Introduction
Renal interstitial fibrosis (RIF) is the
accumulation of scarring within the parenchyma and represents a
common final pathway in the majority of chronic and progressive
kidney diseases (1). The
histopathological characteristics of RIF are extracellular matrix
(ECM) component deposition, tubular cell loss, fibroblast
aggregation and a sparse peritubular microvascular system (2). The process of ECM deposition in the
early stage of tissue injury contributes to the repair of damage,
but in the process of chronic kidney disease (CKD), persistent
tissue damage will lead to local tissue inflammation. Inflammatory
cell-derived free radicals initiate epithelial-mesenchymal
transition (EMT), leading to uncontrolled ECM accumulation and
accelerated fibrogenesis. This pathological cascade ultimately
disrupts organ architecture, impairs perfusion and culminates in
functional deterioration, driving progressive CKD toward end-stage
renal disease (ESRD) (3–6). Epidemiological data have revealed a
marked rise in CKD prevalence, currently affecting 9.1% of the
global population. Furthermore, ~50% of these patients will
progress to ESRD, substantially compromising their quality of life
(7).
Long non-coding RNAs (lncRNAs), typically defined as
RNA transcripts >200 nucleotides without protein-coding
capacity, exhibit precise spatiotemporal and tissue-specific
expression patterns (8–10). These molecules function as key
regulators in cellular processes such as cell cycle control,
differentiation and metabolic homeostasis. Their regulatory
influence extends across multiple tiers of gene expression,
encompassing chromatin organization, transcriptional activation and
post-transcriptional modifications. Notably, emerging evidence has
positioned lncRNAs as upstream master regulators that drive
inflammatory responses and orchestrate fibrotic progression across
numerous organ systems, including hepatic, cardiac, pulmonary and
renal tissues (11–13).
In the present study, the classical TGF-β1 factor
was used to construct an EMT model of HK-2 cells. High-throughput
sequencing was used to screen the most significantly differentially
expressed lncRNAs in the EMT model, and the downstream molecular
mechanisms of the screened results were explored to explore their
possible biological functions and their role in the process of
RIF.
The lncRNA NKILA, an inflammation-related lncRNA
serves a role in numerous disease types, including cardiovascular
and cerebrovascular diseases, neuronal diseases, type 2 diabetes,
acute kidney injury and immune regulation (14,15).
Sequencing results suggest that it may also serve an important role
in the process of renal fibrosis. However, at present,
understanding of its function is limited. The majority of studies
explain the synergistic effect of lncRNA NKILA from the perspective
of the synergistic effect of NKILA and NF-κB pathway (16–19),
but its complete biological and molecular mechanism has not been
fully elucidated. Inflammation is a key process in renal fibrosis
and the JAK2/STAT3 signaling pathway is a notable pathway involved
in the process of inflammation and fibrosis. Previous studies have
shown that lncRNA NKILA has STAT family-associated binding sites in
the promoter region (20,21). Therefore, the present study
hypothesized that lncRNA NKILA is implicated in renal fibrosis by
regulating the JAK2/STAT3 signaling pathway. Subsequently, an in
vitro cell experiment was designed to validate this
hypothesis.
Materials and methods
lncRNA transcriptome sequencing
analysis of HK-2 cells induced by TGF-β1
All cells were purchased from Procell Life Science
& Technology Co., Ltd. and genotyped at the short tandem repeat
and Amelogenin locus. The cell lines used in the present study were
all commercial cell lines and did not involve human experimental
ethics.
HK-2 cells (cat. no. CL-0109) were cultured in RPMI
1640 medium (cat. no. L220KJ; Shanghai Basal Media Technologies
Co., Ltd), supplemented with 10% FBS (cat. no. FBSSR-01021-5,
Suzhou Cyagen Biosciences Inc.) at 37°C in a humified 5%
CO2 atmosphere in a CO2 incubator. The cells
were divided into a normal and a control group. After the cells
adhered to 30–40% confluence in a 75 cm2 flask (cat. no.
430720; Corning, Inc.), the cells were treated with RPMI 1640
medium without FBS for 12 h at 37°C . The normal group was cultured
in RPMI 1640 with FBS for 24 h at 37°C. In the control group, HK-2
cells were induced using 10 ng/ml TGF-β1 (cat. no. 100-21;
PeproTech, Inc; Thermo Fisher Scientific, Inc.) for 24 h at 37°C to
construct the RIF model. Total RNA extraction, quality testing and
high-throughput sequencing of lncRNA were performed by Hangzhou
Lianchuan Biotechnology Co., Ltd.
Stranded RNA libraries were constructed using
ribosomal RNA-depleted RNAs. Libraries were controlled for quality,
quantified using a Qubit™ fluorometer (cat. no. Q32857,
Thermo Fisher Scientific) and diluted to 1 ng/µl. The RNA integrity
was assessed using the Agilent 2100 Bioanalyzer (Agilent
Technologies, Inc.) with RNA integrity number >7.0. The final
library concentration was 12 nM. The average insert size for the
final cDNA library was 300±50 bp. RNA libraries were sequenced
using the Illumina NovaSeq™ 6000 (cat. no. 20012850.
Illumina, Inc.) with a sequencing read length of 2×150 bp
(paired-end 150 bp) at both ends, by 12 cycles. The sequencing
depth was 30× and the quantity of data generated was ≥6GB.
Initial data processing involved adapter trimming
and quality filtering of raw sequencing reads using Cutadapt
(22), eliminating sequences
containing adapter contaminants, low-quality bases or undetermined
nucleotides. Data processing was performed using the R language R
Studio (version 4.1.3; Posit Software, PBC), read quality was
assessed with FastQC (Version 0.11.9;
bioinformatics.babraham.ac.uk/projects/fastqc/). Alignment to the
human reference genome was performed using Bowtie2 (Version 2.4.4)
(23) and HISAT2 (Version HISAT-3N
beta) (24). Finally, transcript
assembly for each sample was performed with StringTie (Version
2.2.0) (25) based on the
successfully mapped reads. Subsequently, the transcriptome was
assembled using the StringTie program. The expression abundances of
mRNAs and lncRNAs were quantified in fragments per kilobase million
(FPKM) units utilizing StringTie. Differential expression analysis
was performed using the ‘edgeR’ package (26) in R Studio (version 4.1.3; Posit
Software, PBC), with significance thresholds set at log2 (fold
change)>1 and adjusted P<0.05. After filtering out known mRNA
and transcript variants (<200 bp in length), putative lncRNAs
were identified by integrating the Coding Potential Calculator
(CPC) (27) and the
Coding-Non-Coding Index (CNCI) (28). The intersections of non-coding
transcripts identified by CNCI and CPC were considered as the
putative lncRNAs. Subsequently, the GENCODE database
(gencodegenes.org/) was used to were aligned to the reference human
genome. The specific transcription factor binding site was
predicted by searching the promoter region of NKILA from 2,000 bp
upstream to 99 bp downstream using the JASPAR database (29).
The Gene Ontology (GO) database (geneontology.org)
was employed for functional annotation of genes. Scatter plots were
generated to visualize the top 20 most significantly enriched GO
terms (ranked by adjusted P-value). Statistical significance was
defined as an adjusted P-value <0.05.
Cell culture, transfection and
treatment
HK-2 cells were cultured in RPMI-1640 medium,
supplemented with 10% FBS at 37°C. According to the results of
lncRNA microarray analyses, the overexpression (OE) lentivirus
(Lv)-NKILA (cat. no. 76304), knockdown (KD) Lv-NKILA-short hairpin
(sh)-RNA 1 (cat. no. 108126) and Lv-NKILA-shRNA 2 (cat. no. 108127)
vectors were constructed by Genechem. Empty vectors were used as
the negative control (NC) groups. Before Lv transfection, HK-2
cells were seeded in 6-well plates at a density of
1×104/ml (1 ml/well), so that the cells were evenly
distributed. When the cell density reached 20–30%, Lv transfection
was performed according to the multiplicity of infection (MOI),
whereby MOI=(virus titer × virus volume)/cell number. Before
transfection, fresh complete medium was replaced and the MOI=5
volume Lv solution and 200 µl HiTransGA solution (cat. no. REVG004,
GeneChem Inc.) was added for transfection. After virus
transfection, cell proliferation status was observed every 4 h.
Furthermore, 10 h after virus transfection, fresh complete medium
was replaced and the cells were continually cultured until cell
density reached 70–80%. The NC lentiviral vectors (LVCON313,
1×109 TU/ML; LVCON335 was 2.5×108 TU/ML), the
oe Lv (Lv-NKILA was 1×109 TU/ML) and the KD Lvs
(Lv-NKILA-shRNA 1was 6×108 TU/ml and 2 was
7×108 TU/ml) were transfected into normally cultured
HK-2 cells at MOI=5. After transfection, the cells were cultured
under in RPMI-1640 medium (cat. no. L220KJ; Shanghai Basal Media
Technologies Co., Ltd), supplemented with 10% FBS (cat. no.
FBSSR-01021-5, Suzhou Cyagen Biosciences Inc.) at 37°C, after 10 h
virus transfection, the cells were cultured in fresh medium until
the cell density was 70–80% and the expression levels of lncRNA
NKILA was detected by reverse transcription-quantitative PCR
(RT-qPCR) to evaluate the effects of OE and KD. To investigate the
role of lncRNA NKILA in RIF, the OE Lv, KD Lv and empty vectors
were transfected into HK-2 cells or TGF-β1 induced HK-2 cells with
a MOI=5. Following a 10-h transduction period, the medium was
replaced with fresh culture medium, and cells were incubated for an
additional 24 h until they reached 70–80% confluence. Subsequently,
cells were harvested for downstream analyses. The group details
were as follows: i) In the normal group, after starvation
treatment, HK-2 cells were replaced with fresh complete medium and
cultured for an additional 24 h; ii) in the OE-NC group, cells were
transfected with a NC virus (LVCON335) and Lv transfection was
performed for 24 h at 37°C; iii) in the control group, 10 ng/ml
TGF-β1 was added to HK-2 cells to construct the RIF model for 24 h
at 37°C; iv) in the OE-NKILA group, Lv-NKILA was used for
transfection and cells were collected after 24 h at 37°C Lv
transfection; v) in the control + KD-NC group, HK-2 cells were
induced by adding 10 ng/ml TGF-β1, transfected with KD-NC virus
(LVCON313) at the same time and collected after 24 h stimulation;
vi) in the control + KD-NKILA2 group, HK-2 cells were induced by
adding 10 ng/ml TGF-β1, transfected with Lv-NKILA-shRNA and cells
were collected after 24 h at 37°C stimulation; vii) in the normal +
DMSO group, 1 µl DMSO (cat. no. D8371; Beijing Solarbio Science
& Technology Co., Ltd) was added to the normal group; viii) in
the control + DMSO group, 10 ng/ml TGF-β1 and 1 µl DMSO were added
and cells were collected after 24 h at 37°C stimulation; ix) in the
control + AG490 group, 10 ng/ml TGF-β1 was added, while 50 µM AG490
(cat. no. S1509; Beyotime Biotechnology) was added for intervention
and cells were collected after a total of 24 h stimulation; and x)
in the OE-NKILA + AG490 group, transfection was performed with OE
Lv-NKILA and 50 µM AG490 was added at the same time and cells were
collected after a total of 24 h at 37°C stimulation. The GV513
vector was used to produce Lv-NKILA (76304). The GV493 viral vector
was used to make Lv-NKILA-shRNA (108126) and Lv-NKILA-shRNA
(108127), and standard negative control sequences (instead of an
empty vector) were used for all control plasmids to ensure that no
specific gene expression was produced during transfection. The gene
specific part: Lv-NKILA-shRNA (108126) was GGAAGATATTGCTGCAGTTTG;
the knockdown virus Lv-NKILA-shRNA (108127) was
GGAGAAGTCACACGTTGATTG. The negative control lentivirus sequence was
TTCTCCGAACGTGTCACGT. Lentiviral construction sequences as well as
vector information are provided in Table I.
 | Table I.Lentiviral construction
sequences. |
Table I.
Lentiviral construction
sequences.
| Name | Lv no. | Sequence | Base pairs |
|---|
| OE-NKILA | Lv-NKILA
(76304) | Forward primer |
AGGTCGACTCTAGAGGATCCAGACCCGGCACCCGCGCAACGGAGGAG |
|
|
| Reverse primer |
ACCGTAAGTTATGTGCTAGCTCCAGTTAAATTGAGATATACTTACAC |
| KD-NKILA1 | Lv-NKILA-shRNA
(108126) | Forward primer |
CCGGGGAAGATATTGCTGCAGTTTGCTCGAGCAAACTGCAGCAATATCTTCCTTTTTG |
|
|
| Reverse primer |
AATTCAAAAAGGAAGATATTGCTGCAGTTTGCTCGAGCAAACTGCAGCAATATCTTCC |
| KD-NKILA2 | Lv-NKILA-shRNA
(108127) | Forward primer |
CCGGGGAGAAGTCACACGTTGATTGCTCGAGCAATCAACGTGTGACTTCTCCTTTTTG |
|
|
| Reverse primer |
AATTCAAAAAGGAGAAGTCACACGTTGATTGCTCGAGCAATCAACGTGTGACTTCTCC |
| KD-NC | LVCON313 | Forward primer |
CCGGTTCTCCGAACGTGTCACGTTTCAAGAGAACGTGACACGTTCGGAGAATTTTTG |
|
|
| Reverse primer |
AATTCAAAAATTCTCCGAACGTGTCACGTTCTCTTGAAACGTGACACGTTCGGAGAA |
| OE-NC | LVCON335 | Forward primer |
CCGGTTCTCCGAACGTGTCACGTTTCAAGAGAACGTGACACGTTCGGAGAATTTTTG |
|
|
| Reverse primer |
AATTCAAAAATTCTCCGAACGTGTCACGTTCTCTTGAAACGTGACACGTTCGGAGAA |
Cell Counting Kit-8 (CCK-8) assay
AG490 was dissolved in DMSO to a stock concentration
of 100 mM. Logarithmically growing HK-2 cells were harvested,
inoculated at 4×104/ml and incubated for 24 h at 37°C
under 5% CO2. AG490 concentration gradients (0, 10, 20,
30, 40, 50, 60, 70, 80, 90 and 100 µM) were set to induce cells for
24 h. CCK-8 (cat. no. BS350B; Biosharp Life Sciences) reagent was
subsequently added away from light and incubated in an incubator at
37°C and 5% CO2 for 2 h. Optical density value (single
wavelength, 450 nm) was used to calculate cell viability and screen
out the most suitable AG490 intervention concentration. The
experiment was repeated three times.
Western blot analysis
Cellular proteins were isolated using RIPA buffer
(cat. no. AR0105; Wuhan Boster Biological Technology, Ltd.) and
quantified using a BCA assay. A standard curve constructed from BSA
absorbance values was used to determine sample concentrations,
which were uniformly adjusted to 3 µg/µl. Samples were combined
with 2X SDS-PAGE loading buffer and PBS, heated to 100°C for 8 min
to denature proteins, cooled and stored at −20°C. Proteins (30
µg/lane) were separated on 8% SDS-PAGE gels (cat. no. AR0138; Wuhan
Boster Biological Technology, Ltd.) and transferred to PVDF
membranes (cat. no. 1212639; GVS S.p.A.). After blocking with 5%
non-fat milk for 2 h at room temperature, membranes were incubated
overnight at 4°C with primary antibodies against fibronectin (FN;
1:1,000; cat. no. ab45688; Abcam), collagen I (Col1; 1:1,000; cat.
no. ab138492; Abcam), vimentin (Vim; 1:1,000; cat. no. 10366-1-AP;
Proteintech Group, Inc.), α-smooth muscle actin (α-SMA; 1:1,000;
cat. no. 14395-1-AP; Proteintech Group, Inc.), JAK2 (1:1,000; cat.
no. 17670-1-AP; Proteintech Group, Inc.), STAT3 (1:1,000; cat. no.
10253-2-AP; Proteintech Group, Inc.), phosphorylated (p)-JAK2
(1:500; cat. no. ab32101; Abcam), p-STAT3 (1:500; cat. no. ab76315;
Abcam) and GAPDH (1:5,000; cat. no. 10494-1-AP; Proteintech Group,
Inc.). Following TBS-1% Tween washes, membranes were incubated with
HRP-conjugated goat anti-rabbit IgG (1:10,000; cat. no. BA1054;
Wuhan Boster Biological Technology, Ltd) for 1 h at 25°C. The blots
were developed using an ECL reagent (cat. no. BL520A; Biosharp Life
Sciences) and imaged using a Chemstudio system (Analytik Jena.).
Band intensities were quantified in ImageJ (ImageJ
Launcher1.4.3.67; National Institutes of Health) with GAPDH as the
loading control. The experiments were repeated 3 times.
Immunofluorescence and imaging
analysis
HK-2 cells were seeded on cell slides 14 mm in
diameter at a density of 2×104/well. HK-2 cells were
rinsed in 1X PBS and subsequently fixed in 4% paraformaldehyde for
30 min at room temperature. After 0.5% Triton-100 permeabilization
for 10 min at room temperature, 5% normal goat serum
(Sigma-Aldrich; Merck KGaA) blocking for 30 min at room temperature
and PBS washing was performed. Samples were incubated with
epithelial (E)-cadherin diluted in 1X PBS (1:50; cat. no. ab231303;
Abcam) primary antibody overnight at 4°C. Subsequently, 1X PBS
diluted CoraLite594-conjugate goat anti-rabbit IgG (1:200; cat. no.
ab6721; Abcam) were incubated in the dark at 37°C for 1 h. The
cells were stained with DAPI in dark for 2 min at room temperature,
then washed with 1X PBS. Stained cells were visualized using a
fluorescence microscope and the exposure time was 4 sec (Leica
Application Suite X3.7.5.24914; Leica Microsystems). The
experiments were repeated three times.
RNA extraction and RT-qPCR
Total RNA was isolated from cells using the RNApure
Tissue & Cell Kit (cat. no. CW0584S; Jiangsu CoWin Biotech Co.,
Ltd.) and reverse transcribed into cDNA with the corresponding
synthesis kit (cat. no. CW2569M; Jiangsu CoWin Biotech Co., Ltd.)
according to the manufacturer's protocol. RNA concentration and
purity were assessed using a NanoDrop™ 2000 system (cat.
no. 840-317500, Thermo Fisher Scientific, Inc.), with A260/A280
ratio of 1.8–2.2. All cDNA samples were standardized to a
concentration of 1,000 ng/µl and prepared as 20 µl reaction systems
for subsequent assays. qPCR was carried using a ProFlex™ PCR System
(cat. no. 4484073, Thermo Fisher Scientific, Inc.) with UltraSYBR
mixture (cat. no. CW2601H; Jiangsu CoWin Biotech Co., Ltd.). The
thermocycling conditions were as follows: Pre-denaturation step was
performed at 95°C for 10 min; denaturation was carried out at 95°C
for 15 sec; annealing and extension were conducted simultaneously
at 60°C for 15 sec; and a total of 40 cycles were completed.
Gene-specific primers employed in the presented study are listed in
Table II. Target gene expression
levels were normalized to GAPDH and relative quantification was
determined using the 2−ΔΔCq method (30). The experiments were repeated three
times.
 | Table II.Gene primers used for reverse
transcription-quantitative PCR. |
Table II.
Gene primers used for reverse
transcription-quantitative PCR.
| Gene | Sequence | Base pairs |
|---|
| Vimentin | Forward primer |
CCTTCGTGAATACCAAGACCTGCTC |
|
| Reverse primer |
AATCCTGCTCTCCTCGCCTTCC |
| α-smooth muscle
actin | Forward primer |
CTTCGTTACTACTGCTGAGCGTGAG |
|
| Reverse primer |
CCATCAGGCAACTCGTAACTCTTCTC |
| Epithelial
cadherin | Forward primer |
GCCATCGCTTACACCATCCTCAG |
|
| Reverse primer |
CTCTCTCGGTCCAGCCCAGTG |
| Fibronectin | Forward primer |
GGCGACAGGACGGACATCTTTG |
|
| Reverse primer |
GGCACAAGGCACCATTGGAATTTC |
| Collagen I | Forward primer |
TGGCAAAGAAGGCGGCAAAGG |
|
| Reverse primer |
AGGAGCACCAGCAGGACCATC |
| Janus kinase-2 | Forward primer |
CCAAAGTGGGCAGAATTAGCAAACC |
|
| Reverse primer |
TCGTATGATGGCTCTGAAAGAAGGC |
| STAT3 | Forward primer |
CACCAAGCGAGGACTGAGCATC |
|
| Reverse primer |
AGCCAGACCCAGAAGGAGAAGC |
| GAPDH | Forward primer |
GGAGCGAGATCCCTCCAAAAT |
|
| Reverse primer |
GGCTGTTGTCATACTTCTCATGG |
Statistical analysis
All data were analyzed and plotted using GraphPad
Prism 10.3 software (Dotmatics). The normality of data distribution
was assessed using the Shapiro-Wilk normality test, P>0.05
indicated that the data were normally distributed. Normally
distributed data are presented as mean ± standard deviation.
Comparisons between two independent groups of were analyzed using
an unpaired, two-tailed Student's t-test. Comparisons among
multiple groups were performed using one-way analysis of variance
followed by Tukey's honestly significant difference test. P<0.05
was considered to indicate a statistically significant difference.
All experiments were repeated at least three times.
Results
LncRNA NKILA is potentially associated
with JAK2/STAT3-mediated RIF
After filtering out the known mRNA and transcript
variant (<200 bp) the HK-2 cell samples included 21,924
transcripts going through quality control of raw data, alignment
and assembly. The raw sequence datasets reported in the present
study have been submitted to the Genome Sequence Archive in the
National Genomics Data Center (China National Center for
Bioinformation/Beijing Institute of Genomics, Chinese Academy of
Sciences; GSA-Human, HRA006966) publicly accessible at https://ngdc.cncb.ac.cn/gsa-human/browse/HRA006966.
Inter-sample variability was observed in both the total number of
detected genes and their expression magnitudes. To quantify this,
genes were binned into discrete FPKM intervals and the count of
genes falling within each interval was tallied for every library
(Table III). According to the
CPC (score ≤0.5), the prediction result of protein-coding potential
was 12,347 (Fig. 1A). According to
the CNCI (length, ≥200; exon numbers ≥1; score, ≤0), the prediction
result of protein-coding potential was 10,617 (Fig. 1B). Using R, the FPKM value of the
gene was taken as a parameter and INF_PVAL was set to
1×10−20. Clustering analysis revealed that 1,380 mRNAs
(897 upregulated and 483 downregulated; Fig. 1C) and 1,147 lncRNAs (687
upregulated and 460 downregulated; Fig. 1D) were significantly differentially
expressed between the two groups.
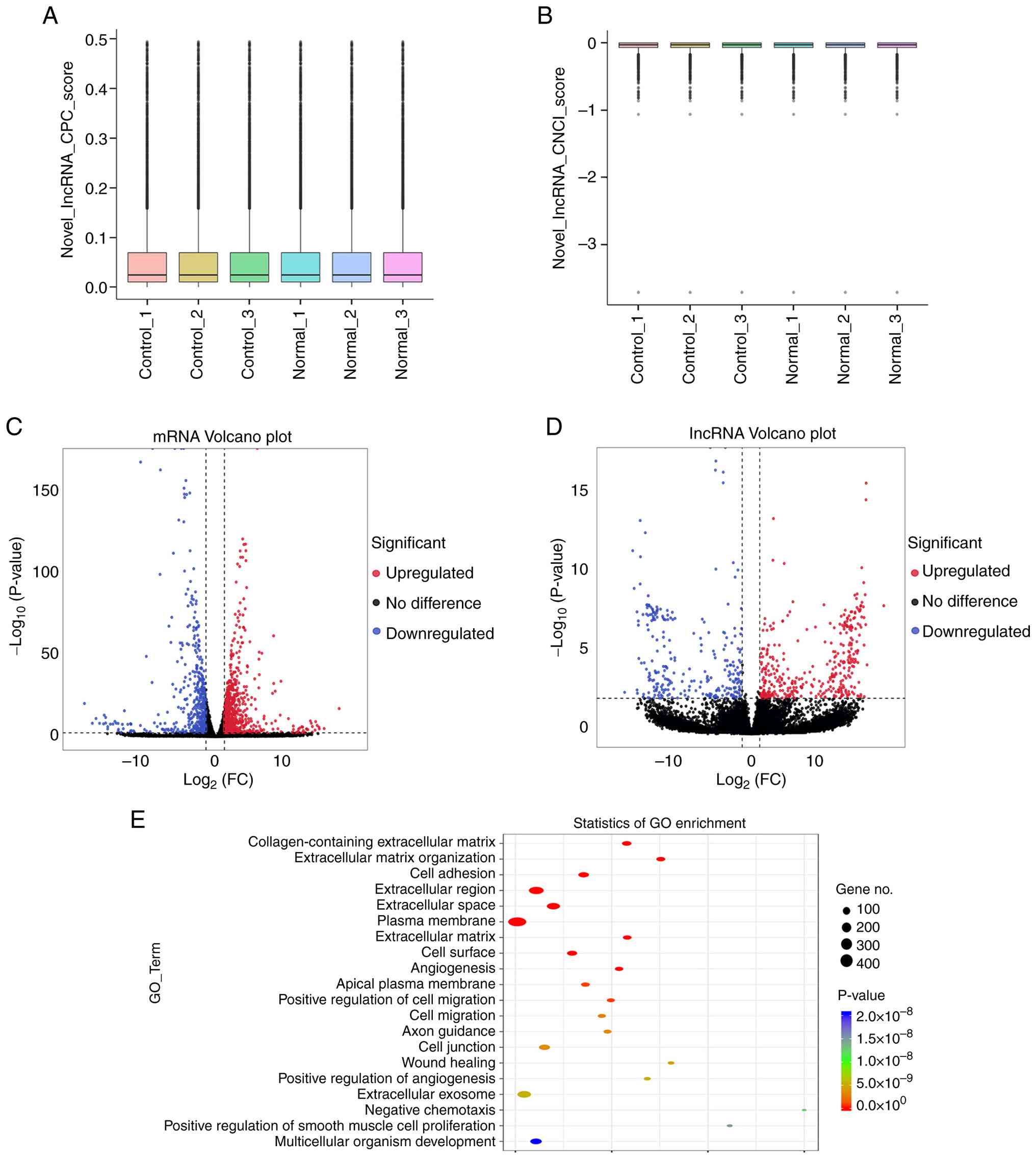 | Figure 1.lncRNA high-throughput data
sequencing. (A) lncRNA CPC score. CPC parameter was set as ≤0.5,
and the predicted result was 12,347. (B) lncRNA CNCI score, CNCI
parameters were set as length ≥200 bp, exon ≥1 and score ≤0 and the
predicted result was 10,617. Volcano plots revealed (C)
differentially expressed mRNAs and (D) differentially expressed
lncRNAs between the normal groups (n=3) and the control group
(n=3). Red represents upregulation, blue represents downregulation
and black represents no significant difference. (E) Top 20 terms in
GO enrichment analysis. lncRNA, long non-coding RNA; CPC, Coding
Potential Calculator CNCI, Coding-Non-Coding Index; GO, Gene
Ontology; FC, fold change. |
 | Table III.Interval distribution statistics of
gene expression values in samples. |
Table III.
Interval distribution statistics of
gene expression values in samples.
| Sample | 0-0.1 FI (%) | 0.1–0.3 FI (%) | 0.3–3.57 FI
(%) | 3.57–15 FI (%) | 15-60 FI (%) | >60 FI (%) |
|---|
| Control_1 | 38058 (56.37) | 5941 (8.80) | 17052 (25.25) | 4795 (7.10) | 1333 (1.97) | 341 (0.51) |
| Control_2 | 38998 (57.76) | 5779 (8.56) | 16566 (24.53) | 4572 (6.77) | 1281 (1.90) | 324 (0.48) |
| Control_3 | 37468 (55.49) | 6291 (9.32) | 17453 (25.85) | 4717 (6.99) | 1276 (1.89) | 315 (0.47) |
| Normal_1 | 37621 (55.72) | 5989 (8.87) | 17491 (25.90) | 4834 (7.16) | 1282 (1.90) | 303 (0.45) |
| Normal_2 | 36799 (54.50) | 6086 (9.01) | 17931 (26.56) | 5044 (7.47) | 1360 (2.01) | 300 (0.44) |
| Normal_3 | 37120 (54.98) | 5925 (8.78) | 17776 (26.33) | 5034 (7.46) | 1349 (2.00) | 316 (0.47) |
High-throughput sequencing identified 1,380
differentially expressed mRNAs, and GO enrichment analysis revealed
significant associations with 1,127 biological functions. Based on
significance rankings, the top 20 enriched functional categories
were visualized, encompassing ‘collagen-containing ECM’, ‘ECM
organization’, ‘cell adhesion’, ‘extracellular region’,
‘extracellular space’, ‘plasma membrane’, ‘cell migration’ and
‘angiogenesis’ (Fig. 1E).
Based on the analysis of differentially expressed
lncRNAs, lncRNA NKILA was the most significantly cis-regulated
target transcripts in TGF-β1-induced HK-2 cells compared with the
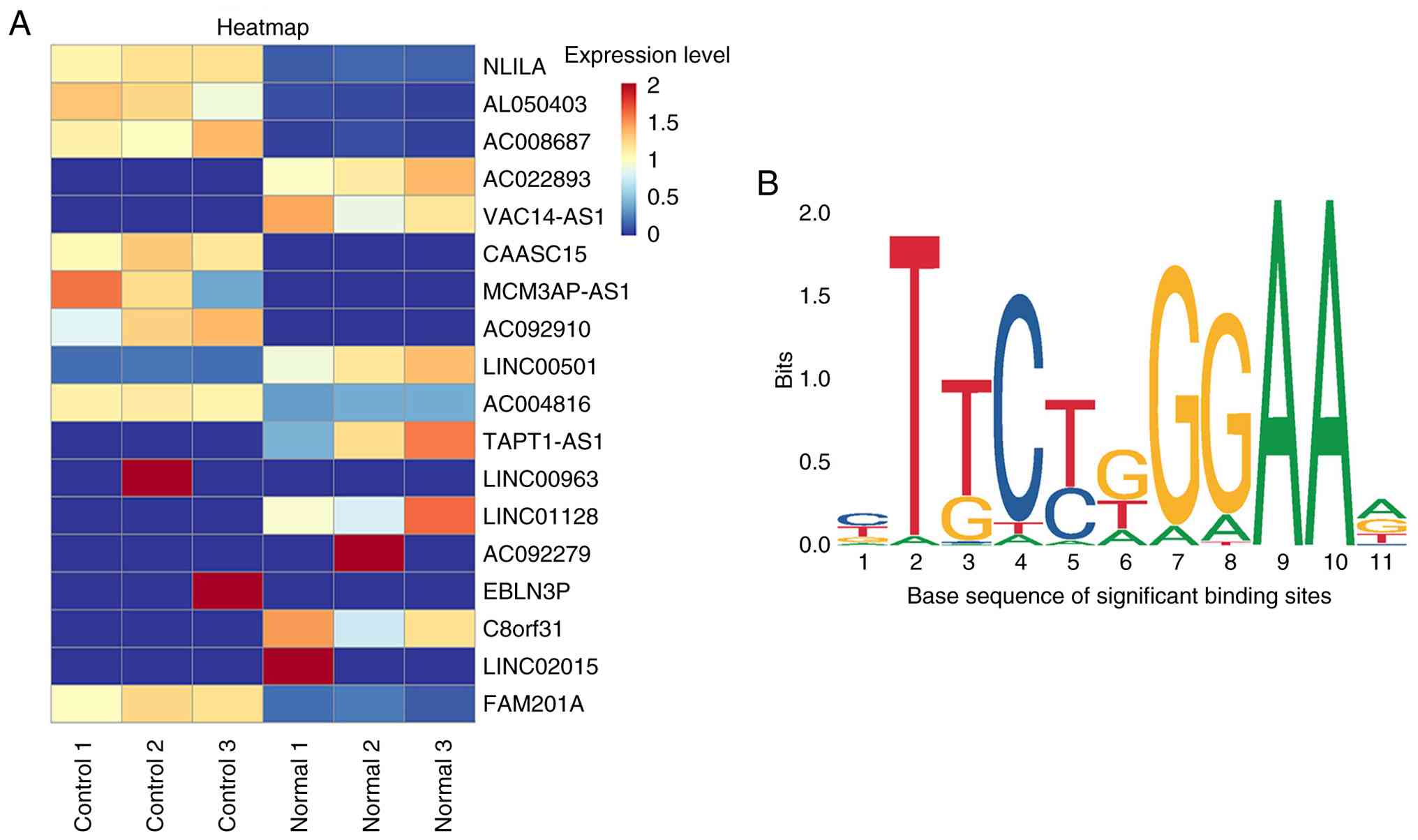
normal groups and was increased in the RIF model (Table IV; Fig. 2A). The binding sites of NKILA
promoter region were successfully predicted by JASPAR database. The
lncRNA NKILA and STAT3 were found to have significant binding sites
(Table V; Fig. 2B). Therefore, JAK2/STAT3 may be the
putative regulatory pathway mediated by lncRNA NKILA.
 | Table IV.Top five significantly cis-regulated
target transcripts in the control vs. normal groups. |
Table IV.
Top five significantly cis-regulated
target transcripts in the control vs. normal groups.
| ID | Length (bp) | Gene name | log2
(FC) | P-value | Regulation |
|---|
|
ENST00000614771 | 2625 | NKILA | 2.926772479 |
1.15×10−34 | Up |
|
ENST00000667822 | 1630 | AL050403 | 3.994934816 |
2.33×10−18 | Up |
|
ENST00000599209 | 598 | AC008687 | 4.053600164 |
9.00×10−18 | Up |
|
ENST00000517664 | 3549 | AC022893 | −13.14250225 |
6.21×10−17 | Down |
|
ENST00000666405 | 2930 | VAC14-AS1 | −13.12616293 |
7.42×10−16 | Down |
 | Table V.Prediction results of binding
sites. |
Table V.
Prediction results of binding
sites.
| Matrix ID | Name | Relative score
(%) | Start | End | Predicted
sequence |
|---|
| MA0144.2 | MA0144.2.STAT3 | 88.33 | 1308 | 1318 | TTGCAGAGAAG |
| MA0144.2 | MA0144.2.STAT3 | 87.19 | 585 | 595 | GAGCTGGGAAC |
| MA0144.2 | MA0144.2.STAT3 | 87.00 | 1551 | 1561 | TTGTCTGAAAG |
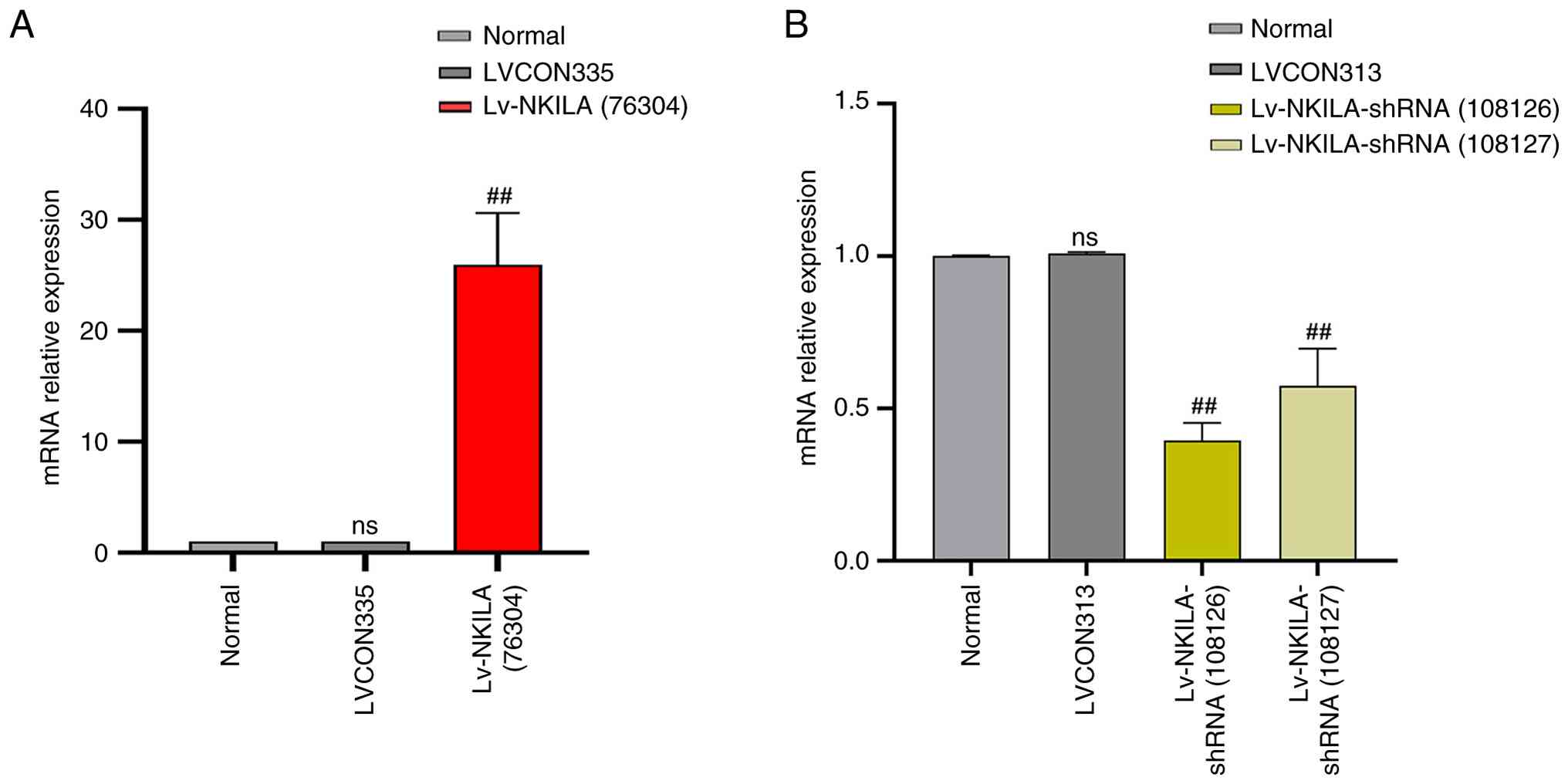
LncRNA NKILA transfection assay
Compared with normal cultured HK-2 cells, the
negative control lentivirus (LVCON335) did not produce biological
effects. Compared with normal HK-2 cells and LVCON335 transfected
cells, the expression level of lncRNA NKILA was significantly
increased after transfected with OE-NKILA lentivirus. Similarly,
transfection of the lentiviral negative control (LVCON313) did not
produce any biological effect in normal cell culture HK-2 cells,
but the expression of lncRNA NKILA was significantly reduced in
normal HK-2 cells transfected with two different fragments of
lncRNA NKILA knockdown virus (Fig.
3).
LncRNA NKILA OE facilitates EMT of
HK-2 cells through the JAK2/STAT3 pathway
To assess the regulatory role of lncRNA NKILA in EMT
through the JAK2/STAT3 pathway, HK-2 cells were transduced with
either lncRNA OE-NKILA or an empty vector control (OE-NC). Compared
with the OE-NC group, OE-NKILA significantly upregulated
mesenchymal markers (FN, Col1, α-SMA and Vim) and downregulated the
epithelial marker E-cadherin, as determined by RT-qPCR and western
blotting (Fig. 4).
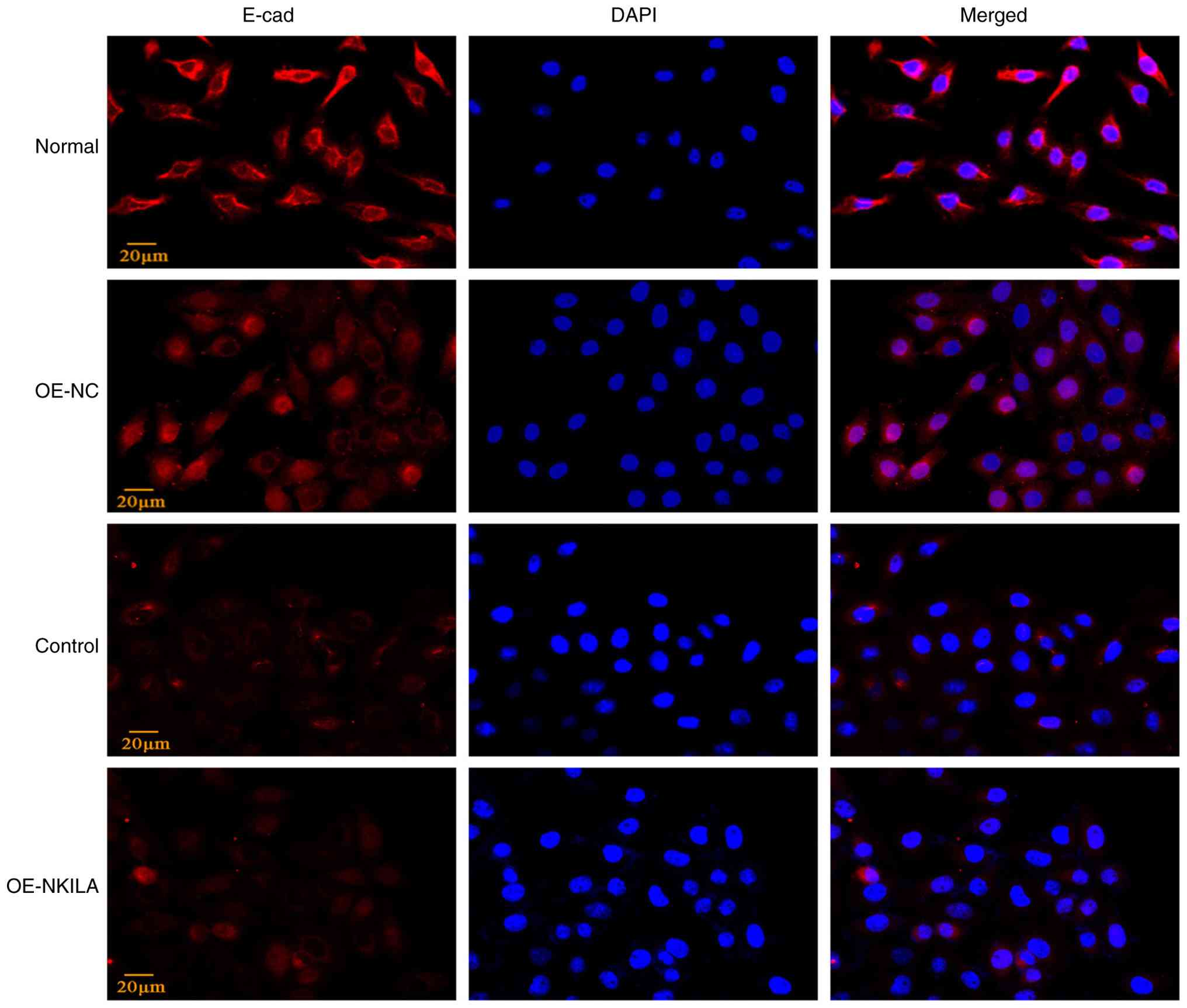
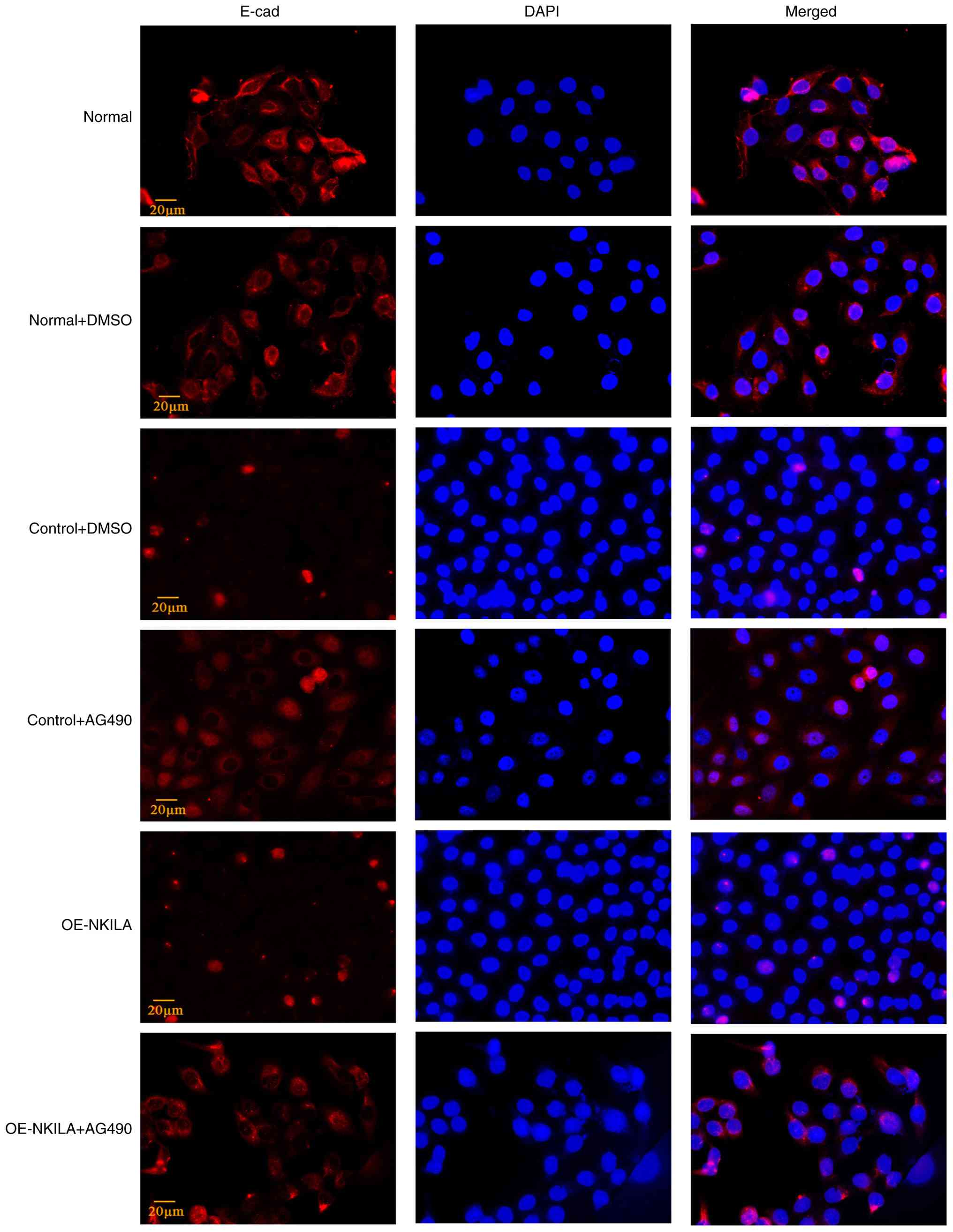
Immunofluorescence staining further demonstrated a reduction in
E-cadherin expression in OE-NKILA-treated cells compared with the
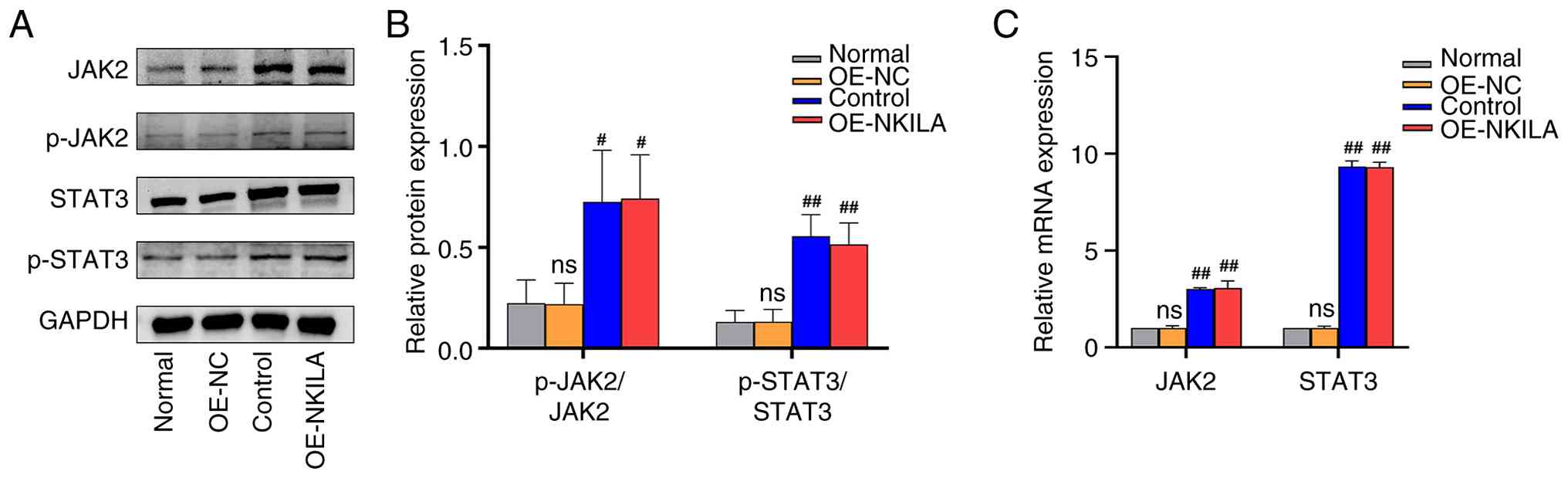
controls, indicating enhanced EMT progression (Figs. 4 and 5). Compared with the normal control group
and the OE-NC group, both the phosphorylation ratio of JAK2
(p-JAK2/JAK2) and that of STAT3 (p-STAT3/STAT3) were significantly
elevated in the control and the OE-NKILA group. Consistent with
these protein-level findings, qPCR analysis revealed a
corresponding upregulation in the expression of relevant downstream
targets, these results suggest that lncRNA NKILA promotes EMT
through activation of the JAK2/STAT3 pathway (Fig. 6).
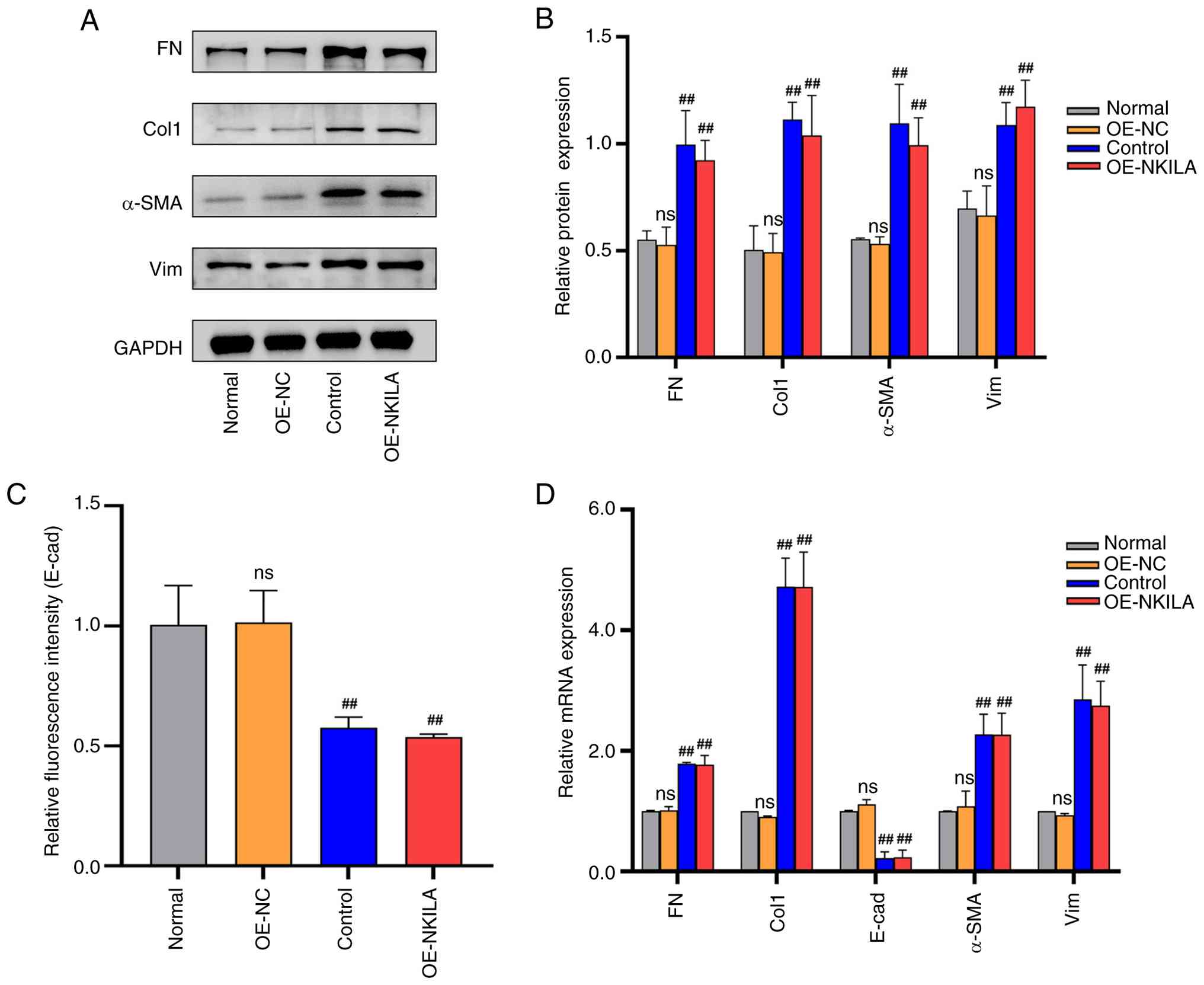 | Figure 4.Partial EMT phenotype proteins in
HK-2 cells transfected with lncRNA NKILA overexpression virus (A)
Representative western blotting images and (B) quantification of
EMT phenotypic indicators. (C) Statistical analysis results of
E-cad fluorescence intensity (n=3). (D) RT-qPCR statistical results
of EMT phenotype indicators (n=3). ‘Normal’ indicates after
starvation treatment, HK-2 cells were replaced with fresh complete
medium and continued to be cultured for 24 h. Control, following
starvation treatment, 10 ng/ml TGF-β1 cytokine diluent was added to
induce HK-2 cells to construct the renal interstitial fibrosis
model for 24 h. ‘OE-NKILA’ indicates after starvation,
overexpressed Lv-NKILA (76304) was used for transfection and cells
were collected after 24 h of lentivirus transfection. Compared with
OE-NC ##P<0.01. E-cad; epithelial-cadherin; RT-qPCR,
reverse transcription quantitative PCR; EMT, epithelial-mesenchymal
transition; NC, negative control; Lv, lentivirus; OE,
overexpression; FN, fibronectin; Col1, collagen I; α-SMA, α-smooth
muscle actin; Vim; vimentin; ns, not significant. |
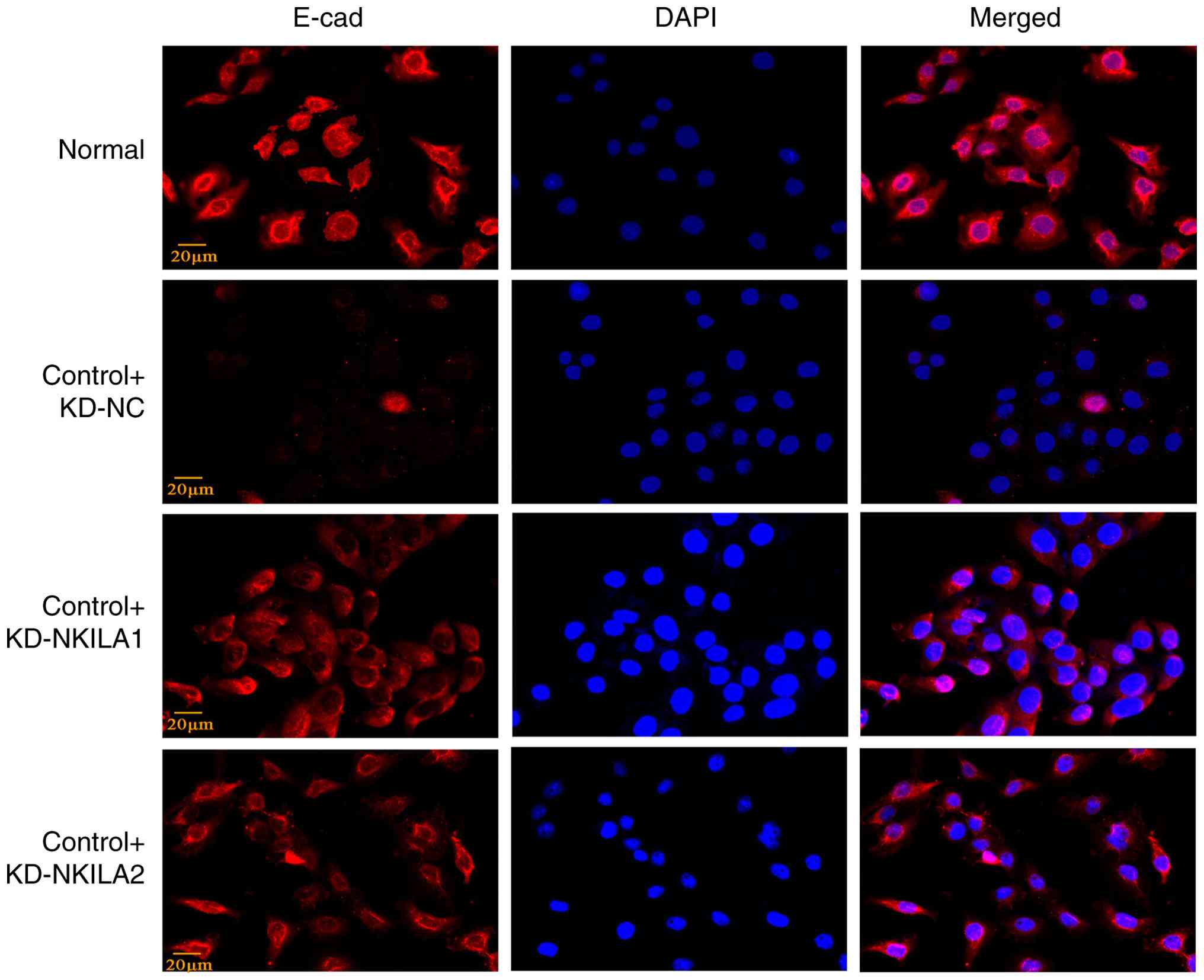
KD of lncRNA NKILA attenuates EMT in
HK-2 cells and inhibits the activation of JAK2/STA3 pathway
In addition, two different lncRNA NKILA-KD
lentiviruses (control + KD-NKILA1 and control + KD-NKILA2) and
their NC were transfected into HK-2 cells to establish the model of
EMT induced by TGF-β1. Compared with control + KD-NC group, NKILA
KD significantly suppressed mesenchymal markers, FN, Col1, α-SMA
and Vim, at both transcript and protein levels. Of note, E-cadherin
expression was consistently elevated upon NKILA depletion, as
demonstrated through RT-qPCR and immunofluorescence analyses
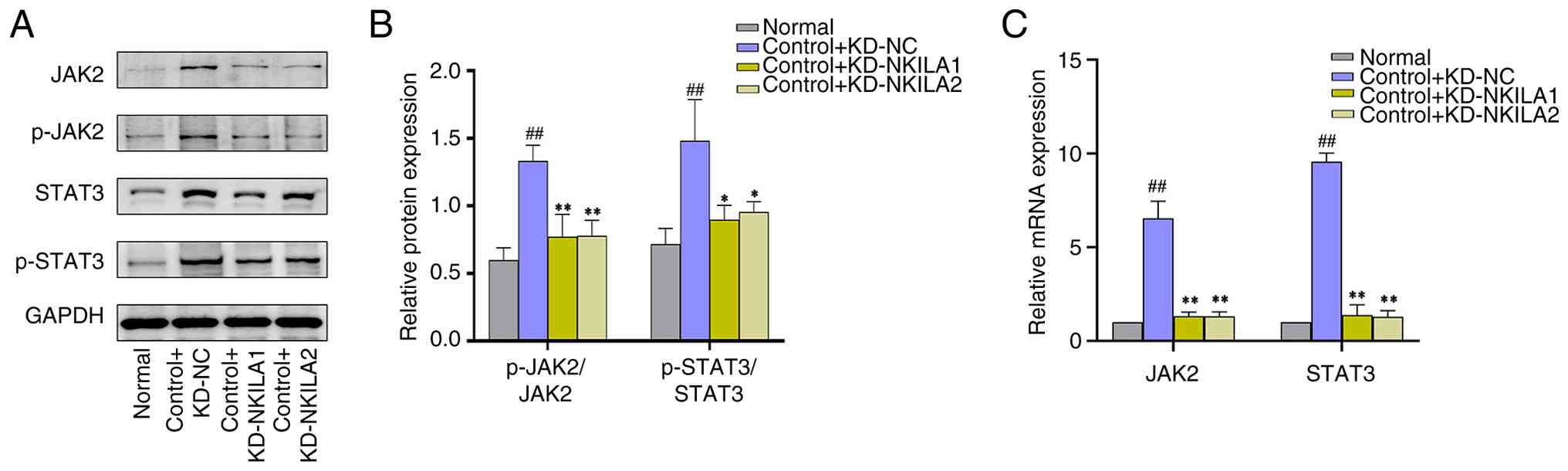
(Figs. 7 and 8). Collectively, Compared with the
control + KD-NC group, The ratio of P-JAK2 to JAK2 protein and
P-STAT3 to STAT3 protein in NKILA KD group (control + KD-NKILA1 and
control + KD-NKILA2) were significantly decreased, these findings
suggest that lncRNA NKILA is key in preventing EMT under TGF-β1
conditions and inhibiting the JAK2/STAT3 pathway (Fig. 9).
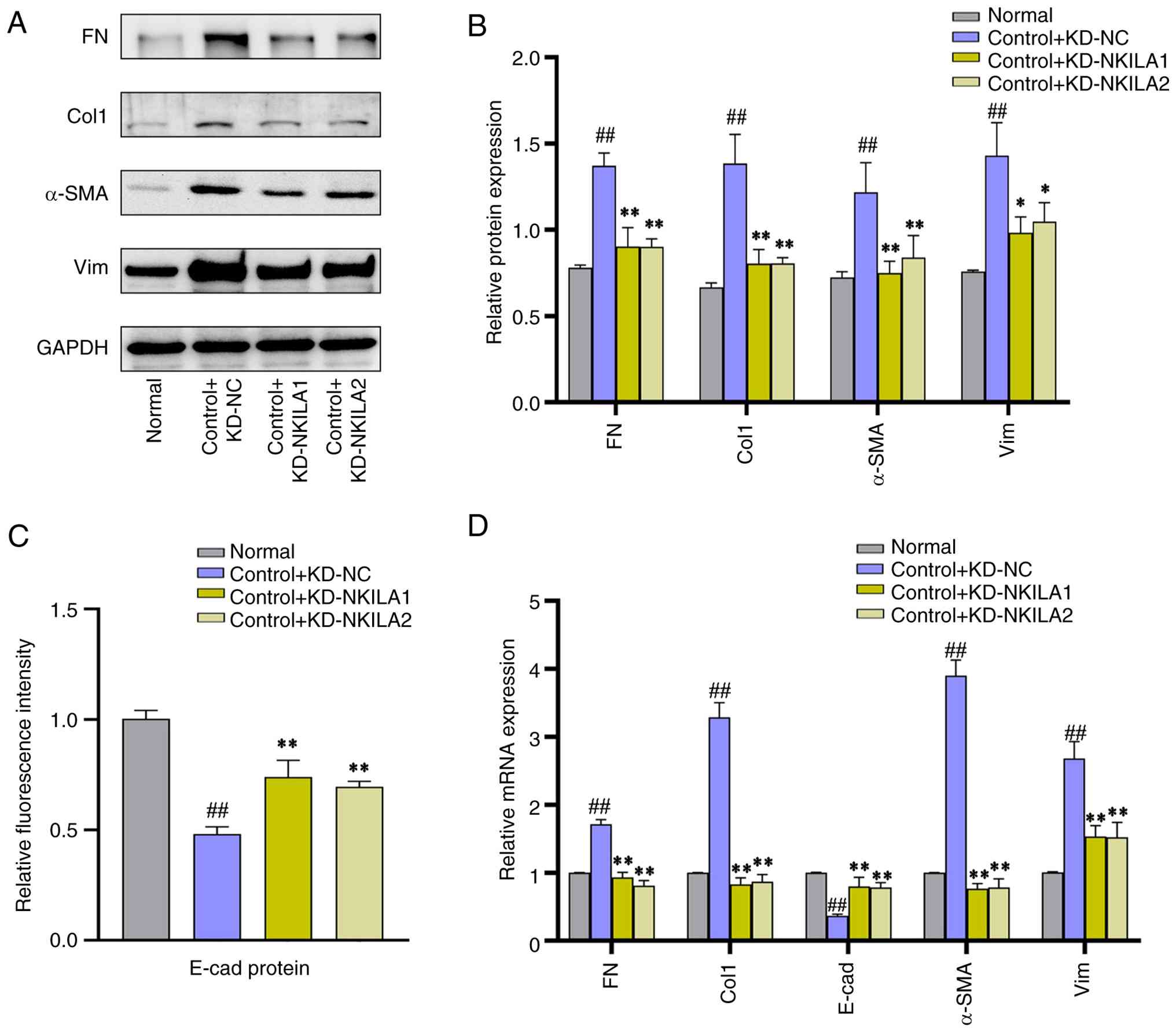 | Figure 7.Partial EMT phenotype proteins in
HK-2 cells transfected with lncRNA NKILA knockdown lentivirus (A)
Representative western blotting bands and (B) semi-quantification
results of EMT phenotypic indicators. (C) E-cad fluorescence
intensity and (D) EMT phenotypic index RT-qPCR (n=3). ‘Normal’
indicates starvation treatment. Compared with normal group
##P<0.01. Compared with control + KD-NC group,
**P<0.01 and *P<0.05. EMT, epithelial-mesenchymal transition;
E-cad, epithelial-cadherin; RT-qPCR, reverse transcription
quantitative PCR; KD, knockdown; Lv, lentivirus; shRNA, short
hairpin RNA; NC, negative control; FN, fibronectin; Col1, collagen
I; α-SMA, α-smooth muscle actin; Vim; vimentin. |
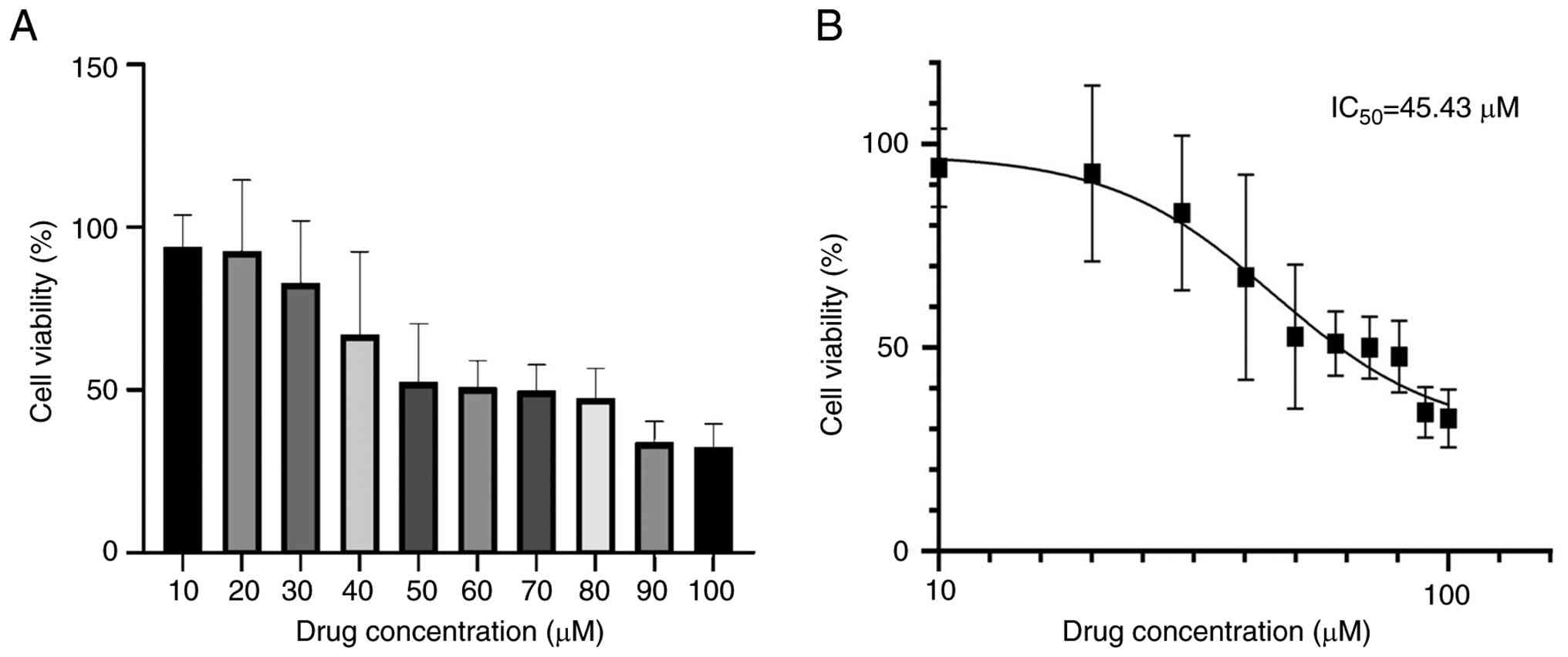
lncRNA NKILA interferes with the
fibrotic process of HK-2 cells by regulating the JAK2/STAT3
pathway
Through the OE and KD of lncRNA NKILA experiments,
lncRNA NKILA was demonstrated to initiate the fibrosis of HK-2
cells and NKILA may serve a fibrogenic role by regulating the
JAK2/STAT3 signaling pathway. In order to validate these findings,
AG490, a specific inhibitor of the JKA2/STAT3 signaling pathway,
was introduced for further experimental verification. The optimal
administration concentration of AG490 was screened using the CCK-8
method. Findings revealed that the IC50 value was 45.43
µM. The present study examined the viability of HK-2 cells in a
concentration gradient ranging from 10 to 100 µM and found no
significant difference in cell viability in a molar concentration
gradient ranging from 50 to 80 µM. However, since AG490 requires
the use of DMSO for dissolution, to avoid excessive damage to HK-2
cells and affect the experimental results, the intervention dose of
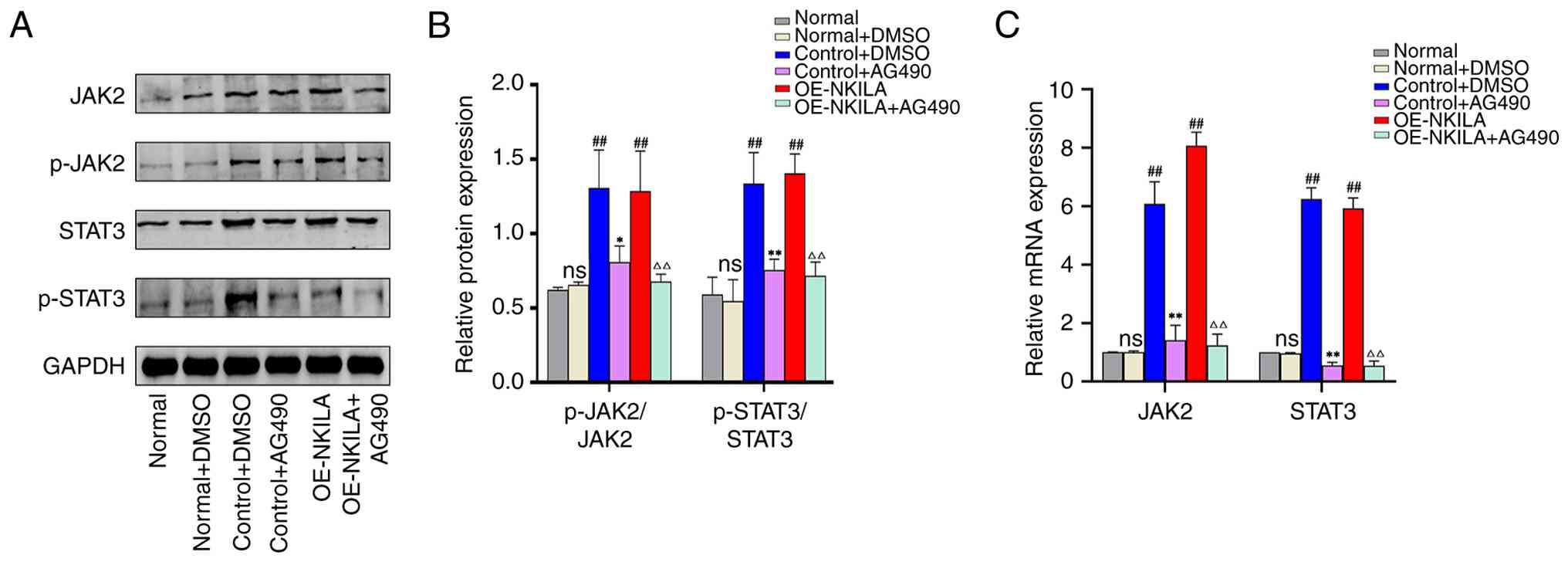
AG490 used in the present experiment was 50 µM (Fig. 10). Subsequently, AG490 was used to
treat the TGF-β1-induced HK-2 cell RIF model and NKILA-induced HK-2
cell EMT model. Compared with the normal control group, no
significant difference was observed in the phosphorylation ratios
of JAK2 and STAT3 in the normal + DMSO group. The phosphorylation
ratios of JAK2 and STAT3 in the control + DMSO group were
comparable to those in the oe-NKILA group. In contrast, both ratios
were significantly elevated in the oe-NKILA group relative to the
control + DMSO group. Treatment with AG490 markedly suppressed JAK2
and STAT3 phosphorylation in both the control and oe-NKILA
backgrounds: specifically, the p-JAK2/JAK2 and p-STAT3/STAT3 ratios
were significantly reduced in the control + AG490 group compared
with the control + DMSO group, and similarly decreased in the
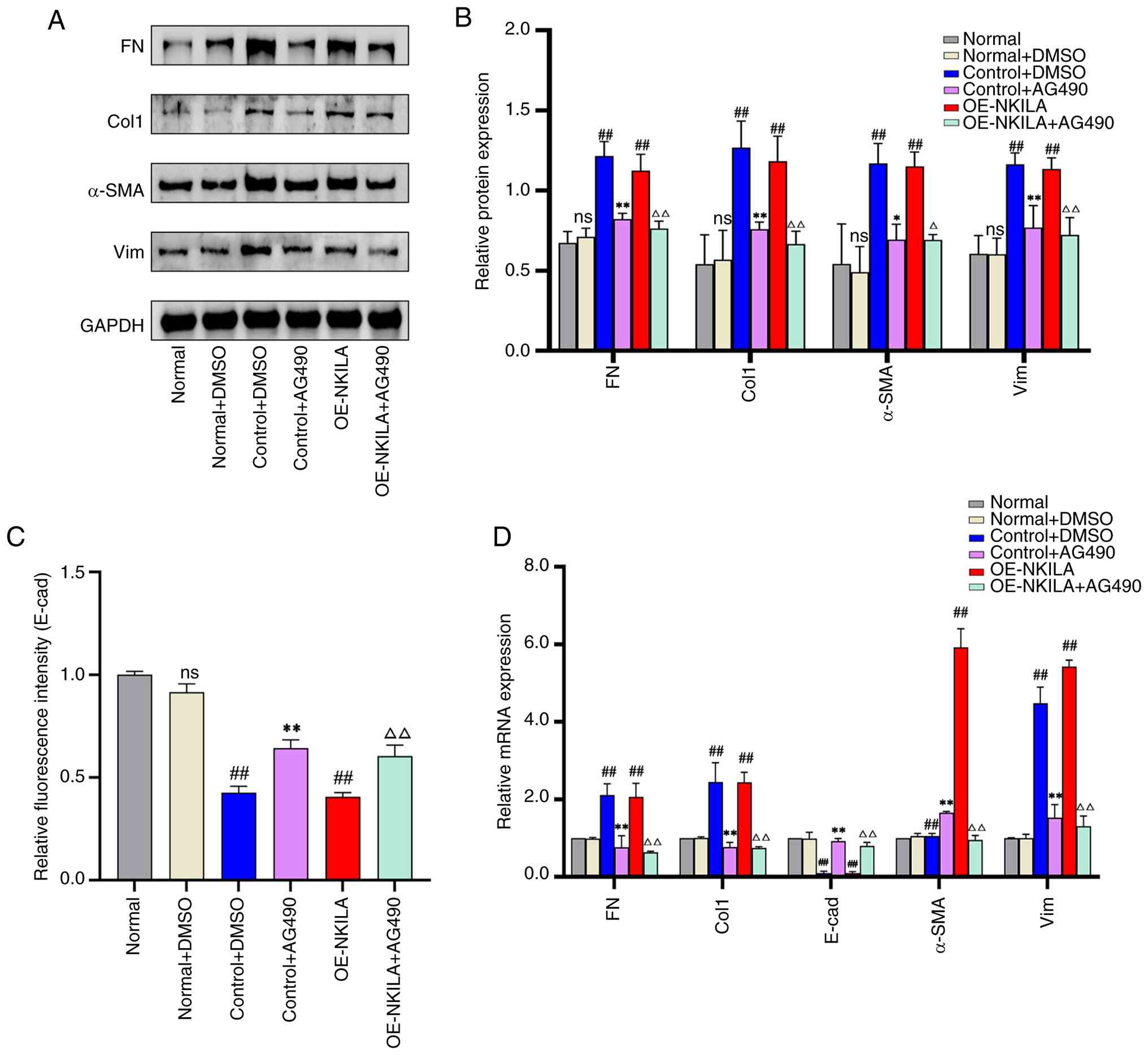
oe-NKILA + AG490 group relative to the oe-NKILA group (Fig. 11). DMSO had no effect on EMT
markers vs. normal controls. The control+ DMSO and oe-NKILA groups
both showed significant EMT induction (upregulated FN, Col1, Vim,
α-SMA; downregulated E-CAD). AG490 treatment markedly reversed this
phenotype, suppressing mesenchymal markers and restoring E-CAD in
both control and oe-NKILA groups (Figs. 12 and 13). Collectively, these data indicate
that NKILA modulates the JAK2/STAT3 axis and thereby attenuates the
fibrogenic response in HK-2 cells.
Discussion
RIF, characterized by abnormal accumulation of
collagen and other ECM proteins in the tubulointerstitial
compartment, is a frequent histological finding in diverse renal
pathologies (31). In CKD
progression, EMT drives ECM remodeling and promotes RIF
development. Notably, evidence has supported the pathogenic role of
EMT in renal repair processes. During initial EMT stages, injured
epithelial cells trigger profibrotic signaling, leading to immune
cell recruitment and subsequent activation. These infiltrating
immune cells exacerbate fibrogenesis through sustained cytokine
release (32).
EMT is orchestrated by complex molecular networks
involving multiple regulatory tiers. Key mechanisms encompass
transcriptional activation, altered expression of specific
cell-surface proteins, cytoskeletal remodeling and production of
ECM-degrading enzymes. With the growing improvement in
transcriptome sequencing technology, large numbers of lncRNAs have
been reported to represent attractive intervention targets for
cardiovascular and cerebrovascular, nervous system, tumor and EMT
processes in previous studies (33–37).
Understanding the functions and spatial structures of lncRNAs could
therefore help to develop potential novel therapies.
lncRNA NKILA was identified and named as a NF-κB
interacting lncRNA in breast cancer, with its main function being
to bind to the NF-κB/IκB complex, inhibiting the NF-κB pathway by
masking the phosphorylation site of IκB and stabilizing the complex
(38–40). However, the interactions between
fragments of NKILA and the NF-κB domain are weak and non-specific.
At present, the studies on NKILA function mainly focus on the role
of NKILA-NF-κB-IκBα in diseases such as cardiovascular and
cerebrovascular disease, lung adenocarcinoma, liver cancer,
esophageal squamous cell carcinoma and osteoarthritis from the
perspective of protecting the NF-κB-IκBα complex from abnormal
activation of the NF-κB signaling pathway (41–43).
However, simultaneously, NKILA can be used as an independent risk
factor for colorectal cancer, coronary heart disease, lung
adenocarcinoma, Parkinson's disease and type 2 diabetes (44–46).
Due to a limited understanding of lncRNA NKILA, there is no unified
standard to assess the change of lncRNA NKILA expression in
different diseases. However, lncRNA NKILA has marked research
potential, which may lead to an improved understanding of lncRNAs
in diseases.
In the present study, EMT markers were assessed in
HK-2 cells following stimulation with TGF-β1 or lncRNA NKILA. The
data revealed that lncRNA NKILA OE elevated FN, Col1, α-SMA and
Vim, while repressing E-cadherin, mirroring a pro-fibrotic EMT
signature. Based on the biological functions of lncRNA NKILA, such
as its involvement in immune regulation and inflammatory response
(47,48), the present results suggest that
NKILA may activate inflammation-related signaling pathways in the
process of fibrosis.
Following previous results of high-throughput
sequencing analysis, the promoter region of lncRNA NKILA was
demonstrated to have a possible binding site with STAT3. The
JAK2/STAT3 pathway is dominant among various subtypes of JAK/STAT
(49,50) and is a key signaling pathway
involved in inflammation with phosphorylation as the main mode of
action. The activation of this signaling pathway is implicated in
driving fibrotic processes across multiple organs, including the
lung, liver and kidneys. The role is mediated through its
involvement in both inflammation and the induction of EMT (51–54).
JAK2/STAT3 can be both an initiating factor and downstream in the
process of renal fibrosis, which simultaneously respond to numerous
cytokine signals to activate myofibroblasts and induce EMT
(55).
The present study also demonstrated that the
JAK2/STAT3 signaling pathway was activated in two different models
of cell fibrosis (TGF-β1-induced and lncRNA NKILA-induced HK-2
cells), accompanied by the expression changes in EMT markers. In
the rescue experiment with AG490, suppressing the expression of
JAK2/STAT3 pathway notably reduced EMT changes in RIF cell models.
On the one hand, the results demonstrated a potential key role of
the JAK2/STAT3 signaling pathway in the progression of renal
fibrosis. On the other hand, the results support the association
between NKILA and the JAK2/STAT3 signaling pathway. Therefore, this
suggests lncRNA NKILA has the potential to act as an independent
stimulator of fibrosis in HK-2 cells through selective activation
of the JAK2/STAT3 signaling pathway.
Although the present HK-2 cell model reveals the
involvement of lncRNA NKILA in EMT-like changes and its interaction
with the JAK2/STAT3 pathway during renal tubular EMT in
vitro, the absence of in vivo validation limits its
ability to fully represent the overall process of renal fibrosis.
In future, the fibrogenic role of lncRNA NKILA may be further
explored using in vivo experiments to evaluate its potential
as an independent fibrogenic factor.
Acknowledgements
Not applicable.
Funding
The present study was funded by the National Famous Traditional
Chinese Medicine Expert Inheritance Studio (grant no. 978022),
Tianjin Health Research Project (grant no. TJWJ2024QN071) and the
Scientific Research Project of the Administration of Traditional
Chinese Medicine of Hebei Province (grant no. T2025046).
Availability of data and materials
The data generated in the present study may be found
in the Genome Sequence Archive under BioProject number PRJCA024511,
dataset number: HRA006966; DAC number: HDAC003863,or at the
following https://ngdc.cncb.ac.cn/gsa-human/browse/HRA006966.
Authors' contributions
YH and YW designed the present study. YH, SY, JZ and
HL performed the experiments and collected the data. YH and SY
wrote the manuscript. YH, JZ and XZ were responsible for the data
analysis of western blot detection. YH and JZ were responsible for
CCK-8 data analysis and processing. YH and SY were responsible for
q-PCR data analysis, immunofluorescence data analysis and
processing, and high-throughput sequencing data visualization
processing. YH and SY were responsible for the production and
proofing of the pictures in the article. All authors read and
approved the final version of the manuscript. YH and SY confirm the
authenticity of all the raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Humphreys BD: Mechanisms of renal
fibrosis. Annu Rev Physiol. 80:309–326. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Liu Y, Bi X, Xiong J, Han W, Xiao T, Xu X,
Yang K, Liu C, Jiang W, He T, et al: MicroRNA-34a promotes renal
fibrosis by downregulation of klotho in tubular epithelial cells.
Mol Ther. 27:1051–1065. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bülow RD and Boor P: Extracellular matrix
in kidney fibrosis: More than just a scaffold. J Histochem
Cytochem. 67:643–661. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Salminen A: Increased immunosuppression
impairs tissue homeostasis with aging and age-related diseases. J
Mol Med (Berl). 99:1–20. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lv JC and Zhang LX: Prevalence and disease
burden of chronic kidney disease. Adv Exp Med Biol. 1165:3–15.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mack M: Inflammation and fibrosis. Matrix
Biol. 68–69. 106–121. 2018.PubMed/NCBI
|
|
7
|
GBD Chronic Kidney Disease Collaboration,
. Global, regional, and national burden of chronic kidney disease,
1990–2017: A systematic analysis for the Global Burden of Disease
Study 2017. Lancet. 395:709–733. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ali T and Grote P: Beyond the
RNA-dependent function of LncRNA genes. Elife. 9:e605832020.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bridges MC, Daulagala AC and Kourtidis A:
LNCcation: LncRNA localization and function. J Cell Biol.
220:e2020090452021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Nojima T and Proudfoot NJ: Mechanisms of
lncRNA biogenesis as revealed by nascent transcriptomics. Nat Rev
Mol Cell Biol. 23:389–406. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yang Z, Jiang S, Shang J, Jiang Y, Dai Y,
Xu B, Yu Y, Liang Z and Yang Y: LncRNA: Shedding light on
mechanisms and opportunities in fibrosis and aging. Ageing Res Rev.
52:17–31. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xu J, Bai J, Zhang X, Lv Y, Gong Y, Liu L,
Zhao H, Yu F, Ping Y, Zhang G, et al: A comprehensive overview of
lncRNA annotation resources. Brief Bioinform. 18:236–249.
2017.PubMed/NCBI
|
|
13
|
Fan XN, Zhang SW, Zhang SY and Ni JJ:
lncRNA_Mdeep: An Alignment-free predictor for distinguishing long
Non-coding RNAs from Protein-coding transcripts by multimodal deep
learning. Int J Mol Sci. 21:52222020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Alaaeldin R, Ali FEM, Bekhit AA, Zhao QL
and Fathy M: Inhibition of NF-kB/IL-6/JAK2/STAT3 pathway and
epithelial-mesenchymal transition in breast cancer cells by
azilsartan. Molecules. 27:78252022. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Bird L: lncRNA NKILA: A killer regulator.
Nat Rev Immunol. 18:666–667. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Dashti S, Ghafouri-Fard S, Esfandi F,
Oskooei VK, Arsang-Jang S and Taheri M: Expression analysis of
NF-κB interacting long noncoding RNAs in breast cancer. Exp Mol
Pathol. 112:1043592020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Soltanmoradi S, Tavakolpour V, Moghadasi
AN and Kouhkan F: Expression analysis of NF-κB-associated long
noncoding RNAs in peripheral blood mononuclear cells from
relapsing-remitting multiple sclerosis patients. J Neuroimmunol.
356:5776022021. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ghafouri-Fard S, Gholipour M, Abak A,
Mazdeh M, Taheri M and Sayad A: Expression analysis of
NF-κB-Related lncRNAs in Parkinson's disease. Front Immunol.
12:7552462021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Tian S, Yu Y, Huang H, Xu A, Xu H and Zhou
Y: Expression level and clinical significance of NKILA in human
cancers: A systematic review and Meta-analysis. Biomed Res Int.
2020:45403122020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li Y, Wang W, Chen K, Ma S and Wang J:
Influence of LncRNA NKILA on bloodstream infection of hypervirulent
klebsiella pneumoniae and its ability to induce delayed neutrophil
apoptosis. Evid Based Complement Alternat Med.
2021:61010782021.PubMed/NCBI
|
|
21
|
Herman AB, Tsitsipatis D and Gorospe M:
Integrated lncRNA function upon genomic and epigenomic regulation.
Mol Cell. 82:2252–2266. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Martin M: Cutadapt removes adapter
sequences from high-throughput sequencing reads. EMBnet J Bioinfo
Action. 17:10–12. 2011. View Article : Google Scholar
|
|
23
|
Langmead B and Salzberg SL: Fast
gapped-read alignment with Bowtie 2. Nat Methods. 9:357–359. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kim D, Langmead B and Salzberg SL: HISAT:
A fast spliced aligner with low memory requirements. Nat Methods.
12:357–360. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Pertea M, Pertea GM, Antonescu CM, Chang
TC, Mendell JT and Salzberg SL: StringTie enables improved
reconstruction of a transcriptome from RNA-seq reads. Nat
Biotechnol. 33:290–295. 2015. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Robinson MD, McCarthy DJ and Smyth GK:
edgeR: A Bioconductor package for differential expression analysis
of digital gene expression data. Bioinformatics. 26:139–140. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kong L, Zhang Y, Ye ZQ, Liu XQ, Zhao SQ,
Wei L and Gao G: CPC: Assess the protein-coding potential of
transcripts using sequence features and support vector machine.
Nucleic Acids Res. 35:W345–W349. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sun L, Luo H, Bu D, Zhao G, Yu K, Zhang C,
Liu Y, Chen R and Zhao Y: Utilizing sequence intrinsic composition
to classify protein-coding and long non-coding transcripts. Nucleic
Acids Res. 41:e1662013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Rauluseviciute I, Riudavets-Puig R,
Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, Lemma RB,
Lucas J, Chèneby J, Baranasic D, et al: JASPAR 2024: 20th
anniversary of the open-access database of transcription factor
binding profiles. Nucleic Acids Res. 52:D174–D182. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Li MT, Tang XH, Cai H, Zhang AH and Guo
ZY: Editorial: Molecular mechanism and therapeutic approach to
renal interstitial fibrosis. Front Med (Lausanne). 9:8799272022.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Herrera J, Henke CA and Bitterman PB:
Extracellular matrix as a driver of progressive fibrosis. J Clin
Invest. 128:45–53. 2018. View
Article : Google Scholar : PubMed/NCBI
|
|
33
|
Tan YT, Lin JF, Li T, Li JJ, Xu RH and Ju
HQ: LncRNA-mediated posttranslational modifications and
reprogramming of energy metabolism in cancer. Cancer Commun (Lond).
41:109–120. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Robinson EK, Covarrubias S and Carpenter
S: The how and why of lncRNA function: An innate immune
perspective. Biochim Biophys Acta Gene Regul Mech. 1863:1944192020.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Slatko BE, Gardner AF and Ausubel FM:
Overview of Next-generation sequencing technologies. Curr Protoc
Mol Biol. 122:e592018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Levy SE and Boone BE: Next-generation
sequencing strategies. Cold Spring Harb Perspect Med.
9:a0257912019. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Rego SM and Snyder MP: High throughput
sequencing and assessing disease risk. Cold Spring Harb Perspect
Med. 9:a0268492019. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Huang D, Chen J, Yang L, Ouyang Q, Li J,
Lao L, Zhao J, Liu J, Lu Y, Xing Y, et al: NKILA lncRNA promotes
tumor immune evasion by sensitizing T cells to activation-induced
cell death. Nat Immunol. 19:1112–1125. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Gupta SC, Awasthee N, Rai V, Chava S,
Gunda V and Challagundla KB: Long non-coding RNAs and nuclear
factor-κB crosstalk in cancer and other human diseases. Biochim
Biophys Acta Rev Cancer. 1873:1883162020. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Liu B, Sun L, Liu Q, Gong C, Yao Y, Lv X,
Lin L, Yao H, Su F, Li D, et al: A cytoplasmic NF-κB interacting
long noncoding RNA blocks IκB phosphorylation and suppresses breast
cancer metastasis. Cancer Cell. 27:370–381. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Singh A, Martinez-Yamout MA, Wright PE and
Dyson HJ: Interactions of a long noncoding RNA with domains of
NF-κB and IκBα: Implications for the inhibition of
Non-Signal-related phosphorylation. Biochemistry. 61:367–376. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Gai X and Li L: Overexpression of Long
Noncoding RNAs (lncRNA) NF-κβ-interacting long Noncoding RNA
(NKILA) in ankylosing spondylitis is correlated with transforming
growth factor β1 (TGF-β1), active disease and predicts length of
treatment. Med Sci Monit. 25:4244–4249. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Li JP, Li R, Liu X, Huo C, Liu TT, Yao J
and Qu YQ: A Seven Immune-related lncRNAs model to increase the
predicted value of lung adenocarcinoma. Front Oncol. 10:5607792020.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Xu G, Yang M, Wang Q, Zhao L, Zhu S, Zhu
L, Xu T, Cao R, Li C, Liu Q, et al: A novel prognostic prediction
model for colorectal cancer based on nine autophagy-Related long
noncoding RNAs. Front Oncol. 11:6139492021. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Jiang P, Han X, Zheng Y, Sui J and Bi W:
Long non-coding RNA NKILA serves as a biomarker in the early
diagnosis and prognosis of patients with colorectal cancer. Oncol
Lett. 18:2109–2117. 2019.PubMed/NCBI
|
|
46
|
Ebadi N, Ghafouri-Fard S, Taheri M,
Arsang-Jang S and Omrani MD: Expression analysis of inflammatory
response-associated genes in coronary artery disease. Arch Physiol
Biochem. 128:601–607. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Liu D and Shi X: Long non-coding RNA NKILA
inhibits proliferation and migration of lung cancer via IL-11/STAT3
signaling. Int J Clin Exp Pathol. 12:2595–2603. 2019.PubMed/NCBI
|
|
48
|
He B, Chen W, Zeng J, Tong W and Zheng P:
Long noncoding RNA NKILA transferred by astrocyte-derived
extracellular vesicles protects against neuronal injury by
upregulating NLRX1 through binding to mir-195 in traumatic brain
injury. Aging (Albany NY). 13:8127–8145. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Butturini E, Carcereri de Prati A and
Mariotto S: Redox regulation of STAT1 and STAT3 Signaling. Int J
Mol Sci. 21:70342020. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Butturini E, Boriero D, Carcereri de Prati
A and Mariotto S: STAT1 drives M1 microglia activation and
neuroinflammation under hypoxia. Arch Biochem Biophys. 669:22–30.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Fu XQ, Chou JY, Li T, Zhu PL, Li JK, Yin
CL, Su T, Guo H, Lee KW, Hossen MJ, et al: The JAK2/STAT3 pathway
is involved in the anti-melanoma effects of atractylenolide I. Exp
Dermatol. 27:201–204. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Li R, Sun N, Chen X, Li X, Zhao J, Cheng
W, Hua H, Fukatsu M, Mori H, Takahashi H, et al: JAK2(V617F)
Mutation Promoted IL-6 production and glycolysis via mediating PKM1
stabilization in macrophages. Front Immunol. 11:5890482020.
View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Chen WD, Zhang JL, Wang XY, Hu ZW and Qian
YB: The JAK2/STAT3 signaling pathway is required for inflammation
and cell death induced by cerulein in AR42J cells. Eur Rev Med
Pharmacol Sci. 23:1770–1777. 2019.PubMed/NCBI
|
|
54
|
Wang B, Liu T, Wu JC, Luo SZ, Chen R, Lu
LG and Xu MY: STAT3 aggravates TGF-β1-induced hepatic
epithelial-to-mesenchymal transition and migration. Biomed
Pharmacother. 98:214–221. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
You X, Jiang X, Zhang C, Jiang K, Zhao X,
Guo T, Zhu X, Bao J and Dou H: Dihydroartemisinin attenuates
pulmonary inflammation and fibrosis in rats by suppressing
JAK2/STAT3 signaling. Aging (Albany NY). 14:1110–1127. 2022.
View Article : Google Scholar : PubMed/NCBI
|