Introduction
Glucocorticoids have been widely used for the
treatment of various pathological conditions, such as allergies,
inflammatory conditions, malignancies and immunological diseases.
Steroid-induced avascular necrosis of the femoral head can be
triggered following steroid-pulse therapy and long-term use of
glucocorticoids (1,2). It has been found that
steroid-induced avascular necrosis of the femoral head is the most
common type of non-traumatic osteonecrosis of the femoral head
(3). It is estimated that
75,000-150,000 new cases of ONFH are diagnosed every year in China
(4). Reduced bone mass and
trabecular fracture may occur after the third stage of ONFH as a
result of cell death in the femoral head (5). Femoral head collapse and
osteoarthritis eventually require artificial joint replacement to
improve patient quality of life, which causes a heavy economic
burden for society. Therefore, early detection of ONFH and
protection of the femoral head are necessary for patient quality of
life. However, as the molecular mechanism underlying the cause of
ONFH is not entirely clear, effective treatment options are still
lacking. The mechanism may be connected with abnormal lipid
distribution, which can induce the formation of microemboli in the
arteries supplying the femoral head (6). Additionally, apoptosis and autophagy
of osteocytes are regulated by glucocorticoids (5,7)
which have multiple effects on bone formation through lipid
metabolism (8). Previous studies
have demonstrated that, following inhibition of the Wnt/β-catenin
pathway, peroxisome proliferator-activated receptor levels are
increased by sclerostin production, causing pluripotent precursor
cells to differentiate into adipocytes, rather than osteoblasts
(9,10). Thus, modulating lipid metabolism
disorders and improving bone mass may be critical factors in the
prevention of ONFH.
Bone marrow-derived mesenchymal stem cells (BMSCs)
are primitive cells that have the potential to differentiate into
adipocytes, chondrocytes, osteoblasts and osteocytes (11,12). Bone resorption and bone formation
are balanced by osteoclasts and osteoblasts in order to maintain
bone mass homeostasis (13).
Matsuya et al (14) found
that autologous BMSC injection into the femoral head can slow down
the progression of ONFH. Lipid metabolism and osteogenic
differentiation are also strongly associated with BMSC function
(15). Another study has
confirmed that glucocorticoids can inhibit the osteogenic
differentiation and proliferation of BMSCs (16), which can reduce BMSC numbers
(16). In addition, in the
long-term, BMSC metabolism and differentiation are reduced as a
result of an imbalance in bone mass homeostasis, thus increasing
bone fragility (17). Tian and Yu
(15) have suggested that the
proliferation of BMSCs is impaired in ONFH. The results of the
aforementioned studies suggest that improved understanding of the
mechanisms underlying BMSC proliferation and differentiation may
provide insight into the pathogenesis of ONFH.
MicroRNA (miRNA/miR) is a type of small non-coding
single-stranded RNA molecule which binds to the 3′-untranslated
region (UTR) of target genes (18). Sedwick and Ambros were the first
to propose that >30% of human genes could be regulated by miRNA
(19). It has also been reported
that miRNA is involved in the regulation of BMSC differentiation
and proliferation (14,16). Moreover, miR-141 has been shown to
inhibit proliferation and migration in several tumor cell types,
such as osteosarcoma cells, SW480 colorectal cancer cells and
hepatocellular carcinoma (HCC) cells (20–22). Overexpression of miR-141 inhibits
the proliferation, migration and invasion of HCC cells (21). miR-141 is also associated with
tissue repair and osteogenic differentiation (23). Indeed, miR-141 inhibits the
proliferation of BMSCs; inhibition of miR-141 can promote the
proliferation of BMSCs (24) and
their osteogenic differentiation (5), possibly by targeting vitamin C
transporter 2 (25). Therefore,
the inhibition of miR-141 could be harnessed for the prevention and
treatment of osteonecrosis caused by glucocorticoids.
E2F transcription factor 3 (E2F3) is a member of the
E2F transcription factor family, which is involved in the
regulation of cell proliferation (26). E2F3 has been identified as the
target of miR-141 in HCC cells, and the overexpression of E2F3 can
partially reverse the tumor-suppressive effects of miR-141
(27). It has been demonstrated
that E2f3+/− Sprague-Dawley rat embryos develop normally
without fatal disease. However, weight loss, delayed growth and
skeletal dysplasia were observed, indicating that E2F3 played a key
role in muscle and skeletal development (28). However, how E2F3 functions in
different tissues and environments remains unclear.
To the best of our knowledge, few studies have
evaluated the role of E2F3 in osteogenic differentiation. It may be
hypothesized that E2F3 could promote osteogenic differentiation of
BMSCs and that miR-141 could inhibit the process by targeting this
transcription factor. A previous study has indicated that E2F3 may
be the target of microRNA (27).
The aim of the present study was to evaluate the role of miR-141
and E2F3 in BMSC proliferation in ONFH. In addition, the role of
miR-141 and E2F3 in the femoral head bone tissue changes induced by
glucocorticoids was also examined.
Materials and methods
Isolation of rat BMSCs
After 7 days adaptive feeding (26°C, 60% humidity,
12 h of light and 12 h of darkness every day, 200 ml water per day,
free access to food), animal health and behavior were monitored
every day. In the present study, two 4-week-old male Sprague Dawley
rats (SCXK2017-0001) were used (29). Their weights were 70.2 and 70.3 g,
respectively. The rats were anesthetized by intraperitoneal
injection of pentobarbital sodium (3% solution for a dose of 40
mg/kg), then sacrificed by cervical dislocation. The rats were
immersed in 75% ethanol for about 5–10 min, and dissection was
carried out under sterile conditions. The bilateral femurs and
tibia were removed with sterile surgical instruments, and the
attached muscles were removed. Three holes were made in the femur
and tibia, and a 10-ml syringe was used to extract medium inserted
into the femurs and tibia. The cells in the bone marrow were
repeatedly flushed into a Petri dish until the femur and tibia were
pale.
The cell suspensions were collected, purified and
centrifuged at 100 × g for 5 min at room temperature, then
re-suspended in α-MEM (HyClone; Cytiva) and incubated at 37°C with
5% CO2. When the cells reached 80–90% confluence, the
culture medium was removed. The cells were trypsinized and washed
in PBS (HyClone; Cytiva). Trypsin (HyClone; Cytiva) was used for
digestion for 3 min. The cells were then sub-cultured at a 1 in 3
ratio.
Transfection and lentiviral
transduction
Transfection was used for luciferase assay.miR-141
mimics were purchased from Shanghai Gemma Pharmaceutical.
Lipofectamine® 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.) was used to transfect BMSCs according to the
manufacturer's instructions. A total of 1 µg plasmid (100 nM miRNA
mimic) was diluted with 50 µl serum-free Opti-MEM. The plasmid was
mixed gently and incubated at room temperature for 5 min. B.
Lipo3000 was diluted with 50 µl serum-free OPti-MEM, mix gently and
incubate at room temperature for 5 min. MiR-141 mimic and lipo3000
were mixed and stood at room temperature for 20 min before
transfection into cells. Subsequent luciferase assay was performed
following transfection for 48 h.
The sequences used were as follows: NC,
5′-UUCUCCGAACGUGUCACGUTT-3′ and 5′-ACGUGACACGUUCGGAGAATT-3′;
miR-141 mimic.
Lentiviral transduction was used in subsequent
experiments including alkaline phosphatase stain and mRNA and
protein expression of E2F3The lentivirus was packaged by Shanghai
GenePharma, using a 3rd-generation lentiviral packaging system. For
lentivirus production, the packaging plasmids (pGag/Pol, pRev,
pVSV-G) and shuttle plasmid pLV2 were transfected into 293T cells
(Genepharma, Shanghai). The supernatant of the 293T cells was
collected by centrifugation at 1,000 g for 4 h at 4°C, then
transferred to a syringe and filtered using a 0.45-µm filter. The
plasmid was mixed with 1.5 ml serum-free DMEM medium and stood at
room temperature for 20 min before transfection. Plasmid ratio 8 µg
LV2 shuttle plasmid, 8 µg pGag/Pol, 4 µg pRev, 6 µg PVSV-g. 300 µl
liposome was mixed in 1.2 ml serum-free medium, and left at room
temperature for 5 min. Plasmid and liposome were mixed and stood at
room temperature for 20 min. Then add into 293T cell culture
medium. After 6 h incubation, medium was removed, and the medium
containing serum was added, and the supernatant was collected after
72 h culture at 37°C The filtrate was centrifuged at 23,000 g for 4
h at 4°C. The plates were shaken evenly, then placed into the
incubator for culture. Positive cells were screened with 0.5 µg/ml
puromycin and maintained at a concentration of 2 µg/ml After 6 h of
cell culture, the medium containing the virus was replaced with
fresh medium for another 72 h before subsequent experiments (cell
proliferation assays and mRNA analysis).
Cell osteogenesis induction
BMSCs were inoculated at a density of
1×105 cells and infected with lentivirus. BMSCs were
collected from the control and the ONFH rat. Cell osteogenesis
induction was carried out as previously reported by Yaghoobi et
al (30). The BMSCs were
digested with trypsin (HyClone; Cytiva) when they reached 80–90%
confluence, then centrifuged at 100 × g for 5 min at room
temperature. The supernatant was removed, and α-MEM (HyClone;
Cytiva) was added to resuspended the BMSCs in a 24-well culture
plate. The BMSCs were cultured to 70% confluence, then the culture
medium was removed and replaced with osteogenic induction culture
medium (Cyagen; cat. no. RASMX-90021). The medium was changed every
3 days thereafter. The osteogenesis levels were examined at day 3,
7 and 14 using alkaline phosphatase staining (31). Images were taken and saved for ALP
assay.
Alkaline phosphatase staining
A total of 1×105 cells were inoculated in
24-well plates. The cells were cultured to 60–70% confluence, then
the culture medium was replaced with osteogenic induction culture
medium. The osteogenic medium was changed every 3 days, and the
cells were washed with PBS at the end of the osteogenic induction
process. On days 3, 7 and 14, the BMSCs were fixed with 10%
formaldehyde solution at room temperature for 30 min, then washed
with PBS twice. The fixed cells were then stained with 50 µl
alkaline phosphatase solution at 37°C for 2 h (32).
Luciferase assays
For luciferase assays, the pGL6-mir reporter plasmid
(Beyotime Institute of Biotechnology; cat. no. D2106) and the
Renilla prL-TK plasmid (Promega; cat. no. E2241) were
transfected using Lipofectamine 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.; cat. no. L30000015). The 3′-UTR of E2F3 was
amplified by PCR as previously described (33), then cloned into the pGL6-mir
reporter vector (Beyotime Institute of Biotechnology; cat. no.
D2106). BMSCs were co-transfected with the wild-type or mutant
pGL6-mir reporter vector (100 ng/well), and either miR-141 mimic or
NC (100 ng/well). The cells were lysed 48 h post-transfection in
order to measure luciferase activity (34) using the Dual Luciferase Reporter
Assay System (Promega Corporation). Renilla luciferase
activity was used for normalization.
RT-qPCR
Total RNA from cells was isolated using
TRIzol® (Invitrogen; Thermo Fisher Scientific, Inc.)
according to the manufacturer's instructions. cDNA was reverse
transcribed using Prime Script RT Reagent kit at 42°C for 60 min
and 70°C for 10 min (Takara Biotechnology Co., Ltd.). qPCR was
carried out using SYBR Premix Ex Taq (Takara Biotechnology Co.,
Ltd.) on an ABI StepOne Plus Real-Time PCR System (Applied
Biosystems; Thermo Fisher Scientific, Inc.). The thermocycling
protocol consisted of an initial denaturation for 5 min at 95°C
followed by 40 cycles at 95°C for 10 sec and 60°C for 34 sec
(35–37). The relative mRNA levels were
obtained using the 2−ΔΔCt method (38). Reverse transcription of miRNA and
quantitative PCR primers were purchased from Guangzhou Ribobio Co.,
Ltd. The experiments were repeated five times. The reference gene
for miR-141 was U6 and the reference gene for E2F3 was GAPDH.
Primers (General Biosystems) were as follows: E2F3 forward,
5′-CGAGAGTGGCCATCAGTACC-3′ and reverse, 5′-CCTCTTCTGCACCTTGAGCA-3′
and GAPDH F-5′ TTCACCACCATGGAGAAGGC 3′ R-5′ AGTGATGGCATGGACTGTG
3′.
Western blot analysis of E2F3 protein
expression
At day 3, 7 and 14, after discarding the RASMX-90021
medium (Cyagen; cat. no. RASMX-90021) and α-MEM (HyClone; Cytiva;
SH30265) the cells were lysed in RIPA lysis buffer at 4°C
overnight. The total protein was harvested via centrifugation at
13,600 g and the protein concentration was determined using the BCA
method (Beyotime Institute of Biotechnology; cat. no. P0010). RIPA
buffer included 50 mmol/l Tris-HCl, pH8.0, 150 mmol/L NaCl, 1%
TritonX-100, 100 µg/ml PMSF.: A total of 30 µg protein per lane was
separated using SDS-PAGE on 8% gels, then transferred to a PVDF
membrane (MilliporeSigma; cat. no. ISEQ00010). The membrane was
then blocked with 5% skim milk in 0.1% Tween-20 Tris-buffered
saline at room temperature for 1 h. The membranes were incubated
with anti-E2F3 (ProteinTech Group, Inc.; cat. no. 12344-1-AP;
diluted 1:1,000) or rabbit anti-GAPDH (Abbkine Scientific Co.,
Ltd.; cat. no. A01020; diluted 1:5,000) antibodies at 4°C
overnight. This was followed by incubation with HRP-conjugated
anti-mouse (Beyotime Institute of Biotechnology; cat. no. A0216;
diluted at 1:5,000) or anti-rabbit (Beyotime Institute of
Biotechnology; cat. no. A0208; diluted 1:5,000) at room temperature
for 2 h. The bands were visualized using ECL reagent (Tanon Science
and Technology Co., Ltd.; cat. no. 180-5001) (39). GAPDH was used as loading control.
Chemiluminescence instrument (Tanon, 5200) was used for
densitometry. The software for densitometry was GelCap 1.0
(Tanon).
Statistical analysis
Each experiment was performed three times, and the
results are shown as the mean ± standard deviation. SPSS 23.0
software (SPSS, Inc.) was used for statistical analysis. The
differences between two groups were compared by unpaired Student's
t test. The differences between three or more groups were compared
using one-way analysis of variance followed by Tukey's post hoc
test. P<0.05 was considered to indicate a statistically
significant difference.
Results
Identification of BMSCs and osteogenic
induction
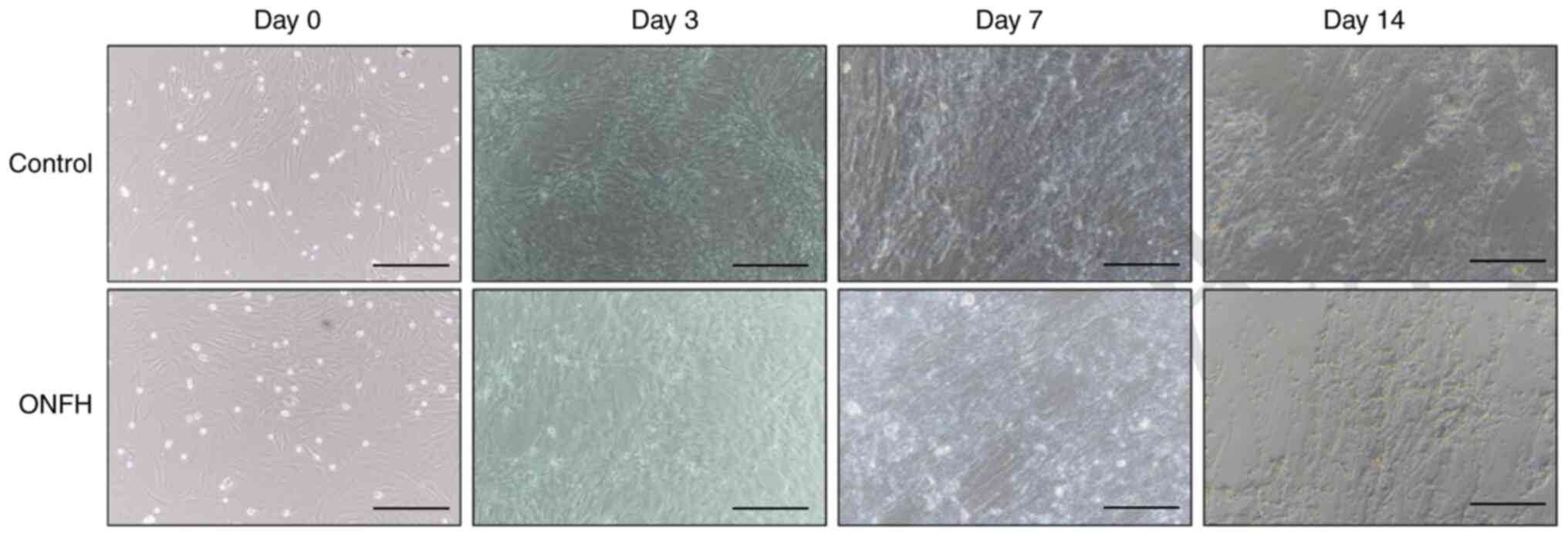
The morphology of BMSCs at day 0 under an optical
microscope is shown in Fig. 1.
BMSCs were round in shape, with strong refraction, and varied in
size. In addition, osteogenesis was examined at days 3, 7 and 14.
Osteogenesis levels were higher in the control than in the ONFH
group, as evidenced by cell density and osteogenic capability
(Fig. 1).
Alkaline phosphatase staining
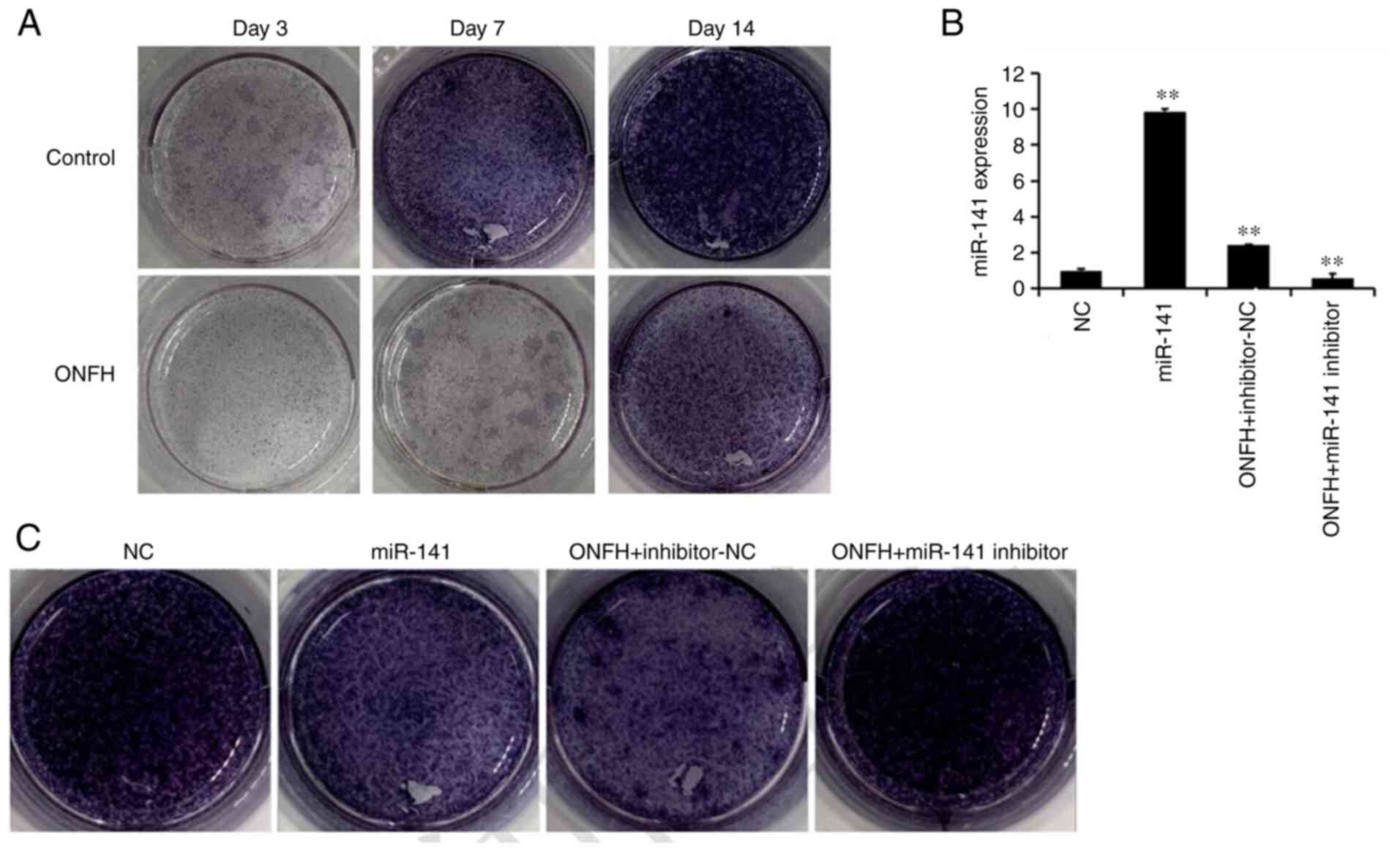
Alkaline phosphatase staining is shown in Fig. 2. The results suggested that the
control group was markedly darker than the ONFH group at days 3, 7
and 14 days (Fig. 2A). BMSCs from
the normal rat were then transduced with, miR-141 mimic lentivirus
(miR-141), whereas those of the ONFH rat were transduced with
miR-141 inhibitor lentivirus. Following transduction with the
miR-141-3p mimic lentivirus, alkaline phosphatase staining in BMSCs
from the normal was reduced compared with the NC. Following
transduction with the miR-141 inhibitor lentivirus, alkaline
phosphatase staining in BMSCs from the ONFH rat was increased
compared with the inhibitor NC. These results suggest that miR-141
expression is associated with osteogenesis.
Association between E2F3 and
miR-141
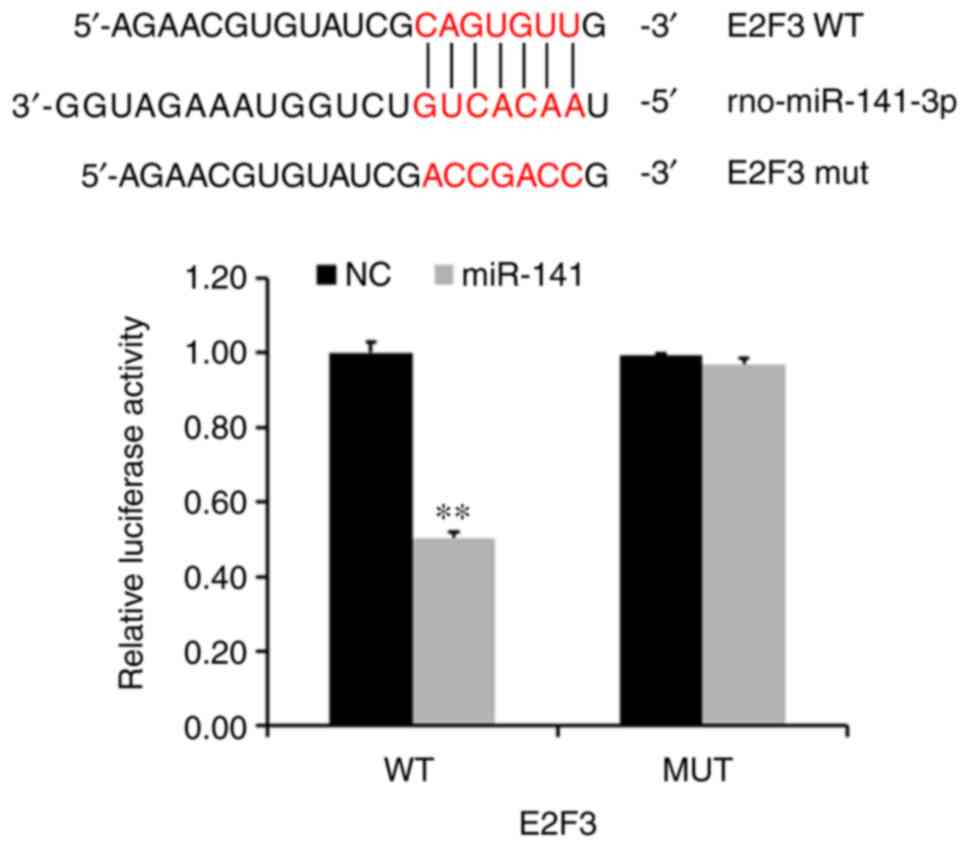
In order to examine the regulatory relationship
between E2F3 and miR-141, luciferase reporter assays were used to
determine whether miR-141 could target E2F3 expression directly
through its UTR. In the wild-type group, the average E2F3
luciferase activity was significantly lower than that of the NC
group (P<0.005; Fig. 3).
However, average E2F3 luciferase activity in the mutant remained
unchanged following transduction the miR-141-3p overexpression
lentivirus.
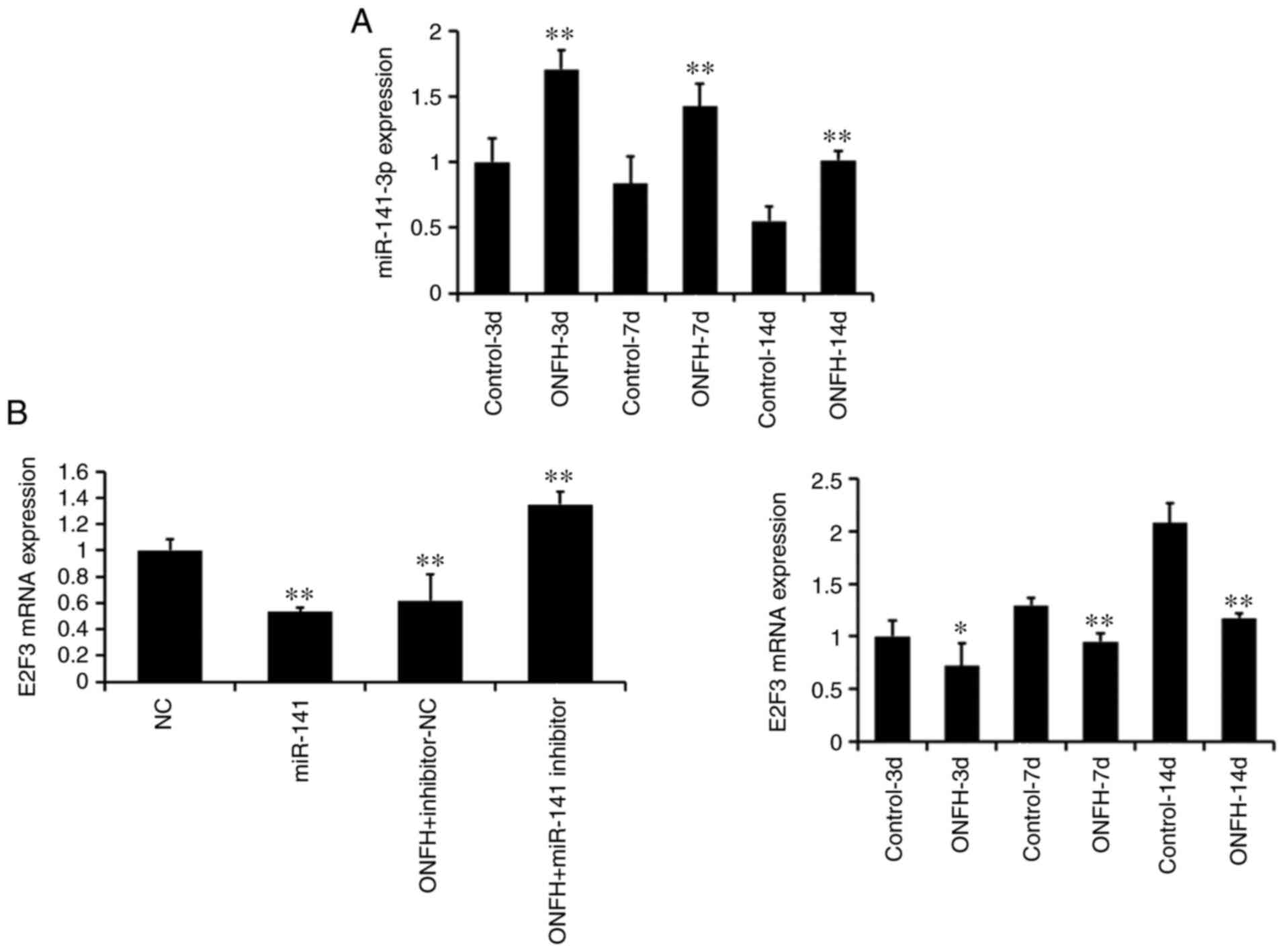
Successful miR-141 mimic transduction in normal
BMSCs and miR-141 inhibitor transduction in BMSCs from the ONFH rat
are shown in Fig. 4A (left; all
P<0.01). Furthermore, RT-qPCR demonstrated that the expression
levels of miR-141 in the ONFH group were significantly higher than
those of the control group at days 3, 7 and 14 (all P<0.01;
Fig. 4A, right). The mRNA
expression of E2F3 was significantly lower in the ONFH group than
in the control group at all time points (all P<0.01; Fig. 4B). This suggests that the
expression of E2F3 is inversely associated with that of
miR-141.
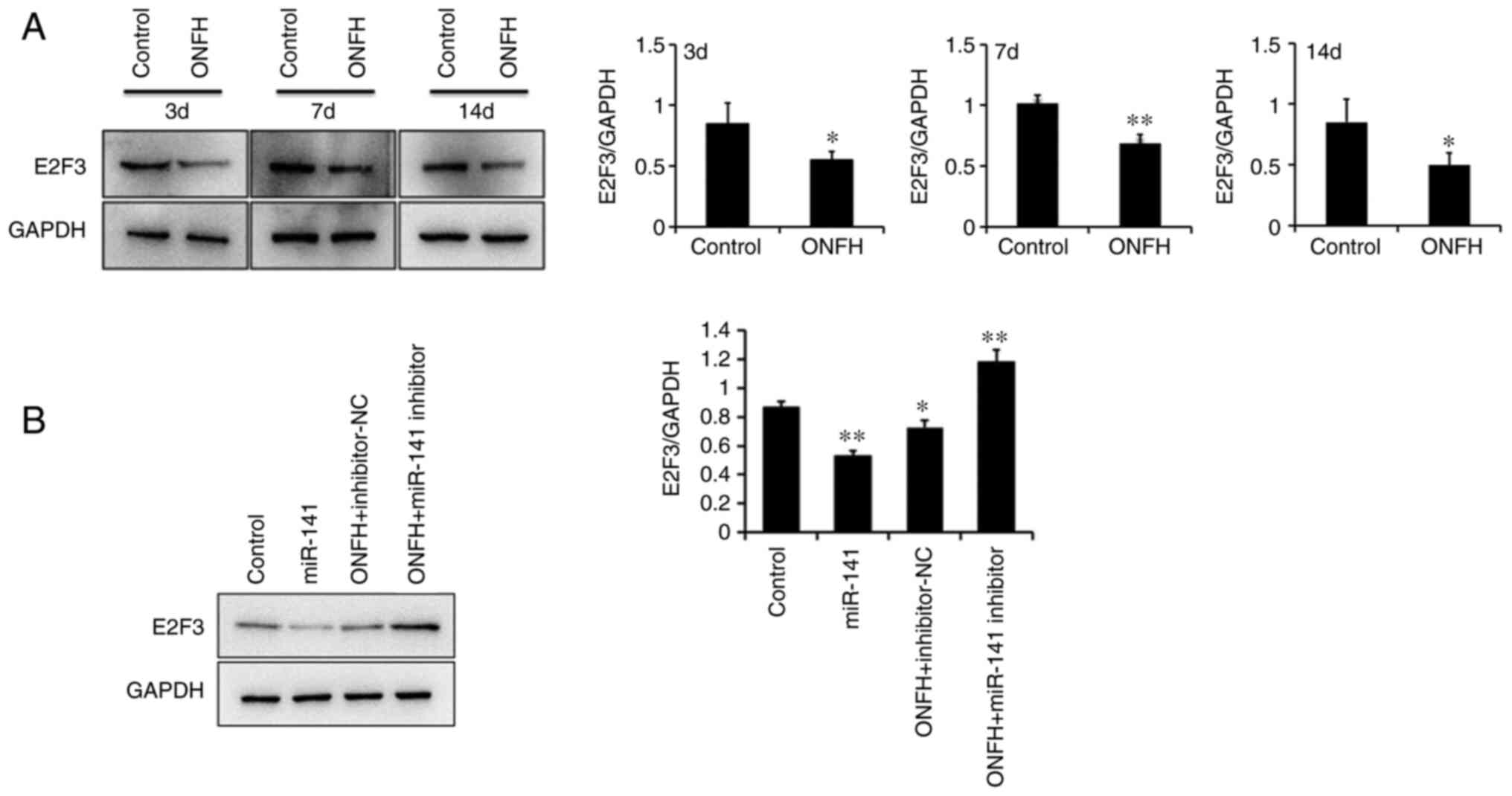
Protein expression levels of E2F3
The protein expression levels of E2F3 were
determined using western blotting. E2F3 expression was
significantly reduced BMSCs from the ONFH rat than those from the
control rat (Fig. 5A). The
protein expression levels of E2F3 were significantly downregulated
following transduction with the miR-141 overexpression lentivirus,
compared with BMSCs transduced with the NC lentivirus. However,
E2F3 was upregulated in BMSCs from the OPNFH rat transduced with
the miR-141 inhibitor lentivirus, compared with miR-141 inhibitor
Nc lentivirus (Fig. 5B). This
confirmed that miR-141 significantly downregulated the protein
expression levels of E2F3.
Discussion
ONFH is a progressive and painful hip joint disorder
affecting individuals in the 30–50 year age range (40). Previous studies have suggested
that glucocorticoids are associated with lipid metabolism.
Glucocorticoids facilitate decomposition of lipid into glycerol and
free fatty acids via direct action. Glucocorticoids inhibit
phosphodiesterase and CMP conversion to 5-terminal AMP.
Glucocorticoids increase fatty acids by upregulating the activity
of phenethylamylmethyltransferase, and adipose tissue. (41). Lipid metabolism and osteogenic
differentiation are associated with BMSCs since BMSCs undergo
adipose differentiation and osteogenic differentiation.
Differentiation of BMSCs is regulated by miRNA (15,42). Lipid metabolism could lead to the
collapse of the trabecular bone, causing empty lacunae and
microfractures in ONFH (43).
Osteocytes, which differentiate from BMSCs, are associated with
ONFH; death of osteocytes lead to ONFH A previous study has
demonstrated that the osteogenic differentiation of BMSCs in cells
from human ONFH tissue is downregulated, whereas adipocyte
differentiation is upregulated (44). The autophagy and apoptosis rates
of osteocytes were also increased in ONFH (45). It has been observed that the
number of osteoblasts, which are cells that differentiate from
BMSCs, are reduced in ONFH, whereas lipid differentiation is
increased (44). Thus, BMSCs play
an important role in the pathological changes associated with ONFH,
and it is important to investigate the factors that may affect
BMSCs in this disease.
miR-141 has been implicated in tissue repair and
osteogenic differentiation (23).
A previous study has indicated that miRNA plays an crucial role in
the osteogenic differentiation of BMSCs, and abnormal expression of
miRNA molecules may affect this process (46). Our previous study revealed that
miR-141 can inhibit proliferation in BMSCs (24). Nevertheless, the role of this
miRNA in osteogenic differentiation is not clear. The inhibition of
miR-141 may promote the osteogenic differentiation of BMSCs
(5) by targeting vitamin C
transporter 2 (25). In acute
kidney injury, it has been reported that miR-141 expression in
mesenchymal stromal cells promotes tissue repair (47). miR-141 overexpression can also
occur as a result of epigenetic regulation during senescence, which
may lead to a declines in physiological function and tissue
regeneration (48). miR-141 is a
member of the miR-200c/141 cluster and expression of the
miR-200c/141 cluster is regulated by DNA methylation, suggesting
epigenetic regulation of this miRNA locus in aggressive breast
cancer cell lines as well as untransformed mammary epithelial
cells. (49). Increasing evidence
suggests that several miRNA molecules participate in the
development and progression of ONFH, acting either as stimulators
or as suppressors (50). In the
present study, miR-141 was overexpressed in BMSCs. The results
demonstrated that miR-141 suppressed BMSC proliferation and
osteogenic differentiation.
E2F3 participates in the regulation of cellular
metastasis (26). Numerous miRNA
molecules have been found to be able to modulate E2F3 expression
(51), including miR-141. Indeed,
E2F3 has been identified as the target of miR-141 in HCC cells, and
the overexpression of E2F3 could partially reverse the
tumor-suppressive effects of miR-141 (27). E2F3 expression is altered in
several tumor types, and this transcription factor plays an
important role in tumor development (52,53). Therefore, it was hypothesized in
the present study that E2F3 and miR-141 could interact to modulate
BMSC differentiation in ONFH and play a crucial role in the
pathogenesis of this disease. The results suggested that miR-141
and E2F3 are potentially relevant factors of BMSC. The
overexpression of miR-141 reduced the expression levels of E2F3 and
inhibited osteogenic differentiation. Following miR-141 inhibition
using lentiviral transduction, E2F3 expression and osteogenic
differentiation significantly increased in ONFH group cells.
Increased E2F3 mRNA levels may partially reverse the suppressive
effects of ONFH on osteogenic differentiation. Altogether, these
observations suggest that miR-141 suppresses osteogenic
differentiation.
In conclusion, ONFH is a complex biological process
involving diverse mechanisms. The present study demonstrated that
miR-141 could suppress the osteogenic differentiation of BMSCs by
reducing E2F3 mRNA expression levels.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the Natural Science
Foundation of Inner Mongolia Autonomous Region (grant no.
2021LHMS08047).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
FX, JW, WF, TH, YL and WW participated in the design
of the study and contributed to drafting and revising the
manuscript. WF, YL and TH collected the data and performed the
statistical analyses. All authors read and approved the manuscript
and agree to be accountable for all aspects of the research in
ensuring that the accuracy or integrity of any part of the work are
appropriately investigated and resolved. FX and WBW confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
All animal experiments were approved by The Inner
Mongolia Medical University Animal Ethics Committee and performed
according to the Guidelines for the Care and Use of Laboratory
Animals.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Vreden SG, Hermus AR, van Liessum PA,
Pieters GF, Smals AG and Kloppenborg PWC: Aseptic bone necrosis in
patients on glucocorticoid replacement therapy. Neth J Med.
39:153–157. 1991.PubMed/NCBI
|
|
2
|
Shigemura T, Nakamura J, Kishida S, Harada
Y, Ohtori S, Kamikawa K, Ochiai N and Takahashi K: Incidence of
osteonecrosis associated with corticosteroid therapy among
different underlying diseases: Prospective MRI study. Rheumatology
(Oxford). 50:2023–2028. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Cui Q, Jo WL, Koo KH, Cheng EY, Drescher
W, Goodman SB, Ha YC, Hernigou P, Jones LC, Kim SY, et al: ARCO
consensus on the pathogenesis of non-traumatic osteonecrosis of the
femoral head. J Korean Med Sci. 36:e652021. View Article : Google Scholar
|
|
4
|
Liu Y, Jia Y, Cao Y, Zhao Y, Du J, An F,
Qi Y, Feng X, Jin T, Shi J and Wang J: MMP9 polymorphism is
associated with susceptibility to non-traumatic osteonecrosis of
femoral head in a Chinese Han population. Oncotarget.
8:82835–82841. 2017. View Article : Google Scholar
|
|
5
|
Liu W, Zhao Z, Na Y, Meng C, Wang J and
Bai R: Dexamethasone-induced production of reactive oxygen species
promotes apoptosis via endoplasmic reticulum stress and autophagy
in MC3T3-E1 cells. Int J Mol Med. 41:2028–2036. 2018.PubMed/NCBI
|
|
6
|
Jones JP Jr: Fat embolism, intravascular
coagulation, and osteonecrosis. Clin Orthop Relat Res. 294–308.
1993. View Article : Google Scholar
|
|
7
|
Xu X, Wen H, Hu Y, Yu H, Zhang Y, Chen C
and Pan X: STAT1-caspase 3 pathway in the apoptotic process
associated with steroid-induced necrosis of the femoral head. J Mol
Histol. 45:473–485. 2014. View Article : Google Scholar
|
|
8
|
Tack L, Tatsi C, Stratakis CA and Lodish
MB: Effects of glucocorticoids on bone: What we can learn from
pediatric endogenous cushing's syndrome. Horm Metab Res.
48:764–770. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wu Q, Xiong X, Zhang X, Lu J, Zhang X,
Chen W, Wu T, Cui L, Liu Y and Xu B: Secondary osteoporosis in
collagen-induced arthritis rats. J Bone Miner Metab. 34:500–516.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wu Z, Bucher NL and Farmer SR: Induction
of peroxisome proliferator-activated receptor gamma during the
conversion of 3T3 fibroblasts into adipocytes is mediated by
C/EBPbeta, C/EBPdelta, and glucocorticoids. Mol Cell Biol.
16:4128–4136. 1996. View Article : Google Scholar
|
|
11
|
Pittenger MF, Mackay AM, Beck SC, Jaiswal
RK, Douglas R, Mosca JD, Moorman MA, Simonetti DW, Craig S and
Marshak DR: Multilineage potential of adult human mesenchymal stem
cells. Science. 284:143. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Prockop DJ: Marrow stromal cells as stem
cells for nonhematopoietic tissues. Science. 276:71–74. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Gao Y, Patil S and Jia J: The development
of molecular biology of osteoporosis. Int J Mol Sci. 22:81822021.
View Article : Google Scholar
|
|
14
|
Matsuya H, Kushida T, Asada T, Umeda M,
Wada T and Iida H: Regenerative effects of transplanting autologous
mesenchymal stem cells on corticosteroid-induced osteonecrosis in
rabbits. Mod Rheumatol. 18:132–139. 2008. View Article : Google Scholar
|
|
15
|
Tian L and Yu X: Lipid metabolism
disorders and bone dysfunction-interrelated and mutually regulated
(Review). Mol Med Rep. 12:783–794. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lee JS, Lee JS, Roh HL, Kim CH, Jin SJ and
Suh KT: Alterations in the differentiation ability of mesenchymal
stem cells in patients with nontraumatic osteonecrosis of the
femoral head: Comparative analysis according to the risk factor. J
Orthop Res. 24:604–609. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Liu K, Wen G, Liu R, Liang XU, Feng LI and
Tao S: Research progress of the treatment of regional osteoporosis
with BMSCs transplantation. Chin J Osteoporos. 11:1203–1206.
2013.(In Chinese).
|
|
18
|
Lewis H, Lance R, Troyer D, Beydoun H,
Hadley M, Orians J, Benzine T, Madric K, Semmes OJ, Drake R and
Esquela-Kerscher A: MiR-888 is an expressed prostatic
secretions-derived microRNA that promotes prostate cell growth and
migration. Cell Cycle. 13:227–239. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ambros V: Victor Ambros: The broad scope
of microRNAs. Interview by Caitlin Sedwick. J Cell Biol.
201:492–493. 2013. View Article : Google Scholar
|
|
20
|
Long ZH, Bai ZG, Song JN, Zheng Z, Li J,
Zhang J, Cai J, Yao HW, Wang J, Yang YC, et al: MiR-141 inhibits
proliferation and migration of colorectal cancer SW480 Cells.
Anticancer Res. 37:4345–4352. 2017.PubMed/NCBI
|
|
21
|
Hou X, Yang L, Jiang X, Liu Z, Li X, Xie
S, Li G and Liu J: Role of microRNA-141-3p in the progression and
metastasis of hepatocellular carcinoma cell. Int J Biol Macromol.
128:331–339. 2019. View Article : Google Scholar
|
|
22
|
Xu H, Mei Q, Xiong C and Zhao J:
Tumor-suppressing effects of miR-141 in human osteosarcoma. Cell
Biochem Biophys. 69:319–325. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yaman Agaoglu F, Kovancilar M, Dizdar Y,
Darendeliler E, Holdenrieder S, Dalay N and Gezer U: Investigation
of miR-21, miR-141, and miR-221 in blood circulation of patients
with prostate cancer. Tumour Biol. 32:583–588. 2011. View Article : Google Scholar
|
|
24
|
Meng CY, Xue F, Zhao ZQ, Hao T, Guo SB and
Feng W: Influence of MicroRNA-141 on inhibition of the
proliferation of bone marrow mesenchymal stem cells in
steroid-Induced Osteonecrosis via SOX11. Orthop Surg. 12:277–285.
2020. View
Article : Google Scholar : PubMed/NCBI
|
|
25
|
Sangani R, Periyasamy-Thandavan S, Kolhe
R, Bhattacharyya MH, Chutkan N, Hunter M, Isales C, Hamrick M, Hill
WD and Fulzele S: MicroRNAs-141 and 200a regulate the SVCT2
transporter in bone marrow stromal cells. Mol Cell Endocrinol.
410:19–26. 2015. View Article : Google Scholar
|
|
26
|
Rady B, Chen Y, Vaca P, Wang Q, Wang Y,
Salmon P and Oberholzer J: Overexpression of E2F3 promotes
proliferation of functional human β cells without induction of
apoptosis. Cell Cycle. 12:2691–2702. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Xue J, Niu YF, Huang J, Peng G, Wang LX,
Yang YH and Li YQ: MiR-141 suppresses the growth and metastasis of
HCC cells by targeting E2F3. Tumour Biol. 35:12103–12107. 2014.
View Article : Google Scholar
|
|
28
|
Kim HR, Rahman FU, Kim KS, Kim EK, Cho SM,
Lee K, Moon OS, Seo YW, Yoon WK, Won YS, et al: Critical roles of
E2F3 in growth and musculo-skeletal phenotype in mice. Int J Med
Sci. 16:1557–1563. 2019. View Article : Google Scholar
|
|
29
|
Farag MR, Anter RGA, Elhady WM, Khalil SR,
Abou-Zeid SM and Hassanen EAA: Diversity, succession pattern and
colonization of forensic entomofauna on indoor rat carrions
concerning the manner of death. Rend Fis Acc Lincei. 32:521–538.
2021. View Article : Google Scholar
|
|
30
|
Yaghoobi M, Hashemi-Najafabadi S,
Soleimani M and Vasheghani-Farahani E: Osteogenic induction of
human mesenchymal stem cells in multilayered electrospun scaffolds
at different flow rates and configurations in a perfusion
bioreactor. J Biosci Bioeng. 128:495–503. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Adibkia K, Ehsani A, Jodaei A, Fathi E,
Farahzadi R and Barzegar-Jalali M: Silver nanoparticles induce the
cardiomyogenic differentiation of bone marrow derived mesenchymal
stem cells via telomere length extension. Beilstein J Nanotechnol.
12:786–797. 2021. View Article : Google Scholar
|
|
32
|
Li X, Wang L, Su Q, Ye L, Zhou X, Zhang L,
Song D and Huang D: Potential roles of bone morphogenetic protein 9
in the odontogenic differentiation of dental pulp cells. J Endod.
47:436–443. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Langkilde A, Raaby L, Johansen C and
Iversen L: MicroRNA normalization candidates for quantitative
reverse-transcriptase polymerase chain reaction in real time in
lesional and nonlesional psoriatic skin. Br J Dermatol.
169:677–681. 2013. View Article : Google Scholar
|
|
34
|
Nicolas FE: Experimental validation of
MicroRNA targets using a luciferase reporter system. Methods Mol
Biol. 732:139–152. 2011. View Article : Google Scholar
|
|
35
|
Wang Y, Sun J, Guo X, Zhang D, Cui Y, Li
W, Liu G, Li Y and Jiang S: TaqMan-based real-time polymerase chain
reaction assay for specific detection of bocavirus-1 in domestic
cats. Mol Cell Probes. 53:1016472020. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Fathi E, Farahzadi R, Vietor I and
Javanmardi S: Cardiac differentiation of bone-marrow-resident
c-kit+ stem cells by L-carnitine increases through
secretion of VEGF, IL6, IGF-1 and TGF-β. as clinical agents in
cardiac regeneration. J Biosci. 45:922020. View Article : Google Scholar
|
|
37
|
Fathi E, Farahzadi R and Valipour B:
Alginate/gelatin encapsulation promotes NK cells differentiation
potential of bone marrow resident C-kit+ hematopoietic stem cells.
Int J Biol Macromol. 177:317–327. 2021. View Article : Google Scholar
|
|
38
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Kurien BT and Scofield RH: Western
blotting. Methods. 38:283–293. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lau RL, Perruccio AV, Evans HM, Mahomed
SR, Mahomed NN and Gandhi R: Stem cell therapy for the treatment of
early stage avascular necrosis of the femoral head: A systematic
review. BMC Musculoskelet Disord. 15:1562014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Peckett AJ, Wright DC and Riddell MC: The
effects of glucocorticoids on adipose tissue lipid metabolism.
Metabolism. 60:1500–1510. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Pansky A, Roitzheim B and Tobiasch E:
Differentiation potential of adult human mesenchymal stem cells.
Clin Lab. 53:81–84. 2011.
|
|
43
|
Yu Y, Wei N, Stanford C, Schmidt T and
Hong L: In vitro effects of RU486 on proliferation and
differentiation capabilities of human bone marrow mesenchymal
stromal cells. Steroids. 77:132–137. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Wang T, Teng S, Zhang Y, Wang F, Ding H
and Guo L: Role of mesenchymal stem cells on differentiation in
steroid-induced avascular necrosis of the femoral head. Exp Ther
Med. 13:669–675. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Bai R, Feng W, Liu WL, Zhao ZH, Zhao AQ,
Wang Y, Wang WX, Sun L, Wu LS and Cui SH: Roles of osteocyte
apoptosis in steroid-induced avascular necrosis of the femoral
head. Genet Mol Res. 15:2016. View Article : Google Scholar
|
|
46
|
Wang J, Liu S, Li J, Zhao S and Yi Z:
Roles for miRNAs in osteogenic differentiation of bone marrow
mesenchymal stem cells. Stem Cell Res Ther. 10:1972019. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
de Almeida DC, Bassi ÊJ, Azevedo H,
Anderson L, Origassa CS, Cenedeze MA, de Andrade-Oliveira V,
Felizardo RJ, da Silva RC, Hiyane MI, et al: A Regulatory
miRNA-mRNA network is associated with tissue repair induced by
mesenchymal stromal cells in acute kidney injury. Front Immunol.
7:6452017. View Article : Google Scholar
|
|
48
|
Yu KR, Lee S, Jung JW, Hong IS, Kim HS,
Seo Y, Shin TH and Kang KS: MicroRNA-141-3p plays a role in human
mesenchymal stem cell aging by directly targeting ZMPSTE24. J Cell
Sci. 126((Pt 23)): 5422–5431. 2013.
|
|
49
|
Neves R, Scheel C, Weinhold S, Honisch E,
Iwaniuk KM, Trompeter HI, Niederacher D, Wernet P, Santourlidis S
and Uhrberg M: Role of DNA methylation in miR-200c/141 cluster
silencing in invasive breast cancer cells. BMC Res Notes.
3:2192010. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Pizzini S, Bisognin A, Mandruzzato S,
Biasiolo M, Facciolli A, Perilli L, Rossi E, Esposito G, Rugge M,
Pilati P, et al: Impact of microRNAs on regulatory networks and
pathways in human colorectal carcinogenesis and development of
metastasis. BMC Genomics. 14:5892013. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Miles WO, Tschop K, Herr A, Ji JY and
Dyson NJ: Pumilio facilitates miRNA regulation of the E2F3
oncogene. Genes Dev. 26:356–368. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zeng X, Yin F, Liu X, Xu J, Xu Y, Huang J,
Nan Y and Qiu X: Upregulation of E2F transcription factor 3 is
associated with poor prognosis in hepatocellular carcinoma. Oncol
Rep. 31:1139–1146. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Bilke S, Schwentner R, Yang F, Kauer M,
Jug G, Walker RL, Davis S, Zhu YJ, Pineda M, Meltzer PS and Kovar
H: Oncogenic ETS fusions deregulate E2F3 target genes in Ewing
sarcoma and prostate cancer. Genome Res. 23:1797–1809. 2013.
View Article : Google Scholar : PubMed/NCBI
|