Introduction
Cancer statistics released in 2018 revealed that
lung cancer is the most frequently diagnosed type of cancer, as
well as a primary cause of cancer-associated mortality worldwide
(1). Non-small cell lung cancer
(NSCLC), the most common type of lung cancer, comprises lung
squamous cell carcinoma and lung adenocarcinoma (LUAD) (2,3).
As a major subtype of lung cancer, LUAD accounts for >40% of
lung cancer cases (4). Although
progress has been made in diagnostic and treatment methods in
recent years, the average 5-year survival rate of patients with
lung cancer is ~18% (5).
Therefore, elucidation of the underlying mechanisms of LUAD and the
identification of more effective therapeutic strategies is of
utmost urgency.
Thioredoxin domain-containing protein 9 (TXNDC9;
also known as ATP-binding protein associated with cell
differentiation or phosducin-like family of proteins 3), belongs to
the small, highly-conserved and ubiquitous TRX family which is
implicated in multiple biological processes via modulating
oxidative stress response (6,7).
TXNDC9 is upregulated in numerous types of cancer, promoting their
development. For example, Feng et al (8) reported that TXNDC9 exerts promotive
effects on cell survival and proliferation of prostate cancer. A
previous study reported that TXNDC9 is upregulated in colorectal
cancer (CRC) and functions as a tumor promoter owing to its
promotive role in cell proliferation and invasiveness (9). Moreover, TXNDC9 accelerates
hepatocellular carcinoma (HCC) cell proliferation, and TXNDC9
overexpression is associated with poor prognosis of patients with
HCC (10). According to results
from The Encyclopedia of RNA Interactomes (ENCORI; http://starbase.sysu.edu.cn/panCancer.php), TXNDC9
expression is increased in LUAD and its high expression is
associated with poor prognosis (Fig.
1A and B); therefore, it was hypothesized that TXNDC9 may
regulate the development of LUAD.
Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase
activation protein γ (YWHAG; also known as 14-3-3γ) is a member of
the 14-3-3 protein family, a family of highly conserved proteins
that regulate signal transduction by binding to
phosphoserine-containing proteins (11,12). YWHAG is upregulated in gastric
cancer (GC) tissue, and YWHAG knockdown has been shown to inhibit
proliferation, migration and invasion of GC cells, and to promote
apoptosis (13). Kim et al
(14) discovered that
overexpression of YWHAG promotes proliferation of breast cancer
cells. Moreover, YWHAG knockdown effectively inhibits
proliferation, migration and invasion of NSCLC (15). Results from Gene Expression
Profiling Interactive Analysis (GEPIA) suggested that TXNDC9 is
positively associated with YWHAG in LUAD; therefore, it was
hypothesized that TXNDC9 may inhibit LUAD progression by targeting
YWHAG.
The present study intended to clarify the effects of
TXNDC9 on the aggressive properties of LUAD cells and to
investigate the underlying mechanism.
Materials and methods
Cell culture, treatment and
transfection
Human type II alveolar epithelial cells (BEAS-2B;
BCRC 60074) and lung cancer cell lines (H1975, HCC827 and A549)
were purchased from the Food Industry Research and Development
Institute (Hsinchu, Taiwan). The cells were cultured in DMEM
(Gibco; Thermo Fisher Scientific, Inc.) containing 10% FBS (Gibco;
Thermo Fisher Scientific, Inc.), 100 U/ml penicillin and 100 µg/ml
streptomycin (Invitrogen; Thermo Fisher Scientific, Inc.) at 37°C
in a humidified atmosphere with 5% CO2. All cell lines
used were verified by STR through Applied Biosystems (Thermo Fisher
Scientific, Inc.).
For transfection, 20 µM small interfering
(si)RNA-negative control (NC; 5′-CACUGAUUUCAAAUGGUGCUAUU-3′),
overexpression (Oe) plasmid-NC, si-TXNDC9
(5′-TTTGGTAGTCTGAAGCAGC-3′) and Oe-YWHAG were obtained from
Shanghai GenePharma Co., Ltd. Cells were incubated with 5%
CO2 at 37°C and were used in subsequent experiments
after 48 h of transfection. Cell transfection was performed using
Lipofectamine 2000® Transfection Reagent (Invitrogen;
Thermo Fisher Scientific, Inc.) according to the manufacturer's
instructions.
Western blot analysis
Total protein extraction from A549 cells was
performed using RIPA lysis buffer (Beijing Solarbio Science &
Technology Co., Ltd.) and the protein concentrations were
quantified using a BCA kit (Beyotime Institute of Biotechnology).
Proteins (40 µg/lane) were separated by 12% SDS-PAGE and then
transferred onto PVDF membranes. After blocking with 5% non-fat
milk for 2 h at room temperature, the membranes were incubated
overnight at 4°C with the following primary antibodies (all
purchased from Abcam): Anti-TXNDC9 (1:5,000; cat. no. ab185959),
anti-YWHAG (1:1,000; cat. no. ab237732), anti-matrix
metalloproteinase (MMP)2 (1:1,000; cat. no. ab92536), anti-MMP9
(1:1,000; cat. no. ab76003), anti-BCL-2 associated X (Bax; 1:1,000;
cat. no. ab32503), anti-B-cell lymphoma-2 (Bcl-2; 1:1,000; cat. no.
ab32124) and anti-GAPDH (1:2,500; cat. no. ab9485). Following
primary incubation, membranes were incubated with goat anti-rabbit
horseradish peroxidase-conjugated IgG secondary antibody (1:5,000;
cat. no. ab6721; Abcam) at room temperature for 2 h. Protein bands
were visualized using enhanced chemiluminescence reagent (cat no.
P0018AS; Beyotime Institute of Biotechnology) and protein
expression levels were detected and semi-quantified using ImageJ
software (version 1.46; National Institutes of Health) with GAPDH
as the loading control.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA from was isolated from the BEAS-2B and
A549 cells using TRIzol® reagent and reverse transcribed
into cDNA using a SuperScript™ Double-Stranded cDNA
Synthesis kit (both Invitrogen; Thermo Fisher Scientific, Inc.).
qPCR for gene quantification was performed using SYBR Premix Ex Taq
(Takara Bio, Inc.) and an ABI 7500 Fast Real-Time PCR System
(Applied Biosystems; Thermo Fisher Scientific, Inc.) according to
the manufacturer's protocol. The following thermocycling conditions
were used for qPCR: 95°C for 10 min; followed by 40 cycles of 95°C
for 10 sec and 60°C for 60 sec. The following primers (purchased
from GenScript) were used for qPCR: TXNDC9 forward,
5′-GTGAAAATGTGGTTTGCCATT-3′ and reverse,
5′-TGCTTTTTCCACATTCAGCTT-3′; YWHAG forward,
5′-GGAGGGTCATCAGTAGCATTG-3′ and reverse, 5′-AGTTATCCAGCAGGCTCAGC-3′
and GAPDH forward, 5′-AGCCACATCGCTCAGACAC-3′ and reverse,
5′-GCCCAATACGACCAAATCC-3′. GAPDH served as the endogenous control
and the calculation of relative gene expression was determined
using 2−ΔΔCq method (16).
Cell Counting Kit-8 (CCK-8) assay
A549 cells (1×103 cells/well) were
inoculated into 96-well plates and incubated at 37°C for 24, 48 and
72 h. Each well was then supplemented with 10 µl CCK-8 reagent
(Beyotime Institute of Biotechnology) and incubated for a further 2
h. The absorbance of each well was detected at a wavelength of 50
nm, using a microplate reader (Bio-Rad Laboratories, Inc.).
Colony formation assay
A549 cells were resuspended in DMEM supplemented
with 10% FBS for colony formation assay; 5×102
cells/well were seeded in 6-well plates and incubated at 37°C with
5% CO2 for 14 days. Subsequently, cells were fixated
using 4% paraformaldehyde for 15 min at 37°C and stained with 0.5%
crystal violet solution for 30 min at room temperature. Finally,
colonies (>50 cells) were counted manually using an inverted
fluorescent microscope (Nikon Corporation; magnification,
×100).
Wound healing assay
A549 cells (1×105 cells/well) were
inoculated in 6-well plates and incubated at 37°C until the cell
confluency reached 90–100%. A linear scratch in the cell monolayer
was then made using a pipette tip. The cells were washed three
times with PBS to remove cellular debris and incubated at 37°C and
5% CO2. Images were captured by a light microscope at 0
and 24 h, and the area of migrated cells in the linear
scratch).
TUNEL assay
The effect of TXNDC9 silencing on A549 cell
apoptosis was detected using TUNEL detection solution (Beyotime
Institute of Biotechnology) according to the manufacturer's
protocol. In brief, A549 cells (1×106 cells/well) were
fixed with 4% paraformaldehyde for 15 min at room temperature and
permeabilized in 0.25% Triton X-100 for 20 min at room temperature.
Subsequently, after cells were rinsed with PBS, TdT solution and
dUTP solution were added and incubated at 37°C for 1 h in the dark.
Cells were treated with 10 µg/ml DAPI for nucleus staining for 5
min at 37°C and mounted in an anti-fade reagent (Beijing Solarbio
Science & Technology Co., Ltd.). In total, three fields of view
were selected at random and an inverted fluorescence microscope
(magnification, ×100; Olympus Corporation) was used to observe the
excitation and emission wavelengths at 450–500 and 515–565 nm,
respectively.
Co-immunoprecipitation (co-IP)
assay
STRING (string-db.org) and GEPIA
(gepia.cancer-pku.cn) databases revealed that TXNDC9 was positively
associated with YWHAG and bound to YWHAG. To verify this, co-IP
assays were performed. Total proteins from the A549 cells were
isolated using RIPA lysis buffer (Beijing Solarbio Science &
Technology Co., Ltd.) and quantified using BCA kit (Beyotime
Institute of Biotechnology). For immunoprecipitation, 500 µg
protein was incubated with 2 µg appropriate antibodies including
TXNDC9 (1:50; cat. no. ab185959; Abcam), YWHAG (1:200; cat. no.
ab237732; Abcam) and IgG (1:2,000; cat. no. A0208; Beyotime
Institute of Biotechnology) overnight at 4°C. Subsequently, 40 µl
Protein G/A agarose beads (Invitrogen; Thermo Fisher Scientific,
Inc.) were added to cell lysate and incubated for 2 h. After beads
were washed with PBS three times, precipitated proteins were
re-suspended in 2X SDS-PAGE loading buffer, boiled for 5 min and
eluted from the beads. Finally, western blot analysis was used to
measure the products from IP as aforementioned.
Bioinformatics tools
ENCORI database (https://starbase.sysu.edu.cn/panCancer.php) was used
to detect TNXDC9 expression in LUAD and to analyze the association
between TNXDC9 and the overall survival rate of LUAD patients.
GEPIA database (http://gepia.cancer-pku.cn/) was used to explore the
correlation between TNXDC9 and YWHAG in LUAD.
Statistical analysis
Data are presented as the mean ± SD. All experiments
were performed in triplicate. Statistical analysis was performed
using SPSS 20.0 software (IBM Corp.). One-way ANOVA was used to
perform statistical analysis followed by Tukey's multiple
comparisons post hoc test. The survival of LUAD patients was
subjected to Kaplan-Meier analysis. Correlation between TNXDC9 and
YWHAG was evaluated using Pearson's correlation analysis. P<0.05
was considered to indicate a statistically significant
difference.
Results
TXNDC9 is upregulated in LUAD
cells
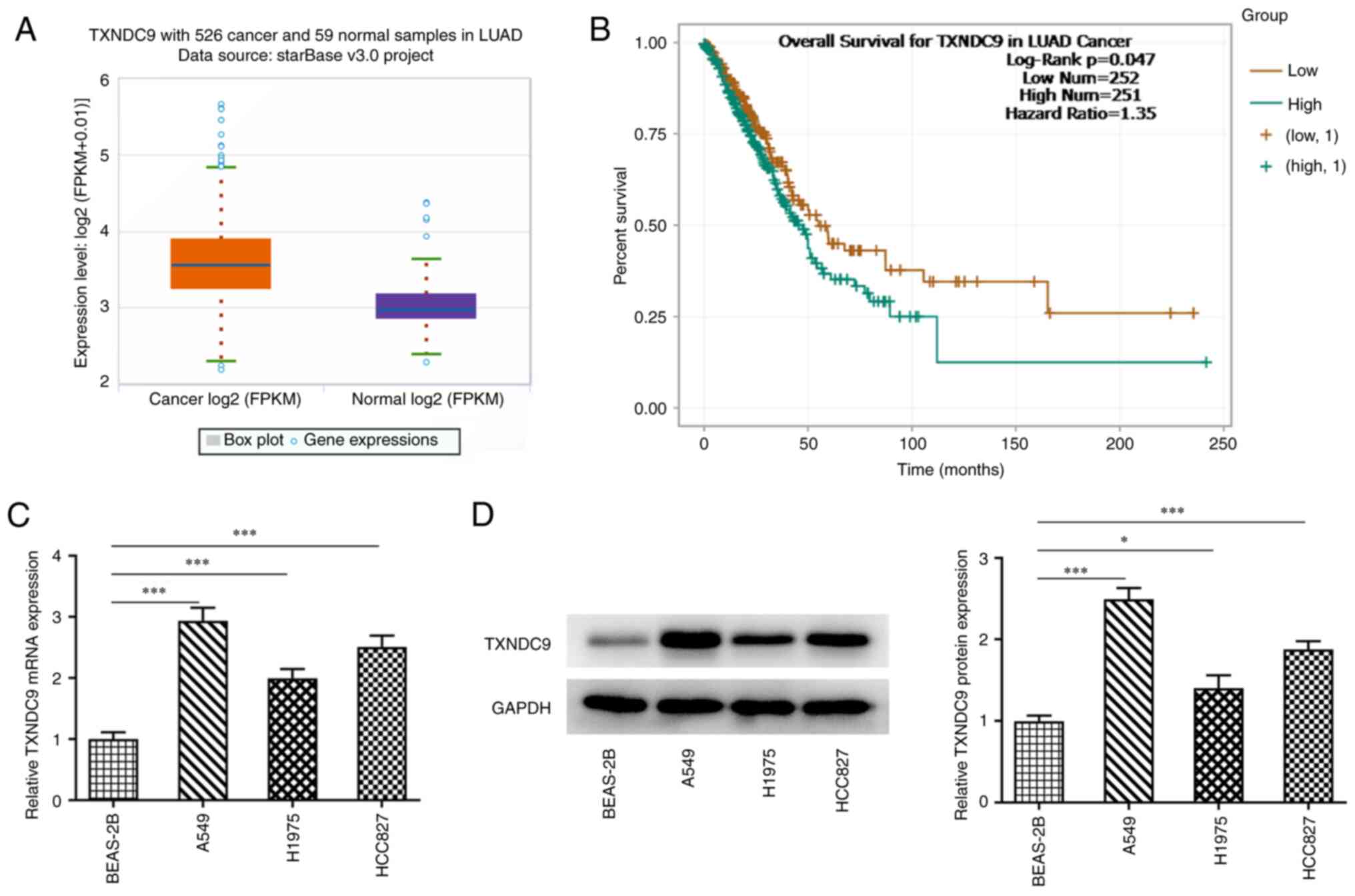
The results from ENCORI suggested that TXNDC9 was
markedly upregulated in LUAD cells (Fig. 1A). Additionally, poor survival of
patients with LUAD was observed in the high TXNDC9 expression group
compared with the low TXNDC9 expression group (Fig. 1B). The relative mRNA and protein
expression levels of TXNDC9 in normal lung epithelial (BEAS-2B0 and
lung cancer cells (A549, H1975 and HCC827 cells) were detected
using RT-qPCR and western blot analysis. Compared with BEAS-2B
normal epithelial cells, relative TXNDC9 mRNA and protein
expression in lung cancer cells was significantly upregulated,
particularly in A549 cells (Fig. 1C
and D, respectively). Therefore, A549 cells were selected for
use in subsequent experiments.
TXNDC9 knockdown inhibits viability
and proliferation of LUAD cells
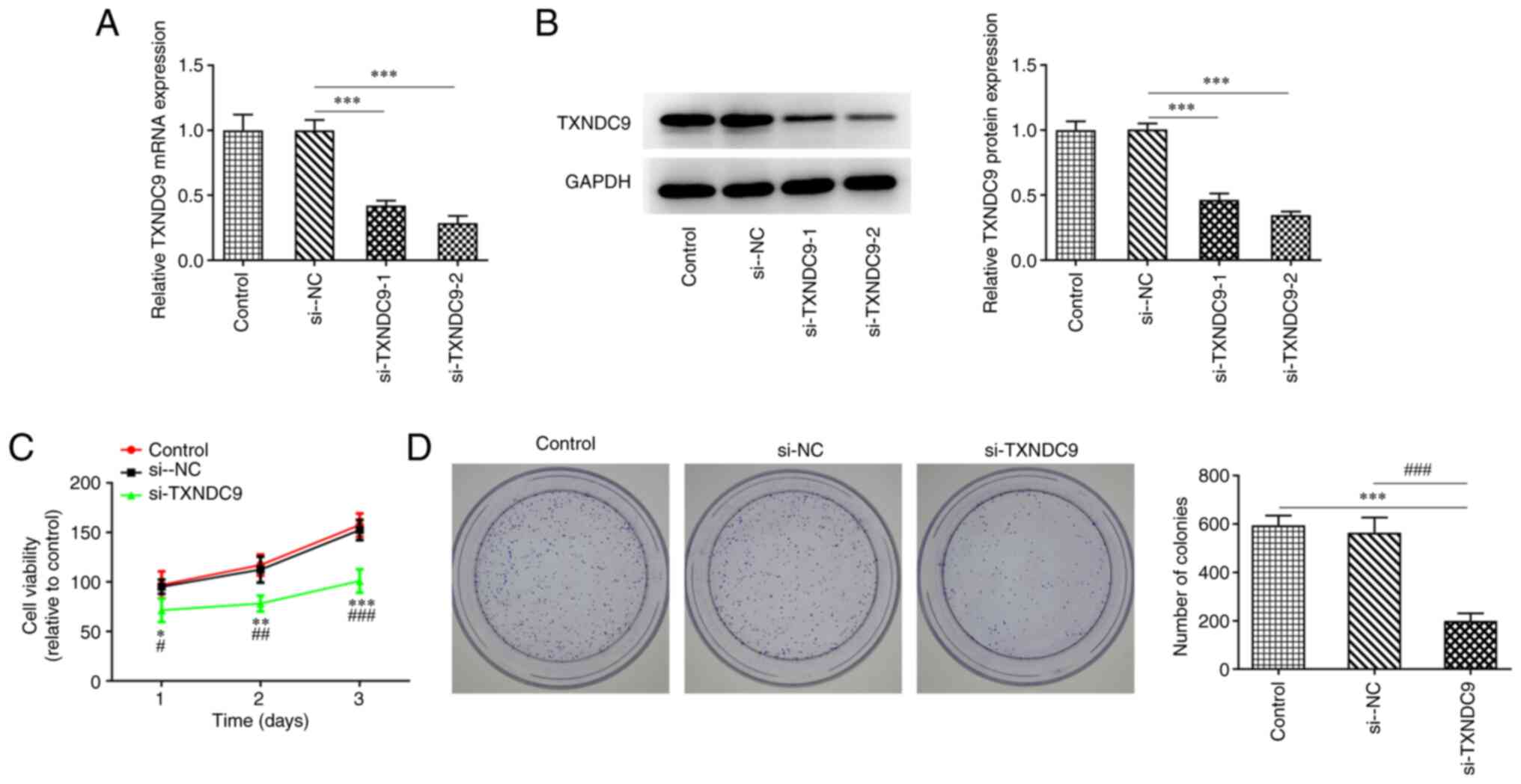
To investigate the effects of TXNDC9 on LUAD cells,
A549 cells were transfected with si-TXNDC9-1/2. mRNA and protein
expression levels of TXNDC9 were significantly decreased in the
TXNDC9-silenced A549 cells compared with the si-NC group (Fig. 2A and B). si-TXNDC9-2 was chosen
for the subsequent experiments as it displayed an improved
interference efficiency compared with si-TXNDC9-1. In addition,
CCK-8 and colony formation assay results demonstrated that the
viability and the proliferation, respectively, of A549 cells were
significantly diminished by TXNDC9 silencing compared with the
controls (Fig. 2C and D),
revealing that TXNDC9 knockdown exerted inhibitory effects on the
proliferation of LUAD cells.
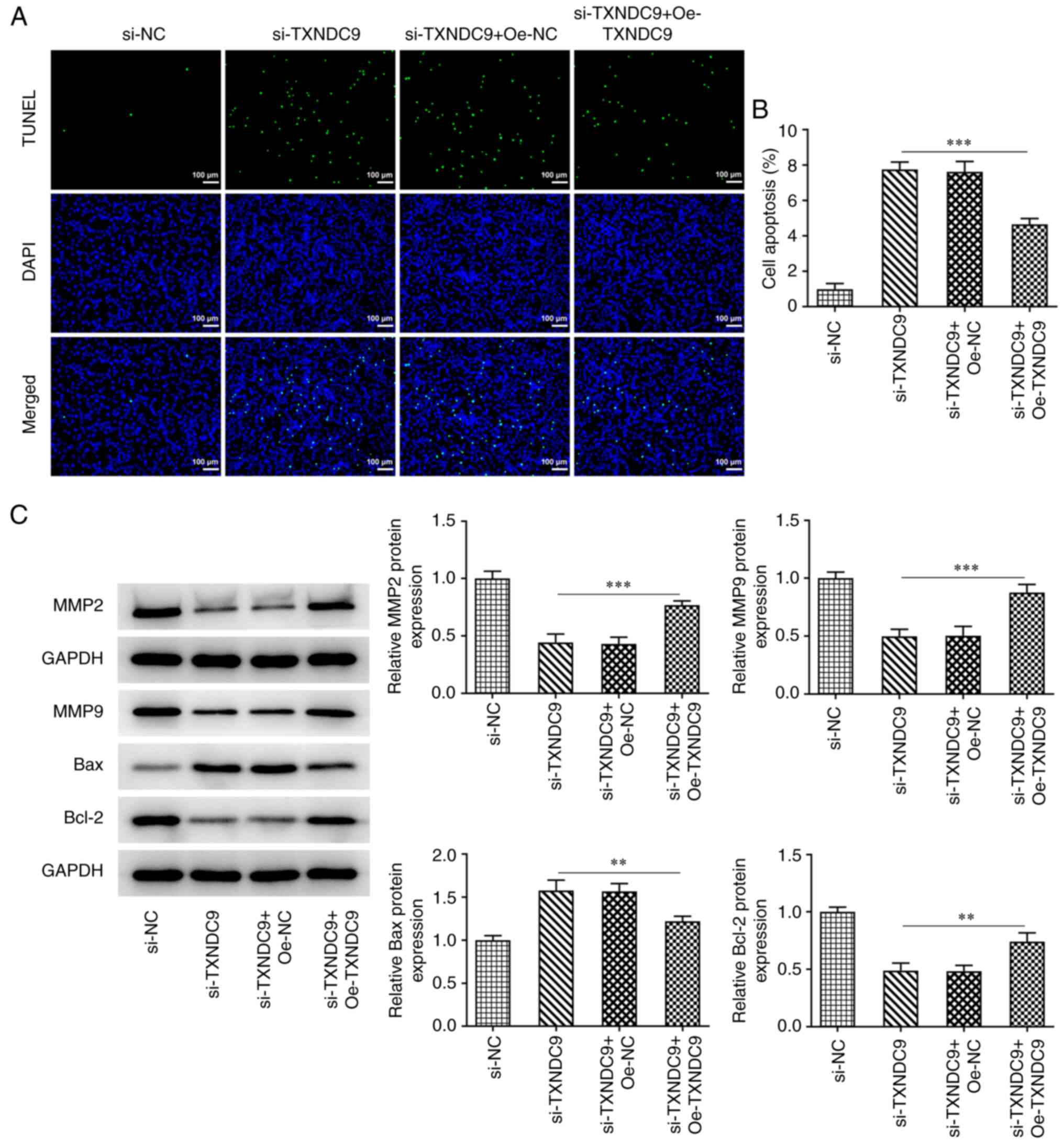
TXNDC9 knockdown inhibits migration
and invasion, and promotes apoptosis, of LUAD cells
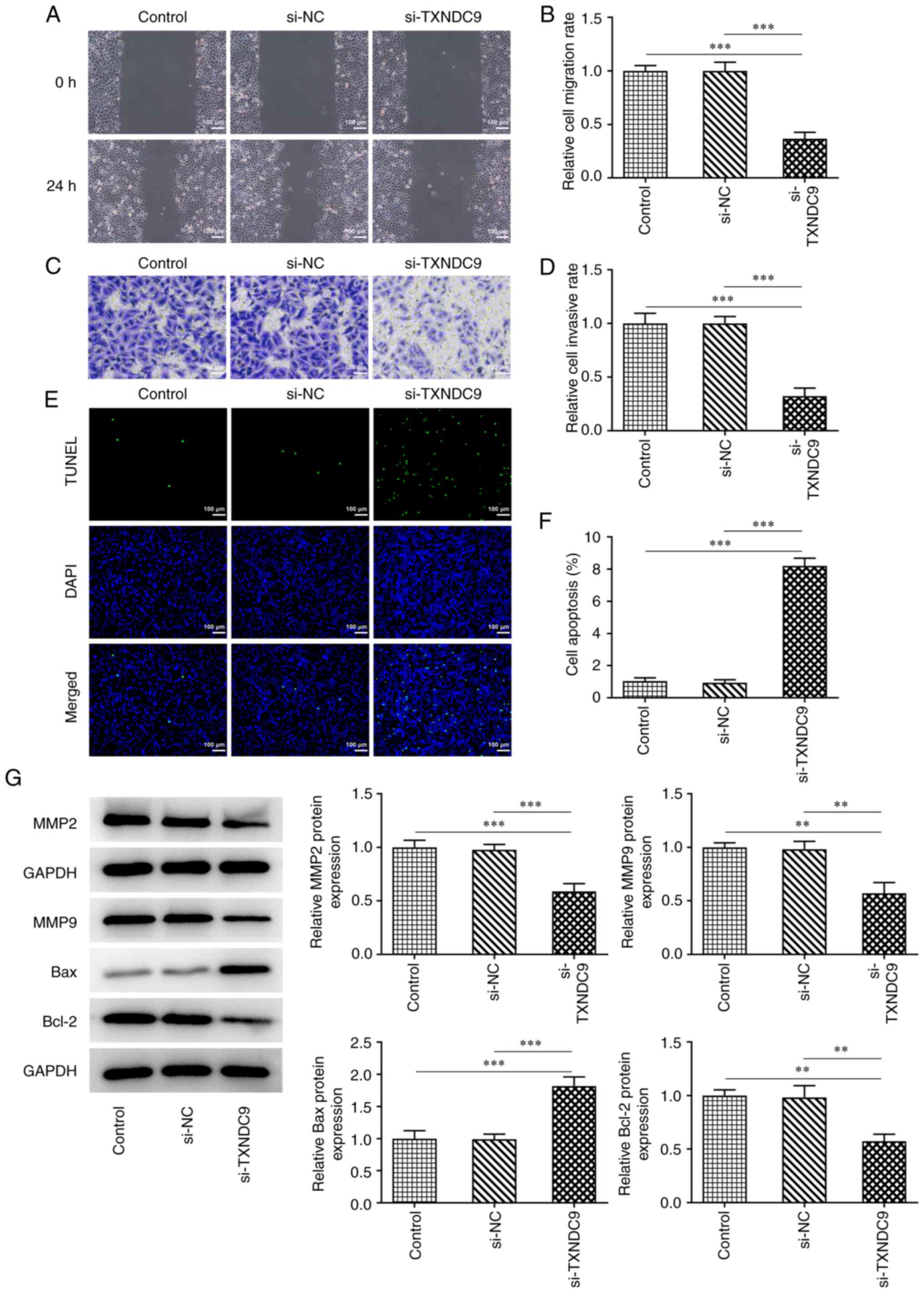
Wound healing and Transwell assays were used to
determine the relative migration rate and invasive ability. TXNDC9
silencing significantly suppressed the migration rate of A549 cells
compared with the si-NC group (Fig.
3A and B); TXNDC9 silencing similarly inhibited A549 cell
invasiveness (Fig. 3C and D).
Additionally, the effects of TXNDC9 knockdown on apoptosis of A549
cells was detected using TUNEL assay. TXNDC9 knockdown
significantly promoted apoptosis of A549 cells (Fig. 3E and F). Moreover, the protein
expression levels of matrix metalloproteinase (MMP)2, MMP9 and
Bcl-2 were significantly decreased by TXNDC9 knockdown, whereas
relative Bax protein expression was significantly increased
(Fig. 3G).
YWHAG is upregulated in LUAD and binds
to TXNDC9
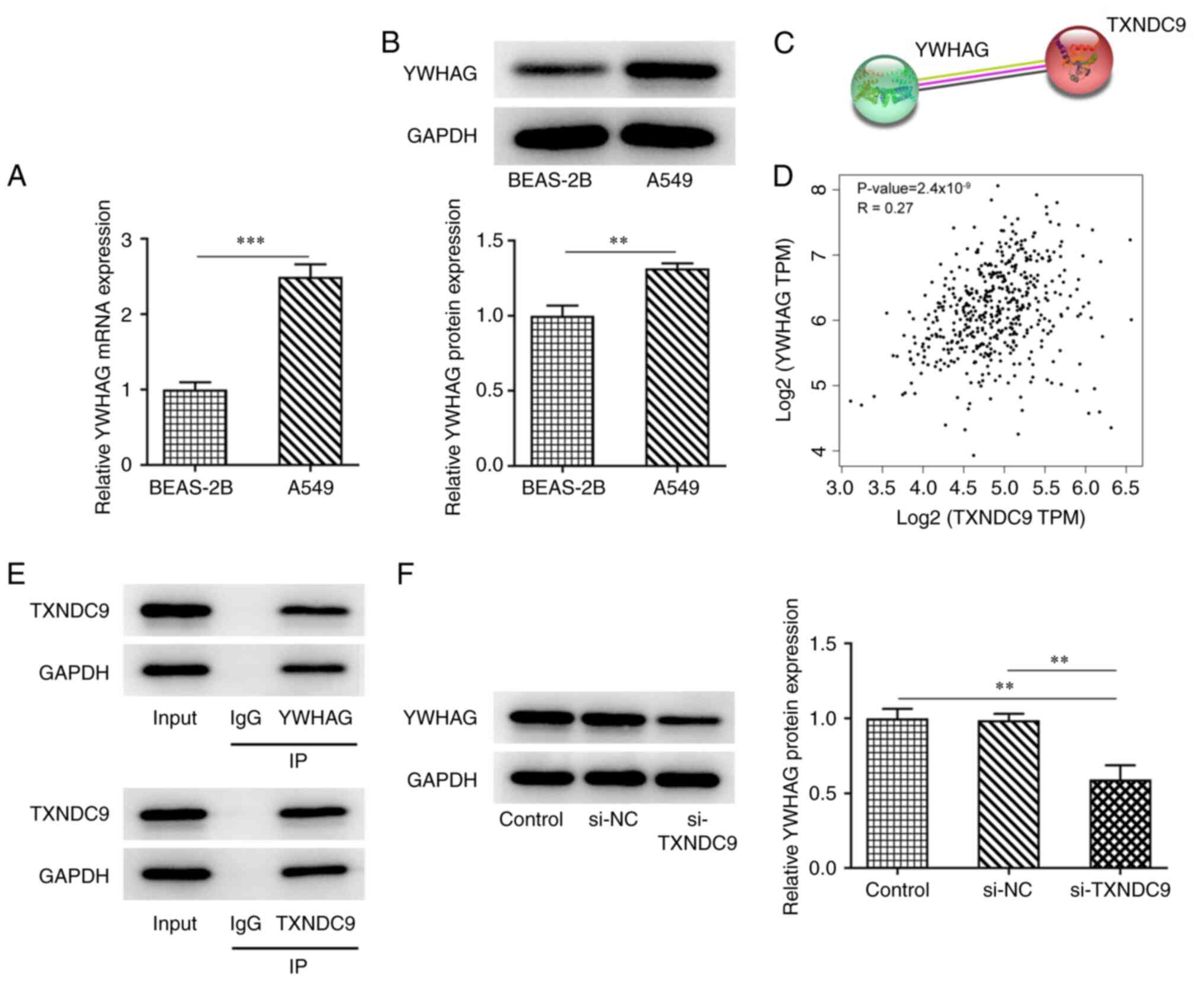
The mRNA and protein expression levels of YWHAG were
higher in A549 LUAD cells compared with BEAS-2B normal lung
epithelial cells (Fig. 4A and B).
According to STRING and GEIPA databases, YWHAG and TXNDC9 were
co-expressed and positively correlated with each other (Fig. 4C and D). Considering the positive
association between YWHAG and TXNDC9, co-IP assay was performed to
verify the binding between YWHAG and TXNDC9. TXNDC9 was observed
with anti-YWHAG and YWHAG was observed with anti-TXNDC9, revealing
that TXNDC9 bound to YWHAG (Fig.
4E). Compared with the si-NC group, the expression of YWHAG was
decreased in TXNDC9-silenced A549 cells (Fig. 4F).
YWHAG overexpression reverses the
inhibitory effect of TXNDC9 silencing on viability, proliferation,
migration and invasion of LUAD cells
A549 cells were transfected with Oe-YWHAG, and the
relative mRNA and protein expression of YWHAG were significantly
upregulated compared with the Oe-NC-transfected group (Fig. 5A and B). Decreased cell viability
and proliferation induced by TXNDC9 knockdown were partially
reversed by YWHAG overexpression (Fig. 5C-E), suggesting that
overexpression of YWHAG decreased the inhibitory effects of TXNDC9
silencing on A549 cell viability and proliferation. Furthermore,
compared with the si-TXNDC9 + Oe-NC group, the decrease in
migratory (Fig. 5F and G) and
invasive rates (Fig. 5H and I) of
the A549 cells were partially reversed following transfection with
YWHAG overexpression vector.
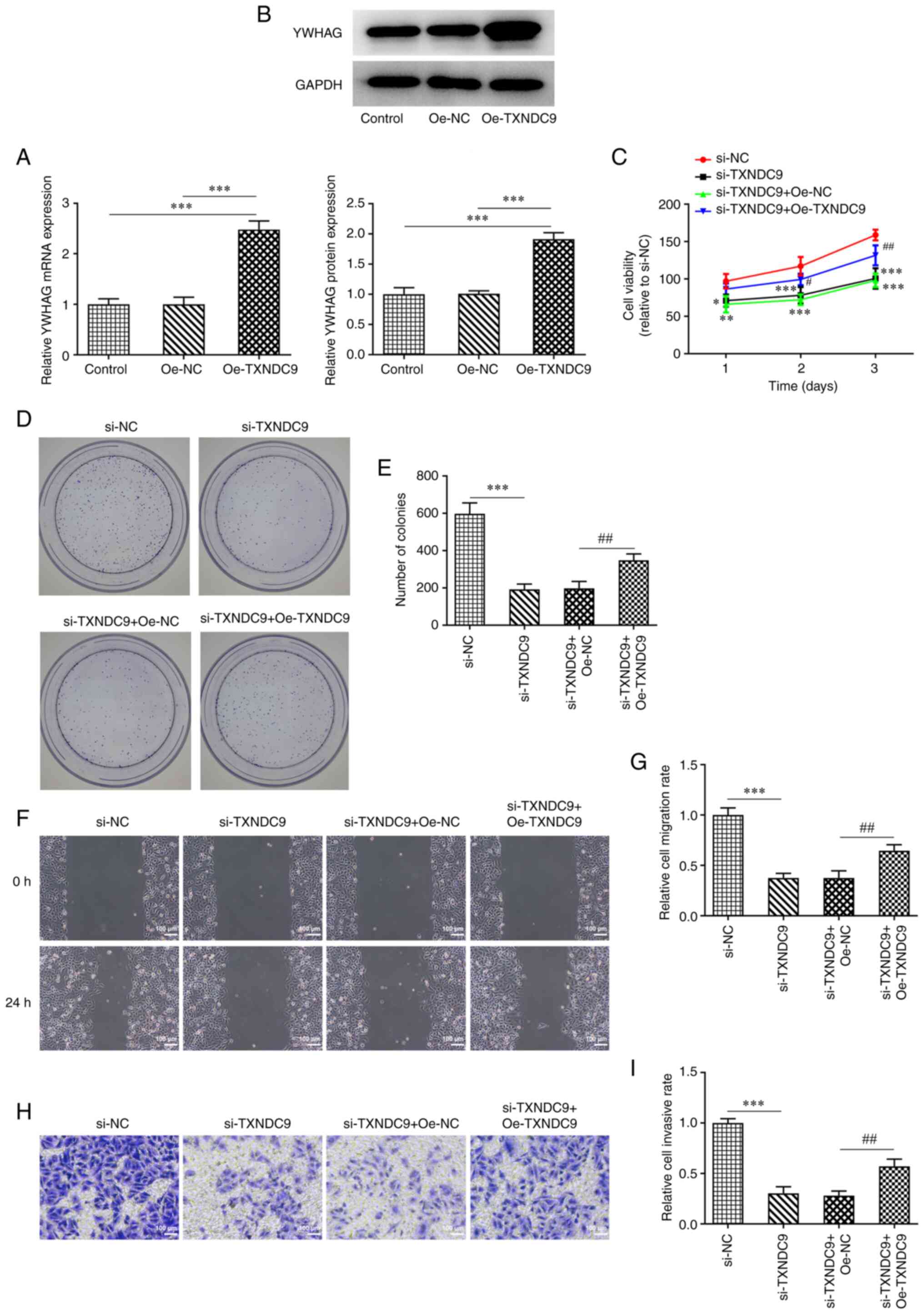 | Figure 5.YWHAG overexpression reverses the
inhibitory effect of TXNDC9 silencing on the viability,
proliferation, migration and invasiveness of lung adenocarcinoma
cells. (A) mRNA and (B) protein expression levels of YWHAG were
measured using reverse transcription-quantitative PCR and western
blotting, respectively. ***P<0.001 vs. Oe-NC or Control. (C)
Cell viability was detected using Cell Counting Kit-8 assay.
*P<0.05, **P<0.01, ***P<0.001 vs. si-NC;
#P<0.05 and ##P<0.01 vs. si-TXNDC9 +
Oe-NC. (D and E) Proliferation was detected using colony formation
assay. Magnification, ×4. ***P<0.001 vs. si-NC;
##P<0.01 vs. si-TXNDC9 + Oe-NC. (F and G) Wound
healing and (H and I) Transwell assay were used to detect migration
and invasiveness, respectively. ***P<0.001 vs. si-NC;
##P<0.01 vs. si-TXNDC9 + Oe-NC. NC, negative control;
Oe, overexpression; si, small interfering RNA; TXNDC9, thioredoxin
domain-containing protein 9; YWHAG, tyrosine
3-monooxygenase/tryptophan 5-monooxygenase activation protein
γ. |
YWHAG overexpression reverses the
promotive effects of TXNDC9 silencing on LUAD cell apoptosis
TUNEL assay was used to detect the effects of YWHAG
overexpression on the apoptosis of TXNDC9-silenced A549 cells. The
results demonstrated that the increased apoptosis observed
following TXNDC9 silencing was decreased by overexpression of YWHAG
compared with the si-TXNDC9 + Oe-NC group (Fig. 6A and B), which indicated that
YWHAG overexpression diminished the promotive effects of TXNDC9 on
cell apoptosis. Furthermore, the protein levels of
apoptosis-related factors including Bax and Bcl-2 and
metastasis-related factors including MMP2 and MMP9 were tested. It
was found that the downregulated expression levels of MMP2, MMP9
and Bcl-2 were upregulated by YWHAG overexpression, whereas
upregulated Bax expression was downregulated (Fig. 6C).
Discussion
Lung cancer poses a serious threat to human health
and is one of the most aggressive and lethal types of cancer
worldwide (17). The recorded
lung cancer mortality rate increased by 464.84% in the past 30
years in China (18). To date,
effective methods for treatment of LUAD are lacking (19). It is well documented that TRX
system protects against oxidative stress and inflammation in lung
diseases (20). In addition,
serum thioredoxin level has been discovered to be elevated in
on-small cell lung carcinoma patients (21).
Several studies have revealed that TXNDC9 has an
abnormal expression in various types of cancer (22,23). A previous study reported that
TXNDC9 expression is upregulated a serves a key role in CRC
(22). Expression of TXNDC9 is
also increased in breast cancer cells (24). Moreover, Chen et al
(10) observed that TXNDC9
promotes HCC progression, whereas TXNDC9 knockdown inhibits
proliferation of HCC cells. Consistent with these findings, in the
present study, TXNDC9 expression was demonstrated to be upregulated
in lung cancer cells compared with normal lung epithelial.
Moreover, TXNDC9 knockdown inhibited the viability and
proliferation of A549 cells in LUAD. Silencing of TXNDC9 abrogated
cell migration and invasion in LUAD, accompanied by downregulated
MMP2 and MMP9 protein levels. By contrast, TXNDC9 knockdown
promoted the apoptosis of A549 cells by increasing Bax expression
and decreasing Bcl-2 expression. These results suggested that
TXNDC9 knockdown inhibited LUAD progression, which implied that
TXNDC9 might primarily serve as an oncogene in multiple
malignancies, LUAD included.
YWHAG has been reported to be involved in various
biological processes, particularly in cancer (25). For example, an effective treatment
for skin cancer is targeting of the interaction of YWHAG with
CDC25A (26). YWHAG has been
shown to promote cell motility in breast cancer and inhibition of
YWHAG may serve as a novel therapeutic target for the treatment of
breast cancer (27). Qi et
al (28) demonstrated that
YWHAG serves a key role in regulating cell cycle progression and
its overexpression contributes to polyploidization in H322 lung
cancer cells. In the present study, the expression of YWHAG was
upregulated in lung cancer cells, which is consistent with a
previous study (15). The
elevated expression of YWHAG in certain types of human cancer
suggests that YWHAG may function as an oncogene (29). Therefore, YWHAG may represent an
effective therapeutic target for the treatment of lung cancer.
According to STRING and GEIPA, TXNDC9 was
co-expressed and was positively correlated with YWHAG; therefore,
further experiments were performed to confirm this prediction.
Considering that TXNDC9 was involved in LUAD progression, it was
hypothesized that TXNDC9 may target YWHAG to inhibit the viability,
proliferation, migration and invasiveness of lung cancer cells and
promote apoptosis, thus exerting inhibitory effects on LUAD
progression. YWHAG overexpression abolished the inhibitory effects
of TXNDC9 silencing on A549 cells. However, the present study only
conducted in vitro experiments, and in vivo
experiments in mice need to be performed. It is also important to
verify the interaction between TXNDC9 and YWHAG for screening other
lung cancer cell lines. Furthermore, the present study only
investigated the effect of TXNDC9 and YWHAG interaction on LUAD;
other mechanisms downstream of YWHAG need to be explored in the
future.
In conclusion, the present study demonstrated that
TXNDC9 and YWHAG were upregulated in LUAD, and that TXNDC9 bound to
YWHAG. TXNDC9 knockdown inhibited LUAD progression, and YWHAG
overexpression reversed these inhibitory effects.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
JW and XP conceived and designed the study. JL and
JZ performed the experiments. XP and JL analyzed the experimental
data. JW and JZ wrote and revised the manuscript. XP and JL confirm
the authenticity of all the raw data. All authors have read and
approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Gridelli C, Rossi A, Carbone DP, Guarize
J, Karachaliou N, Mok T, Petrella F, Spaggiari L and Rosell R:
Non-small-cell lung cancer. Nat Rev Dis Primers. 1:150092015.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Song Q, Shang J, Yang Z, Zhang L, Zhang C,
Chen J and Wu X: Identification of an immune signature predicting
prognosis risk of patients in lung adenocarcinoma. J Transl Med.
17:702019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Travis WD: Lung cancer pathology: Current
concepts. Clin Chest Med. 41:67–85. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ma F, Hou L and Yang L: Txndc9 is required
for meiotic maturation of mouse oocytes. Biomed Res Int.
2017:62658902017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Kang CH, Park JH, Lee ES, Paeng SK, Chae
HB, Hong JC and Lee SY: Redox-dependent structural modification of
nucleoredoxin triggers defense responses against alternaria
brassicicola in arabidopsis. Int J Mol Sci. 21:91962020. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Feng T, Zhao R, Sun F, Lu Q, Wang X, Hu J,
Wang S, Gao L, Zhou Q, Xiong X, et al: TXNDC9 regulates oxidative
stress-induced androgen receptor signaling to promote prostate
cancer progression. Oncogene. 39:356–367. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lu A, Wangpu X, Han D, Feng H, Zhao J, Ma
J, Qu S, Chen X, Liu B and Zheng M: TXNDC9 expression in colorectal
cancer cells and its influence on colorectal cancer prognosis.
Cancer Invest. 30:721–726. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Chen D, Zou J, Zhao Z, Tang X, Deng Z, Jia
J and Liu S: TXNDC9 promotes hepatocellular carcinoma progression
by positive regulation of MYC-mediated transcriptional network.
Cell Death Dis. 9:11102018. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Aitken A: 14-3-3 proteins: A historic
overview. Semin Cancer Biol. 16:162–172. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cho E and Park JY: Emerging roles of
14-3-3γ in the brain disorder. BMB Rep. BMB Rep. 53:500–511. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ni J, Wang J, Fu Y, Yan C, Zhu M, Jiang Y,
Chen J, Ding Y, Fan X, Li G and Jin G: Functional genetic variants
in centrosome-related genes CEP72 and YWHAG confer susceptibility
to gastric cancer. Arch Toxicol. 94:2861–2872. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kim JO, Kim SR, Lim KH, Kim JH, Ajjappala
B, Lee HJ, Choi JI and Baek KH: Deubiquitinating enzyme USP37
regulating oncogenic function of 14-3-3γ. Oncotarget.
6:36551–36576. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang P, Deng Y and Fu X: MiR-509-5p
suppresses the proliferation, migration, and invasion of non-small
cell lung cancer by targeting YWHAG. Biochem Biophys Res Commun.
482:935–941. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Denisenko TV, Budkevich IN and Zhivotovsky
B: Cell death-based treatment of lung adenocarcinoma. Cell Death
Dis. 9:1172018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
She J, Yang P, Hong Q and Bai C: Lung
cancer in China: Challenges and interventions. Chest.
143:1117–1126. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhang C, Zhang Z, Zhang G, Zhang Z, Luo Y,
Wang F, Wang S, Che Y, Zeng Q, Sun N and He J: Clinical
significance and inflammatory landscapes of a novel
recurrence-associated immune signature in early-stage lung
adenocarcinoma. Cancer Lett. 479:31–41. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Xu J, Li T, Wu H and Xu T: Role of
thioredoxin in lung disease. Pulm Pharmacol Ther. 25:154–162. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Fan J, Yu H, Lv Y and Yin L: Diagnostic
and prognostic value of serum thioredoxin and DJ-1 in non-small
cell lung carcinoma patients. Tumour Biol. 37:1949–1958. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhou W, Fang C, Zhang L, Wang Q, Li D and
Zhu D: Thioredoxin domain-containing protein 9 (TXNDC9) contributes
to oxaliplatin resistance through regulation of autophagy-apoptosis
in colorectal adenocarcinoma. Biochem Biophys Res Commun.
524:582–588. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wu Y, Ye H, Peng B, Jiang H, Tang Q, Liu
Y, Xi J and Chen S: MiR-643 functions as a potential tumor
suppressor in gastric cancer by inhibiting cell proliferation and
invasion via targeting TXNDC9. Ann Clin Lab Sci. 51:494–502.
2021.PubMed/NCBI
|
|
24
|
Garcia SA and Nagai MA: Transcriptional
regulation of bidirectional gene pairs by 17-β-estradiol in MCF-7
breast cancer cells. Braz J Med Biol Res. 44:112–122. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Raungrut P, Wongkotsila A,
Lirdprapamongkol K, Svasti J, Geater SL, Phukaoloun M, Suwiwat S
and Thongsuksai P: Prognostic significance of 14-3-3γ
overexpression in advanced non-small cell lung cancer. Asian Pac J
Cancer Prev. 15:3513–3518. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Holmes TR, Al-Matouq J, Holmes M, Nicola
L, Rudd JC, Lovas S and Hansen LA: Targeting 14-3-3ε-CDC25A
interactions to trigger apoptotic cell death in skin cancer.
Oncotarget. 11:3267–3278. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hiraoka E, Mimae T, Ito M, Kadoya T,
Miyata Y, Ito A and Okada M: Correction to: Breast cancer cell
motility is promoted by 14-3-3γ. Breast Cancer. 26:5942019.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Qi W, Liu X, Chen W, Li Q and Martinez JD:
Overexpression of 14-3-3gamma causes polyploidization in H322 lung
cancer cells. Mol Carcinog. 46:847–856. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Qi W, Liu X, Qiao D and Martinez JD:
Isoform-specific expression of 14-3-3 proteins in human lung cancer
tissues. Int J Cancer. 113:359–363. 2005. View Article : Google Scholar : PubMed/NCBI
|