Introduction
Gastric cancer (GC) is the second leading cause of
cancer-related deaths and is a common malignant tumor of the
digestive system, which usually presents with no specific symptoms
(1–3). It has been reported that the risk of
developing GC increases with age. The majority (>60%) of
patients with GC are elderly patients, who are >65 years old
(4). Although the incidence of GC
is decreasing worldwide, this cancer still contributes a high
morbidity and is a tremendous burden to oncology care (5). The mortality rate has been estimated
to be ~70%, while the morbidity rate after surgery declines to 46%
(3). Currently, improvements in
chemoradiotherapy and surgical techniques, as well as novel
molecular targeting therapies have been developed, but
unfortunately, the majority of cases are diagnosed at advanced
stages, and the long-term survival rate and prognosis remain
unsatisfactory in China (6,7). Owing to the complex molecular pathways
involved in the pathogenesis of GC, the underlying mechanisms of
action that are implicated in the tumorigenesis and progression of
GC remain unknown.
Circular RNAs (circRNAs) are a group of special
non-coding RNAs, which lack 5′caps or 3′oly-A tails (8). Accumulating evidence has demonstrated
that circRNAs play an important role in tumor progression and
cellular functions as they are involved in transcriptional and
post-transcriptional regulation (9–11).
hsa-circ-0072309 has been reported to expressed at lower levels in
breast cancer tissue compared with adjacent normal tissue, and the
overexpression of hsa_circ_0072309 significantly suppresses the
proliferative, migratory and invasive capabilities of breast cancer
cells in vitro (12).
Additionally, the expression of hsa_circ_0072309 has also been
reported to be upregulated in kidney cancer cells, having an
anti-tumorigenic role by blocking the PI3K/AKT and mTOR signaling
pathways (13). However, the role
of circ_0072309 in the tumorigenesis and progression of GC remains
unclear. The aim of the present study was to explore the effect of
circ_0072309 on proliferation, invasion and migration of GC cells
and to investigate the underlying mechanisms of action.
In the present study, GC lines were employed to
investigate the role of circ_0072309 in GC progression. It was
demonstrated that circ_0072309 expression in human GC cell lines
was lower than that in normal gastric cells, and overexpression of
circ_0072309 led to inhibition of the proliferation, migration and
invasion of GC cells. As such, hsa_circ_0072309 may serve as a
novel therapeutic target for GC treatment.
Materials and methods
Cell culture and transfection
Human GC cell lines including AGS and MKN-45 cells,
as well as the normal gastric epithelial cell line, GES-1, were
obtained from the American Type Culture Collection. The cell lines
were cultured in RPMI-1640 (Invitrogen; Thermo Fisher Scientific,
Inc.) or DMEM (Gibco; Thermo Fisher Scientific, Inc.) supplemented
with 100 U/ml penicillin/streptomycin and 10% FBS (Gibco; Thermo
Fisher Scientific, Inc.) and incubated at 37°C in a humidified
incubator containing 5% CO2. AGS cells were pretreated
with PPARγ agonist (pioglitazone, 20 µM) or PPARγ antagonist
(GW9662, 2 µM) for 6 h at 37°C.
The coding sequence of hsa_circ_0072309 was cloned
into the PLCDH-cir vector (Guangzhou RiboBio Co., Ltd.) for
hsa_circ_0072309 overexpression. The 100 nM overexpression vector
(Oe)-circ_0072309 or an empty vector, used as negative controls,
(Vector Laboratories, Inc.; Maravai Life Sciences) were transfected
into the AGS cells (2×106/well) using
Lipofectamine® 3000 reagent (Invitrogen; Thermo Fisher
Scientific, Inc.), following the manufacturer's instructions. After
48 h transfection, the AGS cells were used for further experiments
and reverse transcription-quantitative (RT-q) PCR was performed to
confirm the transfection efficiency.
RT-qPCR
According to the manufacturer's instructions,
TRIzol® reagent (Thermo Fisher Scientific, Inc.) and
PrimeScript RT Reagent kit (Takara Bio, Inc.) were employed for RNA
isolation and cDNA synthesis. RT-qPCR was performed using SYBR
Green PCR kits (Roche Diagnostics), according to the manufacturer's
instructions, using a StepOnePlus™ Real-time PCR System (Applied
Biosystems; Thermo Fisher Scientific, Inc.). The qPCR thermocycling
conditions were: 95°C for 30 sec followed by 40 cycles at 95°C for
5 sec and 60°C for 30 sec and the reaction volume was 25 µl. The
gene expression levels were calculated using the 2−ΔΔCq
method (14) and normalized to the
expression levels of GAPDH. The primer sequences used were as
follows: hsa_circ_0072309 forward, 5′-CTCAACCTCTACATTATACCTAA-3′
and reverse, 5′-CCTAGGGACCCTGGTATGGATC-3′; PPARγ forward,
5′-AAAGACAACGGACAAATCAC-3′ and reverse,
5′-GGGATATTTTTGGCATACTCT-3′; PTEN forward,
5′-CTTACAGTTGGGCCCTGTACCATCC-3′ and reverse,
5′-TTTGATGCTGCCGGTAAACTCCACT-3′; PI3K forward,
5′-GCCCAGGCTTACTACAGAC-3′ and reverse, 5′-AAGTAGGGAGGCATCTCG-3′;
AKT forward, 5′-GGAGTGTGTGGACAGTGAAC-3′ and reverse,
5′-CCCACAGTAGAAACATCCTCCC-3′; mTOR forward,
5′-AGTGGGAAGATCCTGCACATT-3′ and reverse,
5′-TGGAAACTTCTCTCGGGTCAT-3′; and β-actin forward,
5′-AGCGAGCATCCCCCAAAGTT-3′ and reverse,
5′-GGGCACGAAGGCTCATCATT-3′.
Cell viability assessment
Cell Counting Kit-8 (CCK-8) assays were performed to
quantify the cell viability of AGS cells transfected with or
without Oe-circ_0072309. AGS cells were seeded at a density of
2×103 cells/well in 96-well plates. Subsequently, AGS
cells were treated with CCK-8 reagent (10 µl per well, Dojindo
Molecular Technologies, Inc.) for 0, 24, 48 or 72 h. After
incubation for 1 h, the optical density (OD) of each well was
measured at 450 nm using a microplate reader (Molecular Devices,
LLC).
Colony formation assay
After transfection with Oe-circ_0072309 or the
control vector, AGS cells (1×103/well)were seeded onto
35 mm culture plates. The cells were cultured for 2 weeks at 37°C.
After cell colonies were formed in culture plates, the cells were
fixed with 4% paraformaldehyde for 15 min at room temperature and
stained with 0.1% crystal violet solution for 0.5 h at room
temperature. Finally, the colonies with diameters >0.5 mm were
imaged and counted using a digital camera (Nikon Corporation).
Migration and invasion assays
Following transfection, cell migratory capabilities
were evaluated using wound-healing assays. AGS cells
(5×105 cells/well) were seeded in a six-well plate and
cultured with RPMI-1640 medium for 24 h. When cells reached ~80%
confluency, a linear wound was created by scraping the monolayers
with a 200 µl sterile pipette tip and the cells were washed twice
with PBS to remove floating cells and debris. The wound monolayers
of AGS cells were cultured in serum-free RPMI-1640 medium. The
wound was captured and measured using a fluorescence microscope
(Leica Microsystems GmbH) at ×100 magnification, from five random
fields at 0 and 48 h. The recovered wound area (%) at the indicated
time point (48 h) was calculated according to the following
formula: (wound width at 0 h) - (wound width at 48 h)/wound width
at 0 h.
In addition, the invasive ability of cells was
analyzed using a Transwell chamber assay. Briefly, cells
(1×105 cells/well) were suspended in RPMI-1640 medium
containing 10% FBS and added to the upper chamber. Matrigel mix was
coated onto the underside of the upper chamber at 37°C for 4 h.
Culture medium (600 µl), supplemented with 10% FBS, was added to
the lower chamber. The non-invaded cells on the upper surface were
removed after 24 h of incubation at 37°C, while the cells on the
bottom of the membrane were fixed with formaldehyde solution for 20
min at 37°C and subsequently stained with 0.1% crystal for 30 min
at room temperature. Finally, the cells were imaged on randomly
selected fields (magnification, ×200) using an Olympus microscope
(Olympus Corporation). The invasion rate was calculated according
to the following formula: Number of cells in tested group/the
number of cells in control group.
Western blotting
Proteins were extracted from the transfected cells
using RIPA lysis buffer containing protease inhibitors (Beyotime
Institute of Biotechnology). The concentration of protein in the
cell lysates was quantified using a BCA assay kit (Bio-Rad
Laboratories, Inc.). Total protein (25 µg) was separated via
SDS-PAGE on 10-12% gels (Beyotime Institute of Biotechnology), and
subsequently transferred onto PVDF membranes. Following blocking
with 5% bovine serum albumin (Sigma-Aldrich; Merck KGaA) for 2 h at
room temperature, membranes were incubated with the appropriate
primary antibodies overnight at 4°C. The primary antibodies used in
the present study were as follows: Anti-matrix metalloproteinase
(MMP)7 (1:1,000; cat. no. ab207299; Abcam), anti-MMP9 (1:1,000;
cat. no. ab76003; Abcam), anti-PPARγ (1:1,000; cat. no. 2430; Cell
Signaling Technology, Inc.), anti-PTEN (1:1,000; cat. no. 9552;
Cell Signaling Technology, Inc.), anti-phosphorylated (p)-PI3K
(1:1,000; cat. no. 4228; Cell Signaling Technology, Inc.),
anti-PI3K (1:500; cat. no. 4292; Cell Signaling Technology, Inc.),
anti-p-AKT (1:1,000; cat. no. 9271; Cell Signaling Technology,
Inc.), anti-AKT (1:1,000; cat. no. 9272; Cell Signaling Technology,
Inc.), anti-p-mTOR (1:500; cat. no. 2974; Cell Signaling
Technology, Inc.) and anti-mTOR (1:1,000; cat. no. 2972; Cell
Signaling Technology, Inc.). After washing, the membranes were
incubated with secondary HRP-conjugated antibodies (1:10,000; cat.
no. A-11046; Pierce; Thermo Fisher Scientific, Inc.) at 25°C for 2
h, which were then visualized and captured by ECL chemiluminescence
(Pierce; Thermo Fisher Scientific, Inc.). The protein band
intensities were semi-quantified using ImageJ software (v1.6;
National Institutes of Health) and normalized to GAPDH (1:1,000;
cat. no. 8884; Cell Signaling Technology, Inc.) expression
levels.
Statistical analysis
Data are expressed as the mean ± SD and were
analyzed using SPSS version 10.0.2 software (SPSS, Inc.) and
GraphPad Prism 5.0 (GraphPad Software, Inc.). All experiments were
performed independently at least three times. ANOVA followed by
Bonferroni's post hoc test and Student's t-tests were performed to
determine the differences in the means between the various
treatment groups. P<0.05 was considered to indicate a
statistically significant difference.
Results
hsa_circ_0072309 is downregulated in
GC cell lines
To identify the role of hsa_circ_0072309 in GC
progression, the expression levels of hsa_circ_0072309 were
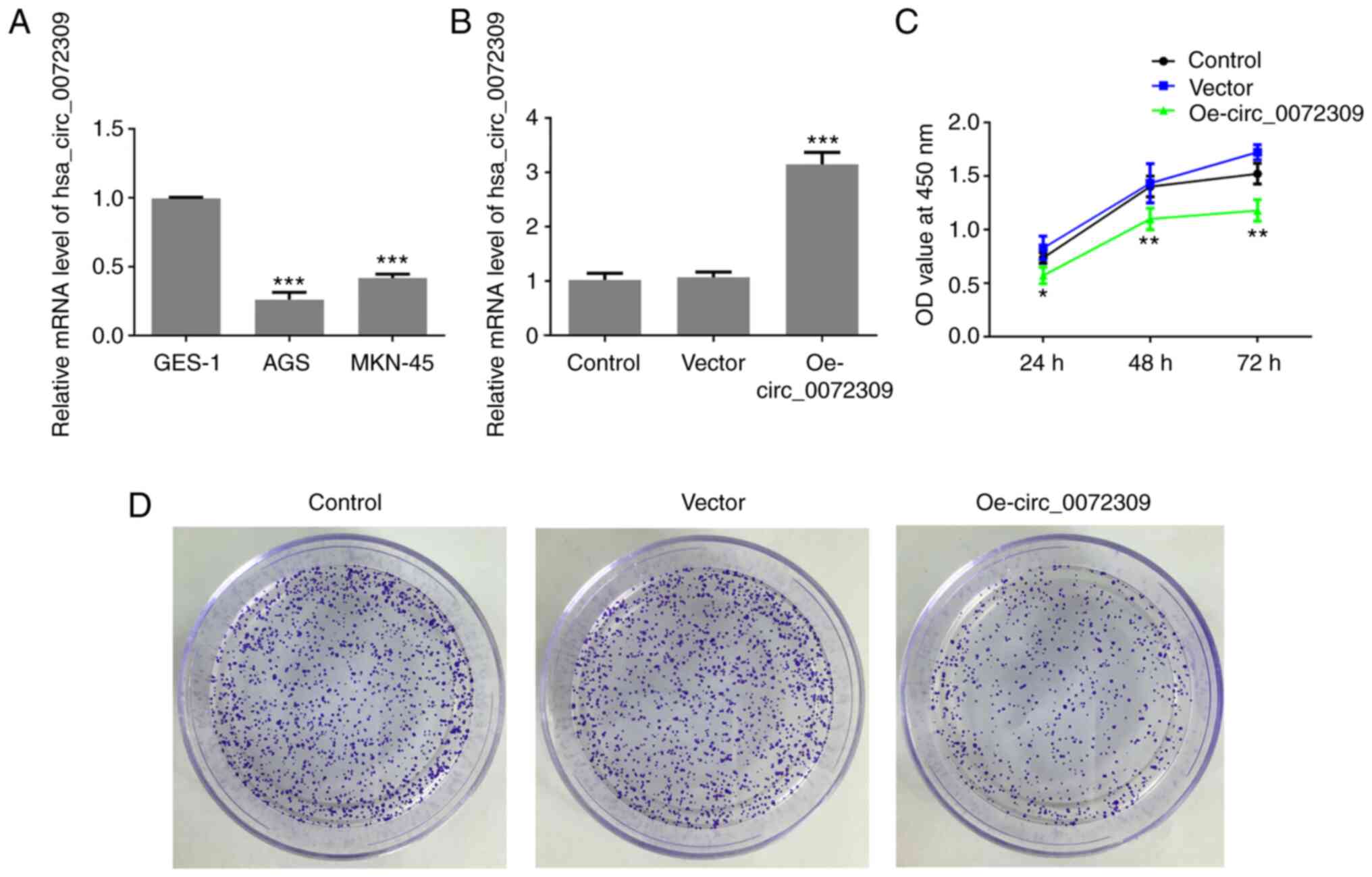
analyzed. According to the RT-qPCR results, hsa_circ_0072309
exhibited a lower expression level in human GC cell lines (AGS and
MKN-45 cells) compared with normal gastric epithelial GES-1 cells
(Fig. 1A). Due to the lowest
expression of hsa_circ_0072309, AGS cells were used for the
following experiments. These results suggested that
hsa_circ_0072309 played a role in GC progression.
hsa_circ_0072309 overexpression
inhibits proliferation of GC cells
To investigate the function of hsa_circ_0072309 in
GC tumorigenesis, Oe-circ_0072309 plasmids were designed to induce
hsa_circ_0072309 overexpression. The RT-qPCR results demonstrated
that Oe-circ_0072309 plasmids significantly upregulated the
expression levels of hsa_circ_0072309, suggesting that
Oe-circ_0072309 plasmids were successfully produced and transfected
into the AGS cells (Fig. 1B). CCK-8
assay results found that hsa_circ_0072309 overexpression led to a
greater reduction in the cell viability of AGS cells in comparison
with control cells (Fig. 1C).
Furthermore, colony formation assays were performed to confirm the
effects of hsa_circ_0072309 on the proliferative ability of AGS
cells. The results indicated that AGS cells transfected with
Oe-circ_0072309 had a reduced proliferative ability compared with
the control group (Fig. 1D). These
data suggested that overexpression of hsa_circ_0072309 reduced the
proliferative ability of AGS cells.
hsa_circ_0072309 overexpression
inhibits the migratory and invasive capabilities of GC cells
To further investigate the function of
hsa_circ_0072309 in GC tumorigenesis, migration and invasion assays
were performed to identify the migratory and invasive abilities of
AGS cells after transfection of Oe-circ_0072309 or control vectors.
As shown in Fig. 2A and B, the
migratory ability of AGC cells was significantly suppressed
following hsa_circ_0072309 overexpression in comparison with the
control group, as demonstrated by wound-healing assays.
Additionally, Transwell chamber assays showed that hsa_circ_0072309
overexpression caused a reduction in the cell invasion rate in the
Oe-circ_0072309 group, compared with the control group (Fig. 2C and D). The expression levels of
MMP7 and MMP9, two molecules involved in tumor invasion and
metastasis, were determined using western blotting. As shown in
Fig. 2E, the expression levels of
MMP7 and MMP9 were significantly downregulated in the AGS cells
transfected with Oe-circ_0072309 compared with the control cells.
These data indicated that raised hsa_circ_0072309 levels obstructed
the migration and invasion of AGS cells.
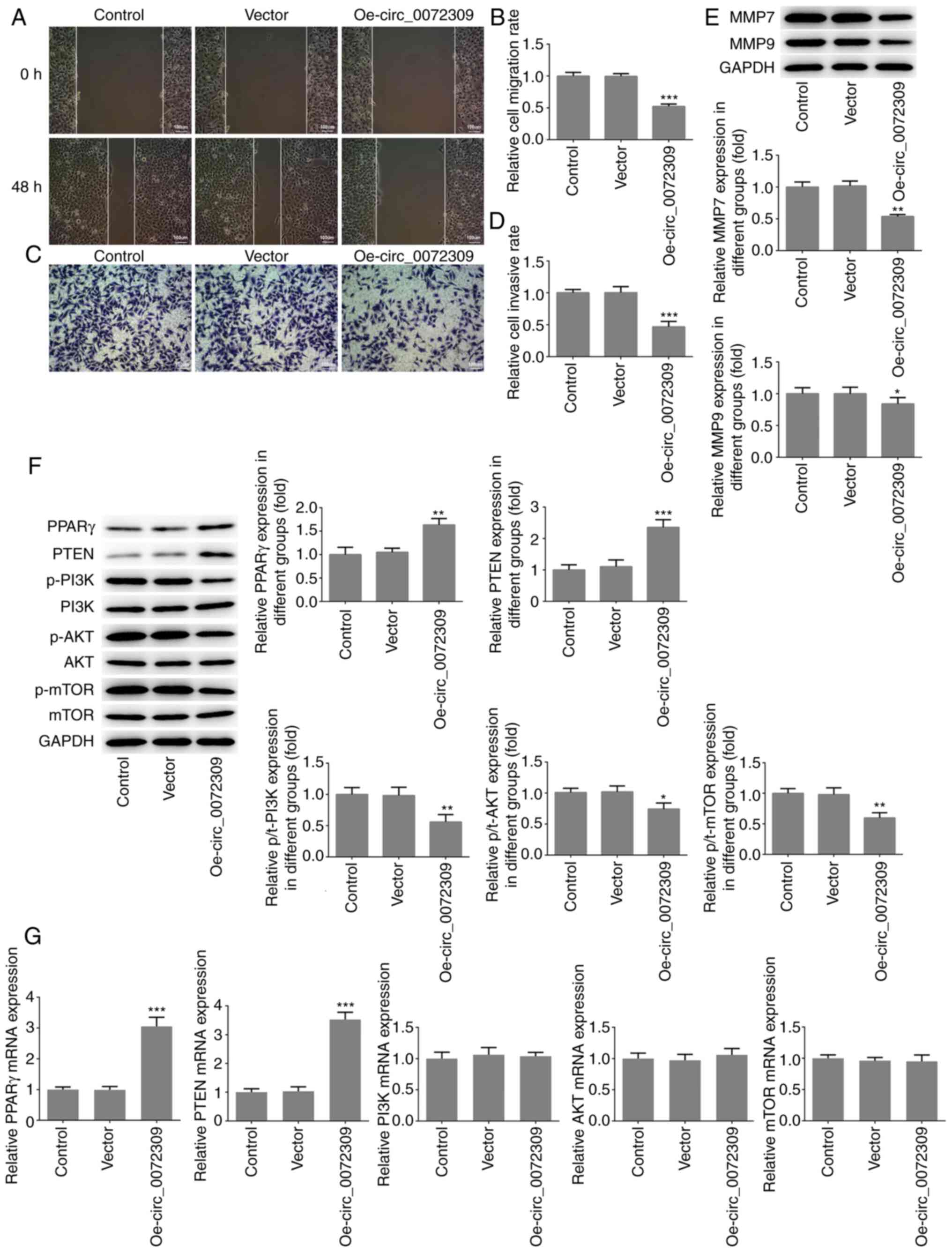 | Figure 2.hsa_circ_0072309 overexpression
induces the inhibition of migration and invasion of gastric cancer
cells. (A) The migratory ability of AGS cells was analyzed using
wound healing assays and (B) quantified. Scale bar, 100 µm. (C) The
invasive ability of AGS cells was evaluated using Transwell assays
and (D) quantified. Scale bar, 100 µm. (E) The protein expression
levels of MMP7 and MMP9, which are related to cell invasion were
detected using western blotting. (F) The expression levels of
proteins including PPARγ, PTEN, p/t-PI3K, p/t-AKT and p/t-mTOR were
detected using western blotting. (G) The mRNA expression levels of
PPARγ, PTEN, PI3K, AKT and mTOR were detected by reverse
transcription-quantitative PCR. Error bars represent the mean ± SEM
from three independent experiments. *P<0.05, **P<0.01,
***P<0.001 vs. control. circ, circular RNA; Oe, overexpression
vector; PPARγ, peroxisome proliferator-activated receptor γ; MMP,
matrix metalloproteinase; p-, phosphorylated. |
Effects of hsa_circ_0072309
overexpression on PPAR γ/PTEN and PI3K/AKT signaling
To further investigate the underlying molecular
mechanisms of hsa_circ_0072309 in GC pathogenesis, western blotting
was performed to analyze the expression levels of proteins involved
in the PPARγ/PTEN and PI3K/AKT signaling pathways. The results
revealed that hsa_circ_0072309 overexpression induced increased
protein and mRNA expression levels of PPARγ and PTEN, as well as a
reduction in p-PI3K, p-AKT and p-mTOR levels, with no impact on the
expression levels of total PI3K, AKT and mTOR (Fig. 2F and G).
To confirm the functional role of hsa_circ_0072309
PPARγ antagonist (GW9662). AGC cells were treated with pioglitazone
to upregulate PPARγ, whereas GW9662 treatment induced a reduction
in PPARγ expression. As shown in Fig.
3A and B, pioglitazone treatment increased the protein and mRNA
expression of PPARγ and PTEN, while GW9662 treatment induced the
downregulation of PPARγ and PTEN expression. Of note, GW9662
treatment blocked the inhibitory effects of hsa_circ_0072309
overexpression on p-PI3K, p-AKT and p-mTOR expression levels,
whereas pioglitazone treatment had further inhibited their
expression. The combination of pioglitazone and GW9662 had no
influence on the expression levels of the aforementioned proteins.
There were no significant changes in total protein expression and
mRNA levels of PI3K, AKT and mTOR. These findings indicated that
hsa_circ_0072309 exerted a suppressive effect on the PI3K/AKT
signaling pathway via the activation of the PPARγ/PTEN signaling
pathway.
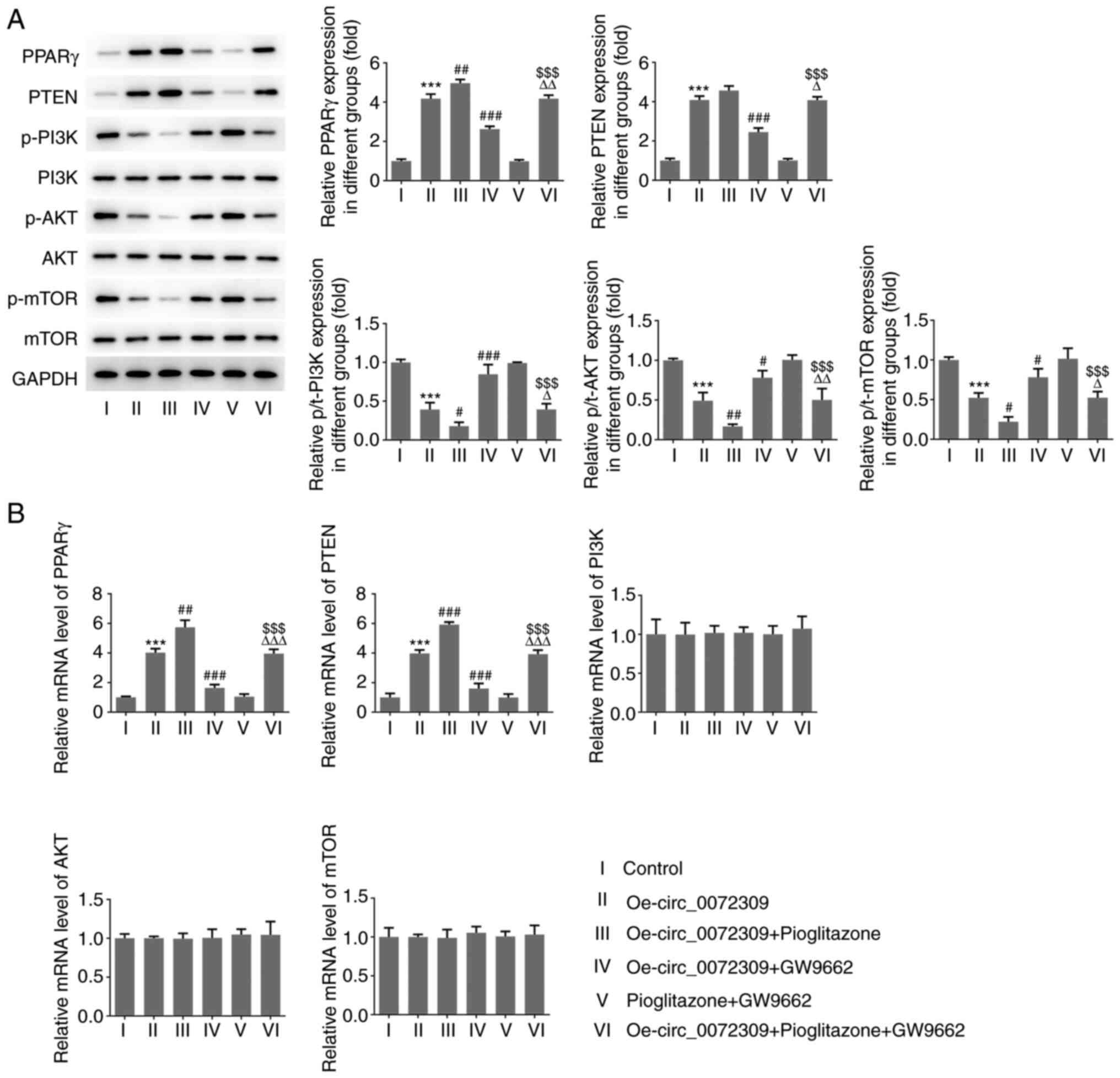 | Figure 3.Effects of hsa_circ_0072309
overexpression on PPARγ/PTEN and PI3K/AKT signaling. (A) The
expression levels of proteins including PPARγ, PTEN, p/t-PI3K,
p/t-AKT and p/t-mTOR were determined using western blotting. (B)
The mRNA expression levels of PPARγ, PTEN, PI3K, AKT and mTOR were
detected by reverse transcription-quantitative PCR. Error bars
represent the mean ± SEM from three independent experiments.
***P<0.001 vs. control; #P<0.05,
##P<0.01, ###P<0.001 vs.
Oe-circ_0072309; $$$P<0.001 vs. pioglitazone +
GW9662; ΔP<0.05, ΔΔP<0.01,
ΔΔΔP<0.001 vs. Oe-circ_0072309 + pioglitazone. circ,
circular RNA; Oe, overexpression vector; PPARγ, peroxisome
proliferator-activated receptor γ; p-, phosphorylated; t-,
total. |
hsa_circ_0072309 overexpression
inhibits the proliferation, migration and invasion of GC cells via
the PPARγ-dependent PTEN pathway
To further determine whether the PPARγ-dependent
PTEN pathway was involved in the function of hsa_circ_0072309 in GC
tumorigenesis, the proliferative, migratory and invasive
capabilities of GC cells were assessed under the treatment of
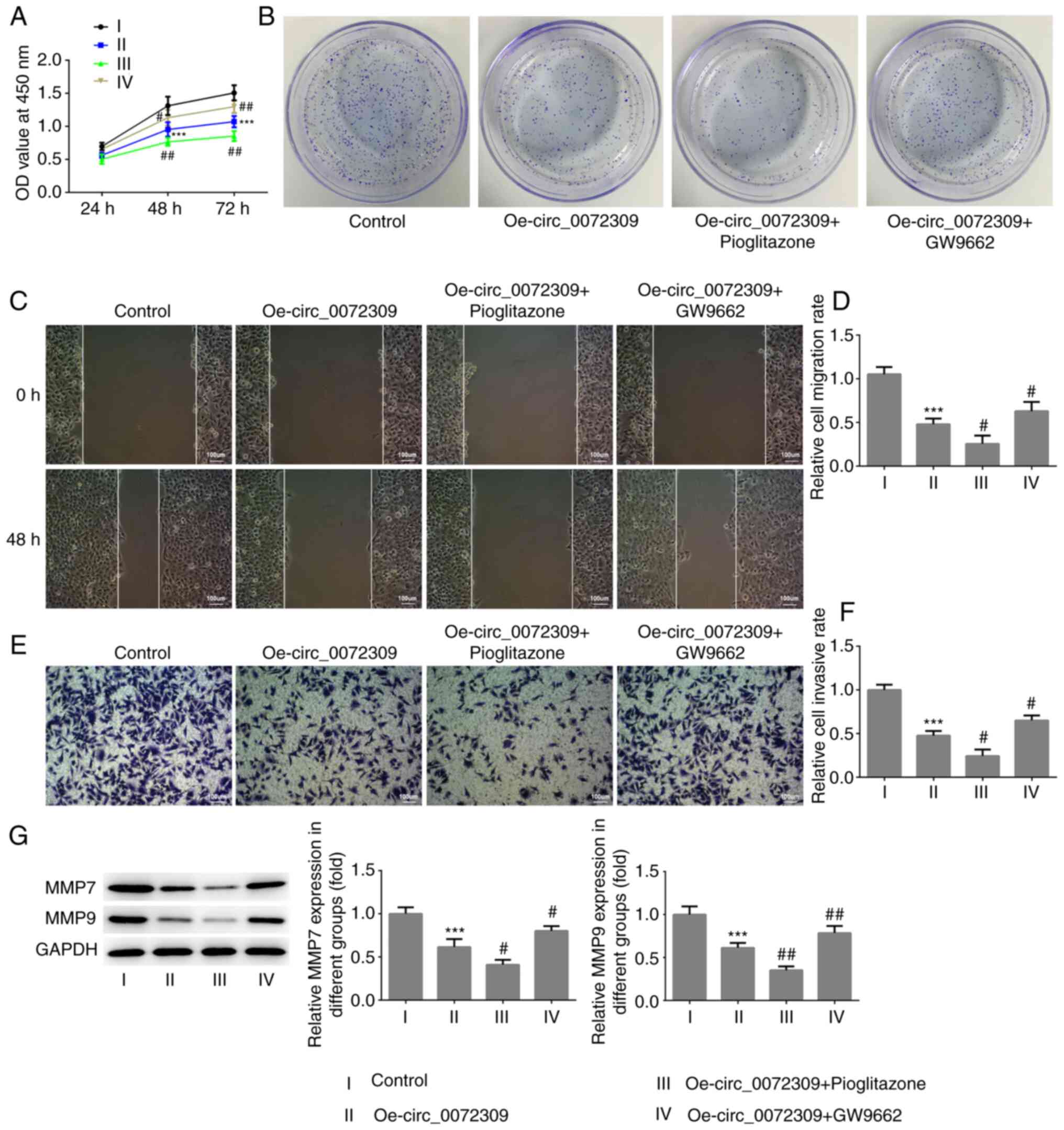
pioglitazone or GW9662. As presented in Fig. 4A-G, treatment with pioglitazone,
strengthened the effect of hsa_circ_0072309 overexpression on cell
viability, and the proliferative, migratory and invasive
capabilities of GC cells, whereas treatment with GW9662, abolished
these effects. The combination of pioglitazone and GW9662 had no
effect on the viability of GC cells (Fig. S1). These data showed that
hsa_circ_0072309 suppressed the proliferation and metastasis of GC
cells via the PPARγ-dependent PTEN pathway.
Discussion
A number of studies have suggested that circRNAs can
be found in multiple tissues and various circRNAs exert specific
roles during tumor progression, with distinctive expression
patterns (15–17). circRNAs may mediate tumor
progression by modulating the cell cycle and metastasis of cancer
cells via distinct mechanisms of action. A previous study reported
that hsa_circ_0072309 plays a tumor-suppressive role in breast
cancer (18). Moreover, Huang et
al (19) illustrated that
hsa_circ_0072309 expression is downregulated in intracranial
aneurysm tissue and in the peripheral blood. Yan et al
(12)also demonstrated that
hsa_circ_0072309 suppresses proliferation, migration and invasion
of breast cancer cells by sponging microRNA (miR)-492. Nonetheless,
the role of hsa_circ_0072309 in GC progression remains unclear. The
aim of the present study was to explore the role of
hsa_circ_0072309 in the pathogenesis of GC and its distinct
mechanism of action in GC cell proliferation, migration and
invasion.
In the present study, it was found that
hsa_circ_0072309 was poorly expressed in GC cells compared with
normal gastric epithelial cells. The overexpression of
hsa_circ_0072309, induced by Oe-circ_0072309 plasmids, produced a
suppressive role on GC cell proliferation, migration and invasion,
which was consistent with the findings of previous studies
investigating the role of hsa_circ_0072309 in other cancer types.
These findings suggested that hsa_circ_0072309 was associated with
GC progression through the regulation of cell proliferation,
migration and invasion. Accordingly, the upregulation of
hsa_circ_0072309 may be a potential target for cancer therapy in
the clinic.
To further investigate the underlying mechanisms of
action of hsa_circ_0072309 in GC progression, further experiments
were carried out to determine the extent of PPARγ/PTEN and PI3K/AKT
signaling following the overexpression of hsa_circ_0072309.
Aberrant activation of with the PPARγ agonist, pioglitazone. A
previous study found that miR-492 promotes the progression of
hepatic cancer through the inhibition of PTEN expression and
activation of the PI3K/AKT cascade (22). Hyun et al (23) reported that 4-O-Methylhonokiol, a
PPARγ activator, enhances PPARγ/PTEN signaling and induces
apoptosis by deactivating the PI3K/Akt signaling pathway in SiHa
cervical cancer cells. Balaglitazone reverses multidrug resistance
via the upregulation of PTEN in a PPARγ-dependent manner in human
myelogenous leukemia cells (21).
Moreover, PPARγ agonists exert tumor-suppressive effects on the
progression of bladder cancer through the inhibition of the
PI3K/AKT signaling pathway (24),
which is consistent with the results of the present study. In the
present study, pioglitazone induced PPARγ upregulation and elevated
the expression level of PTEN, playing a negative role on the
PI3K/AKT signaling cascade, whereas the PPARγ antagonist, GW9662,
led to a reduction in PTEN expression and had the opposite effect
on PI3K/AKT signaling, suggesting that PPARγ controls PTEN
expression. A previous study reported that PPARγ overexpression
caused the upregulation of PTEN expression (25), which is consistent with the present
results. Of note, PPARγ activation with pioglitazone following
hsa_circ_0072309 overexpression significantly inhibited
proliferation, migration and invasion of GC cells. In agreement,
the PPARγ inhibitor GW9662 significantly promoted the
proliferation, migration and invasion of GC cells transfected with
Oe-circ_0072309. Taken together, these data suggested that
hsa_circ_0072309 acted as a tumor-suppressive gene during GC
progression via the inactivation of PI3K/AKT signaling by
activating PPARγ/PTEN signaling. As such, the overproduction of
hsa_circ_0072309 may be a novel therapeutic strategy for the
treatment of GC.
In summary, the present study demonstrated that
hsa_circ_0072309 was downregulated in GC cell lines.
hsa_circ_0072309 overexpression led to a decrease in cell
proliferation, migration and invasion, and significantly inhibited
the phosphorylation of PI3K, AKT and mTOR. Moreover, PPARγ
activation by pioglitazone significantly inhibited proliferation,
migration and invasion of GC cells, whereas the PPARγ inhibitor
GW9662 significantly promoted cell proliferation, migration and
invasion. These findings showed that hsa_circ_0072309 inhibited
proliferation, invasion and migration of GC cells via the
inhibition of PI3K/AKT signaling by activating PPARγ/PTEN
signaling. hsa_circ_0072309 may prove to be an innovative target
for the clinical treatment of GC. As the present study only used
in vitro methods, the role of hsa_circ_0072309 in an animal
model with GC tumor and GC patient tumor samples needs to be
investigated in further studies.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
This work was supported by National Natural Science
Foundation of China (grant no. 81974375).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
XPG and MDQ designed the experiments and drafted the
manuscript. HH, XX, JF and LJ performed the experiments and
analyzed the data. YK and LG designed the experiments and reviewed
the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Mungan I, Dicle ÇB, Bektas S, Sari S,
Yamanyar S, Çavuş M, Turan S and Bostanci EB: Does the preoperative
platelet-to-lymphocyte ratio and neutrophil-to-lymphocyte ratio
predict morbidity after gastrectomy for gastric cancer? Mil Med
Res. 7:42020.PubMed/NCBI
|
|
4
|
Hamilton TD, Mahar AL, Haas B, Beyfuss K,
Law CHL, Karanicolas PJ, Coburn NG and Hallet J: The impact of
advanced age on short-term outcomes following gastric cancer
resection: An ACS-NSQIP analysis. Gastric Cancer. 21:710–719. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Balea AM, Cruce R, Schenker RA, Ionescu
AG, Streba L, Ciurea AM, Ghilusi MC, Pirici D and Vere CE:
Correlations between clinicopathological features and the
vegetative nervous system in gastric cancer. Curr Health Sci J.
45:351–357. 2019.PubMed/NCBI
|
|
6
|
Zong L, Abe M, Seto Y and Ji J: The
challenge of screening for early gastric cancer in China. Lancet.
388:26062016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yao J, Zhang H, Liu C, Chen S, Qian R and
Zhao K: MiR-450b-3p inhibited the proliferation of gastric cancer
via regulating KLF7. Cancer Cell Int. 20:472020. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liu J, Liu T, Wang X and He A: Circles
reshaping the RNA world: From waste to treasure. Mol Cancer.
16:582017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Huang HS, Huang XY, Yu HZ, Xue Y and Zhu
PL: Circular RNA circ-RELL1 regulates inflammatory response by
miR-6873-3p/MyD88/NF-κB axis in endothelial cells. Biochem Biophys
Res Commun. 30:512–519. 2020. View Article : Google Scholar
|
|
10
|
Meng S, Zhou H, Feng Z, Xu Z, Tang Y, Li P
and Wu M: CircRNA: Functions and properties of a novel potential
biomarker for cancer. Mol Cancer. 16:942017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang S, Zhang Y, Cai Q, Ma M, Jin LY, Weng
M, Zhou D, Tang Z, Wang JD and Quan Z: Circular RNA FOXP1 promotes
tumor progression and warburg effect in gallbladder cancer by
regulating PKLR expression. Mol Cancer. 18:1452019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yan L, Zheng M and Wang H: Circular RNA
hsa_circ_0072309 inhibits proliferation and invasion of breast
cancer cells via targeting miR-492. Cancer Manag Res. 11:1033–1041.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen T, Shao S, Li W, Liu Y and Cao Y: The
circular RNA hsa-circ-0072309 plays anti-tumour roles by sponging
miR-100 through the deactivation of PI3K/AKT and mTOR pathways in
the renal carcinoma cell lines. Artif Cells Nanomed Biotechnol.
47:3638–3648. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Sharifi A, Vahedi H, Honarvar MR, Amiriani
T, Nikniaz Z, Rad EY and Hosseinzadeh-Attar MJ: Vitamin D decreases
CD40L gene expression in ulcerative colitis patients: A randomized,
double-blinded, placebo-controlled trial. Turk J Gastroenterol.
31:99–104. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wan B, Liu B and Lv C: Progress of
research into circular RNAs in urinary neoplasms. PeerJ.
8:e86662020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Su Y, Feng W, Shi J, Chen L, Huang J and
Lin T: circRIP2 accelerates bladder cancer progression via
miR-1305/Tgf-β2/smad3 pathway. Mol Cancer. 19:232020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yao J, Xu G, Zhu L and Zheng H:
circGFRA1enhances NSCLC progression by sponging miR-188-3p. Onco
Targets Ther. 13:549–558. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li Z, Ruan Y, Zhang H, Shen Y, Li T and
Xiao B: Tumor-Suppressive circular RNAs: Mechanisms underlying
their suppression of tumor occurrence and use as therapeutic
targets. Cancer Sci. 110:3630–3638. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Huang Q, Huang QY, Sun Y and Wu S:
High-Throughput data reveals novel circular RNAs via competitive
endogenous RNA networks associated with human intracranial
aneurysms. Med Sci Monit. 25:4819–4830. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
de Biase D, Visani M, Pession A and
Tallini G: Molecular diagnosis of carcinomas of the thyroid gland.
Front Biosci (Elite Ed). 6:1–14. 2014.PubMed/NCBI
|
|
21
|
Yousefi B, Azimi A, Majidinia M,
Shafiei-Irannejad V, Badalzadeh R, Baradaran B, Zarghami N and
Samadi N: Balaglitazone reverses P-glycoprotein-mediated multidrug
resistance via upregulation of PTEN in a PPARgamma-dependent manner
in leukemia cells. Tumour Biol. 39:10104283177165012017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Jiang J, Zhang Y, Yu C, Li Z, Pan Y and
Sun C: MicroRNA-492 expression promotes the progression of hepatic
cancer by targeting PTEN. Cancer Cell Int. 14:952014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hyun S, Kim MS, Song YS, Bak Y, Ham SY,
Lee DH, Hong J and Yoon DY: Peroxisome proliferator-activated
receptor-gamma agonist 4-O-methylhonokiol induces apoptosis by
triggering the intrinsic apoptosis pathway and inhibiting the
PI3K/Akt survival pathway in SiHa human cervical cancer cells. J
Microbiol Biotechnol. 25:334–342. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lv S, Wang W, Wang H, Zhu Y and Lei C:
PPARγ activation serves as therapeutic strategy against bladder
cancer via inhibiting PI3K-Akt signaling pathway. BMC Cancer.
19:2042019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Esmaeili S, Safaroghli-Azar A,
Pourbagheri-Sigaroodi A, Salari S, Gharehbaghian A, Hamidpour M and
Bashash D: Stimulation of peroxisome proliferator-activated
receptor-gamma (PPARγ) using pioglitazone decreases the survival of
acute promyelocytic leukemia cells through up-regulation of PTEN
expression. Anticancer Agents Med Chem. 21:108–119. 2021.
View Article : Google Scholar : PubMed/NCBI
|