Introduction
Cancer poses a global health threat, and 8–18% of
individuals with cancer have diabetes as a concurrent health issue
(1,2). Hyperglycemia, characterized by
abnormally high blood glucose levels, is a crucial diagnostic
marker for diabetes, and serves a notable role in both the
initiation and progression of cancer (3). Numerous epidemiological
investigations have revealed that individuals with diabetes are at
a heightened risk of various types of cancer, including liver,
pancreatic, stomach, kidney, breast and colorectal cancer (CRC)
(4,5).
Several studies have revealed that hyperglycemia,
particularly that associated with type II diabetes mellitus, is an
independent risk factor for colon cancer, facilitating colon cancer
incidence, and affecting its recurrence, metastasis and prognosis
(6,7). Our previous study revealed that
patients with colon cancer and diabetes had a higher risk of
recurrence after surgery than those without diabetes (8). However, the mechanisms by which
hyperglycemia mediates poor prognosis in CRC remain unclear.
Exploring the link between hyperglycemia and colon cancer will
improve the understanding of cancer pathogenesis, while offering a
more valuable guide for personalized treatment of colon tumors in
diabetic patients.
Tribble (TRIB) pseudokinases control multiple
aspects of eukaryotic cell biology and have evolved unique features
that distinguish them from other protein kinases (9). The TRIB family comprises TRIB1, TRIB2
and TRIB3, and it has been suggested that TRIB3 serves a crucial
role in diabetes, malignancy and cardiovascular diseases (10,11).
In previous studies, TRIB3 protein levels have been reported to be
markedly elevated in gastric, hepatocellular and lung cancer
(12–14). However, the role of TRIB3 in colon
cancer remains unclear, and its potential to promote colon cancer
progression in high-glucose environments is not well
understood.
The present study aimed to investigate the
expression of TRIB3 in CRC cell lines under high-glucose conditions
and in CRC tissues from patients with diabetes. Furthermore, the
study aimed to explore whether TRIB3 regulates the EMT program via
the PI3K/AKT pathway to mediate the malignant proliferation and
invasion of CRC cells induced by high glucose, and whether
suppressing TRIB3 expression could hinder high glucose-induced
tumor cell biological behaviors. The findings may aid in the
determination of whether targeting TRIB3 could serve as a potential
novel therapeutic strategy for diabetic patients with CRC.
Materials and methods
Cell culture and reagents
The human colorectal adenocarcinoma cell lines
HCT116, SW480, DLD-1, and LoVo, and 293T cell lines were purchased
from the Academy of Medical Sciences. 293T cells were cultured in
DMEM (cat. no. 01-052-1ACS; glucose content, 25 mM; Biological
Industries; Sartorius AG); DLD-1 cells were cultured in RPMI-1640
medium (cat. no. PM150110; Procell Life Science & Technology
Co., Ltd.); LoVo cells were cultured in Ham's F-12K medium (cat.
no. PM150910; Procell Life Science & Technology Co., Ltd.), and
HCT116 and SW480 cell lines were cultured in DMEM with different
glucose concentrations (including 10, 15, 20, 25, 30, 35 and 40
mM). Media containing different glucose concentrations were
prepared by mixing low-glucose DMEM (cat. no. 01-051-1ACS; glucose
content, 5 mM; Biological Industries; Sartorius AG) with a
condensed glucose solution (1 mM). All culture media contained 10%
fetal bovine serum (FBS; cat. no. FSS500; Shanghai ExCell Biology,
Inc.) and 1% penicillin-streptomycin compound (cat. no. C0222;
Beyotime Biotechnology). All cells were cultivated in a humidified
incubator containing 5% CO2 at 37°C. For experimental
treatment, LY294002 (cat. no. S1105; Selleck Chemicals) was diluted
with high-glucose DMEM to a final working concentration of 10 µM,
and CRC or transfected CRC cells were treated with this working
solution at 37°C for 24 h.
Short hairpin RNA (shRNA)-mediated
knockdown of TRIB3
The human TRIB3 shRNA lentiviral vector
[YSH-LV001-hTRIB3(shRNA), EGFP/puro] and the control lentiviral
vector (YSH-LV001-Ctrl) were purchased from Ubigene. A
third-generation lentiviral packaging system was used. Lentiviruses
were produced by transfecting 293T cells in 10 cm dishes with a
total of 20 µg plasmids encoding psPAX2, pMD2.G and lentiviral
vectors at a ratio of transfer plasmid/packaging plasmid/envelope
plasmid of 4:3:1. Transfection was performed using
Lipofectamine® 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.) for 6 h at 37°C in a humidified 5% CO2
incubator. Lentiviral supernatants were collected at 48 and 72 h
post-transfection, filtered through a 0.45-µm filter, and
concentrated if needed. CRC cells were then transduced with
lentivirus particles at a multiplicity of infection of 10 in the
presence of 5 µg/ml polybrene (cat. no. C0351; Beyotime
Biotechnology) for 24 h before the medium was replaced. The
infected CRC cells were cultured for an additional 72 h before
puromycin selection or subsequent experiments. Stably transduced
cells were selected with 2 µg/ml puromycin (cat. no. ST551;
Beyotime Biotechnology) and subsequently maintained in medium
containing 1 µg/ml puromycin.
Transwell migration and invasion
assays
Prior to the migration and invasion assays, HCT116,
SW480, DLD1 and LoVo cell lines were subjected to TRIB3 knockdown,
and HCT116 and SW480 cells were treated with LY294002 as
aforementioned. Cells were cultured under low glucose (10 mM),
control glucose (25 mM) and high glucose (40 mM) conditions
according to the corresponding experimental requirements. For the
migration assay, 5×104 cells/well suspended in
serum-free medium were inoculated into the upper chamber of a
24-well Transwell system (8.0 µm pore polycarbonate membrane). For
invasion assays, the upper chambers were precoated with a 1:8
mixture of reduced growth factor Matrigel (Corning, Inc.) and PBS,
followed by incubation at 37°C for 2 h before cell inoculation. The
lower chambers were filled with complete medium containing 10% FBS
as a chemoattractant. After incubation at 37°C for 24 h, the
migrating cells at the bottom were fixed with 4% paraformaldehyde
at room temperature for 15 min, followed by staining with 0.1%
crystal violet at room temperature for 10–15 min. Using a light
microscope, 3–5 fields/wells were selected for cell counting.
Nuclear protein isolation
For nuclear protein isolation, CRC cells cultured
under normal or high glucose conditions were harvested by scraping
and centrifugation at 1,000 × g for 5 min at 4°C. Nuclear proteins
were extracted using a Nuclear and Cytoplasmic Protein Extraction
Kit (cat. no. P0027; Beyotime Biotechnology). Briefly, cell pellets
were resuspended in ice-cold Cytoplasmic Extraction Buffer A
supplemented with 1 mM phenylmethylsulfonyl fluoride (PMSF),
vortexed thoroughly, and incubated on ice for 15 min. After adding
Cytoplasmic Extraction Buffer B and brief vortexing, samples were
centrifuged at 12,000 × g for 5 min at 4°C, and the cytoplasmic
fraction was discarded. The nuclear pellet was resuspended in
ice-cold Nuclear Extraction Buffer supplemented with 1 mM PMSF,
incubated on ice for 30 min with vigorous vortexing for 15 sec
every 2 min to lyse the nuclear membrane. Finally, nuclear lysates
were centrifuged at 12,000 × g for 15 min at 4°C, and the
supernatant was collected and stored at −80°C.
Protein extraction and western
blotting
Whole-cell protein extraction was performed using
RIPA lysis buffer (cat. no. R00010; Beijing Solarbio Science &
Technology Co., Ltd.) and protein concentration was quantified
using a BCA protein detection kit (Thermo Fisher Scientific, Inc.).
Subsequently, proteins (40 µg/lane) were denatured and separated by
SDS-PAGE on 10% gels. The proteins were then transferred onto PVDF
membranes (MilliporeSigma) using a semi-dry method at a current of
200 mA for a duration of 1.5 h. Membranes were blocked with 8%
non-fat milk at room temperature (25°C) for 1 h with constant
gentle shaking. After overnight incubation with primary antibodies
at 4°C, the membranes were incubated for 1 h at room temperature
with horseradish peroxidase (HRP)-conjugated secondary goat
anti-rabbit or anti-mouse-polyclonal immunoglobulin G antibodies.
Signals were visualized using an enhanced chemiluminescence kit
(Applygen Technologies, Inc.). For semi-quantitative analysis of
the results, ImageJ 1.46r software (National Institutes of Health)
was used. Detailed information regarding all antibodies used in the
present study is provided in Table
SI.
RNA extraction and reverse
transcription-quantitative (RT-qPCR)
Total RNA was extracted from the cells using
TRIeasy™ reagent (cat. no. 10606ES60; Shanghai Yeasen Biotechnology
Co., Ltd.) according to the manufacturer's instructions. Utilizing
1 µg total RNA, cDNA was synthesized in a 20-µl RT reaction using
Hifair® II 1st Strand cDNA Synthesis SuperMix (cat. no.
11137ES60; Shanghai Yeasen Biotechnology Co., Ltd.) according to
the manufacturer's protocol. To assess the mRNA expression levels
of TRIB3 and EMT markers, qPCR analysis was conducted using a 7500
Sequence Detection System (Applied Biosystems; Thermo Fisher
Scientific, Inc.) and the SYBR Green Premix (cat. no. 11203ES;
Shanghai Yeasen Biotechnology Co., Ltd.) under the following
conditions: Initial denaturation at 95°C for 30 sec, followed by 40
cycles of 95°C for 10 sec and 60°C for 30 sec, with a subsequent
melting curve analysis. The primer sequences are listed in Table SII. All experiments were conducted
in triplicate to ensure consistency and reliability. mRNA
expression levels were quantified using the 2−ΔΔCq
method after normalization to β-actin (15).
Cell proliferation assay
Cells were seeded in 96-well plates at a density of
5×103 cells in 100 µl medium/well 1 day before the
experiment. At different time points (0, 6, 24 and 48 h), medium
containing 10% Cell Counting Kit-8 (CCK-8; cat. no. CK001-500T;
Beijing LABLEAD Trading Co., Ltd.) was added to each well and the
cells were cultured at 37°C for 1 h. OD was measured at 450 nm
using a spectrophotometer.
Immunofluorescence
Special glass cover slides (cat. no. abs7027; Absin
Bioscience, Inc.) for cell immunofluorescence were placed in
24-well plates (cat. no. 801006; NEST®; Wuhan NEST
Biotechnology Co., Ltd.). SW480 and HCT116 colon cancer cells were
seeded at a density of 3×104 cells/well on the glass
coverslips. After culturing for 24 h until the cells reached 70%
confluence, they were fixed with 4% formaldehyde in PBS for 15 min
at room temperature. After blocking with 2% bovine serum albumin
(cat. no. 11021029; Gibco; Thermo Fisher Scientific, Inc.) in PBS
(PBA) at room temperature for 1 h, the cells were incubated with
primary antibodies against E-cadherin (1:150; cat. no. 60335-1-Ig;
Proteintech Group, Inc.), vimentin (1:100; cat. no. 10366-1-AP;
Proteintech Group, Inc.) and TRIB3 (1:50; cat. no. ab137526; Abcam)
in 2% PBA overnight at 4°C. The cells were then incubated with
corresponding secondary antibodies, including FITC-conjugated
anti-rabbit (cat. no. SA00003-2; Proteintech Group, Inc.),
TRITC-conjugated anti-rabbit (cat. no. SA00007-2; Proteintech
Group, Inc.), FITC-conjugated anti-mouse (cat. no. SA00003-1;
Proteintech Group, Inc.) and TRITC-conjugated anti-mouse (cat. no.
SA00007-1; Proteintech Group, Inc.), and counterstained with DAPI
(cat. no. ZLI-9557; Beijing Zhongshan Jinqiao Biotechnology Co.,
Ltd.). Images were captured and analyzed using fluorescence
microscopy.
Immunohistochemistry
Pathological tissues were obtained from the surgical
samples of patients with colon cancer at Beijing Chaoyang Hospital
affiliated with Capital Medical University (Beijing, China) between
January 2019 and December 2021. The inclusion criteria included
pathologically confirmed colon cancer, with complete
clinicopathological data and no preoperative neoadjuvant therapy;
the exclusion criteria included concurrent other malignancies,
severe systemic diseases, poor tissue quality or lost follow-up.
The study was approved by the Ethics Committee of Beijing Chaoyang
Hospital affiliated with Capital Medical University. Tumor staging
was determined according to the TNM staging system, and the
detailed clinicopathological information of the patients is
provided in Table SIII (16). Tissues were fixed in 4% formalin
for 24–48 h at room temperature and then embedded in paraffin at
58–60°C for ~10 min. Tissue specimens, each at a thickness of 4 µm,
were then subjected to deparaffinization and hydration.
Deparaffinization was performed with xylene (three changes, 5 min
each), followed by sequential hydration with a gradient ethanol
series (100, 95, 85 and 75%, 3 min each) and a final rinse with
distilled water. Subsequently, heat-induced epitope retrieval was
performed in a pressure cooker with 0.01 mol/l citrate buffer (pH
6.0) for 30 min. The sections were then treated with 3% hydrogen
peroxide for 15 min at room temperature to block endogenous
peroxidase activity, followed by blocking with 5% goat serum (cat.
no. 16210064; Gibco; Thermo Fisher Scientific, Inc.) for 30 min at
room temperature. Subsequently, the sections were incubated
overnight at 4°C with TRIB3 antibody (1:400; cat. no. ab137526;
Abcam), followed by incubation with HRP-conjugated goat anti-rabbit
secondary antibody (1:500; cat. no. SA00001-2; Proteintech Group,
Inc.) for 1 h at room temperature. Immunolabeling was performed
using a DAB detection kit (cat. no. ZLI-9019; Beijing Zhongshan
Jinqiao Biotechnology Co., Ltd.). Following a 10-mi rinse in water,
the sections were re-stained with 0.5% hematoxylin (cat. no.
ZLI-9609; OriGene Technologies, Inc.) for 1.5 min at room
temperature. Immunohistochemical signals were captured using an
optical microscope (Olympus Corporation). Staining intensity was
scored as 0 (negative), 1 (weak), 2 (moderate) and 3 (strong). The
percentage of positive cells was scored as 0 (0%), 1 (1–25%), 2
(26–50%), 3 (51–75%) and 4 (76–100%). The final IHC score was
calculated by multiplying the intensity score by the percentage
score, ranging from 0 to 12.
In vivo subcutaneous tumor formation
assay
Cell suspensions of HCT116/Ctrl and HCT116/shTRIB3
cells were prepared, with the density adjusted to 5×107
cells/ml. Briefly, 5-week-old female BALB/c nude mice (weight:
18–22 g; n=5/subgroup; total, 20 mice; no significant difference in
body weight among groups before the experiment) were purchased from
Beijing Vital River Laboratory Animal Technology Co., Ltd. The mice
were divided into four groups to be subcutaneously injected with
tumor cells into the right inguinal region: Two groups injected
with HCT116/Ctrl cells and two groups injected with HCT116/shTRIB3
cells. Each mouse received a 100-µl cell suspension, corresponding
to 5×106 cells/mouse. The HCT116/Ctrl and HCT116/shTRIB3
groups were each divided into two subgroups receiving either normal
drinking water or 30% high-sugar water [as recommended in multiple
studies (17,18)]. Mice were housed in cages with a
controlled temperature (22±2°C) and relative humidity (40–60%)
under a 12-h light/dark cycle throughout the study. All mice had
unrestricted access to a standard diet (crude protein 16%, crude
fat 4%, crude fiber 12% and ash 8%) and had free access to normal
or 30% high-sugar water as assigned. Tumor size was measured every
5 days. Tumor volume was calculated using the formula: Volume
(mm3)=(length × width2)/2.
Mice were euthanized by CO2 inhalation 21
days after tumor implantation. The maximum tumor diameter did not
exceed 12 mm, the tumors showed no signs of ulceration and mice
exhibited good general health with no signs of cachexia. Briefly,
CO2 was introduced at a displacement rate of 30% chamber
volume/min, in accordance with the AVMA Guidelines for the
Euthanasia of Animals (19).
Confirmation of death required verifying that the mouse was
motionless and exhibited no respiratory activity; after turning off
the CO2, the mice were monitored for an additional 2–3
min to verify death.
The Cancer Genome Atlas (TCGA)
data
The mRNA expression data of TRIB3 (RNAseq) from 430
patients with CRC in TCGA database (cohort: TCGA CRC) were
collected from UCSC Xena (http://xena.ucsc.edu/) to examine the differences in
TRIB3 expression between tumor and adjacent non-tumor tissues. To
investigate the correlation between TRIB3 and the PI3K/AKT pathway,
the ‘ggcorrplot’ R package (version 0.1.4.1; http://www.sthda.com/english/wiki/ggcorrplot-visualization-of-a-correlation-matrix-using-ggplot2)
was used to plot the correlation heat map, and the cor() function
was used to calculate the Spearman correlation coefficient. The R
software used in the present study was R version 4.5.1 (R
Foundation for Statistical Computing; http://www.r-project.org/).
Statistical analysis
Data analysis was performed using GraphPad Prism
software (version 6.0; Dotmatics). Unpaired student's t-test was
used to assess statistical differences for parametric analyses. For
non-parametric comparisons, the Mann-Whitney U test was used.
Multiple group comparisons were analyzed one-way ANOVA, with
inter-group differences further delineated by Tukey's post hoc
test. Quantitative data are presented as the mean ± standard
deviation, with all experiments conducted in triplicate to ensure
consistency and reliability. P<0.05 was considered to indicate a
statistically significant difference.
Results
High glucose activates the malignant
behavior of CRC cells
To validate whether high glucose levels promote
tumor development, a series of in vitro assays were
conducted to examine the effects of high-glucose levels on CRC cell
functions. Emerging research has revealed that when selecting 2 g/l
(11.1 mM) and 5 g/l (27.8 mM) as low- and high-glucose
concentrations, respectively, two standards are satisfied: i) Cell
viability is >80% and ii) the apoptosis rate is <10%
(20). To expand the concentration
differences, in the current study, 10 and 40 were we selected as
the low- and high-glucose concentrations, respectively.
Colon tumor cells were cultured in gradient glucose
conditions (10–40 mM) for 48 h and the proliferative ability of the
two CRC cell lines, SW480 and HCT116, was assessed using the CCK-8
assay. Compared with in cells treated with low glucose (10 mM
glucose), cells treated with high glucose (40 mM glucose) exhibited
a significantly higher proliferative potential (Fig. 1A). To investigate the effect of
high-glucose levels on the migratory and invasive potential of CRC
cells, Transwell assays were performed. The findings revealed that
CRC cells exposed to high-glucose levels exhibited increased
invasion and migration compared with those exposed to low-glucose
levels (Fig. 1B). Alterations in
cancer cell migration and invasion capacity are commonly linked to
epithelial-mesenchymal transition (EMT). Therefore, the expression
levels of EMT-related markers were detected, including the
epithelial markers E-cadherin and ZO1, and the mesenchymal markers
N-cadherin, Vimentin and Snail. RT-qPCR results showed a
significant increase in the expression levels of N-cadherin,
Vimentin and Snail, and a reduction in the expression of E-cadherin
and ZO1 in CRC cells in response to a high-glucose concentration
(Fig. 1C). Subsequently, these
findings were corroborated by western blotting and
immunofluorescence staining, which consistently obtained similar
results (Fig. 1D and E). These
findings indicated that high-glucose levels may induce
tumorigenesis and EMT in CRC cells.
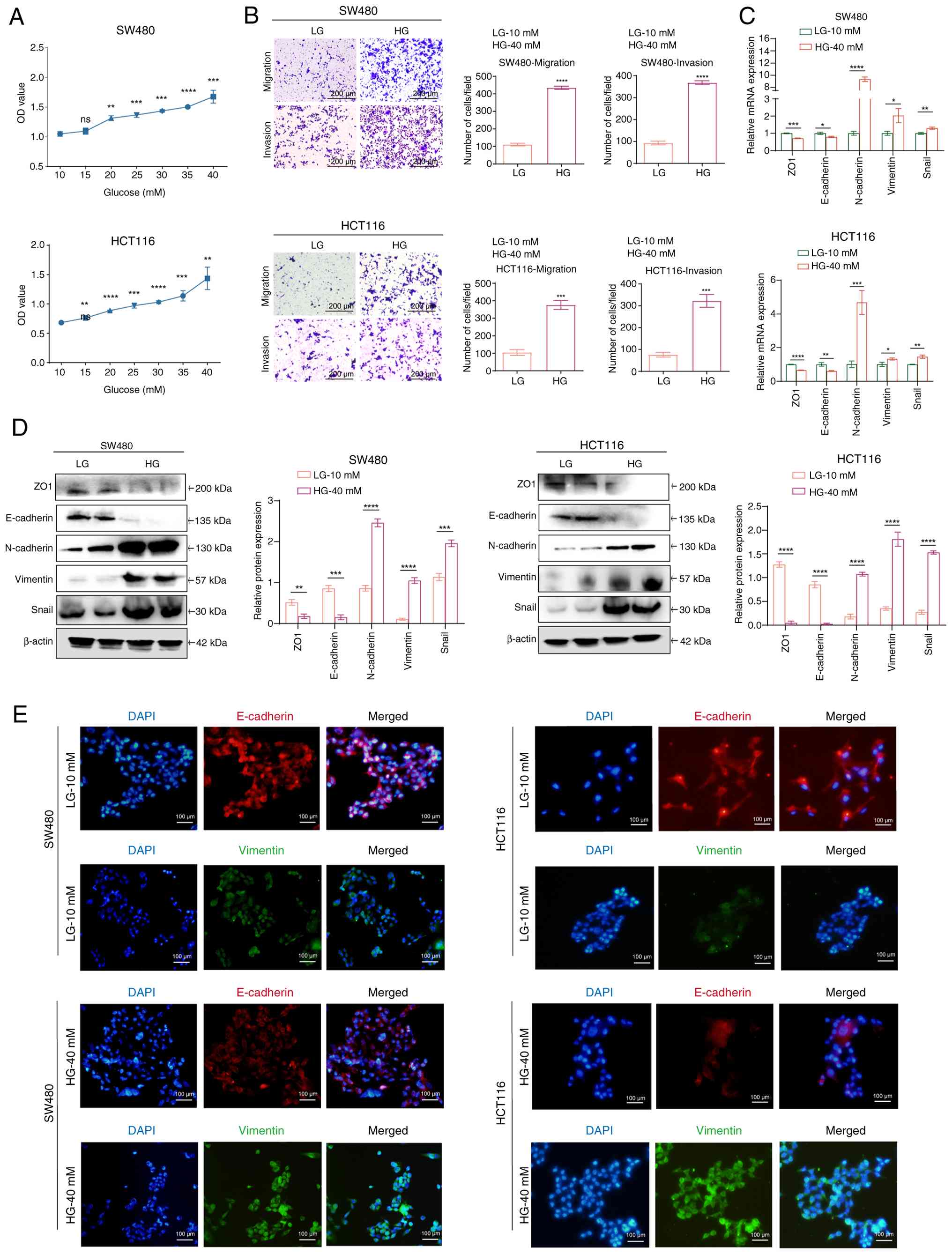 | Figure 1.HG induces tumorigenesis and EMT in
CRC cells. (A) Cell Counting Kit-8 assay was performed to examine
the effects of a gradient glucose concentration within 48 h on the
proliferation of two CRC cell lines, SW480 and HCT116. **P<0.01,
***P<0.001, ****P<0.0001 vs. 0 mM glucose group. (B)
Transwell assays were performed to evaluate the effects of LG and
HG on the migration and invasion of CRC cells. ***P<0.001,
****P<0.0001 vs. LG group. (C) Reverse
transcription-quantitative PCR examination of EMT markers after CRC
cells were treated with LG or HG for 48 h. (D) Western blotting was
utilized to evaluate the levels of EMT marker proteins following
treatment of CRC cells with either LG or HG for 48 h. *P<0.05,
**P<0.01, ***P<0.001, ****P<0.0001 vs. LG group. (E)
Immunofluorescence staining of E-cadherin and Vimentin in two CRC
cell lines treated with LG or HG for 48 h. For all bar-plots, data
are presented as the mean ± SEM, n=3. CRC, colorectal cancer; EMT,
epithelial-mesenchymal transition; HG, high glucose; LG, low
glucose; ns, not significant. |
High glucose induces TRIB3 expression
in CRC
To verify the relationship between high-glucose
levels and TRIB3, the mRNA and protein expression levels of TRIB3
were measured in CRC cells treated with the indicated glucose
concentrations. RT-qPCR results showed that with an increase in
glucose concentration, the expression of TRIB3 exhibited a gradual
upward trend (Fig. 2A). Western
blotting also revealed an apparent increase in TRIB3 protein levels
after treatment with HG compared with LG; in response to a 40-mM
glucose concentration, the expression of TRIB3 was approximately
three times higher than that in the group treated with 10 mM
glucose (Fig. 2B).
Immunofluorescence staining provided corroborative evidence
(Fig. 2C). As shown in the
representative immunohistochemistry images, TRIB3 protein levels in
patients with diabetes and CRC were higher than those in patients
with CRC without diabetes (Fig.
2D). Notably, TRIB3 expression was markedly elevated at the
cellular level under conditions of high-glucose exposure, and this
upregulation was also prominent in patients with CRC and concurrent
diabetes.
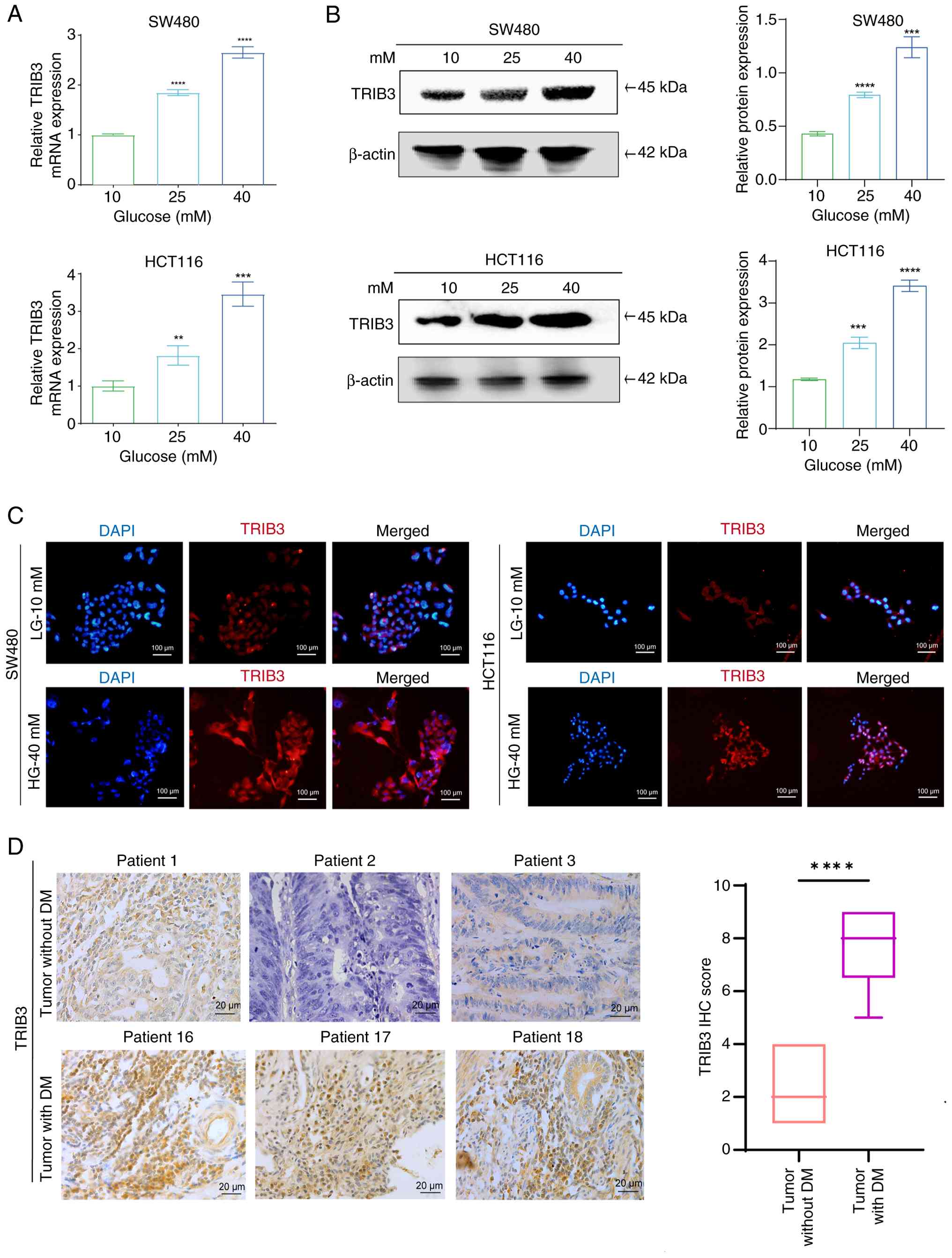 | Figure 2.HG activates TRIB3 expression in CRC.
(A) Reverse transcription-quantitative PCR was conducted to analyze
the mRNA expression levels of TRIB3 in CRC cells subjected to
varying glucose concentrations. (B) Western blot analysis to detect
the protein expression levels of TRIB3 in CRC cells treated with a
gradient of glucose concentrations. **P<0.01, ***P<0.001,
****P<0.0001 vs. 10 mM glucose group. (C) Immunofluorescence
staining of TRIB3 expression in CRC cells exposed to HG or LG
concentrations for 48 h. (D) Representative IHC images of TRIB3
protein levels from patients with CRC with or without DM stratified
by IHC score. ****P<0.0001. Data are presented as the (A-C) the
mean ± SEM, n=3; or (D) median (interquartile range), CRC,
colorectal cancer; DM, diabetes mellitus; HG, high glucose; IHC,
immunohistochemistry; LG, low glucose; TRIB3, tribbles 3. |
TRIB3 knockdown suppresses the
proliferation, invasion and EMT process of CRC cells
The expression levels of TRIB3 were detected in
SW480, HCT116, DLD-1 and LoVo cells (Figs. 3A and S1A). To verify whether TRIB3 mediates
the malignant phenotype of CRC cells, stable TRIB3 knockdown clones
were constructed: SW480/shTRIB3, HCT116/shTRIB3, DLD-1/shTRIB3 and
LoVo/shTRIB3, and knockdown efficiency was confirmed by western
blotting (Figs. 3B and S1B). Notably, CRC cells transduced with
shTRIB3 exhibited significantly decreased proliferation, as
determined by the CCK-8 assay (Figs.
3C and S1C). To investigate
the influence of TRIB3 on the migratory and invasive potential of
CRC cells, Transwell assays were performed and the results revealed
that shTRIB3-transduced CRC cells exhibited decreased invasion and
migration compared with that in the control groups (Figs. 3D and S1D). Subsequently, the effects of TRIB3
knockdown on the expression levels of EMT markers were assessed
using RT-qPCR. Compared with in the control group, an increase in
the expression levels of E-cadherin and ZO1, alongside a decrease
in the levels of N-cadherin and Vimentin, were observed in
shTRIB3-transduced CRC cells (Fig.
3E). Furthermore, similar results were obtained by western
blotting (Fig. 3F). These results
suggest that TRIB3 has the potential to promote the malignant
behavior of CRC cells and drive EMT in tumor cells. However,
whether TRIB3 serves as a key molecule driving the augmented
malignant capabilities in the high-glucose environment of CRC
remains unknown.
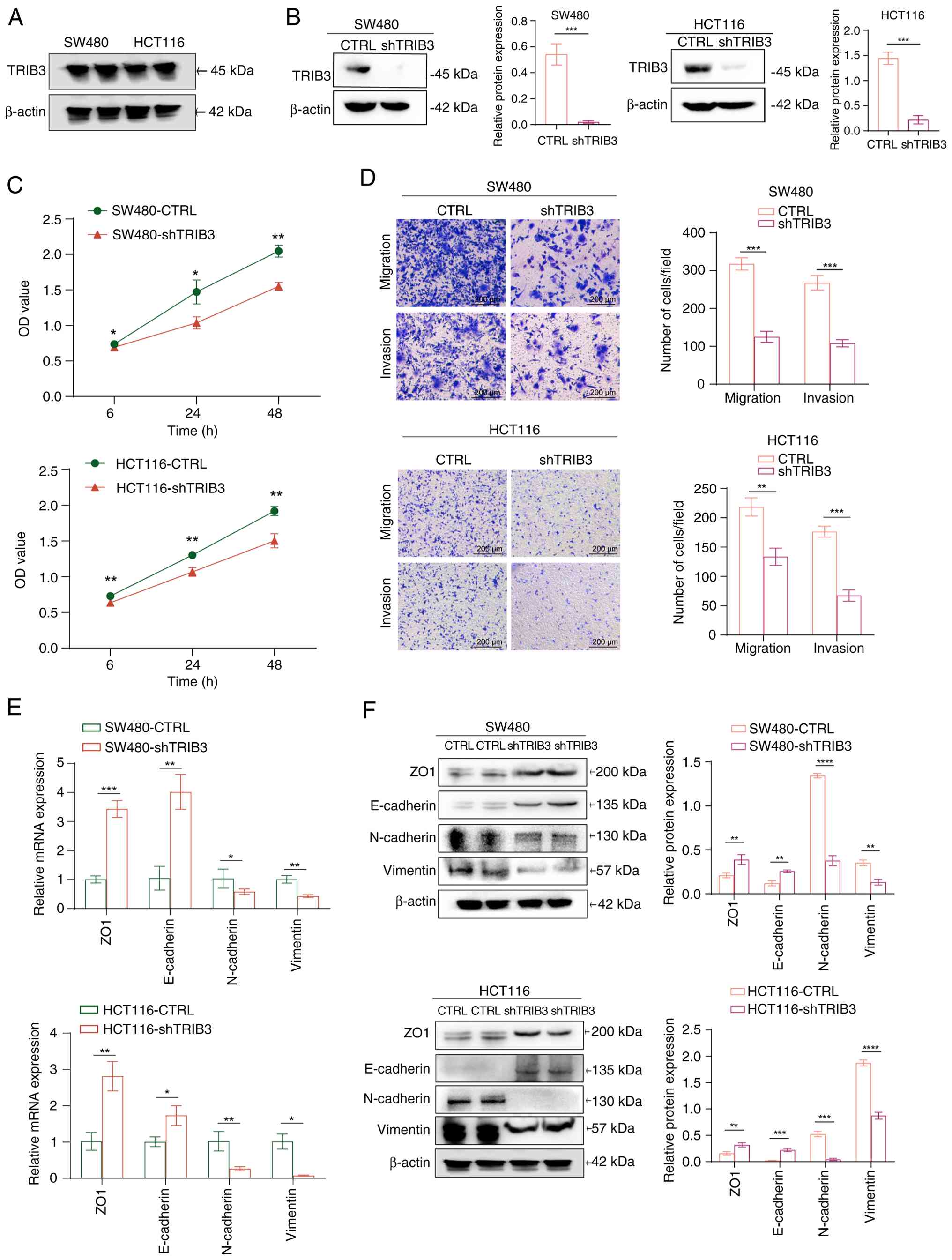 | Figure 3.TRIB3 mediates the proliferation and
invasiveness of CRC cells by remodeling the EMT process. (A)
Western blotting was performed to evaluate the protein expression
levels of TRIB3 in the CRC cell lines SW480 and HCT116. (B) Western
blotting validation of TRIB3 knockdown in two cancer cell lines.
***P<0.001. (C) Cell counting kit-8 assay was used to
investigate the impact of TRIB3 knockdown on the proliferation of
two CRC cell lines. *P<0.05, **P<0.01, vs. control group. (D)
Transwell assays were performed to evaluate the changes in cell
migration and invasion induced by TRIB3 knockdown in two cancer
cell lines. (E) Reverse transcription-quantitative PCR was used to
examine the impact of TRIB3 knockdown on the mRNA expression levels
of EMT markers. (F) Western blotting was utilized to evaluate the
impact of TRIB3 knockdown on the protein expression levels of EMT
markers. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
For all bar-plots, data are presented as the mean ± SEM, n=3. CRC,
colorectal cancer; CTRL, control; EMT, epithelial-mesenchymal
transition; sh, short hairpin; TRIB3, tribbles 3. |
TRIB3 promotes high glucose-induced
malignancy in CRC cells
To validate the role of TRIB3 in promoting
malignancy under high-glucose conditions, a CCK-8 assay was
performed. The results demonstrated that glucose-induced CRC cell
proliferation was significantly reduced by TRIB3 knockdown
(Fig. 4A). In addition, Transwell
assay results showed that TRIB3 knockdown reversed the phenotype of
migration and invasion induced by high glucose in CRC cells
(Fig. 4B). To validate whether the
knockdown of TRIB3 could alleviate high glucose-induced EMT in CRC
cells, RT-qPCR was used to assess the expression levels of EMT
markers; the results revealed that the knockdown of TRIB3 reversed
the trend of HG-induced exacerbation of EMT, as indicated by the
expression levels of ZO-1 and N-cadherin (Fig. 4C). Furthermore, these observations
were corroborated by western blot analysis, which revealed that
inhibiting TRIB3 in a high-glucose environment resulted in a marked
increase in the expression levels of the epithelial marker ZO1.
Furthermore, the expression levels of mesenchymal markers,
including N-cadherin and Vimentin, were markedly decreased
(Fig. 4D).
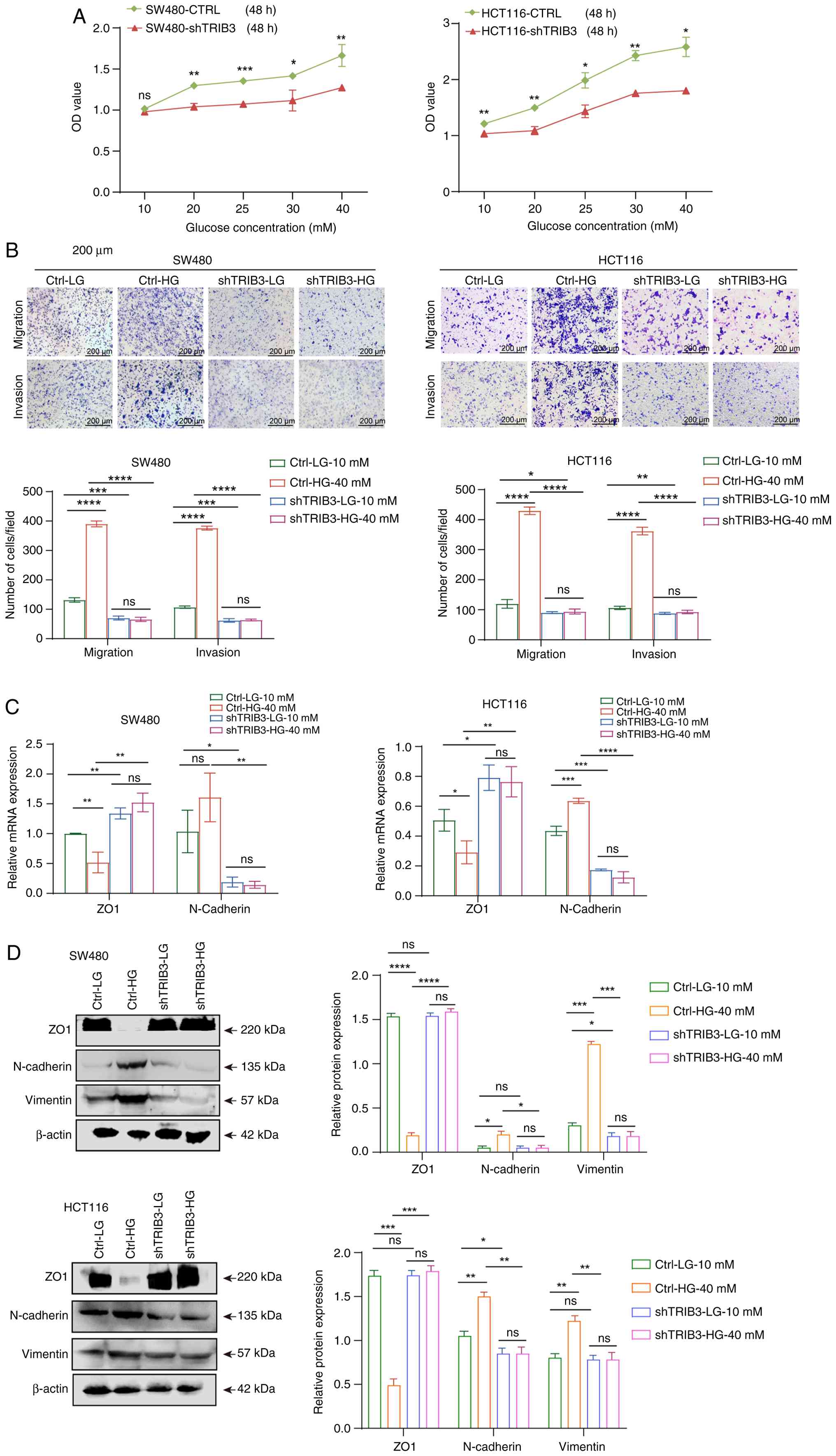 | Figure 4.Knockdown of TRIB3 alleviates
HG-induced malignancy and EMT in CRC cells. (A) Cell counting kit-8
assay was performed to evaluate the effects of TRIB3 knockdown on
CRC cell proliferation induced by different glucose concentrations.
*P<0.05, **P<0.01, ***P<0.001 vs. control group. (B)
Transwell assays were used to assess the effects of TRIB3 knockdown
on the invasive and migratory capabilities of CRC cells under HG
conditions. (C) Reverse transcription-quantitative PCR was
performed to evaluate the effects of shTRIB3 on HG-induced EMT in
two cancer cell lines (SW480 and HCT116). (D) Western blotting was
used to assess the changes in the expression of EMT molecules upon
TRIB3 knockdown in a HG environment. *P<0.05, **P<0.01,
***P<0.001, ****P<0.0001. For all bar-plots, data are
presented as the mean ± SEM, n=3. CRC, colorectal cancer; CTRL,
control; EMT, epithelial-mesenchymal transition; HG, high glucose;
LG, low glucose; ns, not significant; sh, short hairpin; TRIB3,
tribbles 3. |
To further validate the aforementioned conclusions
in vivo, HCT116 control and HCT116 shTRIB3 cells were
subcutaneously inoculated into mice. The HCT116 control group was
divided into two subgroups receiving either normal drinking water
or 30% high-sugar water, with the same treatment applied to the
HCT116 shTRIB3 group. Notably, subcutaneous tumors formed
exclusively in the HCT116 groups, whereas almost no tumors were
observed in the TRIB3-knockdown groups (Fig. S2). Moreover, the subcutaneous
tumor formation in the high-sugar drinking group was significantly
enhanced compared with that that in the normal drinking water
group. Notably, these in vivo data confirmed that TRIB3
knockdown can markedly impair cell tumorigenicity, which is also
consistent with the in vitro experimental results,
confirming the driving role of TRIB3 in tumor formation. However,
its significant upregulation under high-glucose conditions further
exacerbates its regulatory impact on the malignant progression of
tumors (Fig. S2). These findings
illustrate that TRIB3 serves a pivotal role in amplifying the
proliferative and invasive capabilities of CRC cells in a
high-glucose environment. However, the mechanism by which TRIB3
activates the EMT remains unclear.
TRIB3 induces activation of the
PI3K/AKT signaling pathway under high-glucose conditions in CRC
cells
Based on TCGA-Colon Adenocarcinoma dataset, it was
revealed that TRIB3 was closely associated with the PI3K/AKT
pathway in patients with CRC, as evidenced by the increased
expression of PI3K/AKT pathway-related activation markers (AREG,
AKT2, MET, PIK3R2, FGFR2, and EREG) (Fig. 5A). Since the PI3K/AKT pathway is
crucial for cancer cell proliferation and metastasis, the
phosphorylation of proteins in the PI3K pathway was evaluated in
cells under normal glucose conditions and in shTRIB3 CRC cells by
western blotting. The western blotting findings revealed a marked
reduction in PI3K, AKT and mTOR phosphorylation levels in SW480 and
HCT116 cell lines upon TRIB3 knockdown compared with in the control
groups, without notably altering the total PI3K, AKT and mTOR
expression levels (Fig. 5B).
Furthermore, an apparent increase in p-AKT, p-PI3K and p-mTOR
levels was detected after treatment with a high-glucose
concentration compared with that in the low-glucose concentration
group, whereas TRIB3 knockdown significantly inhibited this
phosphorylation (Fig. 5C). In
summary, TRIB3 may activate the PI3K/AKT pathway in response to
high glucose levels.
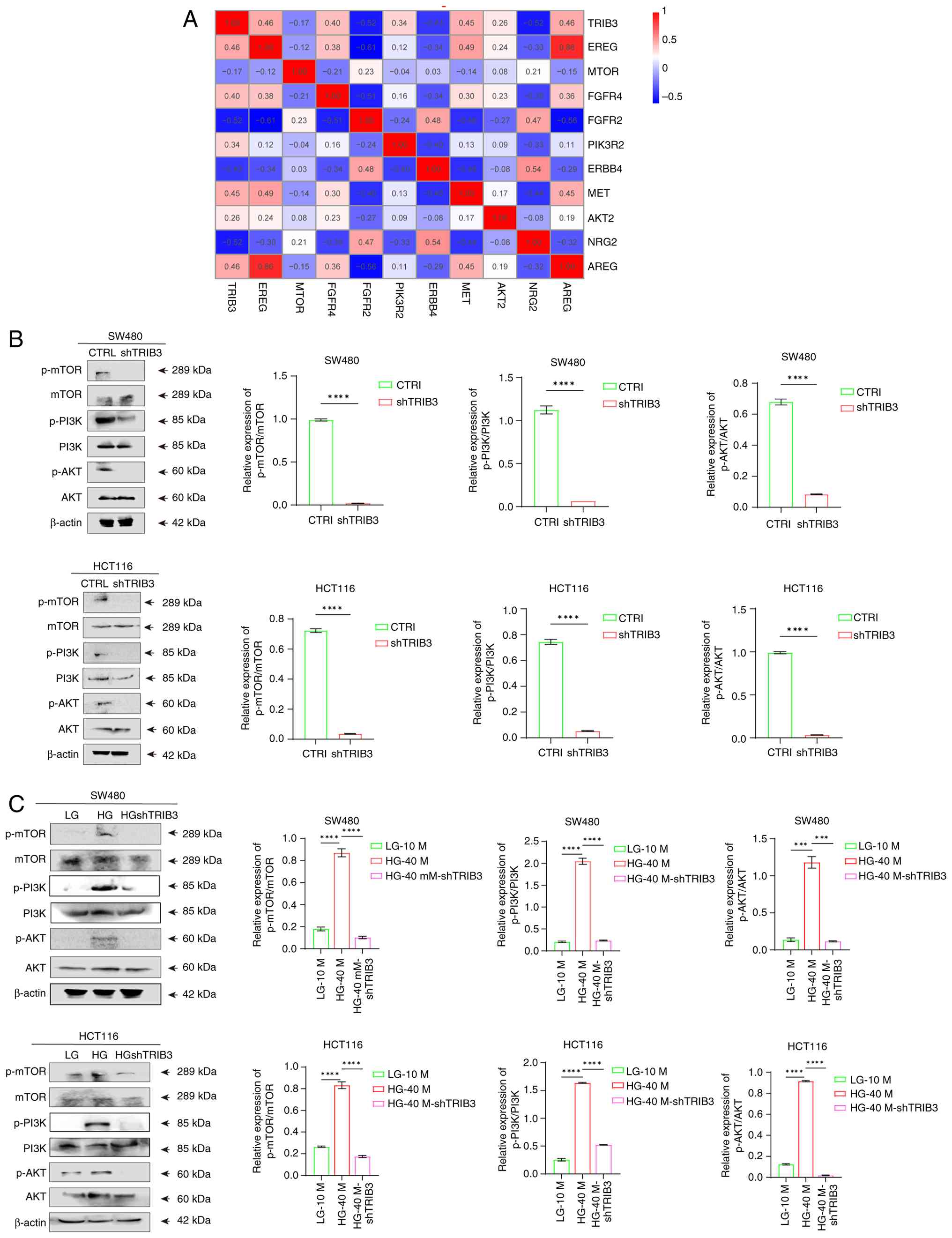 | Figure 5.TRIB3 activates the PI3K/AKT
signaling pathway in high glucose in colorectal cancer cells. (A)
Heatmap analysis demonstrating the correlation between TRIB3
expression levels and PI3K-related pathway. (B) Western blotting
was utilized to evaluate the impact of TRIB3 knockdown on the
PI3K/AKT pathway activation status. (C) Western blotting was
utilized to evaluate the alterations in the PI3K/AKT pathway
activation status following TRIB3 knockdown under HG conditions.
***P<0.001, ****P<0.0001. For all bar-plots, data are
presented as the mean ± SEM, n=3. CTRL, control; HG, high glucose;
LG, low glucose; ns, not significant; p-, phosphorylated; sh, short
hairpin; TRIB3, tribbles 3. |
High glucose-induced TRIB3 activation
drives CRC malignancy via the PI3K/AKT pathway
To investigate whether TRIB3 drives malignant
progression in CRC cells under high-glucose conditions via PI3K/AKT
pathway activation, the PI3K/AKT/mTOR pathway was first inhibited
using LY294002 (10 µM) and then TRIB3 was knocked down. In
high-glucose environments, LY294002 treatment alone led to
decreased phosphorylation levels of PI3K, AKT and mTOR, accompanied
by elevated ZO1 expression and reduced N-cadherin/Vimentin
expression compared with that in the high-glucose control group
(Fig. 6A). Notably, TRIB3
knockdown following pathway inhibition did not further enhance
these effects, indicating that they act on the same regulatory
axis. Furthermore, Transwell assays revealed that LY294002
treatment under high-glucose conditions significantly impaired the
invasion and migration of CRC cells compared with that in the
high-glucose control group, and TRIB3 knockdown elicited identical
phenotypic effects (Fig. 6B).
These findings suggest that TRIB3-driven invasion and migration
under high-glucose conditions are predominantly mediated by the
PI3K/AKT signaling pathway.
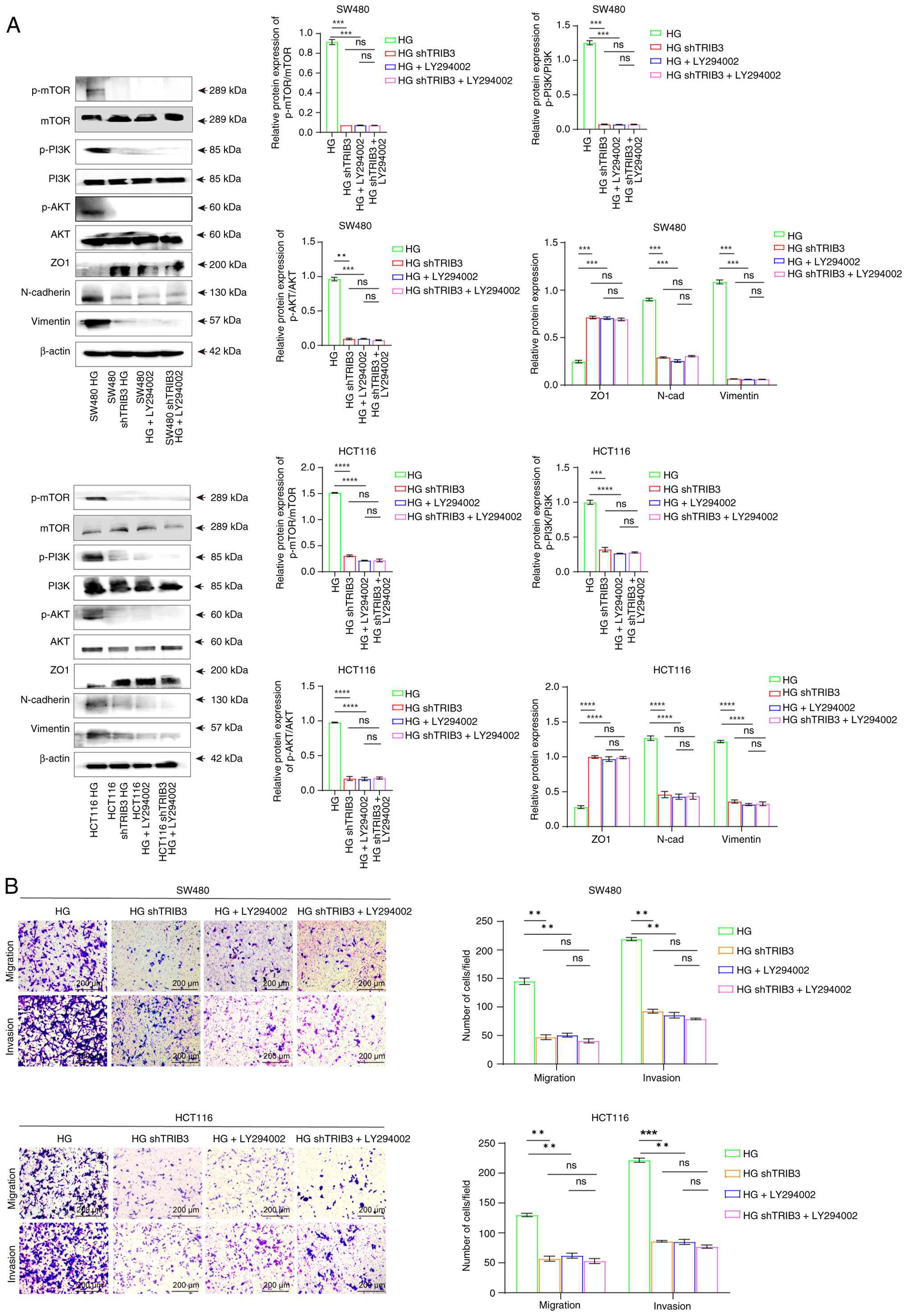 | Figure 6.TRIB3 promotes malignant behavior of
colorectal cancer cells through the PI3K/AKT pathway. (A) Western
blot analysis was performed to evaluate the levels of
epithelial-mesenchymal transition-related proteins and
PI3K/AKT/mTOR pathway proteins in cells treated with the
PI3K/AKT/mTOR pathway inhibitor LY294002 (10 µM), both in the
presence and absence of shTRIB3 in HG conditions. (B) Transwell
assays were used to evaluate the changes in cell migration and
invasion induced by inhibition of the PI3K/AKT signaling pathway
under HG conditions, with or without TRIB3 knockdown. **P<0.01,
***P<0.001, ****P<0.0001. For all bar-plots, data are
presented as the mean ± SEM, n=3. HG, high glucose; ns, not
significant; p-, phosphorylated; sh, short hairpin; TRIB3, tribbles
3. |
In order to confirm the interaction, the western
blot analysis showed that TRIB3 depletion not only reduced PI3K/AKT
phosphorylation but also inhibited GSK-3β phosphorylation at Ser9,
which in turn led to increased β-catenin phosphorylation and its
subsequent degradation in the cytoplasm (Fig. S3A and B). This reduction in
cytoplasmic β-catenin stability resulted in decreased nuclear
translocation of β-catenin, a phenomenon that may suppress
downstream signaling pathways and thereby block tumor progression.
These findings further indicated that TRIB3 is a critical molecular
driver in the development and progression of CRC, exerting its
oncogenic effects by activating multiple signaling pathways.
Targeting the TRIB3 pathway presents a potential option for
reversing the exacerbated malignant behavior of CRC cells induced
by a high-glucose environment.
Discussion
Due to alterations in diet and lifestyle, such as
increased consumption of high-fat, high-calorie food, insufficient
physical activity and smoking, the prevalence of CRC and diabetes
is likely to increase (21–23).
Notably, evidence has indicated an association between diabetes and
various types of cancer, including breast, pancreatic and liver
cancer (24–26), and type 2 diabetes has been
associated with a higher risk of CRC in an observational study
(6). Diabetes exerts a detrimental
influence on the outcomes of CRC treatment, resulting in an
increased risk of metastasis, recurrence and mortality (7). In our previous study, diabetes was
identified as an independent risk factor for postoperative CRC, and
diabetic patients with CRC had a higher risk of recurrence
(8). However, the mechanism
underlying the progression of diabetes into colon cancer remains
unclear.
The TRIB3 protein, categorized within the TRIB
family of pseudokinases, is characterized by the absence of
specific ATP-binding sites and catalytic cores, rendering it devoid
of kinase activity (24). Emerging
evidence has revealed that, as an intracellular pseudokinase, TRIB3
is a pivotal factor in regulating glucose homeostasis and insulin
resistance, and its role as a stress sensor has been firmly
established in response to a wide array of stressors, such as
hypoxia, fasting and high-glucose levels (27–30).
Numerous studies have uncovered a notable role for
TRIB3 in the development of tumors, and TRIB3 has been shown to be
highly expressed in gastric, liver, kidney and non-small cell lung
cancer (31,32). Furthermore, TRIB3 upregulation is
associated with tumor-related staging, lymph node metastasis,
disease recurrence and poor prognosis (33,34).
Previous studies have indicated that TRIB3 increases the expression
of genes associated with cancer stem cells, including CD44, CD133,
OCT4 and SOX2, and facilitates tumor development in CRC, resulting
in unfavorable outcomes (2,35,36).
Although research has provided information on the role of TRIB3 in
patients with CRC and diabetes, the specific underlying mechanisms
are yet to be fully elucidated (36). In the current study, the pivotal
role of TRIB3 in CRC was not only substantiated, but how TRIB3
facilitates the proliferation, invasion and migration of CRC cells
by activating the PI3K/AKT pathway under high-glucose conditions
was clarified.
The present study revealed that high-glucose levels
could stimulate proliferation, migration, invasion and EMT
remodeling in CRC cells, concurrently upregulating the expression
of TRIB3. These findings led to the scientific hypothesis that
high-glucose levels may facilitate malignant biological phenotypes
in CRC through the upregulation of TRIB3. Therefore, in subsequent
experiments in a high-glucose environment, TRIB3 was knocked down;
the results demonstrated that the suppression of TRIB3 expression
significantly counteracted the enhanced cellular proliferation and
invasion induced by high-glucose levels, and markedly inhibited
activation of the EMT process. These findings strongly support the
hypothesis that TRIB3 is a crucial molecule in tumor progression
under high-glucose conditions.
Regarding the mechanism by which TRIB3 regulates
tumor progression, multiple studies have been conducted. Hua et
al (35) demonstrated that
TRIB3 interacts with β-catenin to promote tumor cell malignancy,
which may align with the present findings, as the PI3K/AKT and
β-catenin pathways can converge and crosstalk at the GSK-3β site.
To confirm this interaction, experiments were performed in CRC
cells cultured under high-glucose conditions and with TRIB3
knockdown. The results demonstrated that TRIB3 depletion not only
reduced PI3K/AKT phosphorylation but also inhibited GSK-3β
phosphorylation at Ser9, leading to increased β-catenin
phosphorylation and subsequent degradation in the cytoplasm. This
resulted in reduced nuclear translocation of β-catenin, which may
suppress downstream signaling pathways and block tumor
progression.
Additionally, as demonstrated by Shang et al
(36), TRIB3 can influence the CRC
immune microenvironment and mediate tumor progression by regulating
the STAT1-CXCL10 axis. This may be due to factors downstream of the
PI3K/AKT pathway, such as NF-κB, which can indirectly regulate
STAT1 activity, and TRIB3-mediated inhibition of STAT1 could
further impair CXCL10-mediated T-cell recruitment. These findings
further indicated that TRIB3 is a critical molecular driver in the
development and progression of CRC, exerting its oncogenic effects
by activating multiple signaling pathways.
Moreover, numerous studies have explored the
mechanisms by which high-glucose levels mediate the upregulation of
TRIB3 expression. High-glucose levels are considered to promote
glucose metabolism through the hexosamine biosynthetic pathway,
which elevates O-GlcNAc-modified proteins. These modified proteins
may function as transcription factors or co-regulators that
directly or indirectly influence TRIB3 gene expression (37). For example, O-GlcNAc modification
of Sp1, a well-known transcription factor, has been shown to
upregulate TRIB3 expression (37).
In addition, other factors may contribute to high glucose-induced
TRIB3 upregulation. For example, high-glucose levels can activate
signaling pathways, such as protein kinase C and mitogen-activated
protein kinase (38,39). These pathways could regulate TRIB3
expression through the phosphorylation of transcription factors and
other regulatory proteins.
In conclusion, multiple studies have suggested that
elevated blood glucose levels contribute to the upregulation of
TRIB3 expression, and the present study further indicated that the
upregulation of TRIB3 expression may activate the PI3K/AKT
signaling pathway to promote cell malignant behavior by enhancing
EMT. Conversely, TRIB3 knockdown significantly inhibited the
PI3K/AKT signaling pathway, thereby suppressing oncogenic signaling
and tumor progression. However, the underlying mechanisms remain
highly complex and require further investigation in subsequent
research. Notably, TRIB3 has emerged as a critical therapeutic
target for diabetic CRC.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
This work was supported by the National Natural Science
Foundation of China (grant no. 82303649 to RY) and the Natural
Science Foundation of Beijing Municipality (grant no. 7232064 to
GA).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
DY conceived and designed the study. RY performed
the statistical analysis, revision and final proofreading of the
article. YB and XH acquired the data and performed experiments. XH
analyzed the data. YG and GA interpreted the data. YB and DY
drafted the manuscript. YG and GA revised the manuscript. All
authors read and approved the final manuscript. YB and RY confirm
the authenticity of all the raw data.
Ethics approval and consent to
participate
The use of human tissue samples was approved by the
Ethics Committee of Beijing Chaoyang Hospital affiliated with
Capital Medical University (approval no. 2020-10-13-4), in
accordance with The Declaration of Helsinki and applicable Chinese
regulations. Written informed consent was obtained from all the
participants for their participation. The animal study protocol was
approved by the Animal Ethics Committee of Capital Medical
University (approval no. AEE1-2025-1255). The study strictly
adhered to the ethical principles and standards set forth by the
ethics committee.
Patient consent for publication
All enrolled patients provided written informed
consent for publication.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Harborg S, Kjærgaard KA, Thomsen RW,
Borgquist S, Cronin-Fenton D and Hjorth CF: New Horizons:
Epidemiology of Obesity, Diabetes Mellitus, and Cancer Prognosis. J
Clin Endocrinol Metab. 109:924–935. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Shahid RK, Ahmed S, Le D and Yadav S:
Diabetes and cancer: Risk, challenges, management and outcomes.
Cancers (Basel). 13:57352021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Abudawood M: Diabetes and cancer: A
comprehensive review. J Res Med Sci. 24:942019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Jordt N, Kjærgaard KA, Thomsen RW,
Borgquist S and Cronin-Fenton D: Breast cancer and incidence of
type 2 diabetes mellitus: A systematic review and meta-analysis.
Breast Cancer Res Treat. 202:11–22. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Blaslov K, Bulum T and Duvnjak L:
Pathophysiological factors in the development of diabetic
nephropathy-new insights. Acta Med Croatica. 68:135–140. 2014.(In
Croatian). PubMed/NCBI
|
|
6
|
Murphy N, Song M, Papadimitriou N,
Carreras-Torres R, Langenberg C, Martin RM, Tsilidis KK, Barroso I,
Chen J, Frayling TM, et al: Associations between glycemic traits
and colorectal cancer: A mendelian randomization analysis. J Natl
Cancer Inst. 114:740–752. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Roman D, Saftescu S, Timar B, Avram V,
Braha A, Negru Ș, Bercea A, Serbulescu M, Popovici D and Timar R:
Diabetes mellitus and other predictors for the successful treatment
of metastatic colorectal cancer: A retrospective study. Medicina
(Kaunas). 58:8722022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Yu D, An G and Yao J:
Lymphocyte-to-monocyte ratio combined with CA19-9 for predicting
postoperative recurrence of colorectal cancer in patients with
diabetes. J Clin Lab Anal. 35:e239442021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Eyers PA, Keeshan K and Kannan N: Tribbles
in the 21st Century: The evolving roles of tribbles pseudokinases
in biology and disease. Trends Cell Biol. 27:284–298. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lu G, Li J, Gao T, Liu Q, Chen O, Zhang X,
Xiao M, Guo Y, Wang J, Tang Y and Gu J: Integration of dietary
nutrition and TRIB3 action into diabetes mellitus. Nutr Rev.
82:361–373. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Prudente S, Sesti G, Pandolfi A, Andreozzi
F, Consoli A and Trischitta V: The mammalian tribbles homolog
TRIB3, glucose homeostasis, and cardiovascular diseases. Endocr
Rev. 33:526–546. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cao X, Fang X, Malik WS, He Y, Li X, Xie
M, Sun W, Xu Y and Liu X: TRB3 interacts with ERK and JNK and
contributes to the proliferation, apoptosis, and migration of lung
adenocarcinoma cells. J Cell Physiol. 235:538–547. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang XJ, Li FF, Zhang YJ, Jiang M and Ren
WH: TRIB3 promotes hepatocellular carcinoma growth and predicts
poor prognosis. Cancer Biomark. 29:307–315. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yu JJ, Zhou DD, Yang XX, Cui B, Tan FW,
Wang J, Li K, Shang S, Zhang C, Lv XX, et al: TRIB3-EGFR
interaction promotes lung cancer progression and defines a
therapeutic target. Nat Commun. 11:36602020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Amin MB, Greene FL, Edge SB, Compton CC,
Gershenwald JE, Brookland RK, Meyer L, Gress DM, Byrd DR and
Winchester DP: The eighth edition AJCC cancer staging manual:
Continuing to build a bridge from a population-based to a more
‘personalized’ approach to cancer staging. CA Cancer J Clin.
67:93–99. 2017.PubMed/NCBI
|
|
17
|
Vaziri-Gohar A, Hue JJ, Abbas A, Graor HJ,
Hajihassani O, Zarei M, Titomihelakis G, Feczko J, Rathore M,
Chelstowska S, et al: Increased glucose availability sensitizes
pancreatic cancer to chemotherapy. Nat Commun. 14:38232023.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ding F, Guo Y, Zhang H, Zhong Y, Zhang D,
Huang Q, Zheng Z, Liu G, Zhang X and Weng S: High glucose-induced
mitochondrial fission drives pancreatic cancer progression through
the H3K18la/TTK/BUB1B signal pathway. Cell Signal. 135:1120272025.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Leary S, Underwood W, Anthony R, Cartner
S, Grandin T, Greenacre C, Gwaltney-Bran S, McCrackin MA, Meyer R,
Miller D, et al: AVMA Guidelines for the Euthanasia of Animals:
2020 Edition. American Veterinary Medical Association; Schaumburg,
IL: 2020
|
|
20
|
Li H, Li C, Zhang B and Jiang H:
Lactoferrin suppresses the progression of colon cancer under
hyperglycemia by targeting WTAP/m(6)A/NT5DC3/HKDC1 axis. J Transl
Med. 21:1562023. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sun X, Yan AF, Shi Z, Zhao B, Yan N, Li K,
Gao L, Xue H, Peng W, Cheskin LJ and Wang Y: Health consequences of
obesity and projected future obesity health burden in China.
Obesity (Silver Spring). 30:1724–1751. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Cignarelli A, Genchi VA, Caruso I,
Natalicchio A, Perrini S, Laviola L and Giorgino F: Diabetes and
cancer: Pathophysiological fundamentals of a ‘dangerous affair’.
Diabetes Res Clin Pract. 143:378–388. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhan Z, Chen B, Lin W, Chen X, Huang R,
Yang C and Guo Z: Rising Burden of colon and rectum cancer in
China: An analysis of trends, gender disparities, and projections
to 2030. Ann Surg Oncol. 32:3361–3371. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kang C, LeRoith D and Gallagher EJ:
Diabetes, obesity, and breast cancer. Endocrinology. 159:3801–3812.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Norat T, Scoccianti C, Boutron-Ruault MC,
Anderson A, Berrino F, Cecchini M, Espina C, Key T, Leitzmann M,
Powers H, et al: European code against cancer 4th edition: Diet and
cancer. Cancer Epidemiol. 39 (Suppl 1):S56–S66. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhang Z, Qin W and Sun Y: Contribution of
biomarkers for pancreatic cancer-associated new-onset diabetes to
pancreatic cancer screening. Pathol Res Pract. 214:1923–1928. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Cheng WP, Lo HM, Wang BW, Chua SK, Lu MJ
and Shyu KG: Atorvastatin alleviates cardiomyocyte apoptosis by
suppressing TRB3 induced by acute myocardial infarction and
hypoxia. J Formos Med Assoc. 116:388–397. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Dobens LL, Nauman C, Fischer Z and Yao X:
Control of cell growth and proliferation by the tribbles
pseudokinase: Lessons from drosophila. Cancers (Basel). 13:8832021.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lee SK, Park CY, Kim J, Kim D, Choe H, Kim
JH, Hong JP, Lee YJ, Heo Y, Park HS and Jang YJ: TRIB3 is highly
expressed in the adipose tissue of obese patients and is associated
with insulin resistance. J Clin Endocrinol Metab. 107:e1057–e1073.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ord T and Ord T: Mammalian pseudokinase
TRIB3 in normal physiology and disease: Charting the progress in
old and new avenues. Curr Protein Pept Sci. 18:819–842. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Bowers AJ, Scully S and Boylan JF: SKIP3,
a novel Drosophila tribbles ortholog, is overexpressed in human
tumors and is regulated by hypoxia. Oncogene. 22:2823–2835. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Loo LW, Tiirikainen M, Cheng I, Lum-Jones
A, Seifried A, Church JM, Gryfe R, Weisenberger DJ, Lindor NM,
Gallinger S, et al: Integrated analysis of genome-wide copy number
alterations and gene expression in microsatellite stable, CpG
island methylator phenotype-negative colon cancer. Genes
Chromosomes Cancer. 52:450–466. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Yang K, Li B, Xu X, Yu Z, Lyu X, Ren K,
Liu X, Chen S and Li H: TRIB3 overexpression predicts malignant
progression and poor prognosis in human solid tumors:
Bioinformatics validation and clinical significance. Expert Rev Mol
Diagn. 1–12. 2024.(Epub ahead of print). PubMed/NCBI
|
|
34
|
Zhou H, Luo Y, Chen JH, Hu J, Luo YZ, Wang
W, Zeng Y and Xiao L: Knockdown of TRB3 induces apoptosis in human
lung adenocarcinoma cells through regulation of Notch 1 expression.
Mol Med Rep. 8:47–52. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Hua F, Shang S, Yang YW, Zhang HZ, Xu TL,
Yu JJ, Zhou DD, Cui B, Li K, Lv XX, et al: TRIB3 Interacts With
β-Catenin and TCF4 to increase stem cell features of colorectal
cancer stem cells and tumorigenesis. Gastroenterology.
156:708–721.e15. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Shang S, Yang YW, Chen F, Yu L, Shen SH,
Li K, Cui B, Lv XX, Zhang C, Yang C, et al: TRIB3 reduces CD8(+) T
cell infiltration and induces immune evasion by repressing the
STAT1-CXCL10 axis in colorectal cancer. Sci Transl Med.
14:eabf09922022. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhang W, Liu J, Tian L, Liu Q, Fu Y and
Garvey WT: TRIB3 mediates glucose-induced insulin resistance via a
mechanism that requires the hexosamine biosynthetic pathway.
Diabetes. 62:4192–4200. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Pan D, Xu L and Guo M: The role of protein
kinase C in diabetic microvascular complications. Front Endocrinol
(Lausanne). 13:9730582022. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Schultze SM, Hemmings BA, Niessen M and
Tschopp O: PI3K/AKT, MAPK and AMPK signalling: protein kinases in
glucose homeostasis. Expert Rev Mol Med. 14:e12012. View Article : Google Scholar : PubMed/NCBI
|




















