Introduction
Articular cartilage that is damaged or undergoing
degeneration is unable to repair itself to a healthy state, which
may lead to osteoarthritic alterations (1). Osteoarthritis (OA) is one of the most
common pain-inducing and disabling diseases worldwide (2). Patients living with OA experience
significantly compromised quality of life and increased health-care
costs (3). Age is a primary
contributing factor to OA progression; however, various other
factors are also involved in the development of age-related OA,
including articular cartilage damage, articular space stenosis,
osteophyte formation, subchondral bone alteration and soft tissue
degeneration (4). The prevalence of
OA in China is increasing with the aging of the population
(1). Despite its high prevalence,
the pathogenesis of OA is still relatively unknown, and there are
currently no therapeutic cures for the disease (5,6). Due
to the lack of effective treatments for the reversal of articular
cartilage degradation, current treatments aim to improve pain
symptoms and delay disease progression. Although prosthetic joint
replacement is an effective treatment for end-stage OA, the
functioning of the prosthetic joint is inferior to that of its
healthy counterpart. Accordingly, the development or identification
of conservative treatments that effectively prevent OA progression
is critical (2).
Gene therapy for OA has become a research focus, as
studies continue to identify genes that play important roles in the
development and progression of OA (3,7). Zhou
et al (8) found that
conditional deletion of Indian hedgehog (Ihh) protein in mice
chondrocytes attenuates OA progression, suggesting the possibility
that blocking Ihh signaling can be used as a therapeutic approach
to prevent or delay cartilage degeneration. Therefore,
gene-targeted therapy may be an effective approach to repairing
cartilage damaged by OA.
The forkhead box (FOX)O proteins are a subfamily of
the FOX family of transcription factors. They are transcriptional
activators that are also considered to be longevity factors
(9). This protein family serves
important roles in the regulation of tissue homeostasis through
autophagy and oxidative stress (9,10), and
is essential for maintenance of the hematopoietic stem cell pool
(11). FOX protein O4 (FOXO4) has
been shown to be involved in the cell cycle, apoptosis regulation
and tumorigenesis through transcriptional regulation (12–14).
Baar et al (15) previously
reported increased FOXO4 mRNA expression and a decreased rate of
apoptosis after the induced aging of IMR-90 fibroblast cells with
ionizing radiation. Conversely, inhibiting FOXO4 expression
decreased the expression levels of FOXO4 mRNA and increased the
rate of apoptosis.
In summary, FOXO4 is closely associated with
cellular aging and apoptosis; however, potential associations
between FOXO4 and cartilage degeneration or OA are yet to be
elucidated, and past studies have reported controversial findings.
Akasaki et al (16) revealed
that young articular cartilage expressed FOXO1 and 3 proteins, but
not FOXO4 (16). Additionally,
Ludikhuize et al (17) found
that FOXO4 was expressed and phosphorylated in the knee joint
tissues of patients with rheumatoid arthritis and OA. Matsuzaki
et al (18) also reported
that FOXO1, 3 and 4 may be key to cartilage development and
maturation, and may prevent OA-associated cartilage damage
(18).
In the present study, an effective chondrocyte
degeneration model was established in Sprague-Dawley (SD) rats. The
model was used to experimentally determine the expression levels of
FOXO4 and collagen type II α1 chain (Col2α) in normal and
degenerated rat cartilage cells, and to investigate potential
associations between FOXO4 expression levels and chondrocyte
degeneration.
Materials and methods
Animals
A total of 30 male SD rats (4- and 16-week-old) were
purchased from Hunan Silaike Jingda Laboratory Animal Co., Ltd. The
weight of the SD rats at 4 and 16 weeks were 70–100 g and 380–480
g, respectively. They were housed at 20–22°C and 40–70% humidity
with normal circadian rhythm light/dark cycle. They were fed
sterilized feed (10g/100 g weight) and water (10–15 ml/100g weight)
ad libitum. There were 10 rats in each group. All animal
experiments were approved by the Animal Research and Care Committee
of Nanchang University (approval no. NCXK-2019-21; Hangzhou,
China). All efforts were made to minimize suffering and the number
of animals used in the study. All rats were anesthetized by an
intraperitoneal injection of sodium pentobarbital (40 mg/kg).
Isolation and culture of chondrocytes
from the femoral heads of SD rats
All rats were euthanized by an overdose of
pentobarbital sodium (120 mg/kg, intraperitoneal administration).
The absence of respiration, heartbeat and the corneal/palpebral
reflex indicated animal death. Following routine disinfection, the
femoral head was exposed using surgical forceps and ophthalmic
scissors. The cartilage tissue covering the femoral head was minced
into pieces of ~1.0 mm2, and then digested with 0.25%
trypsin (Gibco; Thermo Fisher Scientific, Inc.) at 37°C for 30 min.
The precipitate was collected after centrifugation at 300 × g for
10 min at room temperature, and further digested with 0.2% type II
collagenase (Beijing Solarbio Science & Technology Co., Ltd.)
at 37°C for 3 h. The resulting precipitate was then agitated for 2
min and transferred to a clean centrifuge tube. After
centrifugation at 300 × g for 10 min at room temperature, the
precipitate containing the chondrocytes was collected and the
entire process was repeated once more.
The dissociated chondrocytes were repeatedly
pipetted, washed with phosphate-buffered saline (PBS; HyClone;
Cytiva), and centrifuged three times at 300 × g for 5 min at room
temperature. The cells were then seeded into 50-cm2
culture flasks filled with Dulbecco's modified Eagle's medium
(DMEM; HyClone; Cytiva) supplemented with 15% fetal bovine serum
(FBS; Gibco; Thermo Fisher Scientific, Inc.). The flasks were
incubated at 37°C (5% CO2) and the medium was changed
every 3–4 days until the cells had reached confluency. The cells
were divided into groups for use in the following experiments, and
proliferation was periodically observed and images were
captured.
Morphological identification of SD rat
chondrocytes
On reaching ~80% confluence, chondrocytes at passage
1 and passage 2 underwent morphological identification.
Chondrocytes were stained using Toluidine blue (Shanghai Macklin
Biochemical Co., Ltd.) and Alcian blue (Beijing Solarbio Science
& Technology Co., Ltd.) for cellular identification, and were
examined under a light microscope (TS100-F; Nikon Corporation).
Cells were fixed with 4% paraformaldehyde in phosphate buffer.
Cells were fixed at room temperature for 10–20 min, washed with
running water for 3 times for 2 min and then stained. Cells were
respectively stained with 1% toluidine blue and 0.1% Alcian blue
for 30 min at 37°C, the excess dye washed away with double
distilled water, dehydrated with absolute ethanol and sealed with
neutral gum.
Col2α immunocytochemical staining
The second generation of chondrocytes was used for
Col2α immunocytochemical staining following adhesion to the flask
wall after culture for 24 h. The second-generation chondrocytes
were cultured in a 6-well plate for 24 h, and then fixed with 4%
paraformaldehyde for 30 min at room temperature. Then, 30%
H2O2 and methanol (1:50) were added for 30
min at room temperature, after which the chondrocytes were blocked
with 5% bovine serum albumin (BSA; Gibco; Thermo Fisher Scientific,
Inc.) in PBS for 1 h at room temperature. Monoclonal rat anti-Col2α
antibody (1:200; cat. no. 28459; ProteinTech Group, Inc.) was added
and the cells were incubated for 2 h at 37°C. After washing with
PBS, the appropriate biotin-conjugated secondary antibodies (1:200;
cat. no. SA00004-8; ProteinTech Group, Inc.) were applied for 30
min at 37°C. The chondrocytes were washed with PBS once more, and
then incubated with avidin-biotin peroxidase conjugate (SABC kit;
Wuhan Boster Biological Technology, Ltd.) for 30 min at 37°C.
Diaminobenzidine tetrahydrochloride (0.3%; DAB kit; Wuhan Boster
Biological Technology, Ltd.) was used as the substrate for the
peroxidase reaction. Staining was visualized and images were
captured with an immunofluorescence microscope (magnification,
×100; TS100-F; Nikon Corporation).
Experimental grouping
The experiments were conducted using three groups of
chondrocytes. Chondrocytes in the control group were obtained from
4-week-old rats, while those in the natural degeneration group were
obtained from 16-week-old rats. Chondrocytes in the induced
degeneration group were obtained from 4-week-old rats and treated
with IL-1β (PeproTech, Inc.) at 37°C for one week. For this
treatment, 10 ng/ml IL-1β solution was added to DMEM/F-12 medium
containing 15% FBS, which was used to culture the chondrocytes of
4-week-old rats from passage 1 until subsequent
experimentation.
β-galactosidase staining
β-galactosidase staining was conducted using the
β-galactosidase staining kit (Beijing Solarbio Science &
Technology Co., Ltd.) per the manufacturer's protocol. Staining at
room temperature for 15 min was performed and images were captured
using a fluorescence microscope (TS100-F; Nikon Corporation).
RNA extraction and reverse
transcription (RT)-semi-quantitative PCR
Chondrocytes from each group were washed with cold
PBS and preserved at −80°C. Total RNA was extracted using
TRIzol® reagent (Thermo Fisher Scientific, Inc.), and
500 ng RNA was reverse transcribed into cDNA using ReverTra Ace™
qPCR RT Master Mix with gDNA Remover (Toyobo Life Science). The RT
kit was used according to the manufacturer's protocol. The
generated cDNA was then used for mRNA screening in
RT-semi-quantitative PCR assays, which were performed in 20 µl with
1 µl prepared cDNA, 0.5 µl each primer and 10 µl SYBR Green PCR
Master Mix (Toyobo Life Science) using the C1000 Touch™ Thermal
Cycler (Bio-Rad Laboratories, Inc.) under the following conditions:
95°C for 1 min, 25 cycles at 95°C for 15 sec, 58°C for 30 sec and
72°C for 30 sec. The relative expression levels of each mRNA were
normalized to those of GAPDH. The primer sequences for Col2α1,
FOXO4 (both PeproTech, Inc.) and GAPDH are listed in Table I.
 | Table I.Primer sequences and product length
of primers used for reverse transcription-semi-quantitative
PCR. |
Table I.
Primer sequences and product length
of primers used for reverse transcription-semi-quantitative
PCR.
| Primer | Primer
sequence | Product length,
bp |
|---|
| GAPDH forward |
5′-GATGCTGGTGCCGAGTAC-3′ | 104 |
| GAPDH reverse |
5′-GCTGAGATGATGACCCTTTTGG-3′ |
|
| Col2α1 forward |
5′-CGAGGTGACAAAGGAGAAGC-3′ | 453 |
| Col2α1 reverse |
5′-CTGGTTGTTCAGCGACTTGA-3′ |
|
| FOXO4 forward |
5′-CCAGAGAATAAGAAGTCAGCCACAGAG-3′ | 147 |
| FOXO4 reverse |
5′-CTCCACCTCGGACGGTTCGG-3′ |
|
Agarose (1 g) was dissolved in 72 ml water and
cooled to 60°C. An RNA sample (3 µg) was prepared with 3X volume of
formaldehyde loading dye solution. Ethidium bromide (EB) was added
to formaldehyde loading dye solution to a final concentration of 10
µg/ml, and heated to 70°C with for 15 min to denature the sample.
The electrophoresis condition was at a voltage of 5–6 V/cm for 2 h,
until the length of bromophenol blue indicator gel was at least 2–3
cm. The gel was observed under ultraviolet transmitted light.
Protein extraction and western blot
analysis
Chondrocytes from each group were washed with cold
PBS and stored at −80°C until use. Total protein was extracted
using the TriPure method (Protein Extraction Reagent; Thermo Fisher
Scientific, Inc.) and quantified using a BCA Protein Assay kit
(Aidlab Biotechnologies., Ltd.). The mass of protein loaded per
lane was 20 µg, following which proteins were separated via
SDS-PAGE on 5% gels and separated proteins were transferred to a
0.45-nm PVDF membrane at 90 V for 120 min. Membranes were then
blocked with 5% BSA at 37°C for 1 h and washed with TBS containing
0.1% Tween-20 buffer. Subsequently, the PVDF membrane was incubated
with anti-GAPDH, anti-Col2α (1:1,000; cat. no. sc-47724 and
sc-52658; Santa Cruz Biotechnology, Inc.) and anti-FOXO4 (1:1,000;
cat. no. ab128908; Abcam) primary antibodies at 4°C overnight,
followed by incubation with the secondary antibody (1:1,000; cat.
no. sc-2005; Santa Cruz Biotechnology, Inc.). The protein bands
were detected using a Bio-Rad Chemiluminescent Imaging System
(Bio-Rad Laboratories, Inc.) and semi-quantified by densitometric
analysis using ImageJ version 1.8.0 software (National Institutes
of Health). The target grey value ratio represents the relative
expression of each target protein.
Statistical analysis
Data analysis was performed using SPSS 22.0
statistical software (IBM Corp). All experiments were repeated at
least three times, and the data are presented as the mean ±
standard deviation. The data between the three groups were compared
using the one-way ANOVA followed by Bonferroni's test. P<0.05
was considered to indicate a statistically significant
difference.
Results
Isolation and culture of SD rat
chondrocytes
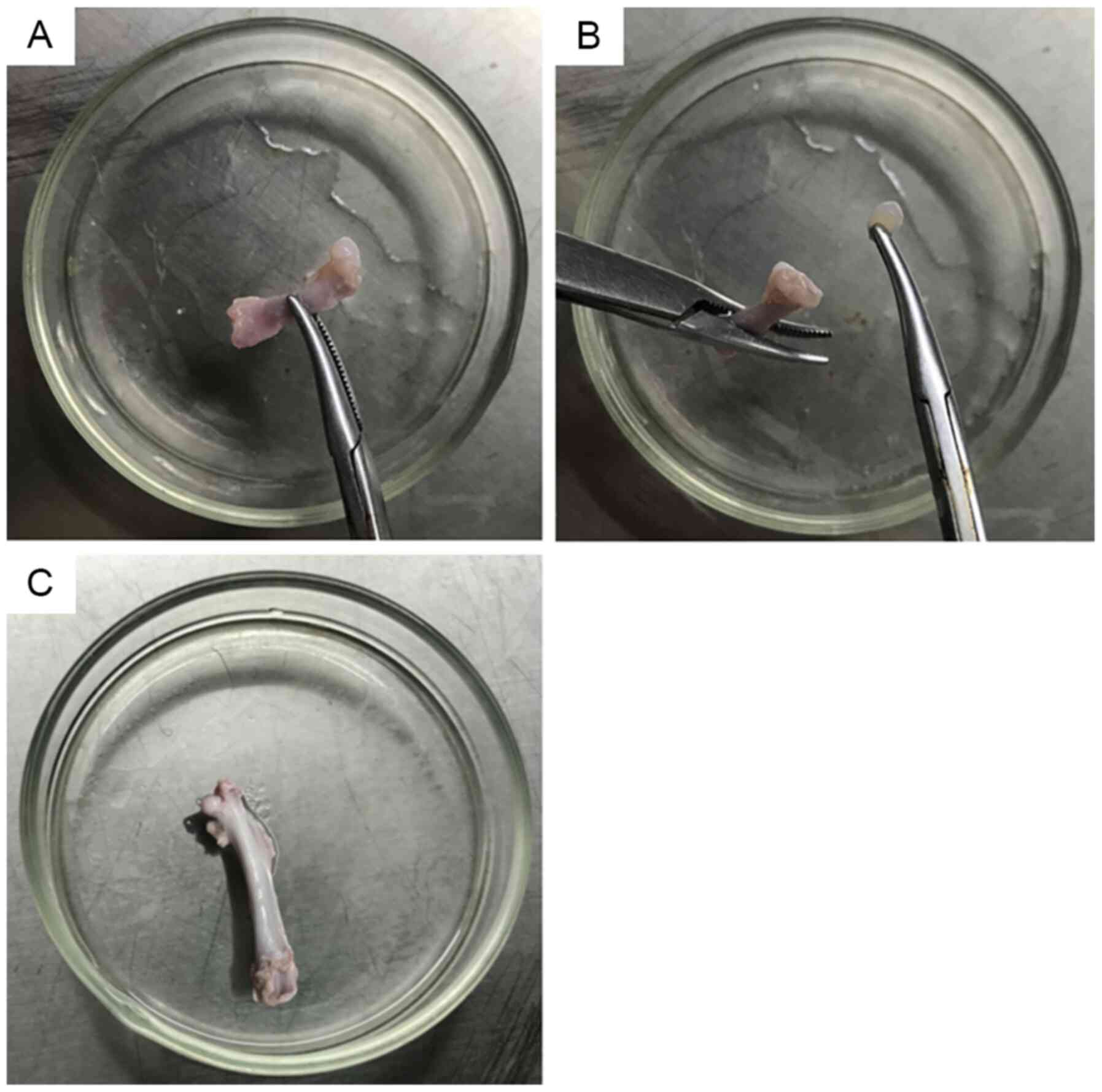
Femoral head cartilage from the 4-week-old rats was
thick, tough, brittle, elastic and easy to completely remove
(Fig. 1A and B). The femoral head
cartilage from the 16-week-old rats was thin and densely combined
with the subchondral bone (Fig.
1C), making it difficult to completely remove from the femoral
bone.
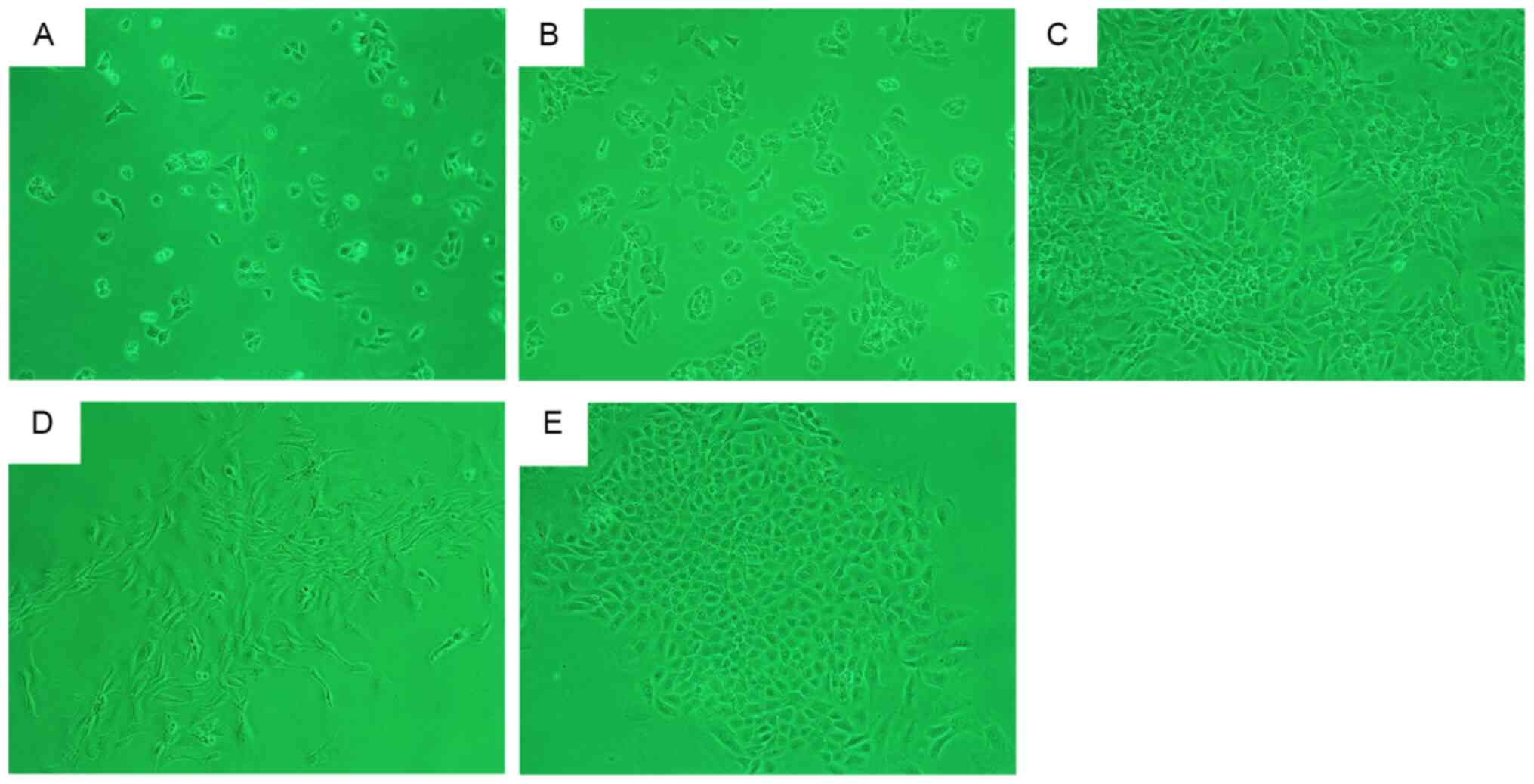
After incubation for 48 h, a large number of
adhering primary chondrocytes were obtained from the 4-week-old
rats. Adherent cells were uniformly distributed (Fig. 2A) with a ‘paving stone’-shaped
appearance (Fig. 2B). After 7 days,
the cells were overconfluent for the first passage (Fig. 2C). After 48 h of incubation, a small
number of primary chondrocytes from the 16-week-old rats had
adhered to the flask wall in a scattered arrangement, before
forming cell clusters and gradually expanding outwards (Fig. 2D and E). At ~2 weeks, the
multi-cluster cell groups merged to form a fully confluent
monolayer, which indicated that the cells were proliferating
correctly (data not shown). The second passage was performed after
3 and 4–5 days on cells from the 4- and 16-week-old rats,
respectively.
Identification and morphological
changes in SD rat chondrocytes
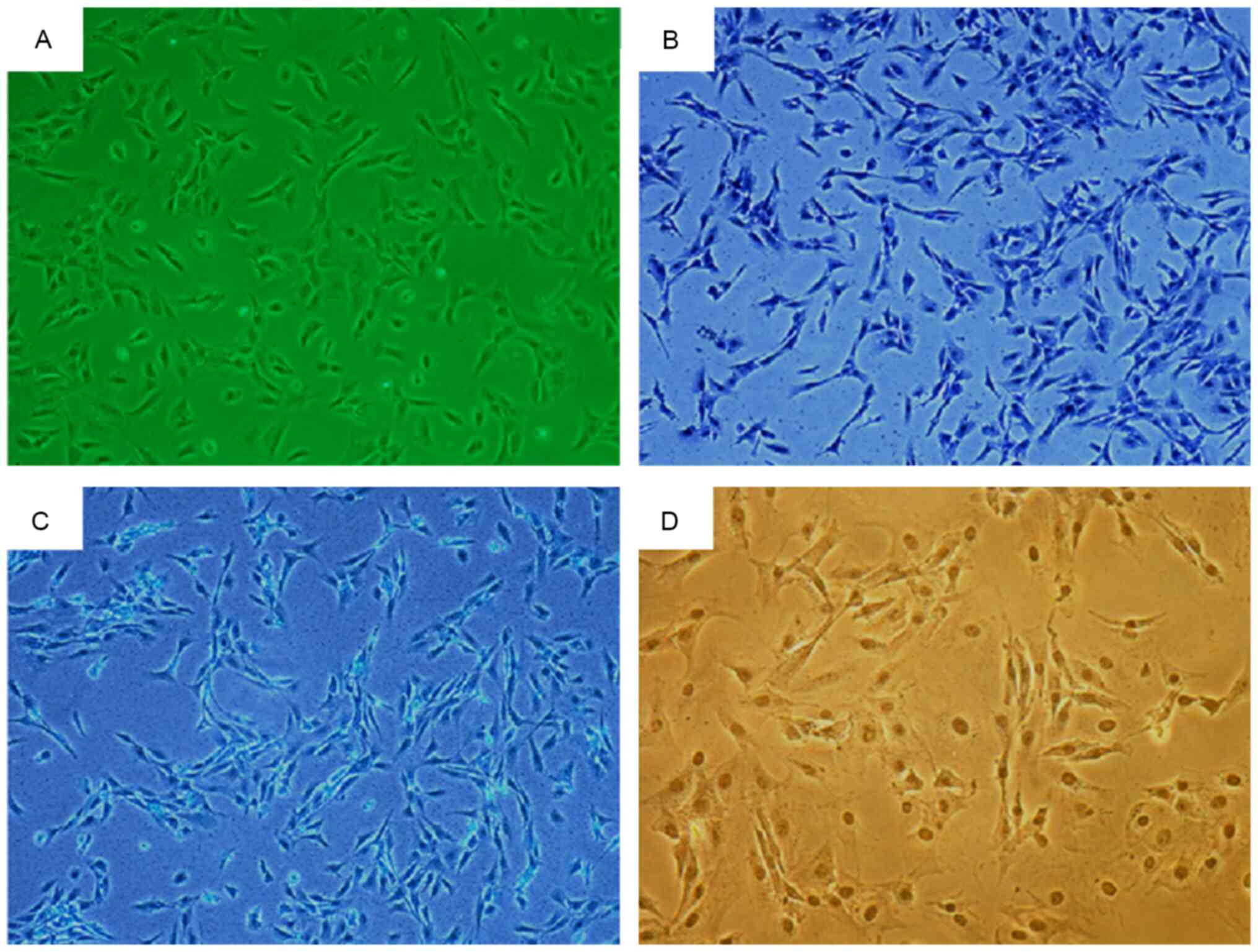
There were no notable morphological differences
between the two groups of chondrocytes, which appeared to be either
triangular or polygonal in shape with an abundant cytoplasm. The
chondrocytes of 4-week-old rats were shown in Fig. 3A. Toluidine blue staining revealed
blue-colored cytoplasm with a darker blue nucleus (Fig. 3B). The cytoplasm appeared pale blue
following Alcian blue staining (Fig.
3C). Immunocytochemical staining revealed cytoplasm with a
brownish-yellow appearance, while the nucleus appeared dark brown,
indicating Col2α staining (Fig.
3D). The three staining methods yielded positive results for
chondrocyte identification.
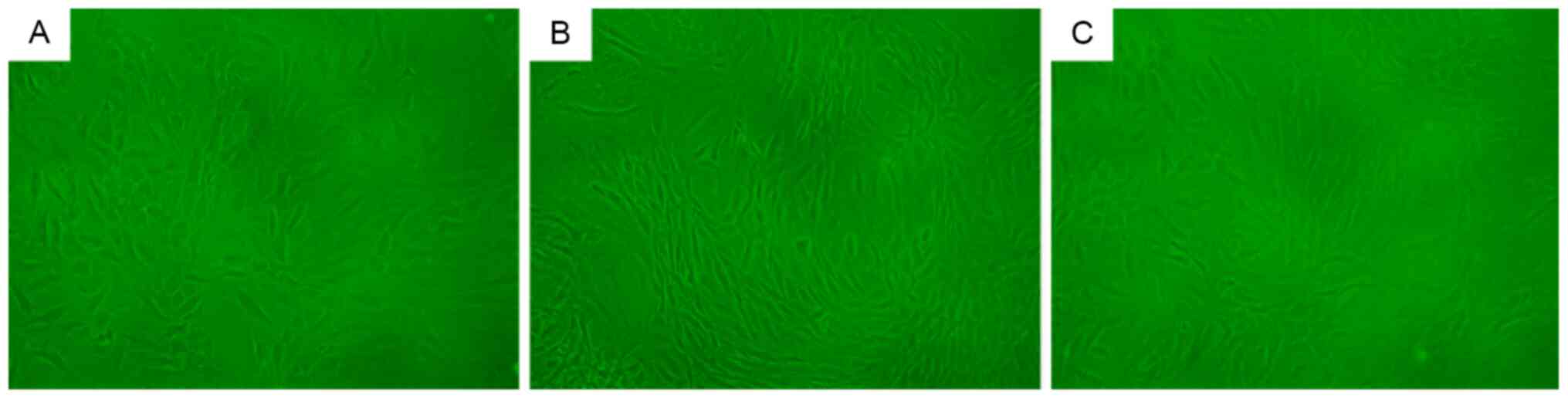
Second-generation cells in the control group
appeared spindle-shaped with a rich cytoplasm and good cell
refractive index (Fig. 4A). By
contrast, second-generation cells from the natural and induced
degeneration groups appeared fusiform or irregular in shape with a
reduced cell refractive index (Fig. 4B
and C).
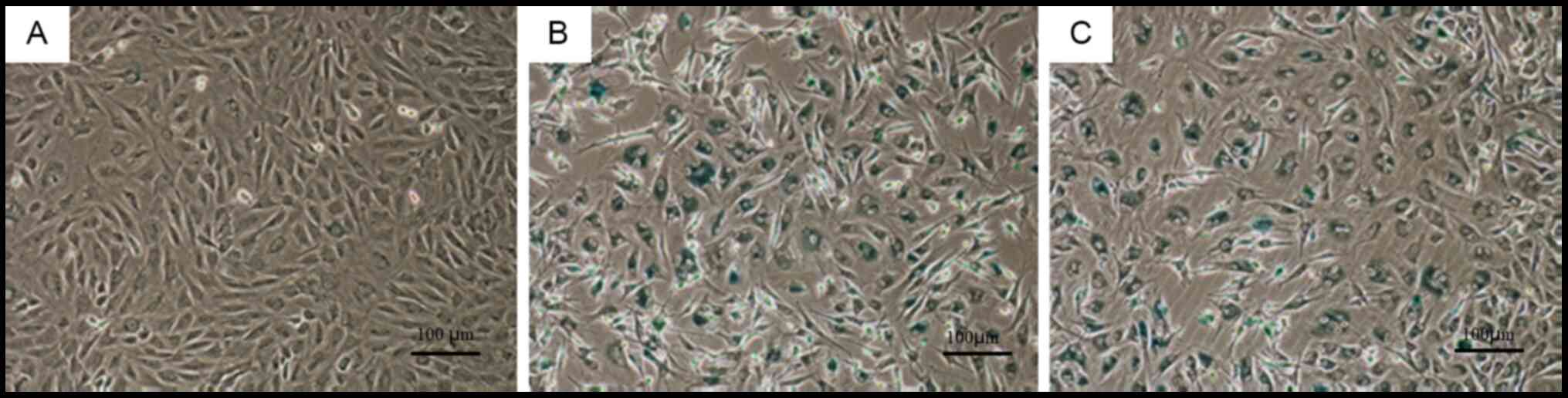
β-galactosidase staining
β-galactosidase staining of cells in the control
group did not elicit a notably positive response (Fig. 5A). By contrast, β-galactosidase
staining of cells in the natural and induced degeneration groups
resulted in a positive reaction, with dark blue staining in a large
number of cells (Fig. 5B and C).
There were no notable differences in β-galactosidase staining
intensity between the natural and induced degeneration groups.
Col2α and FOXO4 transcription and
protein expression
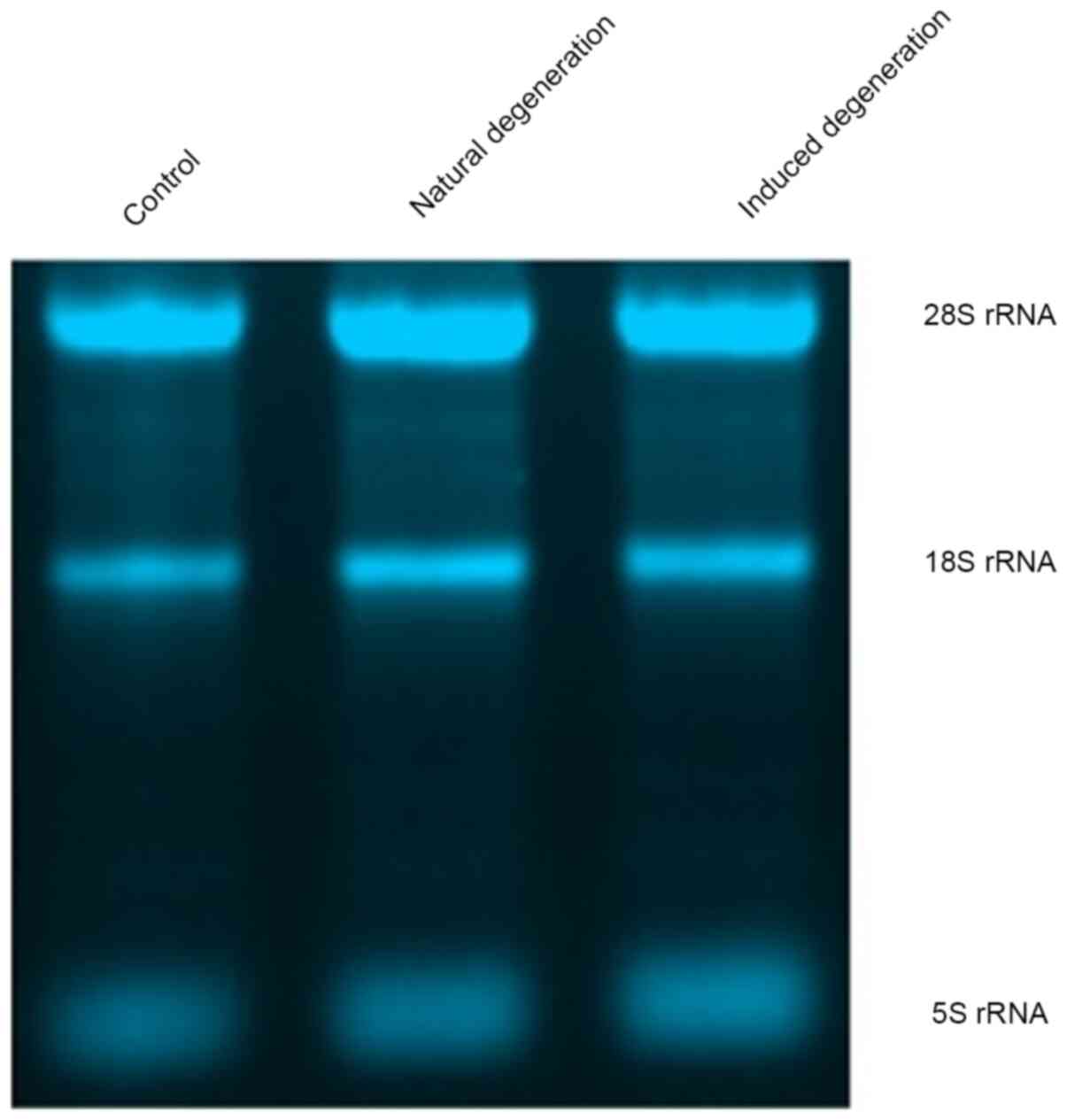
Each group contained 5S, 18S and 28S bands, and the
intensity of the 28S rRNA band was approximately twice that of the
18S rRNA band, indicating sufficient total RNA extraction from
chondrocytes in each group (Fig.
6).
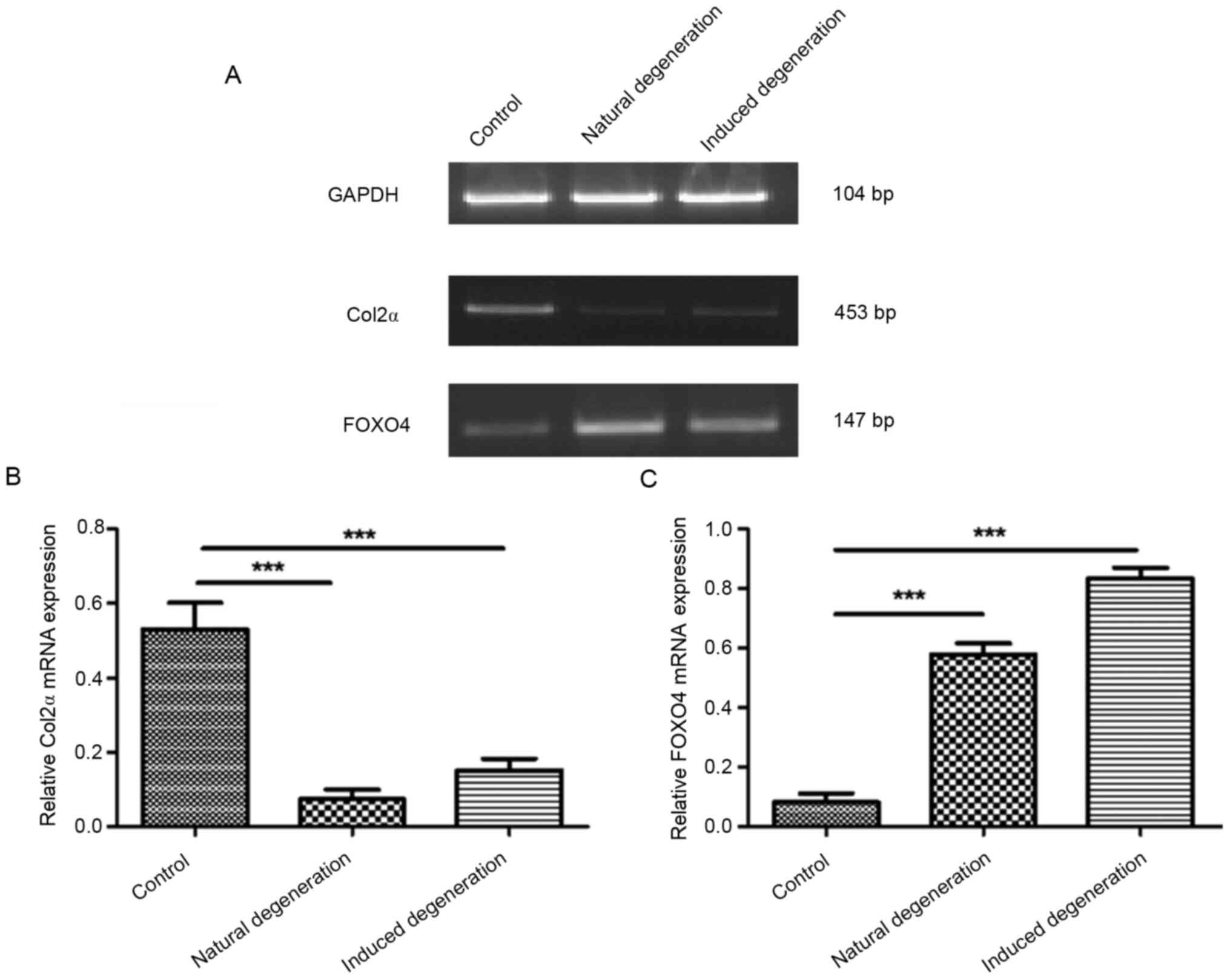
Compared with the control group, Col2α mRNA
expression was significantly reduced in chondrocytes from the
natural and induced degenerative groups (0.07±0.02 vs. 0.53±0.06,
and 0.15±0.03 vs. 0.53±0.06; Fig. 7A
and B). Additionally, FOXO4 mRNA expression was significantly
elevated in both groups compared with the control group (0.58±0.03
vs. 0.08±0.02, and 0.83±0.03 vs. 0.08±0.02; Fig. 7A and C).
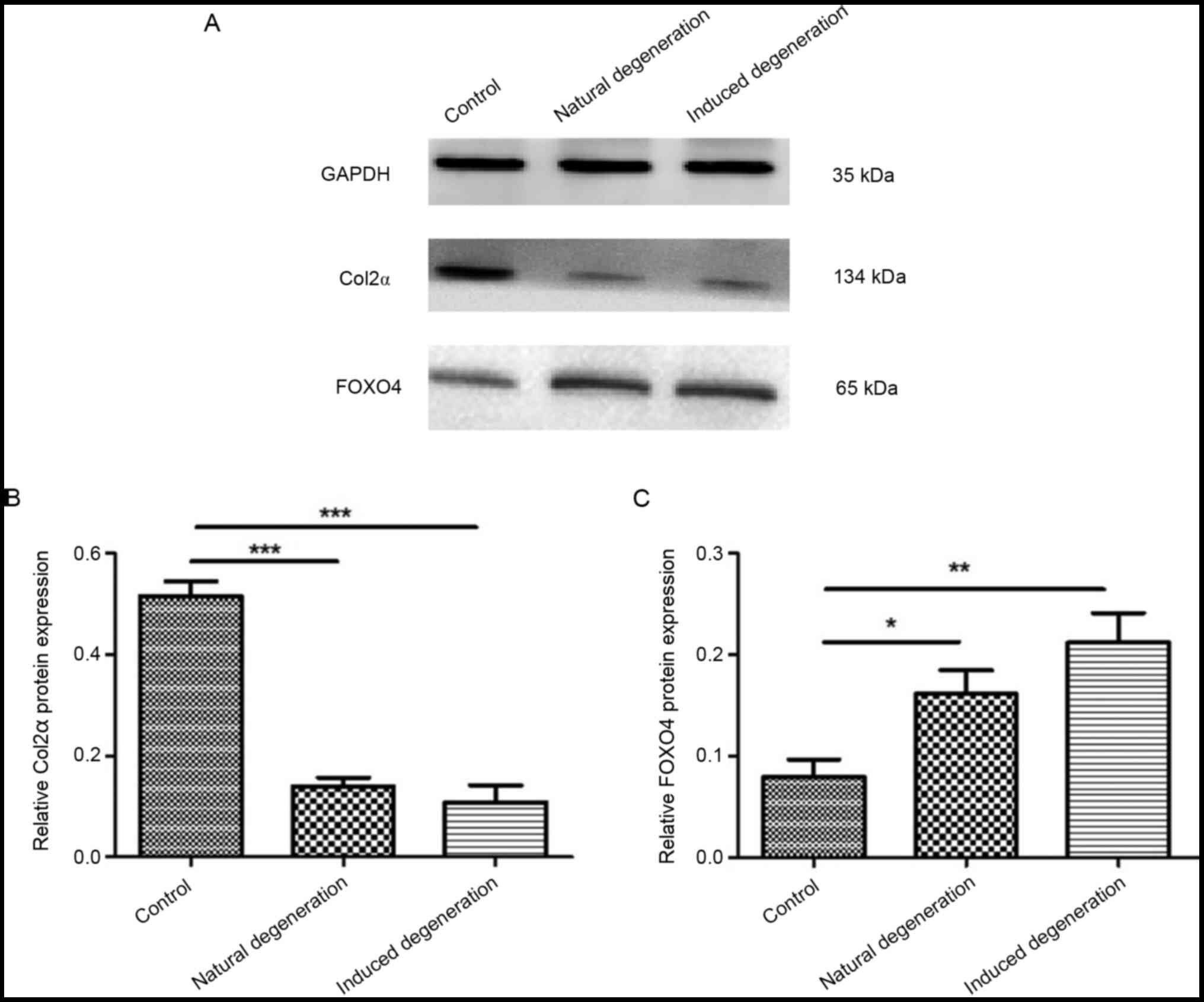
Furthermore, the protein expression levels of Col2α
in chondrocytes from the natural and induced degeneration groups
were significantly lower than those in the control group (0.14±0.01
vs. 0.51±0.02, and 0.11±0.03 vs. 0.51±0.02; Fig. 8A and B). FOXO4 protein expression
was significantly elevated when compared with the control group
(0.16±0.02 vs. 0.08±0.01, and 0.21±0.02 vs. 0.08±0.01; Fig. 8A and C).
Discussion
Aging is a natural process involved in the
pathogenesis of numerous diseases, and is considered a significant
risk factor for the development of OA (7). OA affects >50% of the world's
population >65 years of age (2),
but the association between OA and aging remains to be elucidated.
An enhanced understanding of the mechanisms relevant to OA
development is critical for the investigation of novel treatment
strategies. A well-established hypothesis is that chondrocyte
degeneration causes premature aging due to excessive mechanical
load or oxidative stress, leading to stress-induced aging, and
ultimately, the onset of OA (19).
Therefore, insights into the underlying mechanisms of chondrocyte
degeneration are critical to researching OA development. The
present study introduced the establishment of an effective
chondrocyte degeneration model, and investigated the association
between the expression of related proteins and cartilage
degeneration.
The selection of effective modeling methods is an
important part of disease research (20). In vivo, such animal methods
include artificially-induced and spontaneous models. Relatively
speaking, spontaneous models more adequately simulate the process
of natural articular cartilage degeneration, which was the ideal
model for the present study. Spontaneous OA can occur in guinea
pigs at ~3 months of age, and is characterized by an uneven or
absent cartilage surface and collagen dissolution in chondrocytes
(21). In the present study,
cartilage taken from 16-week-old SD rats displayed observable
thinning of the surface, decreased elasticity, hardening and
deformation in the shape of cultured cells after passage. These
observations suggested that key characteristics of cartilage
degeneration, such as collagen reduction and calcification, can
occur in rats at 16 weeks of age. The in vitro cell model
for cartilage degeneration is commonly established through the use
of inducing agents, such as ascorbate, rat serum and IL-1β
(22). In the present study, the
experiments successfully replicated the model described by Ding
et al (23), which
established a cellular model of arthritis in rabbits using media
with a final concentration of 10 ng/ml IL-1β.
Some aging cells express senescence-associated
β-galactosidase (SA-βgal), which is easily detected at pH 6.0, but
undetectable in young stationary-phase, immortal and tumor cells
(24). SA-βgal activity is closely
associated with aging cells and is widely used in research as a
biomarker for cellular aging (25).
Type II collagen is a major component of cartilage that is
specifically expressed by chondrocytes; together with
proteoglycans, type II collagen responsible for maintaining the
strength and hardness of chondrocytes (26). Sufficient production of type II
collagen is essential to maintaining the biomechanical properties
of cartilage, although its expression gradually decreases as
chondrocyte degeneration progresses (27). In the present study, a large number
of cells in the natural and induced degeneration groups stained
positive for β-galactosidase, while no such staining was observed
in the control cells. As confirmed by RT-semi-quantitative PCR and
western blotting, the mRNA and protein expression levels of Col2α
were decreased in the natural and induced degeneration groups
compared with in the control group.
Due to its role as a transcriptional activator, FOXO
protein is considered to be a pro-longevity factor (28). FOXO4 is a member of the FOXO protein
family that promotes longevity via its subcellular localization and
transcriptional activity, which are primarily regulated by
phosphorylation and acetylation (29). Baar et al (15) demonstrated that FOXO4 mRNA
expression was increased in aging-induced IMR90 fibroblasts,
whereas apoptotic rate was decreased. Conversely, FOXO4 mRNA
expression was decreased and the apoptotic rate was increased
following FOXO4 lentiviral inhibition. Furthermore, apoptosis was
induced by INK-ATTAC gene-mediated or cell-permeable FOXO4-DRI
peptides, which can eliminate aging cells. Overall, this may reduce
aging-related damage and dysfunction, while enhancing cellular
performance and longevity (30).
The aim of the present study was to investigate the
association between FOXO4 expression and chondrocyte degeneration
in rats. The study was predicated by that of Akasaki et al
(16), which revealed that FOXO4
was not expressed in the articular cartilage of young subjects, and
Matsuzaki et al (18), which
indicated that FOXO4 may serve a protective role against OA, and
was critical for cartilage development and maturation. It is
controversial whether FOXO4 expression has a protective effect in
OA. The aim of the present study was to reveal the relationship
between FOXO4 expression and OA. Accordingly, it was hypothesized
that FOXO4 may be associated with the chondrocyte degeneration
process. RT-semi-quantitative PCR and western blotting revealed
elevated FOXO4 mRNA and protein expression levels in the natural
and induced degeneration groups compared with in the control group,
although elucidation of the specific mechanisms requires further
investigation.
The pathogenesis of OA is still being studied, and
the relationship between the matrix metalloproteinase (MMP) family
and OA has received increasing attention. The OA progression was
slowed in MMP13 Col2ER mice 8, 12 and 16 weeks post-surgery.
Cartilage grading by blinded observers confirmed decreased
articular cartilage degeneration in MMP13 Col2ER mice at 8, 12 and
16 weeks compared with negative mice. MMP13 is the main enzyme that
targets cartilage degradation (31). The expression levels of MMP13 in OA
have been reported to be 10 times higher than those in normal
individuals (32). Furthermore, the
increased expression of MMP13 has been shown to be positively
correlated with the development of OA, which may be caused by
inflammation that stimulates synovial hypersecretion of MMP13,
which degrades collagen and breaks the balances of bone matrix
synthesis and decomposition, thus in turn leading to OA occurrence
(31). Studies (33,34)
have shown that FOXO4 may activate the transcription of the MMP9
gene in response to tumor necrosis factor-α signals; the
transactivation domains of FOXO4 are required for FOXO4 to activate
MMP9 transcription. However, to the best of our knowledge, at
present there are no reports on the effects of FOXO4 on the
expression of MMP13, and further research is needed.
In conclusion, the natural and induced degeneration
groups described in the present study represent effective models of
chondrocyte degeneration for use in experiments associated with
cartilage degeneration or OA. The transcription factor FOXO4 is
expressible in, and associated with, the degeneration of rat
chondrocytes, the expression level trend of chondrocytes was
consistent with the degree of cartilage degeneration. As a study
limitation, the effect of FOXO4 on chondrocyte degeneration needs
to be verified by more experiments. We acknowledge that the causal
relationship between FOXO4 and the pathogenesis of OA is not yet
clear. Follow-up work will continue to develop the effects of FOXO4
on the pathogenesis of OA by inducing knockout or overexpression of
FOXO4 in chondrocytes.
Acknowledgements
Not applicable.
Funding
This study was supported by a research grant from
the National Natural Science Foundation of China (grant no.
81860399), the Provincial Youth Science Fund of Jiangxi (grant no.
20142BAB215072), the Science and Technology Key R&D Program of
Jiangxi (grant no. 20202BBG72001) and the Science and Technology
Plan Project of Health Commission of Jiangxi (grant no. 202130504).
The funders had no role in study design, data collection and
analysis, decision to publish, and preparation of the
manuscript.
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors' contributions
JW, HZ and XF planned the experiments; JW, HZ, RD
and LX performed the experiments; JW, LX and MH analyzed the data;
JW, HZ and XF wrote the manuscript. JW, HZ and XF confirm the
authenticity of all the raw data All authors reviewed and approved
the final manuscript.
Ethics approval and consent to
participate
All animal experiments were approved by the Animal
Research and Care Committee of Nanchang University (approval no.
NCXK-2019-21; Hangzhou, China). All efforts were made to minimize
suffering and the number of animals used in the study.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Lu YH and Shi XB: Current status and
progress of epidemiological research on knee osteoarthritis. Chin J
Trad Med Traum Orthop. 20:81–84. 2012.(In Chinese).
|
|
2
|
Zhong L, Huang X, Karperien M and Post JN:
Correlation between Gene Expression and Osteoarthritis Progression
in Human. Int J Mol Sci. 17:E11262016. View Article : Google Scholar
|
|
3
|
Loeser RF, Collins JA and Diekman BO:
Ageing and the pathogenesis of osteoarthritis. Nat Rev Rheumatol.
12:412–420. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Chen Z, Liu D, Wang J, Wu L, Li W, Chen J,
Cai BC and Cheng H: Development of nanoparticles-in-microparticles
system for improved local retention after intra-articular
injection. Drug Deliv. 21:342–350. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Stack J and McCarthy G: Basic calcium
phosphate crystals and osteoarthritis pathogenesis: Novel pathways
and potential targets. Curr Opin Rheumatol. 28:122–126. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Louka ML, Zakaria ZM, Nagaty MM, Elsebaie
MA and Nabil LM: Expression of nucleostemin gene in primary
osteoarthritis. Gene. 587:27–32. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
McAlindon TE, Bannuru RR, Sullivan MC,
Arden NK, Berenbaum F, Bierma-Zeinstra SM, Hawker GA, Henrotin Y,
Hunter DJ, Kawaguchi H, et al: OARSI guidelines for the
non-surgical management of knee osteoarthritis. Osteoarthritis
Cartilage. 22:363–388. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhou J, Wei X and Wei L: Indian Hedgehog,
a critical modulator in osteoarthritis, could be a potential
therapeutic target for attenuating cartilage degeneration disease.
Connect Tissue Res. 55:257–261. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Maiese K: Forkhead Transcription Factors:
Formulating a FOXO Target for Cognitive Loss. Curr Neurovasc Res.
14:415–420. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lee GJ, Lim JJ and Hyun S: Minocycline
treatment increases resistance to oxidative stress and extends
lifespan in Drosophila via FOXO. Oncotarget. 8:87878–87890.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Miyamoto K, Araki KY, Naka K, Arai F,
Takubo K, Yamazaki S, Matsuoka S, Miyamoto T, Ito K, Ohmura M, et
al: Foxo3a is essential for maintenance of the hematopoietic stem
cell pool. Cell Stem Cell. 1:101–112. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen XD, Tang SX and Zhang JH:
Overexpression of FOXO4 promotes apoptosis oflaryngeal carcinoma
cells by regulates Wnt/β-cateninsignaling pathway. Chin Arch
Otolaryngol Head Neck Surg. 24:569–574. 2017.
|
|
13
|
Oteiza A and Mechti N: Control of FoxO4
Activity and Cell Survival by TRIM22 Directs TLR3-Stimulated Cells
Toward IFN Type I Gene Induction or Apoptosis. J Interferon
Cytokine Res. 35:859–874. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wei XW, Ma XW and Guo XH: FoxO4 in
diabetes and its complications. Chin J Diabetes. 3:276–279.
2014.(In Chinese).
|
|
15
|
Baar MP, Brandt RMC, Putavet DA, Klein
JDD, Derks KWJ, Bourgeois BRM, Stryeck S, Rijksen Y, van
Willigenburg H, Feijtel DA, et al: Targeted Apoptosis of Senescent
Cells Restores Tissue Homeostasis in Response to Chemotoxicity and
Aging. Cell. 169:132–147.e16. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Akasaki Y, Hasegawa A, Saito M, Asahara H,
Iwamoto Y and Lotz MK: Dysregulated FOXO transcription factors in
articular cartilage in aging and osteoarthritis. Osteoarthritis
Cartilage. 22:162–170. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ludikhuize J, de Launay D, Groot D, Smeets
TJ, Vinkenoog M, Sanders ME, Tas SW, Tak PP and Reedquist KA:
Inhibition of forkhead box class O family member transcription
factors in rheumatoid synovial tissue. Arthritis Rheum.
56:2180–2191. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Matsuzaki T, Alvarez-Garcia O, Mokuda S,
Nagira K, Olmer M, Gamini R, Miyata K, Akasaki Y, Su AI, Asahara H,
et al: FoxO transcription factors modulate autophagy and
proteoglycan 4 in cartilage homeostasis and osteoarthritis. Sci
Transl Med. 10:eaan07462018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Musumeci G, Szychlinska MA and Mobasheri
A: Age-related degeneration of articular cartilage in the
pathogenesis of osteoarthritis: Molecular markers of senescent
chondrocytes. Histol Histopathol. 30:1–12. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen J, Gu YT, Xie JJ, Wu CC, Xuan J, Guo
WJ, Yan YZ, Chen L, Wu YS, Zhang XL, et al: Gastrodin reduces
IL-1β-induced apoptosis, inflammation, and matrix catabolism in
osteoarthritis chondrocytes and attenuates rat cartilage
degeneration in vivo. Biomed Pharmacother. 97:642–651. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang T, Wen C, Lu WW, Yan CH and Chiu PKY:
A study on the role of subchondral bone change in very early stage
of osteoarthritis with Dunkin-Hartley guinea pigs. The 6th
International Congress of Chinese Orthopaedic Association (COA
2011), Beijing, China, 1–4 December 2011. https://hub.hku.hk/handle/10722/153093
|
|
22
|
Wang XJ, Zhang H, Zhan HS and Ding DF:
Establishment of chondrocyte degeneration model in vitro by rat
serum. Zhejiang Da Xue Xue Bao Yi Xue Ban. 44:308–314. 2015.(In
Chinese). PubMed/NCBI
|
|
23
|
Ding Q, Zhong H, Qi Y, Cheng Y, Li W, Yan
S and Wang X: Anti-arthritic effects of crocin in
interleukin-1β-treated articular chondrocytes and cartilage in a
rabbit osteoarthritic model. Inflamm Res. 62:17–25. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lee BY, Han JA, Im JS, Morrone A, Johung
K, Goodwin EC, Kleijer WJ, DiMaio D and Hwang ES:
Senescence-associated beta-galactosidase is lysosomal
beta-galactosidase. Aging Cell. 5:187–195. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Dimri GP, Lee X, Basile G, Acosta M, Scott
G, Roskelley C, Medrano EE, Linskens M, Rubelj I and Pereira-Smith
O: A biomarker that identifies senescent human cells in culture and
in aging skin in vivo. Proc Natl Acad Sci USA. 92:9363–9367. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Bagi CM, Berryman ER, Teo S and Lane NE:
Oral administration of undenatured native chicken type II collagen
(UC-II) diminished deterioration of articular cartilage in a rat
model of osteoarthritis (OA). Osteoarthritis Cartilage.
25:2080–2090. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ito K and Shinomura T: Development and
application of a new Silent reporter system to quantitate the
activity of enhancer elements in the type II Collagen Gene. Gene.
585:13–21. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Renault VM, Thekkat PU, Hoang KL, White
JL, Brady CA, Kenzelmann Broz D, Venturelli OS, Johnson TM, Oskoui
PR, Xuan Z, et al: The pro-longevity gene FoxO3 is a direct target
of the p53 tumor suppressor. Oncogene. 30:3207–3221. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Bourgeois B and Madl T: Regulation of
cellular senescence via the FOXO4-p53 axis. FEBS Lett.
592:2083–2097. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Valentijn FA, Falke LL, Nguyen TQ and
Goldschmeding R: Cellular senescence in the aging and diseased
kidney. J Cell Commun Signal. 12:69–82. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang M, Sampson ER, Jin H, Li J, Ke QH, Im
HJ and Chen D: MMP13 is a critical target gene during the
progression of osteoarthritis. Arthritis Res Ther. 15:R52013.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Du C, Smith A, Avalos M, South S, Crabtree
K, Wang W, Kwon YH, Vijayagopal P and Juma S: Blueberries Improve
Pain, Gait Performance, and Inflammation in Individuals with
Symptomatic Knee Osteoarthritis. Nutrients. 11:2902019. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Margină D, Ungurianu A, Purdel C,
Tsoukalas D, Sarandi E, Thanasoula M, Tekos F, Mesnage R, Kouretas
D and Tsatsakis A: Chronic Inflammation in the Context of Everyday
Life: Dietary Changes as Mitigating Factors. Int J Environ Res
Public Health. 17:41352020. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Li H, Liang J, Castrillon DH, DePinho RA,
Olson EN and Liu ZP: FoxO4 regulates tumor necrosis factor
alpha-directed smooth muscle cell migration by activating matrix
metalloproteinase 9 gene transcription. Mol Cell Biol.
27:2676–2686. 2007. View Article : Google Scholar : PubMed/NCBI
|