Introduction
Bladder cancer is the most common type of cancer of
the urinary system, and is associated with high morbidity and
mortality rates (1,2). It was estimated that ~430,000 cases
of bladder cancer were diagnosed and there were 165,000 cases of
bladder cancer-associated mortality worldwide in 2012 (3). Although mortality rates of bladder
cancer are decreasing, the trend is moderate, and the genetic
factors contributing to the tumorigenesis and progression of
bladder cancer remain to be fully elucidated (4,5).
MicroRNAs (miRNAs), which are ~20 nucleaotides in
length, are a class of non-coding RNA. Accumulating evidence
suggests that miRNAs are central to several pathological processes
by regulating ~50% of human protein-coding genes at the
post-transcriptional level (6).
Several studies have shown that miR-23a-5p, which is suppressed by
c-Myc and promyelocytic leukemia protein-retinoic acid receptor-α
fusion protein in prostate cancer and acute promyelocytic leukemia,
respectively, functions as a tumor suppressor (7). By contrast, miR-23a-5p has been
reported to be significantly upregulated in bladder cancer via
microchip assays (8,9). Therefore, the expression and function
of miR-23a-5p in bladder cancer requires further investigation.
In the present study, it was found that miR-23a-5p
was upregulated in bladder cancer tissues, and that the inhibition
of miR-23a-5p suppressed the growth and migration abilities of
cells, and induced apoptosis.
Materials and methods
Patient samples
A total of 44 patients pathologically diagnosed with
urothelial carcinoma of the bladder were included in the present
study. All paired bladder cancer tissues and adjacent normal
bladder tissues (located at least 2.0 cm outside of visible bladder
cancer lesions) were immediately immersed in RNAlater®
RNA Stabilization agent (Qiagen GmbH, Hilden, Germany) following
surgical resection, and were snap-frozen in liquid nitrogen and
stored in a cryo freezer at −80°C for further use. All samples were
collected at Department of Urology, Peking University Shenzhen
Hospital from September 2012 to December 2015. The present study
was approved by the Ethics Committee of Peking University Shenzhen
Hospital (Shenzhen, China). Written informed consent was obtained
from all the patients involved. The clinical parameters of patients
are listed in Table I.
 | Table I.Clinicopathological features of
patients with bladder cancer. |
Table I.
Clinicopathological features of
patients with bladder cancer.
| Characteristic | Cases (n) |
|---|
| Median age (range),
years | 67 (38–85) |
| Sex |
|
|
Male/female | 36/8 |
| Histological
type |
|
|
Transitional cell
carcinoma | 44 |
| Grade |
|
|
Low/high | 18/26 |
| Stage |
|
|
I/II/III/IV | 18/21/5/0 |
Cell culture
The T24 and SW780 human bladder cancer cell lines
were purchased from the Institute of Cell Biology, Chinese Academy
of Sciences (Shanghai, China). All cells were cultured in
Dulbecco's modified Eagle's medium (DMEM; Gibco; Thermo Fisher
Scientific, Inc., Waltham, MA, USA), with 10% fetal bovine serum
(Gibco; Thermo Fisher Scientific, Inc.), 1% antibiotics (100 µ/ml
penicillin and 100 mg/ml streptomycin sulfates) and 1% glutamate
(Gibco; Thermo Fisher Scientific, Inc.), at 37°C in a 5%
CO2 atmosphere (10).
Cell transfection
For the suppression of miR-23a-5p, chemically
synthesized miR-23a-5p inhibitor and negative control (NC) were
purchased from GenePharma (Shanghai, China). The sequences were as
follows: Sense, 5′-AAAUCCCAUCCCCAGGAACCCC-3′ and antisense,
5′-CAGUACUUUUGUGUAGUACAA-3′, respectively. Lipofectamine 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) was mixed with the 100
pmol miR-23a-5p inhibitor or NC for transfection, according to the
manufacturer's protocol.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR) analysis
Total RNA was extracted from the tissue samples or
the transfected cells using TRIzol reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) according to the manufacturer's protocol.
The RNA samples with 260/280 ratios of 1.8–2.0 were used for
further experiments. Total RNA was converted into cDNA using the
miScript II RT kit (Qiagen GmbH). The primer sequences were as
follows: miR-23a-5p, forward 5′-GGGGUUCCUGGGGAUGGGAUUU-3′ and
reverse: Universal primer, provided with the miScript
SYBR®-Green PCR kit (Qiagen GmbH); U6 forward,
5′-CTCGCTTCGGCAGCACA-3′ and reverse, 5′-ACGCTTCACGAATTTGCGT-3′. The
qPCR procedure was performed to determine the expression levels of
miR-23a-5p using the miScript SYBR®-Green PCR kit
(Qiagen GmbH), according to the manufacturer's protocol. The
thermocycling conditions for RT-qPCR were as follows: 95°C for 15
min, followed by 40 cycles of 94°C for 15 sec, 55°C for 30 sec and
72°C for 30 sec. The reactions were performed in triplicate using
the LightCycler 480 real-time PCR system (Roche Applied Science,
Mannheim, Germany). U6 was selected as the internal control. The
expression level was determined as the fold difference relative to
U6, which was based on the following equation: Relative expression
= 2−ΔΔCq, ΔΔCq = (meanCqcancer -
meanCqu6) - (meanCqnormal -
meanCqu6) (11).
3-(4,5-dimeth-ylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay
Cell proliferation was analyzed using an MTT (5
mg/ml; Sigma-Aldrich; Merck KGaA, Darmstadt, Germany) assay. The
T24 and SW780 cells, at ~5,000 cells per-well, were plated into
96-well plates, with five replicate wells for each condition. After
24 h, each well was transfected with 5 pmol miR-23a-5p inhibitor or
NC. Cell growth was assessed at 0, 24, 48 and 72 h
post-transfection. The MTT (20 µl) was added into each well and
incubated at 37°C for 6 h. The MTT medium was then discarded and 12
µl dimethyl sulphoxide (Sigma-Aldrich; Merck KGaA) was added.
Following agitation for 30 min at room temperature, the optical
density (OD) value of each sample was measured with an enzyme
immunoassay instrument (Bio-Rad Laboratories, Inc., Hercules, CA,
USA) at a wavelength of 490 nm (with 630 nm as the reference
wavelength).
Wound-healing assay
Cell migration was determined with a wound-healing
assay. Briefly, ~3×105 cells were seeded into 12-well
plates at equal density and grown to 80–90% confluence. Artificial
gaps/wounds were generated using a 200-µl sterile pipette tip
following transfection. The regions of the wounds were marked and
images were captured using a digital camera system (Olympus
Corporation, Tokyo, Japan). The cell migration distance (mm) was
calculated using the HMIAS-2000 (v1.0; Champion Image Co., Ltd.,
Wuhan, China) software program. Each experiment was repeated three
times.
Flow cytometry
Flow cytometry was performed to determine the early
apoptotic rate. The cells were seeded (~3×105 cells per
well) into 6-well plates at equal density. At 48 h
post-transfection, the cells in each well, including floating
cells, were collected and stained with 3 µl propidium iodide (PI)
and 5 µl Annexin V-fluorescein isothiocyanate (Invitrogen; Thermo
Fisher Scientific, Inc.), according to the manufacturer's protocol.
Within 30 min, each sample was measured and analyzed on a flow
cytometer (Coulter Epic XL-4; Beckman Coulter, Inc., Brea, CA, USA)
using EXPO32 ADC software V1.2 (Beckman Coulter).
Statistical analysis
All data are expressed as the mean ± standard
deviation. The significance of differences were determined using
SPSS version 19.0 software (IBM Corps., Armonk, NY, USA). The
differences in the expression of miR-23a-5p between bladder cancer
tissues and adjacent normal tissues were analyzed using a
paired-sample t-test. The independent-samples t-test was used to
analyze other data. P<0.05 was considered to indicate a
statistically significant difference.
Results
miR-23a-5p is upregulated in bladder
cancer
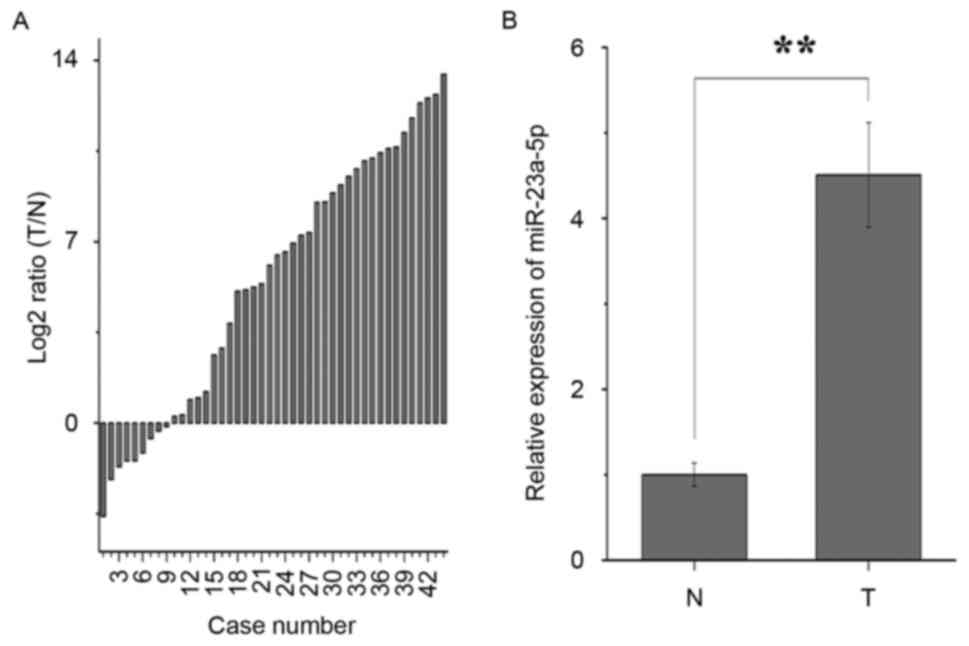
The results of the RT-qPCR analysis showed that
miR-23a-5p was significantly upregulated in 35/44 patients with
bladder cancer (Fig. 1A). As shown
in Fig. 1B, miR-23a-5p was
upregulated in bladder cancer tissues, compared the adjacent normal
tissues, with an average of a 4.509-fold increase in expression, in
accordance with previous studies (8,9).
These results indicated that miR-23a-5p may be have an oncogenic
role in bladder cancer. The clinical parameters of patients are
listed in Table I.
miR-23a-5p inhibitor downregulates the
expression of miR-23a-5p
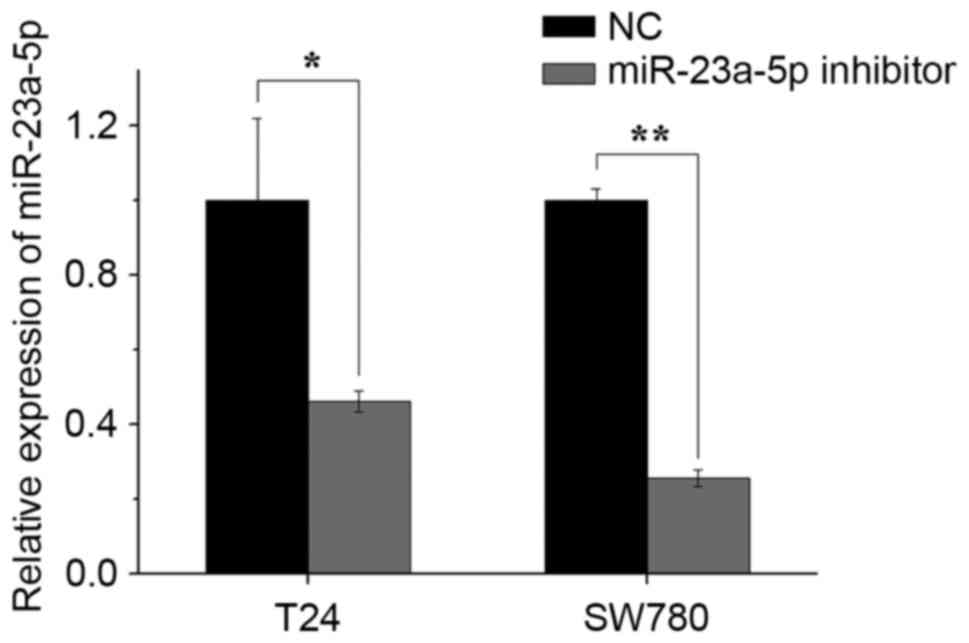
The silencing efficiency of the miR-23a-5p inhibitor
was validated using RT-qPCR analysis 48 h post-transfection. As
shown in Fig. 2, the expression
levels of miR-23a-5p in the miR-23a-5p inhibitor group were
decreased by 53.7% in T24 cells and 74.3% in the SW780 cells,
compared with those in the NC group.
Silencing miR-23a-5p suppresses cell
growth
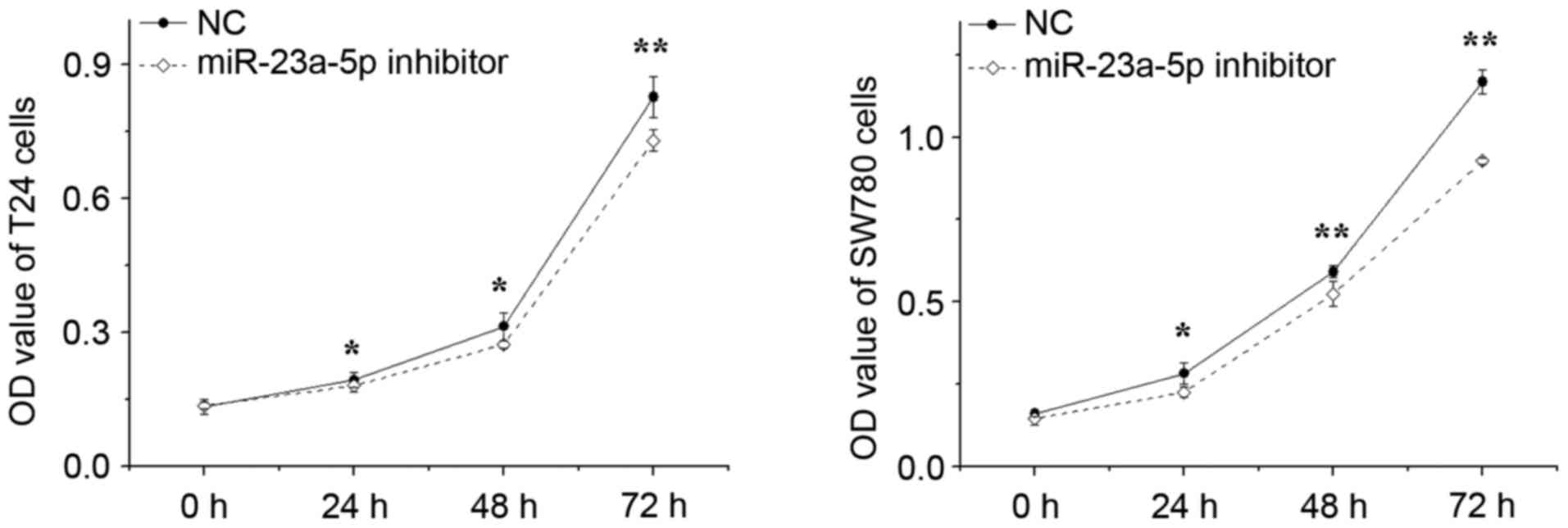
The effect of miR-23a-5p inhibitor on the growth of
bladder cancer cells was determined using MTT assays. The outcomes
revealed that, following silencing of miR-23a-5p, the proliferation
of T24 cells decreased by 6.23% (P<0.05), 12.71% (P<0.05) and
11.72% (P<0.01), and the proliferation of SW780 cells decreased
by 19.91% (P<0.05), 11.24% (P<0.05) and 20.55% (P<0.05) at
24, 48 and 72 h, respectively, compared with proliferation in the
NC group (Fig. 3). These results
indicated that the downregulation of miR-23a-5p significantly
decreased the proliferation of bladder cancer cells.
Silencing miR-23a-5p attenuates cell
migration
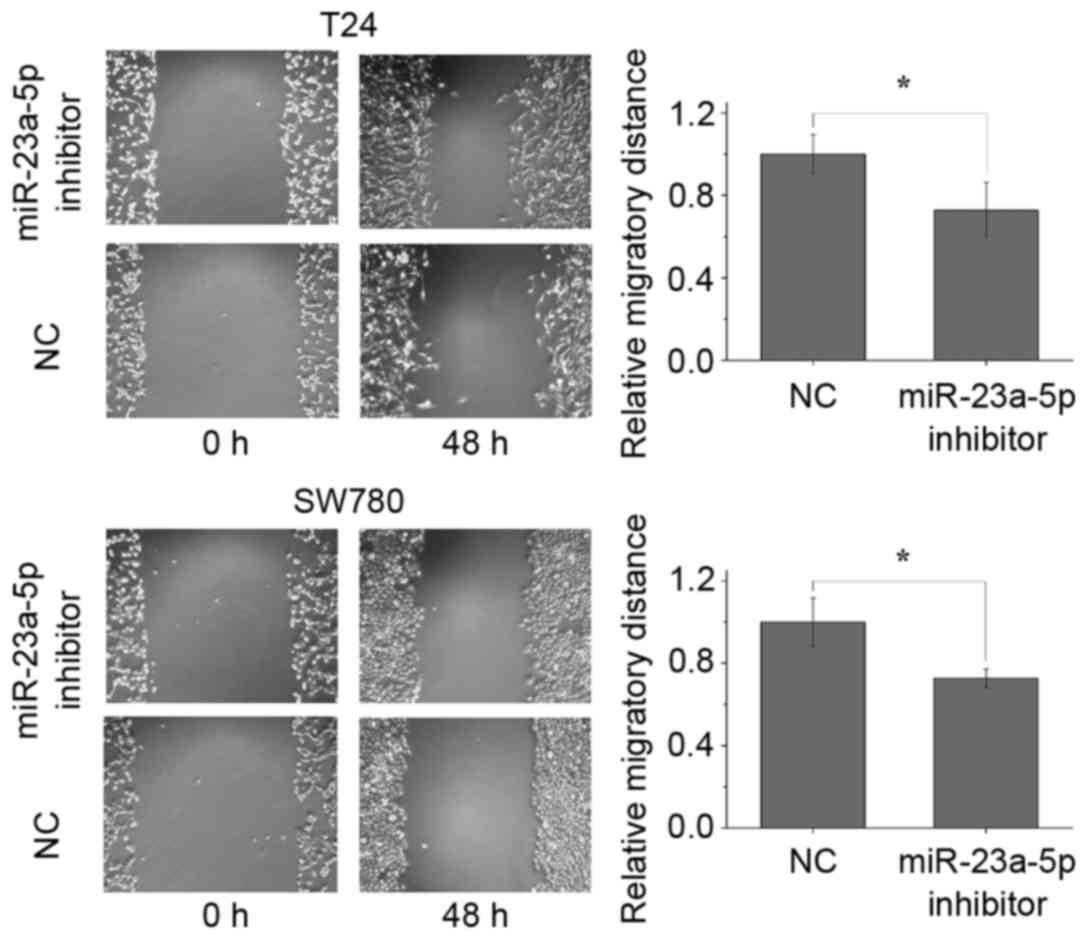
The present study performed wound-healing assays to
observe the effect of miR-23a-5p on cell migratory ability. The
results showed that the migratory distances of the miR-23a-5p
inhibitor group were significantly decreased by 27.2% (P<0.05)
and 27.3% (P<0.05) for the T24 cells and SW780 cells, at 48 h
post-transfection, compared with distances in the NC group
(Fig. 4). This suggested that the
silencing of miR-23a-5p attenuated the migratory abilities of the
bladder cancer cells.
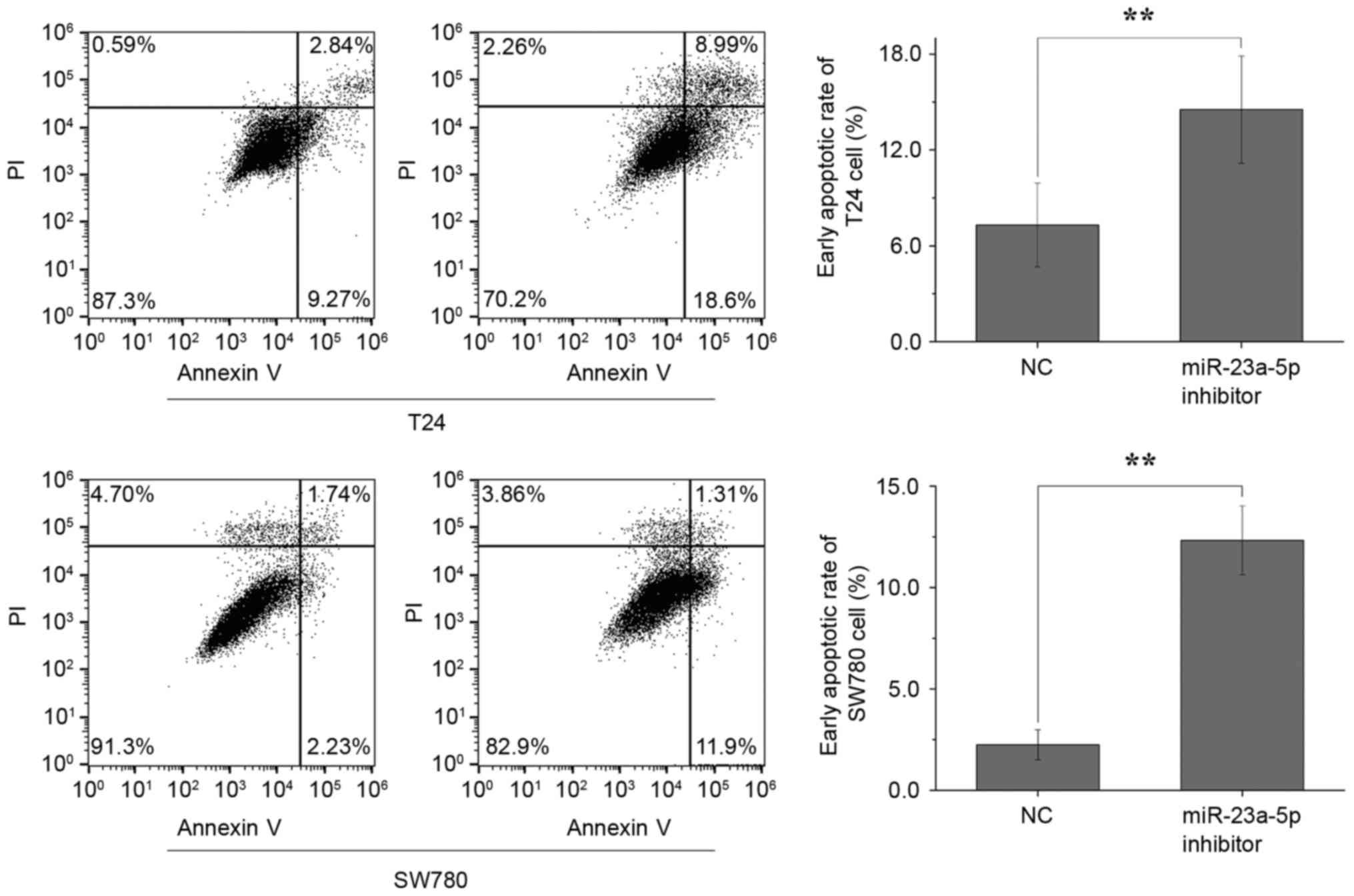
Silencing miR-23a-5p induces
apoptosis
Finally, flow cytometric analysis was performed to
determine whether the miR-23a-5p inhibitor had an effect on bladder
cancer cell apoptosis. The results demonstrated that the average
early apoptotic rates of the T24 cells transfected with miR-23a-5p
inhibitor and NC, were 7.31 and 14.53%, respectively (P<0.01),
whereas the average early apoptotic rates of the SW780 cells
transfected with miR-23a-5p inhibitor and NC were 2.25 and 12.33%,
respectively (P<0.01) (Fig. 5).
These data suggested that the silencing of miR-23a-5p induced
bladder cell apoptosis.
Discussion
miRNAs have emerged as key post-transcriptional
regulators of gene expression and are involved in the regulation of
almost every cellular process (6).
Dysregulation of miRNAs are associated with several human
pathologies, including tumorigenesis and tumor progression
(6). Oncogenic miRNAs targeting
key tumor suppressor genes, including the miR-17-92 cluster and
miR-21, are upregulated in cancer (12). miR-17 and miR-20a, of the miR-17-92
cluster, target E2F transcription factor 1, a cell cycle regulator
involved in cell division and apoptosis (13). miR-21 in breast cancer and colon
cancer leads to increased tumor growth by downregulating programmed
cell death 4, a protein involved in the promotion of cellular
apoptosis (14,15). The miR-200 family is among the most
downregulated tumor suppressive miRNAs in cancer (1). ETS proto-oncogene 1 (ETS1) is one
target of miR-200, and the loss of the miR-200-mediated repression
of ETS1 results in angiogenic responses in cancer cells (16).
miR-23a-5p has been found to be dysregulated in
several types of cancer (7). As a
functional downstream target of cAMP responsive element binding
protein 1, miR-23a-5p represses the tumor suppressor gene
phosphatase and tensin homolog and promotes cell growth and
survival in glioma (17). The
overexpression of miR-23a-5p in pancreatic ductal adenocarcinoma is
involved in increasing the transformation of epithelial mesenchymal
transition-like cell shape and its integration into mesothelial
monolayers through altering the expression of E-cadherin, β-catenin
and Wnt-related genes (18).
Interleukin 6 receptor (IL6R), an evolutionarily conserved
antiproliferative protein, has been confirmed as a direct target
gene for miR-23a in gastric adenocarcinoma (19).
To the best of our knowledge, the present study is
the first to confirm the upregulation of miR-23a-5p in bladder
cancer using RT-qPCR analysis, and to correlate miR-23a-5p with the
development of bladder cancer. In the present study, the relative
expression of miR-23a-5p was quantified in 44 paired bladder cancer
tissues and adjacent normal tissues. The data suggested that
miR-23a-5p was significantly upregulated in the bladder cancer
tissues and that miR-23a-5p may have a functional role in bladder
cancer. To further investigate the role of miR-23a-5p in bladder
cancer, a wound-healing assay, MTT assays and flow cytometric
analysis were performed in bladder cancer-related cells. Cell
proliferation, the suppression of migration and the induction of
apoptosis were observed in the miR-23a-5p inhibitor-transfected T24
and SW780 cells. The resulting data suggested that oncogenic
miR-23a-5p may have a fundamental role in the tumorigenesis and
tumor progression of bladder cancer.
In conclusion, the present study indicated that
miR-23a-5p functions as an oncogene in bladder cancer by affecting
cell proliferation, migration and apoptosis. Further investigations
are required to predict and confirm the upstream and downstream
genes, understand the molecular mechanisms, and examine the
clinical application of miR-23a-5p in bladder cancer.
Acknowledgements
The present study was supported by the Science and
Technology Development Fund Project of Shenzhen (grant no.
JCYJ20150403091443329) and the Science and Technology Development
Fund Project of Yangzhou (grant no. YZ2016130).
References
|
1
|
Kamat AM, Hahn NM, Efstathiou JA, Lerner
SP, Malmström PU, Choi W and Kassouf W: Bladder cancer. Lancet.
23:30512–30518. 2016.
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2016. CA Cancer J Clin. 66:7–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
IARC: Cancer incidence and mortality
worldwide. http://globocan.iarc.fr2016.
|
|
4
|
Antoni S, Ferlay J, Soerjomataram I, Znaor
A, Jemal A and Bray F: Bladder cancer incidence and mortality: A
global overview and recent trends. Eur Urol. 71:96–108. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Babjuk M: Trends in bladder cancer
incidence and mortality: Success or disappointment? Eur Urol.
71:106–110. 2016.
|
|
6
|
Krol J, Loedige I and Filipowicz W: The
widespread regulation of microRNA biogenesis, function and decay.
Nat Rev Genet. 11:597–610. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chhabra R, Dubey R and Saini N:
Cooperative and individualistic functions of the microRNAs in the
miR-23a~27a~24-2 cluster and its implication in human diseases. Mol
Cancer. 9:2322010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gottardo F, Liu CG, Ferracin M, Calin GA,
Fassan M, Bassi P, Sevignani C, Byrne D, Negrini M, Pagano F, et
al: Micro-RNA profiling in kidney and bladder cancers. Urol Oncol.
25:387–392. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Han Y, Chen J, Zhao X, Liang C, Wang Y,
Sun L, Jiang Z, Zhang Z, Yang R, Chen J, et al: MicroRNA expression
signatures of bladder cancer revealed by deep sequencing. PLoS One.
6:e182862011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li Y, Chen D, Jin L, Liu J, Su Z, Li Y,
Gui Y and Lai Y: MicroRNA-20b-5p functions as a tumor suppressor in
renal cell carcinoma by regulating cellular proliferation,
migration and apoptosis. Mol Med Rep. 13:1895–1901. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rupaimoole R, Calin GA, Lopez-Berestein G
and Sood AK: miRNA deregulation in cancer cells and the tumor
microenvironment. Cancer Discov. 6:235–246. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Pickering MT, Stadler BM and Kowalik TF:
miR-17 and miR-20a temper an E2F1-induced G1 checkpoint to regulate
cell cycle progression. Oncogene. 28:140–145. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Asangani IA, Rasheed SA, Nikolova DA,
Leupold JH, Colburn NH, Post S and Allgayer H: MicroRNA-21 (miR-21)
post-transcriptionally downregulates tumor suppressor Pdcd4 and
stimulates invasion, intravasation and metastasis in colorectal
cancer. Oncogene. 27:2128–2136. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Frankel LB, Christoffersen NR, Jacobsen A,
Lindow M, Krogh A and Lund AH: Programmed cell death 4 (PDCD4) is
an important functional target of the microRNA miR-21 in breast
cancer cells. J Biol Chem. 283:1026–1033. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chan YC, Khanna S, Roy S and Sen CK:
miR-200b targets Ets-1 and is down-regulated by hypoxia to induce
angiogenic response of endothelial cells. J Biol Chem.
286:2047–2056. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Tan X, Wang S, Zhu L, Wu C, Yin B, Zhao J,
Yuan J, Qiang B and Peng X: cAMP response element-binding protein
promotes gliomagenesis by modulating the expression of oncogenic
microRNA-23a. Proc Natl Acad Sci USA. 109:15805–15810. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Listing H, Mardin WA, Wohlfromm S, Mees ST
and Haier J: MiR-23a/−24-induced gene silencing results in
mesothelial cell integration of pancreatic cancer. Br J Cancer.
112:131–139. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhu LH, Liu T, Tang H, Tian RQ, Su C, Liu
M and Li X: MicroRNA-23a promotes the growth of gastric
adenocarcinoma cell line MGC803 and downregulates interleukin-6
receptor. Febs J. 277:3726–3734. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Edge SB, Byrd DR, Compton CC, Fritz AG,
Greene FL and Trotti A: AJCC Cancer Staging Handbook. 7th edition.
Springer; New York, NY: pp. 479–489. 2009
|