|
1
|
Cho NH, Shaw JE, Karuranga S, Huang Y, da
Rocha Fernandes JD, Ohlrogge AW and Malanda B: IDF Diabetes Atlas:
Global estimates of diabetes prevalence for 2017 and projections
for 2045. Diabetes Res Clin Pract. 138:271–281. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Morrish NJ, Wang SL, Stevens LK, Fuller JH
and Keen H: Mortality and causes of death in the WHO multinational
study of vascular disease in diabetes. Diabetologia. 44 (Suppl
2):S14–S21. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wang PL, Bao Y, Yee MC, Barrett SP, Hogan
GJ, Olsen MN, Dinneny JR, Brown PO and Salzman J: Circular RNA is
expressed across the eukaryotic tree of life. PLoS One.
9:e908592014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Salzman J, Chen RE, Olsen MN, Wang PL and
Brown PO: Cell-type specific features of circular RNA expression.
PLoS Genet. 9:e10037772013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhang HD, Jiang LH, Sun DW, Hou JC and Ji
ZL: CircRNA: A novel type of biomarker for cancer. Breast Cancer.
25:1–7. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Qu S, Yang X, Li X, Wang J, Gao Y, Shang
R, Sun W, Dou K and Li H: Circular RNA: A new star of noncoding
RNAs. Cancer Lett. 365:141–148. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jiang G, Ma Y, An T, Pan Y, Mo F, Zhao D,
Liu Y, Miao JN, Gu YJ, Wang Y and Gao SH: Relationships of circular
RNA with diabetes and depression. Sci Rep. 7:72852017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Memczak S, Jens M, Elefsinioti A, Torti F,
Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer
M, et al: Circular RNAs are a large class of animal RNAs with
regulatory potency. Nature. 495:333–338. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Fan X, Weng X, Zhao Y, Chen W, Gan T and
Xu D: Circular RNAs in cardiovascular disease: An overview. Biomed
Res Int. 2017:51357812017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J,
Chen D, Gu J, He X and Huang S: Circular RNA is enriched and stable
in exosomes: A promising biomarker for cancer diagnosis. Cell Res.
25:981–984. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Smid M, Wilting SM, Uhr K,
Rodríguez-González FG, de Weerd V, Prager-Van der Smissen WJC, van
der Vlugt-Daane M, van Galen A, Nik-Zainal S, Butler A, et al: The
circular RNome of primary breast cancer. Genome Res. 29:356–366.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
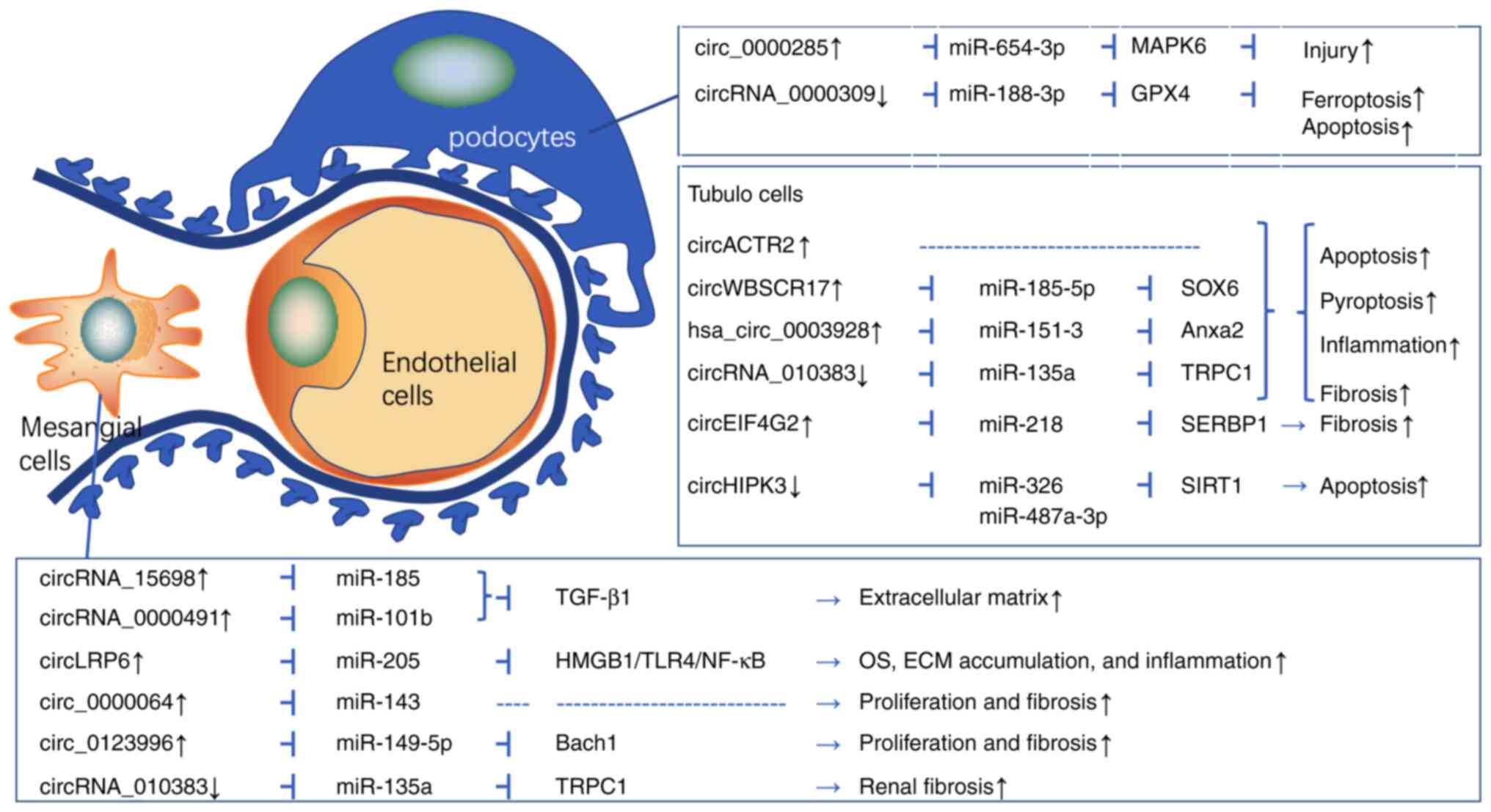
Hu W, Han Q, Zhao L and Wang L: Circular
RNA circRNA_15698 aggravates the extracellular matrix of diabetic
nephropathy mesangial cells via miR-185/TGF-β1. J Cell Physiol.
234:1469–1476. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Shan K, Liu C, Liu BH, Chen X, Dong R, Liu
X, Zhang YY, Liu B, Zhang SJ, Wang JJ, et al: Circular noncoding
RNA HIPK3 mediates retinal vascular dysfunction in diabetes
mellitus. Circulation. 136:1629–1642. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang W, Zhang S, Xu L, Feng Y, Wu X, Zhang
M, Yu Z and Zhou X: Involvement of circHIPK3 in the pathogenesis of
diabetic cardiomyopathy in mice. Diabetologia. 64:681–692. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
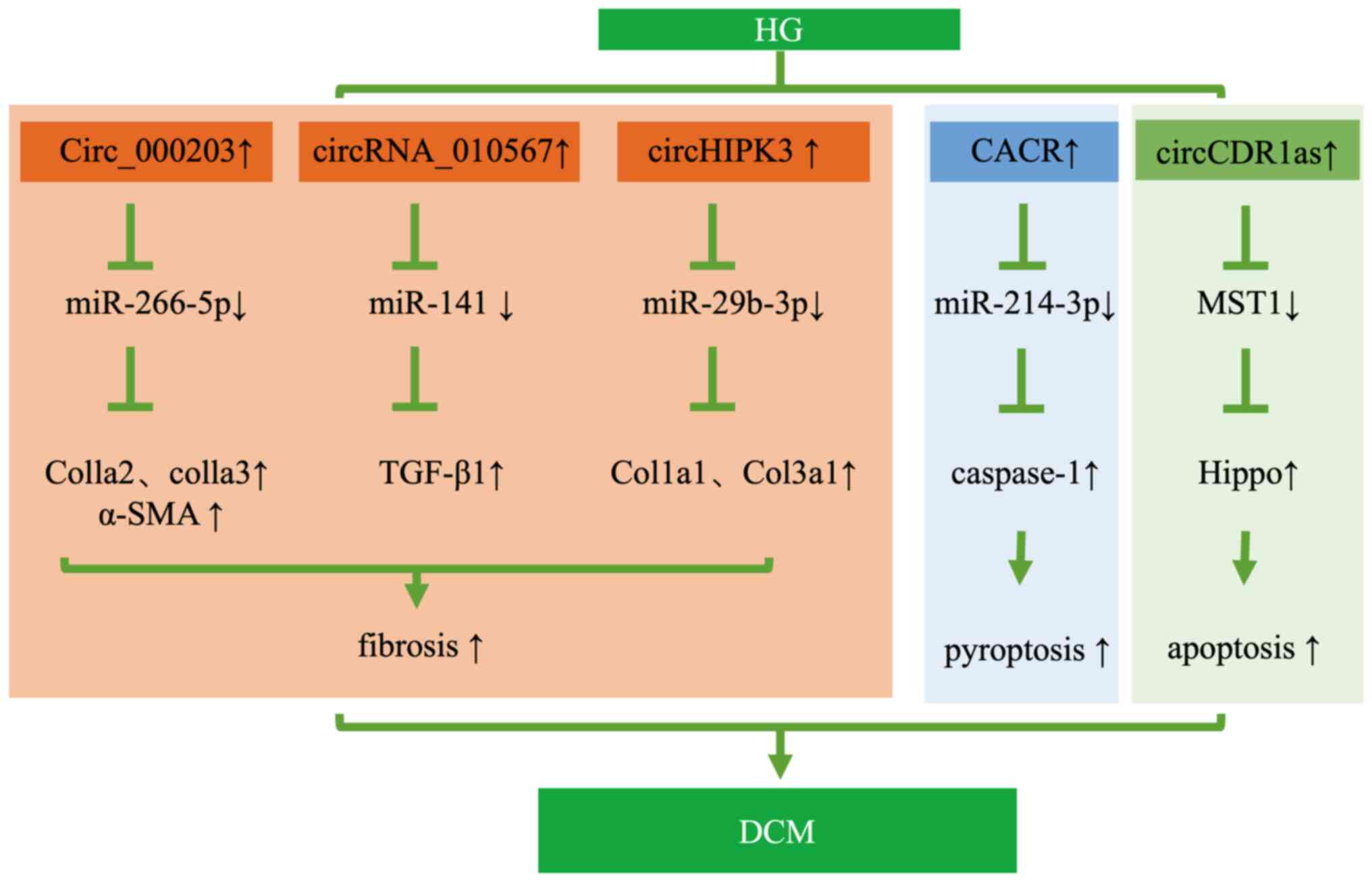
Tang CM, Zhang M, Huang L, Hu ZQ, Zhu JN,
Xiao Z, Zhang Z, Lin QX, Zheng XL, Yang M, et al: CircRNA_000203
enhances the expression of fibrosis-associated genes by
derepressing targets of miR-26b-5p, Col1a2 and CTGF, in cardiac
fibroblasts. Sci Rep. 7:403422017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yang F, Li A, Qin Y, Che H, Wang Y, Lv J,
Li Y, Li H, Yue E, Ding X, et al: A novel circular RNA mediates
pyroptosis of diabetic cardiomyopathy by functioning as a competing
endogenous RNA. Mol Ther Nucleic Acids. 17:636–643. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kristensen LS, Andersen MS, Stagsted LVW,
Ebbesen KK, Hansen TB and Kjems J: The biogenesis, biology and
characterization of circular RNAs. Nat Rev Genet. 20:675–691. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jeck WR, Sorrentino JA, Wang K, Slevin MK,
Burd CE, Liu J, Marzluff WF and Sharpless NE: Circular RNAs are
abundant, conserved, and associated with ALU repeats. RNA.
19:141–157. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Bahn JH, Zhang Q, Li F, Chan TM, Lin X,
Kim Y, Wong DT and Xiao X: The landscape of microRNA,
Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem.
61:221–230. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Memczak S, Papavasileiou P, Peters O and
Rajewsky N: Identification and characterization of circular RNAs As
a new class of putative biomarkers in human blood. PLoS One.
10:e01412142015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ashwal-Fluss R, Meyer M, Pamudurti N,
Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N and
Kadener S: circRNA biogenesis competes with pre-mRNA splicing. Mol
Cell. 56:55–66. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Liu CX, Li X, Nan F, Jiang S, Gao X, Guo
SK, Xue W, Cui Y, Dong K, Ding H, et al: Structure and degradation
of circular RNAs regulate PKR activation in innate immunity. Cell.
177:865–880.e21. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hansen TB, Wiklund ED, Bramsen JB,
Villadsen SB, Statham AL, Clark SJ and Kjems J: miRNA-dependent
gene silencing involving Ago2-mediated cleavage of a circular
antisense RNA. EMBO J. 30:4414–4422. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang Y, Liu J, Ma J, Sun T, Zhou Q, Wang
W, Wang G, Wu P, Wang H, Jiang L, et al: Exosomal circRNAs:
Biogenesis, effect and application in human diseases. Mol Cancer.
18:1162019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL
and Yang L: Complementary sequence-mediated exon circularization.
Cell. 159:134–147. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yoshimoto R, Rahimi K, Hansen TB, Kjems J
and Mayeda A: Biosynthesis of Circular RNA ciRS-7/CDR1as is
mediated by mammalian-wide interspersed repeats. iScience.
23:1013452020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hansen TB, Jensen TI, Clausen BH, Bramsen
JB, Finsen B, Damgaard CK and Kjems J: Natural RNA circles function
as efficient microRNA sponges. Nature. 495:384–388. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Piwecka M, Glažar P, Hernandez-Miranda LR,
Memczak S, Wolf SA, Rybak-Wolf A, Filipchyk A, Klironomos F, Cerda
Jara CA, Fenske P, et al: Loss of a mammalian circular RNA locus
causes miRNA deregulation and affects brain function. Science.
357:eaam85262017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kristensen LS, Okholm TLH, Venø MT and
Kjems J: Circular RNAs are abundantly expressed and upregulated
during human epidermal stem cell differentiation. RNA Biol.
15:280–291. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hsiao KY, Lin YC, Gupta SK, Chang N, Yen
L, Sun HS and Tsai SJ: Noncoding effects of circular RNA CCDC66
promote colon cancer growth and metastasis. Cancer Res.
77:2339–2350. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Cao Y, Yuan G, Zhang Y and Lu R: High
glucose-induced circHIPK3 downregulation mediates endothelial cell
injury. Biochem Biophys Res Commun. 507:362–368. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Barbagallo D, Caponnetto A, Brex D,
Mirabella F, Barbagallo C, Lauretta G, Morrone A, Certo F, Broggi
G, Caltabiano R, et al: CircSMARCA5 Regulates VEGFA mRNA splicing
and angiogenesis in glioblastoma multiforme through the binding of
SRSF1. Cancers (Basel). 11:1942019. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Abdelmohsen K, Panda AC, Munk R,
Grammatikakis I, Dudekula DB, De S, Kim J, Noh JH, Kim KM,
Martindale JL and Gorospe M: Identification of HuR target circular
RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA
Biol. 14:361–369. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Li XX, Xiao L, Chung HK, Ma XX, Liu X,
Song JL, Jin CZ, Rao JN, Gorospe M and Wang JY: Interaction between
HuR and circPABPN1 Modulates autophagy in the intestinal epithelium
by altering ATG16L1 translation. Mol Cell Biol. 40:e004922020.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Abe N, Matsumoto K, Nishihara M, Nakano Y,
Shibata A, Maruyama H, Shuto S, Matsuda A, Yoshida M, Ito Y and Abe
H: Rolling circle translation of circular RNA in living human
cells. Sci Rep. 5:164352015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Meyer KD, Patil DP, Zhou J, Zinoviev A,
Skabkin MA, Elemento O, Pestova TV, Qian SB and Jaffrey SR: 5′ UTR
m(6)A Promotes Cap-Independent Translation. Cell. 163:999–1010.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Li Y, Wang Z, Su P, Liang Y, Li Z, Zhang
H, Song X, Han D, Wang X, Liu Y, et al: Circ-EIF6 encodes
EIF6-224aa to promote TNBC progression via stabilizing MYH9 and
activating Wnt/beta-catenin pathway. Mol Ther. 30:415–430. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Han YN, Xia SQ, Zhang YY, Zheng JH and Li
W: Circular RNAs: A novel type of biomarker and genetic tools in
cancer. Oncotarget. 8:64551–64563. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhang Y, Zhang XO, Chen T, Xiang JF, Yin
QF, Xing YH, Zhu S, Yang L and Chen LL: Circular intronic long
noncoding RNAs. Mol Cell. 51:792–806. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Rines AK, Sharabi K, Tavares CD and
Puigserver P: Targeting hepatic glucose metabolism in the treatment
of type 2 diabetes. Nat Rev Drug Discov. 15:786–804. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Shang FF, Luo S, Liang X and Xia Y:
Alterations of circular RNAs in hyperglycemic human endothelial
cells. Biochem Biophys Res Commun. 499:551–555. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Jin G, Wang Q, Hu X, Li X, Pei X, Xu E and
Li M: Profiling and functional analysis of differentially expressed
circular RNAs in high glucose-induced human umbilical vein
endothelial cells. FEBS Open Bio. 9:1640–1651. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
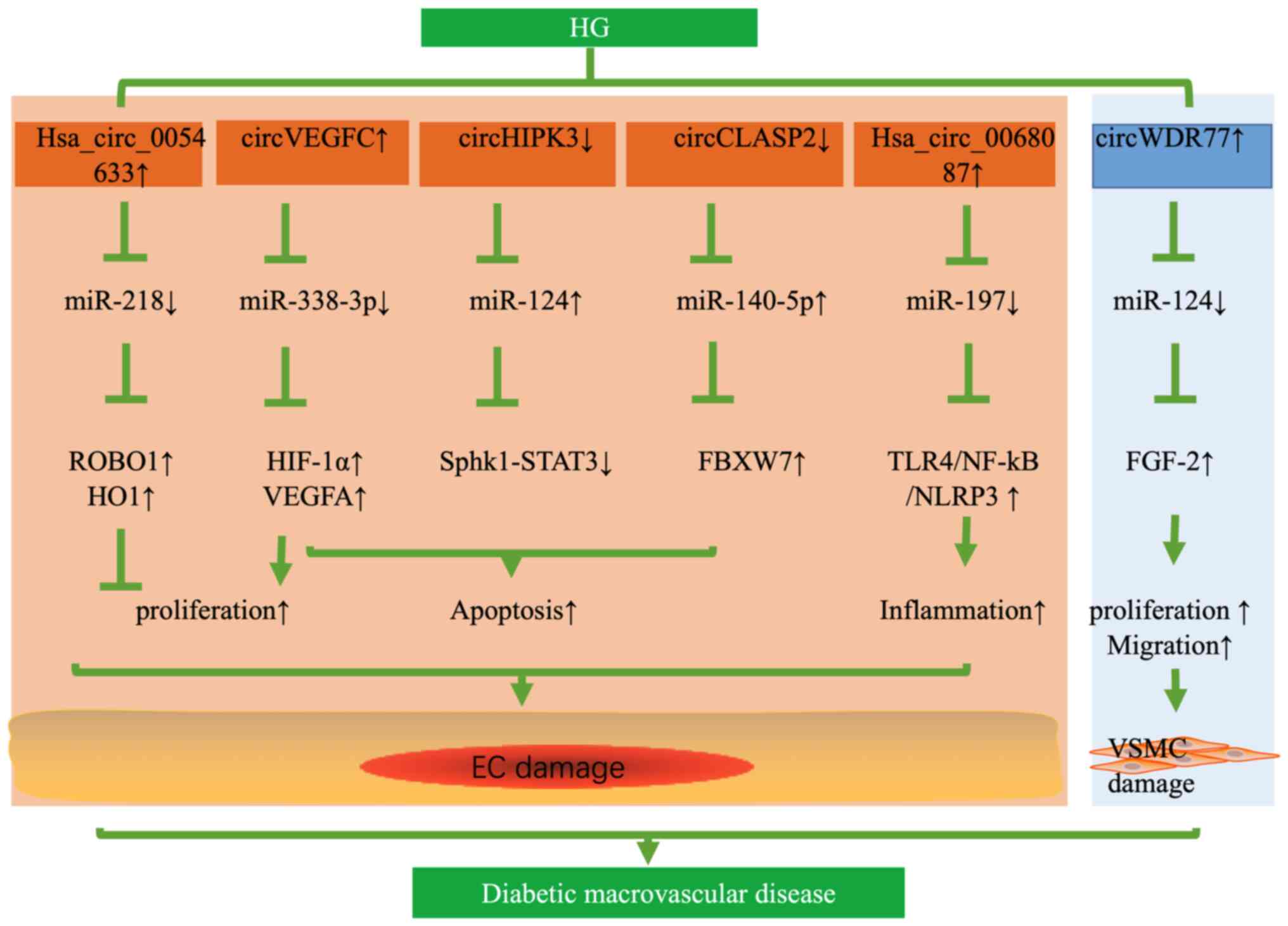
Pan L, Lian W, Zhang X, Han S, Cao C, Li X
and Li M: Human circular RNA-0054633 regulates high glucose-induced
vascular endothelial cell dysfunction through the
microRNA-218/roundabout 1 and microRNA-218/heme oxygenase-1 axes.
Int J Mol Med. 42:597–606. 2018.PubMed/NCBI
|
|
44
|
Zhang Q, Long J, Li N, Ma X and Zheng L:
Circ_CLASP2 regulates high glucose-induced dysfunction of human
endothelial cells through targeting miR-140-5p/FBXW7 Axis. Front
Pharmacol. 12:5947932021. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Wei H, Cao C, Wei X, Meng M, Wu B, Meng L,
Wei X, Gu S and Li H: Circular RNA circVEGFC accelerates high
glucose-induced vascular endothelial cells apoptosis through
miR-338-3p/HIF-1α/VEGFA axis. Aging (Albany NY). 12:14365–14375.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Cheng J, Liu Q, Hu N, Zheng F, Zhang X, Ni
Y and Liu J: Downregulation of hsa_circ_0068087 ameliorates
TLR4/NF-κB/NLRP3 inflammasome-mediated inflammation and endothelial
cell dysfunction in high glucose conditioned by sponging miR-197.
Gene. 709:1–7. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen J, Cui L, Yuan J, Zhang Y and Sang H:
Circular RNA WDR77 target FGF-2 to regulate vascular smooth muscle
cells proliferation and migration by sponging miR-124. Biochem
Biophys Res Commun. 494:126–132. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zaiou M: circRNAs signature as potential
diagnostic and prognostic biomarker for diabetes mellitus and
related cardiovascular complications. Cells. 9:6592020. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Fang Y, Wang X, Li W, Han J, Jin J, Su F,
Zhang J, Huang W, Xiao F, Pan Q and Zou L: Screening of circular
RNAs and validation of circANKRD36 associated with inflammation in
patients with type 2 diabetes mellitus. Int J Mol Med.
42:1865–1874. 2018.PubMed/NCBI
|
|
50
|
An Y, Furber KL and Ji S: Pseudogenes
regulate parental gene expression via ceRNA network. J Cell Mol
Med. 21:185–192. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Peng F, Gong W, Li S, Yin B, Zhao C, Liu
W, Chen X, Luo C, Huang Q, Chen T, et al: circRNA_010383 Acts as a
Sponge for miR-135a, and its downregulated expression contributes
to renal fibrosis in diabetic nephropathy. Diabetes. 70:603–615.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Chen B, Li Y, Liu Y and Xu Z: circLRP6
regulates high glucose-induced proliferation, oxidative stress, ECM
accumulation, and inflammation in mesangial cells. J Cell Physiol.
234:21249–21259. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Wang W, Feng J, Zhou H and Li Q:
Circ_0123996 promotes cell proliferation and fibrosisin mouse
mesangial cells through sponging miR-149-5p and inducing Bach1
expression. Gene. 761:1449712020. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Ge X, Xi L, Wang Q, Li H, Xia L, Cang Z,
Peng W and Huang S: Circular RNA Circ_0000064 promotes the
proliferation and fibrosis of mesangial cells via miR-143 in
diabetic nephropathy. Gene. 758:1449522020. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Yao T, Zha D, Hu C and Wu X: Circ_0000285
promotes podocyte injury through sponging miR-654-3p and activating
MAPK6 in diabetic nephropathy. Gene. 747:1446612020. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Li G, Qin Y, Qin S, Zhou X, Zhao W and
Zhang D: Circ_WBSCR17 aggravates inflammatory responses and
fibrosis by targeting miR-185-5p/SOX6 regulatory axis in high
glucose-induced human kidney tubular cells. Life Sci.
259:1182692020. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
An L, Ji D, Hu W, Wang J, Jin X, Qu Y and
Zhang N: Interference of Hsa_circ_0003928 alleviates high
glucose-induced cell apoptosis and inflammation in HK-2 Cells via
miR-151-3p/Anxa2. Diabetes Metab Syndr Obes. 13:3157–3168. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Mou X, Chenv JW, Zhou DY, Liu K, Chen LJ,
Zhou D and Hu YB: A novel identified circular RNA, circ_0000491,
aggravates the extracellular matrix of diabetic nephropathy
glomerular mesangial cells through suppressing miR-101b by
targeting TGFβRI. Mol Med Rep. 22:3785–3794. 2020.PubMed/NCBI
|
|
59
|
Wen S, Li S, Li L and Fan Q: circACTR2: A
novel mechanism regulating high glucose-induced fibrosis in renal
tubular cells via pyroptosis. Biol Pharm Bull. 43:558–564. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Xu B, Wang Q, Li W, Xia L, Ge X, Shen L,
Cang Z, Peng W, Shao K and Huang S: Circular RNA circEIF4G2
aggravates renal fibrosis in diabetic nephropathy by sponging
miR-218. J Cell Mol Med. 26:1799–1805. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Zhuang L, Wang Z, Hu X, Yang Q, Pei X and
Jin G: CircHIPK3 alleviates high glucose toxicity to human renal
tubular epithelial HK-2 cells through regulation of
miR-326/miR-487a-3p/SIRT1. Diabetes Metab Syndr Obes. 14:729–740.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Jin J, Wang Y, Zheng D, Liang M and He Q:
A novel identified circular RNA, mmu_mmu_circRNA_0000309, involves
in germacrone-mediated improvement of diabetic nephropathy through
regulating ferroptosis by targeting miR-188-3p/GPX4 signaling axis.
Antioxid Redox Signal. 36:740–759. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Antonetti DA, Silva PS and Stitt AW:
Current understanding of the molecular and cellular pathology of
diabetic retinopathy. Nat Rev Endocrinol. 17:195–206. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Gu Y, Ke G, Wang L, Zhou E, Zhu K and Wei
Y: Altered expression profile of circular RNAs in the serum of
patients with diabetic retinopathy revealed by microarray.
Ophthalmic Res. 58:176–184. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Liu C, Ge HM, Liu BM, Dong R, Shan K, Chen
X, Yao MD, Li XM, Yao J, Zhou RM, et al: Targeting
pericyte-endothelial cell crosstalk by circular RNA-cPWWP2A
inhibition aggravates diabetes-induced microvascular dysfunction.
Proc Natl Acad Sci USA. 116:7455–7464. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Jiang Q, Liu C, Li CP, Xu SS, Yao MD, Ge
HM, Sun YN, Li XM, Zhang SJ, Shan K, et al: Circular RNA-ZNF532
regulates diabetes-induced retinal pericyte degeneration and
vascular dysfunction. J Clin Invest. 130:3833–3847. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
He M, Wang W, Yu H, Wang D, Cao D, Zeng Y,
Wu Q, Zhong P, Cheng Z, Hu Y and Zhang L: Comparison of expression
profiling of circular RNAs in vitreous humour between diabetic
retinopathy and non-diabetes mellitus patients. Acta Diabetol.
57:479–489. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Wu Z, Liu B, Ma Y, Chen H, Wu J and Wang
J: Discovery and validation of hsa_circ_0001953 as a potential
biomarker for proliferative diabetic retinopathy in human blood.
Acta Ophthalmol. 99:306–313. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Zhang SJ, Chen X, Li CP, Li XM, Liu C, Liu
BH, Shan K, Jiang Q, Zhao C and Yan B: Identification and
characterization of circular RNAs as a new class of putative
biomarkers in diabetes retinopathy. Invest Ophthalmol Vis Sci.
58:6500–6509. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Liu C, Yao MD, Li CP, Shan K, Yang H, Wang
JJ, Liu B, Li XM, Yao J, Jiang Q and Yan B: Silencing of circular
RNA-ZNF609 ameliorates vascular endothelial dysfunction.
Theranostics. 7:2863–2877. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Zou J, Liu KC, Wang WP and Xu Y: Circular
RNA COL1A2 promotes angiogenesis via regulating miR-29b/VEGF axis
in diabetic retinopathy. Life Sci. 256:1178882020. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Yao MD, Jiang Q, Ma Y, Zhu Y, Zhang QY,
Shi ZH, Zhao C and Yan B: Targeting circular RNA-MET for
anti-angiogenesis treatment via inhibiting endothelial tip cell
specialization. Mol Ther. 30:1252–1264. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Guo J, Xiao F, Ren W, Zhu Y, Du Q, Li Q
and Li X: ViaCircular Ribonucleic Acid circFTO promotes
angiogenesis and impairs blood-retinal barrier targeting the
miR-128-3p/Thioredoxin interacting protein axis in diabetic
retinopathy. Front Mol Biosci. 8:6854662021. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Zhu K, Hu X, Chen H, Li F, Yin N, Liu AL,
Shan K, Qin YW, Huang X, Chang Q, et al: Downregulation of circRNA
DMNT3B contributes to diabetic retinal vascular dysfunction through
targeting miR-20b-5p and BAMBI. EBioMedicine. 49:341–353. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Ye L, Guo H, Wang Y, Peng Y, Zhang Y, Li
S, Yang M and Wang L: Exosomal circEhmt1 released from
hypoxia-pretreated pericytes regulates high glucose-induced
microvascular dysfunction via the NFIA/NLRP3 pathway. Oxid Med Cell
Longev. 2021:88330982021. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Li Y, Cheng T, Wan C and Cang Y:
circRNA_0084043 contributes to the progression of diabetic
retinopathy via sponging miR-140-3p and inducing TGFA gene
expression in retinal pigment epithelial cells. Gene.
747:1446532020. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Sun H and Kang X: hsa_circ_0041795
contributes to human retinal pigment epithelial cells (ARPE 19)
injury induced by high glucose via sponging miR-646 and activating
VEGFC. Gene. 747:1446542020. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Wang L, Luo T, Bao Z, Li Y and Bu W:
Intrathecal circHIPK3 shRNA alleviates neuropathic pain in diabetic
rats. Biochem Biophys Res Commun. 505:644–650. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Zhang HH, Zhang Y, Wang X, Yang P, Zhang
BY, Hu S, Xu GY and Hu J: Circular RNA profile in diabetic
peripheral neuropathy: Analysis of coexpression networks of
circular RNAs and mRNAs. Epigenomics. 12:843–857. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Liu YT, Xu Z, Liu W, Ren S, Xiong HW,
Jiang T, Chen J, Kang Y, Li QY, Wu ZH, et al: The
circ_0002538/miR-138-5p/plasmolipin axis regulates Schwann cell
migration and myelination in diabetic peripheral neuropathy. Neural
Regen Res. 18:1591–1600. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Zhou B and Yu JW: A novel identified
circular RNA, circRNA_010567, promotes myocardial fibrosis via
suppressing miR-141 by targeting TGF-β1. Biochem Biophys Res
Commun. 487:769–775. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Dong S, Tu C, Ye X, Li L, Zhang M, Xue A,
Chen S, Zhao Z, Cong B, Lin J and Shen Y: Expression profiling of
circular RNAs and their potential role in early-stage diabetic
cardiomyopathy. Mol Med Rep. 22:1958–1968. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Shao Y, Li M, Yu Q, Gong M, Wang Y, Yang
X, Liu L, Liu D, Tan Z, Zhang Y, et al: CircRNA CDR1as promotes
cardiomyocyte apoptosis through activating hippo signaling pathway
in diabetic cardiomyopathy. Eur J Pharmacol. 922:1749152022.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Li M, Ding W, Tariq MA, Chang W, Zhang X,
Xu W, Hou L, Wang Y and Wang J: A circular transcript of ncx1 gene
mediates ischemic myocardial injury by targeting miR-133a-3p.
Theranostics. 8:5855–5869. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Sun Y, Yang Z, Zheng B, Zhang XH, Zhang
ML, Zhao XS, Zhao HY, Suzuki T and Wen JK: A novel regulatory
mechanism of smooth muscle α-actin expression by
NRG-1/circACTA2/miR-548f-5p axis. Circ Res. 121:628–635. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Ni H, Li W, Zhuge Y, Xu S, Wang Y, Chen Y,
Shen G and Wang F: Inhibition of circHIPK3 prevents angiotensin
II-induced cardiac fibrosis by sponging miR-29b-3p. Int J Cardiol.
292:188–196. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Zhu Y, Pan W, Yang T, Meng X, Jiang Z, Tao
L and Wang L: Upregulation of circular RNA CircNFIB attenuates
cardiac fibrosis by sponging miR-433. Front Genet. 10:5642019.
View Article : Google Scholar : PubMed/NCBI
|