the
genetic effect of IL7R SNP rs6897932 on MS in
previous studies. |
Table IV
Comparison and meta-analysis of the
genetic effect of IL7R SNP rs6897932 on MS in
previous studies.
| Study
population | Study size (case
vs. control) | MAF | P-value (OR) | Author (year)
(ref.) |
|---|
|
|---|
| Case | Control |
|---|
| Dutch
Caucasian | 697 vs. 174 | ND | ND | 0.0004
(0.61) | Sombekke et
al (2011) (24) |
| Australian and New
Zealandera | 3,874 vs.
5,723 | 0.240 | 0.264 | 0.0013
(0.91) | Australia and New
Zealand Multiple Sclerosis Genetics Consortium (2009) (16) |
| Nordic | 1,210 vs.
1,234 | 0.256 | 0.302 | 0.02
(0.76) | Lundmark et
al (2007) (26) |
| USA | 438 vs. 479 | 0.217 | 0.265 | 0.05
(0.75) | Gregory et
al (2007) (28) |
| European | 1,077 vs.
2,725 | 0.238 | 0.283 | 0.0006
(0.81) | |
| German | 206 vs. 605 | 0.748 | 0.746 | 0.17 (0.88) | Weber et al
(2008) (21) |
| Australian | 1,134 vs.
1,265 | 0.252 | 0.254 | 0.58 (0.96) | Rubio et al
(2008) (22) |
| Canadianc | 1,193 vs.
1,553 | 0.290b | | 0.0002
(0.78) | Ramagopalan et
al (2007) (29) |
| USAa | 207 vs. 413 | 0.227 | 0.303 | 0.0005 | O’Doherty et
al (2008) (25) |
| UKa | 463 vs. 530 | 0.255 | 0.264 | 0.638 | |
| UK and US | 2,322 vs.
2,987 | 0.250b | | 0.00003
(0.85) | Hafler et al
(2007) (6) |
| Germana | 1,267 vs. 868 | 0.249 | 0.275 | 0.054 | Akkad et al
(2009) (23) |
| Finnish | 922 vs. 1,392 | 0.310 | 0.350 | 0.0002
(0.81) | Kallio et al
(2009) (27) |
| Spanish | 599 vs. 594 | 0.225 | 0.274 | 0.003
(0.75) | Alcina et al
(2008) (15) |
| Japanese | 187 vs. 158 | 0.107 | 0.203 | 0.002
(0.47) | Fang et al
(2011) (17) |
| Korean | 82 vs. 298 | 0.146 | 0.187 | 0.06 (0.54) | Present study |
| Meta-analysis |
| Caucasian | 11,056 vs.
13,876 | - | - |
4.60×10−18
(0.82) | - |
| Asian | 269 vs. 456 | - | - | 0.0001
(0.49) | - |
| All | 11,325 vs.
14,332 | - | - |
1.15×10−19
(0.81) | - |
 | Table VMinor allele frequencies and
Chi-square distribution of rs6897932 in different ethnic
groups. |
Table V
Minor allele frequencies and
Chi-square distribution of rs6897932 in different ethnic
groups.
| Race | N | MAF | P-value |
|---|
| African | 510 | 0.073 | - |
| Asian | 733 | 0.182 | - |
| Caucasian | 253 | 0.241 | - |
| AF vs. AS | - | - |
<0.0001 |
| AF vs. CA | - | - |
<0.0001 |
| AS vs. CA | - | - | 0.02 |
Discussion
In the present study, a significant association was
found between rs6897932 and IDD (P=0.0003; OR [95%
confidence interval (CI)]=0.10 [0.01–0.75]). However, this
association may have come from MS and not from NMO, as case MAFs
were lower than control MAFs in both IDD and MS (Table III), while NMO case and control
MAFs were almost the same. The significant association detected for
ht2 is most likely due to rs68797932, as the
haplotype is almost tagged by rs68797932. In the subgroup
analyses for MS and NMO, none of the genetic variants showed a
significant association, including rs6897932 and
ht2.
Numerous studies on the association between MS and
risk genes have led to the identification of how gene variants may
increase the risk of MS. A group of investigators found a
significant association between an IL7R genetic variant and
MS in a Caucasian population, and explained its possible mechanism
via sequence analysis (28). Their
analysis showed that rs6897932 affected the function of the
receptor by inducing the transcripts to skip exon 6 while encoding.
The investigators suspected that the SNP either weakened an exonic
splicing enhancer or strengthened an exonic splicing silencer,
stating that the latter was more likely. The exclusion of exon 6
changed the number of soluble and membrane-bound isoforms of IL7R,
which in turn led to the increased susceptibility for MS.
As shown in Table
IV, significant associations between the risk allele of
rs6897932 and MS were identified in various studies.
Notably, the magnitude of risk (OR) was higher in Asian populations
(Japanese and Korean) than that in Caucasian samples. Thus,
rs6897932 is a potentially stronger risk factor in Asian
populations than in Caucasian populations. This hypothesis was
strengthened by the results of the present study (OR=0.82 in
Caucasian and 0.49 in Asian; Table
IV). Table V and Fig. 2 show that the frequencies and LD
structures of Africans, Asians and Caucasians are different from
each other, which partly explains the different influence of
rs6897932 on MS in Asians compared with that in
Caucasians.
One limitation of our study was the small number of
study samples. However, we only recruited patients whose diagnoses
were clear in order to avoid the possibility of ascertainment
error. In the present study, patients who were enrolled in the NMO
group were seropositive for the AQP4 antibody, as determined by
highly specific assays, and their clinical features were otherwise
typical for NMO. Furthermore, for patients enrolled in the MS
group, a diagnosis was made by expert MS specialists. Therefore,
the possibility of ascertainment error in our study was reduced as
much as possible.
In conclusion, in the present study, we conducted
association studies of 13 IL7R SNPs for MS, NMO and IDD. We
found a significant association between rs6897932 and IDD,
which was likely due to the putative association between the SNP
and MS. Furthermore, the results of the present study were similar
to those from a previous study which identified rs6897932 as
a stronger risk factor in Asian populations than in Caucasian
populations. This finding was also consistent with the results
obtained from the meta-analysis. Therefore, the results from the
present study support the hypothesis that rs6897932 is a
stronger risk factor for IDDs in Asians as compared to
Caucasians.
Acknowledgements
This study was supported by the Korea Science and
Engineering Foundation (KOSEF) funded by the Korea government
(MEST) (no. 2011-0004453), Sogang University Research Grant of 2011
(SRF-201114006.01) and a grant from the Korea Healthcare Technology
R&D Project, Ministry of Health and Welfare, Republic of Korea
(no. A101023). The biospecimens for this study were provided by the
National Biobank of Korea (KOBB-2012-19).
References
|
1
|
Compston A and Coles A: Multiple
sclerosis. Lancet. 372:1502–1517. 2008. View Article : Google Scholar
|
|
2
|
Kenealy SJ, Pericak-Vance MA and Haines
JL: The genetic epidemiology of multiple sclerosis. J Neuroimmunol.
143:7–12. 2003. View Article : Google Scholar
|
|
3
|
Sawcer S, Hellenthal G, Pirinen M, et al:
Genetic risk and a primary role for cell-mediated immune mechanisms
in multiple sclerosis. Nature. 476:214–219. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
De Jager PL, Jia X, Wang J, et al:
Meta-analysis of genome scans and replication identify CD6, IRF8
and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat
Genet. 41:776–782. 2009.PubMed/NCBI
|
|
5
|
Baranzini SE, Wang J, Gibson RA, et al:
Genome-wide association analysis of susceptibility and clinical
phenotype in multiple sclerosis. Hum Mol Genet. 18:767–778. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hafler DA, Compston A, Sawcer S, et al:
Risk alleles for multiple sclerosis identified by a genomewide
study. N Engl J Med. 357:851–862. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wingerchuk DM, Hogancamp WF, O’Brien PC
and Weinshenker BG: The clinical course of neuromyelitis optica
(Devic’s syndrome). Neurology. 53:1107–1114. 1999.
|
|
8
|
Wingerchuk DM, Lennon VA, Pittock SJ,
Lucchinetti CF and Weinshenker BG: Revised diagnostic criteria for
neuromyelitis optica. Neurology. 66:1485–1489. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Fukazawa T, Yamasaki K, Ito H, et al: Both
the HLA-CPB1 and -DRB1 alleles correlate with risk for multiple
sclerosis in Japanese: clinical phenotypes and gender as important
factors. Tissue Antigens. 55:199–205. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lau KK, Wong LK, Li LS, Chan YW, Li HL and
Wong V: Epidemiological study of multiple sclerosis in Hong Kong
Chinese: questionnaire survey. Hong Kong Med J. 8:77–80.
2002.PubMed/NCBI
|
|
11
|
Das A and Puvanendran K: A retrospective
review of patients with clinically definite multiple sclerosis. Ann
Acad Med Singapore. 27:204–209. 1998.PubMed/NCBI
|
|
12
|
Kim HJ, Park HY, Kim E, et al: Common
CYP7A1 promoter polymorphism associated with risk of neuromyelitis
optica. Neurobiol Dis. 37:349–355. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kroemer RT and Richards WG: Homology
modeling study of the human interleukin-7 receptor complex. Protein
Eng. 9:1135–1142. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Xue HH, Bollenbacher J, Rovella V, et al:
GA binding protein regulates interleukin 7 receptor alpha-chain
gene expression in T cells. Nat Immunol. 5:1036–1044. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Alcina A, Fedetz M, Ndagire D, et al: The
T244I variant of the interleukin-7 receptor-alpha gene and multiple
sclerosis. Tissue Antigens. 72:158–161. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Australia and New Zealand Multiple
Sclerosis Genetics Consortium (ANZgene). Genome-wide association
study identifies new multiple sclerosis susceptibility loci on
chromosomes 12 and 20. Nat Genet. 41:824–828. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Fang L, Isobe N, Yoshimura S, et al:
Interleukin-7 receptor alpha gene polymorphism influences multiple
sclerosis risk in Asians. Neurology. 76:2125–2127. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Kurtzke JF: Rating neurologic impairment
in multiple sclerosis: an expanded disability status scale (EDSS).
Neurology. 33:1444–1452. 1983. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Barrett JC, Fry B, Maller J and Daly MJ:
Haploview: analysis and visualization of LD and haplotype maps.
Bioinformatics. 21:263–265. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Stephens M, Smith NJ and Donnelly P: A new
statistical method for haplotype reconstruction from population
data. Am J Hum Genet. 68:978–989. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Weber F, Fontaine B, Cournu-Rebeix I, et
al: IL2RA and IL7RA genes confer susceptibility for multiple
sclerosis in two independent European populations. Genes Immun.
9:259–263. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Rubio JP, Stankovich J, Field J, et al:
Replication of KIAA0350, IL2RA, RPL5 and CD58 as multiple sclerosis
susceptibility genes in Australians. Genes Immun. 9:624–630. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Akkad DA, Hoffjan S, Petrasch-Parwez E,
Beygo J, Gold R and Epplen JT: Variation in the IL7RA and IL2RA
genes in German multiple sclerosis patients. J Autoimmun.
32:110–115. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Sombekke MH, van der Voort LF, Kragt JJ,
et al: Relevance of IL7R genotype and mRNA expression in Dutch
patients with multiple sclerosis. Mult Scler. 17:922–930. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
O’Doherty C, Kantarci O and Vandenbroeck
K: IL7RA polymorphisms and susceptibility to multiple sclerosis. N
Engl J Med. 358:753–754. 2008.PubMed/NCBI
|
|
26
|
Lundmark F, Duvefelt K, Iacobaeus E, et
al: Variation in interleukin 7 receptor alpha chain (IL7R)
influences risk of multiple sclerosis. Nat Genet. 39:1108–1113.
2007. View
Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kallio SP, Jakkula E, Purcell S, et al:
Use of a genetic isolate to identify rare disease variants: C7 on
5p associated with MS. Hum Mol Genet. 18:1670–1683. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gregory SG, Schmidt S, Seth P, et al:
Interleukin 7 receptor alpha chain (IL7R) shows allelic and
functional association with multiple sclerosis. Nat Genet.
39:1083–1091. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ramagopalan SV, Anderson C, Sadovnick AD
and Ebers GC: Genomewide study of multiple sclerosis. N Engl J Med.
357:21992–200. 2007.
|