Introduction
Infantile hemangioma (IH) is one of the most
prevalent benign vascular tumors in infants and young children
(1). It is comprised of benign
proliferating vascular endothelial cells, and the prevalence rate
of IH is 5–10% globally (2).
Preterm birth, low birth weight, multiple pregnancies, progesterone
use during pregnancy and a family history of the disease are
potential risk factors (1–3). IH is characterized by the onset of
bright red papules or nodules 2–4 weeks after birth. This rapid
growth period lasts for ~5 months, reaching a plateau until ~1 year
of age and finally entering a slow regression period; the vast
majority of IHs eventually break down into fibrous fat residues
between 5 and 10 years of age (4).
Although IH is benign and eventually fades, rapid tumor growth
during proliferation can occasionally lead to disfigurement and
life-threatening complications, including ulcers, bleeding,
permanent visual impairment, airway obstruction leading to
respiratory distress and congestive heart failure (5,6).
Therefore, early intervention in the rapid proliferative stage of
IH is key to minimizing or preventing the risk of developing
potential complications. The current clinical treatments for IH
include the use of β-blockers, corticosteroids, interferon,
vincristine, angiotensin-converting enzyme inhibitors, laser
therapy and surgical treatment (7,8).
However, since the pathogenesis of IH remains to be fully
elucidated, these treatments are only partially effective or can
result in severe adverse reactions (7).
MicroRNAs (miRNAs/miRs) are endogenous, small,
non-coding RNA molecules that can regulate gene expression by
directly binding with target mRNAs (9,10).
In our previous study, RNA sequencing screening was performed for
differentially expressed miRNAs related to IH, resulting in the
identification of 97 upregulated and 119 downregulated miRNAs
(11). In addition, the enrichment
and pathways of potential target genes were analyzed, and it was
found that the proteoglycans in cancer signaling pathway exhibited
a notable difference. Common hub genes and miRNAs identified as
being related to proteoglycan signaling pathways in angiogenesis
and cancer, including c-Myc, integrin β1 (ITGB1), Bcl2 and miR-29a,
were screened through an miRNA gene network and protein-protein
interaction analysis. The results of the RNA sequencing screening
suggested that c-Myc, ITGB1, Bcl2 and miR-29a may have served
important roles during the pathogenesis of IH (11).
Integrins are members of the cell surface adhesion
protein receptor family, consisting of 18 specific α subunits and
eight unique β subunits, which form 24 heterodimer structures in a
non-covalent binding manner (12).
Each heterodimer can bind to specific sequences of extracellular
matrix, cell surface or soluble protein ligands. Integrins are
involved in a number of biological processes, such as cell
polarity, cell surface adhesion, proliferation, apoptosis, tumor
formation and metastasis, and are key proteins in the connection
between cells and the surrounding environment (13). ITGB1 has been shown to serve a
carcinogenic role in a variety of types of cancer. The
Akt/Wnt/β-catenin pathway is a key downstream regulator of ITGB1
carcinogenesis in a variety of tumors (14,15),
while c-Myc is a downstream target of the Wnt/β-catenin pathway
(16). c-Myc has also been shown
to be carcinogenic in various types of cancer (17,18).
A previous study revealed the association between the
ITGB1/Wnt/β-catenin/c-Myc axis and ITGB1-divergent transcript
(ITGB1-DT), a long non-coding RNA located on chromosome 10p11.22,
for which the direction of transcription is opposite to that of the
coding gene ITGB1; ITGB1-DT interacts with histone-lysine
N-methyltransferase EZH2 (EZH2), inhibits the binding of EZH2 to
the ITGB1 promoter and reduces the trimethylation level of lysine
27 on histone H3 in the ITGB1 promoter region, resulting in the
activation of ITGB1 expression (19). At the same time, the upregulation
of ITGB1 activates the Wnt/β-catenin/c-Myc pathway, which in turn
directly transcriptionally activates ITGB1-DT expression (6,19).
It has therefore been suggested that ITGB1 can transcriptionally
upregulate c-Myc mRNA expression through the Wnt/β-catenin pathway.
In addition, mucin 1 (MUC1), a transmembrane glycoprotein involved
in cell regeneration, differentiation, adhesion, integration,
signaling and apoptosis, has been reported to activate and regulate
the Wnt/β-catenin pathway (20).
Similar to phosphorylation, O-linked
β-N-acetylglucosaminylation (O-GlcNAcylation) is a broad and
dynamic protein modification regulated by β-N-acetylglucosaminidase
(OGA) and O-linked N-acetylglucosamine transferase 110 kDa subunit
(OGT) and is involved in gene transcription, signal transduction,
proteolytic hydrolysis and other cell activities (21). In addition, a variety of tumor
tissues exhibit high O-GlcNAcylation levels, and O-GlcNAcylation
has been reported to effect cancer cell proliferation, angiogenesis
and metastasis in various cancer types (22,23).
It has also been reported that O-GlcNAcylation can promote the
stability of the c-Myc protein by interfering with ubiquitination
and proteasome-mediated c-Myc degradation (24,25).
It has been reported that the high expression of
c-Myc in IH promotes the proliferation of hemangioma endothelial
cells (11). As such, it is
possible that miR-29a may affect angiogenesis of hemangioma
endothelial cells by regulating the c-Myc pathway. Therefore, the
present study explored the interactions between miR-29a, the c-Myc
pathway and the pathogenesis of angiogenesis using small
interfering RNA (siRNA) interference, reverse
transcription-quantitative PCR (RT-qPCR) and western blot (WB)
analysis with the aim of providing a basis for novel strategies
targeting IH.
Materials and methods
Cells and transfections
Human umbilical vein endothelial cells (HUVECs)
represent a commonly-used in vitro cell model for IH
(26). HUVECs were confirmed to be
mycoplasma free by using a mycoplasma analysis kit (Wuhan
Servicebio Technology Co., Ltd.). As such, HUVECs were purchased
from the American Type Culture Collection (cat. no. CRL-1730) and
cultured in EGM™ Medium (Lonza Group, Ltd.) containing 10% FBS
(Gibco; Thermo Fisher Scientific, Inc.) with 1%
penicillin-streptomycin solution (Beyotime Biotechnology) in a
humidified incubator at 37°C with 5% CO2. HUVECs were
confirmed to be mycoplasma free. Cells were used in the
transfection experiments upon reaching 80% confluency (passage 6).
HUVECs were then transfected with 20 nM of the relevant siRNA. The
siRNA sequences used were as follows: ITGB1 siRNA1, sense
5′-GCCUUGCAUUACUGCUGAUAU-3′, antisense 5′-AUAUCAGCAGUAAUGCAAGGC-3;
ITGB1 siRNA2, sense 5′-GCCCUCCAGAUGACAUAGAAA-3′, antisense
5′-UUUCUAUGUCAUCUGGAGGGC-3′; MUC1 siRNA1, sense
5′-GACACAGUUCAAUCAGUAUAA-3′, antisense 5′-UUAUACUGAUUGAACUGUGUC-3′;
MUC1 siRNA2, sense 5′-CCGGGAUACCUACCAUCCUAU-3′, antisense
5′-AUAGGAUGGUAGGUAUCCCGG-3′; OGT siRNA1, sense
5′-UUUAGCACUCUGGCAAUUAAA-3′, antisense 5′-UUUAAUUGCCAGAGUGCUAAA-3;
OGT siRNA2, sense 5′-GCUGAGCAGUAUUCCGAGAAA-3′, antisense
5′-UUUCUCGGAAUACUGCUCAGC-3′; and siRNA negative control, sense
5′-AUCUUUGAUAUCGCGUCUACG-3, anti-sense 5′-CGUAGACGCGAUAUCAAAGAU-3′.
Cells were transfected for 6 h at 37°C using
Lipofectamine® 2000 reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) and were then cultured for a further 48 h before
subsequent experiments were performed. All siRNAs and the negative
control were provided by Guangzhou RiboBio Co., Ltd.
In addition, HUVECs were also transfected with 20
µg/ml miR-29a mimics, inhibitors or negative controls as
appropriate. The sequences were manufactured by Guangzhou RiboBio
Co., Ltd. and were as follows: miR-29a mimic, sense
5′-UAGCACCAUCUGAAAUCGGUUA-3′, antisense
5′-ACCGAUUUCAGAUGGUGCUAUU-3′; miR-29a mimic negative control, sense
5′-UUCUCCGAACGUGUCACGUUU-3′, antisense 5′-AAACGUGACACGUUCGGAGAA-3′;
miR-29a inhibitor 5′-UAACCGAUUUCAGAUGGUGCUA-3′; and miR-29a
inhibitor negative control, 5′-CAGUACUUUUGUGUAGUACAA-3′. HUVECs
were transfected with the aforementioned miR-29a mimic and
inhibitor sequences for 6 h using Lipofectamine 2000®
and then cultured for a further 48 h before subsequent
experimentation.
Cell viability analysis
A Cell Counting Kit-8 (CCK-8; Nanjing KeyGen Biotech
Co., Ltd.) assay was used to assess cell viability. Cells were
seeded into a 96-well plate and subjected to the aforementioned
transfection protocol; subsequently, transfected HUVECs were
incubated with CCK-8 reagent for 2 h at 37°C. The absorbance of
samples was measured at 450 nm using a microplate reader (Thermo
Fisher Scientific, Inc.).
RT-qPCR
RNA was extracted from the HUVECs using Trizol
(Ambion; Thermo Fisher Scientific, Inc.). Primarily, RNA was
reverse transcribed into cDNA using HiScript Reverse Transcriptase
(Vazyme Biotech Co., Ltd.) according to the manufacturer's
protocol. The mRNA expression level was then determined using SYBR
qPCR Master Mix (Vazyme Biotech Co., Ltd.) alongside specific
primers (Table I) on an eQ9600 PCR
System (Eastwin Life Sciences; Beijing Dongsheng Innovation
Biotechnology Co., Ltd.). The thermocycling conditions were as
follows: Pre-incubation at 95°C for 30 sec; amplification at 95°C
for 10 sec and 60°C for 30 sec, for 40 cycles. The relative mRNA
levels were quantified using the 2−ΔΔCq method with
GAPDH as the housekeeping gene (27).
 | Table I.Primer sequences for reverse
transcription-quantitative PCR. |
Table I.
Primer sequences for reverse
transcription-quantitative PCR.
| Gene | Forward, 5′-3′ | Reverse, 5′-3′ |
|---|
| ITGB1 |
TCCAACCTGATCCTGTGTCC |
CAATTCCAGCAACCACACCA |
| ITGB1-DT |
CAAAACCTGAAGCCCCAAAGA |
CACTGCACCGTCTTCCTAATG |
| c-Myc |
AACACACAACGTCTTGGAGC |
GCACAAGAGTTCCGTAGCTG |
| EZH2 |
CGGCTTCCCAATAACAGTAGC |
ACTCCACTCCACATTCTCAGG |
| MUC1 |
GTGATGTGCCATTTCCTTTCTCT |
TCGCTCATAGGATGGTAGGTATC |
| OGT |
AGGTGTTCTGTTATGCCCTGA |
AGCGCCCTTAGTATAGCCATT |
| OGA |
CAGCCTGATGAAGAACCCATG |
TGTTCATCAGTTTGCATGGGG |
| GAPDH |
TCAAGAAGGTGGTGAAGCAGG |
TCAAAGGTGGAGGAGTGGGT |
| U6 |
CGCTTCGGCAGCACATATAC |
AAATATGGAACGCTTCACGA |
|
hsa-miR-29a-3pa |
TGCGCTAGCACCATCTGAAATCGG |
CCAGTGCAGGGTCCGAGGTATT |
WB
The total protein content of HUVECs was extracted
using RIPA lysate (cat. no. P0013C; Beyotime Biotechnology) and
electrophoresed on 10% SDS-PAGE (10 µg/lane protein), before
proteins were transferred onto PVDF membranes (MilliporeSigma;
Merck KGaA). The protein concentration was measured with the BCA
method (cat. no. P0010; Beyotime Biotechnology). The membranes were
blocked with 5% skimmed milk for 1 h at room-temperature and
incubated with the following primary antibodies overnight at 4°C:
ITGB1 (cat. no. 12594-1-AP; Proteintech Group, Inc.), T-cell factor
1 (TCF1; cat. no. 14464-1-AP; Proteintech Group, Inc.), EZH2 (cat.
no. ab307646; Abcam), c-Myc (cat. no. 67447-1-Ig; Proteintech
Group, Inc.), phosphorylated (p-)GSK3β (cat. no. AF2016; Affinity
Biosciences), GSK3β (cat. no. AF5016; Affinity Biosciences),
p-β-catenin (cat. no. DF2989; Affinity Biosciences), β-catenin
(cat. no. 17565-1-AP; Proteintech Group, Inc.), β-actin (cat. no.
66009-1-Ig; Proteintech Group, Inc.), OGT (cat. no. ab177941;
Abcam) and OGA (cat. no. DF8436; Affinity Biosciences). All primary
antibodies were used at a dilution of 1:1,000. Subsequently, the
membranes were incubated with HRP-labeled goat anti-rabbit
secondary antibody (1:5,000; cat. no. ab6721; Abcam) or goat
anti-mouse secondary antibody (1:5,000; cat. no. ab6789; Abcam) for
1 h at room temperature. Finally, the membrane signals were
detected using an ECL substrate kit (Thermo Fisher Scientific,
Inc.) and quantified using ImageJ software 1.8.0 (National
Institutes of Health).
Dual-luciferase reporter assay
The bioinformatics tool TargetScan (https://www.targetscan.org/vert_80/) was used to
predict the potential association between miR-29a and ITGB1. The
ITBG1 3′-untranslated region sequences containing the wild-type or
mutant miR-29a binding sequence were inserted into the pGL6
luciferase reporter (Beyotime Biotechnology). HUVECs were then
co-transfected with the luciferase reporter and the miR-29a mimic
or mimic negative control using Lipofectamine® 2000
(Invitrogen; Thermo Fisher Scientific, Inc.). After 48 h
transfection at 37°C, the cells were harvested for luciferase
activity detection according to the instruction of the
Dual-Luciferase® Reporter Assay kit used (Promega
Corporation). Renilla luciferase activity was used to
normalize reporter activity.
Statistical analysis
All experiments were performed three times (n=3).
The data are presented as mean ± SD and analyzed using GraphPad
Prism software (version 7; Dotmatics). One-way ANOVA with Tukey's
post hoc test was used to analyze the results when comparing >2
groups in the present study, and a value of P<0.05 was
considered to indicate a statistically significant difference.
Results
Inhibition of ITGB1 downregulates the
β-catenin/c-Myc pathway
The present study first explored the effects of
ITGB1 on the c-Myc pathway in HUVECs using ITGB1 siRNAs. As
indicated by the results of RT-qPCR, both ITGB1 siRNA1 and siRNA2
significantly inhibited the gene expression of ITGB1 in HUVECs
compared with control HUVECs and the control siRNA (Fig. 1A). In addition, both ITGB1 siRNA1
and siRNA2 significantly inhibited the gene expression levels of
ITGB1-DT, EZH2 and c-Myc in HUVECs (Fig. 1B-D). Consistently, the results of
WB analysis demonstrated that both ITGB1 siRNA1 and siRNA2
significantly decreased ITGB1, EZH2, c-Myc, p-GSK3β and p-β-catenin
protein expression levels in HUVECs compared with the controls
(Fig. 1E and F). The CCK-8 assay
also revealed that a significant decrease in cell viability was
observed in HUVECs transfected with ITGB1 siRNA sequences compared
with both controls (Fig. 1G). On
the whole, these data suggest that inhibition of ITGB1 is able to
suppress the β-catenin/c-Myc pathway in HUVECs.
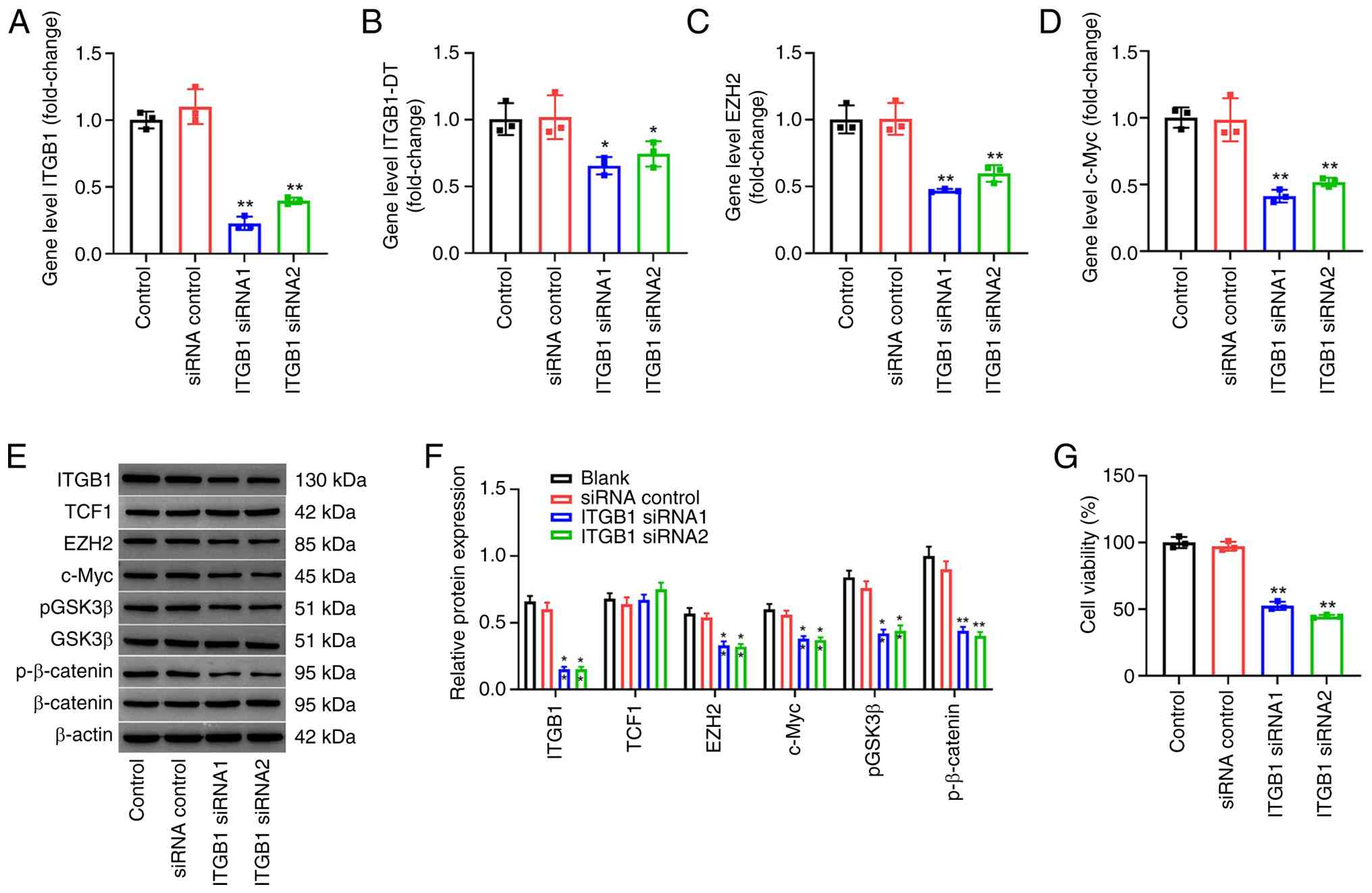 | Figure 1.Inhibition of ITGB1 downregulates the
β-catenin/c-Myc pathway in HUVECs. (A) ITGB1 gene expression after
HUVECs were transfected with 20 nM ITGB1 siRNA1, siRNA2 or siRNA
negative control for 6 h, and cultured for another 48 h. The gene
expressions of (B) ITGB1-DT, (C) EZH2 and (D) c-Myc were assessed
using reverse transcription-quantitative PCR following transfection
with ITGB1 siRNA. (E) The protein expression levels of ITGB1, TCF1,
EZH2, c-Myc, p-GSK3β, GSK3β, p-β-catenin and β-catenin were
evaluated using western blotting following transfection with ITGB1
siRNA. (F) Semi-quantification of western blotting results. p-GSK3β
and p-β-catenin were normalized to the levels of total GSK3β or
β-catenin, respectively. (G) Cell viability was analyzed using a
Cell Counting Kit-8 assay. n=3; *P<0.05 and **P<0.01 compared
with siRNA control group. HUVECs, human umbilical vein endothelial
cells; ITGB1, integrin β1; ITGB1-DT, integrin subunit β1 divergent
transcript; EZH2, histone-lysine N-methyltransferase EZH2; siRNA,
small interfering RNA; TCF1, T-cell factor 1; p-,
phosphorylated. |
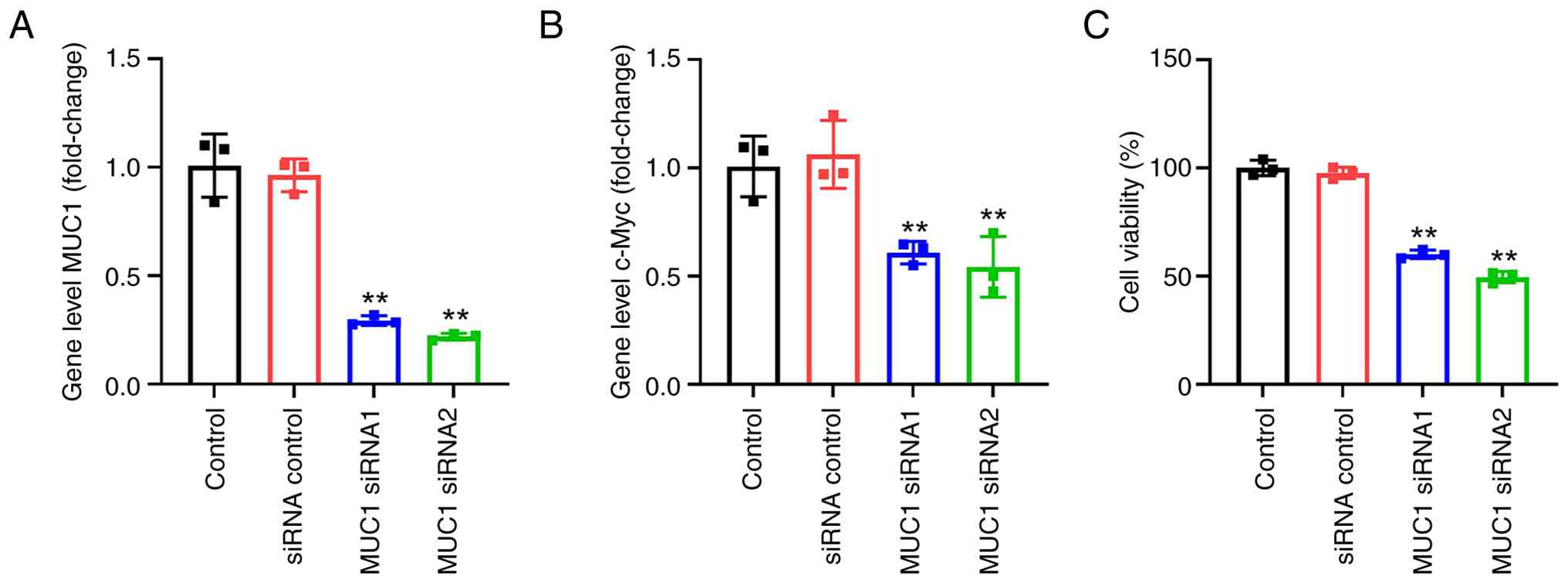
Inhibition of MUC1 suppresses the
c-Myc pathway in HUVECs
The function of MUC1 was subsequently explored using
RT-qPCR and CCK-8 assays. The results indicated that the inhibition
of MUC1 using MUC1-targeting siRNA significantly downregulated
c-Myc gene expression in HUVECs (Fig.
2A and B). In addition, significantly decreased cell viability
was observed in the HUVECs transfected with both MUC1 siRNA
sequences compared with the controls (Fig. 2C). These results suggest that the
inhibition of MUC1 inhibits c-Myc expression and the viability of
HUVECs.
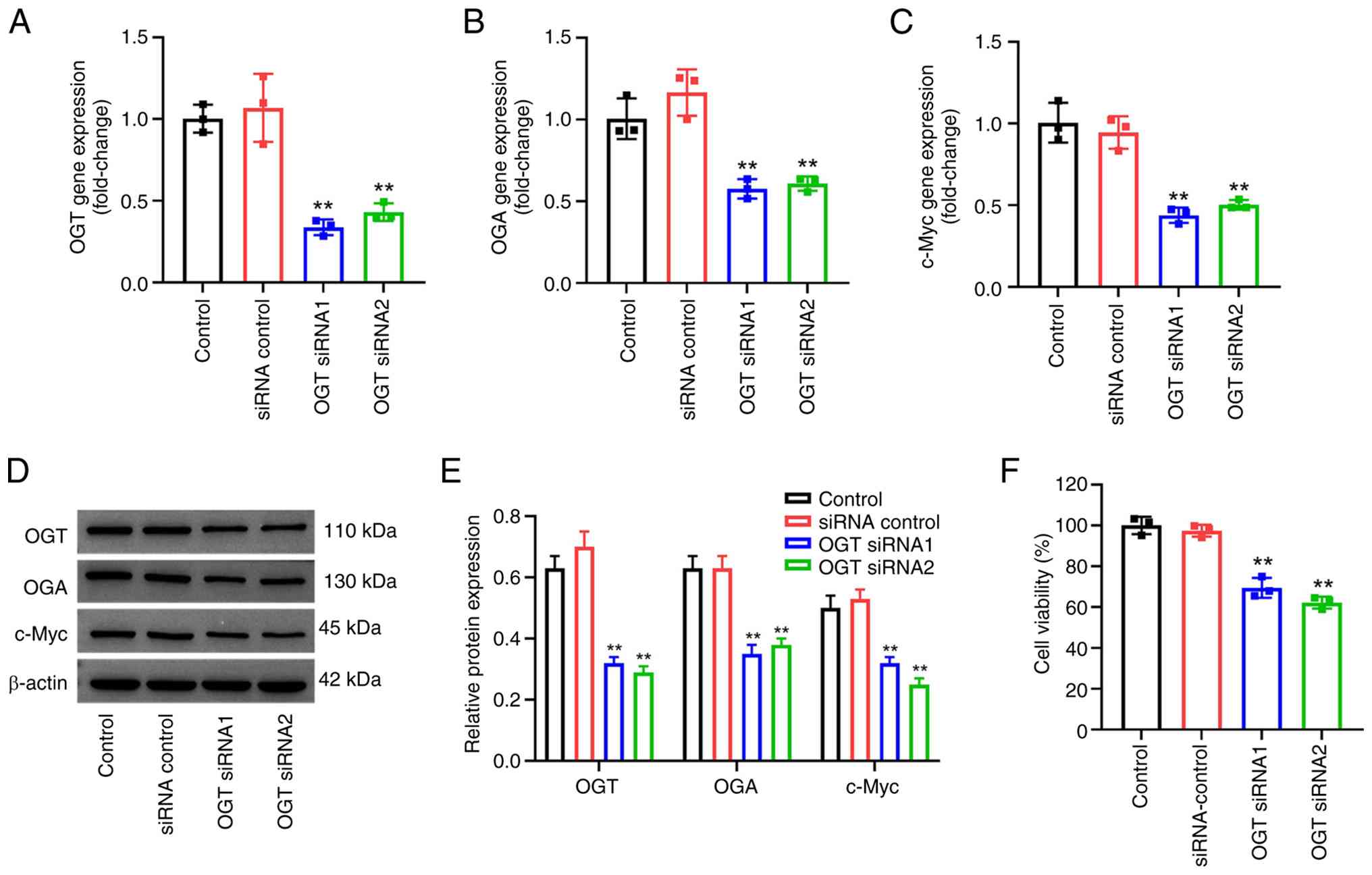
Inhibition of OGT downregulates the
c-Myc pathway in HUVECs
The present study then investigated the role of OGT
in HUVECs using OGT-targeting siRNAs. As shown by the results of
RT-qPCR, both OGT siRNA sequences significantly inhibited the gene
expression of OGT in HUVECs (Fig.
3A). In addition, both OGT siRNA sequences significantly
decreased the gene expression levels of OGA and c-Myc in HUVECs
compared with controls (Fig. 3B and
C). Consistent with the data obtained using RT-qPCR, both OGT
siRNA sequences significantly decreased OGT, OGA and c-Myc protein
expression levels in HUVECs compared with control expression levels
(Fig. 3D and E). The CCK-8 assay
suggested that, compared with control groups, transfection with OGT
siRNA significantly decreased the viability of HUVECs (Fig. 3F). Summarily, these data suggest
that OGT-mediated O-GlcNAcylation is able to stabilize the c-Myc
protein and contribute to the upregulation of c-Myc in HUVECs.
miR-29a inhibits ITGB1 gene
expression
In our previous study, c-Myc, ITGB1, Bcl2 and
miR-29a were identified to be associated with proteoglycan
signaling in angiogenesis (11).
In the present study, miR-29a expression in HUVECs was
significantly upregulated by miR-29a mimics and significantly
downregulated by application of an miR-29a inhibitor compared with
the control groups (Fig. 4A).
Additionally, transfection with the miR-29a mimic significantly
inhibited ITGB1, ITGB1-DT, MUC1 and c-Myc gene expression levels
compared with controls; furthermore, the miR-29a inhibitor exerted
opposing effects to the miR-29a mimic on these proteins by
significantly upregulating their expression compared with the
controls (Fig. 4B-E). The
transfection efficiency of the miR-29a mimic and miR-29a inhibitor
was evaluated compared with specific negative controls for the
mimic and inhibitor respectively in Fig. 4A, but all other experiments were
performed with a negative control for the mimic alone. These
results provide evidence that miR-29a was involved in the
angiogenesis of HUVECs.
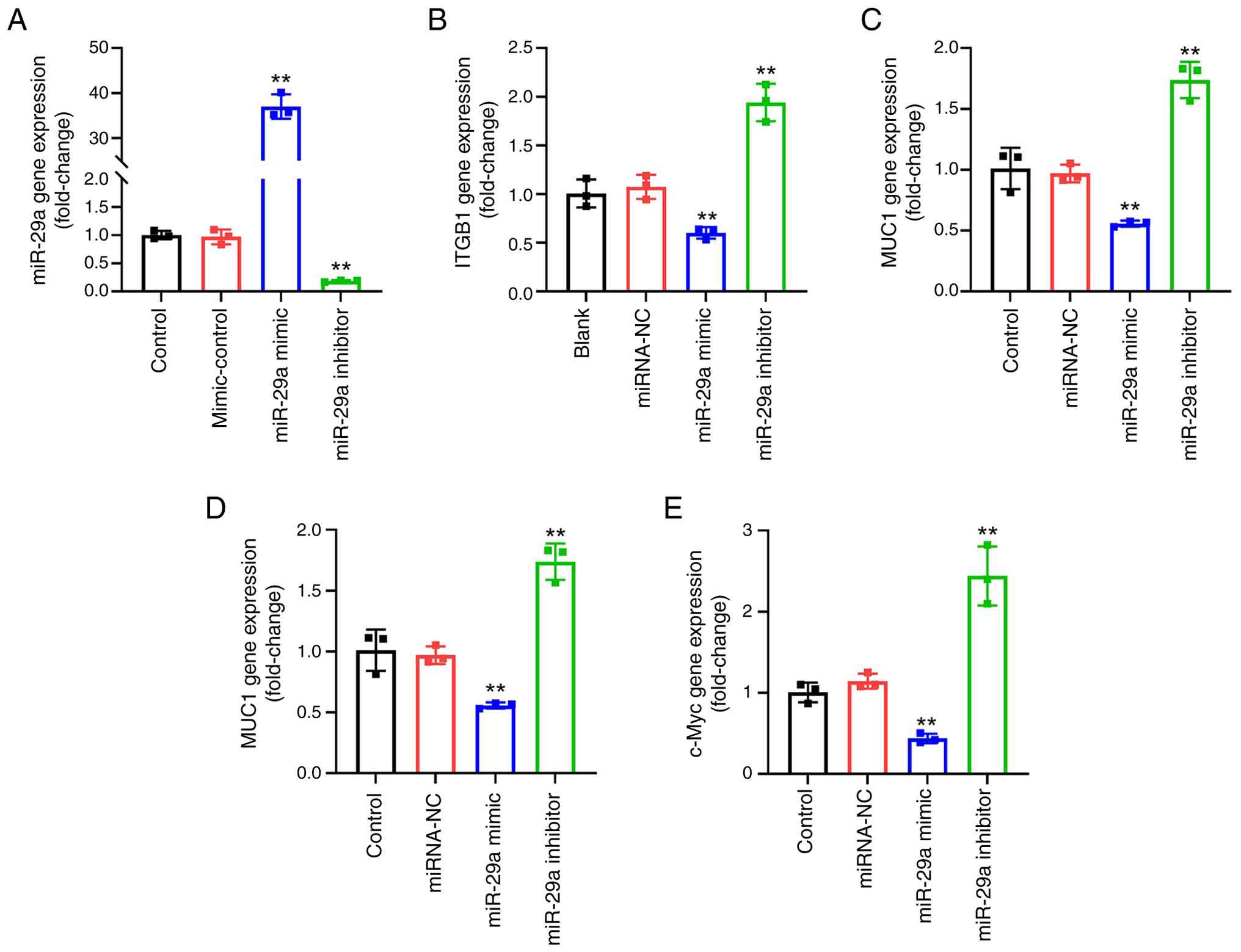 | Figure 4.miR-29a inhibits ITGB1 gene
expression. HUVECs were transfected with 20 µg/ml miR-29a mimic,
mimic NC, miR-29a inhibitor or inhibitor control for 6 h and
cultured for another 48 h. The gene expression levels of (A)
miR-29a (**P<0.01 compared with miRNA-NC or inhibitor-ctrl group
respectively), (B) ITGB1, (C) ITGB1-DT, (D) MUC1 and (E) c-Myc were
assessed using reverse transcription-quantitative PCR. n=3;
**P<0.01 compared with the miRNA-NC group. miR/miRNA, microRNA;
ITGB1, integrin β1; HUVECs, human umbilical vein endothelial cells;
NC, negative control; miRNA-NC, miR-29a mimic negative control;
ITGB1-DT, integrin subunit β1 divergent transcript; MUC1, mucin 1;
inhibitor-ctrl, miR-29a inhibitor negative control. |
miR-29a suppresses the β-catenin/c-Myc
pathway in HUVECs by targeting ITGB1
Finally, the effects of miR-29a on the
β-catenin/c-Myc pathway were explored using WB and dual-luciferase
reporter analysis. The results of the WB revealed that transfection
with the miR-29a mimic significantly reduced the protein expression
levels of OGT, OGA, ITGB1, TCF1, c-Myc, p-GSK3β and p-β-catenin in
HUVECs (Fig. 5A and B); however,
the miR-29a inhibitor exerted opposing effects to the miR-29a mimic
on these RNA levels, significantly upregulating the expression of
OGT, OGA, ITGB1, TCF1, c-Myc and p-β-catenin compared with the
control groups. Additionally, the miR-29a mimic significantly
decreased cell viability, while the miR-29a inhibitor significantly
increased the viability of HUVECs compared with controls (Fig. 5C). The data obtained from the
dual-luciferase reporter assay supported the direct binding of
miR-29a with wild-type ITGB1, as observed by the significant
decrease in luciferase activity in the miR-29a mimic + ITGB1
wild-type group compared with controls (Fig. 5D and E). In summary, these data
suggest that miR-29a suppresses the β-catenin/c-Myc pathway in
HUVECs by targeting ITGB1.
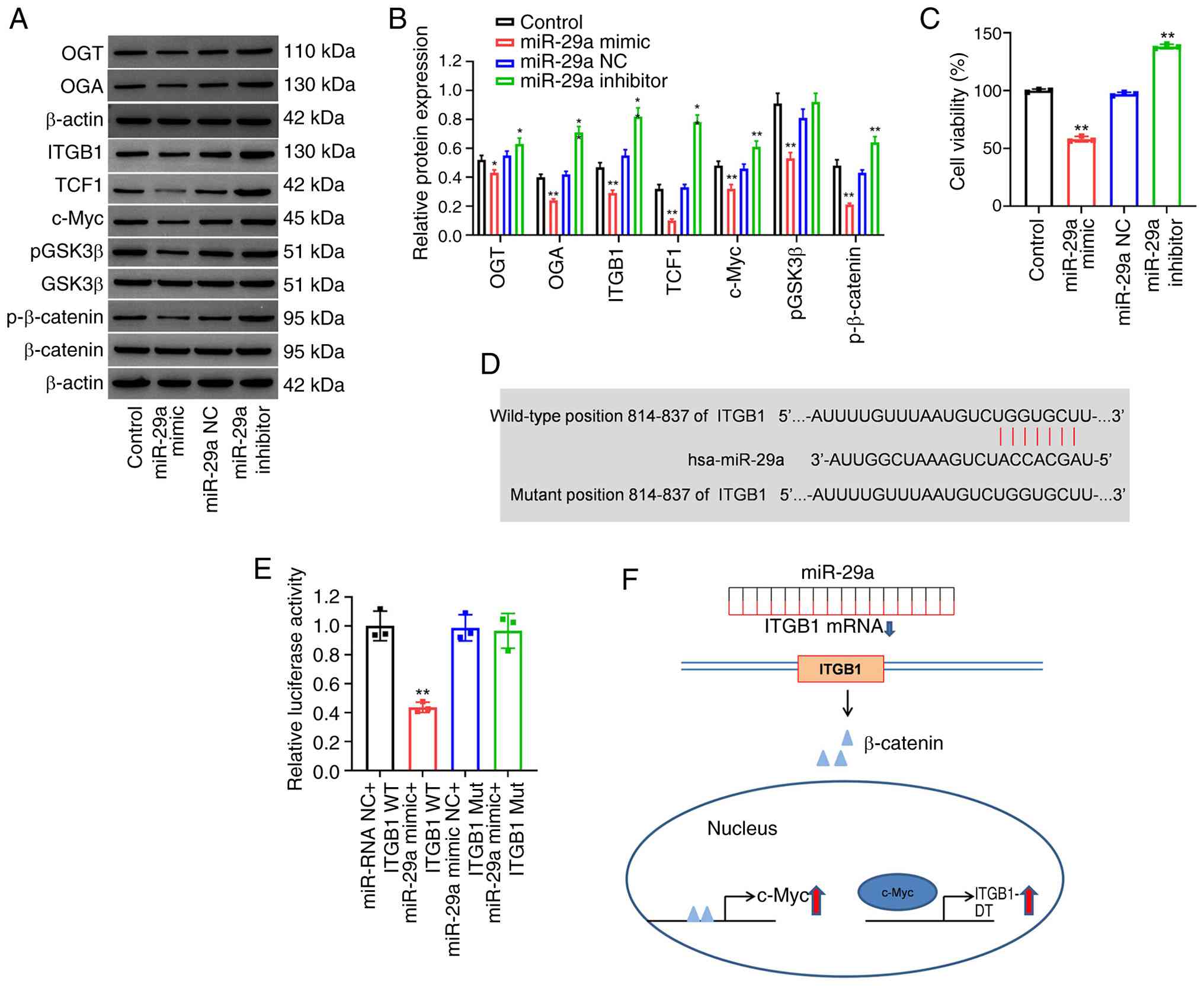 | Figure 5.miR-29a downregulates the
β-catenin/c-Myc pathway in HUVECs by targeting ITGB1. HUVECs were
transfected with 20 µg/ml miR-29a mimics, miR-29a inhibitor or
negative control for 6 h and cultured for another 48 h. (A) The
protein expression levels of OGT, OGA, ITGB1, TCF1, c-Myc, p-GSK3β,
GSK3β, p-β-catenin and β-catenin were assessed using western
blotting and (B) semi-quantified. p-GSK3β and p-β-catenin were
normalized to the levels of total GSK3β or β-catenin, respectively.
(C) Cell viability was analyzed using a Cell Counting Kit-8 assay.
The interaction between miR-29a and ITGB1 was validated using (D)
TargetScan and (E) via dual-luciferase reporter assay (**P<0.01
compared with miR NC + ITGB1 WT group). (F) A schematic figure
illustrating the interactions among miR-29a, ITGB1 and the c-Myc
pathway. n=3; *P<0.05 and **P<0.01 compared with miRNA-NC
group. miR/miRNA, microRNA; HUVECs, human umbilical vein
endothelial cells; ITGB1, integrin β1; OGT, O-linked
N-acetylglucosamine transferase 110 kDa subunit; OGA,
β-N-acetylglucosaminidase; TCF1, T-cell factor 1; p-
phosphorylated; hsa-miR, human microRNA; NC, negative control; WT,
wild-type; Mut, mutant. |
Discussion
miR-29a-3p serves an important role in cell
proliferation and inflammation (28). A study by Qi et al (29) showed that miR-29a-3p inhibited the
development of osteosarcoma through modulating insulin-like growth
factor 1. Additionally, a study by Cao et al (30) reported that miR-29a exerted
apoptotic effects on HUVECs by inhibiting the PI3K/Akt/Bcl2 axis.
Consistent with these reports, the present study demonstrated that
ITGB1 was negatively regulated by miR-29a at the
post-transcriptional level in HUVECs. In addition, miR-29a mimics
significantly reduced cell viability and downregulation of the
c-Myc and p-β-catenin pathways in HUVECs.
ITGB1 is the largest subgroup in the integrin family
and the most important integrin protein expressed by tumor cells,
which is associated with various biological behaviors exhibited by
tumor cells (19,31). The upregulation of ITGB1 activates
the Wnt/β-catenin pathway, which in turn directly transcriptionally
activates ITGB1-DT expression, forming a positive feedback loop of
ITGB1/Wnt/β-catenin/c-Myc (19). A
study by Wang et al (32)
found that Ras-related protein Rab-25 promoted erlotinib resistance
by activating the ITGB1/β-catenin pathway in lung cancer. In
addition, another study by Zou et al (33) reported that platelets facilitated
metastasis of the breast cancer cell line MCF-7 by activating the
integrin α2β1/Wnt/β-catenin pathway. Similarly, the present study
provided evidence that miR-29a mimics inhibited the mRNA level of
ITGB1 via direct binding. Subsequently, ITGB1 was unable to
transcriptionally downregulate c-Myc mRNA through the Wnt/β-catenin
pathway in HUVECs (Fig. 5F).
Mucin is the main component of mucus secretion,
which can be divided into a membrane-associated form, which
interacts with the cell surface, and a secreted subgroup based on
mucin structure and cell localization (20). MUC1, a transmembrane glycoprotein
belonging to membrane-associated subgroup, is abnormally expressed
numerous cancers and functions as a key oncogene in tumorigenesis,
participating in cell regeneration, differentiation, integration
and adhesion by regulating the p53 and β-catenin pathways (20,34).
MUC1 has been reported to activate and regulate the Wnt/β-catenin
pathway and the expression of its key downstream gene, c-Myc in
intrahepatic cholangiocarcinoma (20). Therefore, it was hypothesized that
in IH, the membrane proteins ITGB1 and MUC1 could promote c-Myc
mRNA transcription through the β-catenin pathway. Our previous
study using miRNA sequencing and protein-protein interaction
analyses identified genes and miRNAs related to proteoglycan
signaling pathways in angiogenesis and cancer, including c-Myc,
ITGB1, Bcl2, MUC1 and miR-29a in the HUVEC cell model (11). In the previous study, miR-29a was
upregulated, while c-Myc, ITGB1, Bcl2 and MUC1 were downregulated
in propranolol-treated HUVECs (11). The results of the present study
revealed that inhibition of MUC1 inhibited c-Myc expression and the
viability of HUVECs. Meanwhile, miR-29a mimic transfection
significantly inhibited the expression of the ITGB1, ITGB1-DT,
MUC1, TCF1 and c-Myc genes in HUVECs. TCF1 acts as a key downstream
effector of the β-catenin signaling pathway, confirming the
inhibitory effect of miR-29a mimics on this pathway. However, the
detailed interactions between miR-29a and MUC1 remain ambiguous,
and it is not clear whether miR-29a affects the expression of MUC1
directly or indirectly; the lack of luciferase assay or target
prediction analyses between miR-29a and MUC1 is a limitation of the
present study. Thus, further investigations are warranted.
It was found that inhibition of OGT downregulated
the c-Myc pathway in HUVECs in the present study. In addition,
miR-29a mimics inhibited OGA and OGT expression, while the use of
an miR-29a inhibitor promoted OGA and OGT expression. Thus,
inhibition of OGT and OGA may alleviated the progression of IH.
However, the association between miR-29a and the OGT/OGA pathway
remains to be fully elucidated; proteomics analysis may be required
to further elucidate this association. Additionally, the absence of
an in vivo animal study and the lack of validation using
clinical IH samples limits the translation of the present findings.
Therefore, the expression levels of miR-29a, OGT, OGA and ITGB1 in
clinical IH samples, as well as the role of miR-29a in a hemangioma
subcutaneous xenograft mouse model, should be further explored in
the future.
In the present study, it was found that miR-29a
served an important role in regulating the proliferation of HUVECs.
In addition, miR-29a was able to regulate the β-catenin/c-Myc
pathway and the viability of HUVECs by directly targeting ITGB1.
Furthermore, miR-29a regulated MUC1/c-Myc signaling. Therefore, the
findings of the present study may serve as a theoretical basis for
the clinical research of IH and miR-29a may serve as a potential
therapeutic target for IH.
Acknowledgements
Not applicable.
Funding
The present study was supported by the Suzhou Medical
Application Basic Research Project (grant no. SKY2023184) and the
Postgraduate Research & Practice Innovation Program of Jiangsu
Province (grant no. SJCX24_1824).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
QS and WC were responsible for the study conception
and design. WC, SZ, XZ, YW, YQ, TZ and LZ contributed towards data
collection. QS, WC, LZ, TZ and WL were responsible for the analysis
and interpretation of results. QS and WL were responsible for
drafting the manuscript. WL supervised the entire experimental
process. QS and WL confirm the authenticity of all the raw data.
All authors read and approved the final version of the
manuscript.
Ethics approval and consent to
participate
The use of HUVECs was approved by the ethics
committee of Children's Hospital of Soochow University (approval
no. 2025CS332).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bandera AI, Sebaratnam DF, Wargon O and
Wong LF: Infantile hemangioma. Part 1: Epidemiology, pathogenesis,
clinical presentation and assessment. J Am Acad Dermatol.
85:1379–1392. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Satterfield KR and Chambers CB: Current
treatment and management of infantile hemangiomas. Surv Ophthalmol.
64:608–618. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Nazemian S, Sharif S and Childers ELB:
Infantile hemangioma: A common lesion in a vulnerable population.
Int J Environ Res Public Health. 20:55852023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lee JC, Modiri O, England RW, Shawber CJ
and Wu JK: Propranolol therapy in infantile hemangioma: It is not
just about the beta. Plast Reconstr Surg. 147:875–885. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Hasbani DJ and Hamie L: Infantile
hemangiomas. Dermatol Clin. 40:383–392. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Song J, Zhang J, Wang J, Wang J, Guo X and
Dong W: β1 integrin mediates colorectal cancer cell proliferation
and migration through regulation of the Hedgehog pathway. Tumour
Biol. 36:2013–2021. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Sebaratnam DF, Bandera AL, Wong LF and
Wargon O: Infantile hemangioma. Part 2: Management. J Am Acad
Dermatol. 85:1395–1404. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Muñoz-Garza FZ, Ríos M, Roé-Crespo E,
Bernabeu-Wittel J, Montserrat-García MT, Puig L, Puig L, Gich I and
Baselga E: Efficacy and safety of topical timolol for the treatment
of infantile hemangioma in the early proliferative stage: A
randomized clinical trial. JAMA Dermatol. 157:583–587. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Williams M, Cheng YY, Blenkiron C and Reid
G: Exploring mechanisms of MicroRNA downregulation in cancer.
Microrna. 6:2–16. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Diener C, Keller A and Meese E: Emerging
concepts of miRNA therapeutics: From cells to clinic. Trends Genet.
38:613–626. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang T, Qian Y, Yuan C, Wu Y, Qian H, Lu
H, Hu C and Li W: Propranolol suppresses proliferation and
migration of HUVECs through regulation of the miR-206/VEGFA axis.
Biomed Res Int. 2021:76291762021. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Su C, Mo J, Dong S, Liao Z, Zhang B and
Zhu P: Integrinβ-1 in disorders and cancers: molecular mechanisms
and therapeutic targets. Cell Commun Signal. 22:712024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li S, Sampson C, Liu C, Piao HL and Liu
HX: Integrin signaling in cancer: Bidirectional mechanisms and
therapeutic opportunities. Cell Commun Signal. 21:2662023.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Xie J, Guo T, Zhong Z, Wang N, Liang Y,
Zeng W, Liu S, Chen Q, Tang X, Wu H, et al: ITGB1 Drives
hepatocellular carcinoma progression by modulating cell cycle
process through PXN/YWHAZ/AKT pathways. Front Cell Dev Biol.
9:7111492021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Gu W, Sun H, Zhang M, Mo S, Tan C, Ni S,
Yang Z, Wang Y, Sheng W and Wang L: ITGB1 as a prognostic biomarker
correlated with immune suppression in gastric cancer. Cancer Med.
12:1520–1531. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gao FY, Li XT, Xu K, Wang RT and Guan XX:
c-MYC mediates the crosstalk between breast cancer cells and tumor
microenvironment. Cell Commun Signaling. 21:282023. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Vallée A and Lecarpentier Y: Crosstalk
between peroxisome proliferator-activated receptor gamma and the
canonical WNT/β-catenin pathway in chronic inflammation and
oxidative stress during carcinogenesis. Front Immunol. 9:7452018.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Isagulieva AK, Soshnikova NV and Shtil AA:
Inhibition of the c-Myc oncogene by the aureolic acid group
antibiotics. Dokl Biochem Biophys. 500:308–311. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Chang R, Xiao X, Fu Y, Zhang C, Zhu X and
Gao Y: ITGB1-DT facilitates lung adenocarcinoma progression via
forming a positive feedback loop with ITGB1/Wnt/β-Catenin/MYC.
Front Cell Dev Biol. 9:6312592021. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Song F, Chen FY, Wu SY, Hu B, Liang XL,
Yang HQ, Cheng JW, Wang PX, Guo W, Zhou J, et al: Mucin 1 promotes
tumor progression through activating WNT/β-catenin signaling
pathway in intrahepatic cholangiocarcinoma. J Cancer. 12:6937–6947.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lin CH, Liao CC, Chen MY and Chou TY:
Feedback regulation of O-GlcNAc transferase through translation
control to maintain intracellular O-GlcNAc homeostasis. Int J Mol
Sci. 22:34632021. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ferrer CM, Sodi VL and Reginato MJ:
O-GlcNAcylation in cancer biology: Linking metabolism and
signaling. J Mol Biol. 428:3282–3294. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wu D, Jin J, Qiu Z, Liu D and Luo H:
Functional analysis of O-GlcNAcylation in cancer metastasis. Front
Oncol. 10:5852882020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Luanpitpong S, Rodboon N, Samart P,
Vinayanuwattikun C, Klamkhlai S, Chanvorachote P, Rojanasakul Y and
Issaragrisil S: A novel TRPM7/O-GlcNAc axis mediates tumour cell
motility and metastasis by stabilising c-Myc and caveolin-1 in lung
carcinoma. Br J Cancer. 123:1289–1301. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Jiang T, Yang J, Yang H, Chen W, Ji K, Xu
Y and Yu L: SLC35B4 stabilizes c-MYC protein by O-GlcNAcylation in
HCC. Front Pharmacol. 13:8510892022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yang K, Qiu T, Gong X, Zhou J, Lan Y, Chen
S and Ji Y: Integrated nontargeted and targeted metabolomics
analyses amino acids metabolism in infantile hemangioma. Front
Oncol. 13:11323442023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Horita M, Farquharson C and Stephen LA:
The role of miR-29 family in disease. J Cell Biochem. 122:696–715.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Qi S, Xu L, Han Y, Chen H and Cheng A:
miR-29a-3p mitigates the development of osteosarcoma through
modulating IGF1 mediated PI3k/Akt/FOXO3 pathway by activating
autophagy. Cell Cycle. 21:1980–1995. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Cao Y, Wen H, Leng C and Feng S: MiR-29a
mediates the apoptotic effects of TNF-α on endothelial cells
through inhibiting PI3K/AKT/BCL-2 axis. J Biochem Mol Toxicol.
38:e235982024. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Liu F, Wu Q, Dong Z and Liu K: Integrins
in cancer: Emerging mechanisms and therapeutic opportunities.
Pharmacol Ther. 247:1084582023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wang J, Zhou P, Wang X, Yu Y, Zhu G, Zheng
L, Xu Z, Li F, You Q, Yang Q, et al: Rab25 promotes erlotinib
resistance by activating the β1 integrin/AKT/β-catenin pathway in
NSCLC. Cell Prolif. 52:e125922019. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zuo XX, Yang Y, Zhang Y, Zhang ZG, Wang XF
and Shi YG: Platelets promote breast cancer cell MCF-7 metastasis
by direct interaction: Surface integrin α2β1-contacting-mediated
activation of Wnt-β-catenin pathway. Cell Commun Signal.
17:1422019. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Yasumizu Y, Rajabi H, Jin C, Hata T,
Pitroda S, Long MD, Hagiwara M, Li W, Hu Q, Liu S, et al: MUC1-C
regulates lineage plasticity driving progression to neuroendocrine
prostate cancer. Nat Commun. 11:3382020. View Article : Google Scholar : PubMed/NCBI
|