Introduction
With the improvement of living environment,
individual lifestyle is becoming increasingly westernized (1). Subsequently, the incidence and
mortality rate from colorectal cancer (CRC) has also significantly
increased. For instance, in China, CRC is included in the top five
causes of cancer mortality (2).
Similarly, CRC has the third-highest incidence rates among all
types of cancer and it also ranks fifth in mortality caused by
cancer in the US, in male and females (3). Thus, CRC might be a major threat to
human health worldwide. One main reason for such high morbidity and
mortality of CRC is lack of effective early diagnosis (4). Therefore, it has great significance
for seeking new early tumor diagnostic markers.
DnaJ Heat Shock Protein Family (Hsp40) Member C2
(DNAJC2), also known as zuotin-related factor 1, was a recently
identified epigenetic regulator of gene transcription in stem cells
and cancer (5). DNAJC2 is located
in the key region of 7q22-31.1 and high copy amplification of the
gene has been reported to be involved in the progression of
multiple types of cancer, such as prostate cancer, germ cell,
glioblastoma, head and neck squamous cell carcinoma and gastric
cancer (5–7). These data suggested that DNAJC2 is
significantly related to proliferation and migration of tumor.
Little is known regarding the role of DNAJC2 in CRC,
or its clinical and prognostic significance in patients with CRC.
Therefore, the present study aimed to explore the expression level,
biological function and potential mechanism of DNAJC2 in CRC.
Materials and methods
CRC patient specimens
A total of 84 pairs of human CRC tissues and normal
colon (5 cm away from the tumor) or rectum mucosa adjacent (3 cm
away from the tumor) to the tumor were acquired from patients after
they underwent surgeries between September 2014 and April 2018 in
The First Affiliated Hospital of Gannan Medical University.
Clinical information was collected and presented in Fig. 1. The tissue samples were frozen in
liquid nitrogen and preserved them in a refrigerator with a
temperature of −80°C as soon as possible until RNA extraction was
performed. All of the enrolled patients or their relatives signed
informed consent forms. The National Comprehensive Cancer Network
(https://www.nccn.org/2015.2) was chosen
for the identification of TNM. Patients cured by radiotherapy and
chemotherapy were excluded from the present study. The present
study was approved by the Ethics Committee of The First Affiliated
Hospital of Gannan Medical University (approval no. IRB20170082)
and was conducted in accordance with the Helsinki Declaration. The
expression level of DNAJC2 in Oncomine database was obtained at
https://www.oncomine.org.
Cell lines and culture conditions
CRC cell lines (LoVo, HCT-116, Caco-2, DLD-1 and
HT-29), normal epithelial colon cells (NCM460) and 293T cells were
purchased from the Cell Bank of the Chinese Academy of Sciences
(Shanghai, China). Caco-2 were cultured in DMEM (Gibco; Thermo
Fisher Scientific, Inc.); HCT-116 and HT-29 were cultured in
McCoy's 5A medium (Invitrogen; Thermo Fisher Scientific, Inc.),
DLD-1 was cultured in 1640 medium (Gibco; Thermo Fisher Scientific,
Inc.), LoVo was cultured in F-12K medium (Gibco; Thermo Fisher
Scientific, Inc.). All the media were supplemented with 10% FBS
(WISENT Inc.) and 100 U/ml penicillin and 100 µg/ml streptomycin
(WISENT Inc.). All the cells were cultured in a cell incubator with
humid environment of 5% CO2 and 37°C and all cell lines
were authenticated using STR profiling.
Bioinformatical Analysis
In order to explore the detailed mechanism of DNAJC2
on CRC cell lines, online databases including TargetScan
V3.1(http://www.targetscan.org/mamm_31/) and miRWalk 2021
(http://mirwalk.umm.uni-heidelberg.de/) were used to
predict the potential miRNA for DNAJC2. The 3′untranslated region
(UTR) was used as input to predict potential miRNA that may bind
with DNAJC2.
RNA extraction and reverse
transcription-quantitative (RT-q) PCR
Total RNA, miRNA included, was extracted from
tissues and cells by TRIzol reagent (Thermo Fisher Scientific,
Inc.) in accordance with manufacturer's protocol and cDNA was
produced by means of PrimeScript RT reagent kit (Takara
Biotechnology Co., Ltd.). The RT kit was used according to the
manufacturer's protocol. The RT-qPCR experiment was conducted
through a 10 µl reaction system which consists of 1 µl cDNA, 1 µl
primers, 4 µl dH2O and 4 µl SYBR via SYBR-Green PCR kit
(Roche Diagnostics). StepOne Plus Real-time PCR system (Applied
Biosystems; Thermo Fisher Scientific, Inc.) was used to performed
final reaction. Thermocycling conditions was set as follows:
Initial denaturation at 95°C for 3 min, followed by 35 cycles at
95°C for 15 sec, 60°C for 30 sec, and 72°C for 30 sec, followed by
72°C for 10 min. The method used to standardize the RT-qPCR data
was the 2−ΔΔCq method which facilitates the analysis of
relative changes in gene expression in real-time quantitative PCR
experiments (8). U6 was used as the
internal controls for miRNA quantification while β-actin was used
for mRNA. The primers used were: DNAJC2 forward,
GTTGCGTTGTCTGCTTGAGGT; DNAJC2 reverse, CTGTGAATCTGTTCCCTGCTG;
β-actin forward, AGCGAGCATCCCCCAAAGTT; β-actin reverse,
GGGCACGAAGGCTCATCATT. Bulge-loop miRNA RT-qPCR Primer Sets specific
for miR-672-3p were designed by Guangzhou RiboBio Co., Ltd.
Immunohistochemical assay
The tissue samples were fixed in 4% paraformaldehyde
at 4°C overnight and sectioned. The tissue sections were mount on
slides and were twice dehydrated on a rotor at room temperature
over 10 min using a graded ethanol series: 15, 30, 50, 70, 90 and
100%. After removing the paraffin and rehydration, the 5-mm thick
sections were placed into a pressure cooker for 5 min to restore
the antigen in the nucleus by using the citrate method (9). Then, 3% H2O2 was
used at room temperature for 5 min to suppress the endogenous
peroxidase activity to reduce the background. The samples were
blocked by 10% normal goat serum (WISENT, Inc.) and 5% BSA
(Beyotime Institute of Biotechnology) in TBS for 1 h at room
temperature. The sections were incubated with primary antibody
(1:400 dilution) overnight at 4°C and then washed three times in
PBS (Beyotime Institute of Biotechnology). The primary antibodies
used were DNAJC2, (cat. no. ab134572), Cyclin D1 (cat. no.
ab134175), CDK2 (cat. no. ab235941), AKT (cat. no. ab18785),
phosphorylated (p)-AKT (cat. no. ab38449), P21 (cat. no. ab188224)
and β-actin (cat. no. ab179467). All the antibodies were purchased
from Abcam. After incubation with HRP-conjugated secondary antibody
(1:100 dilution; cat. nos. A0181, A0216 and A0208; Beyotime
Institute of Biotechnology), sections were subjected to the DAB
reaction (Beyotime Institute of Biotechnology). Images of the
sections were captured by using a digital light microscope camera
(Nikon Corporation). For each image, ≥5 fields were randomly
selected and presented with a magnification of ×100 and ×400. The
positive area was analyzed using ImageJ 1.63 software (National
Institutes of Health), as previously described (10).
Knockdown and overexpression of DNAJC2
and miR-672-3p
Small interfering RNA (siRNA) and negative control
of DNAJC2 were Provided by Genomeditech. miR-672-3p mimics and
inhibitors were supplied by Guangzhou RiboBio Co., Ltd. The
sequence for miR-672-3p mimics was 5′-CCGATTCACCAACGA-3′ and the
control was 5′-TTTCATACATTCCAGC-3′. pLenti-Blast-CMV-MCS-V5 1.2
vector was used for overexpression (GeneScript). The empty vector
was applied for control group. Lipofectamine® 3000
(Invitrogen; Thermo Fisher Scientific, Inc.) with 5 µg plasmid
(both control group and overexpression plasmid) was use for
transfection in the DLD-1 and HCT-116 cell lines at room
temperature overnight, with the density of 1×104/ml.
Cells were cultured for 6 h before changing the medium in cell
incubator. After 48 h, the efficiency of knockdown and
overexpression assay was assessed via RT-qPCR detection.
Cell viability assay
To explore the effect of DNAJC2 on the proliferation
of CRC cells, 96-well plates, each well containing 2×103
cells, were cultured in 100 µl nutrient solution (Applied
Biosystems; Thermo Fisher Scientific, Inc.). Cell Counting Kit-8
(CCK-8; Dojindo Molecular Technologies, Inc.) was used to detect
the cell viability. A total of 10 µl CCK-8 was added to each well
at 24, 48, 72 and 96 h and mixed with 90 µl of serum-free medium
per well. After incubation (5% CO2 and 37°C) for 2 h,
the absorbance value was measured by microplate spectrophotometer
with wavelength set as 450 nm.
5-Ethynyl-2′-deoxyuridine (EdU)
assay
EdU assay kit (cat. no. C0071S; Beyotime Institute
of Biotechnology) was used to measure cell proliferation. Cells
were transferred into 96-well plates and cultured for 48 h, each
well containing 5×103 cells. Subsequently, EdU (100 µl;
10 µM) was mixed with nutrient solution and incubated at 4°C for 3
h. After that, cells were immersed in 4% paraformaldehyde at 4°C
for 15 min and then treated with 0.3% Triton X-100 for 10 min at
room temperature, which enhanced the permeability of the cells.
Following washing with 3% BSA, a Nikon TI-DH light microscope
(Nikon Corporation) was used to capture images at ×100
magnification. In total, five independent visual fields were
randomly selected for imaging, and ImageJ was used for optical
density analysis.
Plate colony formation assay
Cells (500) were seeded into 6-well plates in 37°C.
Every 5 days, each well was washed with PBS. After 10 days, cells
were immersed into 0.2% crystal violet dye for staining at room
temperature for 15 min. Following washing with PBS, colonies (≥50
cells/colony) in each well were manually counted via Photoshop CS6
(Adobe Systems, Inc.) and images were captured using a digital
camera (Canon DS126211; Canon, Inc.).
Cell cycle distribution detection
First trypsin was used to digest the cells. The
cells were centrifuged at 2,000 × g for 5 min at room temperature.
Following washing with PBS, the cells were immersed into 75% ethyl
alcohol and stored at 20°C overnight. Subsequently, the cells were
washed with PBS again and stained with PI staining solution (300 µl
per Flow tube; Thermo Fisher Scientific, Inc.) for 10 min at room
temperature in the dark, in accordance with the manufacturer's
protocols. RNase was added for flow cytometry (Beyotime Institute
of Biotechnology). The analysis was performed using a FACSCalibur
flow cytometer (BD Biosciences) and CellQuest software (version
3.0; BD Biosciences).
Western blot analysis
A RIPA kit (Beyotime Institute of Biotechnology) was
used to obtain the whole protein from cells and tissues. The
protein concentration was measured via BCA Protein Assay kit
(Beyotime Institute of Biotechnology). Then, 10% SDS-PAGE was used
for electrophoresis with 20 µg protein in each lane. Protein was
transferred onto polyvinylidene difluoride membranes (EMD
Millipore). The membranes were blocked in 10% BSA solution
(Beyotime Institute of Biotechnology) at room temperature for 2 h
and incubated in an incubation box at 4°C with the primary
antibodies overnight. The membranes then were washed with TBS with
Tween-20 (5% TBST) two or three times every 10–15 min.
Subsequently, the membranes were incubated with immunoglobulin G
(anti-mouse or rabbit) at room temperature for 2 h and washed with
TBS + 0.001% Tween-20 buffer two or three times again, each time
lasted 10–15 min. ECL Plus (EMD Millipore) was adopted for
detection of protein expression levels by Bio-Imaging System. The
primary antibodies included were: DNAJC2 (1:2,000; cat. no.
ab231318), P21 (1:5,000; cat. no. ab109520; both Abcam), AKT
(1:2,000; cat. no. 10176-2-AP) and p-AKT (1:3,000; cat. no.
66444-1-Ig; both from ProteinTech Group, Inc.). Secondary
antibodies were goat anti-rabbit IgG (H+L; 1:5,000; cat. no.
111-035-003; Jackson ImmunoResearch Laboratories, Inc.).
Luciferase reporter assay
HCT-116 cells and 293T cells were planted into
24-well plates (5×103 per well). The overexpressed
plasmid vectors were designed and provided by Genomeditech and
reporter gene plasmid or miR-672-3p mimics were transfected into
cells via Lipofectamine™ 2000. After 48 h, firefly or
Renilla luciferase assay reagent were mixed with cell lysate
in each group. A luciferase Reporter Gene Assay kit (cat. no.
GM-040501A; Genomeditech) was adopted for assessing luciferase
activity on an Infinite M1000 PRO microplate reader (Tecan Group,
Ltd.).
Statistical analysis
Data were presented in forms of mean ± standard
deviation. GraphPad Prism 5.0 software (GraphPad Software Inc.) and
SPSS 20.0 (IBM Corp.) were used to analyze the data. Paired
Student's t-test was used for comparisons expression of DNAJC2 in
paired clinical samples. Moreover, an unpaired Student's t-test was
employed for analyzing difference in the other experiments. One-way
ANOVA followed by Tukey's and repeated measures ANOVA followed by
paired t-test with Bonferroni's correction were used to compare
data sets which contained a number of groups. P<0.05 was
considered to indicate a statistically significant difference.
Results
DNAJC2 is upregulated in CRC and is
associated with clinicopathological factors
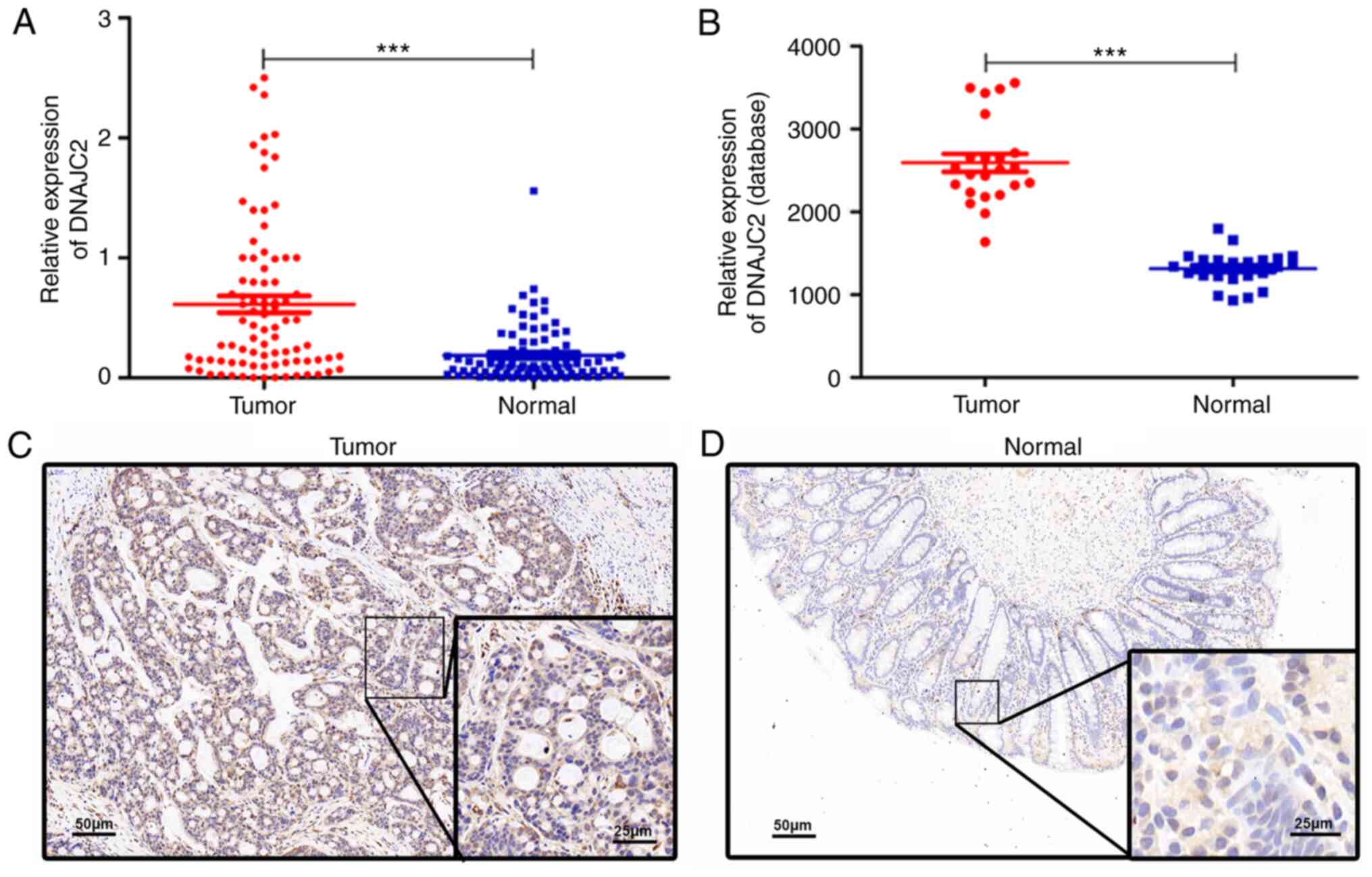
The expression of DNAJC2 was measured by RT-qPCR in
84 pairs of CRC and normal tissues. The expression of DNAJC2 was
higher in tumor tissues compared with normal tissues, in the cohort
enrolled in The First Affiliated Hospital of Gannan Medical
University and the data obtained from the GEO database (Fig. 1A and B). Similarly, the protein
expression was also markedly higher in tumor tissues compared with
normal tissues, which was confirmed by immunohistochemical analysis
(Fig. 1C and D). Next, the
association between DNAJC2 expression level and clinicopathological
features of patients with CRC (Table
I) was ascertained and the 84 cases of CRC tissue samples were
divided into two experimental groups (high, n=42; low, n=42, medium
as the cutoff). The expression level of DNAJC2 was significantly
associated with tumor diameter. However, no significant association
was found between DNAJC2 levels and other factors such as age, sex,
carcinoembryonic antigen, depth of invasion or primary tumor
site.
 | Table I.Association between DNAJC2 expression
and the clinicopathological characteristics of patients with
CRC. |
Table I.
Association between DNAJC2 expression
and the clinicopathological characteristics of patients with
CRC.
| Variable | n | Low n=42 | High n=42 | P-value
χ2 |
|---|
| Age (years) |
|
|
|
|
|
<60 | 32 | 18 | 14 |
|
| ≥60 | 52 | 24 | 28 | 0.833 |
| Sex |
|
|
|
|
| Male | 54 | 26 | 28 |
|
|
Female | 30 | 16 | 14 | 0.649 |
| Tumor diameter
(cm) |
|
|
|
|
|
<5 | 42 | 26 | 16 |
|
| ≥5 | 42 | 16 | 26 |
0.029a |
| Lymph node
invasion |
|
|
|
|
|
Negative | 34 | 16 | 18 |
|
|
Positive | 50 | 26 | 24 | 0.657 |
| Depth of
invasion |
|
|
|
|
|
T1+T2 | 19 | 9 | 10 |
|
|
T3+T4 | 65 | 33 | 32 | 0.794 |
| Primary tumor
site |
|
|
|
|
|
Colon | 41 | 22 | 19 |
|
|
Rectum | 43 | 20 | 23 | 0.513 |
| CEA |
|
|
|
|
|
<4.7 | 40 | 22 | 18 |
|
|
≥4.7 | 44 | 20 | 24 | 0.382 |
DNAJC2 regulates the proliferative
capacities of CRC cells in vitro
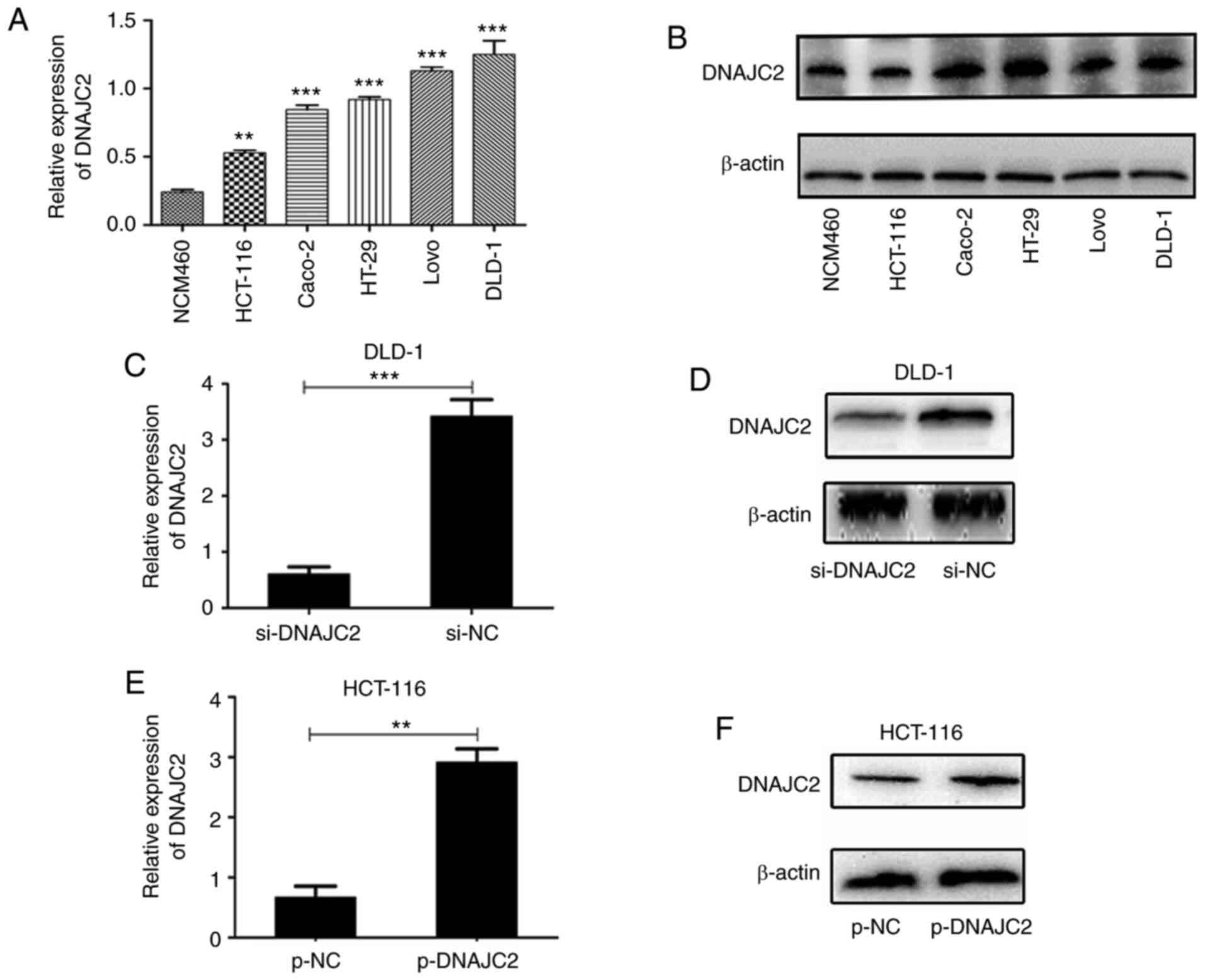
To detect the expression level of DNAJC2 in CRC
cells, five CRC cell lines (HCT-116, LoVo, Caco-2, HT-29 and DLD-1)
and the normal colon epithelial cell line (NCM460) were examined by
RT-qPCR and western blotting. It was found that the mRNA and
protein levels of DNAJC2 were significantly increased in the CRC
cell lines compared with that in the NCM460 cells (Fig. 2A and B). Based on the endogenous
expression of DNAJC2, the DLD-1 cell line was chosen for
transfection with si-DNAJC2 while HCT-116 cell line was chosen for
transfection with overexpression plasmid. According to the results
of RT-qPCR and western blotting, the expression of DNAJC2 was
markedly decreased in DLD-1 cells (Fig.
2C and D) and was overexpressed by transfection with the
DNAJC2-overexpressing plasmid in HCT116 cells (Fig. 2E and F). To further clarify the role
of DNAJC2 in CRC, the DNAJC2-overexpressing plasmid and si-DNAJC2
were selected to perform further in vitro experiments.
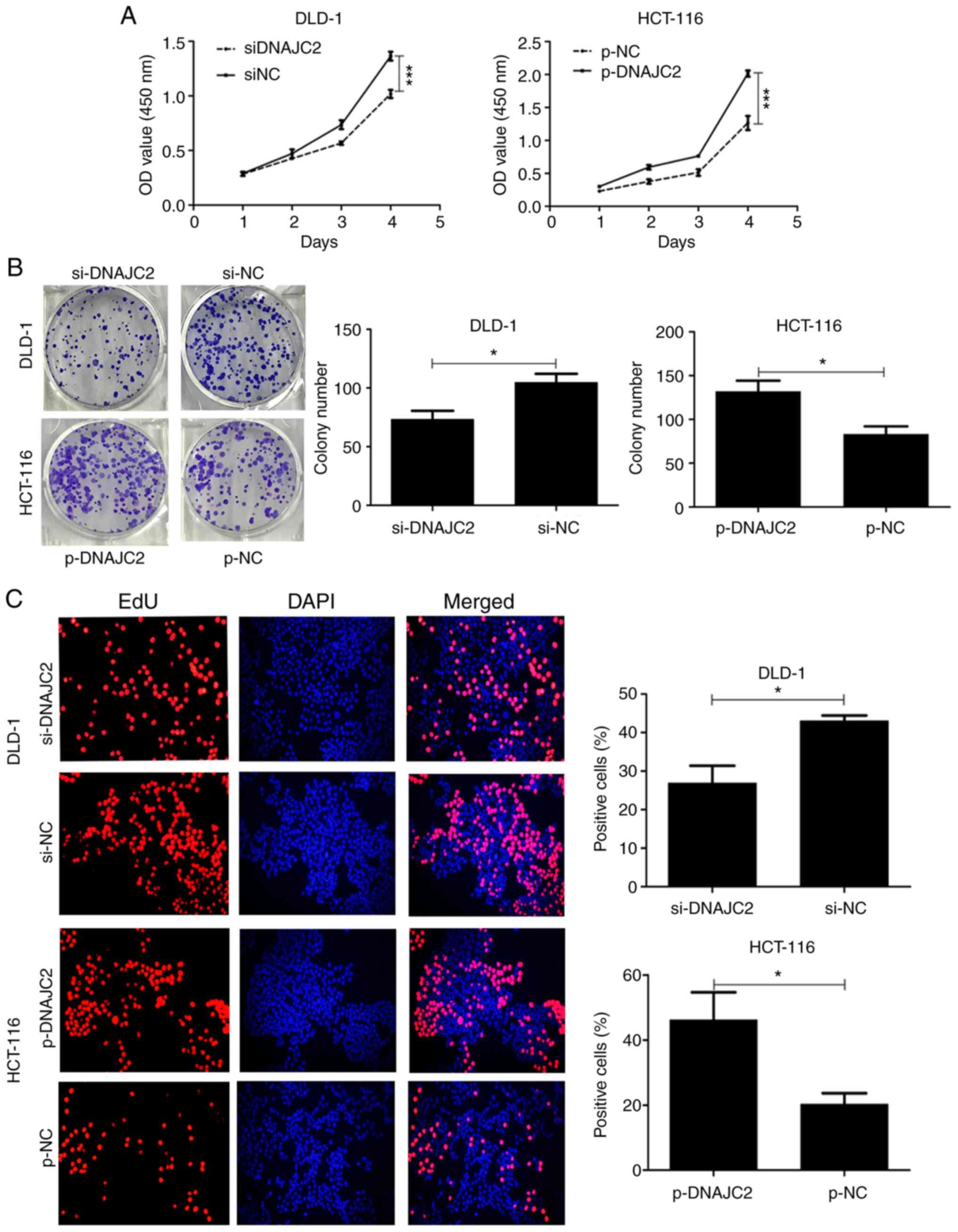
Initially, it was hypothesized that DNAJC2 may serve
an important role in CRC cell proliferation. Cells treated with
si-DNAJC2 revealed markedly decreased proliferation compared with
the control group (Fig. 3A).
Colony-formation assay results also confirmed that DNAJC2 knockdown
significantly reduced formation of colonies of DLD-1 cells
(Fig. 3B). EdU and DAPI staining
also showed that decreased expression of DNAJC2 suppressed the
percentage of proliferative cells (Fig.
3C). Conversely, DNAJC2-overexpressing plasmid HCT-116 cells
displayed prominently increased cell proliferation.
DNAJC2 influences CRC cell cycle
progression to promote tumor growth
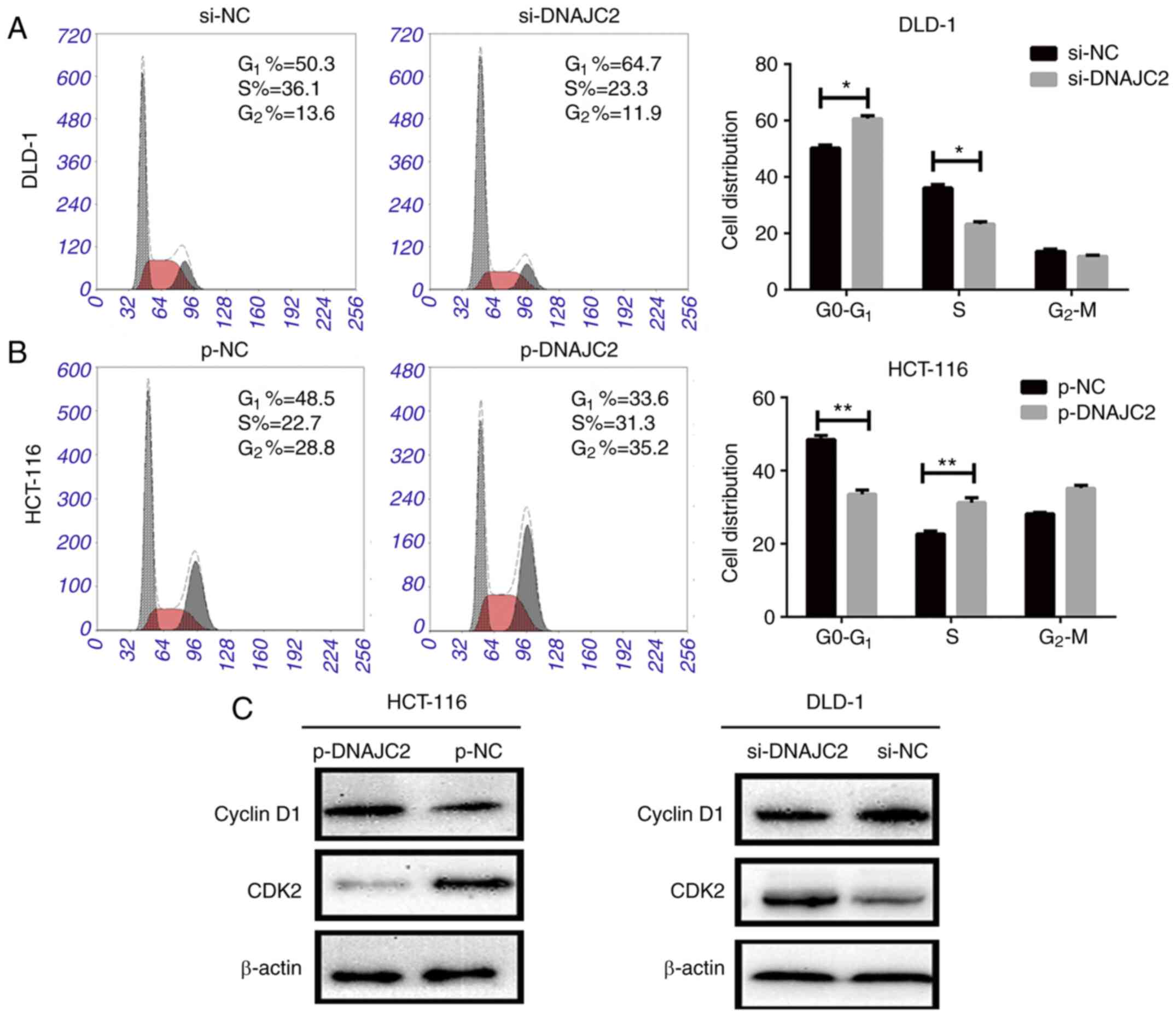
To clarify the role of DNAJC2 in CRC cell cycle
progression, the cell cycle of DLD-1 and HCT-116 cell lines were
analyzed with flow cytometry. It was observed that the DLD-1 cell
line transfected with si-DNAJC2 apparently induced a G1
phase arrest compared with the control group. DNAJC2-overexpressing
plasmid HCT-116 cells could also accelerate cell cycle progression
(Fig. 4A and B). It was also
discovered that increased DNAJC2 could promote the expression of
cyclinD1 while attenuate the level of CDK2, otherwise, suppression
of DNAJC2 could decease the level of cyclinD1 while promote CDK2
expression. (Fig. 4C). Therefore,
the results showed that DNAJC2 might influence the cell cycle
progression to promote the growth of CRC cells.
miR-672-3p reversely targets DNAJC2
through binding with the 3′UTR
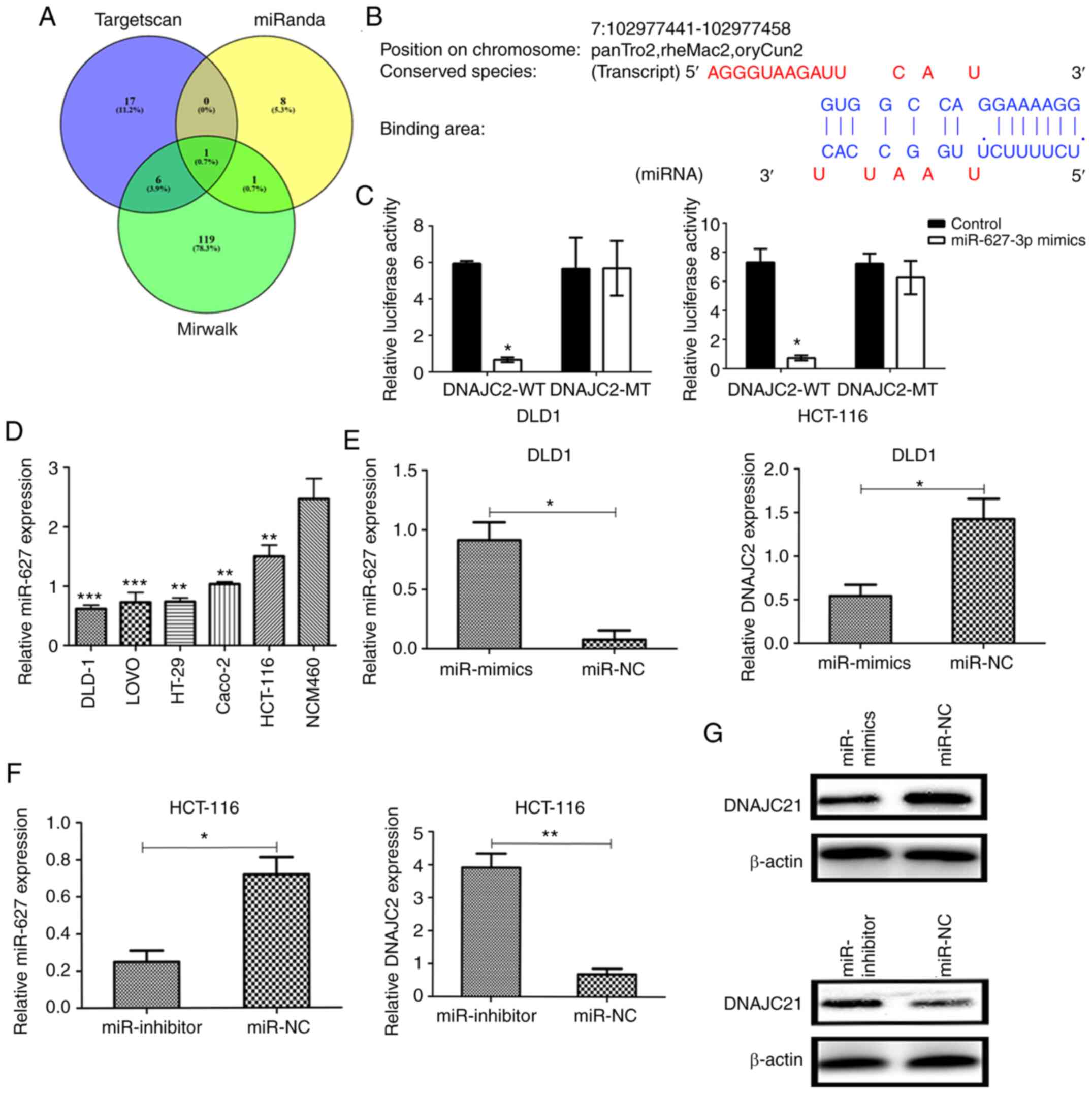
In order to explore the detailed mechanism of DNAJC2
on CRC cell lines, online databases including TargetScan V3.1
(http://www.targetscan.org/mamm_31/)
and miRWalk 2021 (http://mirwalk.umm.uni-heidelberg.de/) were used to
predict the potential miRNA for DNAJC2. The 3′UTR region was used
as input to predict potential miRNA that may bind with DNAJC2.
miR-672-3p was predicted with the highest binding score according
to the database (Fig. 5A and B).
Next luciferase reporter assay was adopted to confirm the
prediction. Mutant-type (MT) and wild-type (WT) DNAJC2 3′UTR
sequences (site-directed mutations in the putative miR-672-3p
target sites were included in the former) were cloned into reporter
plasmids. The luciferase activity of DNAJC2 3′UTR was inhibited by
over-expression of miR-672-3p. By contrast, the luciferase activity
of DNAJC2-3′UTR-MUT was unaffected and the same phenomenon was also
observed in HCT-116 cell lines (Fig.
5C). The above conclusions demonstrated that miR-672-3p is a
potential upstream regulator of DNAJC2.
In CRC cell lines and the normal epithelial colon
cell line the expression of miR-672-3p was downregulated in the CRC
cell lines compared with NCM460 (normal epithelial colon cell;
Fig. 5D). In transfected CRC cells,
RT-qPCR and western blotting demonstrated that expression level of
DNAJC2 was lower in DLD-1 miR-672-3p-mimics and higher in HCT-116
miR-672-3p-inhibitor compared with the negative control (Fig. 5E-G). These findings indicated that
miR-672-3p can directly negatively control DNAJC2.
miR-672-3p attenuates cell
proliferation and regulates DNAJC2 through AKT/P21 pathway in CRC
cell
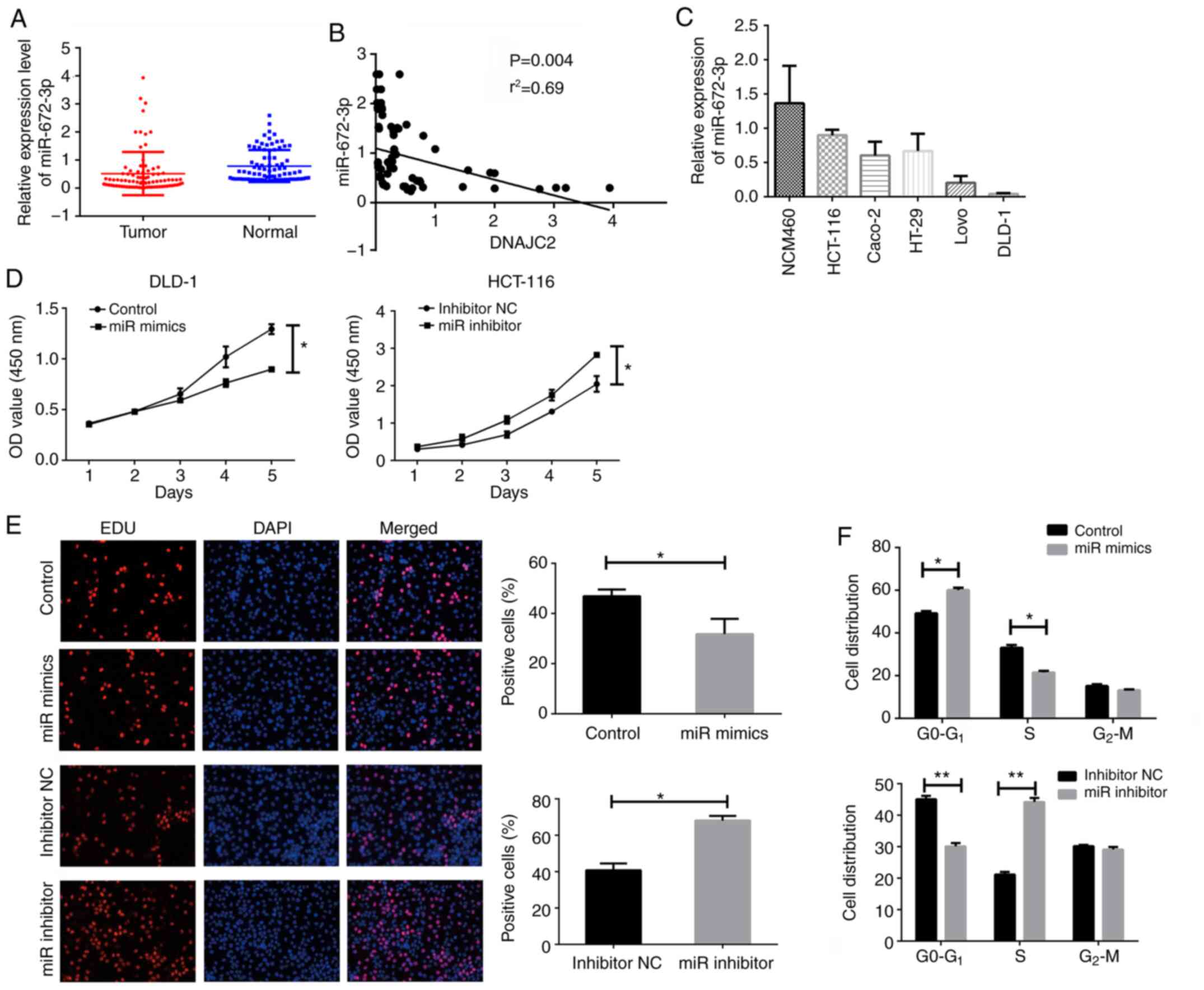
The expression of miR-672-3p was next investigated
in the same 84 patients with CRC. As presented in Fig. 6A, a downregulated level of
miR-672-3p was observed in tumor tissues of patients with CRC.
Meanwhile, the correlation of miR-672-3p and DNAJC2 in tissues
samples was also analyzed and there was an inverse correlation
between miR-672-3p and DNAJC2 (Fig.
6B).
To investigate whether miR-672-3p regulated cell
proliferation, the endogenous expression of miR-672-3p in CRC cells
lines was detected. The expression of miR-672-3p was relatively
higher in HCT-116 and lower in DLD-1 (Fig. 6C). The function of miR-672-3p in CRC
cells was next detected. CCK8 and EDU assay were used for detecting
the cell proliferation. It was found that increased miR-672-3p
could attenuate the proliferation of CRC cells. Following
miR-672-3p knockdown, cell growth could also be promoted (Fig. 6D and E). Cells overexpressed with
miR-672-3p presented an increased G0-G1 and
decreased S phase percentage. The opposite result was observed with
miR-672-3p knockdown (Fig. 6F).
The PI3K/AKT pathway serves an essential role in
cell proliferation and cell cycle activities (11). Accordingly, P21 is positively
correlated with malignancies by mediating cellular senescence
(12,13). The present study explored whether
the PI3K/AKT pathway and P21 were closely connected to the cellular
response which is mediated by DNAJC2 in CRC. Western blotting
demonstrated decreased levels of p-AKT in the cells transfected
with si-DNAJC2. The opposite phenomenon can be seen in the HCT-116
cell line, which was transfected by plasmid (Fig. 7A). The effects of miR-672-3p on
CCND1, CDK2, p-AKT and P21 in CRC cells was also investigated. It
was found that cells overexpressed with miR-672-3p mimics
suppressed Cyclin D1 while increasing CKD2 in CRC cell lines. The
opposite result was observed following miR-672-3p knockdown. It was
also found that cell treated with miR-672-3p mimics could decreased
the phosphorylation of AKT with the accumulation of P21; however,
when cells were treated with miR-672-3p inhibitor, the
phosphorylation of AKT was increased accompanied by the suppression
of P21 (Fig. 7B). According to the
above results, it was concluded that DNAJC2 facilitates the
proliferation of colorectal cancer via the p-AKT pathway.
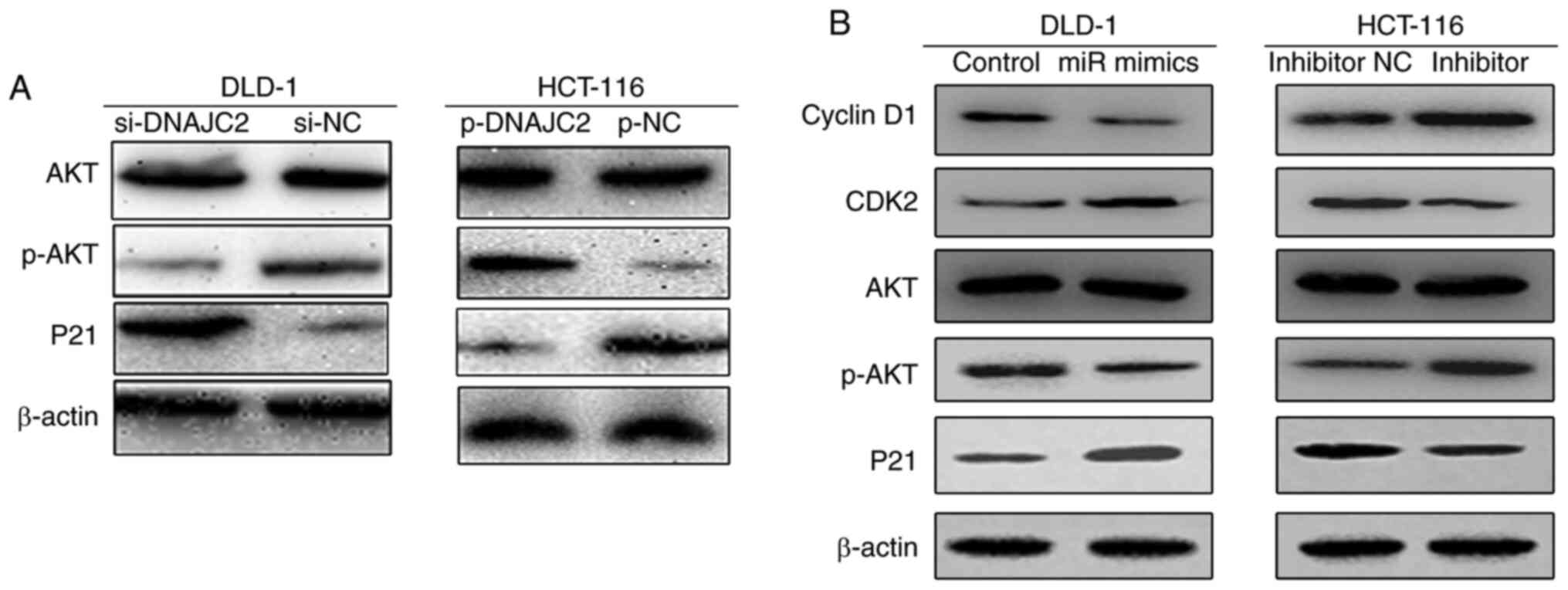 | Figure 7.miR-672-3p and DNAJC2 influences the
activation of the p-AKT pathways in CRC cells. (A) Expression of
AKT, p-AKT and P21 was evaluated by western blotting in DLD-1 and
HCT-116 cell lines compared with NC after the transfection of siRNA
or plasmid. (B) Expression of Cyclin D1, CDK2, AKT, p-AKT and P21
was evaluated by western blotting in DLD-1 and HCT-116 cell lines
after treating with miR-672-3p mimics or inhibitor. miR/miRNA,
microRNA; DNAJC2, DnaJ Heat Shock Protein Family (Hsp40) Member C2;
CRC, colorectal cancer; p-, phosphorylated; NC, negative control;
si, small interfering. |
Discussion
An increased level of DNAJC2 was found in tumor
tissues as well as the CRC cell lines. It has been shown that
DNAJC2 can accelerate transcriptional activation by particularly
replacing polycomb-repressive complex 1 from chromatin at the
beginning of differentiation (14).
DNAJC2 has also been reported to be closely associated with
different types of cancer. For example, in gastric cancer, DNAJC2
promotes cell proliferation, migration and invasion and induced
apoptosis in a pathway mediated by p53 (15). In acute myeloid leukemia DNAJC2 can
regulate the INK4-ARF locus during cellular proliferation and
senescence (14). Knockdown of
DNAJC2 can strongly inhibit leukemia progression by obtaining
command of genes related to retinoic acid (RA) by means of its
connection with the RA receptor α and its binding to genes relevant
to RA, both of which describe the mechanism of DNAJC2 in acute
myeloid leukemia at the molecular level (14). In breast cancer, DNAJC2 can be
considered as potential molecular marker in the sera of patients
with breast cancer (16). Another
study verified that depletion of DNAJC2 promotes endocrine
resistance via deregulation of cell death and cell survival related
pathways (16). All these findings
are consistent with the present study which indicated DNAJC2
function as a cancer promotor.
As already noted, increasing evidence demonstrates
that miRNAs serve a crucial role in various biological processes
and their unusual expression is implicated in a number of diseases,
such as cancer and autoimmune disorders. miRNAs can act as a key
role in several regulatory pathways involving CRC biology, such as
epithelial-mesenchymal transition, angiogenesis and metastasis. For
example, miR-1258 is a suppressive factor in a E2F8-mediated
pathway. indicating that miR-1258 can be used as a therapeutic
target for human CRC (17). Another
study showed that miR-195-5p can downregulate the expression of
YAP1 and miR-195-5p-mediated downregulation of YAP1, which impedes
genesis and development of tumors in CRC (18). For miR-672-3p, it has been noted
that miR-672-3p can serve a role of tumor suppressor in a number of
types of cancer (19). However, its
expression, function and mechanism in CRC remains to be elucidated.
Overexpression of miR-672 could promote the angiogenesis of
adipose-derived mesenchymal stem cells by suppressing tissue
inhibitor of metalloproteinase 2 (20). miR-672 also induces osteoblast
differentiation and mineralization through inhibition of Smad
ubiquitin regulatory factor 1 with enhanced Runt-related
transcription factor 2 transcriptional activation in mice (21). However, little has been reported on
the function and mechanism in colorectal cancer.
Nevertheless, there are still a number of questions
regarding the association between DNAJC2 and CRC remaining to be
explored. First, as a transcription factor, the downstream targets
of DNAJC2 need further examination. Second, whether miR-672-3p
promotes or inhibits the development of colorectal cancer remains
to be elucidated. Meanwhile, the role of DNAJC2 in invasion and
metastasis of CRC is still unclear. Third, one of the limitations
of the present study was not being able to provide results of in
vivo studies it is hoped to provide more evidence with the help
of a xenotransplantation model or other methods in future
studies.
In conclusion, the present study verified that
DNAJC2 can promote CRC cell proliferation and cell cycle
progression by mediating cyclinD1, CDK2, p-AKT and p21 and
miR-672-3p can regulate the expression of DNAJC2, all of which
proved that DNAJC2 may serve a vital role as a potential
therapeutic target of CRC.
Acknowledgements
Not applicable.
Funding
The present study was funded by a grant of The First
Affiliated Hospital of Gannan Medical University (grant no.
GMU1011).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
XY conceived the idea. HL designed experimental
technology. HL and JL performed the majority of the experiments,
analyzed and interpreted the results, produced figures and wrote
the manuscript. HZ and XL performed the remainder of the
experiments and analyzed the data. HL, JL, HZ and XL helped design
the experimental studies and edited the manuscript. HL interpreted
the results and wrote the manuscript. All authors reviewed and
approved the final manuscript. HZ and XL confirm the authenticity
of all the raw data.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of The First Affliated Hospital of Gannan Medical
University (approval no. IRB20170082)
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD, Fedewa SA, Ahnen DJ,
Meester RGS, Barzi A and Jemal A: Colorectal cancer statistics,
2017. CA Cancer J Clin. 67:177–193. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Luo H, Zhang NQ, Huang J, Zhang X, Feng
XL, Pan ZZ, Chen YM, Fang YJ and Zhang CX: Dietary intakes of
different forms and sources of iron and colorectal cancer risk: A
case-control study in China. Br J Nutr. 121:735–747. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Inadomi J and Jung B: Colorectal
cancer-recent advances and future challenges. Gastroenterology.
158:289–290. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Dekker E, Tanis PJ, Vleugels JLA, Kasi PM
and Wallace MB: Colorectal cancer. Lancet. 394:1467–1480. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Aloia L, Demajo S and Di Croce L: ZRF1: A
novel epigenetic regulator of stem cell identity and cancer. Cell
Cycle. 14:510–515. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Imamura T, Komatsu S, Ichikawa D, Miyamae
M, Okajima W, Ohashi T, Kiuchi J, Nishibeppu K, Kosuga T, Konishi
H, et al: Overexpression of ZRF1 is related to tumor malignant
potential and a poor outcome of gastric carcinoma. Carcinogenesis.
39:263–271. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Demajo S, Uribesalgo I, Gutiérrez A,
Ballaré C, Capdevila S, Roth M, Zuber J, Martín-Caballero J and Di
Croce L: ZRF1 controls the retinoic acid pathway and regulates
leukemogenic potential in acute myeloid leukemia. Oncogene.
33:5501–5510. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Waldvogel HJ, Curtis MA, Baer K, Rees MI
and Faull RL: Immunohistochemical staining of post-mortem adult
human brain sections. Nat Protoc. 1:2719–2732. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Varghese F, Bukhari AB, Malhotra R and De
A: IHC Profiler: An open source plugin for the quantitative
evaluation and automated scoring of immunohistochemistry images of
human tissue samples. PLoS One. 9:e968012014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Huang Y, Lin L, Shen Z, Li Y, Cao H, Peng
L, Qiu Y, Cheng X, Meng M, Lu D, et al: CEBPG promotes esophageal
squamous cell carcinoma progression by enhancing PI3K-AKT
signaling. Am J Cancer Res. 10:3328–3344. 2020.PubMed/NCBI
|
|
12
|
Liu Y, Marin A, Ejlerskov P, Rasmussen LM,
Prinz M and Issazadeh-Navikas S: Neuronal IFN-beta-induced
PI3K/Akt-FoxA1 signalling is essential for generation of FoxA1+Treg
cells. Nat Commun. 8:147092017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Liu P, Begley M, Michowski W, Inuzuka H,
Ginzberg M, Gao D, Tsou P, Gan W, Papa A, Kim BM, et al:
Cell-cycle-regulated activation of Akt kinase by phosphorylation at
its carboxyl terminus. Nature. 508:541–545. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ribeiro JD, Morey L, Mas A, Gutierrez A,
Luis NM, Mejetta S, Richly H, Benitah SA, Keyes WM and Di Croce L:
ZRF1 controls oncogene-induced senescence through the INK4-ARF
locus. Oncogene. 32:2161–2168. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Katsoulas A, Rachid Z, McNamee JP,
Williams C and Jean-Claude BJ: Combi-targeting concept: An
optimized single-molecule dual-targeting model for the treatment of
chronic myelogenous leukemia. Mol Cancer Ther. 7:1033–1043. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Dyachenko L, Havrysh K, Lytovchenko A,
Dosenko I, Antoniuk S, Filonenko V and Kiyamova R: Autoantibody
response to ZRF1 and KRR1 SEREX antigens in patients with breast
tumors of different histological types and grades. Dis Markers.
2016:51287202016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhang Z, Li J, Huang Y, Peng W, Qian W, Gu
J, Wang Q, Hu T, Ji D, Ji B, et al: Upregulated miR-1258 regulates
cell cycle and inhibits cell proliferation by directly targeting
E2F8 in CRC. Cell Prolif. 51:e125052018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liu X, Zhou Y, Ning YE, Gu H, Tong Y and
Wang N: MiR-195-5p inhibits malignant progression of cervical
cancer by targeting YAP1. OncoTargets Ther. 13:931–944. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zierau O, Helle J, Schadyew S, Morgenroth
Y, Bentler M, Hennig A, Chittur S, Tenniswood M and Kretzschmar G:
Role of miR-203 in estrogen receptor-mediated signaling in the rat
uterus and endometrial carcinoma. J Cell Biochem. 119:5359–5372.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen M, Zhou M, Fu Y, Li J and Wang Z:
Effects of miR-672 on the angiogenesis of adipose-derived
mesenchymal stem cells during bone regeneration. Stem Cell Res
Ther. 12:852021. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ahmad N, Kushwaha P, Karvande A, Tripathi
AK, Kothari P, Adhikary S, Khedgikar V, Mishra VK and Trivedi R:
MicroRNA-672-5p identified during weaning reverses osteopenia and
sarcopenia in ovariectomized mice. Mol Ther Nucleic Acids.
14:536–549. 2019. View Article : Google Scholar : PubMed/NCBI
|