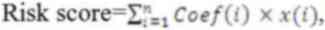Introduction
Bladder cancer is one of the most common malignant
tumours of the genitourinary system, the annual incidence of which
is increasing worldwide, with ~550,000 new cases annually (1,2).
Radical cystectomy is the primary treatment method for bladder
cancer. However, ~50% of patients experience recurrence and cancer
metastasis. Therefore, the 5-year survival rate of patients has not
notably improved (3,4). The metastasis of bladder cancer is
one of the primary factors that leads to the poor prognosis of
patients. Previous studies have reported that the mechanism
underlying bladder cancer metastasis is associated with tumour
angiogenesis (5–7). Angiogenesis is the generation of
microvessels on existing blood vessels and serves an important role
in tumour metastasis (8).
In the human genome, <2% of the DNA sequence
encodes proteins and the remaining sequences that do not encode
proteins are known as non-coding sequences. Compared with simpler
eukaryotes, the proportion of non-coding sequences is higher in the
genome of more complex lifeforms (9,10).
Long non-coding (lnc)RNAs belong to a class of non-protein-coding
transcripts that are >200 base pairs in length and regulate gene
expression on multiple levels. This can occur by modulation of
chromatin modification, RNA splicing and protein activity, which
influence the occurrence and progression of bladder cancer
(11).
LncRNAs and angiogenesis serve important roles in
the regulation of the progression of tumours. Therefore, in the
present study, angiogenesis-associated lncRNAs were screened using
The Cancer Genome Atlas (TCGA) database and the angiogenesis gene
set from The Molecular Signatures Database. Using these data, an
angiogenesis-associated lncRNA signature was constructed to predict
the prognosis of patients with bladder cancer. Furthermore, the
role and underlying mechanism of the angiogenesis-associated long
intergenic non-coding RNA (LINC)02321 in the progression of bladder
cancer was evaluated. The present study provided a new idea for the
understanding of the mechanism of bladder cancer malignant
progression and a new potential target for the treatment of bladder
cancer patients.
Materials and methods
Data acquisition
Bladder cancer RNA-sequencing and clinical data
(Table SI) were downloaded from
TCGA database (https://portal.gdc.cancer.gov/. TCGA-BLCA). Bladder
cancer cases were randomly divided into a training set (n=201) and
a validation set (n=201) using a simple random grouping method. The
signature was constructed with the training set and verified with
the validation set and the whole set (n=402). A set of
angiogenesis-associated genes (Table
SII) were downloaded from the gene set ‘Angiogenesis, M14493’
in The Molecular Signatures Database (www.broadinstitute.org/gsea/msigdb).
Construction of the
angiogenesis-associated lncRNA prognostic signature
The lncRNA expression data in bladder cancer from
TCGA database were assessed using scripts written using the ‘Perl’
(version 5.32.1.1) programming language (https://www.perl.org/). Angiogenesis-associated
lncRNAs were identified using Pearson's correlation analysis
(BiocManager and limma packages; R software, version
4.0.2, http://cran.r-project.org/mirrors.html) (coefficient
|R2|>0.5 and P<0.05). A total of 390
angiogenesis-associated lncRNAs were screened (Table SIII). Univariate/multivariate Cox
regression analysis (survival package; R software, version
4.0.2) was performed to identify the patient prognosis-associated
lncRNAs (Table SIV). A total of
six angiogenesis-associated lncRNAs were identified as candidates
for use in the prognostic signature (Table I). Cox regression analysis was
used to analyze the relationship between the expression of lncRNAs
and the survival and prognosis of patients with bladder cancer.
Only the lncRNAs related to the prognosis of bladder cancer
patients were screened. The six lncRNAs screened were closely
related to the prognosis of bladder cancer patients).
 | Table I.Angiogenesis-associated lncRNAs
significantly associated with the overall survival of patients with
bladder cancer. |
Table I.
Angiogenesis-associated lncRNAs
significantly associated with the overall survival of patients with
bladder cancer.
| lncRNA | Ensemble ID | Location | β-value | Hazard ratio | P-value |
|---|
| USP30-AS1 |
ENSG00000256262 |
chr12:109,051,790-109,054,033 | −0.1012 | 0.9036 | 0.0015 |
| LINC02321 |
ENSG00000258884 |
chr14:90,819,380-90,828,206 | 0.1271 | 1.1356 | 0.0011 |
| PSMB8-AS1 |
ENSG00000204261 |
chr6:32,811,863-32,814,272 | −0.0938 | 0.9104 | 0.0007 |
| KRT7-AS |
ENSG00000257671 |
chr12:52,638,832-52,641,232 | 0.01966 | 1.0198 | 0.0046 |
| LINC01767 |
ENSG00000223956 |
chr1:56,414,930-56,420,384 | −0.19960 | 0.8190 | 0.0101 |
| OCIAD1-AS1 |
ENSG00000248256 |
chr4:48,852,007-48,860,646 | −0.57417 | 0.5631 | 0.0005 |
Evaluation and verification of the
accuracy of the prognostic signature
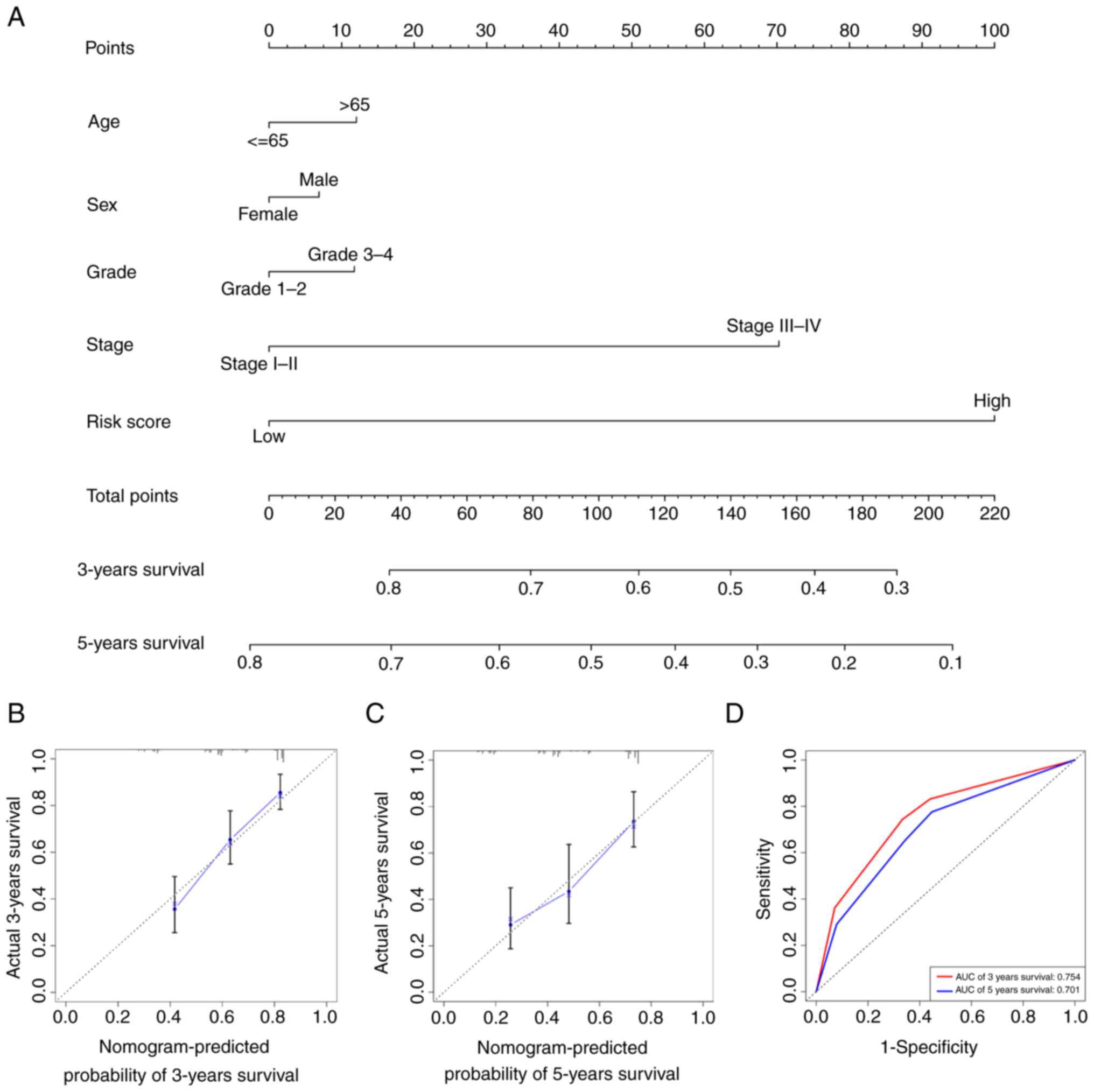
The risk score for each patient was calculated based
on the following formula:
where ‘Coef(i)’ represents the
estimated regression coefficient and ‘x(i)’
represents the expression of each angiogenesis-associated lncRNA.
The patients with bladder cancer were categorised into low- and
high-risk groups according to the median risk score (training set,
1.82462; validation set, 1.79457; whole set, 1.85822). The
prediction efficiency of the angiogenesis-associated lncRNA
signature was evaluated by plotting receiver operating
characteristic (ROC) curves. The independent prediction ability of
the signature was quantified using Cox regression analysis
(survival package of R software, version 4.0.2).
Establishment of the nomogram and gene
set enrichment analysis (GSEA)
We carried out COX regression analysis (rms
package of R software, version 4.0.2) to construct nomogram. The
nomogram was constructed by integrating clinical variables [age,
sex, BLCA grade (12), American
Joint Committee on Cancer (AJCC) stage (13)] and risk score to evaluate the
overall survival (OS) of patients with bladder cancer. Calibration
plots and time-dependent ROC curves were constructed to assess the
nomogram. GSEA (4.0.3) was performed using the
‘c2.cp.kegg.v7.2.symbols.gmt’ Kyoto Encyclopaedia of Genes and
Genomes gene sets (http://www.gsea-msigdb.org/gsea/index.jsp) to assess
the LINC02321-associated pathways. Expression data for VEGFA in
bladder cancer were obtained from the Human Protein Atlas database
(www.proteinatlas.org; accession no.
M-00100 and M812033).
Cell culture and tissue samples
The human bladder cancer cell lines 5637, T24 and
UM-UC-3, and the ureteral epithelial cell line SV-HUC-1 were
purchased from American Type Culture Collection. The bladder cancer
cell lines were cultured in RPMI-1640 medium (Gibco; Thermo Fisher
Scientific, Inc.) containing 10% FBS (PAN-Biotech GmbH) and 1%
penicillin-streptomycin (Beyotime Institute of Biotechnology).
SV-HUC-1 cells were cultured in DMEM (Corning, Inc.) containing 10%
FBS and 1% penicillin-streptomycin. All cells were cultured in 5%
CO2 at 37°C. Fresh bladder cancer and normal adjacent
tissues (distance between tumor and adjacent tissue ≤3 cm) were
collected from patients (n=10; age, 53–81 years; male, 8; female,
2) who underwent radical cystectomy at The First Affiliated
Hospital of Chongqing Medical University between April 2021 and
February 2022. Inclusion criteria used were as follows: i)
Diagnosed with myometrial invasive bladder cancer or high-risk
patients with non-muscular invasive bladder cancer who failed to
respond to Bacillus Calmette-Guerin vaccine (BCG) therapy and
underwent total cystectomy according to the 2020 European
Urological Association Guidelines for Diagnosis and Treatment of
Myometrial Invasive bladder cancer (14); ii) patients with basic language
communication and understanding ability; iii) patients knew of
their own conditions, were able to provide informed consent and
volunteered to participate in the study. Patients with other
malignant tumors involving other systems at the same time were
excluded. Each patient signed a written informed consent form and
the present study was approved by the Ethics Committee of the First
Affiliated Hospital of Chongqing Medical University (approval no.
2021069; Chongqing, China).
Small interfering (si)RNA
All experiments using siRNA were performed in T24
cells. siRNA and negative control siRNA (si-NC) against LINC02321
were designed and synthesised by Shanghai Genepharma Co., Ltd.
After the T24 cells reached 80% confluence, the medium was replaced
with 1.5 ml basal medium containing 500 µl
Lipofectamine® 2000 (Invitrogen, Thermo Fisher
Scientific, Inc.) and 0.2 nmol siRNA, and the cells were incubated
at 37°C for 6 h. The subsequent experiments related to lncRNA
expression were performed 24 h later and the cell function
experiments were performed 48 h later. The sequences of the siRNAs
were as follows: si-LINC02321-#1,
5′-UUAUUCUGAATUATTUGUCAUUAUUCUCAAUGAGTTGAACAC-3′; si-LINC02321-#2,
5′-UATTCCUCUGAGGTTACUCACUAUUUAAAGUGAGUUAUUAAA-3′; si-LINC02321-#3,
5′-UUGGGAGTTUACAAAGAATTCUCCCUUCUAUCAGUGAGACUU-3′ and si-NC,
5′-UUCUCUCAACGUCGUTCCGAATGUGACGUGAAATCUCGGAGT-3′.
Construction of
LINC02321-overexpressing cell lines
The lenti-vector and lenti-OE were cloned into the
Gemma gene (Shanghai Jima Pharmaceutical Technology Co., Ltd.). The
recombinant pGag/Pol-pRev-pVSV-G viral plasmid (3rd generation
system) encoding lentivirus particles and the carrier plasmids of
the three auxiliary packaging components were prepared and
high-purity endotoxin free extraction was performed. 293T cells
were co-transfected with 10 µg of the lentivirus (lentivirus,
packaging, envelope plasmids=4:3:1) at 37°C using Shanghai Jima
Pharmaceutical Technology Co., Ltd. company's transfection reagent
RNAi Mate. Media was replaced with a complete medium 6 h after
transfection. After 72 h of culture, the cell supernatant, was
collected and concentrated to obtain a lentivirus concentrate with
high titre for infection of target cells [multiplicity of infection
(MOI)=30]. The infected target cells were screened using the
resistance gene (Puromycin; 2 ng/ml for selection; 1 ng/ml for
maintenance) and fluorescent fusion protein on the lentivirus
vector. The plasmids, cell lines and other reagents were purchased
from Shanghai Jima Pharmaceutical Technology Co., Ltd. The 5637
cells in a normal growth state were evenly seeded (using the cell
count plate) into six-well plates and cultured in a 37°C incubator
containing 5% CO2 when 5637 cells reached ~60%
confluence. A basic RPMI-1640 medium (Gibco; Thermo Fisher
Scientific, Inc.) containing the overexpressing lentivirus or
control virus (MOI=30) was added to the culture for 24 h when the
cells reached 30–50% confluency. After the virus was transfected
into cells, puromycin (2 ng/ml) was added to the medium to screen
and maintain the stably transfected cells for subsequent
experiments. Subsequent experiments were carried out 24 h after
successful screening. LINC02321 overexpression was compared with
the empty vector control.
Reverse transcription-quantitative PCR
(RT-qPCR)
RNA from cells (T24, 5637 and UM-UC-3 bladder cancer
cell lines and the SV-HUC-1 ureteral epithelial cell line) and
tissues (bladder cancer and normal adjacent tissues) was extracted
using TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocols. The
extracted RNA was reverse-transcribed into cDNA using a Primescript
RT kit (Takara Bio, Inc.) according to the manufacturer's protocol.
qPCR (PCR kit, Takara Bio, Inc.) was performed using the following
primer pairs (: LINC02321 forward (F), 5′-GGAGATGAGGACTGGGAGGT-3′
and reverse I, 5′-GGACAGCTGGGAAAGGAGTC-3′; and GAPDH F,
5′-GCAGCGAGATCCCTCCAAAAT-3′ and R, 5′-CTGTTGTCATACTTCTCATGG-3′, and
thermocycling conditions of 95°C for 30 sec, followed by 40 cycles
of 95°C for 5 sec and 65°C for 15 sec. The mRNA expression of GAPDH
was used as the reference to calculate the expression level of the
target gene. The 2-ΔΔCq method was used to calculate
relative gene expression after normalization to GAPDH (15).
Wound healing assay
T24 cells were inoculated into a six-well plate and
after reaching 100% confluency, a sterile pipette tip was used to
scratch cells perpendicular to the plate (two lines horizontally
and vertically) (16).
Subsequently, serum-free RPMI-1640 medium (Gibco; Thermo Fisher
Scientific, Inc.) was added before images were captured using an
inverted light microscope. The time at which the images were
captured was recorded as 0 h. Finally, serum-free RPMI-1640 medium
(Gibco; Thermo Fisher Scientific, Inc.) was added and the cells
were cultured at 37°C for 18 h. The migration of cells in the
control and experimental groups was assessed at 18 h using an
inverted light microscope (magnification, ×100). ImageJ software
(National Institutes of Health, version 1.8.0) was used to assess
the images.
Transwell experiments
Matrigel was used for invasion experiments and was
placed in the upper layer of the Transwell chamber. Precoating was
performed at 37°C for 4 h. A total of 8×104 T24 and 5637
cells were inoculated into each chamber and 600 µl RPMI-1640 medium
with 10% FBS was added into the lower chamber for culture. The
cells were incubated at 37°C for 12 h and fixed using 4%
paraformaldehyde at 37°C for 15 min. The fixed cells were stained
using 0.1% crystal violet solution at 37°C for 15 min. Following
this, cells were imaged using a light microscope and quantified
using ImageJ software (National Institutes of Health, version
1.8.0). The addition of Matrigel was not performed for Transwell
migration experiments, but all subsequent procedures were identical
to those aforementioned.
Cell counting kit-8 (CCK-8)
proliferation assay
T24 and 5637 cells were inoculated into 96-well
plates with 4,000 cells/well. A total of 10 µl CCK-8 reagent
(Dojindo Molecular Technologies, Inc.) was added after 24, 48 and
72 h of incubation at 37°C for 1 h, before absorbance was assessed
at 450 nm.
CCK-8 assay of cisplatin
sensitivity
T24 and 5637 cells were inoculated into 96-well
plates with 4,000 cells/well. After cells attached, cisplatin
(Beijing Solarbio Science & Technology Co., Ltd.) was added at
concentrations of 0.0, 0.5, 1.0, 2.0, 5.0, 10.0 and 20.0 µg/ml and
cells were cultured at 37°C for 48 h. Subsequently, 10 µl CCK-8
reagent was added to each well and incubated for 1 h. Finally, the
absorbance was assessed at 450 nm using a microplate reader.
Western blotting
RIPA buffer (Beyotime Institute of Biotechnology)
containing PMSF (Beyotime Institute of Biotechnology) was used to
extract cellular proteins. Protein concentration was assessed using
a BCA protein analysis kit. Protein samples from each group (30
µg/lane) were separated using 10% SDS-PAGE and transferred onto
PVDF membranes. PVDF membranes were blocked with QuickBlock™
Western rapid sealing solution (cat. no. P0252, Beyotime Institute
of Biotechnology,) at room temperature for 30 min. Primary rabbit
anti-VEGFA (1:1,000; cat. no. 50661; Cell Signaling Technology,
Inc.) and anti-GAPDH (1:8,000; cat. no. 60004-1-Ig; ProteinTech
Group, Inc.) antibodies were then added to the membranes, which
were incubated at 4°C for ≥12 h. The membranes were incubated at
37°C for 1 h with HRP-labelled secondary antibody (1:3,000, cat.
no. PR30011; ProteinTech Group, Inc; http://www.ptgcn.com/). Finally, a fusion imaging
system QuantityOne (Bio-Rad Laboratories, Inc.) was used to assess
the relative density of the band. ImageJ software (National
Institutes of Health, version 1.8.0) was used for analysis.
Statistical analysis
Statistical analysis was performed using SPSS 20.0
software (IBM Corp.). Graphs were generated using GraphPad Prism
8.0 (GraphPad Software, Inc.). All data are presented as the mean ±
standard deviation of experiments that were repeated three times.
Paired Student's t test was used when comparing matched samples;
unpaired Student's t test was used when comparing non-matched
samples. Comparisons between multiple groups were performed using
two-way ANOVA and post hoc Dunnett's test for multiple comparisons.
The survival curves were produced and analysed using the
Kaplan-Meier method and parallel log-rank inspection. P<0.05 was
considered to indicate a statistically significant difference.
Results
Construction and verification of the
angiogenesis-associated lncRNA signature
A total of 14,142 lncRNAs were identified using the
‘Perl’ programming language. A total of 390 angiogenesis-associated
lncRNAs were screened using Pearson's correlation analysis
(Table SIII). In the training
set, 64 angiogenesis-associated lncRNAs were screened using
univariate Cox regression analysis (Table SIV), of which six were identified
as candidates for the prognostic signature using multivariate Cox
regression analysis: USP30-AS1, LINC02321, PSMB8-AS1, KRT7-AS,
LINC01767 and OCIAD1-AS1 (Table
I). The patients in the training, validation and whole sets
were divided into two groups according to the median risk score
(Fig. 1A). The OS rate was
significantly shorter in the high-risk group compared with that in
the low-risk patients in each group (Fig. 1B). The sensitivity and specificity
of the signature were evaluated based on the ROC. In the three
sets, the highest area under the curve (AUC) value was 0.807 for
the whole set at 5 years (Fig.
1C). These results demonstrated that the
angiogenesis-associated lncRNA signature may accurately predict
survival and prognosis of patients with bladder cancer.
Angiogenesis-associated lncRNA
signature is an independent prognostic factor
The independent predictive ability of the risk score
based on the angiogenesis-associated lncRNA signature and
clinicopathological parameters on the prognosis of patients with
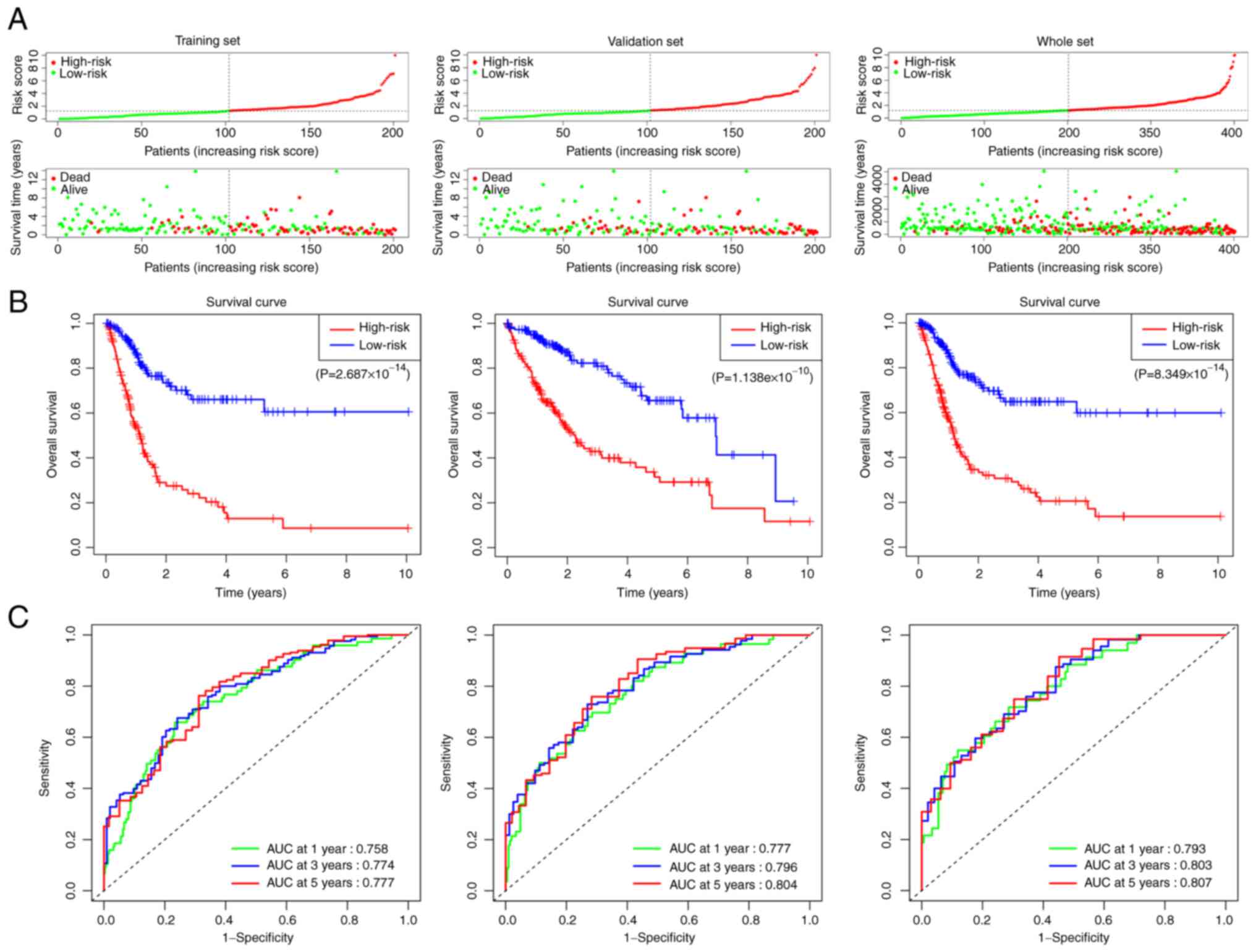
bladder cancer was evaluated using Cox regression analysis. The
results demonstrated a significant association between risk score
of the signature and the OS time of patients (Fig. 2A and B). ROC curve results
demonstrated that the AUC of the risk score, 0.844, was higher
compared with those of the other clinicopathological parameters
(Fig. 2C). These results
suggested that the risk score based on the angiogenesis-associated
lncRNA signature may be used as an independent factor to predict
the prognosis of patients with bladder cancer.
Establishment of the nomogram
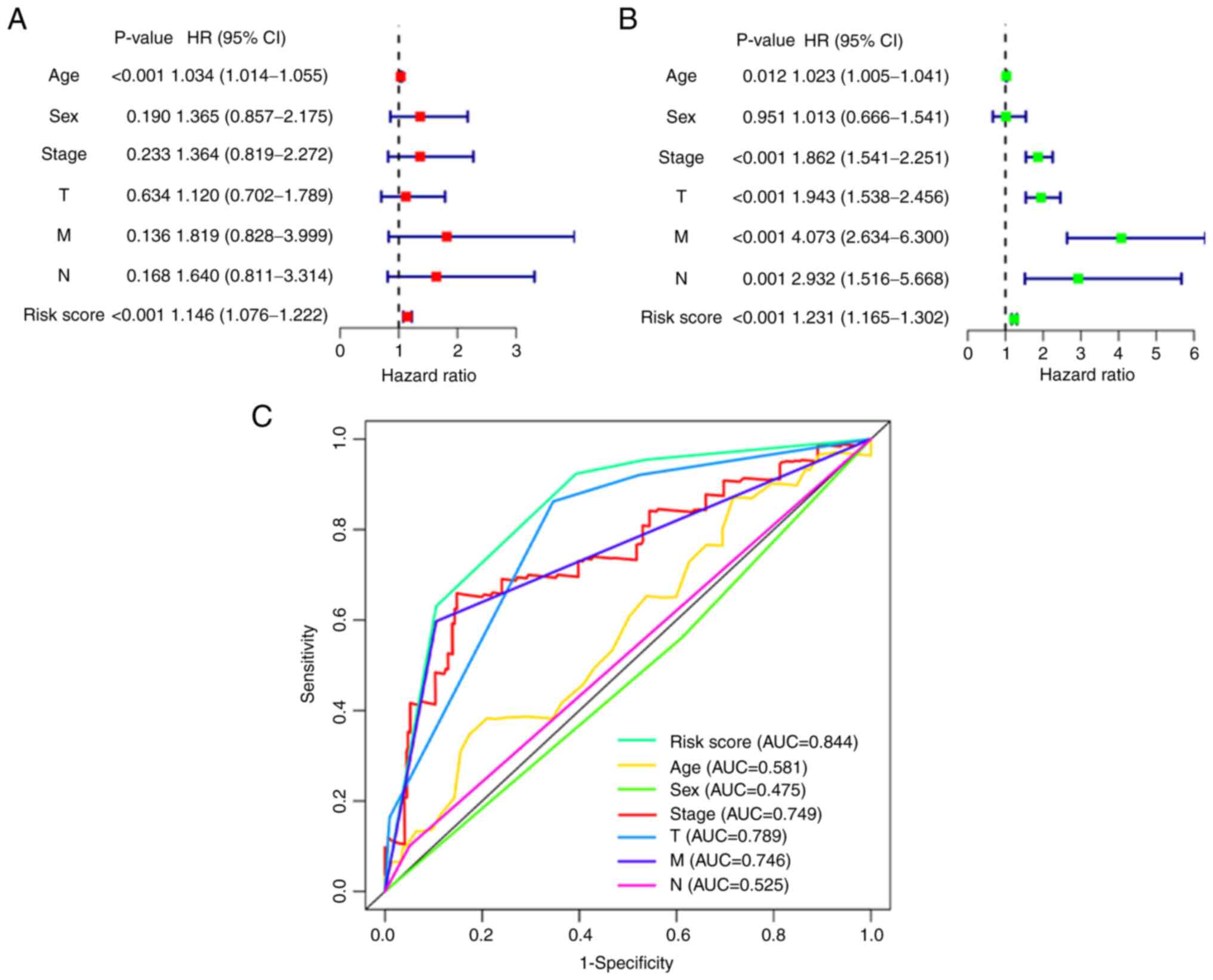
A nomogram combines multiple indicators to diagnose
or predict the onset and progression of disease (17). Therefore, a nomogram was
constructed based on the multivariate Cox regression analysis
results by considering the risk score of the signature, age, sex,
grade and AJCC stage of the tumour (Fig. 3A). Furthermore, calibration using
Cox regression analysis (Fig. 3B)
and ROC curves (Fig. 3C) were
plotted to assess the predictive ability of the nomogram, which
yielded 3-year and 5-year AUC values of 0.754 and 0.701,
respectively (Fig. 3D).
LINC02321 is highly expressed in
bladder cancer
Analysis of the angiogenesis-associated lncRNA
signature was performed. The key lncRNAs of the signature were
screened using prediction of their expression profiles from TCGA
dataset and RT-qPCR validation of significant lncRNAs. In TCGA
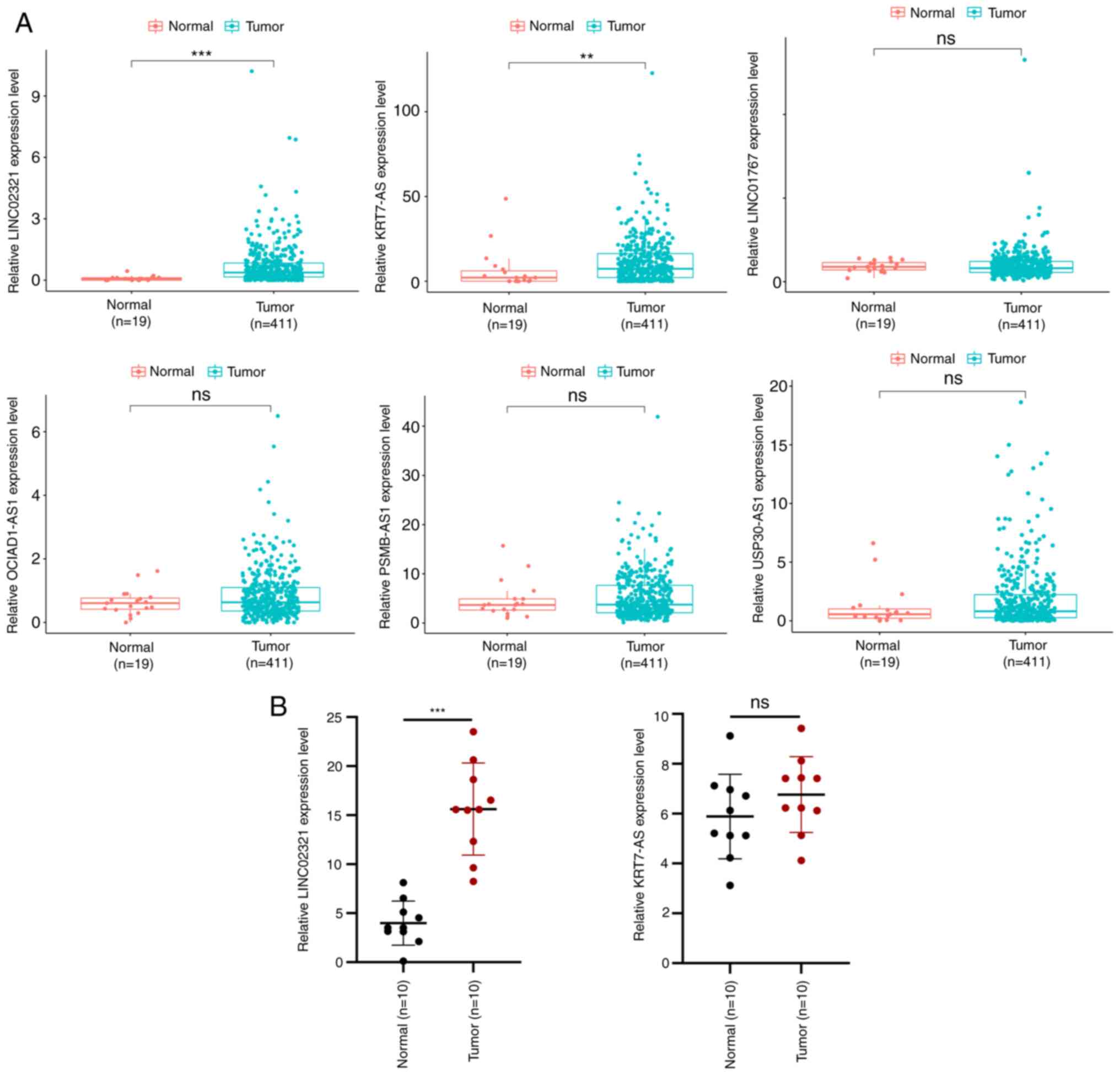
database, the expression profiles of the six lncRNAs in the
signature were evaluated; only the expression levels of LINC02321
and KRT7-AS were significantly different when compared between the
tumour and normal groups (Fig.
4A). Therefore, the expression levels of LINC02321 and KRT7-AS
in tissues collected in the present study were evaluated by
RT-qPCR. LINC02321 expression levels were significantly increased
in bladder cancer tissue, whereas there was no statistically
significant difference in the expression levels of KRT7-AS compared
with the normal adjacent tissue (Fig.
4B). Therefore, LINC02321 was used as the target for subsequent
experiments.
LINC02321 silencing in bladder cancer
cells suppresses proliferation, migration and invasion, and
enhances chemosensitivity to cisplatin
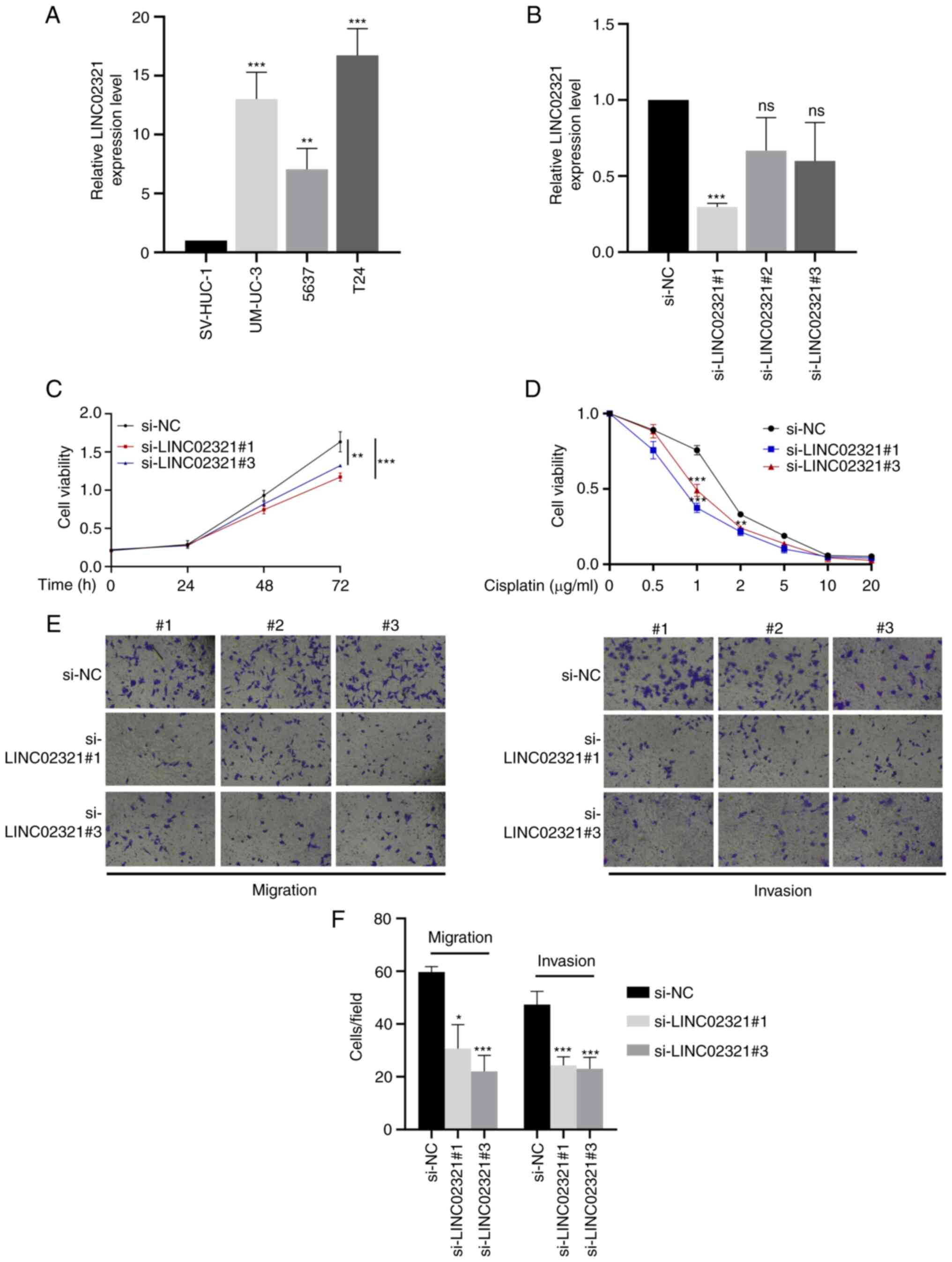
Expression of LINC02321 in bladder cancer cell lines
was assessed using RT-qPCR. Compared with SV-HUC-1 normal
uroepithelial immortalised cells, LINC02321 expression levels were
significantly increased in the bladder cancer cell lines T24, 5637
and UM-UC-3, with T24 and 5637 demonstrating the highest and the
lowest expression levels, respectively (Fig. 5A). Therefore, T24 was used for
knockdown of LINC02321 expression, for which si-LINC02321#1 and
si-LINC02321#3 were demonstrated to be the most efficient and were
selected for subsequent use (Fig.
5B). CCK-8 proliferation assay results demonstrated that cell
proliferation was significantly decreased after LINC02321 knockdown
compared with si-NC (Fig. 5C).
Furthermore, CCK-8 drug sensitivity assay demonstrated that the
sensitivity of the cells to cisplatin (1.0 and 2.0 µg/ml) was
increased significantly following LINC02321 knockdown compared with
the si-NC (Fig. 5D). Transwell
assays demonstrated that the invasive and migratory capability of
the cells decreased significantly after LINC02321 knockdown
compared with the si-NC (Fig. 5E and
F).
LINC02321 overexpression in bladder
cancer cells enhances proliferation, migration and invasion and
suppresses chemosensitivity to cisplatin
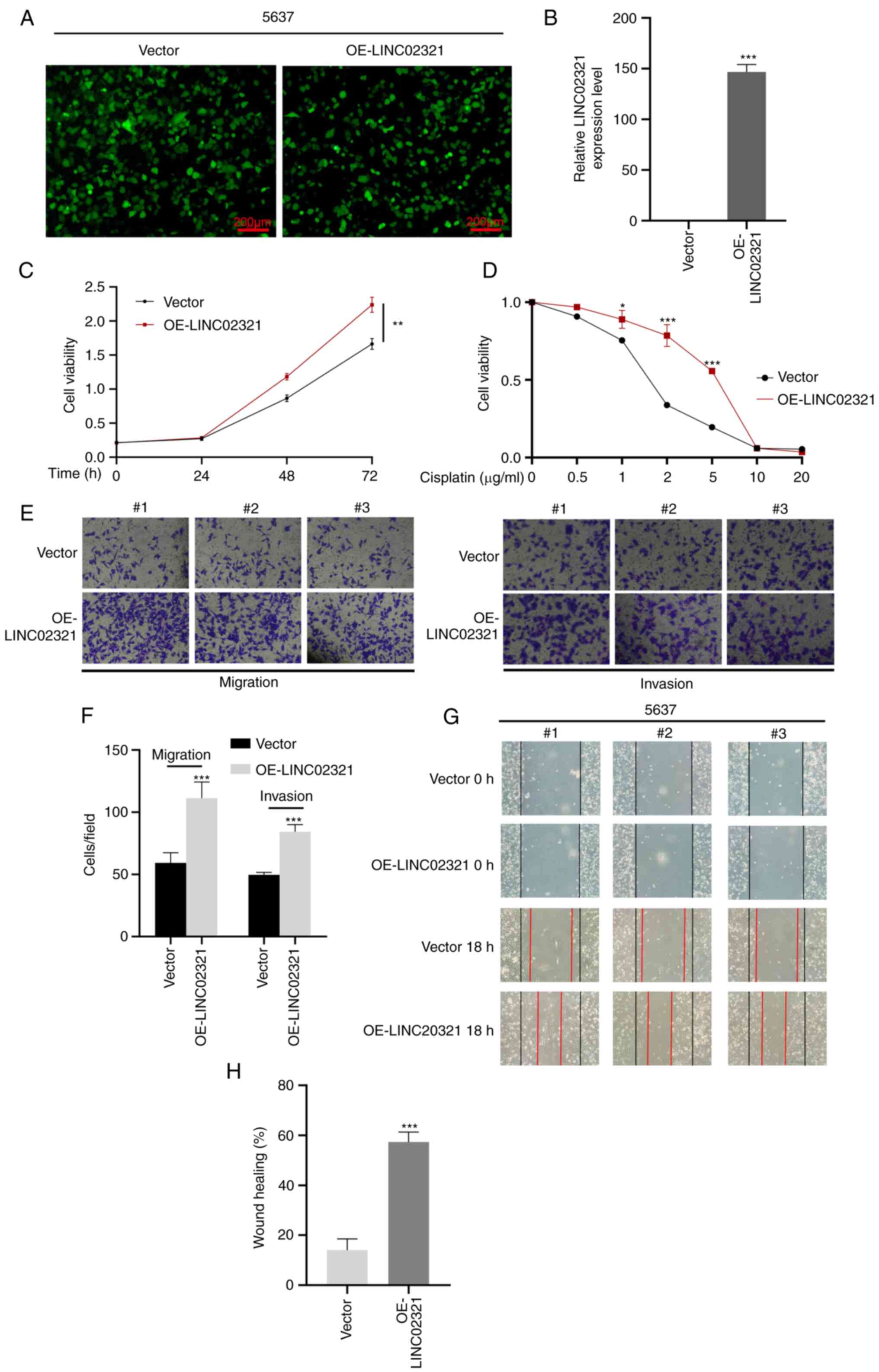
LINC02321 was stably overexpressed in the 5637
bladder cancer cells following transfection with the lentiviral
vector, which demonstrated high efficiency (Fig. 6A and B). CCK-8 proliferation assay
demonstrated that cell proliferation increased significantly after
LINC02321 overexpression compared with the empty vector control
(Fig. 6C). CCK-8 drug sensitivity
assay demonstrated that the sensitivity of the cells to 1, 2 and 5
µg/ml cisplatin decreased significantly after LINC02321
overexpression compared with the control (Fig. 6D). Transwell assays demonstrated
that the invasive and migratory abilities of cells significantly
increased after LINC02321 overexpression compared with the vector
control (Fig. 6E and F). Since
siRNA can affect the cell state, it is unsuitable for use in wound
healing experiments. Therefore, wound healing assays were performed
following LINC02321 overexpression, and the results demonstrated
significantly increased wound healing rate following LINC02321
overexpression compared with the control (Fig. 6G and H).
VEGFA signaling is involved in
LINC02321-regulated progression of bladder cancer
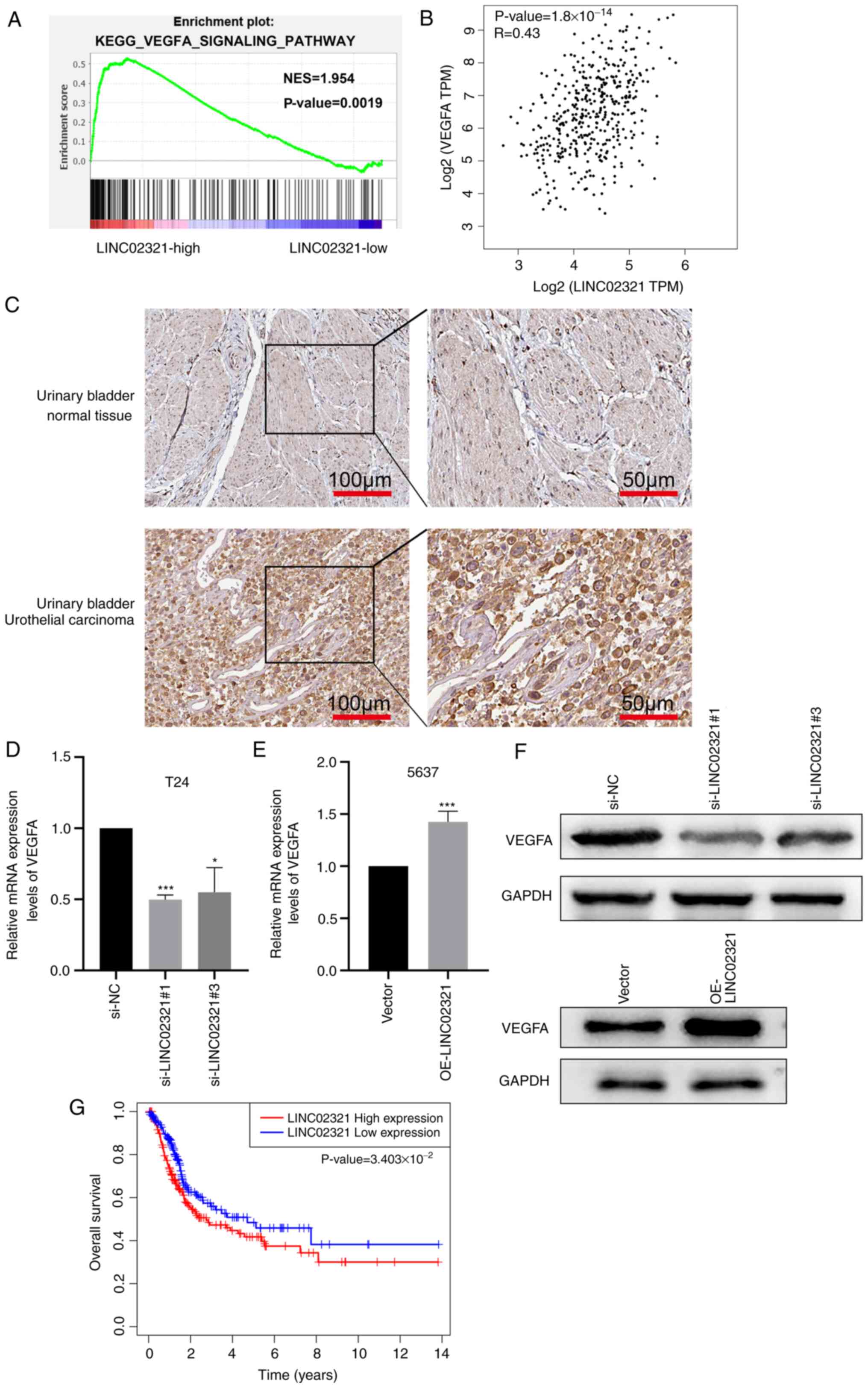
The downstream regulatory mechanism of LINC02321 was
evaluated. GSEA demonstrated that the expression levels of
LINC02321 in TCGA bladder cancer dataset was associated with the
VEGFA signalling (Fig. 7A).
Pearson's correlation analysis demonstrated that LINC02321
expression levels correlated positively with expression levels of
VEGFA (Fig. 7B).
Immunohistochemical staining images from the Human Protein Atlas
database (www.proteinatlas.org) demonstrated that the intensity
of VEGFA in bladder cancer tissues was markedly stronger compared
with that in the adjacent mucosa (Fig. 7C). LINC02321 regulated mRNA and
protein expression levels of VEGFA, which were assessed following
transfection with si-LINC02321 and OE-LINC02321 using RT-qPCR
(Fig. 7D and E) and western
blotting (Fig. 7F), respectively.
Furthermore, the association between LINC02321 expression and OS
were analysed in 402 patients with bladder cancer from TCGA
dataset. The survival time of patients with high LINC02321
expression levels was significantly shorter compared with that of
patients with low expression levels (Fig. 7G). These results suggested that
the VEGFA signalling pathway may be involved in LINC02321-regulated
progression of bladder cancer and that LINC02321 may be a potential
target for the treatment of this malignancy.
Discussion
Numerous biomarkers, including DNA methylation,
templates with single-nucleotide polymorphisms, protein or
metabolic changes and mRNA and non-coding RNA expression changes
are associated with the occurrence of disease in the body (18). In clinical practice, tumour
biomarkers are used to identify primary tumours and to screen
high-risk populations, and they are key for the prognosis and
prediction of the efficacy of treatments. Over the past decade,
numerous bladder cancer-associated tumour markers, such as bladder
tumour antigen series (19),
nuclear matrix protein 22 (20)
and fibrin degradation products (21), have been reported in clinical
practice. These tumour markers facilitate the detection of occult
bladder cancer in the clinic. To the best of our knowledge,
however, there are currently no specific markers of malignancy that
predict the survival and prognosis of patients with bladder cancer.
Therefore, novel biomarkers associated with the prognosis of
bladder cancer may have beneficial clinical applicability.
Bladder cancer metastasis of is one of the primary
factors that leads to the poor prognosis of patients. Previous
studies have reported that the mechanism of metastasis in bladder
cancer is associated with tumour angiogenesis (5,22).
Tumours become angiogenic through numerous methods, including
outgrowth and sleeve-in angiogenesis, angiogenesis with the
recruitment of endothelial progenitor cells and angiogenic mimicry
(23–26). Vasculogenesis, which is associated
with the recruitment of endothelial progenitor cells, is the main
mechanism of tumour angiogenesis, alongside vascular sprouting. The
initiation of neovascularisation requires activation of the
angiogenic switch, followed by expression of matrix
metalloproteinases to degrade the basement membrane (27). The endothelial cells proliferate
after migration to the corresponding locations, where blood vessels
are to be generated, under the action of chemokines and angiogenic
factors produced by autocrine and paracrine mechanisms and develop
into blood vessels with the support of tumour stromal cells
(6). During this process, tumour
and stromal cells interact to form a microenvironment suitable for
angiogenesis. The main biological function of angiogenesis in
malignant tumours is to enhance the ability of cells to metastasise
(28). Investigation of the key
components in angiogenesis and angiogenesis-driven processes in
bladder cancer may enable evaluation of the malignant features of
bladder cancer, such as invasion and metastasis, and the
identification of novel therapeutic targets.
lncRNAs occupy a large proportion of the genomes of
more complex forms of life and have been reported to serve a more
important role than the sequences that encode proteins (9). lncRNAs regulate the expression of
genes on multiple levels by regulation of chromatin modification,
RNA splicing and protein activity, which regulate occurrence,
development, prognosis and chemotherapy resistance of tumours
(10). Previous studies have
reported that lncRNAs can be used as novel biomarkers to predict
occurrence, development and prognosis of bladder cancer. Higher
expression levels of the serum exosome-derived lncRNA LNMAT2 have
been reported to be associated with shorter survival time, which
suggests that it may serve as a potential diagnostic biomarker and
therapeutic target for lymph node metastasis in bladder cancer
(29). Furthermore, normal
bladder tissue-derived exosomes have been reported to inhibit
malignant progression of bladder cancer through the lncRNA PTENP1,
which may be used as a biomarker for the prediction of the survival
of patients with bladder cancer (30). Another study reported that an
angiogenesis-associated lncRNA signature based on data from TCGA
database effectively predicted the prognosis of patients with
bladder cancer, (31) which was
similar to the results of the present study; therefore, these
findings may provide a new perspective and novel antiangiogenic
targets for clinical diagnosis and treatment strategies of bladder
cancer. Furthermore, an epithelial-mesenchymal
transition-associated 14-lncRNA signature was reported to
effectively predict progression of bladder cancer and prognosis of
patients (32). Another
extracellular matrix-associated six-lncRNA signature was previously
proposed to serve as a novel marker for the prediction of prognosis
of patients with bladder cancer (33). With further study into the
mechanism of lncRNAs that underlie tumorigenesis and development,
the diagnostic performance of lncRNAs in progression and prognosis
of bladder cancer will likely continue to improve.
Both lncRNAs and angiogenesis serve important roles
in the prediction of the progression of bladder cancer. Therefore,
in the present study, bioinformatics was used to analyse the
predictive performance of an angiogenesis-associated lncRNA
signature in the prognosis of bladder cancer, and six lncRNAs were
identified to be significantly associated with OS of patients with
bladder cancer. Therefore, a signature based on these six
angiogenesis-associated lncRNAs was constructed. Patients were
divided based on risk score of the signature, and the OS of
patients in the low-risk group was significantly longer compared
with that of patients in the high-risk group; this suggested that
the six angiogenesis-associated lncRNA signature in the present
study aided the prediction of the prognosis of patients with
bladder cancer. A nomogram was constructed, and compared with the
other clinical indicators, the risk score of the signature
demonstrated markedly higher predictive power for patient
prognosis.
From the six angiogenesis-associated lncRNA
signature, RT-qPCR and TCGA data analysis demonstrated that
LINC02321 expression levels differed significantly between bladder
cancer and normal tissue. It was subsequently demonstrated that
LINC02321 expression correlated positively with that of VEGFA, a
key member of the VEGF family (34). VEGFA has been reported to serve an
important role in the promotion of tumour progression (34). It has been reported that certain
lncRNAs promote tumour progression through regulation of VEGFA. For
example, lncRNA PVT1 was previously reported to promote gastric
cancer progression by activation of the STAT3/VEGFA axis (35). Furthermore, lncRNA TUSC8 has been
reported to affect proliferation and migration of oesophageal
cancer cells through the regulation of VEGFA (36). In another study, LINC00707 was
reported to regulate the VEGFA pathway to enhance the progression
of cervical cancer (37). The
lncRNA SNHG16 has been reported to drive the proliferation,
migration and invasion of lung cancer cells by modulating of the
VEGF axis (38). In the present
study, it was demonstrated that LINC02321 positively regulated
VEGFA mRNA and protein expression levels. Previous studies have
reported that VEGFA directly promotes malignant progression of
bladder cancer (39,40). Therefore, based on results from
the present study and those of previous studies, it was
hypothesised that LINC02321 promotes the malignant progression of
bladder cancer via the VEGFA signalling pathway. Finally, the
prognostic role of LINC02321 in patients with bladder cancer was
analysed. The prognosis of patients with high LINC02321 expression
levels was significantly poorer compared with those of patients
with low LINC02321 expression levels. These results suggested that
LINC02321 may promote the malignant progression of bladder cancer
and may serve as a novel therapeutic target.
In conclusion, the present study evaluated the
predictive role of an angiogenesis-associated lncRNA signature in
the prognosis of patients with bladder cancer. The role of
LINC02321 in the progression of bladder cancer was also assessed.
Results from the present study provide a novel path for the
exploration of the mechanism that underlies bladder cancer
progression and potential therapeutic targets.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was funded by Zhao Kang and Tao Li.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
ZK and TL designed the study; ZK and QD performed
the experiments and wrote the manuscript; ZK, QD, TH, MTT, YPZ and
MW analyzed the results. QD revised the manuscript according to the
review comments. All authors confirmed the authenticity of all the
raw data. All authors agree to be accountable for all aspects of
the research in ensuring that the accuracy or integrity of any part
of the work are appropriately assessed and resolved. All authors
have read and approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of The First Affiliated Hospital of Chongqing Medical
University (Chongqing, China; approval no. 2021069). Written
informed consent of patients was obtained before the
experiment.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Tran L, Xiao JF, Agarwal N, Duex JE and
Theodorescu D: Advances in bladder cancer biology and therapy. Nat
Rev Cancer. 21:104–121. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Richters A, Aben KKH and Kiemeney LALM:
The global burden of urinary bladder cancer: An update. World J
Urol. 38:1895–1904. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Patel VG, Oh WK and Galsky MD: Treatment
of muscle-invasive and advanced bladder cancer in 2020. CA Cancer J
Clin. 70:404–423. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lenis AT, Lec PM, Chamie K and Mshs MD:
Bladder cancer: A review. JAMA. 324:1980–1991. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Fus ŁP and Górnicka B: Role of
angiogenesis in urothelial bladder carcinoma. Cent European J Urol.
69:258–263. 2016.PubMed/NCBI
|
|
6
|
Sonpavde G and Bellmunt J: Bladder cancer:
Angiogenesis as a therapeutic target in urothelial carcinoma. Nat
Rev Urol. 13:306–307. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wigner P, Grębowski R, Bijak M,
Saluk-Bijak J and Szemraj J: The interplay between oxidative
stress, inflammation and angiogenesis in bladder cancer
development. Int J Mol Sci. 22:44832021. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhang K, Liu D, Zhao J, Shi S, He X, Da P,
You Y and You B: Nuclear exosome HMGB3 secreted by nasopharyngeal
carcinoma cells promotes tumour metastasis by inducing
angiogenesis. Cell Death Dis. 12:5542021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Goodall GJ and Wickramasinghe VO: RNA in
cancer. Nat Rev Cancer. 21:22–36. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yan H and Bu P: Non-coding RNA in cancer.
Essays Biochem. 65:625–639. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li Y, Li G, Guo X, Yao H, Wang G and Li C:
Non-coding RNA in bladder cancer. Cancer Lett. 485:38–44. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kirkali Z, Chan T, Manoharan M, Algaba F,
Busch C, Cheng L, Kiemeney L, Kriegmair M, Montironi R, Murphy WM,
et al: Bladder cancer: Epidemiology, staging and grading, and
diagnosis. Urology. 66 (6 Suppl 1):S4–S34. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang G and McKenney JK: Urinary bladder
pathology: World Health Organization classification and American
joint committee on cancer staging update. Arch Pathol Lab Med.
143:571–577. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Witjes JA, Bruins HM, Cathomas R, Compérat
EM, Cowan NC, Gakis G, Hernández V, Linares Espinós E, Lorch A,
Neuzillet Y, et al: European association of urology guidelines on
muscle-invasive and metastatic bladder cancer: Summary of the 2020
guidelines. Eur Urol. 79:82–104. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gao S, Yin H, Tong H, Zhan K, Yang G,
Hossain MA, Li T, Gou X and He W: Nucleolar and spindle associated
protein 1 (NUSAP1) promotes bladder cancer progression through the
TGF-β signaling pathway. Onco Targets Ther. 13:813–825. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhan X, Jiang M, Deng W, Liu X, Chen L and
Fu B: Development and validation of a prognostic nomogram for
predicting cancer-specific survival in patients with lymph node
positive bladder cancer: A study based on SEER database. Front
Oncol. 12:7890282022. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Cao Z, Zhao K, Jose I, Hoogenraad NJ and
Osellame LD: Biomarkers for cancer cachexia: A mini review. Int J
Mol Sci. 22:45012021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Patel SP and Kurzrock R: PD-L1 expression
as a predictive biomarker in cancer immunotherapy. Mol Cancer Ther.
14:847–856. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang J, Zhao X, Jiang XL, Lu D, Yuan Q and
Li J: Diagnostic performance of nuclear matrix protein 22 and urine
cytology for bladder cancer: A meta-analysis. Diagn Cytopathol.
50:300–312. 2022. View
Article : Google Scholar : PubMed/NCBI
|
|
21
|
Li X, Shu K, Zhou J, Yu Q, Cui S, Liu J,
Zhou R and Ding D: Preoperative plasma fibrinogen and D-dimer as
prognostic biomarkers for non-muscle-invasive bladder cancer. Clin
Genitourin Cancer. 18:11–19. e12020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chan TC, Hsing CH, Shiue YL, Huang SK,
Hsieh KL, Kuo YH and Li CF: Angiogenesis driven by the
CEBPD-hsa-miR-429-VEGFA signaling axis promotes urothelial
carcinoma progression. Cells. 11:6382022. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Viallard C and Larrivée B: Tumor
angiogenesis and vascular normalization: Alternative therapeutic
targets. Angiogenesis. 20:409–426. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Oguntade AS, Al-Amodi F, Alrumayh A,
Alobaida M and Bwalya M: Anti-angiogenesis in cancer therapeutics:
The magic bullet. J Egypt Natl Canc Inst. 33:152021. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hariprabu KNG, Sathya M and Vimalraj S:
CRISPR/Cas9 in cancer therapy: A review with a special focus on
tumor angiogenesis. Int J Biol Macromol. 192:913–930. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Armani G, Pozzi E, Pagani A, Porta C,
Rizzo M, Cicognini D, Rovati B, Moccia F, Pedrazzoli P and Ferraris
E: The heterogeneity of cancer endothelium: The relevance of
angiogenesis and endothelial progenitor cells in cancer
microenvironment. Microvasc Res. 138:1041892021. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Vallée A, Guillevin R and Vallée JN:
Vasculogenesis and angiogenesis initiation under normoxic
conditions through Wnt/β-catenin pathway in gliomas. Rev Neurosci.
29:71–91. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Schaaf MB, Houbaert D, Meçe O and
Agostinis P: Autophagy in endothelial cells and tumor angiogenesis.
Cell Death Differ. 26:665–679. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chen C, Luo Y, He W, Zhao Y, Kong Y, Liu
H, Zhong G, Li Y, Li J, Huang J, et al: Exosomal long noncoding RNA
LNMAT2 promotes lymphatic metastasis in bladder cancer. J Clin
Invest. 130:404–421. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zheng R, Du M, Wang X, Xu W, Liang J, Wang
W, Lv Q, Qin C, Chu H, Wang M, et al: Exosome-transmitted long
non-coding RNA PTENP1 suppresses bladder cancer progression. Mol
Cancer. 17:1432018. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Li X, Zhang C, Peng X, Li Y, Chen G, Gou
X, Zhou X and Ma C: A novel risk score model based on five
angiogenesis-related long non-coding RNAs for bladder urothelial
carcinoma. Cancer Cell Int. 22:1572022. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tong H, Li T, Gao S, Yin H, Cao H and He
W: An epithelial-mesenchymal transition-related long noncoding RNA
signature correlates with the prognosis and progression in patients
with bladder cancer. Biosci Rep. 41:BSR202039442021. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Qing L, Gu P, Liu M, Shen J, Liu X, Guang
R, Ke K, Huang Z, Lee W and Zhao H: Extracellular matrix-related
six-lncRNA signature as a novel prognostic biomarker for bladder
cancer. Onco Targets Ther. 13:12521–12538. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Claesson-Welsh L and Welsh M: VEGFA and
tumour angiogenesis. J Intern Med. 273:114–127. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhao J, Du P, Cui P, Qin Y, Hu C, Wu J,
Zhou Z, Zhang W, Qin L and Huang G: LncRNA PVT1 promotes
angiogenesis via activating the STAT3/VEGFA axis in gastric cancer.
Oncogene. 37:4094–4109. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Hu R, Bi R, Jiang L, Yang X, Zhong Y and
Xie X: LncRNA TUSC8 suppresses the proliferation and migration of
esophageal cancer cells by downregulation of VEGFA. J Cancer.
12:6393–6400. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Guo H, Li J, Fan F and Zhou P: LINC00707
regulates miR-382-5p/VEGFA pathway to enhance cervical cancer
progression. J Immunol Res. 2021:55246322021. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Chen L, Qiu CH, Chen Y, Wang Y, Zhao JJ
and Zhang M: LncRNA SNHG16 drives proliferation, migration, and
invasion of lung cancer cell through modulation of miR-520/VEGF
axis. Eur Rev Med Pharmacol Sci. 24:9522–9531. 2020.PubMed/NCBI
|
|
39
|
Cao W, Zhao Y, Wang L and Huang X:
Circ0001429 regulates progression of bladder cancer through binding
miR-205-5p and promoting VEGFA expression. Cancer Biomark.
25:101–113. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wang H, Niu X, Jiang H, Mao F, Zhong B,
Jiang X and Fu G: Long non-coding RNA DLX6-AS1 facilitates bladder
cancer progression through modulating miR-195-5p/VEGFA signaling
pathway. Aging (Albany NY). 12:16021–16034. 2020. View Article : Google Scholar : PubMed/NCBI
|