Introduction
Urethral injury is a common form of urinary tract
trauma, often caused by external injury, surgical intervention or
infection (1,2). This type of injury frequently leads
to urethral stricture and fibrosis, resulting in severe functional
impairment (1). Due to the
complexity of tissue repair and lack of effective therapeutic
options, urethral injury presents notable clinical challenges
(3). Urethral injury not only
affects physiological functions but also markedly compromises the
quality of life of patients in the long term (4). Therefore, developing effective
therapeutic approaches for urethral repair and inhibiting fibrosis
remains an important clinical and research challenge (5).
The repair process following urethral injury
involves complex biological responses, including immune regulation,
cellular proliferation and migration and extracellular matrix
deposition (5,6). The immune response and fibrosis
progression serve notable roles in these processes (7,8).
Activation of the immune system following urethral injury often
triggers excessive inflammation, which subsequently leads to
fibrosis (9). Fibrosis impairs the
structural and functional integrity of the urethra, exacerbating
urethral stricture and contributing to chronic urinary dysfunction.
Thus, precise regulation of immune responses, cellular metabolism
and proliferation is important for effective inhibition of fibrosis
(10,11). Therefore, therapeutic strategies
simultaneously modulating immune responses and preventing the
progression of fibrosis are required.
Rapamycin, a commonly used immunosuppressant, is
widely utilized in organ transplantation and immune-related
diseases (12). Previously,
considerable attention has been paid to the potential therapeutic
benefits of rapamycin in fibrotic conditions. Rapamycin primarily
inhibits the mTOR signaling pathway, thereby modulating immune
responses, regulating cell proliferation and effectively inhibiting
fibrosis (13). Nonetheless,
notable clinical limitations, including side effects and drug
resistance, restrict the broader application of Rapamycin (14–16).
Therefore, studies (17–19) have increasingly focused on
alternative therapeutic agents or combination therapy, including
DNA nanostructure-based immunomodulators and anti-fibrotic
interventions targeting the TGF-β/Smad pathway, either alone or
combined with immune regulatory strategies.
The present study focused on novel immunomodulatory
molecules, tetrahedral DNA nanostructures (TDNs), self-assembled
tetrahedral framework nucleic acids characterized by high
structural stability, notable biocompatibility and efficient
cellular uptake (20). Due to
these unique properties, TDNs have attracted considerable attention
in biomedical research (21,22).
Previous studies have shown that TDN exhibits significant
therapeutic effects in inflammatory diseases, cancer and fibrosis.
Mechanistically, TDN regulates immune responses by modulating
macrophage polarization and suppressing excessive inflammatory
signaling (23,24), while also exerting anti-fibrotic
effects by inhibiting fibroblast activation and reducing
extracellular matrix deposition (25,26).
However, the potential role of TDN and its underlying mechanisms in
urethral injury repair remain to be fully elucidated. Therefore,
the present study investigated the therapeutic efficacy of TDN in
urethral injury and elucidated the underlying mechanisms.
Specifically, the present study assessed the role of TDN in immune
modulation, regulation of cellular metabolism and inhibition of
fibrosis.
To investigate the therapeutic potential of TDN in
urethral injury, the present study established a rat model of
urethral injury via mechanical trauma (27). This model effectively mimicked the
pathological conditions observed in clinical urethral injury,
including localized inflammation and subsequent fibrosis, providing
a robust experimental platform for assessing therapeutic efficacy
(28). TDN was administered to
rats with ureteral injury to observe its effects on immune
responses, cell proliferation, fibrosis and tissue repair.
Additionally, transcriptomic profiling and differential gene
network analysis were conducted to unravel the molecular mechanisms
through which TDN regulates the immune response, cell metabolism
and fibrosis.
Overall, the novelty of the present study lies in
the application of TDN in the treatment of urethral injury and the
comprehensive assessment of the underlying mechanisms.
Particularly, the present study examined the role of TDN in immune
regulation and fibrosis inhibition.
Materials and methods
Animals and grouping
The present study used 24 healthy male
Sprague-Dawley rats (age, 6–8 weeks; body weight, 180–220 g at the
start of the study), which were purchased from SPF Biotechnology
Co., Ltd. [license no. SCXK (Beijing) 2019–0010]. The rats were
housed under standard environmental conditions (temperature,
22±2°C; humidity, 50±10%; 12-h light/dark cycle) with free access
to food and water, and were acclimated for 1 week before the
experiment. The rats were randomly divided into four groups:
Control (n=6; intraperitoneal injection of an equal volume of
saline every other day; injection volume, 0.2 ml per rat), model
(n=6; urethral injury followed by intraperitoneal injection of
saline every other day; injection volume, 0.2 ml per rat), model +
rapamycin [n=6; 2.0 mg/kg of rapamycin (cat. no. HY-10219;
MedChemExpress) injected intraperitoneally every other day after
injury] and model + TDN (n=6; 10 nmol/day TDN administered via tail
vein injection daily after injury; injection volume, 0.2 ml per
rat). TDN was freshly prepared in TM buffer (10 mM Tris-HCl, 5 mM
MgCl2; pH 8.0), and its successful assembly and purity
were confirmed. Animal experiments were approved by the
Experimental Animal Welfare and Ethics Committee of Guizhou Medical
University (approval no. 2502311; Guiyang, China).
Preparation and verification of
TDN
The four single-stranded DNAs used for TDN assembly
were chemically synthesized by Sangon Biotech Co., Ltd. TDN was
synthesized by self-assembly of four single-stranded DNAs at a
final concentration of 30 µM in TM buffer (10 mM Tris-HCl, 5 mM
MgCl2; pH 8.0; cat. no. T10420; Shanghai Shangbao
Biotechnology Co., Ltd.), followed by denaturation at 95°C for 10
min and rapid cooling to 4°C. The sequences of the four
single-stranded DNAs were as follows: S1,
5′-ATTTATCACCCGCCATAGTAGACGTATCACCAGGCAGTTGAGACGAACATTCCTAAGTCTGAA-3′;
S2,
5′-ATTTATCACCCGCCATAGTAGACGTATCACCAGGCAGTTGAGACGAACATTCCTAAGTCTGAA-3′;
S3,
5′-ACTACTATGGCGGGTGATAAAACGTGTAGCAAGCTGTAATCGACGGGAAGAGCATGCCCATCC-3′;
and S4,
5′-ACGGTATTGGACCCTCGCATGACTCAACTGCCTGGTGATACGAGGATGGGCATGCTCTTCCCG-3′.
The successful formation of TDN was verified by 1% agarose gel
electrophoresis (cat. no. A8201; Beijing Solarbio Science &
Technology Co., Ltd.) using SerRed nucleic acid stain (cat. no.
G3606; Wuhan Servicebio Technology Co., Ltd.), with staining
performed at room temperature for 20 min, and by UV-visible
spectrophotometry (NanoPhotometer® N50; Implen GmbH). A
distinct absorption peak was observed at ~260 nm, with a
concentration of 184.35 ng/µl, confirming successful tetrahedral
assembly and high purity (Fig.
S1).
Urethral injury model
The urethral injury model was established based on
previous studies with slight modifications (27,29–31).
Rats were anesthetized with 1% pentobarbital sodium (40 mg/kg;
intraperitoneal; cat. no. P3761; MilliporeSigma) and then fixed on
a surgical platform (RWD Life Science Co., Ltd.). The surrounding
hair of the penis was shaved using an electric clipper (Flyco), and
the surgical field was disinfected with iodine tincture (Wuhan
Servicebio Technology Co., Ltd.). A longitudinal incision was made
on the ventral side of the penis to expose the urethra. Mechanical
injury was induced using an 18-G needle (BD Biosciences), forming
an injury of ~5 mm in length. After inducing injury, the urethral
sponge and skin were sutured (Fig.
S2) using 6–0 absorbable sutures (Ethicon, Inc.; Johnson &
Johnson). The wound was disinfected daily with povidone-iodine
solution (Wuhan Servicebio Technology Co., Ltd.), and body weight
and wound healing were monitored.
At the end of the experiment, all rats were
euthanized by intraperitoneal injection of sodium pentobarbital at
a dose of 100 mg/kg (cat. no. P3761; MilliporeSigma). Death was
confirmed by the absence of thoracoabdominal movements, a lack of
corneal reflex upon gentle touch and no withdrawal response to a
firm toe pinch after loss of consciousness.
Retrograde urethrography
After successful anesthesia with 1% pentobarbital
sodium (40 mg/kg, intraperitoneal), as aforementioned, rats were
fixed on a digital fluoroscopic table (DRX-Ascend; Carestream
Health, Inc.). Experimental staff wore protective clothing (Wuhan
Servicebio Technology Co., Ltd.) and inserted a 24-G intravenous
catheter (BD Biosciences) into the anterior urethra via the
urethral opening (32). Using
real-time X-ray monitoring, one hand stabilized the urethra and
catheter, while the other hand slowly injected the iodinated
contrast agent (iohexol; 300 mg iodine/ml; GE Healthcare) into the
bladder via the catheter. X-ray images were obtained to assess the
narrowing of the urethral lumen.
Hematoxylin and eosin (H&E)
staining
The urethral tissue was fixed in 4% paraformaldehyde
solution [cat. no. NH250218; Nuohai Life Science (Shanghai) Co.,
Ltd.] at room temperature for 24–48 h. Following fixation, the
tissues were rinsed with PBS (cat. no. G1101; Wuhan Servicebio
Technology Co., Ltd.), dehydrated through a graded ethanol series
[70, 80, 95% and absolute ethanol (CAS no. 64-17-5; analytical
grade;; Chengdu Kelong Chemical Co., Ltd.)], and cleared with
xylene (cat. no. 33535; Xilong Scientific Co., Ltd.). Samples were
subsequently embedded in paraffin (Shanghai Huayong Paraffin Co.,
Ltd.) and sectioned at a thickness of 5 µm using a rotary microtome
[HistoCore MULTICUT; cat. no. 149MULTIGC1; Leica Biosystems Co.,
Ltd.].
For H&E staining, paraffin-embedded sections
were deparaffinized with xylene and rehydrated through descending
concentrations of ethanol, followed by staining with hematoxylin
solution (cat. no. G1004-100ML; Wuhan Servicebio Technology Co.,
Ltd.) at room temperature for 5 min, rinsed with running water, and
counterstained with eosin staining solution (cat. no. G1108;
Beijing Solarbio Science & Technology Co., Ltd.) at room
temperature for 1–2 min. After dehydration and clearing with
xylene, the sections were mounted using neutral resin mounting
medium (cat. no. WG10004160; Wuhan Servicebio Technology Co., Ltd.)
and histological images were captured using an inverted light
microscope (TS2; Nikon Corporation).
Histological staining
Masson's trichrome staining was conducted to
evaluate collagen deposition in the urethral tissue (33). The urethral tissue was fixed in 4%
paraformaldehyde solution (cat. no. G1101; Wuhan Servicebio
Technology Co., Ltd.) at room temperature for 24 h, dehydrated and
embedded in paraffin. Sections with a thickness of 4 µm were
prepared using a rotary microtome. Masson's trichrome staining was
conducted at room temperature using a commercial kit (cat. no.
G1006; Wuhan Servicebio Technology Co., Ltd.) according to the
manufacturer's protocol, with the staining duration following the
standard instructions. Images were captured at a magnification of
×200 using a light microscope. Collagen fibers were stained blue,
whereas muscle fibers, fibrin and red blood cells appeared red.
Immunohistochemical staining (34) was conducted to evaluate the
expression of fibrosis markers, including α-smooth muscle actin
(α-SMA; cat. no. AF1032; Affinity Biosciences), TGF-β1 (1:5,000;
cat. no. 21898-1-AP; Proteintech Group, Inc.), collagen I (cat. no.
AF7001; Affinity Biosciences), collagen III (1:1,000; cat. no.
AF5457; Affinity Biosciences) and Smad3 (cat. no. 87035-1-RR;
Proteintech Group, Inc.). Briefly, paraffin-embedded tissue
sections prepared as aforementioned were sectioned at a thickness
of 4 µm. Sections were then deparaffinized, rehydrated through a
descending ethanol series and subjected to heat-induced antigen
retrieval using citrate buffer (pH 6.0; cat. no. G1202; Wuhan
Servicebio Technology Co., Ltd.). Subsequently, the sections were
incubated with 3% hydrogen peroxide (Wuhan Servicebio Technology
Co., Ltd.) to quench endogenous peroxidase activity and were
blocked with 5% BSA (cat. no. CR2302110; Beijing Solarbio Science
& Technology Co., Ltd.) for 30 min at room temperature; after
which, the sections were incubated overnight at 4°C with the
corresponding primary antibodies diluted in 2% BSA at a dilution of
1:100. This was followed by incubation with a HRP-conjugated
secondary antibody (cat. no. G1215; Wuhan Servicebio Technology
Co., Ltd.) for 50 min at 37°C. Color development was achieved using
a DAB chromogenic kit (cat. no. G1211; Wuhan Servicebio Technology
Co., Ltd.), and cell nuclei were counterstained with hematoxylin
(cat. no. G1040-500ML; Wuhan Servicebio Technology Co., Ltd.) at
room temperature for 2 min. Images were captured at a magnification
of ×200 using a light microscope (CX23; Olympus Corporation).
Image-Pro Plus software (version 6.0; Media Cybernetics, Inc.) was
employed to quantify the percentage and intensity of positive
staining to assess the degree of fibrosis.
Western blotting
To evaluate the protein expression levels of α-SMA,
TGF-β1, collagen I, Smad3 and collagen III, total protein was
extracted from rat urethral tissues using RIPA lysis buffer (cat.
no. G2002; Wuhan Servicebio Technology Co., Ltd.) containing
protease and phosphatase inhibitor cocktail (cat. no. G2007; Wuhan
Servicebio Technology Co., Ltd.). The protein concentration was
determined using a BCA assay kit (cat. no. G2026; Wuhan Servicebio
Technology Co., Ltd.) (35).
Proteins (30 µg per lane) were separated using SDS-PAGE (10% gel;
cat. no. P0012A; Beyotime Biotechnology) and transferred onto PVDF
membranes (cat. no. IPVH00010; MilliporeSigma) using the wet
transfer method. The membranes were blocked with 5% BSA for 1 h at
room temperature, and incubated overnight at 4°C with the following
primary antibodies: α-SMA (1:1,000; cat. no. AF1032; Affinity
Biosciences), TGF-β1 (1:5,000; cat. no. 21898-1-AP; Proteintech
Group, Inc.), collagen I (1:1,000; cat. no. AF7001; Affinity
Biosciences), collagen III (1:1,000; cat. no. AF5457; Affinity
Biosciences) and Smad3 (1:10,000; cat. no. 66516-1-Ig; Proteintech
Group, Inc.), and β-actin (1:25,000; cat. no. 66009-1-Ig;
Proteintech Group, Inc.), which was used as a loading control.
After washing three times with TBS containing 0.1%
Tween-20 (cat. no. G0004-500M; Wuhan Servicebio Technology Co.,
Ltd.), the membranes were incubated for 1 h at 37°C with
HRP-conjugated goat anti-rabbit IgG secondary antibody (1:3,000;
cat. no. GB23303; Wuhan Servicebio Technology Co., Ltd.) or
HRP-conjugated goat anti-mouse IgG secondary antibody (1:5,000;
cat. no. GB23301; Wuhan Servicebio Technology Co., Ltd.), as
appropriate. Protein bands were visualized using an enhanced
chemiluminescence kit (cat. no. G2014; Wuhan Servicebio Technology
Co., Ltd.) and images were captured with a ChemiDoc™ MP Imaging
System (cat. no. 12003154; Bio-Rad Laboratories, Inc.). Band
intensities were semi-quantified using Image-Pro Plus software
(version 6.0).
Transcriptome sequencing and
analysis
To explore the molecular mechanisms underlying
urethral injury and repair, total RNA was extracted from rat
urethral tissues using TRIzol® reagent (cat. no.
15596026; Invitrogen; Thermo Fisher Scientific, Inc.) according to
the manufacturer's protocol.
The RNA concentration and purity were assessed using
a NanoDrop™ 2000 spectrophotometer (cat. no. ND-2000; Thermo Fisher
Scientific, Inc.), and RNA integrity was verified by 1% agarose gel
electrophoresis (cat. no. A8201; Beijing Solarbio Science &
Technology Co., Ltd.). High-throughput sequencing libraries were
prepared using the NEBNext® Ultra™ RNA Library Prep Kit
for Illumina (cat. no. E7530L; New England BioLabs, Inc.) and
sequencing was performed on an Illumina NovaSeq 6000 platform using
the NovaSeq 6000 S4 Reagent Kit v1.5 (300 cycles; cat. no.
20028312; Illumina, Inc.) (36).
Paired-end RNA sequencing (2×150 bp) was performed in a
forward-reverse orientation, with the final libraries loaded at a
concentration of 300 pM. The quality of raw sequencing data was
evaluated using FastQC (version 0.11.9; Babraham Bioinformatics),
and adaptor trimming and filtering were conducted using Trimmomatic
(version 0.39; http://www.usadellab.org/cms/?page=trimmomatic). Clean
reads were aligned to the rat reference genome (Rnor_6.0) using
HISAT2 (version 2.2.1; http://daehwankimlab.github.io/hisat2/). Principal
component analysis (PCA) was performed to evaluate global
transcriptional differences among samples and to assess the
consistency of biological replicates. Sample-to-sample correlation
analysis was performed using Pearson correlation coefficients based
on normalized gene expression values to examine intra-group
repeatability and inter-group variability. The results were
visualized using PCA for dimensionality reduction and correlation
heatmaps generated using the pheatmap package (version 1.0.12;
http://cran.r-project.org/package=pheatmap) in R
software (version 4.2.2; The R Foundation for Statistical
Computing; http://www.r-project.org/).
Differential gene expression analysis was performed using the
DESeq2 package (version 1.38.0; http://bioconductor.org/packages/DESeq2/) in R
software, with thresholds of |log2(fold change)|≥0.5 and P<0.05
to identify significantly differentially expressed genes (DEGs).
Functional enrichment analyses were conducted using the Kyoto
Encyclopedia of Genes and Genomes (KEGG) database (https://www.kegg.jp) to identify pathways related to
fibrosis, inflammation, immune regulation, metabolism, cell cycle
regulation and tissue repair. Specifically, pathways were selected
based on their direct relevance to immune regulation, inflammation,
metabolism, cell cycle regulation and tissue repair, which are key
processes associated with urethral injury and TDN-mediated
therapeutic effects. Disease-related pathways with limited
mechanistic relevance to urethral pathology were excluded from the
final analysis. Statistical analyses and visualization were
conducted in R software using the pheatmap package and ggplot2
package (version 3.4.0; http://cran.r-project.org/package=ggplot2) to display
gene expression patterns and enrichment results.
Reverse transcription-quantitative PCR
(RT-qPCR)
To validate the transcriptome sequencing results,
total RNA was extracted from rat urethral tissues using
TRIzol® reagent (cat. no. 15596026; Invitrogen; Thermo
Fisher Scientific, Inc.) according to the manufacturer's
instructions. Reverse transcription was performed using the
PrimeScript™ RT reagent kit with gDNA eraser (cat. no. RR047A;
Takara Bio, Inc.) to synthesize cDNA. Reverse transcription was
performed at 42°C for 15 min, followed by enzyme inactivation at
85°C for 5 sec. qPCR was conducted using TB Green®
Premix Ex Taq™ II (cat. no. RR820A; Takara Bio, Inc.) on a
QuantStudio™ 5 Real-Time PCR System (cat. no. A28574; Applied
Biosystems; Thermo Fisher Scientific, Inc.). Each reaction (20 µl)
contained 10 µl SYBR Green mix, 1 µl cDNA template, 0.4 µl of each
primer (10 µM) and 8.2 µl nuclease-free water. The PCR
amplification conditions were as follows: 95°C for 30 sec, followed
by 40 cycles of 95°C for 5 sec and 60°C for 30 sec. The expression
levels of the cell cycle-related genes cyclin B1 (Ccnb1),
ribonucleotide reductase regulatory subunit M2 (Rrm2),
polo-like kinase 1 (Plk1) and cyclin-dependent kinase 1
(Cdk1), as well as the inflammatory cytokines IL-6,
IL-1β and TNF-α, were analyzed, with GAPDH used
as the internal control. The relative gene expression levels were
calculated using the 2−ΔΔCq method for quantification,
as previously described by Livak and Schmittgen (37). Primer sequences are listed in
Table SI.
Differential gene network
analysis
After identifying DEGs from the transcriptome data,
protein-protein interaction (PPI) network analysis was performed
using the Search Tool for the Retrieval of Interacting
Genes/Proteins database (version 12.0; http://string-db.org/) to evaluate gene-gene
associations. A confidence score threshold of 0.7 (high confidence)
was applied to filter significant interactions and the resulting
interaction data were imported into Cytoscape software (version
3.10.0; Cytoscape Consortium) for visualization. The PPI network
was optimized using the cytoHubba plug-in (version 0.1; http://apps.cytoscape.org/apps/cytohubba) to identify
hub genes with the highest degree of connectivity.
These key genes were analyzed in the context of
fibrosis, extracellular matrix remodeling and cell cycle regulation
to explore their biological significance and potential roles in the
pathogenesis of urethral injury.
ELISA
The serum concentrations of IL-6, IL-1β and TNF-α
were measured using ELISA kits: IL-6 (cat. no. SEA079Ra;
CLOUD-CLONE CORP), IL-1β (cat. no. SEA563Ra; CLOUD-CLONE CORP) and
TNF-α (cat. no. SEA133Ra; CLOUD-CLONE CORP). The assays were
conducted according to the manufacturer's protocols. Absorbance was
read at 450 nm using a SpectraMax® iD3 microplate reader
(iD3; Molecular Devices, LLC). Cytokine levels were quantified
based on standard curves and expressed as pg/ml. These data were
used to evaluate the systemic immune and inflammatory responses
following urethral injury.
Single-sample gene set enrichment
analysis (ssGSEA)
ssGSEA was performed to evaluate the immune
microenvironment and quantify immune cell infiltration in rat
urethral tissues.
The analysis was based on the normalized
transcriptome expression matrix. The GSVA package (version 1.46.0;
http://bioconductor.org/packages/GSVA/) in R software
was used to calculate the enrichment scores of immune-related gene
sets for each sample. Immune cell-related gene sets representing 24
immune cell types were derived directly from the immune cell
signature gene sets defined in a previous study by Bindea et
al (38) and were curated for
application in ssGSEA. These gene sets included signatures for
macrophages, T cells, natural killer cells, dendritic cells and
neutrophils. The enrichment scores were visualized using the
ggplot2 package and the pheatmap package n R., enabling
quantitative assessment of immune cell infiltration and
characterization of immune landscape alterations associated with
urethral injury and TDN treatment.
Fibroblast isolation and culture
Primary fibroblasts were isolated from rat urethral
tissues under sterile conditions. Briefly, freshly harvested
urethral tissues were rinsed 3–5 times with PBS supplemented with
5% penicillin-streptomycin solution (Dalian Meilun Biology
Technology Co., Ltd.), with each wash lasting 5 min. The tissues
were then minced into small fragments using sterile scissors and
digested with 0.25% trypsin (Dalian Meilun Biology Technology Co.,
Ltd.) at 37°C in a humidified incubator with 5% CO2 for
60 min.
Following enzymatic digestion, 6 ml complete
Dulbecco's modified Eagle's medium (DMEM; Dalian Meilun Biology
Technology Co., Ltd.) supplemented with 10% fetal bovine serum
(FBS; Gibco; Thermo Fisher Scientific, Inc.) and 1%
penicillin-streptomycin was added to terminate digestion. The cell
suspension was collected and centrifuged at 1,000 rpm for 5 min.
The resulting cell pellet was resuspended in complete DMEM and
transferred to culture flasks for incubation at 37°C in a
humidified atmosphere containing 5% CO2.
The culture medium was replaced every 2–3 days. When
cells reached ~80% confluence, fibroblasts were passaged using
0.25% trypsin digestion. Cells were washed twice with PBS, digested
until most cells became rounded and detached, and digestion was
terminated with complete DMEM. After centrifugation at ~200 × g for
5 min at room temperature, the cells were resuspended in fresh
complete medium and passaged at a ratio of 1:2. Primary rat
urethral fibroblasts were cultured in complete DMEM supplemented
with 10% FBS and 1% penicillin-streptomycin at 37°C in a humidified
incubator containing 5% CO2. When cells reached 70–80%
confluence, in vitro treatments were performed. Cells were
divided into the following three groups: Control, TGF-β-treated and
TGF-β + TDN-treated groups. For induction of vimentin expression,
fibroblasts were treated with TGF-β (5 ng/ml; cat. no. 80116-R08H;
Beijing Sino Technology Co., Ltd.) for 24 h. For the intervention
group, TDN (1 µM) was added simultaneously with TGF-β, and the
cells were co-treated for 24 h at 37°C. After treatment, the cells
were collected for immunohistochemical analysis of vimentin
expression, as described in the Materials and methods section.
Immunocytochemical staining for
vimentin
To evaluate vimentin expression in fibroblasts,
immunocytochemical staining was performed. After treatment, the
cells were washed with PBS and fixed with 4% paraformaldehyde at
room temperature overnight. Following fixation, the cells were
washed with PBS, subjected to antigen retrieval using pepsin
solution at 37°C for 30 min, and endogenous peroxidase activity was
quenched with 3% hydrogen peroxide for 20 min at room temperature.
The cells were then blocked with 5% bovine serum albumin (cat. no.
CR2302110; Beijing Solarbio Science & Technology Co., Ltd.) for
30 min at 37°C and incubated overnight at 4°C with a primary
antibody against vimentin (1:150; cat. no. AFRM0062; Hunan Aifang
Biotechnology Co., Ltd.), diluted in 2% bovine serum albumin. After
washing, the cells were incubated with a polymer-based secondary
antibody using a universal two-step detection kit (cat. no.
PV-9000; OriGene Technologies, Inc.) according to the
manufacturer's instructions. Color development was achieved using a
DAB chromogenic kit, and cell nuclei were counterstained with
hematoxylin for 5 min at room temperature. Representative images
were captured using a light microscope, and quantitative analysis
was performed using Image-Pro Plus software, as aforementioned.
H&E staining of fibroblasts
Primary rat urethral fibroblasts were isolated from
urethral tissues and cultured under standard conditions as
aforementioned. After expansion, fibroblasts were enzymatically
dissociated and seeded onto glass coverslips for cell attachment.
Following culture, fibroblast samples (n=6) were provided for
histological evaluation, including H&E staining and
immunocytochemical staining. H&E staining was performed as
aforementioned to assess fibroblast morphology, and representative
images were acquired using a light microscope.
Immunofluorescence identification of
fibroblasts
To confirm the purity and identity of the
fibroblasts isolated from rat urethral tissues, immunofluorescence
staining for α-SMA and CD90 was performed. Cells were fixed with 4%
paraformaldehyde (cat. no. G1101; Wuhan Servicebio Technology Co.,
Ltd.) for 15 min at room temperature, permeabilized with 0.3%
Triton X-100 (cat. no. G1203; Wuhan Servicebio Technology Co.,
Ltd.) and blocked with 5% BSA for 30 min at room temperature. The
cells were incubated with primary antibodies against α-SMA (1:100;
cat. no. AF1032; Affinity Biosciences) and CD90 (1:100; cat. no.
DF4804; Affinity Biosciences) overnight at 4°C. After washing with
PBS, cells were incubated with goat anti-rabbit IgG H&L (Alexa
Fluor™ 488-conjugated; 1:200; cat. no. ab150077; Abcam) secondary
antibodies for 40 min at 37°C. Nuclei were counterstained with DAPI
(cat. no. G1012; Wuhan Servicebio Technology Co., Ltd.) at room
temperature for 8 min in the dark. Fluorescence images were
captured using a Nikon TS2-S-SM inverted fluorescence microscope
(Nikon Corporation). The results showed negative α-SMA staining
together with positive CD90 expression, indicating that the
isolated cells were predominantly fibroblasts and excluding
myofibroblast contamination.
Statistical analysis
All quantitative data are presented as the mean ±
standard deviation from at least three independent experiments.
Statistical analyses were performed using GraphPad Prism software
(version 9.0; Dotmatics). This approach was used for multi-group
quantitative analyses presented in the figures. Comparisons among
three or more groups were performed using one-way analysis of
variance followed by Tukey's post hoc test. P<0.05 was
considered to indicate a statistically significant difference.
Transcriptome data were analyzed using R software. The ggplot2
(version 3.4.0), pheatmap (version 1.0.12) and ClusterProfiler
(version 4.6.0; http://bioconductor.org/packages/clusterProfiler/)
packages were used for data visualization and functional enrichment
analysis. Sample correlation analysis was performed using Pearson
correlation analysis based on normalized gene expression levels,
and the results are presented as a sample-to-sample correlation
heatmap. All figures were generated using GraphPad Prism and R and
assembled in Adobe Illustrator (version 2023; Adobe Systems, Inc.)
for final presentation.
Results
Validation of the rat model of
urethral injury
To ensure reliability, the urethral injury model was
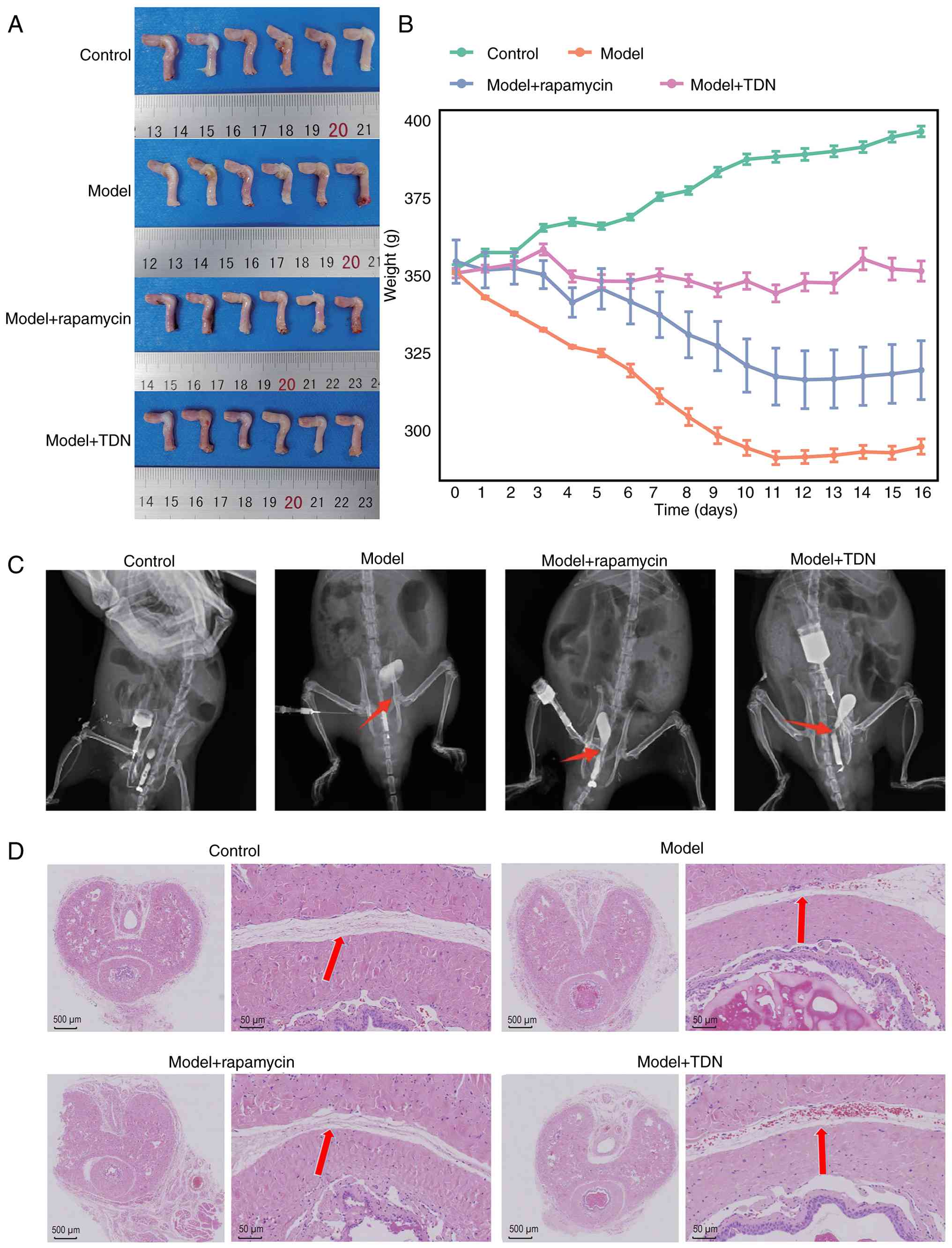
validated from multiple aspects. Fig.
1A shows the tissue collection images of rat urethras in
different groups. Postoperative body weight recordings showed that
the control group gradually gained weight, whereas the model group
experienced marked weight loss. Notably, body weight in the
rapamycin and TDN groups exhibited a slight, non-significant
decrease and remained relatively stable overall (Fig. 1B). Urethrography showed that the
urethras of rats were healthy in the control group, while urethral
stenosis was successfully developed in the model group. Urethral
stenosis improved after treatment with rapamycin and TDN (Fig. 1C). H&E staining showed that in
the model group, the mucosal layer of the urethra was notably
thickened and the lumen was narrower, with marked inflammatory cell
infiltration compared with the control group. Consistent with the
results of urethrography, the rapamycin and TDN groups showed
notable improvements in urethral histological structure, with
reduced luminal narrowing and improved tissue organization,
suggesting an attenuation of urethral injury following rapamycin
and TDN treatment (Fig. 1D).
These results suggested that the urethral injury
model was successfully established, and different treatment groups
exhibited notable differences in terms of urethral damage and
fibrosis. Particularly, the results supported the therapeutic
effects of Rapamycin and TDN, further supporting their efficacy in
alleviating urethral injury.
Differential gene expression induced
by urethral injury in rats
Transcriptome sequencing revealed the relationship
between different treatments and gene expression. PCA (Fig. 2A) showed that samples from the same
group clustered closely together, whereas samples from different
groups were clearly separated, indicating distinct transcriptomic
profiles among the control model, and treatment groups.
Consistently, sample correlation analysis (Fig. 2B) revealed high correlation
coefficients among biological replicates within each group and
lower correlations between different groups, demonstrating good
intra-group reproducibility and clear inter-group differentiation.
Differential expression analysis identified 3,008 DEGs, including
1,248 upregulated and 819 downregulated genes between the model and
control groups, 194 upregulated and 45 downregulated genes between
the Rapamycin and model groups, and 206 upregulated and 496
downregulated genes between the TDN and model groups (Fig. 2C and D). All three comparisons
shared 41 genes (Fig. 2E). The
slight discrepancy in gene numbers between the Venn diagram
(Fig. 2E) and the DEG counts shown
in Fig. 2C and D is due to
differences in gene inclusion criteria, as one gene located at the
significance threshold was included in the Venn analysis but
excluded from the final DEG count. These genes, including genes
related to metabolism, keratin, acute-phase response, inflammation
and immune function, exhibited differential expression patterns
after different treatments (Fig.
2F).
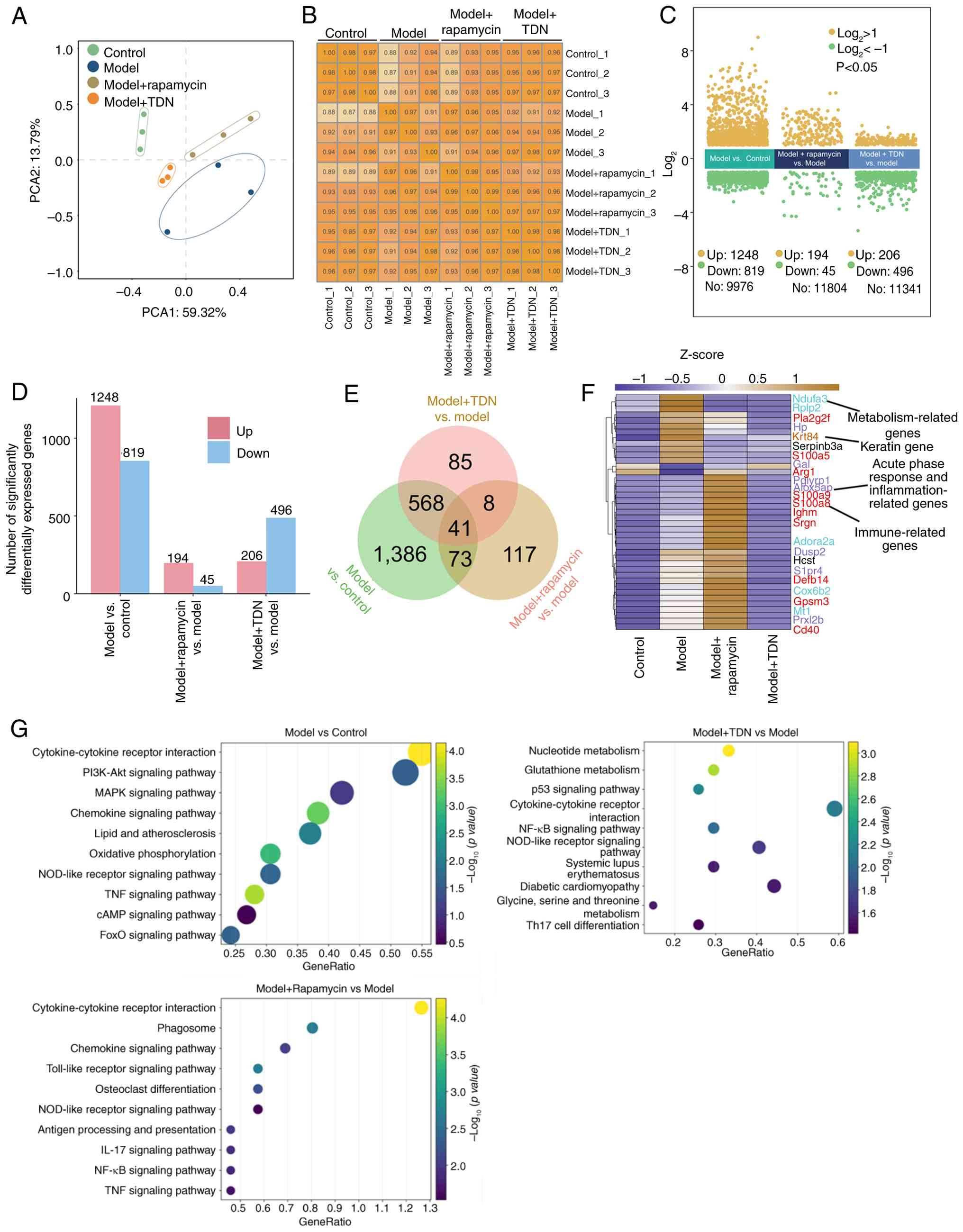 | Figure 2.(A) PCA results. Different colors
represent different treatment groups. (B) Sample correlation
heatmap. The color intensity corresponds to correlation values. (C)
Combined volcano plot showing the distribution of FCs in
differentially expressed genes in the three group comparisons
(model vs. control; model + rapamycin vs. model; model + TDN vs.
model), with yellow dots representing upregulated genes and green
dots representing downregulated genes. (D) Bar chart of
differential gene counts showing the number of differential genes
in the three group comparisons. (E) Venn diagram of differential
genes displaying the distribution of differential genes in the
three group comparisons. The numbers in different areas represent
specific intersections or unique differential genes. (F) Heatmap
showing the expression patterns of 25 common differentially
expressed genes identified from three pairwise comparisons,
displayed across four experimental groups (Control, Model, Model +
rapamycin, and Model + TDN). Each row represents one gene and each
column represents an individual sample. Color gradients indicate
normalized gene expression levels. (G) KEGG pathway enrichment
analysis of differentially expressed genes from the three pairwise
comparisons (model vs. control; model + rapamycin vs. model; model
+ TDN vs. model). Enrichment results are presented as dot plots.
The x-axis represents the GeneRatio, and the size of each dot
reflects the proportion of genes enriched in the corresponding
pathway. Dot color indicates the statistical significance expressed
as -log10(P-value). KEGG pathways are displayed consistently across
the three comparisons to facilitate direct visual comparison. KEGG,
Kyoto Encyclopedia of Genes and Genomes; TDN, tetrahedral DNA
nanostructure; FC, fold change; PCA, principal component
analysis. |
KEGG pathway enrichment analysis of all differential
genes revealed that differential genes between the model and
control groups were predominantly enriched in: i) Immune-related
and inflammation-related pathways, including ‘Cytokine-cytokine
receptor interaction’, ‘Chemokine signaling pathway’, ‘NOD-like
receptor signaling pathway’ and ‘TNF signaling pathway’; ii)
survival, proliferation and metabolism-related pathways, such as
‘PI3K-Akt signaling pathway’, ‘MAPK signaling pathway’, ‘cAMP
signaling pathway’ and ‘FoxO signaling pathway’; and iii)
cardiovascular and metabolic pathways, including the ‘Lipid and
atherosclerosis’ and ‘Oxidative phosphorylation’ pathways.
Differential genes between the Rapamycin and model groups were
mainly enriched in: i) Immune-related and inflammation-related
pathways, such as ‘Cytokine-cytokine receptor interaction’,
‘Chemokine signaling pathway’, ‘Toll-like receptor signaling
pathway’, ‘NF-κB signaling pathway’, ‘TNF signaling pathway’ and
‘IL-17 signaling pathway’; and ii) antigen presentation and immune
recognition pathways, including the ‘Antigen processing and
presentation’ and ‘Phagosome’ pathways. Differential genes between
the TDN and model groups were predominantly enriched in: i)
Immune-related and inflammation-related pathways, such as
‘Cytokine-cytokine receptor interaction’, ‘NF-κB signaling pathway’
and ‘NOD-like receptor signaling pathway’; ii) metabolism-related
and cell function-related pathways, such as the ‘Nucleotide
metabolism’, ‘Glutathione metabolism’ and ‘Glycine, serine and
threonine metabolism’ pathways; and iii) cell cycle-related
pathways such as the ‘p53 signaling pathway’ (Fig. 2G).
Urethral fibrosis induced by urethral
injury in rats
Masson's trichrome staining showed a notable
increase in collagen fibers in the urethral tissue of the model
group compared with the control group. By contrast, the content of
collagen fibers decreased after treatment with rapamycin and TDN.
Immunohistochemical staining also demonstrated upregulated
expression of fibrosis markers, such as α-SMA, TGF-β1, collagen I,
collagen III and Smad3, in the model group compared with the
control group, but their expression was subsequently decreased in
the rapamycin and TDN groups compared with that in the model group
(Fig. 3A).
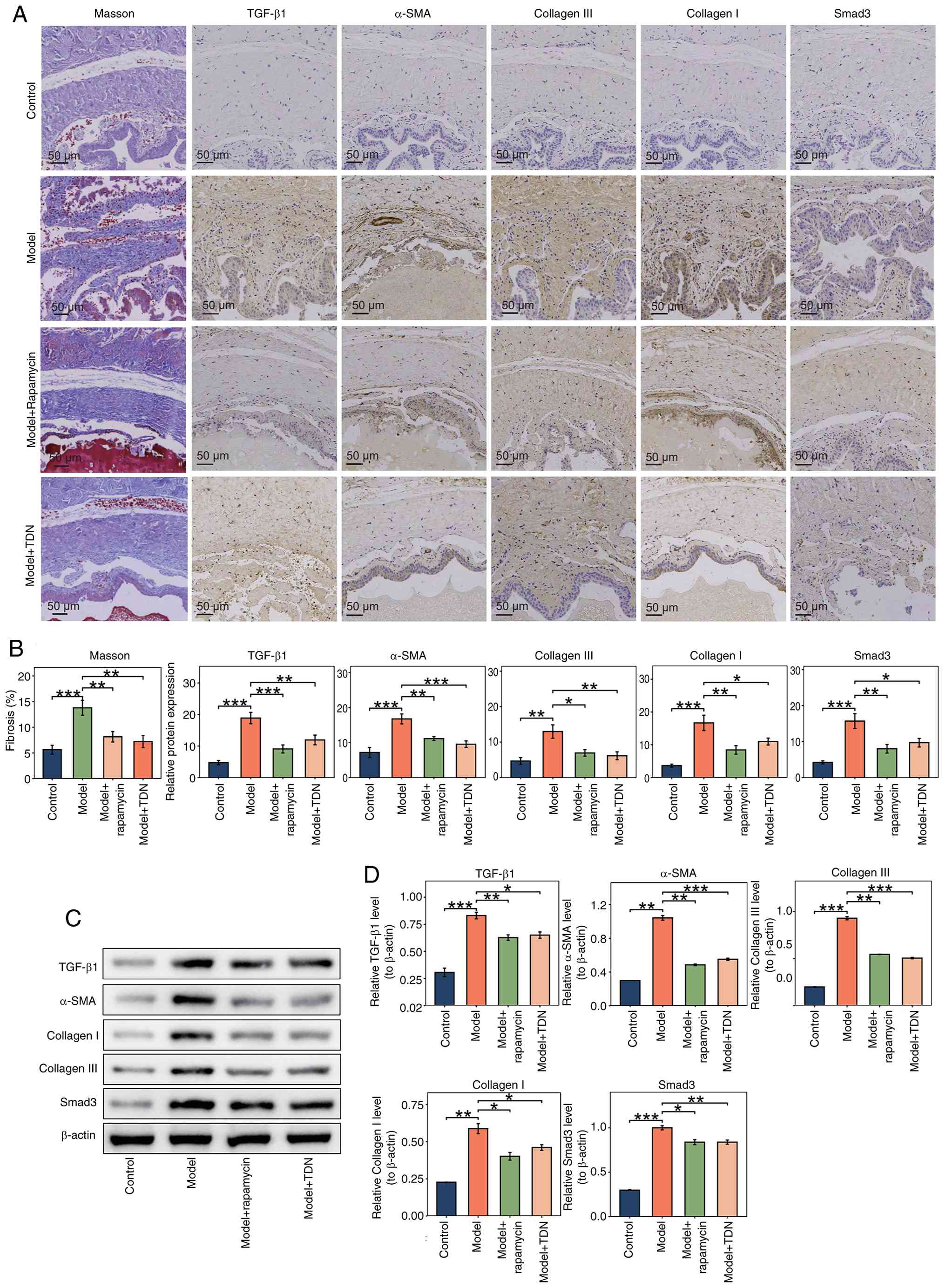 | Figure 3.(A) Masson's trichrome staining and
immunohistochemistry results. Masson's trichrome staining detected
changes in collagen fibers in urethral tissues, with collagen
fibers appearing blue and muscle fibers, fibrin and red blood cells
appearing red. Immunohistochemistry staining showed the cell nuclei
in blue and positive protein staining for the fibrosis markers
α-SMA, TGF-β1, collagen I, collagen III and Smad3 in brown. Images
are presented at a magnification of ×200. (B) Statistical analysis
of Masson's trichrome staining and immunohistochemistry results.
Staining results were analyzed with Image-Pro Plus software(version
6.0; Media Cybernetics, Inc.), followed by bar chart construction
using GraphPad Prism. *P<0.05, **P<0.01 and ***P<0.001.
(C) Western blot analysis results. Western blot analysis was used
to detect the protein expression of fibrosis markers in urethral
tissue. (D) Statistical analysis of western blotting results. Band
densities were analyzed using Image-Pro Plus software(version 6.0;
Media Cybernetics, Inc.), and bar charts were drawn with GraphPad
Prism. *P<0.05, **P<0.01 and ***P<0.001. α-SMA, α-smooth
muscle actin; TDN, tetrahedral DNA nanostructure. |
Quantitative analysis of Masson's trichrome-stained
sections using Image-Pro Plus software revealed a significant
increase in collagen fiber content in the model group compared with
that in the control group (P<0.001), along with a significant
increase in the expression of fibrosis markers based on
immunohistochemical analyses (P<0.01 or P<0.001). By
contrast, the rapamycin and TDN groups exhibited a significant
decrease in collagen content and reduced expression of fibrosis
markers compared with the model group (P<0.05, P<0.01 or
P<0.001), suggesting that rapamycin and TDN treatment exerted
inhibitory effects on fibrosis (Fig.
3B).
Western blot analysis supported these findings, with
significant increases observed in the protein expression levels of
α-SMA, TGF-β1, collagen I, Smad3 and collagen III in the model
group compared with the control group (P<0.01 or P<0.001).
Compared with the model group, significant decreases in the
expression of these markers were observed in the Rapamycin and TDN
groups (P<0.05, P<0.01 or P<0.001) (Fig. 3C and D).
Overall, TDN effectively mitigated the progression
of urethral fibrosis in the rat model by downregulating the
expression levels of the fibrosis markers α-SMA, TGF-β1, collagen
I, collagen III and Smad3, as well as by inhibiting collagen
deposition as evidenced by Masson's trichrome staining Masson's
trichrome staining and immunohistochemical staining indicated that
TDN significantly suppressed the accumulation of collagen fibers
and downregulated fibrosis markers in the model group, suggesting
that TDN displayed protective effects against fibrosis. These
results provide a theoretical basis for exploring the potential
application of TDN.
Immune response induced by urethral
injury in rats
Immune infiltration scores in rats were analyzed by
ssGSEA to unravel the relationship between urethral injury and the
immune system. The model group showed marked immune cell
infiltration of innate and adaptive immune cells, particularly
macrophages, monocytes, and activated CD4+ and
CD8+ T cells, whereas neutrophil infiltration was
reduced in the model group compared with in the control group,
indicating that urethral injury induced a shift from acute
neutrophil-dominated inflammation toward a macrophage- and
lymphocyte-driven immune response, suggesting the development of a
sustained inflammatory and immune remodeling process (Fig. 4A). Furthermore, the immune
infiltration score heatmap showed marked differences in immune cell
infiltration across the groups, with TDN treatment markedly
reducing immune cell infiltration compared with that in the model
group (Fig. 4B).
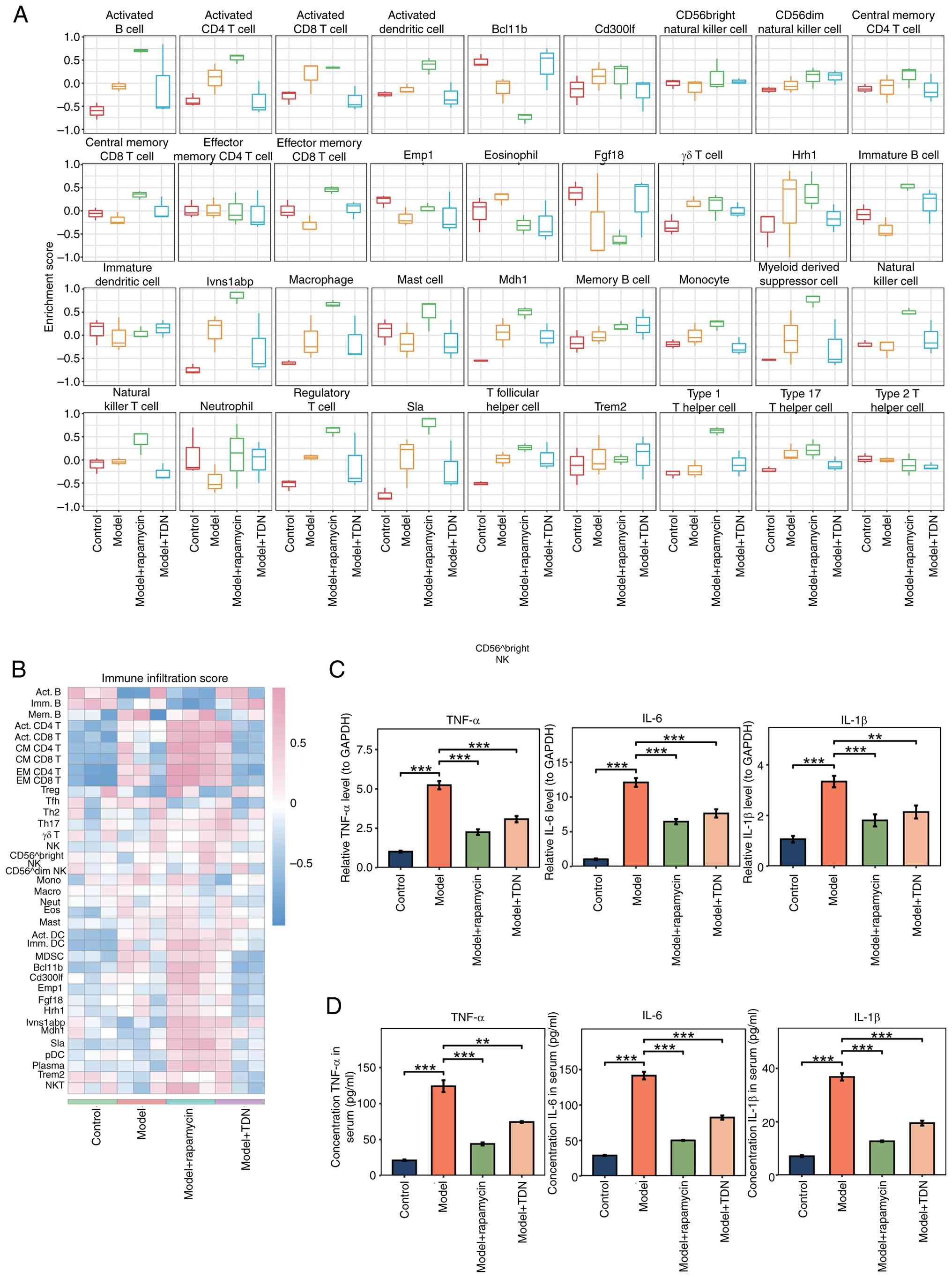 | Figure 4.(A) Immune infiltration score
boxplot. Single-sample gene set enrichment analysis was used to
calculate the immune cell infiltration scores in each group and
results were displayed as a boxplot, with the box representing the
interquartile range, the median indicated by a horizontal line and
the whiskers showing the distribution range of the data. (B) Immune
infiltration score heatmap. After standardization, the heatmap
displays the infiltration scores of different immune cells across
groups. Color changes represent high to low immune infiltration
scores, with red indicating high scores and blue indicating low
scores, rows represent different immune cell types, and columns
represent experimental groups. (C) RT-qPCR detection of
inflammatory cytokines. RT-qPCR analysis was used to measure the
mRNA expression levels of IL-6, IL-1β and TNF-α. **P<0.01 and
***P<0.001. (D) ELISA detection of inflammatory cytokines in
serum. An ELISA was used to measure the levels of IL-6, IL-1β and
TNF-α in serum. **P<0.01 and ***P<0.001. RT-qPCR, reverse
transcription-quantitative PCR; TDN, tetrahedral DNA nanostructure;
NK, natural killer; NKT, natural killer T cell; DC, dendritic cell;
pDC, plasmacytoid dendritic cell; MDSC, myeloid-derived suppressor
cell; Act., activated; Imm., immature; CM, central memory; EM,
effector memory; Treg, regulatory T cell; Tfh, T follicular helper
cell; Th1, type 1 T helper cell; Th2, type 2 T helper cell; Th17,
type 17 T helper cell; Mono, monocyte; Macro, macrophage; Neut,
neutrophil; Eos, eosinophil; Mast, mast cell; Plasma, plasma
cell. |
RT-qPCR showed that the mRNA expression levels of
the inflammatory cytokines IL-6, IL-1β and TNF-α were significantly
increased in the model group compared with the control group
(P<0.001). Compared with those in the model group, rapamycin and
TDN significantly downregulated the levels of these inflammatory
cytokines (P<0.01 or P<0.001), indicating that after urethral
injury, both rapamycin and TDN can modulate the immune response at
the transcriptional level (Fig.
4C). An ELISA also supported these findings. The serum levels
of inflammatory cytokines were significantly higher in the model
group compared with the control group (P<0.001). Compared with
those in the model group, the serum levels of inflammatory
cytokines were significantly decreased in the treatment groups
(P<0.01 or P<0.001) (Fig.
4D).
These results suggested that urethral injury
significantly enhanced the immune response, while TDN effectively
inhibited immune cell infiltration and lowered the expression
levels of inflammatory cytokines, reducing the immune response
triggered by urethral injury. TDN modulated the immune response at
the transcriptional level and prevented the release of inflammatory
cytokines, indicating its potential anti-inflammatory and
immunoregulatory properties after urethral injury.
Expression levels of Ccnb1, Rrm2, Plk1
and Cdk1 in urethral injury
Transcriptomic analysis revealed that the expression
levels of cell cycle-related genes, such as Ccnb1, Rrm2,
Plk1 and Cdk1, were notably upregulated in the model
group compared with the control group, suggesting that urethral
injury may have accelerated cell proliferation by promoting cell
cycle progression Compared with those in the model group, the
expression levels of most of these genes were decreased in the
rapamycin and TDN groups, whereas Rrm2 showed no marked reduction
in the rapamycin group, indicating that rapamycin and TDN may have
regulated cell proliferation by inhibiting the expression of cell
cycle-related genes. Heatmap and boxplot analyses (Fig. 5A and B) visualized between-group
differences in the expression of these genes, showing that after
urethral injury, the expression of cell cycle-related genes was
effectively controlled in the intervention groups.
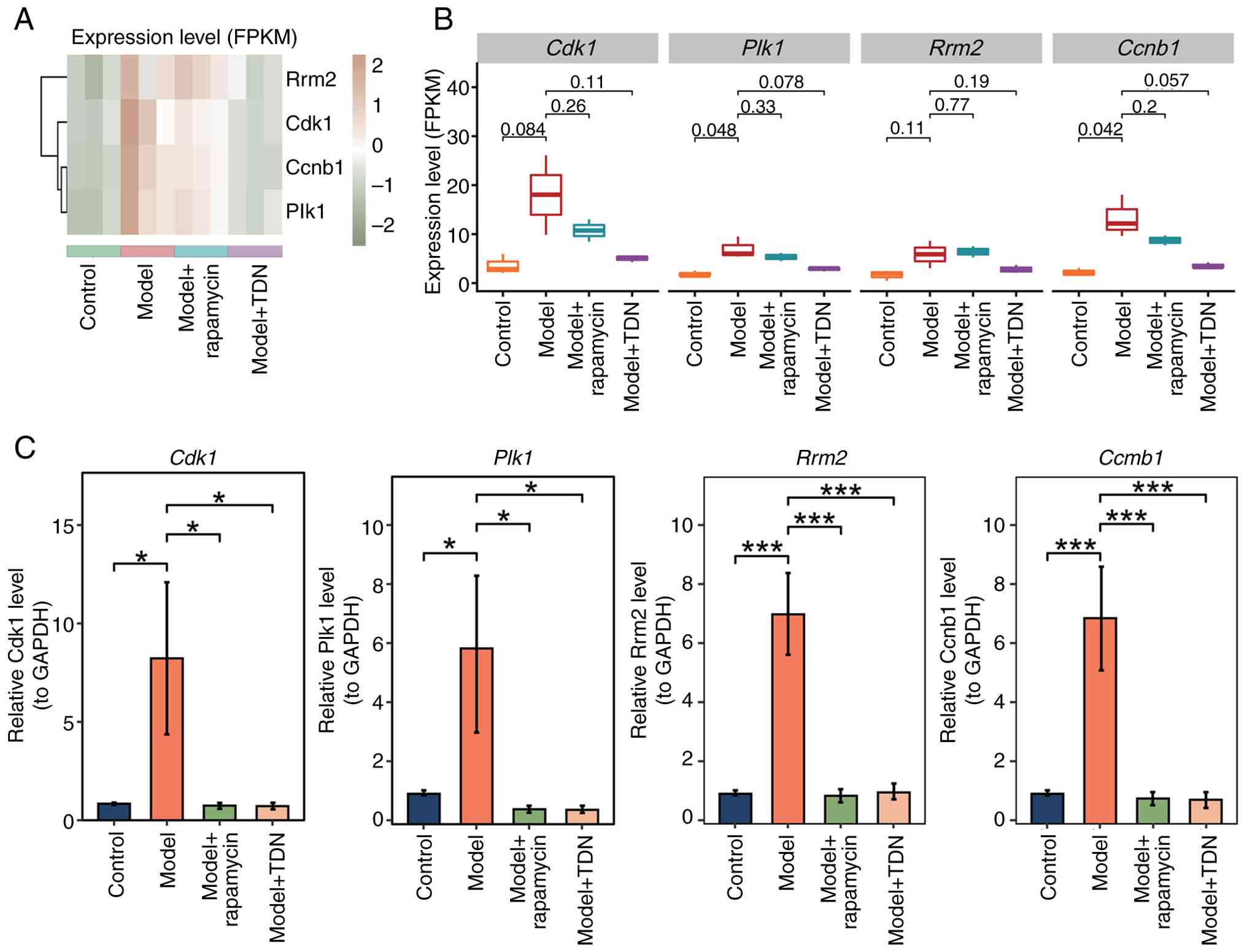 | Figure 5.(A) Gene expression heatmap and (B)
boxplots for Ccnb1, Rrm2, Plk1 and Cdk1. The heatmap
shows the expression level changes of Ccnb1, Rrm2, Plk1 and
Cdk1 across groups, with color changes reflecting the
difference in gene expression, where orange indicates high
expression and green indicates low expression. The boxplot
illustrates the distribution of gene expression levels among
groups. (C) RT-qPCR detection of gene expression. RT-qPCR was used
to measure the mRNA expression levels of Ccnb1, Rrm2, Plk1
and Cdk1 in urethral tissue, with results displayed in a bar
chart. *P<0.05 and ***P<0.001. RT-qPCR, reverse
transcription-quantitative PCR; TDN, tetrahedral DNA nanostructure;
Ccnb1, cyclin B1; Rrm2, ribonucleotide reductase
regulatory subunit M2; Plk1, polo-like kinase 1;
Cdk1, cyclin-dependent kinase 1; FPKM, fragments per
kilobase million. |
RT-qPCR further supported the results of the
transcriptomic analysis. The analysis (Fig. 5C) showed that, compared with those
in the control group, the mRNA expression levels of Ccnb1, Rrm2,
Plk1 and Cdk1 were significantly enhanced in the model
group (P<0.05 or P<0.001). Furthermore, the expression levels
of cell cycle genes were significantly lower in the rapamycin and
TDN groups compared with the model group (P<0.05 or P<0.001),
providing evidence that rapamycin and TDN regulated the expression
of genes related to the cell cycle to inhibit cell
proliferation.
Differential gene network
analysis
Differential gene network analysis revealed the
relationships between genes in the rat model of urethral injury.
The network illustrated genes with significant interactions after
urethral injury, with gene nodes connected by edges. The density of
the network reflected the strength of gene interactions. By
analyzing the interaction networks of these genes, the present
study divided them into several functional modules, each
representing a specific biological characteristic or function
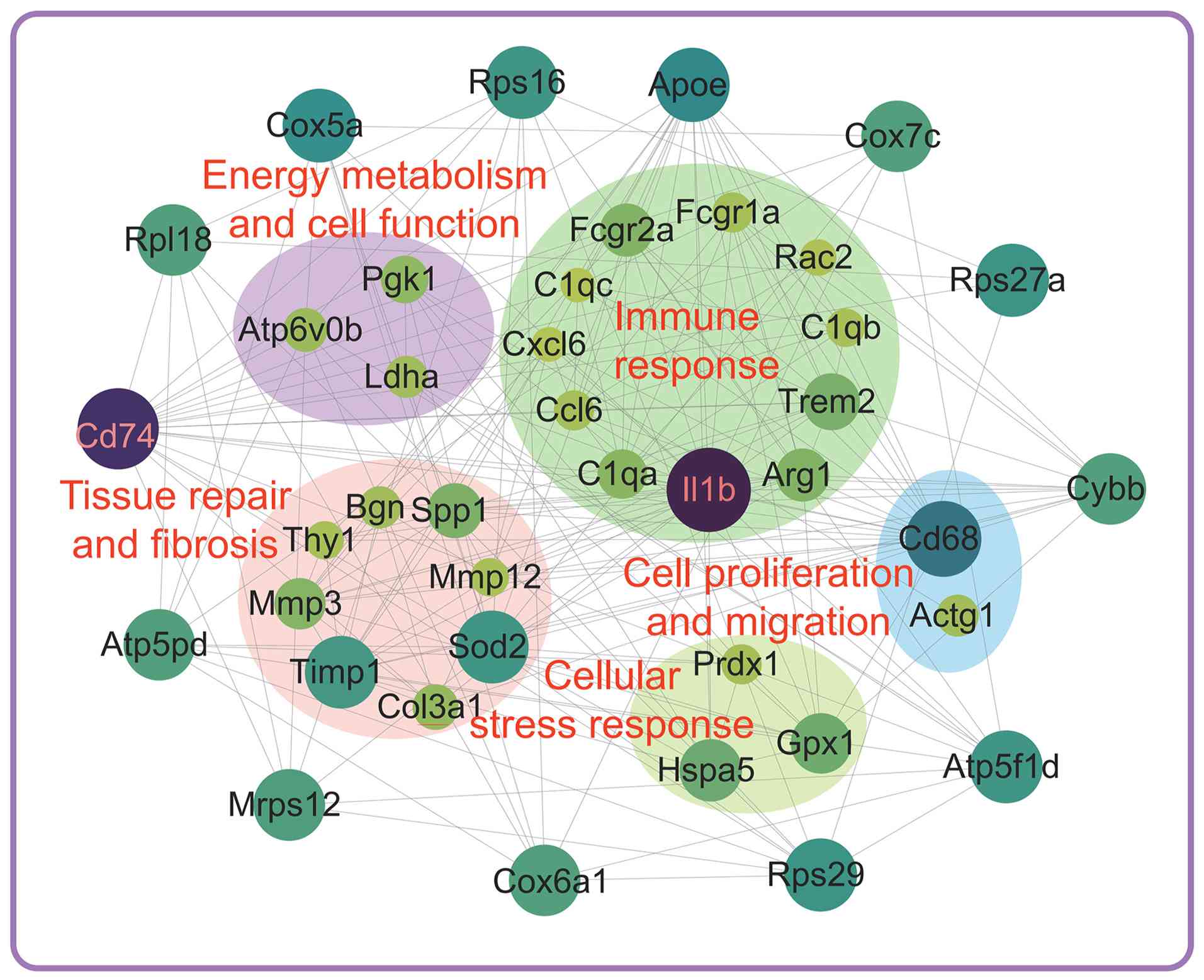
(Fig. 6).
Energy metabolism and cell
function
This module contained genes related to cell energy
metabolism, such as phosphoglycerate kinase 1 (Pgk1) and
lactate dehydrogenase A (Ldha), indicating that urethral
injury reprogrammed cell metabolism to support the repair and
recovery of damaged cells. The changes in genes in this module
suggested that cell energy metabolism may have served a key role
after injury, providing sufficient energy for the repair
process.
Tissue repair and fibrosis
Changes in the expression of genes related to tissue
repair and fibrosis, such as biglycan (Bgn) and matrix
metallopeptidase 3 (Mmp3), revealed that urethral injury may
have triggered a fibrotic response, changing the tissue structure.
The initiation of fibrosis may have affected normal urethral
function, with gene regulatory changes indicating that the repair
process after injury may have been accompanied by unwanted fibrotic
reactions.
Immune response
This module involved immune-related genes, such as
IL-1β complement C1q A chain (C1qa) and complement
C1q B chain (C1qb), whose expression was significantly
upregulated after urethral injury. Activation of these immune genes
supported the important role of the immune system in the repair
process after injury.
Cell proliferation and migration
Genes related to cell proliferation and migration,
such as Cd68 and actin γ1 (Actg1), may have promoted cell
proliferation and migration at the injury site, thereby
facilitating tissue repair after urethral injury. Regulation of
cell proliferation and migration is a critical step in the tissue
repair process, as it directly influences cellular turnover and
structural remodeling during wound healing (28), and changes in gene expression in
this module may reflect the activation of repair-associated
cellular programs. Cellular stress response. Genes involved
in cellular stress responses, such as peroxiredoxin 1, and heat
shock protein family A member 5, may have helped cells to cope with
injury and maintain normal function by regulating stress responses.
The regulation of stress responses is important for cell survival
and functional recovery after injury (27).
After urethral injury, activation of the immune
response was supported by the differential expression of genes,
such as IL-1β, C1qa and C1qb, suggesting that the
immune response may have served an important role in the repair
process. The immune system may have regulated inflammation and
served a key role in fibrosis and tissue repair. Differential
expression of Cd68 and Actg1 suggested that cell
proliferation and migration served key roles in the repair process
after injury. Changes in the expression of genes involved in cell
energy metabolism, such as Pgk1 and Ldha, suggested
that cell metabolism was important for repair after injury by
providing sufficient energy to support cellular activities involved
in tissue repair. The differential expression of genes associated
with fibrosis, such as Bgn and Mmp3, suggested that
fibrosis may have occurred after urethral injury and affected
tissue repair.
Effect of TDN on vimentin
expression
To explore the effect of TDN on vimentin expression,
fibroblasts were successfully isolated from rat urethral tissue and
cultured using an enzymatic digestion method. The cells exhibited
typical adherent growth characteristics (Fig. 7A). H&E staining revealed that
these fibroblasts displayed typical morphological features,
including deeply stained nuclei, lightly stained cytoplasm and
clearly defined cell boundaries (Fig.
7B). To ensure the purity of fibroblasts used for vimentin
regulation assays, immunofluorescence staining of α-SMA and CD90
was performed. Cells exhibited negative α-SMA staining together
with positive CD90 expression, confirming fibroblast identity and
excluding myofibroblast contamination (Fig. 7C). Immunohistochemical analysis
showed high expression of vimentin in all three repeats (Fig. 7D and E).
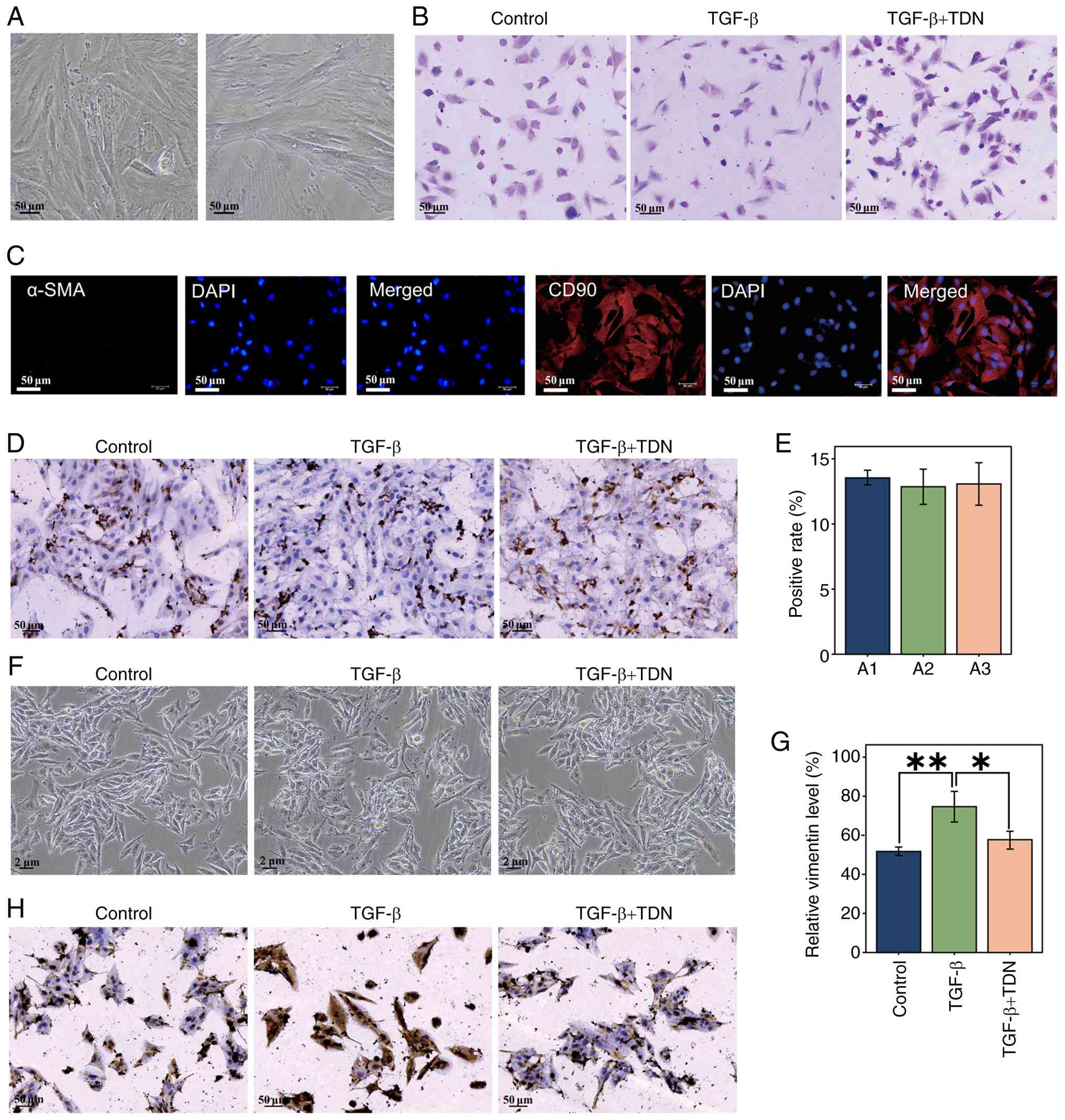 | Figure 7.(A) Fibroblast isolation and
cultivation. Representative phase-contrast images of primary rat
urethral fibroblasts following enzymatic digestion are shown. The
two images represent different microscopic fields from the same
primary culture, illustrating progressive cell adhesion and the
typical spindle-shaped, adherent morphology with elongated cell
bodies and prominent cytoplasmic extensions. (B) H&E staining.
Cell nuclei were stained darkly, with the cytoplasm stained lightly
and clear cell boundaries visible, with cells displaying the normal
morphological characteristics of fibroblasts after H&E
staining. (C) Immunofluorescence staining identification of
fibroblasts. Immunofluorescence staining for α-SMA (green) and CD90
(red) was performed to verify fibroblast purity. Nuclei were
counterstained with DAPI (blue) and imaged at a magnification of
×200 using a Nikon TS2-S-SM fluorescence microscope. Cells
exhibited strong CD90 expression, α-SMA staining was negative,
whereas CD90 staining was clearly positive, confirming fibroblast
identity and excluding myofibroblast contamination. (D)
Immunohistochemical detection of vimentin. Vimentin expression in
fibroblasts was detected by immunohistochemistry. The nuclei were
stained blue, and vimentin-positive cytoplasm appeared
brownish-yellow. Images were obtained at a magnification of ×200
using a digital slide scanner. (E) Immunohistochemical detection of
vimentin. Quantitative analysis among three replicates was
performed using Image-Pro Plus, and statistical evaluation with
GraphPad Prism 9.0 showed no significant differences among groups
(P>0.05), confirming reproducibility. (F) Cell culture under
different treatments. Representative phase-contrast images
(magnification, ×100) showed fibroblasts in the control, TGF-β and
TGF-β + TDN groups, all displaying sufficient adherence and
morphology. (G) Quantitative analysis of vimentin expression under
different treatments. Vimentin expression levels in fibroblasts
from the control, TGF-β and TGF-β + TDN groups were quantified
based on immunohistochemical staining using Image-Pro Plus.
Statistical analysis was performed using GraphPad Prism 9.0
(*P<0.05, **P<0.01). (H) Immunohistochemical detection of
vimentin under different treatments. Representative
immunohistochemical images showed vimentin expression in
fibroblasts from the Control, TGF-β and TGF-β + TDN groups. α-SMA,
α-smooth muscle actin; TDN, tetrahedral DNA nanostructure. |
Subsequently, three experimental conditions were
established: Control, TGF-β-induced vimentin expression and
TGF-β-induced vimentin expression + TDN treatment. Cells showed
satisfactory growth in all groups (Fig. 7F). Immunohistochemical analysis was
conducted to detect vimentin expression after different treatments.
Cell images showed that the nuclei were stained blue, with vimentin
stained brownish-yellow (Fig. 7H).
Quantitative analysis indicated that, compared with the control
group, treatment with TGF-β significantly increased the expression
of vimentin (P<0.01). Treatment with TDN significantly reduced
the expression of vimentin compared with that in the TGF-β group
(P<0.05) (Fig. 7G). These
results suggested that TDN may have regulated urethral fibrosis by
inhibiting TGF-β-induced expression of vimentin.
Discussion
In the present study, a rat model of urethral injury
was successfully established, and the therapeutic effects of
rapamycin and TDN on urethral injury, fibrosis and the immune
response were evaluated. The results indicated that TDN markedly
alleviated urethral injury, inhibited the progression of fibrosis
and modulated immune responses, highlighting its potential as a
therapeutic agent. The key findings and their possible mechanisms
are discussed subsequently.
In the present study, a rat model of urethral injury
was established using a mechanical method closely mimicking
traumatic urethral injury and allowing precise control of the
injury (27,39). The model was validated by multiple
indicators, including retrograde urethral angiography, H&E
staining and pathological analysis. These methods confirmed
urethral stenosis, fibrosis and inflammatory infiltration in the
model group. Both Rapamycin and TDN notably improved pathological
changes, supporting the reliability of the model.
Excessive immune activation is an important driver
of fibrosis after urethral injury (40–42).
Transcriptomic analysis and RT-qPCR revealed increased immune cell
infiltration and elevated levels of the pro-inflammatory cytokines
IL-6, IL-1β and TNF-α in the model group (10). Treatment with TDN significantly
reduced cytokine expression and immune cell infiltration,
indicating effective immunomodulation. Pathway enrichment analysis
also suggested that TDN suppressed the ‘NF-κB signaling pathway’,
‘NOD-like receptor signaling pathway’ and ‘Cytokine-cytokine
receptor interaction’ (43–47),
thereby attenuating immune-inflammatory responses. TDN mitigated
excessive inflammation and fibrosis by limiting macrophage and
T-cell infiltration (48–50).
Fibrosis is the predominant cause of urethral
stenosis and dysfunction (51). In
the present study, TDN significantly reduced collagen deposition
and the expression of fibrosis markers, including α-SMA, collagen
I/III and Smad3, compared with that in the model group.
Transcriptomic analyses indicated that TDN may have prevented the
progression of fibrosis via pathways associated with cellular
metabolism and fibroblast activation. The altered expression levels
of Pgk1 and Ldha suggested that TDN improved energy
metabolism to support repair (52,53),
while changes in Cd74, Bgn and Mmp3 implied direct
regulation of fibrosis-associated genes (54,55).
Inhibition of TGF-β-induced vimentin expression supported the
anti-fibrotic potential of TDN (56).
Tissue repair after urethral injury necessitates
balanced cell proliferation (57–59).
Aberrant upregulation of cell cycle genes, such as Ccnb1, Rrm2,
Plk1 and Cdk1, may enhance tissue fibrosis (60,61).
Treatment with TDN significantly reduced the expression of some of
these genes, whereas others showed a decreasing trend, thereby
preventing excessive fibroblast proliferation and limiting collagen
deposition. Furthermore, by improving energy metabolism via genes
such as Pgk1 and Ldha, TDN indirectly modulated the
cell cycle and maintained controlled proliferation (52,62,63).
Immune responses serve both protective and
pathogenic roles in fibrosis (41). In the present study, transcriptomic
analysis revealed excessive activation of immune-related genes,
including Il-1β, C1qa and C1qb, indicating excessive
activation of pro-inflammatory and complement-associated immune
responses in the model group., consistent with previous reports
showing that immune activation contributes to fibrosis (40,50,64).
TDN downregulated pro-inflammatory cytokines, such as IL-6 and
TNF-α, and prevented immune cell infiltration, thereby attenuating
fibrosis-promoting immune reactions (41,65–68).
These findings suggested that the anti-fibrotic effects of TDN were
closely linked to its immunomodulatory properties.
Although TDN showed significant therapeutic efficacy
in the rat model, further studies are needed to confirm its
potential for clinical application. Additionally, the present study
primarily focused on immune regulation and the inhibition of
fibrosis. Future studies should further investigate the effects of
TDN on specific immune cell subtypes, such as macrophage
polarization states (M1/M2) and distinct lymphocyte subsets, using
more targeted experimental approaches, including flow cytometry,
immunofluorescence co-staining and single-cell RNA sequencing.
Additionally, future work may explore potential synergistic
applications of TDN with other therapeutic agents. In summary, the
present study revealed that TDN promoted repair after urethral
injury by regulating immune responses, suppressing fibrosis and
improving cellular metabolism. These findings not only highlight
the value of TDN as a promising therapeutic candidate for urethral
injury but also provide novel insights into immunoregulatory
strategies for fibrotic diseases.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
The present research was funded by the Guizhou Provincial
Natural Science Foundation [grant no. Qiankehebasis-ZK (2024)
general 241], the Start-up Fund for Doctoral Research at the
Affiliated Hospital of Guizhou Medical University (grant no.
gyfybsky-2022-38), the National Natural Science Foundation of China
Cultivation Program of Affiliated Hospital of Guizhou Medical
University [grant no. gyfynsfc (2023)-62], the Key Medical
Discipline Construction Project of Guizhou Provincial Health
Commission during 2025–2026 and the 2025 Annual Hospital-Level
Scientific Research Fund of Guizhou Hospital of Beijing Jishuitan
Hospital [grant no. JGYYK(2025)-24].
Availability of data and materials
The RNA sequencing data generated in the present
study have been deposited in the NCBI Gene Expression Omnibus (GEO)
database under accession number GSE314890 or at the following URL:
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE314890.
The raw sequencing data (FASTQ files) have been submitted to the
NCBI Sequence Read Archive under BioProject accession number
PRJNA1393091 or at the following URL: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1393091.
Other data generated in this study are available from the
corresponding author upon reasonable request.
Authors' contributions
CG conceived and designed the experiments, performed
the experiments, analyzed the data and drafted the manuscript. JL
contributed to the conceptualization and experimental design of the
study, critically interpreted the data, revised the manuscript for
important intellectual content, supervised the research process and
secured funding. CG and JL confirm the authenticity of all the raw
data. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Experimental
Animal Welfare and Ethics Committee of Guizhou Medical University
(Guiyang, China; approval no. 2502311).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Mundy AR and Andrich DE: Urethral trauma.
Part I: Introduction, history, anatomy, pathology, assessment and
emergency management. BJU Int. 10:310–327. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Rosenstein DI and Alsikafi NF: Diagnosis
and classification of urethral injuries. Urol Clin North Am.
33:73–85. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Verla W, Oosterlinck W, Spinoit AF and
Waterloos M: A comprehensive review emphasizing anatomy, etiology,
diagnosis, and treatment of male urethral stricture disease. Biomed
Res Int. 2019:90464302019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Payne SR, Anderson P, Spasojević N,
Demilow TL, Teferi G and Dickerson D: Male urethral stricture
disease: Why management guidelines are challenging in low-income
countries. BJU Int. 130:157–165. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Rahardjo HE, Märker V, Tsikas D, Kuczyk
MA, Ückert S and Bannowsky A: Fibrotic strategies diseases of the
human urinary and genital tract: Current understanding and
potential for treatment. J Clin Med. 12:47702023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Doersch KM, Barnett D, Chase A, Johnston D
and Gabrielsen JS: The contribution of the immune system to
genitourinary fibrosis. Exp Biol Med (Maywood). 247:765–778. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Metcalfe PD, Wang J, Jiao H, Huang Y, Hori
K, Moore RB and Tredget EE: Bladder outlet obstruction: Progression
from inflammation to fibrosis. BJU Int. 106:1686–1694. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hirano Y, Horiguchi A, Ojima K, Azuma R,
Shinchi M, Ito K and Miyai K: Myofibroblast-dominant proliferation
associated with severe fibrosis in bulbar urethral strictures. Int
J Urol. 30:107–112. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wynn TA: Cellular and molecular mechanisms
of fibrosis. J Pathol. 214:199–210. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lupher ML Jr and Gallatin WM: Regulation
of fibrosis by the immune system. Adv Immunol. 89:245–288. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tiwari P, Verma S, Washimkar KR and
Nilakanth Mugale M: Immune cells crosstalk Pathways, and metabolic
alterations in Idiopathic pulmonary fibrosis. Int Immunopharmacol.
135:1122692024. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cangemi M: Multiparametric immune
profiling to predict the risk of cancer development in chronic
immune suppressed solid organ transplant patients. Doctoral
dissertation. University of Udine; 2019
|
|
13
|
Gedaly R, De Stefano F, Turcios L, Hill M,
Hidalgo G, Mitov MI, Alstott MC, Butterfield DA, Mitchell HC, Hart
J, et al: mTOR inhibitor everolimus in regulatory T cell expansion
for clinical application in transplantation. Transplantation.
103:705–715. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Blagosklonny MV: Rapamycin for longevity:
Opinion article. Aging (Albany NY). 11:8048–8067. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Stallone G, Infante B, Grandaliano G and
Gesualdo L: Management of side effects of sirolimus therapy.
Transplantation. 87 (8 Suppl):S23–S26. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li J, Kim SG and Blenis J: Rapamycin: One
drug, many effects. Cell Metab. 19:373–379. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Keller A and Linko V: Challenges and
perspectives of DNA nanostructures in biomedicine. Angew Chem Int
Ed Engl. 59:15818–15833. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Györfi AH, Matei AE and Distler JHW:
Targeting TGF-β signaling for the treatment of fibrosis. Matrix
Biol. 68–69. 8–27. 2018.PubMed/NCBI
|
|
19
|
Zhao X, Kwan JYY, Yip K, Liu PP and Liu
FF: Targeting metabolic dysregulation for fibrosis therapy. Nat Rev
Drug Discov. 19:57–75. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang W, Lin M, Wang W, Shen Z and Wu S:
DNA tetrahedral nanostructures for the biomedical application and
spatial orientation of biomolecules. Bioactive Materials.
33:279–310. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang Q, Jiang Q, Li N, Dai L, Liu Q, Song
L, Wang J, Li Y, Tian J, Ding B and Du Y: DNA origami as an in vivo
drug delivery vehicle for cancer therapy. ACS Nano. 8:6633–6643.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Shi S, Lin S, Shao X, Li Q, Tao Z and Lin
Y: Modulation of chondrocyte motility by tetrahedral DNA
nanostructures. Cell Proliferation. 50:e123682017. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hong S, Jiang W, Ding Q, Lin K, Zhao C and
Wang X: The Current Progress of Tetrahedral DNA Nanostructure for
Antibacterial Application and Bone Tissue Regeneration. Int J
Nanomedicine. 18:3761–3780. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gu J, Liang J, Tian T and Lin Y: Current
Understanding and Translational Prospects of Tetrahedral Framework
Nucleic Acids. JACS. 5:486–520. 2025.
|
|
25
|
Liang S, Li J, Zou Z, Mao M, Ming S, Lin
F, Zhang Z, Cao, Cao C, Zhou J, et al: Tetrahedral DNA
nanostructures synergize with MnO2 to enhance antitumor
immunity via promoting STING activation and M1 polarization. Acta
Pharmaceutica Sinica B. 12:2494–2505. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Cui W, Yang X, Dou Y, Du Y, Ma X, Hu L and
Lin Y: Effects of tetrahedral DNA nanostructures on the treatment
of osteoporosis. Cell Proliferation. 57:e136252024. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wu Z, Tang Z, Zheng Z and Tan S: A novel
trauma induced urethral stricture in rat model. Sci Rep.
14:63252024. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Hyuga T, Fujimoto K, Hashimoto D, Tanabe
K, Kubo T, Nakamura S, Ueda Y, Fujita-Jimbo E, Muramatsu K, Suzuki
K, et al: Wound healing responses of urinary extravasation after
urethral injury. Sci Rep. 13:106282023. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
She L, Niu B, Wu X, Yin Y, Wen K, Li H,
Xie D, Zheng X and Dai Y: Iatrogenic injury recapitulated:
Electroexcision technique for urethral stricture modeling in rats.
J Vis Exp. 212:668642024.
|
|
30
|
Abdelkhalek AS, Clarke PD, Sommers MA, Oe
T, Andersen TM, Andersen CT, Hejbøl EK, Schrøder HD and Zvara P:
Validation of a new rat model of urethral sphincter injury and leak
point pressure measurements. Scand J Urol. 55:498–504. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Aydın A, Sönmez MG, Oltulu P, Kocabaş R,
Öztürk Sönmez L, Özcan S, Boğa MS and Balasar M: Histopathologic
evaluation of the effects of intraurethral platelet rich plasma in
urethral trauma experimentally induced in rat model. Urology.
141:187.e9–187.e14. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zhang SL and Wong AW: A novel technique
for atraumatic transurethral catheterisation of male rats. Biol
Open. 13:bio0604762024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Bauman TM, Nicholson TM, Abler LL,
Eliceiri KW, Huang W, Vezina CM and Ricke WA: Characterization of
fibrillar collagens and extracellular matrix of glandular benign
prostatic hyperplasia nodules. PLoS One. 9:e1091022014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Shen H, Wang J, Min J, Xi W, Gao Y, Yin L,
Yu Y, Liu K, Xiao J, Zhang YF and Wang ZN: Activation of
TGF-β1/α-SMA/Col I profibrotic pathway in fibroblasts by galectin-3
contributes to atrial fibrosis in experimental models and patients.
Cell Physiol Biochem. 47:851–863. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Taylor SC and Posch A: The design of a
quantitative western blot experiment. Biomed Res Int.
2014:3615902014. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Modi A, Vai S, Caramelli D and Lari M: The
Illumina sequencing protocol and the NovaSeq 6000 system. Methods
Mol Biol. 2242:15–42. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Bindea G, Mlecnik B, Tosolini M,
Kirilovsky A, Waldner M, Obenauf AC, Angell H, Fredriksen T,
Lafontaine L, Berger A, et al: Spatiotemporal dynamics of
intratumoral immune cells reveal the immune landscape in human
cancer. Immunity. 39:782–795. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Ergün O, Tepebaşi MY, Onaran İ, Öztürk SA,
Baltik M and Koşar PA: Standardizing urethral stricture models in
rats: A comprehensive study on histomorphologic and molecular
approach. Int Urol Nephrol. 56:2945–2954. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Antar SA, Ashour NA, Marawan ME and
Al-Karmalawy AA: Fibrosis: Types, effects, markers, mechanisms for
disease progression, and its relation with oxidative stress,
immunity, and inflammation. Int J Mol Sci. 24:40042023. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Huang E, Peng N, Xiao F, Hu D, Wang X and
Lu L: The roles of immune cells in the pathogenesis of fibrosis.
Int J Mol Sci. 21:52032020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ising C and Heneka MT: Functional and
structural damage of neurons by innate immune mechanisms during
neurodegeneration. Cell Death Dis. 9:1202018. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Qiu D, Zhang D, Yu Z, Jiang Y and Zhu D:
Bioinformatics approach reveals the critical role of the NOD-like
receptor signaling pathway in COVID-19-associated multiple
sclerosis syndrome. J Neural Transm (Vienna). 129:1031–1038. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Tieri P, Termanini A, Bellavista E,
Salvioli S, Capri M and Franceschi C: Charting the NF-κB pathway
interactome map. PLoS One. 7:e326782012. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Shawky E, Nada AA and Ibrahim RS:
Potential role of medicinal plants and their constituents in the
mitigation of SARS-CoV-2: Identifying related therapeutic targets
using network pharmacology and molecular docking analyses. RSC Adv.
10:27961–27983. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wang M, Li X, Yang Z, Chen Y, Shu T and
Huang Y: LncRNA MEG3 alleviates interstitial cystitis in rats by
upregulating Nrf2 and inhibiting the p38/NF-κB pathway. Cytokine.
165:1561692023. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Naiyila X, Li J, Huang Y, Chen B, Zhu M,
Li J, Chen Z, Yang L, Ai J, Wei Q, et al: A novel insight into the
immune-related interaction of inflammatory cytokines in benign
prostatic hyperplasia. J Clin Med. 12:18212023. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Barron L and Wynn TA: Fibrosis is
regulated by Th2 and Th17 responses and by dynamic interactions
between fibroblasts and macrophages. Am J Physiol Gastrointest
Liver Physiol. 300:G723–G728. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhang M and Zhang S: T cells in fibrosis
and fibrotic diseases. Front Immunol. 11:11422020. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Wynn TA and Barron L: Macrophages: Master
regulators of inflammation and fibrosis. Semin Liver Dis.
30:245–257. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Lima NA, Vasconcelos CC, Filgueira PH,
Kretzmann M, Sindeaux TA, Feitosa Neto B, Silva Junior GB and Daher
EF: Review of genitourinary tuberculosis with focus on end-stage
renal disease. Rev Inst Med Trop Sao Paulo. 54:57–60. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Gou R, Hu Y, Liu O, Dong H, Gao L, Wang S,
Zheng M, Li X and Lin B: PGK1 is a key target for anti-glycolytic
therapy of ovarian cancer: Based on the comprehensive analysis of
glycolysis-related genes. Front Oncol. 11:6824612021. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Wang P, Wang Q, Yang X, An Y, Wang J, Nie
F, Pan B, Bi H and Qin Z: Targeting the glycolytic enzyme PGK1 to
inhibit the warburg effect: A new strategy for keloid therapy.
Plast Reconstr Surg. 151:970e–980e. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Lv W, Wu M, Ren Y, Luo X, Hu W, Zhang Q
and Wu Y: Treatment of keloids through Runx2 siRNA-induced
inhibition of the PI3K/AKT signaling pathway. Mol Med Rep.
23:552021. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Giannandrea M and Parks WC: Diverse
functions of matrix metalloproteinases during fibrosis. Dis Model
Mech. 7:193–203. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Cheng F, Shen Y, Mohanasundaram P,
Lindström M, Ivaska J, Ny T and Eriksson JE: Vimentin coordinates
fibroblast proliferation and keratinocyte differentiation in wound
healing via TGF-β-Slug signaling. Proc Natl Acad Sci USA.
113:E4320–E4327. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Diaz-Moralli S, Tarrado-Castellarnau M,
Miranda A and Cascante M: Targeting cell cycle regulation in cancer
therapy. Pharmacol Ther. 138:255–271. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Campos A and Clemente-Blanco A: Cell cycle
and DNA repair regulation in the damage response: Protein
phosphatases take over the reins. Int J Mol Sci. 21:4462020.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Vermeulen K, Van Bockstaele DR and
Berneman ZN: The cell cycle: A review of regulation, deregulation
and therapeutic targets in cancer. Cell Prolif. 36:131–149. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Baek A, Shin JC, Lee MY, Kim SH, Kim J and
Cho SR: Parasympathetic effect induces cell cycle activation in
upper limbs of paraplegic patients with spinal cord injury. Int J
Mol Sci. 20:59822019. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Darby IA and Hewitson TD: Hypoxia in
tissue repair and fibrosis. Cell Tissue Res. 365:553–562. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Icard P, Fournel L, Wu Z, Alifano M and
Lincet H: Interconnection between metabolism and cell cycle in
cancer. Trends Biochem Sci. 44:490–501. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
He Y, Luo Y, Zhang D, Wang X, Zhang P, Li
H, Ejaz S and Liang S: PGK1-mediated cancer progression and drug
resistance. Am J Cancer Res. 9:2280–2302. 2019.PubMed/NCBI
|
|
64
|
Absinta M, Maric D, Gharagozloo M, Garton
T, Smith MD, Jin J, Fitzgerald KC, Song A, Liu P, Lin JP, et al: A
lymphocyte-microglia-astrocyte axis in chronic active multiple
sclerosis. Nature. 597:709–714. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Wick G, Grundtman C, Mayerl C,
Wimpissinger TF, Feichtinger J, Zelger B, Sgonc R and Wolfram D:
The immunology of fibrosis. Annu Rev Immunol. 31:107–135. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Gieseck RL III, Wilson MS and Wynn TA:
Type 2 immunity in tissue repair and fibrosis. Nat Rev Immunol.
18:62–76. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Möller B and Villiger PM: Inhibition of
IL-1, IL-6, and TN. Semin Immunopathol. 27:391–408. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Wang T and He CJ: TNF-α and IL-6: The link
between immune and bone system. Curr Drug Targets. 21:213–227.
2020. View Article : Google Scholar : PubMed/NCBI
|