Introduction
Coronary artery disease (CAD), affecting ~10.9% of
adults aged 45 years or older and responsible for over 800,000
myocardial infarctions annually in the United States, represents
the primary cause of morbidity and mortality globally, with
pathogenesis involving complex interactions among genetic,
environmental and lifestyle factors (1,2).
Despite notable advances in understanding traditional
cardiovascular risk factors, notable inter-individual variations in
CAD susceptibility propose the involvement of additional genetic
determinants that may contribute to disease development and
progression (3,4). Among these emerging biomarkers,
haptoglobin (HP), an acute-phase glycoprotein primarily synthesized
in the liver, has garnered considerable attention for its potential
role in cardiovascular pathophysiology (5–7). HP,
a plasma glycoprotein, serves an important role in preventing
oxidative damage by binding free hemoglobin released during
intravascular hemolysis (8). This
binding mechanism protects vascular tissues from hemoglobin-induced
oxidative stress and facilitates the clearance of hemoglobin-HP
complexes through the CD163 receptor on macrophages (9).
The HP gene exhibits a common structural
polymorphism resulting in three major phenotypes, namely Hp1-1,
Hp2-1 and Hp2-2, which are determined by the inheritance of HP1 and
HP2 alleles (10). Beyond the
common Hp1-1, Hp2-1 and Hp2-2 phenotypes, rare HP deletion (HPdel)
genotypes, resulting from an ~28-kb deletion spanning the HP
promoter to HP-related protein exon 5, are prevalent in East Asian
populations. Hp1-del/Hp2-del compound heterozygotes and Hpdel-del
homozygotes show ethnic frequency variations, comprising ~0.025,
0.067, 0.100 and 2.000% of Japanese, Korean, Chinese and Vietnamese
populations respectively (11).
The phenotypic variants of HP demonstrate distinct functional
properties, with Hp1-1 showing superior antioxidant capacity
compared with Hp2-2, whereas Hp2-1 exhibits intermediate
functionality (12). Genome-wide
association studies have previously identified specific genetic
loci such as rs2000999 as strong determinants of circulating HP
levels, providing molecular insights into the genetic regulation of
HP expression (13). Hpdel-del
homozygosity causes complete anhaptoglobinemia, whereas compound
heterozygotes demonstrate reduced serum HP levels. Clinically,
anhaptoglobinemia raises anaphylactic transfusion reaction risk
through anti-HP antibodies, and these variants may affect
antioxidant capacity and cardiovascular risk assessment,
particularly in East Asian populations (14). This functional heterogeneity has
notable implications for cardiovascular health, as oxidative stress
has been shown to serve a notable role in the development and
progression of atherosclerosis (15).
Previous studies have demonstrated notable
associations between HP polymorphisms and CAD risk, with particular
emphasis on ethnic-specific variations (16–18).
Notably, a comprehensive meta-analysis of 8,632 individuals
revealed that HP polymorphisms were notably associated with CAD
susceptibility, with stronger associations observed in Asian
populations than in Caucasians (19). These findings underscore the
importance of population-specific investigations, as genetic
architecture and allele frequencies may vary substantially across
ethnic groups (20).
In Chinese populations, preliminary studies have
suggested distinct patterns of HP phenotype distribution and their
cardiovascular implications. A prospective study of Chinese
patients with type 2 diabetes demonstrated that the Hp1-1 phenotype
was associated with a 43% increased risk of incident acute
myocardial infarction (21).
Furthermore, Mendelian randomization analysis in Chinese patients
with diabetes provided evidence for a causal relationship between
the serum levels of HP in patients and macroangiopathic incidence
(22). These findings indicate
that HP may serve as both a biomarker and a potential therapeutic
target in Chinese populations.
However, despite these promising preliminary
findings, to the best of our knowledge, no previous study has
comprehensively examined the relationship between HP phenotypes,
serum concentrations and CAD risk specifically in Chinese
populations. Most studies have focused predominantly on Western
populations or have examined HP in the context of diabetes-related
complications rather than primary CAD (23,24).
Additionally, the simultaneous assessment of HP phenotypes and
serum concentrations in relation to CAD risk has been
insufficiently explored, particularly in Asian populations, where
genetic background may substantially influence these associations
(25).
The present study addressed this notable knowledge
gap by examining the relationship between HP phenotypes and serum
concentrations with CAD risk in a Chinese population. By employing
rigorous phenotyping methodologies and a well-defined case-control
design, the present study aimed to provide valuable insights into
the role of HP polymorphisms in CAD susceptibility among Chinese
individuals. This may help improve risk stratification and
personalize cardiovascular medicine approaches in this
population.
Materials and methods
Study participants
The present case-control study enrolled genetically
unrelated patients treated at the Division of Cardiology, Bethune
International Peace Hospital (Shijiazhuang, China), between August
and December 2021. All patients suspected of CAD underwent coronary
angiography (CAG). CAG images were independently evaluated by two
experienced interventional cardiologists who were blinded to the
clinical characteristics and laboratory data of the patients.
Disagreements were resolved by consensus or by a third senior
interventional cardiologist. CAD severity was assessed according to
American Heart Association guidelines (26), with CAD defined as ≥50% luminal
stenosis in at least one major epicardial coronary artery or its
major branches. The number of diseased vessels was determined by
examining the left anterior descending artery, left circumflex
artery and right coronary artery (range, 0–3 vessels).
Two control groups were established: Group 1
comprised patients with coronary luminal stenosis ≤40% on
angiography, normal left ventricular ejection fraction (LVEF) and
normal regional wall motion, while group 2 consisted of healthy
individuals randomly selected from those undergoing routine health
examinations at the Bethune International Peace Hospital. Clinical
and demographic data, including baseline characteristics and
laboratory parameters, were extracted from electronic medical
records. Serum lipid profiles, including total cholesterol,
triglycerides, high-density lipoprotein-cholesterol (HDL-C), and
low-density lipoprotein-cholesterol (LDL-C), were measured using
standard enzymatic colorimetric methods on a Beckman Coulter AU5800
automated clinical chemistry analyzer (Beckman Coulter, Inc., Brea,
CA, USA).
Retrospectively-collected residual clinical blood
specimens obtained within 24 h of admission were utilized; serum
samples were allocated for HP electrophoresis-based phenotyping and
quantitative determination, and anticoagulated venous blood samples
were reserved for HP PCR-based genotyping analysis.
Serum HP concentration
measurement
Serum HP concentrations were quantified using a BN™
II nephelometer system (Siemens Healthineers) employing
immunoturbidimetric methodology. Detection was performed using N
Antiserum to Human Haptoglobin (Cat. No. OSAS09; Siemens
Healthineers) with N Protein Standard SL (Cat. No. OQIM13) for
calibration. Quality control was performed using N/T Protein
Control SL (Cat. No. OQIN13) and N/T Protein Control L (Cat. No.
OQIO13; Siemens Healthineers). The lower detection limit of the
assay was 0.08 g/l. Routine instrument maintenance, calibration and
quality control procedures were performed according to manufacturer
specifications and ISO9001 quality management standards (27).
HP genotyping and phenotyping
HP genotypes were determined using two complementary
methods: Reverse transcription-quantitative PCR (RT-qPCR) and
native polyacrylamide gel electrophoresis, ensuring result
validation and accuracy. HP genotyping was performed using a
TaqMan-based RT-qPCR assay adapted from previously validated
methods (28,29). Genomic DNA was extracted from
EDTA-anticoagulated peripheral blood samples using a GeneRotex 96
automated nucleic acid extraction system with the Magnetic
Bead-based Whole Blood Genomic DNA Extraction kit (cat. no. T148H;
Xi'an Tianlong Science and Technology Co., Ltd.,) according to the
manufacturer's instructions.
This assay determined HP genotypes by evaluating the
relative copy number of the HP2 allele-specific duplication
junction region. The HP-5′ promoter region, which is present in all
individuals regardless of genotype (28,29),
was used as an internal genomic control for normalization. Primers
and TaqMan probes were synthesized by Sangon Biotech Co., Ltd.,
with the following sequences: HP5, forward
5′-CACATTTACTGATTTCAGGCTGGA-3′, reverse
5′-CCTTTTCACAGTAATTTTCTCCACCT-3′, TaqMan probe
5′-FAM-AGCTTTTAAGCAATAGGGAGATGGCCACA-BHQ1-3′; HP2, forward
5′-GGAGCTGCTCTGCACATCAA-3′, reverse 5′-CCCTTTCAATGAATTTCAGGGA-3′,
TaqMan probe 5′-VIC-ACCCCGAATAGAAGCTCGCGAACTGTA-BHQ1-3′; and HPdel,
forward 5′-TCTTTATGGCACTGGGGAACA-3′, reverse
5′-AGCAAGACACTCGTGAGTGGAA-3′, TaqMan probe
5′-ROX-TGTGCAAGAGCCTTTCCAATTTTGATCA-BHQ2-3′. Each 20 µl PCR
reaction contained 2 µl genomic DNA template, 10 µl Premix Ex Taq™
(Perfect Real Time; cat. No. RR039A; Takara Bio Inc.,), 1 µl each
of HP2 forward and reverse primers (300 nM), 1 µl each of HPdel
forward and reverse primers (300 nM), 0.5 µl each of HP5 forward
and reverse primers (150 nM), 1 µl each of HP2-TaqMan and
HPdel-TaqMan probes (83 nM), 0.5 µl of the HP5-TaqMan probe (42 nM)
and nuclease-free water to reach the final volume. An AGS4800
Real-Time PCR System (AGS Technologies Co., Ltd) was used for
amplification. The PCR cycling conditions were as follows: Initial
denaturation at 95°C for 10 min, followed by 40 cycles of
denaturation at 95°C for 10 sec and annealing/extension at 60°C for
1 min.
HP genotypes were determined using the comparative
Cq method (2−ΔΔCq method) (30). For each sample, the following
calculations were performed: ΔCq=Cq(HP5′)-Cq(HP2);
ΔΔCq=ΔCq(sample)-ΔCq(reference); HP2/HP5′ ratio=2−ΔΔCq;
where the reference was genomic DNA from a confirmed HP2/HP2
homozygotic individual. Genotypes were assigned based on the
HP2/HP5′ ratio: i) HP1/HP1 was assigned when no notable HP2 signal
was detected; ii) HP2/HP1 was defined as a ratio ranging from
0.34–0.50; and iii) HP2/HP2 individuals displayed ratios ranging
from 0.79–0.98. The presence of HPdel was identified by detection
of the HPdel-specific signal, and samples showing HPdel signals
were further categorized as HP1/HPdel, HP2/HPdel or HPdel/HPdel
based on the combination of HP2 and HPdel signal patterns.
Gel electrophoresis phenotyping
HP phenotypes were confirmed by native
polyacrylamide gel electrophoresis of HP-hemoglobin complexes
(10,31,32).
Briefly, 10 µl serum samples were mixed with 2 µl 10% (v/v)
hemoglobin solution and incubated at room temperature for 10 min to
allow formation of HP-hemoglobin complexes. Samples were then mixed
with an equal volume of non-denaturing loading buffer, consisting
of 125 mM Tris-HCl, 20% glycerol and 0.001% bromophenol blue at pH
6.8.
Electrophoresis was performed using a discontinuous
polyacrylamide gel system, which consisted of a 4% stacking gel
(125 mM Tris-HCl; pH 6.8) and a 4.7% resolving gel (360 mM
Tris-HCl; pH 8.8). Initial electrophoresis was conducted at 120 V
until the bromophenol blue front approached the resolving gel,
followed by 150 V until completion. Gels were stained using a
chromogenic substrate solution containing 5 ml 0.2% (w/v)
3,3′,5,5′-tetramethylbenzidine in methanol, 0.5 ml dimethyl
sulfoxide, 10 ml 5% (v/v) glacial acetic acid, 1 ml 1% (w/v)
potassium ferricyanide and 150 µl 30% (w/w) hydrogen peroxide.
Staining was performed at room temperature for 5–15 min.
Statistical analysis
Statistical analyses were conducted using Python
(version 3.9; Python Software Foundation) with pandas (33), NumPy (34), scipy.stats (35) and scikit-learn libraries (36). Continuous variables were presented
as mean ± standard deviation and categorical variables as
frequencies (percentages). Between-group comparisons were conducted
using χ2 test for categorical variables and one-way
ANOVA) or Kruskal-Wallis tests for continuous variables, depending
on data distribution assessed by the Shapiro-Wilk test. Following
significant ANOVA results, post hoc pairwise comparisons were
performed using Bonferroni correction. For Kruskal-Wallis tests,
Mann-Whitney U tests with Bonferroni correction were applied for
post hoc pairwise comparisons. For χ2 tests involving
multiple groups, post-hoc pairwise comparisons were performed using
individual χ2 tests with Bonferroni correction for
multiple comparisons (adjusted α=0.05/number of comparisons).
Correlation analyses employed Pearson's or
Spearman's coefficients as appropriate. Multivariate logistic
regression identified independent CAD risk factors, including
variables with P<0.2 from the univariate analysis. Model
performance was evaluated using receiver operating characteristic
(ROC) curve analysis and area under the curve (AUC)
calculations.
Genotype-stratified analyses explored HP
genotype-specific associations by performing the aforementioned
correlation analysis methods within each genotype subgroup (n≥10).
Between-genotype HP concentration comparisons were conducted using
Kruskal-Wallis tests with η2 effect sizes.
Missing data were handled by complete case analysis,
for example the quantity of LVEF missing data was 18.3%.
Statistical significance was set at α=0.05.
Results
Baseline characteristics of the study
participants
The present study enrolled 230 patients with CAD, 83
participants in the control 1 group and 192 participants in the
control 2 group. Key baseline characteristics are summarized in
Table I. The CAD group was
predominantly male (74.8%), significantly more compared with the
control groups 1 (56.6%; P=0.003) and 2 (53.1%; P<0.001).
Smoking prevalence was significantly higher in the CAD group
(30.0%) compared with the control groups 1 (16.9%; P=0.029) and 2
(15.1%; P<0.001). Hypertension was significantly more frequent
in the CAD group (62.6%) compared with in the control 2 group
(26.0%; P<0.001). Similarly, diabetes mellitus was significantly
more prevalent in the CAD group (21.3%) compared with in control
group 2 (8.3%; P<0.001). Genetic history was rare but present
only in the CAD and control 1 groups.
 | Table I.Baseline characteristics of the study
participants. |
Table I.
Baseline characteristics of the study
participants.
|
| Group | Statistical
analysis |
|---|
|
|
|
|
|---|
| Variables | CAD (n=230) | Control 1
(n=83) | Control 2
(n=192) |
P-valuea |
P-valueb |
|---|
| Age, years | 61.0
(53.0–68.75) | 59.0
(51.0–63.0) | 62.0
(50.75–71.0) | 0.067 | 0.714 |
| Sex |
|
|
| 0.003 | <0.001 |
|
Male | 172 (74.8) | 47 (56.6) | 102 (53.1) |
|
|
|
Female | 58 (25.2) | 36 (43.4) | 90 (46.9) |
|
|
| Smoking | 69 (30.0) | 14 (16.9) | 29 (15.1) | 0.029 | <0.001 |
| Drinking | 29 (12.6) | 11 (13.3) | 28 (14.6) | >0.999 | 0.654 |
| Hypertension | 144 (62.6) | 46 (55.4) | 50 (26.0) | 0.309 | <0.001 |
| Diabetes
mellitus | 49 (21.3) | 15 (18.1) | 16 (8.3) | 0.640 | <0.001 |
| Genetic
history | 7 (3.0) | 2 (2.4) | 0 (0.0) | >0.999 | 0.040 |
| Lipid profile,
mg/dl |
|
|
|
|
|
| TC | 4.22±1.02 | 4.34±0.85 | 4.38±0.57 | 0.350 | 0.064 |
| TG | 1.62±0.88 | 1.59±1.08 | 1.12±0.32 | 0.853 | <0.001 |
|
HDL | 1.04±0.24 | 1.12±0.16 | 1.33±0.27 | 0.004 | <0.001 |
|
LDL | 2.74±0.78 | 2.84±0.72 | 2.79±0.42 | 0.308 | 0.418 |
| No. of diseased
vessels |
|
|
|
|
|
| 1 | 75 (32.6) | - | - | - | - |
| 2 | 71 (30.9) | - | - | - | - |
| 3 | 84 (36.5) | - | - | - | - |
| LVEFc | 0.619±0.077 | 0.647±0.044 | - | 0.003 | - |
CAD group demonstrated significantly lower
high-density lipoprotein-cholesterol (HDL-C) levels (1.04±0.24
mg/dl) compared with control groups 1 (1.12±0.16 mg/dl; P=0.004)
and 2 (1.33±0.27 mg/dl; P<0.001) and significantly higher
triglyceride levels (1.62±0.88 mg/dl) than control group 2
(1.12±0.32 mg/dl; P<0.001). LVEF was significantly lower in the
CAD group (0.619±0.077) than in control group 1 (0.647±0.044;
P=0.003). No significant differences were observed in median age,
drinking prevalence, total cholesterol or low-density
lipoprotein-cholesterol between the CAD group and either control
group.
HP phenotype and serum levels
Of 505 samples analyzed, 450 showed concordant
results between the two genotyping methods, yielding an overall
concordance rate of 89.11% (450/505; data not shown). The remaining
55 samples required complementary use of both methods for accurate
genotyping. A representative electrophoretic typing profile of HP
is presented in Fig. S1.
Serum HP levels demonstrated significant genotypic
and group variations (Table II).
Within each genotype, the CAD group consistently displayed the
highest HP concentrations. Specifically, Hp2-2 carriers in the CAD
group demonstrated significantly higher serum HP levels (1.50±0.84
g/l) compared with those in control groups 1 (1.11±0.54 g/l;
P=0.004) and 2 (0.85±0.42 g/l; P<0.001). Similarly, Hp2-1
carriers in the CAD group (2.01±0.86 g/l) had significantly higher
HP levels than their counterparts in control groups 1 (1.39±0.39
g/l; P<0.001) and 2 (1.19±0.47 g/l; both P<0.001). The Hp1-1
genotype also showed significantly higher levels in the CAD group
than in control group 2 (2.03±0.79 vs. 1.31±0.61 g/l; P=0.001).
 | Table II.Distribution of HP genotypes and
serum levels. |
Table II.
Distribution of HP genotypes and
serum levels.
|
| CAD (n=230) | Control 1
(n=83) | Control 2
(n=192) | Statistical
analysis |
|---|
|
|
|
|
|
|
|---|
| HP genotypes | n (%) | HP level, g/l | n (%) | HP level, g/l | n (%) | HP level, g/l |
P-valuea |
P-valueb |
|---|
| 1-1 | 21 (9.1) | 2.03±0.79 | 5 (6.0) | 1.88±0.93 | 26 (13.5) | 1.31±0.61 | 0.715 | 0.001 |
| 2-1 | 69 (30.0) | 2.01±0.86 | 28 (33.7) | 1.39±0.39 | 75 (39.1) | 1.19±0.47 | <0.001 | <0.001 |
| 2-2 | 126 (54.8) | 1.50±0.84 | 45 (54.2) | 1.11±0.54 | 79 (41.1) | 0.85±0.42 | 0.004 | <0.001 |
| 1-del | 2 (0.9) | 0.56±0.68 | 5 (6.0) | 0.73±0.49 | 2 (1.0) | 0.08±0.00 | 0.713 | 0.423 |
| 2-del | 12 (5.2) | 0.66±0.92 | 0 (0.0) | - | 10 (5.2) | 0.41±0.28 | - | 0.425 |
| del-del | 0 (0.0) | - | 0 (0.0) | - | 0 (0.0) | - | - | - |
HP genotype distribution differed significantly
among CAD, control 1 and control 2 groups (χ2=25.014;
degrees of freedom, 8; P=0.002; data not shown). In CAD vs. control
2 group comparisons (Table III),
Hp2-2 was significantly more prevalent in patients with CAD than in
control group 2 [54.8 vs. 41.1%; χ2=7.254; P=0.007; odds
ratio (OR), 1.73; 95% confidence interval (CI), 1.18–2.55]. Hp2-1
showed a non-significant trend toward lower frequency in the CAD
group compared with control group 2 (30.0 vs. 39.1%;
χ2=3.431; P=0.064; OR, 0.67; 95% CI, 0.45–1.00). No
significant differences were observed for Hp1-1 (P=0.201) or
deletion genotypes (both P>0.999). Detailed statistical
comparisons are listed in Table
III. This comparison was selected for detailed analysis because
control group 2 provided greater statistical power (n=192 vs. n=83
for control group 1) and demonstrated the most substantial
differences in genotype distribution compared with the CAD
group.
 | Table III.χ2 test results for HP
genotype distribution comparing CAD vs. control 2 groups. |
Table III.
χ2 test results for HP
genotype distribution comparing CAD vs. control 2 groups.
| Genotype | CAD, n (%) | Control 2, n
(%) | χ2 | df | P-value | OR | 95% CI |
|---|
| Hp1-1 | 21 (9.1) | 26 (13.5) | 1.636 | 1 | 0.201 | 0.64 | 0.35–1.18 |
| Hp2-1 | 69 (30.0) | 75 (39.1) | 3.431 | 1 | 0.064 | 0.67 | 0.45–1.00 |
| Hp2-2 | 126 (54.8) | 79 (41.1) | 7.254 | 1 | 0.007a | 1.73 | 1.18–2.55 |
| Hp1-del | 2 (0.9) | 2 (1.0) | 0.000 | 1 | >0.999 | 0.83 | 0.12–5.97 |
| Hp2-del | 12 (5.2) | 10 (5.2) | 0.000 | 1 | >0.999 | 1.00 | 0.42–2.37 |
| Total | 230 (100.0) | 192 (100.0) | 8.386 | 4 | 0.078 | - | - |
Multivariate logistic regression
analysis of HP and CAD risk
Table IV presents
univariate and multivariate logistic regression analyses to
identify independent CAD risk factors. In the multivariate
analysis, HP concentration emerged as the strongest independent
predictor (OR, 4.556; P<0.001), followed by hypertension (OR,
3.772; P<0.001), triglycerides (OR, 2.682; P=0.002) and HDL-C as
a protective factor (OR, 0.011; P<0.001). Among HP phenotypes,
Hp2-2 showed an independent association with CAD risk (OR, 1.781;
P=0.045) compared with a Hp1-1 reference, representing a 78%
increased risk even after adjusting for HP concentration and
traditional cardiovascular risk factors. Other HP phenotypes showed
no significant associations.
 | Table IV.Multivariable logistic regression
analysis of variables associated with coronary artery disease
risk. |
Table IV.
Multivariable logistic regression
analysis of variables associated with coronary artery disease
risk.
|
| Univariate
analysisb | Multivariate
analysis |
|---|
|
|
|
|
|---|
|
Variablesa | OR | 95% CI | P-value | OR | 95% CI | P-value |
|---|
| Male sex | 2.617 | 1.735–3.947 | <0.001 | 1.407 | 0.770–2.571 | 0.267 |
| Age, years | 0.998 | 0.982–1.013 | 0.752 |
|
|
|
| Family history | 4.285 | 0.881–20.832 | 0.071 | 3.58 | 0.627–20.453 | 0.152 |
| Hypertension | 4.755 | 3.129–7.227 | <0.001 | 3.772 | 2.139–6.653 | <0.001 |
| Diabetes
mellitus | 2.978 | 1.632–5.433 | <0.001 | 2.068 | 0.881–4.858 | 0.095 |
| Smoking | 2.409 | 1.483–3.914 | <0.001 | 0.954 | 0.468–1.947 | 0.898 |
| Alcohol
consumption | 0.845 | 0.483–1.478 | 0.555 |
|
|
|
| HP concentration
(g/l) | 3.178 | 2.331–4.335 | <0.001 | 4.556 | 2.923–7.101 | <0.001 |
| Total cholesterol
(mmol/l) | 0.806 | 0.641–1.014 | 0.065 | 1.356 | 0.880–2.090 | 0.168 |
| Triglycerides
(mmol/l) | 4.731 | 2.962–7.557 | <0.001 | 2.682 | 1.435–5.013 | 0.002 |
| HDL cholesterol
(mmol/l) | 0.013 | 0.005–0.034 | <0.001 | 0.011 | 0.002–0.047 | <0.001 |
| LDL cholesterol
(mmol/l) | 0.883 | 0.654–1.192 | 0.417 |
|
|
|
| Hp2-2 vs.
Hp1-1 | 1.975 | 1.041–3.746 | 0.037 | 1.781 | 1.013–3.130 | 0.045 |
| Hp2-1 vs.
Hp1-1 | 1.139 | 0.588–2.207 | 0.700 |
|
|
|
| Hp1-del vs.
Hp1-1 | 1.238 | 0.161–9.546 | 0.838 |
|
|
|
| Hp2-del vs.
Hp1-1 | 1.486 | 0.537–4.109 | 0.446 |
|
|
|
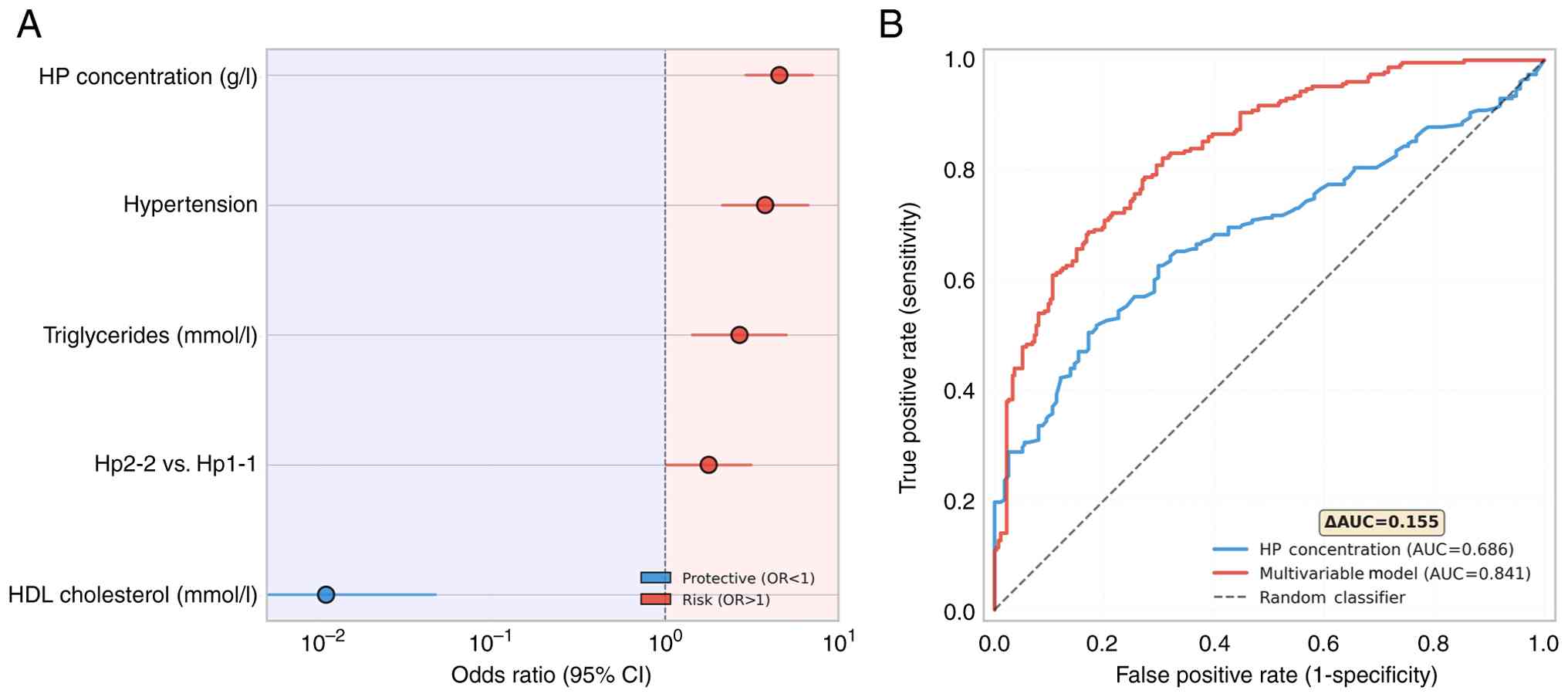
Visualization by forest plot demonstrated the
magnitude and direction of associations for the previously
identified significant predictors of CAD, with HP concentration and
hypertension showing the strongest positive associations with CAD
risk, whereas HDL-C levels demonstrated the most pronounced
protective effect (Fig. 1A). ROC
curve analysis demonstrated the superior discriminative ability of
the multivariate model (AUC=0.841) compared with HP concentration
alone (AUC=0.686), representing a clinically meaningful improvement
of 0.155 in AUC value (Fig. 1B).
The multivariate model achieved good discriminative performance,
indicating its potential clinical utility for the assessment of CAD
risk.
Genotype-specific analysis of HP with
CAD severity
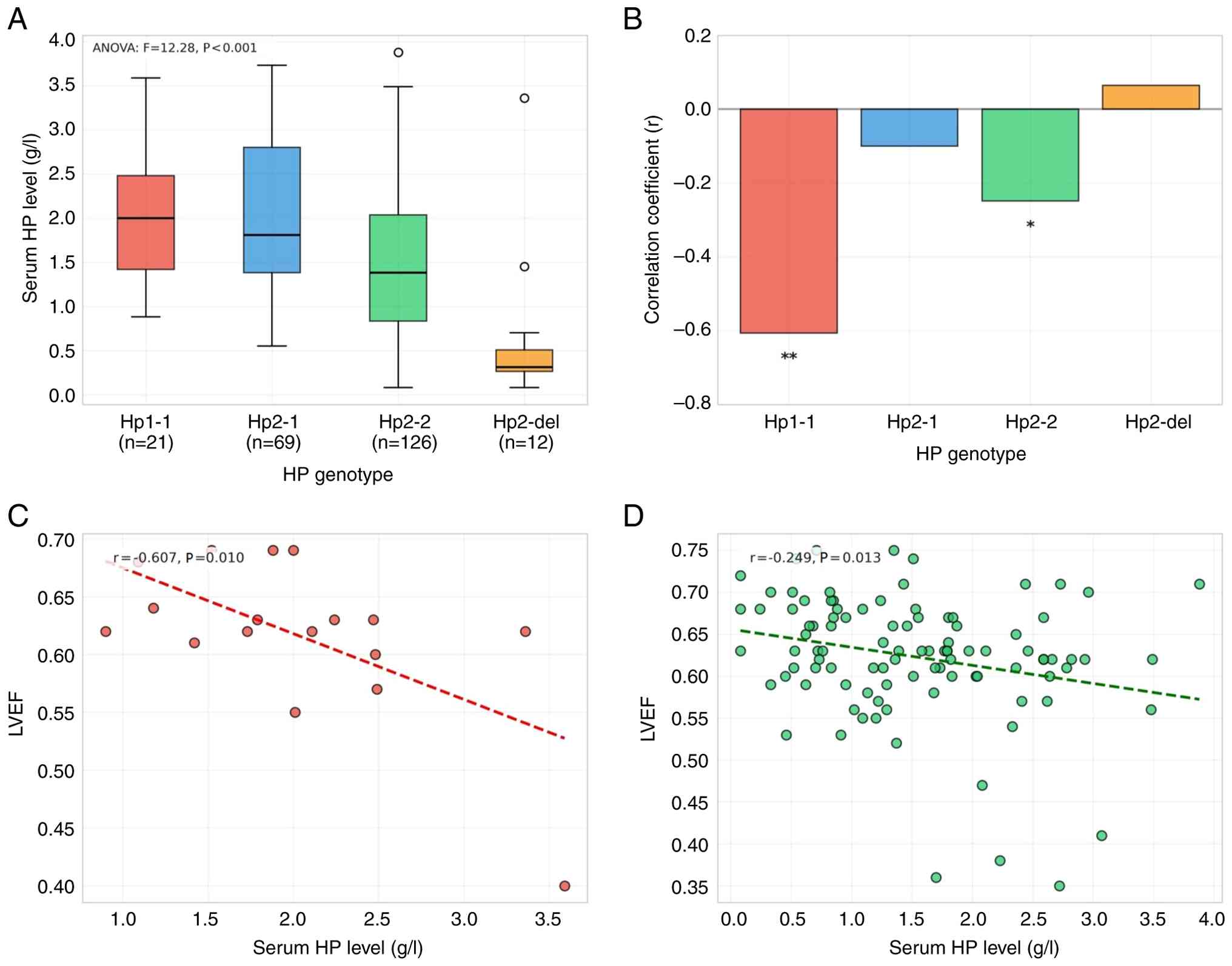
Significant differences in serum HP levels were
observed across genotypes (ANOVA F=12.28; P<0.001). Hp1-1 and
Hp2-1 genotypes exhibited the highest HP concentrations (2.03±0.79
and 2.01±0.86 g/l, respectively), whereas Hp2-2 showed intermediate
levels (1.50±0.84 g/l), and Hp2-del demonstrated the lowest
concentrations (0.66±0.92 g/l) (Fig.
2A).
Genotype-stratified correlation analysis revealed
distinct patterns of association between HP levels and cardiac
function (Fig. 2B). In the Hp1-1
group (n=17), serum HP concentration showed a moderate negative
correlation with LVEF (r=−0.607; P=0.010), explaining 36.8% of the
variance in cardiac function (Fig.
2C). Similarly, the Hp2-2 group (n=99) demonstrated a notable
negative correlation with cardiac function (r=−0.249; P=0.013)
(Fig. 2D). By contrast, Hp2-1 and
Hp2-del genotypes showed no significant associations with LVEF
(P>0.05).
Regarding CAD severity, only the Hp1-1 group showed
a trending positive correlation between HP levels and vessel count
(r=0.410; P=0.065), whereas other genotypes demonstrated no
significant associations (data not shown). These findings suggest
that HP concentration serves as a clinically relevant biomarker
primarily in specific genotype subgroups, supporting the
implementation of genotype-stratified HP assessment in
cardiovascular risk evaluation.
Discussion
The present study is a comprehensive investigation
of HP polymorphisms and serum concentrations in Chinese patients
with CAD, revealing significant genotype-specific associations with
cardiac function and disease incidence. The findings of the present
study provide novel insights into the clinical utility of HP
parameters as biomarkers for CAD risk assessment in the Chinese
population.
The 54.8% prevalence of the Hp2-2 genotype in the
present CAD cohort was consistent with findings from previous
multi-ethnic cardiovascular studies (37,38).
This proportion was not only higher than that of the control groups
established in the present study but also exceeded the Hp2-2
frequency reported in the general Chinese populations in earlier
studies (39,40). Multivariate analysis identified HP
concentration and the Hp2-2 genotype (compared with Hp1-1) as
robust independent predictors of CAD risk. The multivariable model
demonstrated superior discriminative ability compared with HP
concentration alone
The most notable finding of the present study was
the genotype-specific associations observed between HP serum levels
and cardiac function. The Hp1-1 group exhibited a moderate negative
correlation between HP concentration and LVEF, which was
particularly noteworthy due to the superior antioxidant capacity of
Hp1-1 protein (12), suggesting
that high HP levels in these patients may reflect severe oxidative
stress and inflammatory burden. The notable negative association
found in the Hp2-2 group aligned with the results of previous
mechanistic studies indicating that Hp2-2 protein demonstrates
lower antioxidant efficiency and hemoglobin-binding capacity
compared with Hp1-1 protein (41,42).
This functional difference may clarify why HP concentration showed
more sensitivity as a biomarker of cardiovascular dysfunction in
specific genotypes than others.
The genotype-specific associations observed in the
present study likely reflected fundamental differences in HP
protein structure and function. Hp1-1 forms stable dimers with
superior hemoglobin-binding affinity, whereas Hp2-2 creates large
multimeric complexes with reduced antioxidant capacity (14,43).
In cases of acute coronary syndrome, increased hemolysis and
oxidative stress may exceed the antioxidant capacity of different
HP variants to varying degrees, leading to genotype-specific HP
concentration elevation patterns (8,44).
The positive correlation between HP concentration
and CAD severity in the Hp1-1 group indicates that HP may have
acted as both a protective antioxidant and an inflammatory marker,
with the balance shifting to pathological signaling when
concentrations exceeded physiological ranges. Multiple lines of
evidence support this dual role: i) HP acts as the primary plasma
hemoglobin scavenger, mitigating heme-induced oxidative damage
(8,45); ii) HP is an acute-phase protein
that increases expression during systemic inflammation, with
dose-dependent promotion of monocyte chemotaxis (46,47);
iii) clinical studies show a U-shaped relationship between HP
concentration and cardiovascular outcomes, where both very low and
very high levels associate with higher patient mortality (48); and iv) proteomic analyses in
patients with acute coronary syndrome consistently identify a high
HP level as an inflammatory biomarker (44,49).
This concentration-dependent shift from protection to pathology
reflected the finite hemoglobin-binding capacity of HP: Adequate HP
levels relative to free hemoglobin confer antioxidant protection,
however high HP levels driven primarily by acute-phase
inflammation, rather than those observed by compensatory
upregulation against hemoglobin, mark disease severity rather than
protection (8,50).
The findings of the present study support the
implementation of genotype-stratified HP assessment in clinical
practice. For Hp1-1 genotypes, HP concentration monitoring could
provide valuable prognostic information regarding cardiac function,
whereas routine HP measurement may demonstrate limited utility in
the Hp2-1 group. This personalized approach aligns with precision
medicine principles and could improve the cost-effectiveness of
biomarker-based CAD management. Given the 3-fold difference in
baseline HP concentrations between genotypes, the establishment of
genotype-specific reference ranges appears warranted (40). Current clinical laboratories
typically use population-based reference intervals that may
misclassify individuals with specific genotypes (39).
Several study limitations warrant consideration. The
cross-sectional design of the present study prevented causal
inference; thus, longitudinal studies are required to establish the
temporal relationship between HP parameters and CAD progression.
Although the sample sizes used in the present study provided
adequate statistical power for the primary analyses, a larger
cohort would have enhanced the robustness of findings, particularly
for subgroup analyses. Furthermore, the single-center design of the
present study may have limited generalizability. Missing LVEF data
(18.3%) may have introduced selection bias, although sensitivity
analyses suggested minimal effect on the conclusions drawn in the
present study. Additionally, relatively small sample sizes observed
for rare genotypes (Hp1-del, n=9; Hp2-del, n=22) limited the
precision of estimates for these subgroups. Future multicenter
studies with larger cohorts are needed to validate the
genotype-specific associations observed in the present study and to
explore their prognostic value in diverse clinical settings.
Future prospective cohort studies should evaluate
the prognostic utility of genotype-stratified HP assessment for
predicting cardiovascular events and treatment responses. Studies
of HP dynamics in patients with acute coronary syndrome may uncover
additional clinical applications. Mechanistic investigations
through in vitro cellular assays, protein functional studies
and animal models are needed to clarify the pathogenic pathways
linking HP genotypes to CAD. Furthermore, therapeutic studies
targeting HP-mediated oxidative pathways could explore novel
treatment approaches for genotype-specific CAD management (51).
In conclusion, HP genotypes significantly modified
the clinical relevance of serum HP concentrations in patients with
CAD. Thus, genotype-stratified HP assessment could enhance
cardiovascular risk evaluation and support personalized medicine
approaches in Chinese populations.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by the National Key R&D
Program of China (grant no. 2019YFF0216502).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
ZH and FW conceived and planned the present study.
ZH, HL and KJ performed the experiments and analyzed data. DL and
LA contributed to sample preparation. HL, DL and JW contributed to
the interpretation of results. FW was responsible for writing the
manuscript. All authors provided critical feedback and helped shape
the research, analysis and manuscript. ZH and FW confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
The present study was conducted in accordance with
the Declaration of Helsinki and approved by the Medical Ethics
Committee of Bethune International Peace Hospital (Shijiazhuang,
China; approval no. 2021-KY-152). Written informed consent was
waived due to the retrospective nature of the study using residual
clinical specimens.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Malakar AK, Choudhury D, Halder B, Paul P,
Uddin A and Chakraborty S: A review on coronary artery disease, its
risk factors, and therapeutics. J Cell Physiol. 234:16812–16823.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Duggan JP, Peters AS, Trachiotis GD and
Antevil JL: Epidemiology of coronary artery disease. Surg Clin
North Am. 102:499–516. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jia Q, Han Y, Huang P, Woodward NC,
Gukasyan J, Kettunen J, Ala-Korpela M, Anufrieva O, Wang Q, Perola
M, et al: Genetic determinants of circulating glycine levels and
risk of coronary artery disease. J Am Heart Assoc. 8:e0119222019.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
McPherson R and Tybjaerg-Hansen A:
Genetics of coronary artery disease. Circ Res. 118:564–578. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Mewborn EK, Tolley EA, Wright DB, Doneen
AL, Harvey M and Stanfill AG: Haptoglobin genotype is a risk factor
for coronary artery disease in prediabetes: A case-control study.
Am J Prev Cardiol. 17:1006252023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chang X, Dorajoo R, Han Y, Wang L, Liu J,
Khor CC, Low AF, Chan MY, Yuan JM, Koh WP, et al: Interaction
between a haptoglobin genetic variant and coronary artery disease
(CAD) risk factors on CAD severity in Singaporean Chinese
population. Mol Genet Genomic Med. 8:e14502020. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Orchard TJ, Backlund JC, Costacou T,
Cleary P, Lopes-Virella M, Levy AP and Lachin JM; DCCT/EDIC
Research Group, : Haptoglobin 2–2 genotype and the risk of coronary
artery disease in the Diabetes control and complications
trial/epidemiology of diabetes interventions and complications
study (DCCT/EDIC). J Diabetes Complications. 30:1577–1584. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Andersen CBF, Stødkilde K, Sæderup KL,
Kuhlee A, Raunser S, Graversen JH and Moestrup SK: Haptoglobin.
Antioxid Redox Signal. 26:814–831. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Etzerodt A, Mikkelsen JH, Torvund-Jensen
M, Hennig D, Boesen T, Graversen JH, Moestrup SK, Kollman JM and
Andersen CBF: The Cryo-EM structure of human CD163 bound to
haptoglobin-hemoglobin reveals molecular mechanisms of hemoglobin
scavenging. Nat Commun. 15:108712024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Carter K and Worwood M: Haptoglobin: A
review of the major allele frequencies worldwide and their
association with diseases. Int J Lab Hematol. 29:92–110. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Soejima M, Agusa T, Iwata H, Fujihara J,
Kunito T, Takeshita H, Lan VT, Minh TB, Takahashi S, Trang PT, et
al: Haptoglobin genotyping of Vietnamese: Global distribution of HP
del, complete deletion allele of the HP gene. Leg Med (Tokyo).
17:14–16. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cooper CE, Schaer DJ, Buehler PW, Wilson
MT, Reeder BJ, Silkstone G, Svistunenko DA, Bulow L and Alayash AI:
Haptoglobin binding stabilizes hemoglobin ferryl iron and the
globin radical on tyrosine β145. Antioxid Redox Signal.
18:2264–2273. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Froguel P, Ndiaye NC, Bonnefond A,
Bouatia-Naji N, Dechaume A, Siest G, Herbeth B, Falchi M, Bottolo
L, Guéant-Rodriguez RM, et al: A genome-wide association study
identifies rs2000999 as a strong genetic determinant of circulating
haptoglobin levels. PLoS One. 7:e323272012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Delanghe JR, Delrue C, Speeckaert R and
Speeckaert MM: Unlocking the link between haptoglobin polymorphism
and noninfectious human diseases: Insights and implications. Crit
Rev Clin Lab Sci. 61:275–297. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Schaer CA, Deuel JW, Bittermann AG, Rubio
IG, Schoedon G, Spahn DR, Wepf RA, Vallelian F and Schaer DJ:
Mechanisms of haptoglobin protection against hemoglobin
peroxidation triggered endothelial damage. Cell Death Differ.
20:1569–1579. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Moussa A, Rejeb J, Omezzine A, Rebhi L,
Boumaiza I, Kacem S, Ben Rejeb N, Boughzala E, Ben Abdelaziz A and
Bouslama A: Association between haptoglobin 2–2 genotype and
coronary artery disease and its severity in a tunisian population.
Biochem Genet. 52:269–282. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Christensen JH, Krarup HB, Riahi S, Toft E
and Schmidt EB: Heart rate variability is associated with
haptoglobin phenotype in patients with coronary artery disease. Eur
J Cardiovasc Prev Rehabil. 12:221–225. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Cahill LE, Warren RA, Carew AS, Levy AP,
Ginsberg HN, Sapp J, Lache O and Rimm EB: The relationship between
time-varying achieved HbA1c and risk of coronary events depends on
haptoglobin phenotype among white and black ACCORD participants.
Diabetes Care. 46:1941–1948. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wang J, Zhou X, Su Y, Chai D, Ruan Y and
Wang J: Association between haptoglobin polymorphism and coronary
artery disease: A meta-analysis. Front Genet. 15:14349752024.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tishkoff SA and Kidd KK: Implications of
biogeography of human populations for ‘race’ and medicine. Nat
Genet. 36 (11 Suppl):S21–S27. 2004. View
Article : Google Scholar : PubMed/NCBI
|
|
21
|
Gurung RL, Yiamunaa M, Liu S, Liu JJ, Chan
C, Choo RWM, Ang K, Sum CF, Tavintharan S and Lim SC: Association
of haptoglobin phenotype with incident acute myocardial infarction
in Chinese patients with type 2 diabetes. Cardiovasc Diabetol.
18:652019. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang S, Wang J, Zhang R, Wang T, Yan D, He
Z, Jiang F, Hu C and Jia W: Mendelian randomization analysis to
assess a causal effect of haptoglobin on macroangiopathy in Chinese
type 2 diabetes patients. Cardiovasc Diabetol. 17:142018.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Costacou T, Ferrell RE and Orchard TJ:
Haptoglobin genotype: A determinant of cardiovascular complication
risk in type 1 diabetes. Diabetes. 57:1702–1706. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Levy AP, Hochberg I, Jablonski K, Resnick
HE, Lee ET, Best L and Howard BV; Strong Heart Study, : Haptoglobin
phenotype is an independent risk factor for cardiovascular disease
in individuals with diabetes: The strong heart study. J Am Coll
Cardiol. 40:1984–1990. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Nakamura H, Soejima M, Munkhtulga L,
Iwamoto S and Koda Y: Haptoglobin polymorphism in Mongolian
population: Comparison of the two genotyping methods. Clin Chim
Acta. 408:110–113. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Scanlon PJ, Faxon DP, Audet AM, Carabello
B, Dehmer GJ, Eagle KA, Legako RD, Leon DF, Murray JA, Nissen SE,
et al: ACC/AHA guidelines for coronary angiography. A report of the
American College of Cardiology/American Heart Association Task
Force on practice guidelines (Committee on Coronary Angiography).
Developed in collaboration with the Society for Cardiac Angiography
and Interventions. J Am Coll Cardiol. 33:1756–1824. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Standardization ISO, . ISO 9001:2015
Quality management systems - Requirements. ISO; Geneva: 2015
|
|
28
|
Soejima M and Koda Y: TaqMan-based
real-time PCR for genotyping common polymorphisms of haptoglobin
(HP1 and HP2). Clin Chem. 54:1908–1913. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Soejima M and Koda Y: Rapid real-time PCR
detection of HPdel directly from diluted blood samples. Clin Chem.
54:1095–1096. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Hochberg I, Roguin A, Nikolsky E,
Chanderashekhar PV, Cohen S and Levy AP: Haptoglobin phenotype and
coronary artery collaterals in diabetic patients. Atherosclerosis.
161:441–446. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Smithies O: Zone electrophoresis in starch
gels: Group variations in the serum proteins of normal human
adults. Biochem J. 61:629–641. 1955. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
McKinney W: Data Structures for
Statistical Computing in Python. In. SciPy2010.
|
|
34
|
Harris CR, Millman KJ, van der Walt SJ,
Gommers R, Virtanen P, Cournapeau D, Wieser E, Taylor J, Berg S,
Smith NJ, et al: Array programming with NumPy. Nature. 585:357–362.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Virtanen P, Gommers R, Oliphant TE,
Haberland M, Reddy T, Cournapeau D, Burovski E, Peterson P,
Weckesser W, Bright J, et al: SciPy 1.0: Fundamental algorithms for
scientific computing in Python. Nat Methods. 17:261–272. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Pedregosa F, Varoquaux G, Gramfort A,
Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R,
Dubourg V, et al: Scikit-learn: Machine Learning in Python. JMLR.
12:2825–2830. 2011.
|
|
37
|
Dalan R, Liew H, Goh LL, Gao X, Chew DE,
Boehm BO and Leow MK: The haptoglobin 2–2 genotype is associated
with inflammation and carotid artery intima-media thickness. Diab
Vasc Dis Res. 13:373–376. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Adams JN, Cox AJ, Freedman BI, Langefeld
CD, Carr JJ and Bowden DW: Genetic analysis of haptoglobin
polymorphisms with cardiovascular disease and type 2 diabetes in
the Diabetes Heart Study. Cardiovasc Diabetol. 12:312013.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Shi X, Sun L, Wang L, Jin F, Sun J, Zhu X,
Tang L, Qu Y and Yang Z: Haptoglobin 2–2 genotype is associated
with increased risk of type 2 diabetes mellitus in Northern
Chinese. Genet Test Mol Biomarkers. 16:563–568. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lei D, Hu S, Guo M, Wang J, Ma X, Wang F
and He Z: Genotype-specific reference interval of haptoglobin tests
in a Chinese population on the BN II System. Sci Rep. 13:5772023.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Delanghe JR, Langlois MR and De Buyzere
ML: Haptoglobin polymorphism: A key factor in the proatherogenic
role of B cells? Atherosclerosis. 217:80–82. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Edwards O, Burris A, Lua J, Wilkie DJ,
Ezenwa MO and Doré S: Influence of haptoglobin polymorphism on
stroke in sickle cell disease patients. Genes (Basel). 13:1442022.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Tamara S, Franc V and Heck AJR: A wealth
of genotype-specific proteoforms fine-tunes hemoglobin scavenging
by haptoglobin. Proc Natl Acad Sci USA. 117:15554–15564. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Pekayvaz K, Losert C, Knottenberg V, Gold
C, van Blokland IV, Oelen R, Groot HE, Benjamins JW, Brambs S,
Kaiser R, et al: Multiomic analyses uncover immunological
signatures in acute and chronic coronary syndromes. Nat Med.
30:1696–1710. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Thomsen JH, Etzerodt A, Svendsen P and
Moestrup SK: The haptoglobin-CD163-heme oxygenase-1 pathway for
hemoglobin scavenging. Oxid Med Cell Longev. 2013:5236522013.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
di Masi A, De Simone G, Ciaccio C, D'Orso
S, Coletta M and Ascenzi P: Haptoglobin: From hemoglobin scavenging
to human health. Mol Aspects Med. 73:1008512020. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Kolattukudy PE and Niu J: Inflammation,
endoplasmic reticulum stress, autophagy, and the monocyte
chemoattractant protein-1/CCR2 pathway. Circ Res. 110:174–189.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Tan QH, Huang YQ, Liu XC, Liu L, Lo K,
Chen JY and Feng YQ: A U-shaped relationship between selenium
concentrations and all-cause or cardiovascular mortality in
patients with hypertension. Front Cardiovasc Med. 8:6716182021.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Mohamed Bakrim N, Mohd Shah ANS, Talib NA,
Ab Rahman J and Abdullah A: Identification of haptoglobin as a
potential biomarker in young adults with acute myocardial
infarction by proteomic analysis. Malays J Med Sci. 27:64–76. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Ascenzi P and Coletta M: Peroxynitrite
detoxification by human haptoglobin:hemoglobin complexes: A
comparative study. J Phys Chem B. 122:11100–11107. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Bruse N, Pardali K, Kraan M, Kox M and
Pickkers P; REVIVAL investigators, : Phenotype-specific therapeutic
efficacy of ilofotase alfa in patients with sepsis-associated acute
kidney injury. Crit Care. 28:502024. View Article : Google Scholar : PubMed/NCBI
|