Introduction
Intervertebral disc degeneration (IDD) is a major
pathological basis of spinal degenerative diseases, and a key
factor leading to lumbar disc herniation, spinal stenosis and
chronic low back pain (1).
Epidemiological studies have shown that the prevalence of IDD
increases with age, and is influenced by genetic factors,
mechanical load, inflammatory responses and environmental
conditions (2,3). Due to the complex pathogenesis of
IDD, there is currently no effective treatment to reverse or halt
its progression. Existing therapeutic approaches primarily focus on
symptom relief, including pain medications, physical therapy and
surgical interventions. However, with advancements in molecular
biology, the role of epigenetic regulation in IDD has gained
increasing attention and is considered a promising avenue for
potential therapeutic targets at present, as it offers
opportunities to modulate gene expression involved in extracellular
matrix (ECM) metabolism, inflammation and cell senescence without
altering the underlying DNA sequence. By targeting these reversible
epigenetic changes, such as DNA methylation, histone modifications
and non-coding RNAs, future therapies may be able to slow or even
reverse disc degeneration rather than merely alleviate symptoms
(4).
Epigenetics refers to regulatory mechanisms that
influence gene expression without altering the DNA sequence, mainly
through reversible chemical modifications or changes in nucleosome
structure. These mechanisms include DNA methylation, histone
modifications, non-coding RNA (ncRNA)-mediated regulation and
chromatin remodeling (5). These
regulatory pathways serve crucial roles in the onset and
progression of IDD (6). For
example, DNA methylation can influence the expression of genes such
as MMPs and type II collagen (COL2A1), histone modifications can
alter the transcriptional state of intervertebral disc cells, and
ncRNAs are involved in regulating apoptosis, inflammatory responses
and ECM metabolism (7–9). Additionally, IDD is often associated
with chronic inflammation and oxidative stress, where epigenetic
modifications serve a key role in regulating inflammatory mediators
and metabolic pathways (6).
The present review systematically explores the role
of epigenetic regulation in IDD, focusing on how mechanisms such as
DNA methylation, histone modifications, and ncRNAs-mediated
regulation influence IDD, and how metabolic disorders may modulate
these epigenetic processes. Furthermore, the present review
discusses the potential therapeutic implications of targeting these
epigenetic mechanisms in IDD treatment.
Role of DNA methylation in IDD
Overall association between DNA
methylation and IDD
A previous study employing genome-wide methylation
analysis identified hypermethylation in advanced degenerative discs
compared with early-stage discs (10). Further evidence has demonstrated
that numerous genes related to stress responses, matrix catabolism
and cell death are regulated by aberrant DNA methylation in
degenerative discs (11).
In particular, the upregulation of the DNA
methyltransferase (DNMT)3B has received considerable attention.
DNMT3B expression is regarded as a prominent feature of
degenerative nucleus pulposus (NP) cells (12). In both in vitro and in
vivo models, blocking DNMT activity [e.g., using the inhibitor
5-azacytidine (5-AZA)] can reduce ferroptosis and oxidative stress,
ultimately slowing NP cell degeneration and ECM breakdown (12), as schematically illustrated in
Fig. 1, which depicts the
regulatory roles of DNMTs and TET enzymes in methylation-mediated
ferroptosis and ECM metabolism.
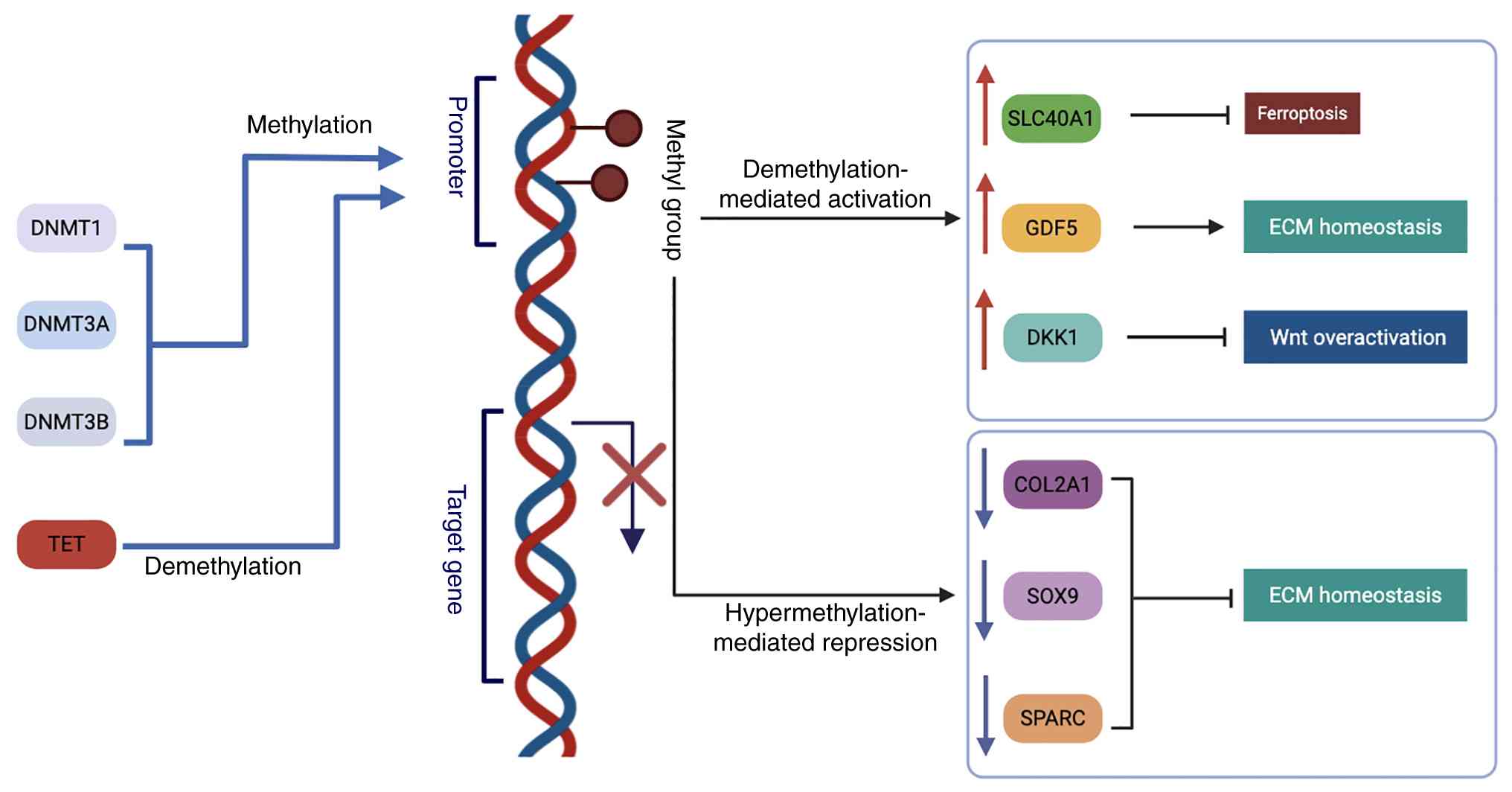 | Figure 1.Critical role of DNA methylation
regulation in IDD. DNA methyltransferases (DNMT1, DNMT3A and
DNMT3B) mediate promoter methylation and repress target genes,
whereas TET enzymes promote demethylation and transcriptional
activation. Demethylation of SLC40A1 and GDF5 restores their
expression, thereby regulating ferroptosis resistance and ECM
homeostasis. Demethylation-mediated upregulation of DKK1, a Wnt
antagonist, inhibits Wnt signaling activity. By contrast,
hypermethylation of COL2A1, SOX9 and SPARC represses their
expression, leading to ECM degradation and accelerating IDD
progression. COL2A1, type II collagen; DKK1, dickkopf Wnt signaling
pathway inhibitor 1; DNMT, DNA methyltransferase; ECM,
extracellular matrix; GDF5, growth differentiation factor 5; IDD,
intervertebral disc degeneration; SLC40A1, solute carrier family 40
member 1; SPARC, secreted protein acidic and cysteine rich; TET,
ten-eleven translocation. |
Abnormal methylation of key genes
Research has indicated that secreted protein acidic
and cysteine rich (SPARC), an important regulator of ECM
homeostasis, is hypermethylated in aging and degenerative discs,
leading to suppressed SPARC expression, which disrupts matrix
integrity and is closely associated with chronic pain (13). Growth differentiation factor 5
(GDF5) has a critical CpG site in its 5′ untranslated region (UTR),
where increased methylation alters transcription factor (SP1/SP3)
binding, reducing GDF5 expression and increasing the risk of
multiple musculoskeletal disorders, including lumbar disc
degeneration (14).
Regarding ferroptosis and oxidative stress,
silencing of the solute carrier family 40 member 1 (SLC40A1) gene
by DNMT3B-mediated hypermethylation makes NP cells more vulnerable
to ferroptosis and oxidative stress (12). Conversely, restoring SLC40A1
expression by inhibiting DNMT3B can ease disc damage (15). In addition, epigenetic regulation
of apoptotic pathways is evident in the methylation-dependent
upregulation of certain microRNAs (miRNAs/miRs; for example,
miR-143), which suppress BCL2 and increase NP cell apoptosis
(16).
Autophagy and senescence pathways also display
altered methylation patterns. For instance, miR-129-5p is
downregulated in degenerative discs due to hypermethylation in its
promoter region; consequently, its reduced suppression of Beclin-1
affects normal autophagy homeostasis (17). E4F transcription factor 1, another
key factor in cell senescence, is also epigenetically inhibited
when DNMT3B is upregulated by ALKBH5-mediated
N6-methyladenosine (m6A) demethylation,
compounding the degenerative process (18).
Regulatory mechanisms and network
interactions
DNA methylation not only affects gene transcription
but also interacts with other epigenetic processes, such as histone
modifications, ncRNA-mediated regulation and RNA methylation, to
amplify injury from senescence, inflammation and oxidative stress
(6). Inflammatory signals can
stimulate DNMT and ten-eleven translocation (TET) enzyme
expression, while oxidative stress can modulate the activity of
methyltransferases via the p38/MAPK pathway, increasing catabolic
enzyme expression (19).
Histone modifications and IDD
Histone modifications, including acetylation,
methylation, phosphorylation and ubiquitination, are essential
epigenetic mechanisms that influence chromatin structure and gene
expression (6). In the context of
IDD, growing evidence indicates that dysregulation of these
modifications contributes to pathological processes such as ECM
degradation, inflammation, cellular senescence and apoptosis
(20,21). Among these various modifications,
histone acetylation/deacetylation and histone
methylation/demethylation have been most prominently studied
(18). This section summarizes the
findings on these two major modifications and their regulatory
factors in IDD (Fig. 2), which are
supported by recent studies (21,22).
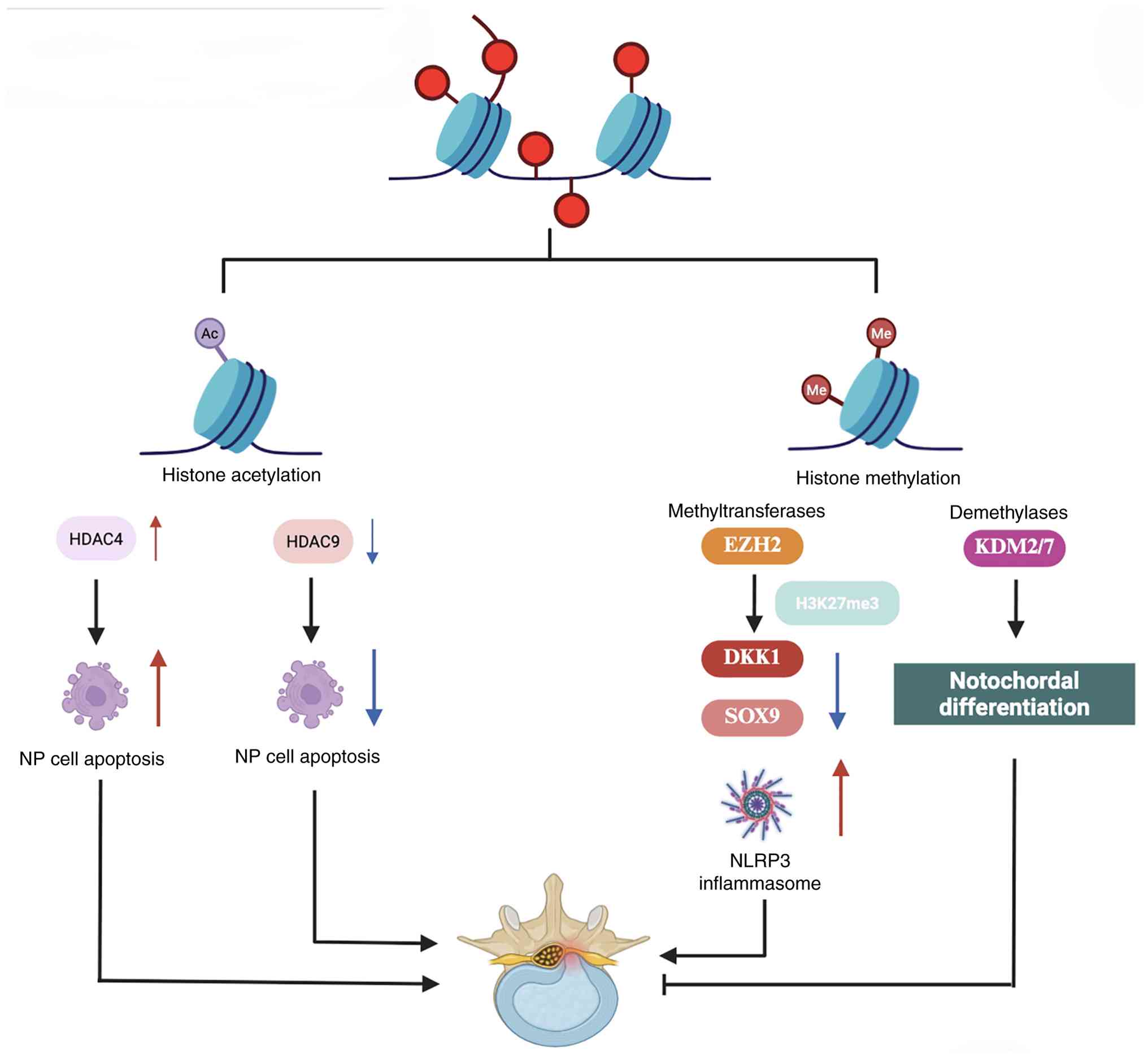 | Figure 2.Role of histone modifications in IDD.
Histone acetylation and methylation influence IDD pathogenesis.
HDAC4 promotes NP cell apoptosis, whereas HDAC9 exerts protective
effects. EZH2-mediated H3K27me3 suppresses SOX9 expression,
enhancing NLRP3 inflammasome activation and extracellular matrix
degradation. By contrast, KDM2/7 promotes notochordal
differentiation, suggesting a potential therapeutic approach for
IDD. DKK1, dickkopf Wnt signaling pathway inhibitor 1; EZH2,
enhancer of zeste homolog 2; H3K27me3, trimethylation of histone H3
lysine 27; HDAC, histone deacetylase; IDD, intervertebral disc
degeneration; KDM2/7, lysine demethylase 2/7; NLRP3, NLR family
pyrin domain containing 3; NP, nucleus pulposus; Me, methyl
group. |
Role of histone
acetylation/deacetylation [histone acetyltransferases
(HATs)/histone deacetylases (HDACs)] in IDD
Histone acetylation, mediated by HATs, reduces the
positive charge of lysine residues, leading to chromatin relaxation
and promoting gene transcription (23). Conversely, HDACs remove acetyl
groups, resulting in chromatin condensation and transcriptional
repression. Emerging evidence suggests that dysregulation of HDACs
serves a crucial role in IDD (22).
Studies have demonstrated that HDAC9 expression is
reduced in degenerative NP cells (22,24).
Under normal conditions, HDAC9 deacetylates and stabilizes RUNX
family transcription factor 3 (RUNX3) to maintain cell viability.
Loss of HDAC9 expression in IDD diminishes RUNX3 stability, thereby
promoting NP cell apoptosis (22).
By contrast, HDAC4 has been shown to promote IDD progression by
upregulating the Krüppel-like factor 5 (KLF5) and apoptosis
signal-regulating kinase 1 signaling axis, which accelerates ECM
degradation (24). These findings
highlight the complex role of histone acetylation/deacetylation in
IDD pathogenesis and suggest that selective HDAC inhibition may
offer novel therapeutic potential for mitigating disc
degeneration.
Role of histone methylation and its
regulatory factors in IDD
In the pathogenesis and progression of IDD, histone
methylation has been shown to serve a crucial regulatory role in
key processes such as ECM homeostasis, cellular senescence,
autophagy and inflammation (21).
This modification is primarily mediated by methyltransferases [such
as enhancer of zeste homolog 2 (EZH2), lysine methyltransferase 2A
(KMT2A) and SUV39H2] and demethylases [such as lysine demethylase
(KDM)2/7, KDM4B and KDM3A], which add or remove methyl groups on
specific lysine or arginine residues of histones, thereby
influencing chromatin conformation and gene transcription (25,26).
Multiple studies have demonstrated that EZH2, a
methyltransferase responsible for trimethylation of histone H3
lysine 27 (H3K27), is elevated in degenerative discs (27,28).
By enriching trimethylation of histone H3 lysine 27 (H3K27me3) in
specific gene promoter regions, EZH2 suppresses gene expression,
accelerating the senescence of NP cells and promoting ECM
degradation (28). For example,
inhibition of EZH2 can remove H3K27me3 marks at the SOX9 promoter,
upregulate cartilage phenotype-related genes such as SOX9, and curb
ECM breakdown, thus counteracting endplate and NP degeneration
(27). Additionally, EZH2-mediated
H3K27me3 can silence key regulatory factors such as dickkopf Wnt
signaling pathway inhibitor 1 or miR-129-5p, thereby activating the
NLRP3, NLR family apoptosis inhibitory protein/NLR family CARD
domain containing 4 or MAPK1 pathways, and further exacerbating
pyroptosis and inflammation in NP cells (21,28).
These findings suggest that inhibiting EZH2 expression or activity
may be a potential strategy to delay disc degeneration.
Besides EZH2, other methyltransferases also serve
essential roles in the pathological progression of IDD. SUV39H2 can
specifically monomethylate protein phosphatase 1 catalytic subunit
α at K141, disrupting its interaction with transcription factor EB
(TFEB), and hampering TFEB nuclear translocation and autophagy gene
activation, ultimately inducing cell senescence and disc
degeneration (29). KMT2A (MLL1)
enhances METTL3 expression via H3K4me3 enrichment at the METTL3
promoter, which increases m6A modification and
downregulates autophagy related (ATG)4a transcription. This leads
to impaired autophagy, GATA binding protein 4 activation and a
senescent phenotype in NP cells (30). Such findings highlight the
interplay among histone methylation, autophagy and cellular
senescence, offering novel insights into IDD mechanisms.
In contrast to methyltransferases, histone
demethylases (for example, the KDM2, KDM4 and KDM3 families) remove
methyl groups from specific residues, thus either activating or
repressing target genes (31).
Research has shown that targeted inhibition of KDM2/7, using either
gene editing or pharmacological approaches, may promote the
differentiation of human induced pluripotent stem cells into
notochord-like cells, potentially opening novel avenues for
regenerative therapy against IDD (32). Furthermore, in a 400 mOsm
environment, KDM4B upregulates NP marker genes and enhances stem
cell differentiation into NP-like cells (33). This finding highlights the crucial
role of histone demethylation in maintaining disc cell identity
under osmotic stress, a key pathological factor in IDD. By
promoting notochordal and NP-like differentiation, KDM4B-mediated
demethylation may help restore the regenerative potential of
degenerated discs. Therefore, targeting KDM4B activity or mimicking
its epigenetic effects could represent a promising therapeutic
approach for disc repair and regeneration. However, excessive
demethylase activity could trigger abnormal regulation of
downstream genes. For instance, increased KDM3A may elevate
hypoxia-inducible factor 1α levels, provoking disturbances in
autophagy and apoptosis among NP cells; thus, precisely regulating
demethylase activities remains a crucial therapeutic goal (34).
Role of ncRNAs in IDD
ncRNAs have attracted attention in the study of IDD.
ncRNAs, including miRNAs, long non-coding RNAs (lncRNAs) and
circular RNAs (circRNAs), regulate various cellular processes
related to IDD, including ECM metabolism, inflammation, oxidative
stress, apoptosis and ferroptosis (35).
Role of miRNAs in IDD
miRNAs are endogenous small RNAs, 20–22 nucleotides
in length, and regulate gene expression by binding to the 3′UTR of
target mRNAs, thereby inhibiting translation or promoting
degradation (36,37). During IDD, miRNAs serve key
regulatory roles in NP cells, annulus fibrosus cells and
inflammation-related processes (38). Studies have demonstrated that
miRNAs, through their specific targeting mechanisms, influence ECM
synthesis, autophagy and inflammatory pathways (39,40).
For example, a regulatory network involving multiple miRNAs has
been identified as crucial in IDD progression (41).
During IDD pathogenesis, some miRNAs affect matrix
metabolism and apoptosis. For instance, miR-141-5p promotes IDD
progression by regulating circRNA_0000253 (42). Additionally, miR-141 has been shown
to accelerate IDD by inhibiting the sirtuin (SIRT)1/NF-κB pathway
(43).
In IDD-related inflammation regulation, studies have
indicated that miRNAs influence the expression of inflammatory
factors by modulating specific pathways (44,45).
For example, miRNA-222 serves a role in IDD progression by
regulating MMP1 expression (46).
Furthermore, the interaction between miRNAs and autophagy is also
considered a key factor in IDD progression (47).
Notably, the function of certain miRNAs may depend
on different stages of IDD or specific microenvironments (38). This regulatory complexity suggests
that miRNAs could serve as potential therapeutic targets for IDD
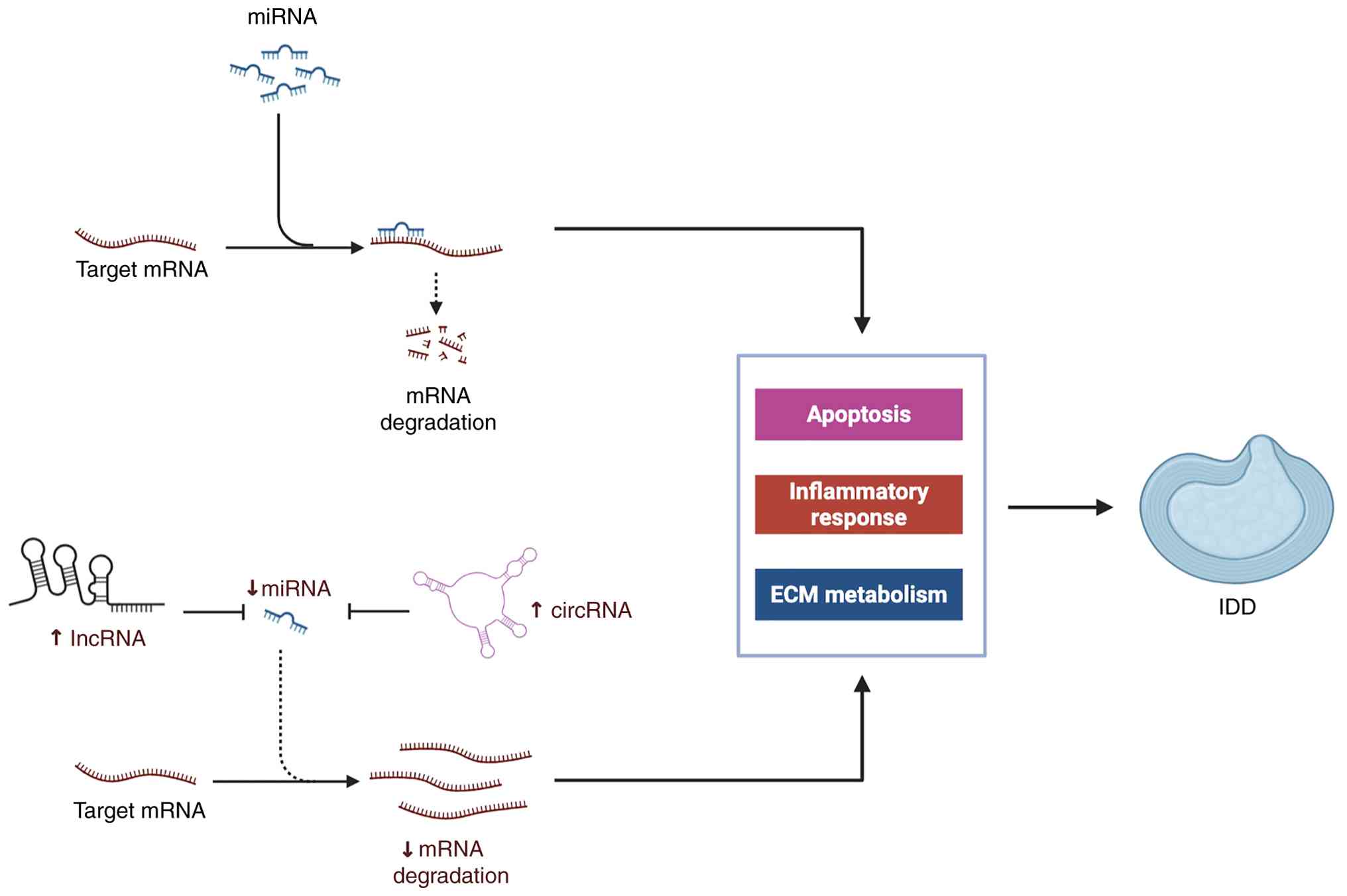
(48). As illustrated in Fig. 3, miRNAs repress target messenger
RNAs (mRNAs), thereby regulating key biological processes such as
apoptosis, inflammation, and ECM metabolism. In addition, long
non-coding RNAs (lncRNAs) and circular RNAs (circRNAs) act as
competing endogenous RNAs (ceRNAs) that modulate miRNA activity,
further influencing IDD pathogenesis (49).
lncRNAs in IDD
lncRNAs, which are transcripts longer than 200
nucleotides without significant protein-coding potential, serve
pivotal roles in IDD. By acting as competing endogenous RNAs
(ceRNAs), interacting with proteins or regulating RNA modifications
(such as m6A), lncRNAs influence key processes such as
NP cell proliferation, apoptosis, ECM catabolism and inflammatory
responses (50,51). Current studies have identified a
variety of lncRNAs that either promote or ameliorate IDD (52,53).
Several lncRNAs have been shown to aggravate IDD by
promoting NP cell apoptosis, inflammation or ECM degradation. For
instance, OIP5-AS1 is highly expressed in degenerative discs and
exacerbates these pathological changes by sponging miR-25-3p
(54). LINC01121 is also induced
by IL-1 and TNF-α, and accelerates disc degeneration through the
miR-150-5p/MMP16 axis, elevating inflammation-related enzymes such
as MMP-3 and ADAM metallopeptidase with thrombospondin type 1 motif
5 (ADAMTS5) (55). Similarly,
LINC00324 is upregulated in IDD and enhances Fas ligand expression,
thereby exacerbating NP cell apoptosis (56). HOTAIR promotes pro-degenerative
events by targeting miR-130b and inhibiting the PTEN/AKT pathway,
leading to reduced CyclinD1 expression and impaired NP-cell
proliferation (57). In addition,
HCG18 augments apoptosis and inflammation via the
miR-495-3p/follistatin like 1 axis (58), while FAF1 (referring to a lncRNA
transcript in that region) is associated with advanced degeneration
grades and activates the Erk signaling pathway (59). Beyond ceRNA interactions, aberrant
m6A modification also contributes to IDD. WTAP-mediated
hypermethylation of NORAD accelerates its degradation via YTH
N6-methyladenosine RNA binding protein F2, reducing the
ability of NORAD to restrain PUMILIO and thereby promoting cellular
senescence (51).
By contrast, some lncRNAs exhibit protective
functions against IDD. KLF3-AS1 is downregulated in degenerative
discs but can improve NP cell viability and suppress apoptosis and
ECM breakdown by sequestering miR-10a-3p, thereby upregulating zinc
finger and BTB domain containing 20 (60). RP11-81H3.2 also serves a protective
role by binding miR-1539, relieving its inhibitory effect on COL2A1
and mitigating NP cell apoptosis (61). Additionally, LINC00689 counters
disc degeneration by sponging miR-3127-5p and activating the
autophagy-related gene ATG7, which promotes autophagy and
diminishes apoptosis in NP cells (62).
circRNAs in IDD
ncRNAs, particularly circRNAs, have attracted
attention in studies of IDD. With their high stability and
resistance to degradation, circRNAs can bind to proteins or miRNAs
in ways that influence NP-cell proliferation, apoptosis, ECM
metabolism and inflammatory responses, thereby contributing to the
development and progression of IDD (63,64).
Multiple studies have demonstrated that circRNAs
frequently show aberrant expression in degenerative NP tissues and
can aggravate IDD by regulating crucial pathways. For example, Chen
et al (65) found that
circGPATCH2L was highly expressed in degenerated NP tissues when in
a hypomethylated (m6A) state, enabling it to bind to and
block the phosphorylation of tripartite motif containing 28,
thereby inhibiting P53 degradation and promoting DNA damage and
apoptosis. Du et al (66)
reported that hsa_circ_0083756 was upregulated in degenerative NP
tissues and cells and, by sponging miR-558, it upregulated
triggering receptor expressed on myeloid cells 1 expression,
ultimately suppressing NP cell proliferation and ECM formation.
Meng and Xu (67) demonstrated
that hsa_circ_0001658 exerted a protective effect in IDD by
sponging miR-181c-5p and enhancing FAS expression, thereby
inhibiting degeneration. Wang et al (68) identified circEYA3 as a key
regulator of intervertebral disc degeneration. circEYA3 acts as a
sponge for miR-196a-5p to upregulate EBF1 (early B-cell factor 1),
thereby activating NF-κB signaling and promoting NP cell apoptosis
and ECM degradation. Yan et al (69) reported that hsa_circ_0134111 was
upregulated in IDD tissues and could bind to miR-578, thereby
increasing the levels of ADAMTS5 and MMP-9 to promote ECM
degradation and inflammation.
These findings collectively indicate that circRNAs
serve pivotal regulatory roles in IDD. They often serve as miRNA
‘sponges’ or interact with specific proteins to disrupt the dynamic
balance of ECM metabolism, cell proliferation and apoptosis, and
inflammatory responses (70). At
the same time, they offer novel avenues for potential clinical
interventions. Strategies, such as inhibiting the upregulation of
particular circRNAs or enhancing their physiological degradation,
could represent promising approaches for slowing or preventing disc
degeneration in the future.
Interaction between metabolic regulation and
epigenetics in IDD
In the development and progression of IDD, cellular
metabolic disorders often interact with epigenetic modifications,
forming a complex regulatory network that jointly drives
degeneration (6). Studies have
shed light on the close relationship between the two in the
following aspects (Fig. 4).
Specifically, recent research indicates that reactive oxygen
species (ROS), NAD+, lactate, and α-ketoglutarate
regulate histone and DNA modifications, thereby influencing
cellular senescence, ECM homeostasis, and inflammation, ultimately
driving the initiation and progression of IDD (6,71).
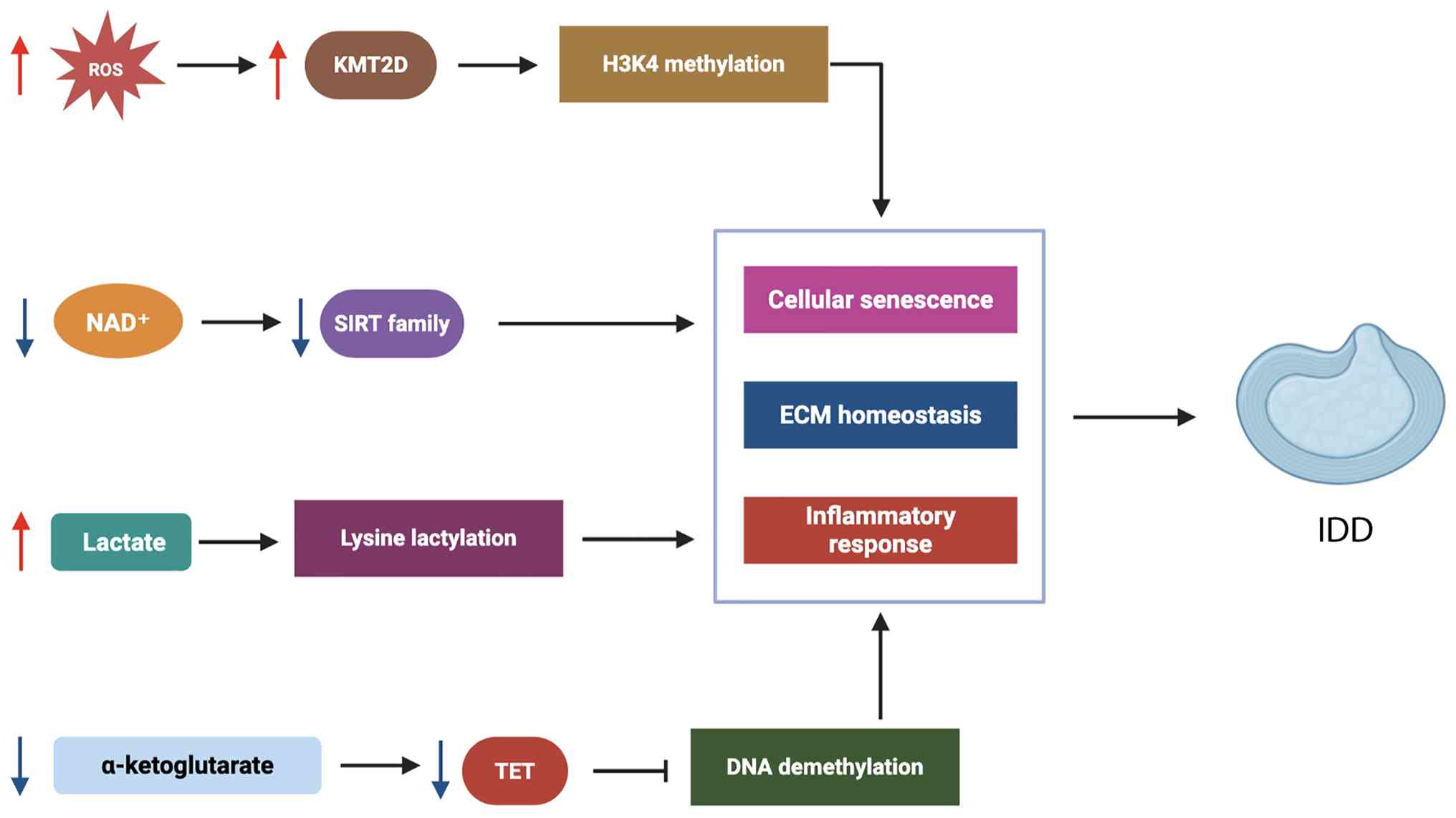 | Figure 4.Interaction between metabolic
regulation and epigenetics in IDD. ROS, NAD+, lactate
and α-ketoglutarate regulate histone and DNA modifications, thereby
affecting cellular senescence, ECM homeostasis and inflammation.
These metabolism-epigenetics interactions drive the initiation and
progression of IDD. ECM, extracellular matrix; H3K4, histone H3
lysine 4; IDD, intervertebral disc degeneration; KMT2D, lysine
methyltransferase 2D; ROS, reactive oxygen species; SIRT, sirtuin;
TET, ten-eleven translocation. |
Oxidative stress and epigenetic
modifications
Excessive reactive oxygen species (ROS) can activate
p38/MAPK, ERK and other signaling pathways, thereby affecting the
stability and transcriptional activity of histone
methyltransferases (such as KMT2D) and other epigenetic factors
(for example, DNMTs and HDACs). This leads to the upregulation of
matrix-degrading genes (MMPs and ADAMTS) and exacerbates ECM
degradation (19). Notably,
oxidative stress is not only a product of metabolic imbalance but
also amplifies degeneration via DNA and histone modifications
(19).
Regulation of epigenetic enzyme
activity by key metabolic substrates
NAD+ is a cofactor for the SIRT family,
directly influencing the catalytic efficiency of deacetylases such
as SIRT1 and SIRT6. When NAD+ levels decline, SIRT
activity is diminished, weakening the resilience of intervertebral
disc cells against external stress. Conversely, maintaining higher
NAD+ levels helps regulate autophagy and ECM homeostasis
(72). Additionally,
α-ketoglutarate (α-KG) can serve as a cofactor for TET
demethylases, influencing DNA and histone demethylation (73). Recent findings suggest that α-KG
inhibits excessive expression of inflammatory or fibrotic genes in
disc cells and possibly slows degenerative processes by
upregulating demethylase activity (74).
Novel modifications linked to
metabolic products
Lysine lactylation (Kla), a histone modification
dependent on the intracellular lactate concentration, is
increasingly recognized (75). In
the hypoxic environment of the disc, anaerobic glycolysis leads to
lactate accumulation, and the resulting Kla modifications may
regulate key biological processes such as cellular aging, ECM
metabolic imbalance and inflammatory responses. Recent evidence
from NP cells further confirms that Kla is dynamically regulated
under hypoxic and high-lactate conditions, highlighting its
critical role in intervertebral disc degeneration (76). Additionally, m6A RNA
modification intersects with various metabolic pathways (including
ferroptosis and oxidative stress): ALKBH5 and FTO can alter the
stability or translation of key gene mRNAs, thereby affecting the
cellular stress response and degenerative progression (18).
Taken together, IDD is no longer viewed merely as a
structural degradation resulting from mechanical loading and
inflammation, but rather as a disease driven by the bidirectional
regulatory network involving metabolic intermediates and pathways,
including those associated with NAD+ metabolism,
α-ketoglutarate-dependent reactions, and lactate-mediated
signaling, as well as epigenetic modifications (including
DNA/histone methylation, m6A, and Kla) (76). A deeper understanding of these
‘metabolism-epigenetics’ interactions will aid in identifying novel
molecular targets and strategies for early intervention or even
potential reversal of disc degeneration (77).
Potential clinical translational
applications
Drug and epigenetic intervention
strategies
Targeting epigenetic modifications offers novel
therapeutic approaches for IDD. DNA methylation inhibitors, such as
5-AZA, have been shown to restore the expression of key genes
involved in disc homeostasis by reversing hypermethylation-induced
silencing (12). Similarly,
histone modification regulators, such as EZH2 inhibitors, have
demonstrated the ability to attenuate inflammation and ECM
degradation, offering another promising avenue for treatment
(6,27).
RNA modifications, particularly m6A
methylation, serve a crucial role in IDD progression by influencing
mRNA stability and translation (78). The modulation of key m6A
enzymes, such as METTL3, FTO and ALKBH5, may provide an effective
means to regulate NP cell apoptosis and ECM homeostasis (18). RNA-based therapies, including small
interfering RNA and antisense oligonucleotides, could offer precise
interventions to correct abnormal RNA modifications and restore
gene function (79).
Combined biological therapies
Epigenetic regulation can be combined with
biological therapies, such as stem cell transplantation and gene
editing, to enhance IDD treatment outcomes. For instance,
mesenchymal stem cell (MSC) transplantation has been explored as a
regenerative therapy for IDD, but its efficacy is often limited by
the harsh degenerative microenvironment (80,81).
Preconditioning stem cells with epigenetic modulators, such as
DNMT3A or EZH2 inhibitors, could improve their survival and
therapeutic potential in IDD treatment (6,27).
Another promising approach involves integrating
epigenetic regulation with nanotechnology-based drug delivery
systems. Nanocarriers, such as liposomes and biodegradable
polymers, could facilitate the targeted delivery of epigenetic
drugs, reducing systemic side effects while enhancing local
efficacy (82). Additionally,
gene-editing technologies, such as clustered regularly interspaced
short palindromic repeats (CRISPR)/CRISPR-associated protein 9
(Cas9), offer the potential to directly modify epigenetic
regulatory genes, providing a long-term therapeutic solution for
IDD (83).
Beyond these approaches, bioengineered nanomaterials
have been applied to co-deliver MSCs and epigenetic modulators for
disc regeneration. Novel nanofibrous spongy microspheres (NF-SMS)
enhanced MSC seeding, proliferation and differentiation compared
with conventional microcarriers. A hyperbranched polymer (HP) with
strong miRNA-binding affinity was used to complex with
anti-miR-199a, forming ‘double shell’ polyplexes with high
transfection efficiency. Encapsulation of these polyplexes within
biodegradable nanospheres (NS) enabled sustained release of
anti-miR-199a. The composite system
(MSC/HP-anti-miR-199a/NS/NF-SMS) promoted NP-like phenotypes,
resisted calcification in vitro and in vivo, and, in
a rabbit lumbar degeneration model, preserved disc height,
maintained ECM function and prevented IDD calcification (84).
Personalized and precision
medicine
Advances in epigenomic profiling have paved the way
for personalized IDD treatment strategies. By analyzing epigenetic
signatures, such as DNA methylation patterns and histone
modification profiles, clinicians can stratify patients based on
their molecular characteristics and tailor treatment plans
accordingly (13,27,85).
For example, patients with elevated EZH2 expression and suppressed
SOX9 levels may benefit from EZH2 inhibitors in combination with
MSC therapy, whereas those with increased DNMT3b activity might
respond better to DNA methylation inhibitors (18).
Dynamic monitoring of epigenetic modifications
during treatment could enable real-time adjustments to therapeutic
strategies. This ‘precise-dynamic-reversible’ approach refers to
the concept of precisely targeting disease-related epigenetic
modifications, dynamically monitoring their changes during
treatment and reversibly modulating these marks in response to
therapeutic needs. In the context of IDD, this strategy emphasizes
individualized, adaptable interventions that can fine-tune gene
expression and cellular responses in real time, potentially
improving both safety and efficacy in clinical management (15,49).
To synthesize the aforementioned translational
insights, Table I (6,81,86–88)
summarizes the major candidate epigenetic interventions for IDD.
Table I compiles current
strategies, their targets, experimental systems, therapeutic
outcomes, adverse effects and delivery options, providing a
comprehensive overview of the preclinical landscape and guiding
future clinical development.
 | Table I.Candidate epigenetic interventions
for IDD. |
Table I.
Candidate epigenetic interventions
for IDD.
| Drug/strategy | Primary target | Model system (in
vitro/in vivo) | Outcomes | Reported adverse
effects | Delivery
options |
|---|
| 5-AZA | DNMTs (DNA
methylation) | NP cells; rat IDD
models | Restores SLC40A1
expression, reduces ferroptosis, attenuates ECM degradation | Cytotoxicity at
high doses | Intraperitoneal
injection; potential local hydrogel-based delivery |
| EZH2 inhibitors
(such as GSK126 and EPZ6438) | EZH2
(H3K27me3) | Human NP cells;
cartilage endplate degeneration models | Increases SOX9
expression, improves ECM integrity, reduces inflammatory
signaling | Off-target effects;
altered immune responses | Systemic
administration; nanoparticle delivery explored |
| HDAC inhibitors
(such as trichostatin A and SAHA/vorinostat) | Class I/II
HDACs | Human NP cells; rat
puncture IDD model | Suppresses
apoptosis, restores ECM markers, reduces inflammation | Broad inhibition
leads to risk of toxicity | Systemic injection;
possible local delivery |
| SIRT1/SIRT6
activators (such as resveratrol and NAD+ boosters) | SIRT1, SIRT6
(deacetylases) | Human NP cells;
rabbit IDD models | Enhances autophagy,
reduces oxidative stress, preserves disc structure | Generally low
toxicity; bioavailability issues | Oral; intradiscal
injection; NAD+ precursor supplementation |
| siRNA/antisense
oligonucleotides | miRNAs, lncRNAs,
circRNAs (such as miR-143, LINC01121 and circ_0001658) | Human NP cells;
mouse/rat IDD models | Suppresses
apoptosis, inflammation or ECM catabolism depending on target | Off-target
silencing; immune activation | Local hydrogel,
liposome or viral vector delivery |
| CRISPR/dCas9-based
epigenetic editing | Gene-specific
methylation/acetylation marks | Proof-of-concept in
NP cells (pre-clinical) | Precise modulation
of target gene expression (e.g., SOX9 and COL2A1) | Off-target genome
editing; safety not established | Viral vectors;
nanoparticle systems (under development) |
Translational bottlenecks
Despite encouraging preclinical findings, several
translational bottlenecks hinder the clinical application of
epigenetic therapies for IDD. First, off-target effects remain a
major concern. Epigenetic modulators such as DNA methylation
inhibitors or HDAC inhibitors often act broadly rather than in a
locus-specific manner, potentially leading to dysregulation of
unrelated genes and adverse biological outcomes in non-disc tissues
(89). Second, immune responses
induced by RNA-based therapies or CRISPR/deactivated Cas9
epigenetic editors can activate innate immunity or exacerbate
inflammation, which is particularly problematic in the already
inflamed disc microenvironment (90,91).
Third, long-term safety is insufficiently understood. While
epigenetic modifications are theoretically reversible, persistent
or cumulative alterations may increase the risk of tumorigenesis,
ectopic tissue remodeling or interference with normal aging
pathways (92). Finally, the lack
of standardized outcome measures complicates translational
progress. Preclinical studies employ heterogeneous endpoints, from
disc height index and MRI signals to histological and molecular
markers, making cross-study comparisons difficult and limiting
their predictive value for clinical outcomes such as pain relief
and functional improvement (93).
Overcoming these challenges will require the
development of precision-targeted delivery systems, immune-evasive
biomaterials, long-term safety monitoring protocols and
consensus-driven standardized evaluation criteria to guide both
preclinical and clinical research (94).
Conclusion
Epigenetic modifications serve a crucial role in
maintaining intervertebral disc homeostasis, regulating NP cell
fate and modulating local inflammatory responses. Targeting DNA
methylation, histone modifications and RNA methylation at the
molecular level has been demonstrated to have the potential to
mitigate IDD progression, offering novel therapeutic strategies
(6,95).
Further research is needed to fully understand the
mechanisms underlying epigenetic regulation in IDD and to identify
optimal intervention timing and dosages. Multidisciplinary
collaboration will be essential to translate these findings into
clinical applications, ensuring that epigenetic therapies are both
effective and safe.
Epigenetic therapies should be integrated with
existing treatments, including stem cell transplantation, gene
editing, biomaterial scaffolds and physical therapy, to enhance
overall treatment efficacy. With continuous advancements in
high-throughput sequencing, gene editing and regenerative medicine,
personalized epigenetic therapies hold great promise for the future
of IDD treatment, potentially offering long-term relief for
patients suffering from chronic low back pain.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
Not applicable.
Authors' contributions
YH and LL drafted the manuscript and prepared the
figures and tables. YG conducted an in-depth literature review,
contributed to the conception and design of the review structure,
and participated substantially in manuscript revision, including
critical evaluation of the scientific content, organization of
updated references, and language refinement after peer-review. JS
conceived and supervised the study, provided critical revisions for
important intellectual content, and approved the final version of
the manuscript. Data authentication is not applicable. All authors
read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Diwan AD and Melrose J: Intervertebral
disc degeneration and how it leads to low back pain. JOR Spine.
6:e12312022. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Teraguchi M, Yoshimura N, Hashizume H,
Yamada H, Oka H, Minamide A, Nagata K, Ishimoto Y, Kagotani R,
Kawaguchi H, et al: Progression, incidence, and risk factors for
intervertebral disc degeneration in a longitudinal population-based
cohort: The Wakayama Spine Study. Osteoarthritis Cartilage.
25:1122–1131. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hassett G, Hart DJ, Manek NJ, Doyle DV and
Spector TD: Risk factors for progression of lumbar spine disc
degeneration: The Chingford Study. Arthritis Rheum. 48:3112–3117.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Li G, Zhang W, Liang H and Yang C:
Epigenetic regulation in intervertebral disc degeneration. Trends
Mol Med. 28:803–805. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Clark J and Rager JE: Chapter
1-Epigenetics: An overview of CpG methylation, chromatin
remodeling, and regulatory/noncoding RNAs. Fry RC: Environmental
Epigenetics in Toxicology and Public Health. Academic Press; pp.
3–32. 2020, View Article : Google Scholar
|
|
6
|
Kang L, Zhang H, Jia C, Zhang R and Shen
C: Epigenetic modifications of inflammation in intervertebral disc
degeneration. Ageing Res Rev. 87:1019022023. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Razmara E, Bitaraf A, Yousefi H, Nguyen
TH, Garshasbi M, Cho WC and Babashah S: Non-coding RNAs in
cartilage development: An updated review. Int J Mol Sci.
20:44752019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kiełbowski K, Herian M, Bakinowska E,
Banach B, Sroczyński T and Pawlik A: The role of genetics and
epigenetic regulation in the pathogenesis of osteoarthritis. Int J
Mol Sci. 24:116552023. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ma Z, Liu X, Zhang X, Li S, An J and Luo
Z: Research progress on long non-coding RNAs in non-infectious
spinal diseases (Review). Mol Med Rep. 30:1642024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ikuno A, Akeda K, Takebayashi SI, Shimaoka
M, Okumura K and Sudo A: Genome-wide analysis of DNA methylation
profile identifies differentially methylated loci associated with
human intervertebral disc degeneration. PLoS One. 14:e02221882019.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hong JY, Kim H, Jeon WJ, Lee J, Yeo C, Lee
YJ and Ha IH: Epigenetic changes within the annulus fibrosus by DNA
methylation in rat intervertebral disc degeneration model. Cells.
11:35472022. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen J, Yang X, Li Q, Ma J, Li H, Wang L,
Chen Z and Quan Z: Inhibiting DNA methyltransferase DNMT3B confers
protection against ferroptosis in nucleus pulposus and ameliorates
intervertebral disc degeneration via upregulating SLC40A1. Free
Radic Biol Med. 220:139–153. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tajerian M, Alvarado S, Millecamps M,
Dashwood T, Anderson KM, Haglund L, Ouellet J, Szyf M and Stone LS:
DNA methylation of SPARC and chronic low back pain. Mol Pain.
7:652011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Reynard LN, Bui C, Syddall CM and Loughlin
J: CpG methylation regulates allelic expression of GDF5 by
modulating binding of SP1 and SP3 repressor proteins to the
osteoarthritis susceptibility SNP rs143383. Hum Genet.
133:1059–1073. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Esteras N, Blacker TS, Zherebtsov EA,
Stelmashuk OA, Zhang Y, Wigley WC, Duchen MR, Dinkova-Kostova AT
and Abramov AY: Nrf2 regulates glucose uptake and metabolism in
neurons and astrocytes. Redox Biol. 62:1026722023. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhao K, Zhang Y, Kang L, Song Y, Wang K,
Li S, Wu X, Hua W, Shao Z, Yang S and Yang C: Epigenetic silencing
of miRNA-143 regulates apoptosis by targeting BCL2 in human
intervertebral disc degeneration. Gene. 628:259–266. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhao K, Zhang Y, Kang L, Song Y, Wang K,
Li S, Wu X, Hua W, Shao Z, Yang S and Yang C: Methylation of
microRNA-129-5P modulates nucleus pulposus cell autophagy by
targeting Beclin-1 in intervertebral disc degeneration. Oncotarget.
8:86264–86276. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li G, Luo R, Zhang W, He S, Wang B, Liang
H, Song Y, Ke W, Shi Y, Feng X, et al: m6A hypomethylation of
DNMT3B regulated by ALKBH5 promotes intervertebral disc
degeneration via E4F1 deficiency. Clin Transl Med. 12:e7652022.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Xu W, Zhang X, Liu G, Zhu M, Wu Y, Jie Z,
Xie Z, Wang S, Ma Q, Fan S and Fang X: Oxidative stress abrogates
the degradation of KMT2D to promote degeneration in nucleus
pulposus. Biochim Biophys Acta Mol Basis Dis. 1866:1658882020.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zheng J, Ma Z, Liu P, Wei J, Min S, Shan
Y, Zhang J, Li Y, Xue L, Tan Z and Wang D: EZH2 inhibits
senescence-associated inflammation and attenuates intervertebral
disc degeneration by regulating the cGAS/STING pathway via
H3K27me3. Osteoarthritis Cartilage. 33:548–559. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yao Q, Lei Y, Zhang Y, Chen H, Dong X, Ye
Z and Liang H: EZH2-H3K27me3-mediated epigenetic silencing of DKK1
induces nucleus pulposus cell pyroptosis in intervertebral disc
degeneration by activating NLRP3 and NAIP/NLRC4. Inflammation.
48:902–918. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Lei M, Lin H, Shi D, Hong P, Song H,
Herman B, Liao Z and Yang C: Molecular mechanism and therapeutic
potential of HDAC9 in intervertebral disc degeneration. Cell Mol
Biol Lett. 28:1042023. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kouzarides T: Chromatin modifications and
their function. Cell. 128:693–705. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Xiao L, Gong D, Liang L, Liang A, Liang H,
Xu X and Teng H: Inhibition of HDAC4 by GSK3β leads to
downregulation of KLF5 and ASK1 and prevents the progression of
intravertebral disc degeneration. Clin Epigenetics. 13:532021.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Bennett RL: The epigenetic underpinnings
of lower back pain. Clin Transl Med. 12:e8682022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhu D, Liang H, Tong B, Du Z, Li G, Zhang
W, Wu D, Zhou X, Lei J, Zhang X, et al: DDX1 methylation mediated
MATR3 splicing regulates intervertebral disc degeneration by
initiating chromatin reprogramming. Nat Commun. 16:61532025.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Jiang C, Guo Q, Jin Y, Xu JJ, Sun ZM, Zhu
DC, Lin JH, Tian NF, Sun LJ, Zhang XL and Wu YS: Inhibition of EZH2
ameliorates cartilage endplate degeneration and attenuates the
progression of intervertebral disc degeneration via demethylation
of Sox-9. EBioMedicine. 48:619–629. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zhou M, He SJ, Liu W, Yang MJ, Hou ZY,
Meng Q and Qian ZL: EZH2 upregulates the expression of MAPK1 to
promote intervertebral disc degeneration via suppression of
miR-129-5p. J Gene Med. 24:e33952022. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Liang H, Luo R, Li G, Zhang W, Zhu D, Wu
D, Zhou X, Tong B, Wang B, Feng X, et al: Lysine methylation of
PPP1CA by the methyltransferase SUV39H2 disrupts TFEB-dependent
autophagy and promotes intervertebral disc degeneration. Cell Death
Differ. 30:2135–2150. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Wu O, Jin Y, Zhang Z, Zhou H, Xu W, Chen
L, Jones M, Kwan KYH, Gao J, Zhang K, et al: KMT2A regulates the
autophagy-GATA4 axis through METTL3-mediated m6A
modification of ATG4a to promote NPCs senescence and IVDD
progression. Bone Res. 12:672024. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kooistra SM and Helin K: Molecular
mechanisms and potential functions of histone demethylases. Nat Rev
Mol Cell Biol. 13:297–311. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Diaz-Hernandez ME, Murakami K, Murata S,
Khan NM, Shenoy SPV, Henke K, Yamada H and Drissi H: Inhibition of
KDM2/7 promotes notochordal differentiation of hiPSCs. Cells.
13:14822024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang Y, Wang Y, Zhou X, Wang J, Shi M,
Wang J, Li F and Chen Q: Osmolarity controls the differentiation of
adipose-derived stem cells into nucleus pulposus cells via histone
demethylase KDM4B. Mol Cell Biochem. 472:157–171. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhou X, Li J, Teng J, Liu Y, Zhang D, Liu
L and Zhang W: microRNA-155-3p attenuates intervertebral disc
degeneration via inhibition of KDM3A and HIF1α. Inflamm Res.
70:297–308. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li Z, Li X, Chen C, Li S, Shen J, Tse G,
Chan MTV and Wu WKK: Long non-coding RNAs in nucleus pulposus cell
function and intervertebral disc degeneration. Cell Prolif.
51:e124832018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Lytle JR, Yario TA and Steitz JA: Target
mRNAs are repressed as efficiently by microRNA-binding sites in the
5′ UTR as in the 3′ UTR. Proc Natl Acad Sci USA. 104:9667–9672.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Haas U, Sczakiel G and Laufer SD:
MicroRNA-mediated regulation of gene expression is affected by
disease-associated SNPs within the 3′-UTR via altered RNA
structure. RNA Biol. 9:924–937. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Wang C, Cui L, Gu Q, Guo S, Zhu B, Liu X,
Li Y, Liu X, Wang D and Li S: The mechanism and function of miRNA
in intervertebral disc degeneration. Orthop Surg. 14:463–471. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Yu Z, Fan C, Mao Y, Wu X and Mao H:
Autophagy activation alleviates annulus fibrosus degeneration via
the miR-2355-5p/mTOR pathway. J Orthop Surg Res. 20:862025.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Chen D, Jiang X and Zou H: hASCs-derived
exosomal miR-155-5p targeting TGFβR2 promotes autophagy and reduces
pyroptosis to alleviate intervertebral disc degeneration. J Orthop
Translat. 39:163–176. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Hu P, Feng B, Wang G, Ning B and Jia T:
Microarray based analysis of gene regulation by microRNA in
intervertebral disc degeneration. Mol Med Rep. 12:4925–4930. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Song J, Chen ZH, Zheng CJ, Song KH, Xu GY,
Xu S, Zou F, Ma XS, Wang HL and Jiang JY: Exosome-transported
circRNA_0000253 competitively adsorbs MicroRNA-141-5p and increases
IDD. Mol Ther Nucleic Acids. 21:1087–1099. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Ji ML, Jiang H, Zhang XJ, Shi PL, Li C, Wu
H, Wu XT, Wang YT, Wang C and Lu J: Preclinical development of a
microRNA-based therapy for intervertebral disc degeneration. Nat
Commun. 9:50512018. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Lin H, Zhang W, Zhou T, Li W, Chen Z, Ji
C, Zhang C and He F: Mechanism of microRNA-21 regulating IL-6
inflammatory response and cell autophagy in intervertebral disc
degeneration. Exp Ther Med. 14:1441–1444. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Gu SX, Li X, Hamilton JL, Chee A, Kc R,
Chen D, An HS, Kim JS, Oh CD, Ma YZ, et al: MicroRNA-146a reduces
IL-1 dependent inflammatory responses in the intervertebral disc.
Gene. 555:80–87. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Ge C and Li C: Expression and regulatory
role of miRNA-222 in intervertebral disc degeneration (IDD).
Biotechnol Biotechnol Equip. 33:1553–1559. 2019. View Article : Google Scholar
|
|
47
|
Lan T, Shiyu-Hu Shen Z, Yan B and Chen J:
New insights into the interplay between miRNAs and autophagy in the
aging of intervertebral discs. Ageing Res Rev. 65:1012272021.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Li Z, Sun Y, He M and Liu J:
Differentially-expressed mRNAs, microRNAs and long noncoding RNAs
in intervertebral disc degeneration identified by RNA-sequencing.
Bioengineered. 12:1026–1039. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Cazzanelli P and Wuertz-Kozak K: MicroRNAs
in intervertebral disc degeneration, apoptosis, inflammation, and
mechanobiology. Int J Mol Sci. 21:36012020. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Zhu J, Zhang X, Gao W, Hu H, Wang X and
Hao D: lncRNA/circRNA-miRNA-mRNA ceRNA network in lumbar
intervertebral disc degeneration. Mol Med Rep. 20:3160–3174.
2019.PubMed/NCBI
|
|
51
|
Li G, Ma L, He S, Luo R, Wang B, Zhang W,
Song Y, Liao Z, Ke W, Xiang Q, et al: WTAP-mediated m6A
modification of lncRNA NORAD promotes intervertebral disc
degeneration. Nat Commun. 13:14692022. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Chen HW, Zhou JW, Zhang GZ, Luo ZB, Li L
and Kang XW: Emerging role and therapeutic implication of mTOR
signalling in intervertebral disc degeneration. Cell Prolif.
56:e133382023. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Sun Z, Tang X, Wang H, Sun H, Chu P, Sun L
and Tian J: LncRNA H19 aggravates intervertebral disc degeneration
by promoting the autophagy and apoptosis of nucleus pulposus cells
through the miR-139/CXCR4/NF-κB axis. Stem Cells Dev. 30:736–748.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Che Z, Xueqin J and Zhang Z: LncRNA
OIP5-AS1 accelerates intervertebral disc degeneration by targeting
miR-25-3p. Bioengineered. 12:11201–11212. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Chen X, Li Z, Xu D and Li S: LINC01121
induced intervertebral disc degeneration via modulating
miR-150-5p/MMP16 axis. J Gene Med. 22:e32312020. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Chen Y, Wu Y, Chen R, Xu C and Chen Q:
LncRNA LINC00324 is upregulated in intervertebral disk degeneration
and upregulates FasL in nucleus pulposus cells. Mol Cell Biochem.
476:1995–2000. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Chen WK, Zhang HJ, Zou MX, Wang C, Yan YG,
Zhan XL, Li XL and Wang WJ: LncRNA HOTAIR influences cell
proliferation via miR-130b/PTEN/AKT axis in IDD. Cell Cycle.
21:323–339. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Luo Y, He Y, Wang Y, Xu Y and Yang L:
LncRNA HCG18 promotes inflammation and apoptosis in intervertebral
disc degeneration via the miR-495-3p/FSTL1 axis. Mol Cell Biochem.
479:171–181. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Mi D, Cai C, Zhou B, Liu X, Ma P, Shen S,
Lu W and Huang W: Long non-coding RNA FAF1 promotes intervertebral
disc degeneration by targeting the Erk signaling pathway. Mol Med
Rep. 17:3158–3163. 2018.PubMed/NCBI
|
|
60
|
Chen S, Zhuang Q, Li P, Zeng J, Peng Y,
Ding Z, Cao H, Zheng R and Wang W: The long non-coding RNA
KLF3-AS1/ miR-10a-3p/ZBTB20 axis improves the degenerative changes
in human nucleus pulposus cells. Cell Tissue Res. 393:97–109. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Qiao L, Peng SY, Zhou YP, Yin J, Xu JP,
Chen B, Zhang H, Zhu C and Yu XD: Long non-coding RNA RP11-81H3.2
suppresses apoptosis by targeting microRNA-1539/COL2A1 in human
nucleus pulposus cells. Exp Ther Med. 22:8842021. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Wang C, Chen R, Zhu X and Zhang X: Long
non-coding RNAs LINC00689 inhibits the apoptosis of human nucleus
pulposus cells via miR-3127-5p/ATG7 axis-mediated autophagy. Open
Med (Wars). 17:1821–1832. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Li Y, Zhou S, Hu X and Lu S: The
pathological mechanisms of circRNAs in mediating intervertebral
disc degeneration. Noncoding RNA Res. 8:633–640. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Xie G, Wu T, Ji G, Wu H, Lai Y, Wei B and
Huang W: Circular RNA and intervertebral disc degeneration:
Unravelling mechanisms and implications. Front Mol Biosci.
10:13020172023. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Chen Z, Song J, Xie L, Xu G, Zheng C, Xia
X, Lu F, Ma X, Zou F, Jiang J and Wang H: N6-methyladenosine
hypomethylation of circGPATCH2L regulates DNA damage and apoptosis
through TRIM28 in intervertebral disc degeneration. Cell Death
Differ. 30:1957–1972. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Du X, Chen S, Cui H, Huang Y, Wang J, Liu
H, Li Z, Liang C, Zheng Z and Wang H: Circular RNA hsa_circ_0083756
promotes intervertebral disc degeneration by sponging miR-558 and
regulating TREM1 expression. Cell Prolif. 55:e132052022. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Meng GD and Xu BS: Circular RNA
hsa_circ_0001658 inhibits intervertebral disc degeneration
development by regulating hsa-miR-181c-5p/FAS. Comput Math Methods
Med. 2021:78533352021. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Wang T, Yan X, Song D, Li Y, Li Z and Feng
D: CircEYA3 aggravates intervertebral disc degeneration through the
miR-196a-5p/EBF1 axis and NF-κB signaling. Commun Biol. 7:3902024.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Yan P, Sun C, Luan L, Han J, Qu Y, Zhou C
and Xu D: Hsa_circ_0134111 promotes intervertebral disc
degeneration via sponging miR-578. Cell Death Discov. 8:552022.
View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Cheng X, Zhang L, Zhang K, Zhang G, Hu Y,
Sun X, Zhao C, Li H, Li YM and Zhao J: Circular RNA VMA21 protects
against intervertebral disc degeneration through targeting miR-200c
and X linked inhibitor-of-apoptosis protein. Ann Rheum Dis.
77:770–779. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Zhu D, Liang H, Du Z, Liu Q, Li G, Zhang
W, Wu D, Zhou X, Song Y and Yang C: Altered metabolism and
inflammation driven by post-translational modifications in
intervertebral disc degeneration. Research (Wash D C).
7:03502024.PubMed/NCBI
|
|
72
|
Shi C, Wu H, Du D, Im HJ, Zhang Y, Hu B,
Chen H, Wang X, Liu Y, Cao P, et al: Nicotinamide
phosphoribosyltransferase inhibitor APO866 prevents IL-1β-induced
human nucleus pulposus cell degeneration via autophagy. Cell
Physiol Biochem. 49:2463–2482. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Sandalova E, Goh J, Lim ZX, Lim ZM,
Barardo D, Dorajoo R, Kennedy BK and Maier AB: Alpha-ketoglutarate
supplementation and BiologicaL agE in middle-aged adults
(ABLE)-intervention study protocol. Geroscience. 45:2897–2907.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Xu HW, Fang XY, Liu XW, Zhang SB, Yi YY,
Chang SJ, Chen H and Wang SJ: α-Ketoglutaric acid ameliorates
intervertebral disk degeneration by blocking the IL-6/JAK2/STAT3
pathway. Am J Physiol Cell Physiol. 325:C1119–C1130. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Zhang D, Tang Z, Huang H, Zhou G, Cui C,
Weng Y, Liu W, Kim S, Lee S, Perez-Neut M, et al: Metabolic
regulation of gene expression by histone lactylation. Nature.
574:575–580. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Sheng L, Xu H, Wang Y, Ni J, Xiang T, Xu
H, Zhou X, Wei K and Dai J: Systematic analysis of lysine
lactylation in nucleus pulposus cells. iScience. 27:1111572024.
View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Xu WN, Yang RZ, Zheng HL, Yu W, Zheng XF,
Li B, Jiang SD and Jiang LS: PGC-1α acts as an mediator of Sirtuin2
to protect annulus fibrosus from apoptosis induced by oxidative
stress through restraining mitophagy. Int J Biol Macromol.
136:1007–1017. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Lei Y, Zhan E, Chen C, Hu Y, Lv Z, He Q,
Wang X, Li X and Zhang F: ALKBH5-mediated m6A
demethylation of Runx2 mRNA promotes extracellular matrix
degradation and intervertebral disc degeneration. Cell Biosci.
14:792024. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Hofman CR and Corey DR: Targeting RNA with
synthetic oligonucleotides: Clinical success invites new
challenges. Cell Chem Biol. 31:125–138. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Zhang W, Wang D, Li H, Xu G, Zhang H, Xu C
and Li J: Mesenchymal stem cells can improve discogenic pain in
patients with intervertebral disc degeneration: A systematic review
and meta-analysis. Front Bioeng Biotechnol. 11:11553572023.
View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Zhao Y, He J, Qiu T, Zhang H, Liao L and
Su X: Epigenetic therapy targeting bone marrow mesenchymal stem
cells for age-related bone diseases. Stem Cell Res Ther.
13:2012022. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Li T, Chen Y and Li S: The advances in the
development of epigenetic modifications therapeutic drugs delivery
systems. Int J Nanomedicine. 19:10623–10637. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Fadul SM, Arshad A and Mehmood R:
CRISPR-based epigenome editing: Mechanisms and Applications.
Epigenomics. 15:1137–1155. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Feng G, Zhang Z, Dang M, Rambhia KJ and Ma
PX: Nanofibrous spongy microspheres to deliver rabbit mesenchymal
stem cells and anti-miR-199a to regenerate nucleus pulposus and
prevent calcification. Biomaterials. 256:1202132020. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Cheng P, Wei HZ, Chen HW, Wang ZQ, Mao P
and Zhang HH: DNMT3a-mediated methylation of PPARγ promote
intervertebral disc degeneration by regulating the NF-κB pathway. J
Cell Mol Med. 28:e180482024. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Yan X, Wang X, Che Y, Feng P and Zhang T:
Immunological mechanisms and immunoregulatory strategies in
intervertebral disc degeneration. arXiv. 2506.09336. 2025.
|
|
87
|
Milheiro C, Moura ML, Amendola M, Barbosa
MA and Caldeira J: Harnessing CRISPR potential for intervertebral
disc regeneration strategies. Front Bioeng Biotechnol.
13:15624122025. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Zhang Y, Wang J, Hua D, Fan C, He W, Deng
Y, Tang M, Geng D, Wu X and Mao H: Histone modifications: Unveiling
the epigenetic enigma of degenerative skeletal diseases. J Orthop
Translat. 55:245–266. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Roh EJ, Darai A, Kyung JW, Choi H, Kwon
SY, Bhujel B, Kim KT and Han I: Genetic therapy for intervertebral
disc degeneration. Int J Mol Sci. 22:15792021. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Keshavarz S, Alavi CE, Aghayan H,
Jafari-Shakib R and Vojoudi E: Advancements in degenerative disc
disease treatment: a regenerative medicine approach. Stem Cell Rev
Rep. 21:1252–1282. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Ma W, Wang W, Zhao L, Fan J, Liu L, Huang
L, Peng B, Wang J, Xu B, Liu H, et al: Reprogramming to restore
youthful epigenetics of senescent nucleus pulposus cells for
mitigating intervertebral disc degeneration and alleviating low
back pain. Bone Res. 13:352025. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Li X, Chang H, Cai F, Zhang Y, Li A, Yang
X, Cai Z, Cui W and Liu X: Progress of gene-functionalized
regenerative material repair intervertebral disc degeneration.
Small Sci. 4:23003552024. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Samanta A, Lufkin T and Kraus P:
Intervertebral disc degeneration-current therapeutic options and
challenges. Front Public Health. 11:11567492023. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Ma L, Pan J, Zhang J and Liu F: Innovative
strategies in combating intervertebral disc degeneration:
pathological mechanisms and biomaterial advancements. Front Bioeng
Biotechnol. 13:16432222025. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Zhu B, Chen HX, Li S, Tan JH, Xie Y, Zou
MX, Wang C, Xue JB, Li XL, Cao Y and Yan YG: Comprehensive analysis
of N6-methyladenosine (m6A) modification during the
degeneration of lumbar intervertebral disc in mice. J Orthop
Translat. 31:126–138. 2021. View Article : Google Scholar : PubMed/NCBI
|