Introduction
Best vitelliform macular dystrophy (BVMD) is one of
the most common forms of autosomal dominant macular dystrophy,
characterized by the presence of yellowish lesions in the macular
area (1–4).
BVMD has been classified into five phenotypic
stages: The previtelliform, vitelliform, pseudohypopyon,
vitelliruptive and atrophic stages. Not all patients will progress
through all five stages, and the stages do not always occur
consecutively. Furthermore, BVMD lesions may simultaneously display
characteristics of various BVMD stages (5–8).
BVMD is associated with mutations in the
bestrophin 1 gene (BEST1), on chromosome 11q12, which
encodes a 585 amino acid transmembrane protein and is selectively
expressed in the retinal pigment epithelium (RPE) (9–11).
The BEST1 gene is the founding member of a family of four
paralogs, which also includes BEST2, BEST3 and
BEST4. BEST1 is expressed in the RPE cells of the
developing and adult eye cells (1,9).
Greater than 300 distinct BEST1 mutations
have been identified in families or sporadic patients affected by
BVMD (12–17). It has been hypothesized that
mutations in BEST1 may result in channel dysfunction which
leads to abnormal fluid and ion transport through the RPE,
resulting in defects during ocular growth and later-onset retinal
dystrophy (18,19).
Mutations in the BEST1 gene are detected in
the majority of cases of BVMD with a positive family history. The
current study aimed to conduct mutational analysis of two Chinese
families with BVMD at the gene level, in addition to analyzing the
associated clinical features.
Patients and methods
Ethical approval
Approval for the current study was provided by the
ethics committee of Zhongshan Ophthalmic Center, Sun Yat-Sen
University (Guangzhou, China). Two patients were diagnosed with
BVMD at the Zhongshan Ophthalmic Center.
Ophthalmic examinations
A variety of techniques were used for the ophthalmic
examinations, which are outlined as follows: Visual acuity was
examined using the Early Treatment Diabetic Retinopathy Study chart
(Precision Vision, LaSalle, IL, USA). Anterior segment photographs
were captured using a BX 900 Slit Lamp (Haag-Streit AG, Köniz,
Switzerland). Anterior segment measurements were taken with
Pentacam® HR version 70700 (OCULUS Optikgeräte GmbH,
Wetzlar, Germany). Fundus photography and fundus fluorescein
angiography (FFA) imaging was performed using a Heidelberg Retina
Angiograph (Heidelberg Engineering GmbG, Heidelberg, Germany). In
addition, physical examinations, including blood examination, a
urine test, electrocardiogram, chest X-ray, blood biochemistry
test, blood lipid and blood coagulation tests were conducted to
exclude systemic diseases.
Sample collection
Two affected families were identified as described.
A total of 100 subjects who exhibited no diagnostic features of
BVMD were recruited from the same population to serve as normal
controls. Written informed consent was obtained from all
participating individuals prior to the commencement of the current
study. According to the principles of the Declaration of Helsinki,
venous blood samples were collected from peripheral blood
leukocytes for genomic DNA extraction using a DNA extraction kit
(Qiagen, Hilden, Germany).
Mutation detection
Exons of the BEST1 gene were amplified with
the polymerase chain reaction (PCR), using the primers (Beijing
Genomics Institute, Guangzhou, China) presented in Table I (20). In brief, PCR was performed in a
50-µl total volume reaction. All reagents used for PCR were
purchased from (Takara Bio, Inc., Tokyo, Japan). The PCR cycling
profile was as follows: One cycle at 94°C for 5 min, followed by 40
cycles at 94°C for 45 sec, 52–66°C for 45 sec and 72°C for 45 sec,
in addition to one cycle at 72°C for 10 min. The PCR products were
sequenced from both directions using an ABI3730 Automated Sequencer
(Applied Biosystems Life Technologies, Foster City, CA, USA). The
sequencing results were analyzed using SeqMan, version 2.3
(Technelysium Pty Ltd., Brisbane, Australia) and they were compared
with the reference sequences in the database at the National Center
for Biotechnology Information (NC_000011.9; http://www.ncbi.nlm.nih.gov/nuccore/224589802).
 | Table IPrimers used for the polymerase chain
reaction. |
Table I
Primers used for the polymerase chain
reaction.
| Exon | Forward (5′-3′) | Reverse (5′-3′) | Product size
(bp) | Annealing temperature
(°C) |
|---|
| 2 |
AGTCTCAGCCATCTCCTCGC |
TGGCCTGTCTGGAGCCTG | 212 | 61 |
| 3 | GGGACAGTCTCAGCC
ATCTC |
CAGCTCCTCGTGATCCTCC | 238 | 58 |
| 4 |
AGAAAGCTGGAGGAGCCG |
GCGGCAGCCCTGTCTGTAC | 1408 | 59 |
| 5 |
GGGGCAGGTGGTGTTCAGA |
GGCAGCCTCACCAGCCTAG | 150 | 59 |
| 6 |
GGGCAGGTGGTGTTCAGA |
CCTTGGTCCTTCTAGCCTCAG | 181 | 59 |
| 7 |
CATCCTGATTTCAGGGTTCC |
CTCTGGCCATGCCTCCAG | 257 | 59 |
| 8 |
AGCTGAGGTTTAAAGGGGGA |
TCTCTTTGGGTCCACTTTGG | 215 | 59 |
| 9 |
ACATACAAGGTCCTGCCTGG |
GCATTAACTAGTGCTATTCTAAGTTCC | 298 | 59 |
| 10A |
GGTGTTGGTCCTTTGTCCAC |
CTCTGGCATATCCGTCAGGT | 591 | 59 |
| 10B |
CTTCAAGTCTGCCCCACTGT |
TAGGCTCAGAGCAAGGGAAG | 457 | 59 |
| 11 |
CATTTTGGTATTTGAAATGAAGG |
CCATTTGATTCAGGCTGTTG | 216 | 59 |
Results
Clinical data
The patients evaluated in the current study were
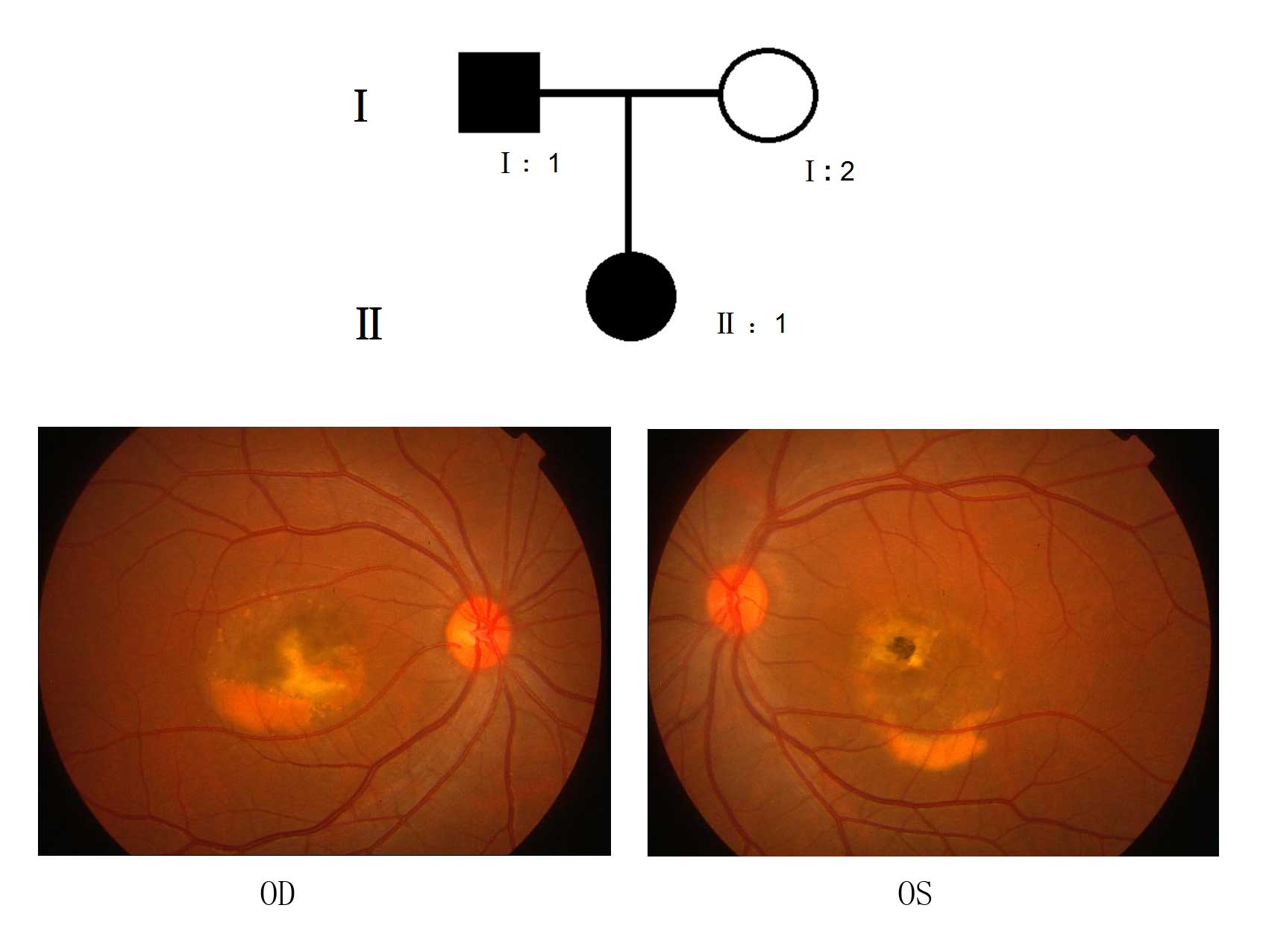
from the southern area of China. Case 1 (II:1) of Family 1 was a
23-year old female with a family history (Fig. 1) of ocular disease, diagnosed with
BVMD at the age of 21 years. Her best-corrected visual acuity, as
measured by the Logarithm of the Minimum Angle of Resolution
(LogMAR) using the following formula: LogMAR visual acuity = 0.1 +
LogMAR value of the best line read −0.02 X (number of letters
read), was 0.1 in the right eye and 0.4 in the left eye.
Examination of the fundus identified vitelliruptive lesions with a
scrambled egg-like appearance and dispersion of the vitelliform
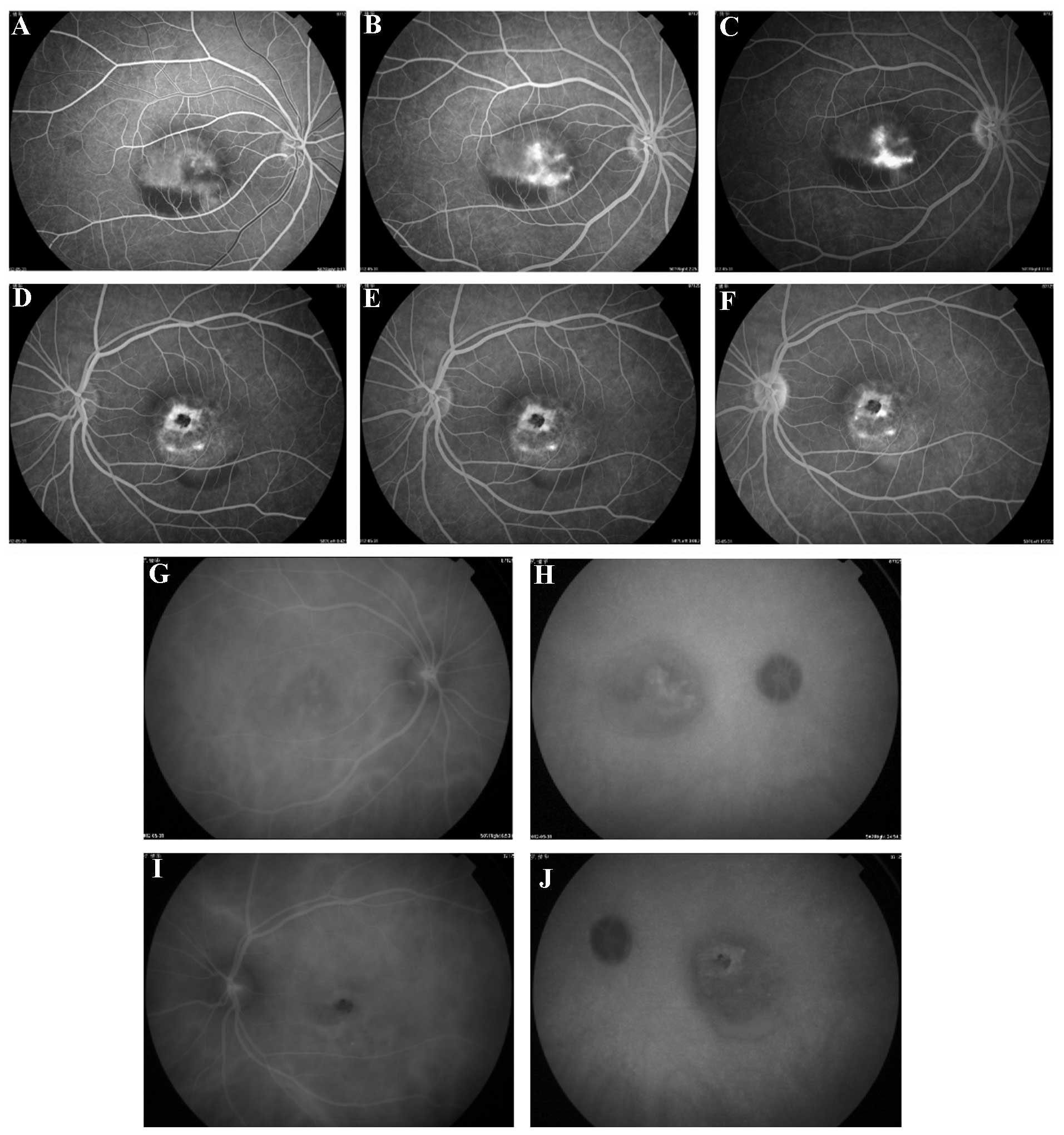
material, including signs of atrophy, in the left eye (Fig. 1; OS). Fluorescein angiography
(Fig. 2) demonstrated significant
early hyperfluorescence (Fig. 2A)
that increased in intensity at the late stage of the angiographic
sequence, and was associated with moderate leakage (Fig. 2B and C) in the right eye, and
macular lesions that simulated a pattern of dystrophy in the left
eye (Fig. 2D–F). Indocyanine green
chorioangiography (ICG; Fig. 2G–J)
detected hypofluorescence in the early stages (Fig. 2G and I) of the angiographic
sequence in the left and right eyes and subsequently detected
little abnormal hyperfluorescence the late stages (Fig. 2H and J). The father of Case 1 was
also diagnosed with ocular disease during his teenage years, and at
the time of the present study, exhibited marked macular
degeneration and cataracts.
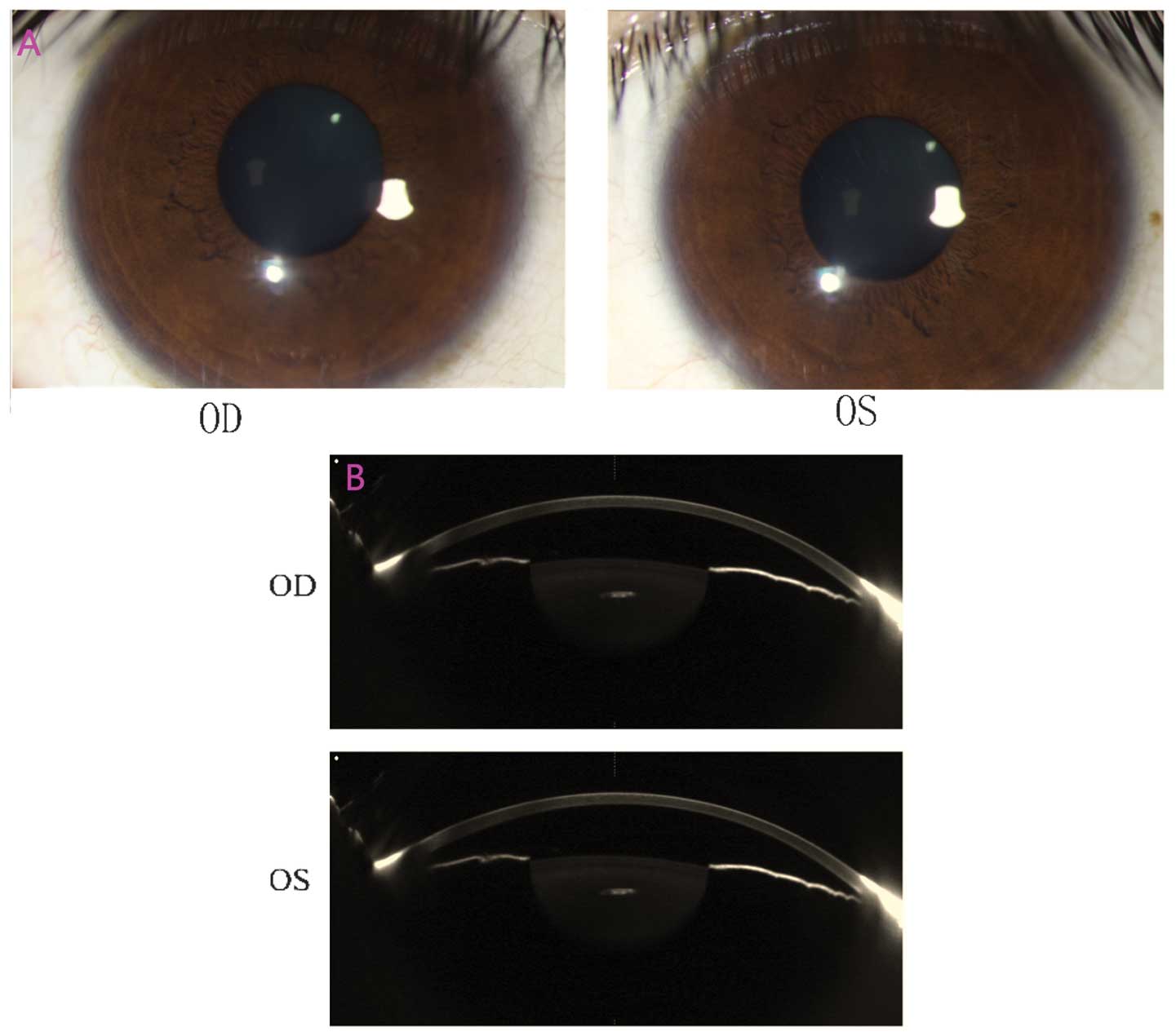
The anterior segment photograph captured using a BX
900 Slit Lamp is presented in Fig.
3A, and the images presented in Fig. 3B were captured using Pentacam. The
anterior chamber depths were 2.09 mm [oculus dexter (OD)] and 2.12
mm [oculus sinister (OS)], and the central corneal thickness was
identified to be 548 µm (OD) and 558 µm (OS).
Case 2 of Family 2 was a 22 year-old male with no
known familial history of ocular disease at the time of diagnosis
of BVMD aged 19 years. His best-corrected visual acuity, as
measured by LogMAR, was 0.7 in the right eye and 0.4 in the left
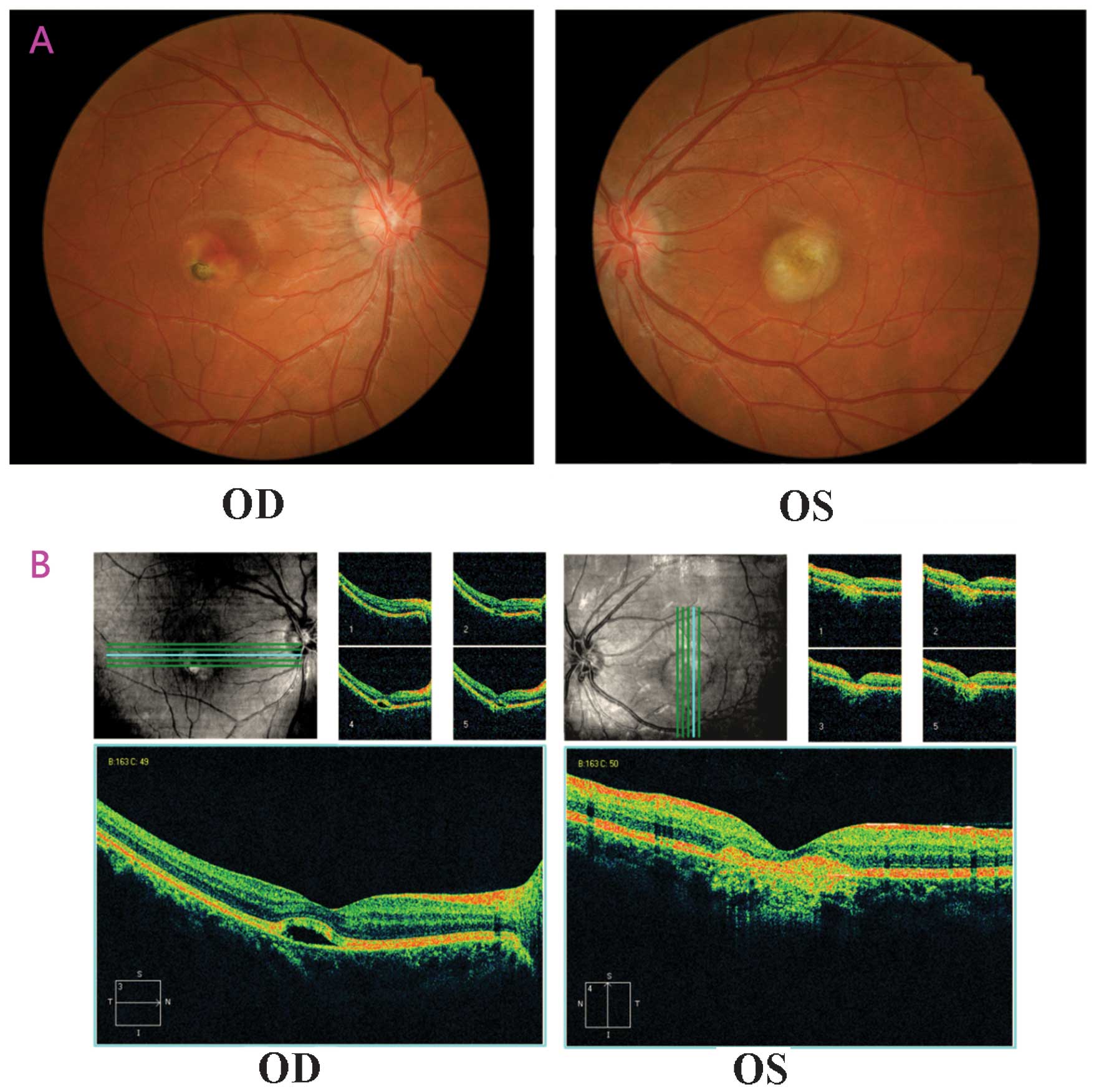
eye. Fundus examination (Fig. 4A)
indicated vitelliruptive lesions with a scrambled egg-like
appearance and dispersion of the vitelliform material with signs of
atrophy of the right eye. This patient was identified to be
supersensitive to fluorescein, therefore no FFA or ICGA results
were obtained.
The anterior chamber depths were 3.15 mm (OD) and
3.17 mm (OS), and the central corneal thickness was identified to
be 564 µm (OD) and 568 µm (OS). Spectral domain
optical coherence tomography (OCT) scans revealed clear
abnormalities, including the absence of the foveal pit, serous
retinal detachment, cystoid macular edema and interruption of the
outer limiting membrane (Fig. 4B).
The right eye was observed to exhibit prominent yellow-white
subretinal scarring with pigmented borders, surrounded by a serous
retinal detachment. Additionally, the left eye presented with an
atrophic lesion.
OCT revealed that the foveal region of the right eye
was unusually thick due to the presence of abnormal neuroretinal
detachment from the RPE, which was suggested to have been triggered
by an aberrant accumulation of fluid within the choriocapillaris
and between the RPE and the fovea. In addition, OCT indicated that
the foveal region of the left eye was abnormally thick due to
irregularities in the deep retinal layers, with aberrant junctions
between the inner and outer segents, as well as a prominent and
hyperreflective structure in contact with the overlying retina,
which was surrounded by elevated retina with underlying spots of
increased reflectivity.
Mutation screening
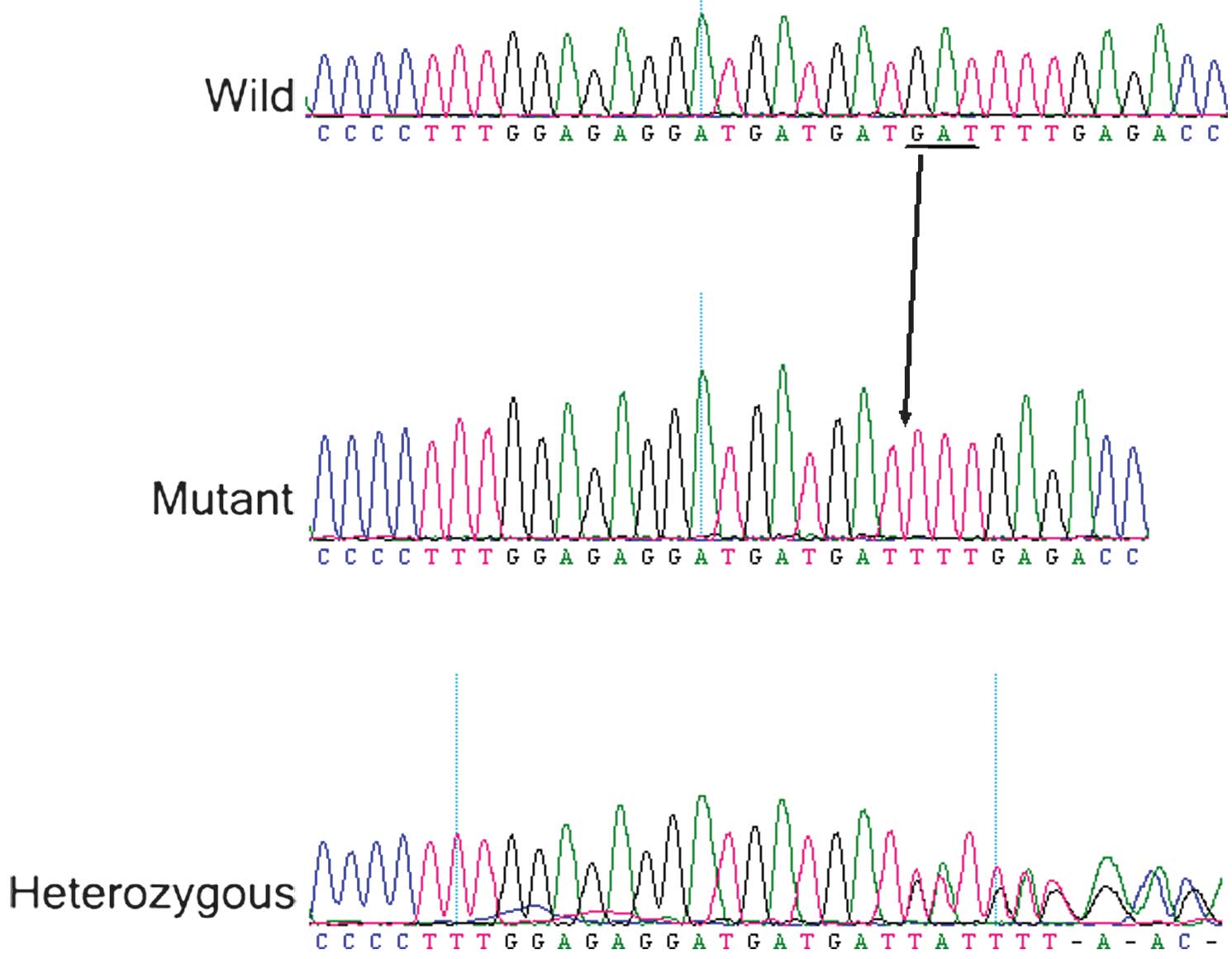
A heterozygous mutation c.910_912delGAT (p.304del
Asp) was identified in exon 7 in the affected family members of
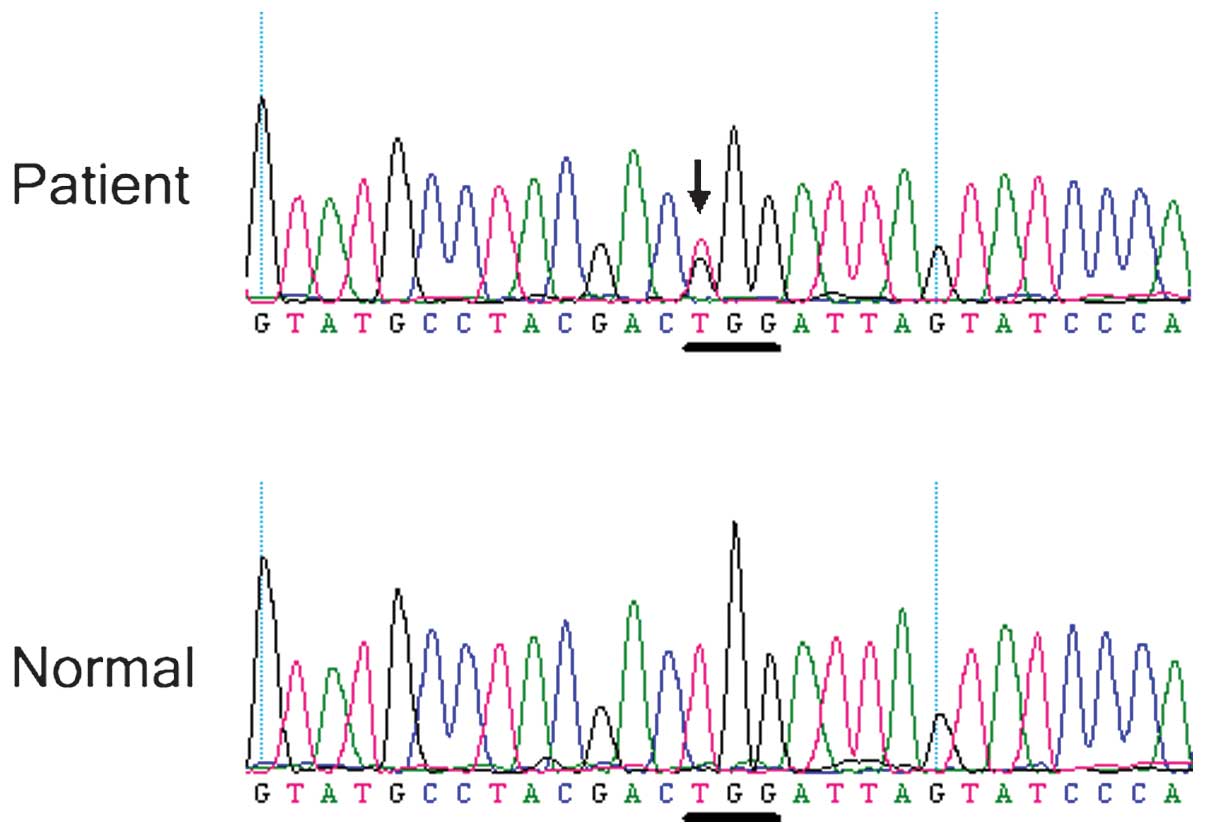
Family 1 (Fig. 5). A heterozygous
BEST1 missense mutation c.685T>G (p.Trp229Gly) in exon 5
was identified in the affected Case 2 (Fig. 6), but not the unaffected members of
Family 2 or the normal controls. These identified mutations had not
previously been reported.
Discussion
BVMD is associated with mutations in the
BEST1 gene, which was formerly known as the VMD2
gene. The BEST1 gene encodes bestrophin-1, a transmembrane
protein located in the basolateral membrane of the RPE (21–23).
A large number of allelic variants of the
BEST1 gene have previously been identified and are recorded
in the Regensburg University (Bavaria, Germany) database (24).
In the current study, two mutations were identified
in the BEST-1 gene, which are associated with Best Syndrome:
c.910_912delGAT (p.304del Asp) and c.685T>G (p.Trp229Gly). These
mutations, rather than a rare polymorphism in the normal
population, are suggested to be the causative mutations in the two
sporadic BVMD patients.
Although >300 distinct BEST1 allelic
variants have been identified in BVMD, a comprehensive database
summarizing the known associations between BEST1 mutations
and the prognosis of patients with BVMD worldwide remains to be
developed. This is due to the fact that systematic whole
BEST1 gene sequencing and clinical examinations have not
been completed in a sufficiently large cohort of affected
patients.
BVMD is a bilateral, symmetric, progressive disease
of the macular area, associated with long-term loss of vision.
Onset of the disease is predominantly early in life, often
occurring by 10 years of age (7,25,26).
Amongst certain patients, BVMD rapidly progresses towards the loss
of central vision (22,26); thus, a thorough investigation of
the age of onset of this disease may aid the development of
valuable diagnostic/prognostic techniques. Conducting studies on
independent cohorts of children belonging to families with a known
history of BVMD may also be beneficial.
In conclusion, the current study identified two
novel mutations of the BEST1 gene in two Chinese families
with BVMD. The results of the current study expanded the library of
known mutations of BEST1 and are valuable for the
development of genetic counseling and prenatal diagnosis in
families with BVMD.
Acknowledgments
The authors would like to thank all patients,
families and normal control volunteers for their participation in
the present study. The current study was supported by the National
Natural Science Foundation of China (grant no. 30973277); the
Science and Technology Planning Project of Guangdong Province,
China (grant nos. 2010B090400416 and 2011B080701033); the Medical
Scientific Research Foundation of Guangdong Province (grant no.
B2012126); the Key Clinical Program of the Ministry of Health
(grant no. 2010.439); the Youth Project of Fundamental Research
Funds of the State Key Laboratory of Ophthalmology (grant no.
2011Q09) and the Project of Fundamental Research Funds of the State
Key Laboratory of Ophthalmology (grant no. 2012KF03).
References
|
1
|
White K, Marquardt A and Weber BH: VMD2
mutations in vitelliform macular dystrophy (Best disease) and other
maculopathies. Hum Mutat. 15:301–308. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Sun H, Tsunenari T, Yau KW and Nathans J:
The vitelliform macular dystrophy protein defines a new family of
chloride channels. Proc Natl Acad Sci USA. 99:4008–4013. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Sodi A, Passerini I, Murro V, Caputo R,
Bacci GM, et al: BEST1 sequence variants in Italian patients with
vitelliform macular dystrophy. Mol Vis. 18:2736–2748.
2012.PubMed/NCBI
|
|
4
|
MacDonald IM and Lee T: Best vitelliform
macular dystrophy. GeneReviews® (Internet). Pagon RA,
Adam MP, Ardinger HH, Bird TD, Dolan CR, et al: University of
Washington; Seattle, WA: 2003
|
|
5
|
Hayami M, Decock C, Brabant P, Van
Kerckhoven W, Lafaut BA, et al: Optical coherence tomography of
adult-onset vitelliform dystrophy. Bull Soc Belge Ophtalmol.
289:53–61. 2003.PubMed/NCBI
|
|
6
|
Eksandh L, Bakall B, Bauer B, Wadelius C
and Andréasson S: Best’s vitelliform macular dystrophy caused by a
new mutation (Val89Ala) in the VMD2 gene. Ophthalmic Genet.
22:107–115. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Boon CJ, Theelen T, Hoefsloot EH, van
Schooneveld MJ, Keunen JE, et al: Clinical and molecular genetic
analysis of best vitelliform macular dystrophy. Retina. 29:835–847.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Furino C, Boscia F, Cardascia N, Sborgia L
and Sborgia C: Fundus autofluorescence, optical coherence
tomography and visual acuity in adult-onset foveomacular dystrophy.
Ophthalmologica. 222:240–244. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Caldwell GM, Kakuk LE, Griesinger IB,
Simpson SA, Nowak NJ, et al: Bestrophin gene mutations in patients
with best vitelliform macular dystrophy. Genomics. 58:98–101. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Krämer F, Mohr N, Kellner U, Rudolph G and
Weber BH: Ten novel mutations in VMD2 associated with best macular
dystrophy (BMD). Hum Mutat. 22:4182003. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Krämer F, White K, Pauleikhoff D, Gehrig
A, Passmore L, et al: Mutations in the VMD2 gene are associated
with juvenile-onset vitelliform macular dystrophy (Best disease)
and adult vitelliform macular dystrophy but not age-related macular
degeneration. Eur J Hum Genet. 8:286–292. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
MacDonald IM, Gudiseva HV, Villanueva A,
Greve M, Caruso R, et al: Phenotype and genotype of patients with
autosomal recessive bestrophinopathy. Ophthalmic Genet. 33:123–129.
2012. View Article : Google Scholar
|
|
13
|
Marchant D, Gogat K, Boutboul S, Pequignot
M, Sternberg C, et al: Identification of novel VMD2 gene mutations
in patients with best vitelliform macular dystrophy. Hum Mutat.
17:2352001. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Marchant D, Yu K, Bigot K, Roche O,
Germain A, et al: New VMD2 gene mutations identified in patients
affected by best vitelliform macular dystrophy. J Med Genet.
44:e702007. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Musarella MA: Molecular genetics of
macular degeneration. Doc Ophthalmol. 102:165–177. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sodi A, Passerini I, Simonelli F, Testa F,
Menchini U, et al: A novel mutation in the VMD2 gene in an Italian
family with best maculopathy. J Fr Ophtalmol. 30:616–620. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhao L, Grob S, Corey R, Krupa M, Luo J,
et al: A novel compound heterozygous mutation in the BEST1 gene
causes autosomal recessive best vitelliform macular dystrophy. Eye
(Lond). 26:866–871. 2012. View Article : Google Scholar
|
|
18
|
Shamshinova AM: Local ERG for clinical
examination of eye diseases. Doc Ophthalmol. 76:1–11. 1990.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Querques G, Zerbib J, Santacroce R,
Margaglione M, Delphin N, et al: Functional and clinical data of
best vitelliform macular dystrophy patients with mutations in the
BEST1 gene. Mol Vis. 15:2960–2972. 2009.
|
|
20
|
Lin Y, Liang X, Ai S, Chen C, Liu X, Luo
L, et al: FGFR2 molecular analysis and related clinical findings in
one Chinese family with Crouzon syndrome. Mol Vis. 18:449–454.
2012.PubMed/NCBI
|
|
21
|
Allikmets R, Seddon JM, Bernstein PS,
Hutchinson A, Atkinson A, et al: Evaluation of the best disease
gene in patients with age-related macular degeneration and other
maculopathies. Hum Genet. 104:449–453. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Brecher R and Bird AC: Adult vitelliform
macular dystrophy. Eye (Lond). 4:210–215. 1990. View Article : Google Scholar
|
|
23
|
Preising MN, Pasquay C, Friedburg C, Bowl
W, Jager M, et al: Autosomal recessive bestrophinopathy (ARB): a
clinical and molecular description of two patients at childhood.
Klin Monbl Augenheilkd. 229:1009–1017. 2012.In German. PubMed/NCBI
|
|
24
|
Lacassagne E, Dhuez A, Rigaudière F,
Dansault A, Vêtu C, Bigot K, Vieira V, Puech B, Defoort-Dhellemmes
S and Abitbol M: Phenotypic variability in a French family with a
novel mutation in the BEST1 gene causing multifocal best
vitelliform macular dystrophy. Mol Vis. 17:309–322. 2011.PubMed/NCBI
|
|
25
|
Booij JC, Boon CJ, van Schooneveld MJ, ten
Brink JB, Bakker A, et al: Course of visual decline in relation to
the Best1 genotype in vitelliform macular dystrophy. Ophthalmology.
117:1415–1422. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Fishman GA, Baca W, Alexander KR, Derlacki
DJ, Glenn AM, et al: Visual acuity in patients with best
vitelliform macular dystrophy. Ophthalmology. 100:1665–1670. 1993.
View Article : Google Scholar : PubMed/NCBI
|