Introduction
Cervical cancer is the second leading cause of all
cancer-associated mortalities in females after breast cancer
(1). As carcinogenesis is a
complex biological process involving a large variety of factors and
comprising multiple steps, it is necessary to enhance the current
understanding of the mechanisms involved in the spread of
carcinomas to identify novel therapeutic targets. MicroRNAs (miRs)
are small non-coding RNAs that are 18–24 nucleotides (nt) in length
and regulate gene expression at the post-transcriptional level
(2). miRs have been implicated in
the regulation of tissue morphogenesis and cellular
differentiation, and direct link between miRNAs and the pathology
of various diseases has been identified, including the occurrence,
development, invasion and metastasis of tumors (3–5).
The members of the miR-200 family have been reported
to have various functions (6),
among which the downregulation of tumor progression and inhibition
of epithelial-mesenchymal transition (EMT) are prominent (7,8). The
anti-tumor effects of miR-200 include repression of cancer stem
cell self-renewal (9), inhibition
of cell division (10) as well as
induction of apoptosis (11,12).
Mounting evidence has indicated that the EMT is a means by which
solid tissue epithelial cancers invade and metastasize (13–15).
Neoplastic cells undergoing EMT are characterized by an increased
potential to invade surrounding tissues and disseminate to distant
sites (16,17). During EMT, epithelial cells lose
phenotypic features, including apical-basal polarity and cell-cell
contacts, and acquire mesenchymal cell properties such as increased
cell mobility (18). The EMT is
characterized by downregulation of epithelial markers, such as
E-cadherin, which has a central role in maintaining normal
epithelial morphology, and gain of mesenchymal markers, including
fibronectin, vimentin and N-cadherin, accompanied by a loss of
cellular polarity (19). The
expression of miR-200b has been found to be suppressed in numerous
tumor types and, of note, miR-200b is a suppressor of the EMT
(20).
The present study assessed the role of miR-200b in
cervical cancer, focusing on its effects on the EMT and migration.
It was demonstrated that miR-200b acts as a tumor suppressor gene
in cervical cancer, and upregulation of miR-200b repressed the
migration and progression of EMT. The present study contributed to
the present understanding of the pathogenesis of cervical cancer
and provided an experimental and theoretical basis for the
treatment and inhibition of metastasization of cervical cancer.
Materials and methods
Ethics statement
Human samples used in the present study were
obtained according to the principles of the Declaration of
Helsinki, and all procedures were approved by the Ethics Committee
of the Renmin Hospital of Wuhan University (Wuhan, China). Written
informed consent was obtained from all patients.
Clinical specimens
The study cohort consisted of 13 patients with
invasive carcinoma of the cervix (ICC) at the early stage (termed
stage IA-IIA) identified by pathological diagnosis (age range,
29–66 years; mean age, 46 years), who were treated at Renmin
Hospital of Wuhan University (Wuhan, China) between February and
July 2014. A total of 13 cervical cancer tissues and 12 normal
cervical epithelial tissues were collected at the same time points.
Staging was performed according to the standard of the
International Federation of Gynecology and Obstetrics (FIGO)
(21).
Cell transfection
The human cervical carcinoma cell line HeLa was
purchased from the Type Culture Collection of the Chinese Academy
of Sciences (Shanghai, China). HeLa cells were cultured in RPMI
1640 medium (Thermo Fisher Scientific, Inc., Waltham, MA, USA)
supplemented with 10% fetal bovine serum (Gibco; Thermo Fisher
Scientific, Inc.), 100 U/ml penicillin and 100 µg/ml
streptomycin (Gibco; Thermo Fisher Scientific, Inc.) and cultured
in an incubator at 37°C with 5% CO2. The cells were then
seeded into six-well plates and grown overnight to reach ~40%
confluence. Cells were transfected with miR-200b mimics (Ribobio,
Guangzhou, China), miR-200b inhibitors (Ribobio) and negative
control miR (Ribobio) using Lipofectamine 2000 (Invitrogen, Thermo
Fisher Scientific, Inc.) in Opti-MEM (Invitrogen) according to the
manufacturer's protocol. After 6 h of transfection, the medium
(DMEM; Invitrogen) was removed and replaced with complete growth
medium. At 48 h after transfection, cells were harvested for RNA or
protein analysis.
Reverse-transcription quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was extracted with TRIzol regent
(Invitrogen) in accordance with the manufacturer's protocol. For
analysis of miR-200b expression, cDNA was synthesized using
specific reverse transcription primers (miR-200b, F 5′-GCG GCT AAT
ACT GCC TGG TAA-3′; R 5′-GTG CAG GGT CCG AGGT-3′; U6, F 5′-CGC TTC
GGC AGC ACA TAT ACTA-3′; R 5′-CGC TTC ACG AAT TTG CGT GTCA-3′).
purchased from Ribobio and the miRNA First-strand cDNA Synthesis
kit (Fermentas, Vilnius, Lithuania). Real-time PCR was performed in
the Step One Plus Real-Time system (Applied Biosystems, Thermo
Fisher Scientific, Inc.) with ABI SYBR Green Master Mix (Applied
Biosystems), and U6 was used as the endogenous reference gene. The
thermocycling conditions were as follows: 95°C for 10 min, followed
by 40 cycles of 95°C for 15 sec and 60°C for 1 min. The gene
expression levels were calculated relative to the expression of U6
using the 2−ΔΔCq method (22).
Cell migration assays
Cell migration was examined by a Transwell migration
assay. Transwell insert chambers with 8-µm porous membranes
(Corning Inc., Corning, NY, USA) were used. Following 24 h of
transfection, the cells were dislodged from the culture vessel
surface with trypsin (Gibco, Thermo Fisher Scientific, Inc.)
suspended at 2×105 cells/ml. For each group, 200
µl cell suspension was seeded into the upper Transwell
chambers. The lower compartment contained 10% fetal calf serum
(Gibco). After 24 h of incubation, cells on the upper side of the
membrane were removed, followed by washing with phosphate-buffered
saline (PBS) three times. The migrated cells on the bottom side of
the membrane were then fixed with methanol (Goodbio Technology,
Wuhan, China) for 15 min at room temperature and stained with 0.5%
crystal violet (Sigma-Aldrich, St. Louis, MO, USA) in 25% methanol
for 5 min. Images of migrated cells in at least six random selected
fields were captured under a microscope at 100× magnification and
cells were counted. Experiments were performed in technical and
biological triplicates. One representative high-power field per
Transwell membrane was counted (using six randomly selected
high-power fields per treatment condition).
Western blot analysis
Following transfection, cells were lysed in
radioimmunoprecipitation assay buffer (Beyotime Institute of
Biotechnology, Haimen, China). The obtained protein extracts were
centrifuged at 16,099 × g for 15 min. The protein concentrations
were determined using a bicinchoninic acid protein assay kit
(Beyotime Institute of Biotechnology). For western blot analysis,
equal quantity (30 µg) of protein was loaded in each lane
and subjected to 10% sodium dodecyl sulfate polyacrylamide gel
electrophoresis. Proteins were then transferred onto a
polyvinylidene difluoride membrane (EMD Millipore, Billerica, MA,
USA). After blocking, the membranes were probed with antibodies
against monoclonal anti-rabbit human E-cadherin (1:1,000; cat. no.
3195), monoclonal anti-rabbit human vimentin (1:1,000; cat. no.
5741) or anti-rabbit human GAPDH (1:1,000; cat. no. 2118; Cell
Signaling Technology, Inc., Danvers, MA, USA), or monoclonal
anti-mouse human matrix metalloproteinase (MMP-9) antibody (1:800;
cat. no. ab58803; Abcam, Cambridge, UK) overnight at 4°C. After
washing in Tris-buffered saline containing Tween 20 (TBST), the
membranes of E-cadherin, vimentin and GAPDH were incubated with
bovine anit-rabbit horseradish peroxidase-conjugated secondary
antibody (1:8,000; cat. no. sc-2370; Santa Cruz Biotechnology,
Inc., Dallas, TX, USA) and MMP-9 membranes were incubated with
bovine anti-mouse horseradish peroxidase-conjugated secondary
antibody (1:8,000; cat. no. sc2371; Santa Cruz Biotechnology, Inc.)
at 37°C for 1 h. After washing of the membranes in TBST, the
proteins were visualized using an enhanced chemiluminescence
detection kit (EMD Millipore) and specific antibody binding was
visualized and quantified using Image J software (version 1.46;
National Institutes of Health, Bethesda, MD, USA). Experiments were
performed three times.
Immunofluorescence analysis
Cells grown on coverslips in a culture plate were
washed three times with cold PBS and fixed with 4% paraformaldehyde
for 15 min. Coverslips were then washed three times with PBS and
rinsed with 0.5% Triton X-100 in PBS for 20 min. After blocking
with goat serum (Goodbio Technology) for 30 min at room
temperature, cells were incubated with primary antibody overnight
at 4°C. Coverslips were rinsed three times with PBS containing
Tween 20 (PBST) and then labeled with the secondary antibody for 1
h in the dark at room temperature. After incubation, coverslips
were then washed three times with PBST and incubated with DAPI
(Beyotime Institute of Biotechnology) for 5 min in the dark and
washed again with PBST. Following mounting with Antifade Mounting
Medium (EMD Millipore), images were acquired with a fluorescence
microscope (CKX31; Olympus Corp., Tokyo, Japan).
Statistical analysis
Statistical analyses were performed with SPSS 19.0
software (International Business Machines, Armonk, NY, UA). Values
are expressed as the mean ± standard deviation of at least three
independent experiments. The normality test was performed using the
K-S test and the homogeneity test for variance was conducted by
Levene's test. Statistical analysis was performed using the
independent-samples t-test (normal distribution, equal
variances) and one-way analysis of variance followed by the
least-significant differences test (equal variances) or Dunnett's
T3 test (unequal variances) when appropriate. P<0.05 was
considered to indicate a statistically significant difference
between values.
Results
miR-200b is downregulated in human
ICC
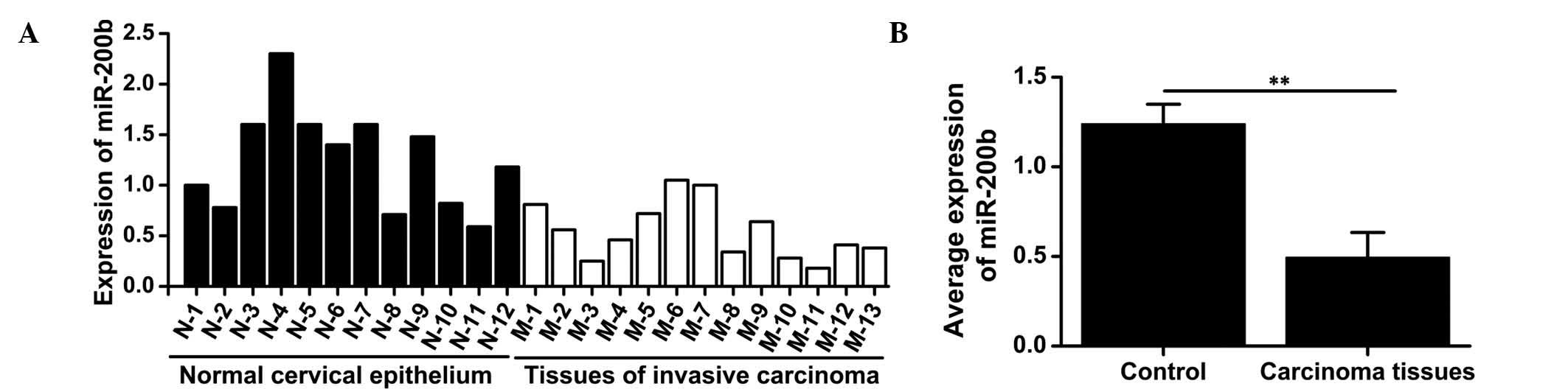
To investigate the expression of miR-200b in the
cervical cancer tissues and normal adjacent tissues, RT-qPCR was
performed on 13 tumor specimens and 12 normal cervical epithelial
samples. While miR-200b was expressed in all samples (Fig. 1A), the ICC tissues exhibited a
significantly lower expression of miR-200b compared with that in
normal tissues (t=7.353; P=0.002) (Fig. 1B), suggesting that miR-200b may
have a tumor suppressor role in ICC.
Effects of the transfection of miR-200b
mimics or inhibitors on miR-200b expression
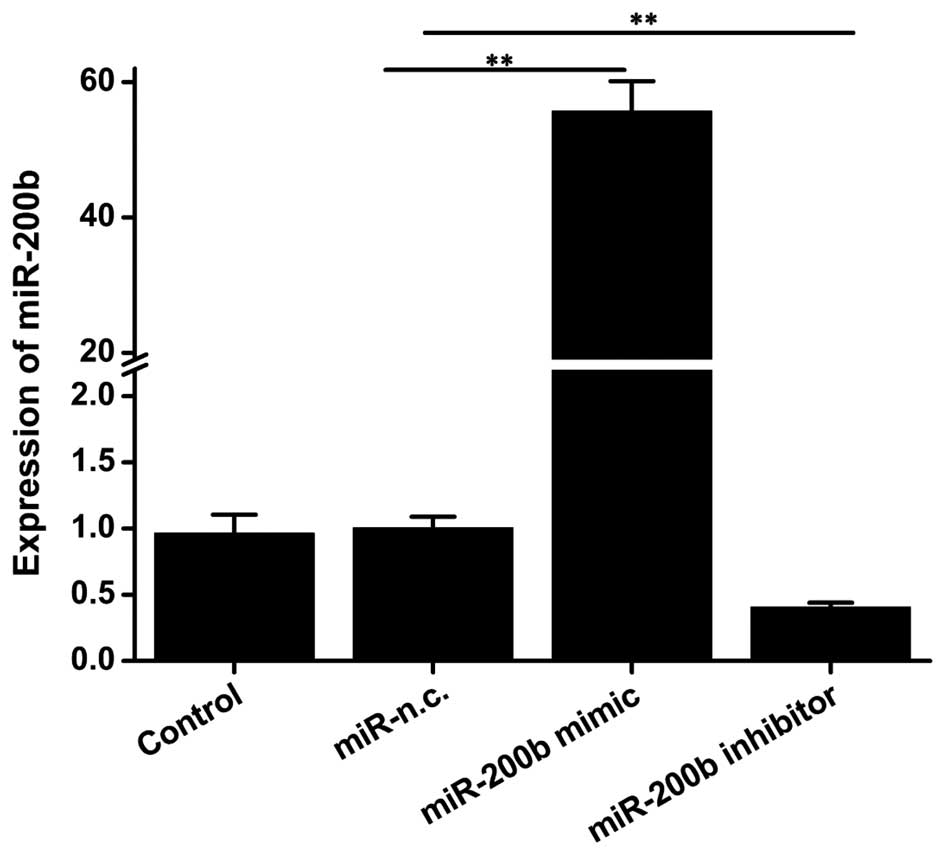
For loss-and gain-of-function studies of miR-200b in
ICC, HeLa cells were transfected with miR-200b mimics and
inhibitor, and the resulting upregulation or knockdown of miR-200b
gene expression were confirmed by RT-qPCR. Transfection of miR-200b
mimics significantly increased the expression of miR-200b to 50
fold of that of the negative control group (P=0.007) (Fig. 2), while a significant decrease by
0.4-fold was achieved in cells transfected with inhibitor (P=0.01).
These results suggested the successful overexpression and
knockdown, respectively, of miR-200b.
miR-200b exerts an anti-migratory effect
on ICC
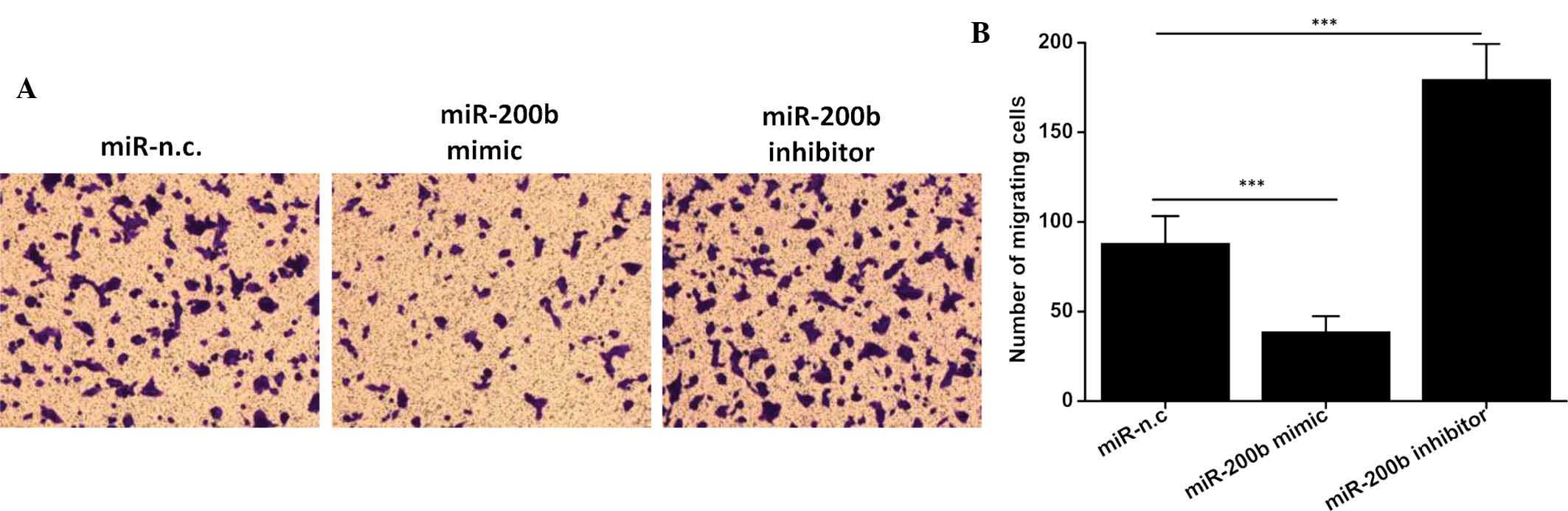
The migratory capacity of neoplastic cells is linked
with their degree of malignancy due to its association with
metastasis and represents an important potential therapeutic
target. Therefore, the present study examined the ability of HeLa
cells transfected with miR-200b mimics or miR-200b inhibitor to
migrate to the lower compartment. As shown in Fig. 3, transfection with miR-200b mimics
significantly inhibited the cell migratory ability compared with
that of the negative control group (P<0.001). Conversely, a
significant elevation of the cell migratory ability was found in
cells transfected with miR-200b inhibitor (P<0.001). These
results suggested that miR-200b has an inhibitory effect on the
migratory ability of ICC, explaining for the downregulation of this
tumor suppressor miR in this invasive cancer type.
miR-200b suppresses the EMT of cervical
carcinoma cells
To test whether miR-200b affects the expression of
proteins involved in the EMT, HeLa cells were transfected with
miR-200b mimics or inhibitor. In cells transfected with miR-200b
mimics, a substantial elevation of E-cadherin (P<0.001) and a
reduction in vimentin (P=0.002) and MMP-9 (P=0.002) was observed
compared to that in the negative control group (Fig. 4A–D). Conversely, inhibition of
miR-200b reduced the expression of E-cadherin (P=0.002), but
increased the expression of vimentin (P<0.001) and MMP-9
(P<0.001) (Fig. 4A–D). These
results indicated that miR-200b was able to suppress the EMT
process of HeLa cells, and to therefore potentially inhibit
cervical carcinoma cell metastasis. To further verify the effects
of miR-200b on the EMT process of cervical carcinoma cells, the
expression of E-cadherin, which mediates cell-cell adhesion and has
a pivotal role in epithelial cell behavior and tissue
morphogenesis/remodelling, was determined by immunofluorescence
analysis. As shown in Fig. 4E,
miR-200b mimics elevated the expression of E-cadherin, while
inhibition of miR-200b reduced E-cadherin. These findings provided
further evidence to support the causal role of miR-200b in the EMT
process of cervical carcinoma.
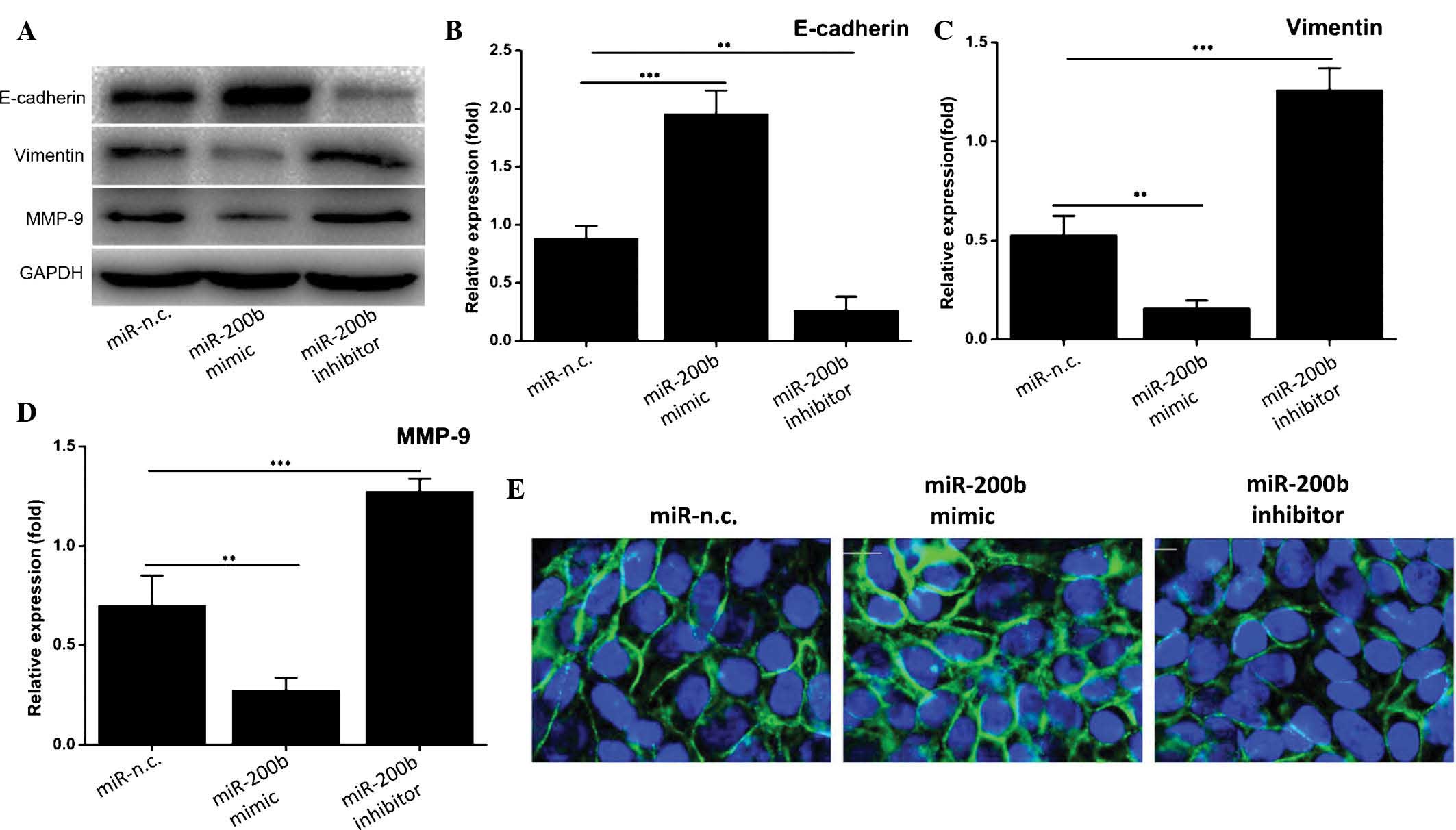 | Figure 4miR-200b suppresses the process of
epithelial-mesenchymal transition of cervical carcinoma cells. (A)
Western blot analysis of the levels of E-cadherin, vimentin and
MMP-9. GAPDH was used as a loading control. A representative blot
of three experiments with similar results is shown. (B-D) The
expression of E-cadherin, vimentin and MMP-9, respectively, was
densitometrically quantified and normalized to GAPDH. Values are
expressed as the mean ± standard deviation (n=3).
**P≤0.01, ***P≤0.001. (E) The expression of
E-cadherin (green) was determined by immunofluorescence analysis.
Nuclei were counterstained with DAPI (blue). Magnification, ×400.
MMP, matrix metalloproteinase; n.c., negative control; GAPDH,
glyceraldehyde-3-phosphate dehydrogenase; miR, microRNA. |
Discussion
A large number of miRs have expression patterns that
are unique within certain types of cancer tissue (23). It has been suggested that these
miRs are potential biomarkers and powerful diagnostic tools for
tumor classification and diagnosis (23,24).
The aberrant expression of certain miRs in cervical carcinoma is
associated with the proliferation, apoptosis, invasion and
metastasis of tumor cells (3–5).
miR-200b expression has been shown to be decreased in various tumor
types, including meningeoma, nasopharyngeal carcinoma, anaplastic
thyroid carcinoma and ovarian cancer (24,25).
Therefore, the present study assessed miR-200b expression in early
ICC by RT-qPCR. miR-200b was found to be downregulated in ICC
tissues compared with that in normal cervical epithelium,
suggesting that miR-200b may be involved in tumor cell metastasis
and invasion. Furthermore, HeLa cells were used to explore the role
of miR-200b in cervical carcinoma progression and the underlying
molecular mechanisms by transfecting them with miR-200b mimics,
inhibitor or negative control miR. A Transwell assay revealed that
cells transfected with miR-200b mimics exhibited a reduced
migratory potential compared with that of cells transfected with
negative control miR, while inhibition of miR-200b enhanced cell
migration. This result suggested that miR-200b is involved in the
migration and metastasis of HeLa cells.
EMT is a critical process in embryogenesis and a
particularly important process during early tumor invasion and
metastasis. EMT facilitates the metastasis and progression of
tumors by enhancing the migratory and invasive potential of cells
(17). Metastatic tumor cells show
characteristics of mesenchymal cells as a result of EMT. During
cancer progression, transcriptional E-cadherin reprogramming in
epithelial cells leads to EMT with decreased adhesion and enhanced
migration/invasion. The present study found that overexpression of
miR-200b in HeLa cells led to the upregulation of the epithelial
marker E-cadherin and downregulation of the mesenchymal marker
vimentin, while inhibition of miR-200b had the opposite effects.
These results suggested that miR-200b inhibits cervical carcinoma
cell metastasis by suppressing EMT. Reduction of E-cadherin
expression is a crucial step during EMT. In the present study,
immunofluorescence microscopy indicated that E-cadherin expression
in HeLa cells was increased following transfection with miR-200b
mimics, while it was decreased following miR-200b inhibition. It is
known that miR-200 family members regulate EMT by targeting
zinc-finger E-box binding homeobox (ZEB) proteins, particularly
ZEB1 and ZEB2, and that ZEB2 inhibits the transcription of proteins
including E-cadherin and cytokeratin by combining with the
E-cadherin gene promoter at the E-box sequence (26–28).
ZEB1 and ZEB2 are involved in the EMT through the transforming
growth factor-β (29), nuclear
factor-κB (30) and Notch
(31) pathways. The degradation of
extracellular matrix and basement membrane are key steps in tumor
invasion and metastasis, and MMP-9 has a central role in this
process (32). The present study
revealed that overexpression of miR-200b inhibited the expression
of MMP-9, leading to the suppression of cervical carcinoma cell
metastasis, while the miR-200b inhibitor had the opposite effect.
These results further confirmed the involvement of miR-200b in the
regulation of the EMT of cervical cancer cells.
In conclusion, the present study demonstrated that
miR-200b has a tumor suppressor role in cervical carcinoma.
miR-200b was significantly downregulated in ICC tissues compared
with normal adjacent tissues. Upregulation of miR-200b in HeLa
cells reduced their migratory potential and inhibited EMT, as
indicated by enhanced E-cadherin expression and reduced vimentin
and MMP-9 expression, while inhibition of miR-200b had the opposite
effect. These results indicated that miR-200b may be able to
inhibit the progression and metastasization of cervical cancer.
miR-200b mimics may therefore be utilized for targeting of cervical
carcinoma and other tumor types.
Acknowledgments
The present study was supported by the Natural
Science Foundation of China (nos. 81302273 and 81201196), the
Science and Technology Department of Hubei Province, China (no.
ZRY039), the Health and Family Planning Commission of Hubei
Province, China (no. 2012ZY02) and the Chinese Postdoctoral Science
Foundation (no. 2011M500857).
References
|
1
|
Cai HB, Ding XH and Chen CC: Prevalence of
single and multiple human papillomavirus types in cervical cancer
and precursor lesions in Hubei, China. Oncology. 76:157–161. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Granados López AJ and López JA: Multistep
model of cervical cancer: Participation of miRNAs and coding genes.
Int J Mol Sci. 15:15700–15733. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kloosterman WP and Plasterk RH: The
diverse functions of microRNAs in animal development and disease.
Dev Cell. 11:441–450. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kestens C, Siersema PD and van Baal JW:
Current understanding of the functional roles of aberrantly
expressed microRNAs in esophageal cancer. World J Gastroenterol.
22:1–7. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Frixa T, Donzelli S and Blandino G:
Oncogenic microRNAs: Key players in malignant transformation.
Cancers (Basel). 7:2466–2485. 2015. View Article : Google Scholar
|
|
6
|
O'Day E and Lal A: MicroRNAs and their
target gene networks in breast cancer. Breast Cancer Res.
12:2012010. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lim YY, Wright JA, Attema JL, Gregory PA,
Bert AG, Smith E, Thomas D, Lopez AF, Drew PA, Khew-Goodall Y and
Goodall GJ: Epigenetic modulation of the miR-200 family is
associated with transition to a breast cancer stem-cell-like state.
J Cell Sci. 126:2256–2266. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Schliekelman MJ, Gibbons DL, Faca VM,
Creighton CJ, Rizvi ZH, Zhang Q, Wong CH, Wang H, Ungewiss C, Ahn
YH, et al: Targets of the tumor suppressor miR-200 in regulation of
the epithelial-mesenchymal transition in cancer. Cancer Res.
71:7670–7682. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Iliopoulos D, Lindahl-Allen M, Polytarchou
C, Hirsch HA, Tsichlis PN and Struhl K: Loss of miR-200 inhibition
of Suz12 leads to polycomb-mediated repression required for the
formation and maintenance of cancer stem cells. Mol Cell.
39:761–772. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Uhlmann S, Zhang JD, Schwäger A,
Mannsperger H, Riaz-alhosseini Y, Burmester S, Ward A, Korf U,
Wiemann S and Sahin O: miR-200bc/429 cluster targets PLCgamma1 and
differentially regulates proliferation and EGF-driven invasion than
miR-200a/141 in breast cancer. Oncogene. 29:4297–4306. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Schickel R, Park SM, Murmann AE and Peter
ME: miR-200c regulates induction of apoptosis through CD95 by
targeting FAP-1. Mol Cell. 38:908–915. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Magenta A, Cencioni C, Fasanaro P,
Zaccagnini G, Greco S, Sarra-Ferraris G, Antonini A, Martelli F and
Capogrossi MC: miR-200c is upregulated by oxidative stress and
induces endo-thelial cell apoptosis and senescence via ZEB1
inhibition. Cell Death Differ. 18:1628–1639. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yang J and Weinberg RA:
Epithelial-mesenchymal transition: At the crossroads of development
and tumor metastasis. Dev Cell. 14:818–829. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Thompson EW, Newgreen DF and Tarin D:
Carcinoma invasion and metastasis: A role for
epithelial-mesenchymal transition? Cancer Res. 65:5991–5995. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
von Burstin J, Eser S, Paul MC, Seidler B,
Brandl M, Messer M, von Werder A, Schmidt A, Mages J, Pagel P, et
al: E-cadherin regulates metastasis of pancreatic cancer in vivo
and is suppressed by a SNAIL/HDAC1/HDAC2 repressor complex.
Gastroenterology. 137:361–371. 371.e1–e5. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Haslehurst AM, Koti M, Dharsee M, Nuin P,
Evans K, Geraci J, Childs T, Chen J, Li J, Weberpals J, et al: EMT
transcription factors snail and slug directly contribute to
cisplatin resistance in ovarian cancer. BMC Cancer. 12:912012.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Kang Y and Massagué J:
Epithelial-mesenchymal transitions: Twist in development and
metastasis. Cell. 118:277–279. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Lamouille S, Xu J and Derynck R: Molecular
mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell
Biol. 15:178–196. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen Y, Xiao Y, Ge W, Zhou K, Wen J, Yan
W, Wang Y, Wang B, Qu C, Wu J, et al: miR-200b inhibits
TGF-β1-induced epithelial-mesenchymal transition and promotes
growth of intestinal epithelial cells. Cell Death Dis. 4:e5412013.
View Article : Google Scholar
|
|
21
|
Pecorelli S: Revised FIGO staging for
carcinoma of the vulva, cervix, and endometrium. Int J Gynaecol
Obstet. 105:103–104. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
23
|
Lim J and Thiery JP:
Epithelial-mesenchymal transitions: Insights from development.
Development. 139:3471–3486. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Vergara D, Merlot B, Lucot JP, Collinet P,
Vinatier D, Fournier I and Salzet M: Epithelial-mesenchymal
transition in ovarian cancer. Cancer Lett. 291:59–66. 2010.
View Article : Google Scholar
|
|
25
|
Park SM, Gaur AB, Lengyel E and Peter ME:
The miR-200 family determines the epithelial phenotype of cancer
cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes
Dev. 22:894–907. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Burk U, Schubert J, Wellner U, Schmalhofer
O, Vincan E, Spaderna S and Brabletz T: A reciprocal repression
between ZEB1 and members of the miR-200 family promotes EMT and
invasion in cancer cells. EMBO Rep. 9:582–589. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Gregory PA, Bert AG, Paterson EL, Barry
SC, Tsykin A, Farshid G, Vadas MA, Khew-Goodall Y and Goodall GJ:
The miR-200 family and miR-205 regulate epithelial to mesenchymal
transition by targeting ZEB1 and SIP1. Nat Cell Biol. 10:593–601.
2008. View
Article : Google Scholar : PubMed/NCBI
|
|
28
|
Korpal M, Lee ES, Hu G and Kang Y: The
miR-200 family inhibits epithelial-mesenchymal transition and
cancer cell migration by direct targeting of E-cadherin
transcriptional repressors ZEB1 and ZEB2. J Biol Chem.
283:14910–14914. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Postigo AA: Opposing functions of ZEB
proteins in the regulation of the TGFbeta/BMP signaling pathway.
EMBO J. 22:2443–2452. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chua HL, Bhat-Nakshatri P, Clare SE,
Morimiya A, Badve S and Nakshatri H: NF-kappaB represses E-cadherin
expression and enhances epithelial to mesenchymal transition of
mammary epithelial cells: Potential involvement of ZEB-1 and ZEB-2.
Oncogene. 26:711–724. 2007. View Article : Google Scholar
|
|
31
|
Wang Z, Li Y, Kong D, Banerjee S, Ahmad A,
Azmi AS, Ali S, Abbruzzese JL, Gallick GE and Sarkar FH:
Acquisition of epithelial-mesenchymal transition phenotype of
gemcitabine-resistant pancreatic cancer cells is linked with
activation of the notch signaling pathway. Cancer Res.
69:2400–2407. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Xu E, Xia X, Lü B, Xing X, Huang Q, Ma Y,
Wang W and Lai M: Association of matrix metalloproteinase-2 and -9
promoter polymorphisms with colorectal cancer in Chinese. Mol
Carcinog. 46:924–929. 2007. View
Article : Google Scholar : PubMed/NCBI
|