Psoriasis is a chronic, immune-mediated inflammatory
disorder primarily affecting the skin, characterized by recurring
episodes of erythematous plaques covered with silvery scales,
typically on extensor surfaces such as the elbows and knees
(1). Clinical variants include
palmoplantar, pustular, erythrodermic and guttate types, each
presenting with distinct morphological and systemic features
(2). Once considered a
dermatological condition, psoriasis is increasingly recognized as a
systemic disease, with ~75% of patients experiencing at least one
comorbidity and a number of affected by multiple concurrent
conditions (2).
The pathogenesis of psoriasis is complex and
immune-mediated, with high heritability estimated at 60–90%
(3). Epidemiological evidence
supports this, showing that first- and second-degree relatives of
psoriatic patients exhibit markedly higher rates of the disease
compared with the general population. Monozygotic twins show even
higher concordance rates than dizygotic twins, with ~30% of
patients having a first-degree relative also affected (4–6).
These findings strongly suggest a heritable basis for psoriasis
susceptibility. Extensive research has identified multiple genetic
susceptibility loci, further elucidating the genetic architecture
of the disease (7).
In addition to genetic predisposition, environmental
and epigenetic factors play pivotal roles in disease onset and
progression. Modifiable triggers, such as infections, medications,
trauma and stress, can influence disease activity via epigenetic
modifications, including DNA methylation and histone modifications
(8,9). Advancements in high-throughput omics
technologies have facilitated integrative analyses of
transcriptomic, proteomic and metabolomic data, revealing molecular
alterations in psoriatic lesions and highlighting dysregulated
pathways as potential therapeutic targets (9–11).
Furthermore, emerging evidence uncovers the role of the skin and
gut microbiome in modulating immune responses in psoriasis,
prompting investigations into host-microbiota interactions as
potential contributors to disease pathogenesis and novel
therapeutic avenues (12). The
present review summarized recent advances in understanding
psoriasis pathogenesis, focusing on genetic, epigenetic and
systems-level insights that inform improved diagnostic and
therapeutic strategies.
The substantial heritability of psoriasis highlights
a significant genetic component in its pathogenesis (13). Linkage studies, which identify
genetic markers co-inherited with the trait within families, have
led to the discovery of 15 psoriasis susceptibility loci (PSORS)
(14). The primary susceptibility
locus, PSORS1 (15), located in
the major histocompatibility complex (MHC) on chromosome 6p21.3,
accounts for 35–50% of the genetic risk for psoriasis (15,16).
The PSORS1 region includes several candidate genes, notably HLA-C
(specifically the HLA-C06:02 allele, also known as HLA-Cw6),
CCHCR1 and CDSN (17). HLA-Cw6 is
particularly significant as it promotes cytotoxic immune responses
in the skin by presenting self-antigens, such as ADAMTS-like
protein 5, to CD8+ T cells, thereby exacerbating disease
onset and severity (18).
The completion of the Human Genome Project and
advances in sequencing technologies have greatly expanded genomic
datasets, offering valuable resources for investigating
genotype-phenotype correlations. Genome-wide association studies
(GWAS) using these datasets have markedly advanced our
understanding of psoriasis genetics. GWAS have not only confirmed
the findings from linkage studies but also facilitated new
discoveries. A recent meta-analysis identified 109 distinct PSORS
loci, 46 of which had not been previously reported (19). While the number of identified loci
continues to increase, it is noteworthy that ethnicity-specific
loci have also been identified (20,21).
A trans-ethnic GWAS of Caucasian and Chinese populations, for
example, revealed 11 population-specific loci (21), prompting recent studies to focus on
psoriasis genetics across different ethnic groups and regions
(Table I).
While genomic factors are central to psoriasis risk,
they are not the sole determinants. This is evident from the 64%
discordance rate observed among monozygotic twins, who share an
identical DNA sequence (22). This
suggests that mechanisms beyond genetics contribute to psoriasis
pathogenesis. Extensive research has highlighted the critical role
of epigenetic factors in psoriasis development (8,23).
Epigenetics involves changes in gene expression without alterations
to the underlying DNA sequence, typically mediated by DNA
methylation, histone modifications and noncoding RNA regulation
(24). These epigenetic
modifications influence gene expression patterns, further reshaping
the transcriptome, proteome and metabolome, all of which contribute
to psoriasis. The present review discussed how these epigenetic
mechanisms play a role in psoriasis and summarizes recent
discoveries in the field.
Methylation of cytosines or adenines is a key
epigenetic modification that contributes to chromatin repression
and gene silencing, playing a significant role in various diseases
(25). Typically, DNA methylation
in the promoter or enhancer regions of genes is catalyzed by DNA
methyltransferases (26). This
modification, known as hypermethylation, reduces the binding
affinity of transcription factors (TFs) and recruits methyl-CpG
binding domain (MBD) proteins such as MeCP2 and MBD2, which help
establish a repressive chromatin structure and silence gene
expression. Conversely, the removal of methylation marks
(hypomethylation), either passively during DNA replication or
actively through the action of ten-eleven translocation enzymes,
which convert 5-methylcytosine (5-mC) to 5-hydroxymethylcytosine
(5-hmC), relieves this repression (26–28).
In psoriasis, a self-immune disease, DNA methylation changes have
been observed in immune cells. Li et al (29) investigated DNA methylation
differences in immune cells from the peripheral blood of
psoriasis-discordant twins and identified four genes, PTPN6, CCL5,
NFATC1 and PRF1, showing differential methylation between psoriasis
and unaffected individuals, suggesting their involvement in
pathogenesis. In another study focusing on specific cell types, Han
et al (30) found that
naïve CD4+ T cells from psoriatic patients exhibited
stronger hypomethylation in 26 pericentromeric genomic regions and
hypermethylation in the promoter regions of 121 X chromosome genes.
While the differential methylation of immune cells has been
established, further research is needed to systematically map
regulatory targets and gain an improved understanding of their role
in psoriasis development (31).
Skin cells, by contrast, exhibit distinct DNA
methylation profiles compared with immune cells (32). Specific sites of interest in
epidermal cells include hypermethylation of the p16INK4a
promoter region, observed in ~30% of psoriatic patients. This
modification is associated with reduced gene expression and
increased keratinocyte proliferation, associated with higher
Psoriasis Area and Severity Index scores (33). Hypomethylation, on the other hand,
affects loci within the epidermal differentiation complex, such as
S100A7, S100A12 and OAS2, leading to enhanced
expression of pro-inflammatory genes (34). Additionally, intragenic
hypomethylation in the enhancer region of CYP2S1 has been
shown to increase its expression in psoriatic keratinocytes,
influencing cell proliferation and immune signaling pathways
(35). These methylation
alterations disrupt keratinocyte homeostasis and reinforce the
chronic inflammatory cycle characteristic of psoriasis (36).
Histone proteins undergo various post-translational
modifications (PTMs) that influence their interaction with DNA. To
date, at least nine distinct types of histone modifications have
been identified, with acetylation, methylation, phosphorylation and
ubiquitylation being the most extensively studied (37).
In psoriasis, histone methylation and acetylation
have been particularly well studied, revealing a distinct histone
modification landscape between skin cells and immune cells. In
psoriatic keratinocytes, acetylation of Histone H3K27 at the
RPL22 promoter leads to its overexpression, which
upregulates Cyclin D1, promotes keratinocyte proliferation,
inhibits apoptosis and recruits CD4+ T cells (38). Wilms tumor 1 further enhances
IL-1β expression by increasing histone acetylation in
keratinocytes (39). Additionally,
Sirtuin 1, a class III histone deacetylase (HDAC) crucial for
keratinocyte differentiation, is suppressed by IFN-γ in psoriatic
lesions, making the cells more responsive to IL-22 (40). Regarding methylation, reduced
levels of H3K9 methylation in keratinocytes are associated with
elevated IL-23 expression, which can trigger IL-17-driven
inflammation (41). EZH2, a
histone methyltransferase and its product H3K27me3 are both
upregulated in the psoriatic epidermis and in keratinocytes
stimulated with psoriatic cytokines (42). Inhibition or knockdown of EZH2
markedly reduces keratinocyte proliferation and alleviates
psoriasis-like symptoms in mouse models (42). Moreover, CD147 promotes psoriasis
onset by modifying H3K9me3 in keratinocytes (43).
Immune cells in psoriasis also exhibit profound
changes in histone modifications. In peripheral blood cells, global
acetylation of H3 and H4 is reduced in psoriatic patients and
inversely correlates with disease severity (44). On the methylation side, increased
levels of H3K4me are observed in peripheral blood mononuclear cells
from moderate-to-severe plaque psoriatic patients (45). Memory T cells, which are implicated
in disease recurrence, display distal H3K27ac at enhancers,
supporting persistent transcription of psoriasis-associated genes
enriched with GRHL TF-binding motifs (26). Glutaminase 1 enhances H3
acetylation at the IL-17A promoter in γδ T17 and Th17 cells,
promoting their differentiation and inflammatory activity (46). Furthermore, HDAC activity is
essential for the conversion of Tregs into IL-17-producing cells, a
process that can be blocked by the HDAC inhibitor trichostatin A
(47). While HDAC1 is upregulated
in psoriatic skin, SIRT1 is downregulated, reflecting an overall
imbalance in histone acetylation in immune cells (48,49).
Critically, Jmjd3-mediated H3K27 demethylation regulates Th17
differentiation and its inhibition suppresses IL-17 production by
downregulating Th17-related genes (50). Finally, infliximab treatment
restores the expression of genes regulated by histone lysine
demethylase KDM5B, further supporting a functional link between
histone methylation and therapeutic outcomes in psoriasis (51). These cell-type-specific histone
modification patterns highlight the divergent roles in psoriasis
pathogenesis, with immune cells undergoing changes that affect T
cell differentiation, cytokine expression and immune memory, while
skin cells primarily exhibit modifications that promote
hyperproliferation and cytokine production.
Transcriptional heterogeneity in psoriasis is
evident from the variability in gene expression both between
individual cells and across different patients, even within
clinically similar lesions (57).
This variability contributes to differences in disease severity,
lesion characteristics and treatment responses.
Early bulk RNA sequencing (RNA-seq) studies hinted
at inter-individual differences in gene expression profiles,
suggesting underlying molecular diversity (58,59).
More recent advancements in single-cell (sc) RNA-seq have provided
detailed insights into transcriptional heterogeneity at a cellular
resolution. For example, scRNA-seq analyses of both human and
murine psoriatic skin have revealed distinct keratinocyte subtypes,
some highly proliferative, others expressing inflammatory genes or
undergoing epithelial-to-mesenchymal transition, as well as diverse
fibroblast and immune cell populations with unique transcriptional
identities (60–62).
This heterogeneity is also apparent within immune
cells. scRNA-seq of psoriatic lesions has identified varying
activation states of Th17 and tissue-resident memory T cells, with
some subsets showing elevated IFN-γ, while others are enriched in
chromatin remodeling pathways, highlighting patient-specific immune
signatures (60).
Notably, heterogeneity is observed not only between
patients or cell types but also within different lesions of the
same individual. Non-lesional skin may already exhibit
‘pre-psoriatic’ transcriptional priming and distinct plaques within
a single patient can show variable expression of cytokines and
inflammatory mediators (60,62–64).
This spatial diversity suggests that transcriptional heterogeneity
correlates with lesion evolution, location and even therapeutic
outcomes.
Another important feature is transcriptional
plasticity: Cell states in psoriasis are dynamic, shifting in
response to cytokines and targeted therapies. Longitudinal
scRNA-seq studies have demonstrated that IL-23 blockade with agents
such as risankizumab induces rapid reprogramming of skin cell
populations (65,66). Within days, pro-inflammatory
fibroblast subsets, specifically WNT5A+/IL24+
cells, decline sharply, followed by reductions in activated T cells
and keratinocytes. By two weeks post-treatment, keratinocyte and
myeloid populations exhibit marked downregulation of IL-17/TNF
signaling pathways, highlighting the substantial plasticity and
reversibility of pathogenic cell states following therapeutic
intervention (66).
TFs are crucial in regulating gene expression
related to immune responses, epidermal differentiation and
inflammation, markedly contributing to the transcriptional
heterogeneity observed in psoriasis. Transcriptomic analyses of
psoriatic skin have revealed differentially expressed genes (DEGs),
a number of which encode cytokines and chemokines. Key TFs, such as
STAT1, STAT2 and STAT3, have consistently been linked to these DEGs
(67–69). For instance, STAT1 is activated in
lesional skin of psoriatic patients, where its expression represses
IL-22. The imbalance between STAT1 and STAT3 disrupts IL-22
expression, contributing to psoriasis pathogenesis (68,70).
STAT2 activates the expression of cytokines CXCL11 and CCL5 in
keratinocytes (67). Additionally,
lesser-known TFs, such as FOSL1 and FOXC1, have been implicated in
regulating psoriasis-associated gene networks (71,72).
Notably, FOXC1 negatively correlates with most immune-related DEGs,
suggesting its role as a transcriptional repressor in the
inflammatory environment of psoriatic lesions (72).
The Th17 immune axis is central to psoriasis
pathogenesis, with RORγt acting as the lineage-determining TF for
Th17 cells (73,74). RORγt regulates the expression of
IL-17A, IL-17F, IL-22 and other pro-inflammatory cytokines that
drive epidermal hyperplasia and immune infiltration (75). Inhibition of RORγt by small
molecules has emerged as a potential therapeutic approach to
simultaneously suppress multiple inflammatory signals (76). The broader balance of T helper cell
subsets is also important: psoriatic patients show reduced
expression of GATA-3 and IL-4, indicating an impaired Th2 response,
while T-bet and IFN-γ expression remain unchanged, reflecting a
Th1/Th17-skewed immune profile (77).
Transcriptional regulation in keratinocytes plays a
direct role in the pathological features of psoriasis.
Dysregulation of the Hedgehog signaling pathway, through increased
GLI1 expression, is observed in lesional skin and can be triggered
by reduced neurofibromin (NF1) levels. This activation drives
keratinocyte proliferation, establishing a mechanistic link between
NF1 deficiency and the psoriatic phenotype (78). Other keratinocyte-specific TFs,
such as Ovol1 and HES1, influence neutrophil recruitment and the
amplification of inflammation, further highlighting the crosstalk
between epidermal and immune compartments (79,80).
While NF-κB has been extensively studied in
inflammatory pathways, its exact role in psoriasis remains under
investigation (69,71). Elevated levels of phosphorylated
NF-κB are found in psoriatic skin and therapies such as TNF
inhibitors and corticosteroids are thought to partially exert their
effects through NF-κB suppression (69). However, due to NF-κB's broad role
in immunity, targeting it may require cell-specific strategies to
avoid systemic immunosuppression (81). Similarly, metabolic regulators like
PPAR-γ are emerging as transcriptional modulators with therapeutic
potential. PPAR-γ influences keratinocyte differentiation, skin
barrier integrity and inflammation and agonists approved for
metabolic diseases may be repurposed to treat psoriatic
inflammation and associated comorbidities (82).
Recent advances in single-cell transcriptomics have
facilitated the identification of TFs contributing to psoriasis.
For example, distinct pathways were activated in different
fibroblast cell clusters, governed by specific TFs (83). A recent study using regulon module
analysis revealed differential regulon activity between lesional
and non-lesional skin in psoriasis, highlighting IRF7 as a key
transcriptional regulator. IRF7 was markedly downregulated
following guselkumab treatment and its role in modulating the IL-17
pathway highlights its potential as both a disease driver and a
therapeutic response marker (84).
Collectively, these findings demonstrate that psoriasis is driven
by a complex, multi-layered transcriptional hierarchy, where TFs,
in conjunction with epigenetic factors, reshape the transcriptomic
landscape.
Proteins are central to the pathophysiology of
psoriasis, serving not only as structural components of the skin
barrier but also as dynamic mediators of inflammation, immune
signaling and cellular proliferation. In psoriatic lesions, the
dysregulated expression of key proteins, including cytokines such
as IL-17A and IL-36γ, chemokines and antimicrobial peptides such as
LL37, drives both the initiation and chronic progression of
inflammation (85,86). These proteins act as both effectors
and biomarkers of immune activation and epidermal dysfunction,
making them key targets for mechanistic investigation.
Advances in LC-MS technologies have enabled the
development of proteomic studies in psoriasis. Unlike
transcriptomics, which captures gene expression potential,
proteomics reflects the functional output of cells, including PTMs,
protein-protein interactions and secreted mediators (87). Extensive proteomic studies have
been conducted on psoriatic patients, primarily using skin and
serum samples (88,89).
Human samples and animal models have both been
employed in proteomic research, revealing distinct molecular
differences between psoriatic and healthy tissues. In murine
models, Schonthaler et al (90) identified calprotectin as the most
upregulated protein in lesional psoriatic skin. Further validation
showed that deletion of the S100A9 subunit of calprotectin markedly
inhibited psoriasis and inflammation. Human cell lines, such as
HaCaT keratinocytes stimulated with TNF-α to model psoriasis, have
also been used in proteomics studies. Exosomal proteomes from these
models revealed 131 differentially expressed proteins associated
with angiogenesis, epigenetic regulation and inflammation (91). Human skin and serum samples further
support these findings, with Møller et al (92) identifying the inner epidermis as
exhibiting the most distinct proteomic alterations, primarily
related to innate immunity and cholesterol biosynthesis. Lu et
al (93) used untargeted
proteomics combined with ELISA validation to identify serum pigment
epithelium-derived factor as a potential biomarker for diagnosing
peripheral psoriatic arthritis (PsA).
Collectively, these proteomic studies provide
valuable insights into the pathogenic mechanisms of psoriasis.
Interpreting these findings not only facilitates the discovery of
novel diagnostic biomarkers but also deepens our understanding of
disease-related cellular behaviors at the protein level. Moreover,
integrating multi-omics data, such as genomics, transcriptomics and
metabolomics, will offer a more comprehensive view of psoriasis'
molecular landscape, enabling the identification of key regulatory
networks and therapeutic targets (94). For example, by integrating
large-scale plasma proteomics with psoriasis GWAS data, Liu et
al (95) identified AIF1,
FCGR3 and HSPA1A as novel, druggable protein targets for psoriasis
through proteome-wide Mendelian randomization (MR) and
colocalization analysis.
The multi-layered genetic regulation of psoriasis
leads to a global reshaping of the functional proteome, which
alters cellular behavior and, in turn, rewires metabolic processes.
This cascade results in a shifted metabolome that reflects the
biochemical landscape of the disease. As the most downstream layer
of cellular regulation, the metabolome serves as a valuable
indicator for understanding the mechanisms of pathogenesis and for
identifying diagnostic biomarkers. Beyond their diagnostic value,
certain metabolites also play regulatory roles in psoriasis
progression. Notably, quinolinic acid, a tryptophan-derived
metabolite, has been shown to suppress NOD-like receptor pyrin
domain-containing protein 3 inflammasome activation and its
supplementation markedly alleviated psoriasiform skin inflammation
in murine models (96).
Metabolomic studies in psoriasis have revealed
widespread alterations in key biochemical pathways, with consistent
findings across various biological samples, including serum,
plasma, skin, urine and mononuclear cells. These changes reflect
the systemic and localized metabolic disruptions associated with
psoriatic disease and offer potential insights for both diagnostic
and therapeutic applications.
In serum and plasma, altered levels of amino acids,
lipids, carnitines and organic acids have been consistently
reported. Amino acids such as arginine, glutamine, cysteine and
asparagine show significant variation in psoriatic patients
compared with healthy controls (11,97–99).
Lipid profiles also exhibit dysregulation, including decreased
levels of polyunsaturated fatty acids such as linoleic acid and
arachidonic acid, alongside increased lipid peroxidation products
(100). Notably,
trimethylamine-N-oxide levels correlate with disease severity and
comorbidities like cardiovascular conditions (101). Some of these metabolic shifts
correlate with treatment response, suggesting a role for
metabolomics in monitoring therapeutic outcomes (102).
Skin-targeted metabolomics has provided additional
insights into disease-specific metabolic alterations. Lesional skin
exhibits altered levels of amino acid derivatives, with
concentrations correlating with plaque severity scores (103). A key feature of psoriasis skin is
the rewiring of lipid metabolism (104). For example, ceramides, critical
metabolites for the skin barrier, are found at altered levels in
psoriatic skin compared with healthy tissue. Additionally, elevated
concentrations of electrophilic fatty acids and hepoxilins in
psoriatic lesions indicate an imbalance in lipid mediator pathways
(105,106).
Urinary metabolomics, although less extensively
studied, presents an interesting area for research due to the ease
of urine collection, making it ideal for diagnostic method
development. Researchers have identified potential biomarkers such
as reduced citrate levels in PsA patients and elevated
tetranor-12(S)-HETE, which reflect neutrophil activity in psoriatic
lesions (107,108). Overall, metabolomics studies
across different sample types consistently highlight disturbances
in amino acid metabolism and lipid pathways in psoriasis. These
systemic metabolic shifts may contribute to disease pathogenesis.
By integrating proteomic and transcriptomic data, these insights
are advancing biomarker discovery for diagnostic purposes (Table II) and will inform the development
of therapeutic strategies.
Microbial organisms at the skin surface interact
with the epithelial barrier and immune systems, contributing to the
progression of skin barrier diseases (109). In psoriasis, commensal fungi like
Candida albicans can trigger pathogenic immune responses,
particularly Th17 activation and neutrophil recruitment, which
directly drive psoriatic skin inflammation (110).
The skin microbiome in psoriasis is marked by
reduced microbial diversity and significant compositional shifts
compared with healthy skin. Lesional skin exhibits decreased α- and
β-diversity, with reductions in key commensals such as
Cutibacterium, Lactobacillus, Burkholderia and
Corynebacterium, alongside an increase in potentially
pro-inflammatory taxa like Streptococcus and
Firmicutes (111–114). These microbial changes correlate
with disease severity, as increased abundance of
Corynebacterium and Staphylococcus has been
associated with higher psoriasis severity in some studies (115). However, other research suggests a
more varied and less consistent pattern in the psoriatic skin
microbiome: While healthy skin is predominantly colonized by
Malassezia fungi, fungal diversity among psoriatic patients
differs and Malassezia populations do not appear to differ
between healthy and psoriatic skin (116).
Extensive studies have also highlighted the
correlation between the gut microbiome and skin barrier diseases,
suggesting interactions within the gut-skin axis (117,118). In psoriasis, intestinal barrier
biomarkers correlate with disease severity, further supporting the
interaction between the skin condition and the gut environment
(119,120). Significant alterations in the gut
microbiome have been observed in psoriatic patients, though,
similar to skin microbiome findings, causality remains unclear.
While a number of studies report significant changes in β-diversity
(the diversity of microbial communities between samples), most
indicate minimal change in α-diversity (the variety of microbes
within individual samples) (121). The variability in findings across
studies is notable: Some report a decrease in Bacteroidaceae,
Erysipelotrichaceae, Veillonellaceae and
Bifidobacteriaceae, while others observe the opposite trend
(122–124).
Overall, both the gut and skin microbiomes in
psoriasis show significant changes in microbial composition, but
current data do not reveal clear patterns. This heterogeneity may
stem from variations in study design, such as sampling and
sequencing methods, as well as geographic or ethnic differences
(125). To address these
discrepancies, future research should focus on large, standardized,
longitudinal studies with consistent methodology and clear clinical
stratification to better define the role of the gut and skin
microbiomes in psoriasis.
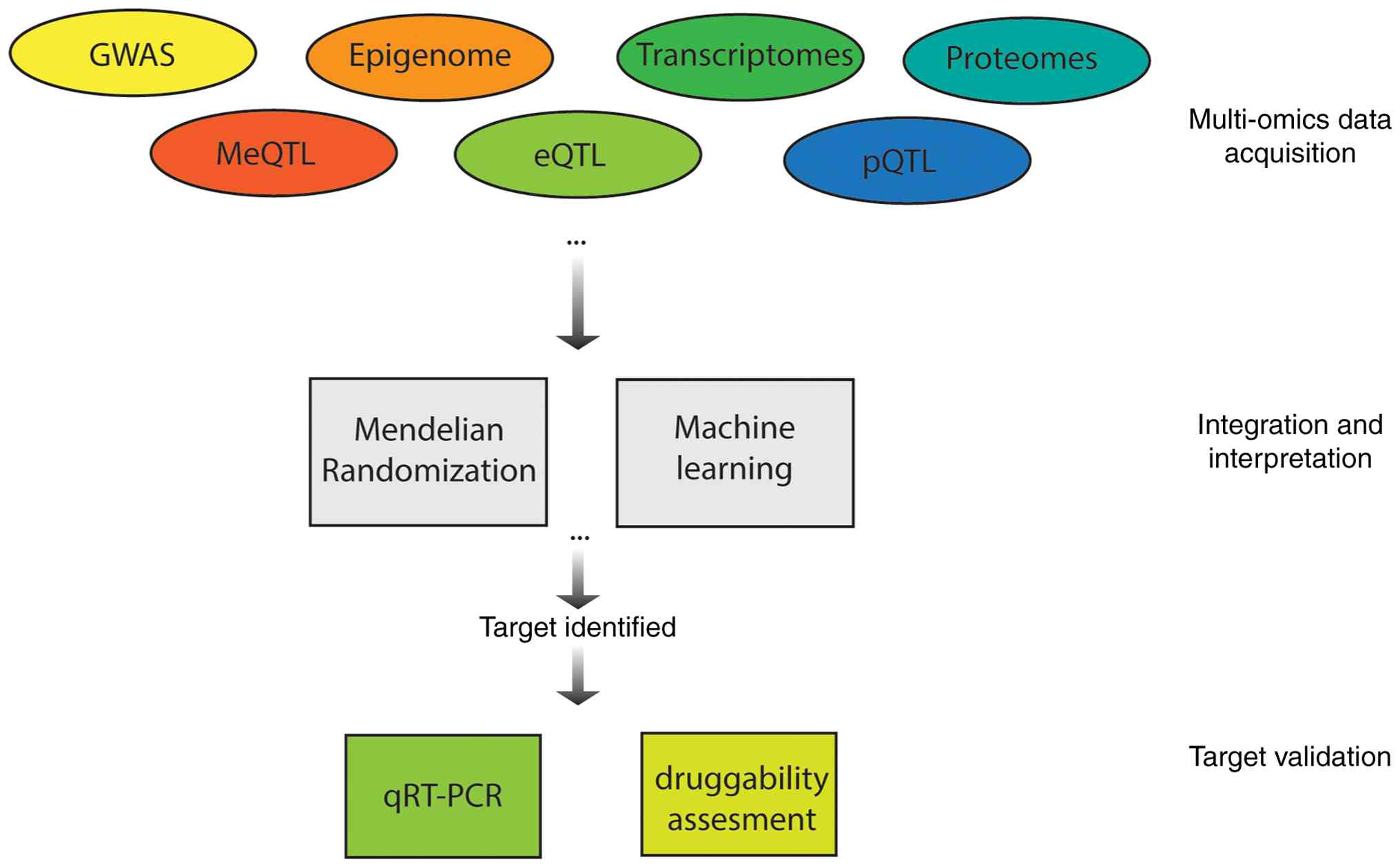
Advances in omics methodologies and the growing body
of knowledge in psoriasis have facilitated efforts to bridge
discovery and translation in psoriatic diseases by integrating
various omics layers to identify biomarkers and therapeutic
targets. Methodologies such as MR have enabled the integration of
genomic data with other omics datasets, using genetic variants as
instrumental variables to infer causal relationships between
molecular traits and disease risk. For example, Cai et al
(94) developed a proteome-wide MR
framework by aligning plasma protein quantitative trait loci data
with GWAS summary statistics for PsA, identifying seven proteins,
NEO1, IL23R, ERAP2, IFNLR1, KIR2DL3, CLSTN3 and POLR2F, that are
causally associated with the disease. Significant MR hits were
further analyzed through PPI networks to assess connectivity with
known drug targets. High-confidence Tier 1 and Tier 2 targets were
validated across multiple omics layers, including expression
quantitative trait locus (eQTL)-MR for genetic regulation,
drug-gene databases for therapeutic tractability, single-cell
transcriptomics for tissue specificity and PheWAS for pleiotropic
disease relevance, providing insights for potential therapy
development (94). Similar
MR-based frameworks have been applied in other studies. Notably,
Guo et al (126) applied
MR to integrate DNA methylation, gene expression and protein
abundance, identifying a regulatory axis involving lncRNA
RP11-977G19.11 and APOF as mediators in psoriasis pathogenesis,
thus offering a new potential therapeutic target. They also
validated the involvement of TNFAIP3 and MX1, which are critical
components of druggable pathways in psoriasis (126).
Machine learning has also emerged as a powerful tool
in multi-omics-based psoriasis research. For instance, Xing et
al (127) combined
transcriptomic and DNA methylation datasets from psoriasis lesions
and controls to identify differentially expressed methylated genes.
Machine learning algorithms were then used to pinpoint GJB2 as a
hub gene with strong diagnostic value, which was validated through
reverse transcription-quantitative PCR and immunohistochemistry. In
parallel, Deng et al (128) applied a multi-omics approach
integrating serum proteomics and skin transcriptomics, alongside
machine learning classification, to identify PI3 as a psoriasis
biomarker linked to disease severity and keratinocyte
hyperproliferation (128).
Integrating data across multiple omics layers offers a more
comprehensive understanding of disease mechanisms and facilitates
the discovery of robust biomarkers and therapeutic targets
(Fig. 1). Continued advances in
data integration methods will be critical for translating
multi-omics insights into biomarkers for precision diagnostics.
Moreover, robust models for target-guided drug screening will help
validate multi-omics findings and support the development of
effective psoriasis treatments (129).
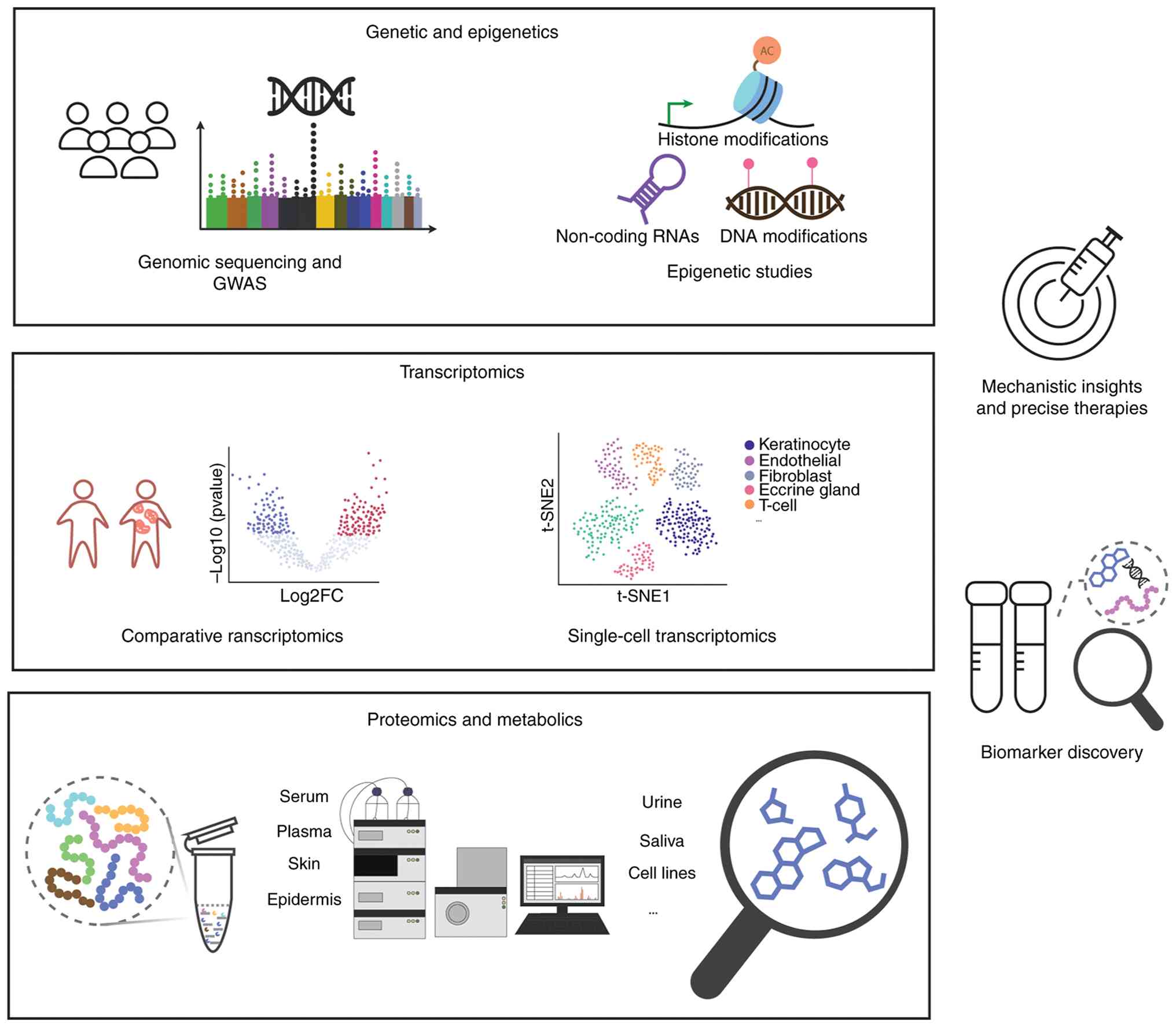
Traditionally regarded as an immune-mediated
disease, psoriasis pathogenesis has been broadened by multi-omics
investigations, extending beyond immune cell activation and
keratinocyte proliferation, emphasizing the roles of complex immune
signaling, transcriptional heterogeneity, dynamic proteomic
changes, metabolic rewiring and host-microbiome interactions. These
multi-layered insights not only enhance our understanding of
disease mechanisms but also identify promising biomarkers and
therapeutic targets (Fig. 2). In
the future, integrating omics data across spatial, temporal and
cellular dimensions will be crucial for advancing precision
diagnostics and targeted therapies.
Despite these advances, several challenges remain in
translating multi-omics findings into clinical applications. First,
at the single-omic level, inconsistencies in findings across
studies are prevalent, especially in microbiome and metabolome
research, where results often vary due to technical differences,
limited sample sizes, lack of longitudinal data and heterogeneity
of patient cohorts. These inconsistencies hinder the establishment
of robust biomarkers and call for standardized experimental
protocols and reference datasets.
Second, integrating multiple omics layers introduces
its own limitations. Although computational integration methods,
such as MR and machine learning, have enhanced target
identification, inconsistent data quality and the lack of
open-access, harmonized datasets may limit reproducibility and
cross-study comparability. Moreover, a number of current studies
suffer from limited sample sizes, particularly when combining
multi-omics data from the same individuals, which reduces
statistical power and increases the risk of overfitting in
computational models. Small and imbalanced datasets can also
obscure subtle biological signals, limiting the generalizability of
findings across diverse patient populations. Future endeavors
should prioritize the generation of large, well-annotated,
multi-omics cohorts with matched clinical metadata, as well as the
development of standardized pipelines and open-access platforms to
facilitate reproducibility, benchmarking and collaborative
discovery in psoriasis research.
Meanwhile, it is noteworthy that a number of
computationally identified biomarkers and therapeutic targets
remain unvalidated. A number of studies stop at correlation-based
insights without performing in vitro or in vivo
functional assays to establish causality or druggability, creating
a translational bottleneck. Bridging this gap will require robust
experimental follow-up and the integration of clinical metadata to
contextualize findings.
In conclusion, while multi-omics approaches have
shown great promise in unraveling the molecular complexity of
psoriasis, future efforts should prioritize methodological
standardization, cross-layer validation and translational research
to fully realize the potential of these insights in precision
diagnostics and targeted therapies.
Not applicable.
The present study was supported by Yunnan Provincial Department
of Science and Technology Applied Basic Research Joint Special
Project (grant no. 202101AZ070001-168), as well as Yunnan
University of Chinese Medicine Joint University-Hospital Fund
(grant no. XYLH2024053).
Not applicable.
HZ, DL, LZ, HY, LY, XY and YZ contributed to the
literature review and manuscript writing. HZ, DL and LZ also
critically reviewed and revised the manuscript for important
intellectual content. Data authentication is not applicable. All
authors read and approved the final manuscript.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
|
1
|
Boehncke WH and Schön MP: Psoriasis.
Lancet. 386:983–994. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Rendon A and Schäkel K: Psoriasis
pathogenesis and treatment. Int J Mol Sci. 20:14752019. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lønnberg AS, Skov L, Skytthe A, Kyvik KO,
Pedersen OB and Thomsen SF: Heritability of psoriasis in a large
twin sample. Br J Dermatol. 169:412–416. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Farber EM, Nall ML and Watson W: Natural
history of psoriasis in 61 twin pairs. Arch Dermatol. 109:207–211.
1974. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Griffiths CE and Barker JN: Pathogenesis
and clinical features of psoriasis. Lancet. 370:263–271. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kimmel GW and Lebwohl M: Psoriasis:
Overview and diagnosis. Evidence-Based Psoriasis. Springer; New
York, NY: pp. 1–16. 2018, View Article : Google Scholar
|
|
7
|
Tsoi LC, Spain SL, Knight J, Ellinghaus E,
Stuart PE, Capon F, Ding J, Li Y, Tejasvi T, Gudjonsson JE, et al:
Identification of 15 new psoriasis susceptibility loci highlights
the role of innate immunity. Nat Genet. 44:1341–1348. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mateu-Arrom L and Puig L: Genetic and
epigenetic mechanisms of psoriasis. Genes (Basel). 14:16192023.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Botchkarev VA, Fessing MY, Botchkareva NV,
Westgate G and Tobin DJ: First international symposium ‘Epigenetic
control of skin development and regeneration’: How Chromatin
regulators orchestrate skin functions. J Invest Dermatol.
133:1918–1921. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sobolev VV, Soboleva AG, Denisova EV,
Pechatnikova EA, Dvoryankova E, Korsunskaya IM and Mezentsev A:
Proteomic studies of psoriasis. Biomedicines. 10:6192022.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen C, Hou G, Zeng C, Ren Y, Chen X and
Peng C: Metabolomic profiling reveals amino acid and carnitine
alterations as metabolic signatures in psoriasis. Theranostics.
11:754–767. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Tett A, Pasolli E, Farina S, Truong DT,
Asnicar F, Zolfo M, Beghini F, Armanini F, Jousson O, De Sanctis V,
et al: Unexplored diversity and strain-level structure of the skin
microbiome associated with psoriasis. NPJ Biofilms Microbiomes.
3:142017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mahil SK, Capon F and Barker JN: Genetics
of psoriasis. Dermatol Clin. 33:1–11. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bowcock AM, Elder JT and Trembath R: The
international psoriasis genetics study: Assessing linkage to 14
candidate susceptibility loci in a cohort of 942 affected sib
pairs. Am J Human Genet. 73:430–437. 2003. View Article : Google Scholar
|
|
15
|
Allen MH, Ameen H, Veal C, Evans J,
Ramrakha-Jones VS, Marsland AM, Burden AD, Griffiths CE, Trembath
RC and Barker JN: The major psoriasis susceptibility locus PSORS1
is not a risk factor for Late-onset psoriasis. J Invest Dermatol.
124:103–106. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Nair RP, Henseler T, Jenisch S, Stuart P,
Bichakjian CK, Lenk W, Westphal E, Guo SW, Christophers E, Voorhees
JJ and Elder JT: Evidence for two psoriasis susceptibility loci
(HLA and 17q) and two novel candidate regions (16q and 20p) by
Genome-wide scan. Hum Mol Genet. 6:1349–1356. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zalesak M, Danisovic L and Harsanyi S:
Psoriasis and psoriatic Arthritis-associated genes, cytokines, and
human leukocyte antigens. Medicina (Kaunas). 60:8152024. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Owczarek W: The role of HLA-Cw6 in
psoriasis and psoriatic arthritis. Reumatologia. 60:303–305. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Dand N, Stuart PE, Bowes J, Ellinghaus D,
Nititham J, Saklatvala JR, Teder-Laving M, Thomas LF, Traks T, Uebe
S, et al: GWAS meta-analysis of psoriasis identifies new
susceptibility alleles impacting disease mechanisms and therapeutic
targets. Nat Commun. 16:20512025. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhang XJ, Huang W, Yang S, Sun LD, Zhang
FY, Zhu QX, Zhang FR, Zhang C, Du WH, Pu XM, et al: Psoriasis
genome-wide association study identifies susceptibility variants
within LCE gene cluster at 1q21. Nat Genet. 41:205–210. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yin X, Low HQ, Wang L, Li Y, Ellinghaus E,
Han J, Estivill X, Sun L, Zuo X, Shen C, et al: Genome-wide
meta-analysis identifies multiple novel associations and ethnic
heterogeneity of psoriasis susceptibility. Nat Commun. 6:69162015.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Xiang Z, Yang Y, Chang C and Lu Q: The
epigenetic mechanism for discordance of autoimmunity in monozygotic
twins. J Autoimmun. 83:43–50. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Pan J, Chen S, Chen X, Song Y and Cheng H:
Histone modifications and DNA methylation in psoriasis: A cellular
perspective. Clin Rev Allergy Immunol. 68:62025. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang M, Hu T, Ma T, Huang W and Wang Y:
Epigenetics and environmental health. Front Med. 18:571–596. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Siegfried Z and Simon I: DNA methylation
and gene expression. Wiley Interdiscip Rev Syst Biol Med.
2:362–371. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zeng C, Tsoi LC and Gudjonsson JE:
Dysregulated epigenetic modifications in psoriasis. Exp Dermatol.
30:1156–1166. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ichiyama K, Chen T, Wang X, Yan X, Kim BS,
Tanaka S, Ndiaye-Lobry D, Deng Y, Zou Y, Zheng P, et al: The
methylcytosine dioxygenase Tet2 promotes DNA demethylation and
activation of cytokine gene expression in T cells. Immunity.
42:613–626. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Moore LD, Le T and Fan G: DNA methylation
and its basic function. Neuropsychopharmacology. 38:23–38. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li X, Zhao X, Xing J, Li J, He F, Hou R,
Wang Q, Yin G, Li X and Zhang K: Different epigenome regulation and
transcriptome expression of CD4+ and CD8+ T cells from monozygotic
twins discordant for psoriasis. Australas J Dermatol. 61:e388–e394.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Han J, Park SG, Bae JB, Choi J, Lyu JM,
Park SH, Kim HS, Kim YJ, Kim S and Kim TY: The characteristics of
genome-wide DNA methylation in naïve CD4+ T cells of patients with
psoriasis or atopic dermatitis. Biochem Biophys Res Commun.
422:157–163. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Park GT, Han J, Park SG, Kim S and Kim TY:
DNA methylation analysis of CD4+ T cells in patients with
psoriasis. Arch Dermatol Res. 306:259–268. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Natoli V, Charras A, Hofmann SR, Northey
S, Russ S, Schulze F, McCann L, Abraham S and Hedrich CM: DNA
methylation patterns in CD4+ T-cells separate psoriasis patients
from healthy controls, and skin psoriasis from psoriatic arthritis.
Front Immunol. 14:12458762023. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Chen M, Chen ZQ, Cui PG, Yao X, Li YM, Li
AS, Gong JQ and Cao YH: The methylation pattern of p16INK4a gene
promoter in psoriatic epidermis and its clinical significance. Br J
Dermatol. 158:987–993. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhou F, Shen C, Xu J, Gao J, Zheng X, Ko
R, Dou J, Cheng Y, Zhu C, Xu S, et al: Epigenome-wide association
data implicates DNA methylation-mediated genetic risk in psoriasis.
Clin Epigenetics. 8:1312016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhou F, Wang W, Shen C, Li H, Zuo X, Zheng
X, Yue M, Zhang C, Yu L, Chen M, et al: Epigenome-wide association
analysis identified nine skin DNA methylation loci for psoriasis. J
Invest Dermatol. 136:779–787. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Terui T, Ozawa M and Tagami H: Role of
neutrophils in induction of acute inflammation in T-cell-mediated
immune dermatosis, psoriasis: A neutrophil-associated
inflammation-boosting loop. Exp Dermatol. 9:1–10. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Cao J and Yan Q: Histone ubiquitination
and deubiquitination in transcription, DNA damage response, and
cancer. Front Oncol. 2:201502012. View Article : Google Scholar
|
|
38
|
Zeng J, Zhang Y, Zhang H, Zhang Y, Gao L,
Tong X, Xie Y, Hu Q, Chen C, Ding S and Lu J: RPL22 Overexpression
promotes Psoriasis-like lesion by inducing keratinocytes abnormal
biological behavior. Front Immunol. 12:6999002021. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Liao Y, Su Y, Wu R, Zhang P and Feng C:
Overexpression of Wilms tumor 1 promotes IL-1β expression by
upregulating histone acetylation in keratinocytes. Int
Immunopharmacol. 96:1077932021. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Sestito R, Madonna S, Scarponi C,
Cianfarani F, Failla CM, Cavani A, Girolomoni G and Albanesi C:
STAT3-dependent effects of IL-22 in human keratinocytes are
counterregulated by sirtuin 1 through a direct inhibition of STAT3
acetylation. FASEB J. 25:916–927. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Li H, Yao Q, Mariscal AG, Wu X, Hülse J,
Pedersen E, Helin K, Waisman A, Vinkel C, Thomsen SF, et al:
Epigenetic control of IL-23 expression in keratinocytes is
important for chronic skin inflammation. Nat Commun. 9:14202018.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zhang T, Yang L, Ke Y, Lei J, Shen S, Shao
S, Zhang C, Zhu Z, Dang E and Wang G: EZH2-dependent epigenetic
modulation of histone H3 lysine-27 contributes to psoriasis by
promoting keratinocyte proliferation. Cell Death Dis. 11:8262020.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Chen C, Yi X, Liu P, Li J, Yan B, Zhang D,
Zhu L, Yu P, Li L, Zhang J, et al: CD147 facilitates the
pathogenesis of psoriasis through glycolysis and H3K9me3
modification in keratinocytes. Research (Wash D C).
6:01672023.PubMed/NCBI
|
|
44
|
Liu R, Zhang L and Zhang K: Histone
modification in psoriasis: Molecular mechanisms and potential
therapeutic targets. Exp Dermatol. 33:e151512024. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Ovejero-Benito MC, Reolid A,
Sánchez-Jiménez P, Saiz-Rodríguez M, Muñoz-Aceituno E,
Llamas-Velasco M, Martín-Vilchez S, Cabaleiro T, Román M, Ochoa D,
et al: Histone modifications associated with biological drug
response in moderate-to-severe psoriasis. Exp Dermatol.
27:1361–1371. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Xia X, Cao G, Sun G, Zhu L, Tian Y, Song
Y, Guo C, Wang X, Zhong J, Zhou W, et al: GLS1-mediated
glutaminolysis unbridled by MALT1 protease promotes psoriasis
pathogenesis. J Clin Invest. 130:5180–5196. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Koenen HJPM, Smeets RL, Vink PM, Van
Rijssen E, Boots AMH and Joosten I: Human CD25highFoxp3pos
regulatory T cells differentiate into IL-17 producing cells. Blood.
112:2340–2352. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Rasheed H, El-Komy MHM, Hegazy RA, Gawdat
HI, AlOrbani AM and Shaker OG: Expression of sirtuins 1, 6, tumor
necrosis factor, and interferon-γ in psoriatic patients. Int J
Immunopathol Pharmacol. 29:764–768. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Hwang YJ, Na JI, Byun SY, Kwon SH, Yang
SH, Lee HS, Choi HR, Cho S, Youn SW, Park KC, et al: Histone
deacetylase 1 and sirtuin 1 expression in psoriatic skin: A
comparison between guttate and plaque psoriasis. Life (Basel).
10:1572020.PubMed/NCBI
|
|
50
|
Liu Z, Cao W, Xu L, Chen X, Zhan Y, Yang
Q, Liu S, Chen P, Jiang Y, Sun X, et al: The histone H3 lysine-27
demethylase Jmjd3 plays a critical role in specific regulation of
Th17 cell differentiation. J Mol Cell Biol. 7:505–516. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Reolid A, Muñoz-Aceituno E, Abad-Santos F,
Ovejero-Benito MC and Daudén E: Epigenetics in Non-tumor
Immune-mediated skin diseases. Mol Diagn Ther. 25:137–161. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Djebali S, Davis CA, Merkel A, Dobin A,
Lassmann T, Mortazavi A, Tanzer A, Lagarde J, Lin W, Schlesinger F,
et al: Landscape of transcription in human cells. Nature.
489:101–108. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Shi R, Ma R, Jiang X, Tang X, Gong Y, Yu Z
and Shi Y: Implications of LncRNAs and CircRNAs in psoriasis: A
review. RNA Biol. 20:334–347. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Sonkoly E, Bata-Csorgo Z, Pivarcsi A,
Polyanka H, Kenderessy-Szabo A, Molnar G, Szentpali K, Bari L,
Megyeri K, Mandi Y, et al: Identification and characterization of a
novel, psoriasis susceptibility-related noncoding RNA gene, PRINS.
J Biol Chem. 280:24159–24167. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Jiang X, Shi R, Ma R, Tang X, Gong Y, Yu Z
and Shi Y: The role of microRNA in psoriasis: A review. Exp
Dermatol. 32:1598–1612. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Ghosh D, Ganguly T and Chatterjee R:
Emerging roles of non-coding RNAs in psoriasis pathogenesis. Funct
Integr Genomics. 23:1292023. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Bigler J, Rand HA, Kerkof K, Timour M and
Russell CB: Cross-study homogeneity of psoriasis gene expression in
skin across a large expression range. PLoS One. 8:e522422013.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Quaranta M, Knapp B, Garzorz N, Mattii M,
Pullabhatla V, Pennino D, Andres C, Traidl-Hoffmann C, Cavani A,
Theis FJ, et al: Intraindividual genome expression analysis reveals
a specific molecular signature of psoriasis and eczema. Sci Transl
Med. 6:244ra902014. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Palau N, Julià A, Ferrándiz C, Puig L,
Fonseca E, Fernández E, López-Lasanta M, Tortosa R and Marsal S:
Genome-wide transcriptional analysis of T cell activation reveals
differential gene expression associated with psoriasis. BMC
Genomics. 14:8252013. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Liu Y, Wang H, Cook C, Taylor MA, North
JP, Hailer A, Shou Y, Sadik A, Kim E, Purdom E, et al: Defining
Patient-level molecular heterogeneity in psoriasis vulgaris based
on Single-Cell transcriptomics. Front Immunol. 13:8426512022.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Guo D, Li X, Wang J, Liu X, Wang Y, Huang
S and Dang N: Single-cell RNA-seq reveals keratinocyte and
fibroblast heterogeneity and their crosstalk via
epithelial-mesenchymal transition in psoriasis. Cell Death Dis.
15:2072024. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Ma F, Plazyo O, Billi AC, Tsoi LC, Xing X,
Wasikowski R, Gharaee-Kermani M, Hile G, Jiang Y, Harms PW, et al:
Single cell and spatial sequencing define processes by which
keratinocytes and fibroblasts amplify inflammatory responses in
psoriasis. Nat Commun. 14:34552023. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Solvin ÅØ, Chawla K, Jenssen M, Olsen CL,
Furberg AS, Danielsen K, Saunes M, Hveem K, Sætrom P and Løset M:
Meta-analysis of RNA sequencing data from 534 skin samples shows
substantial IL-17 effects in non-lesional psoriatic skin. medRxiv.
Nov 04–2023.doi: 10.1101/2023.11.03.23298021.
|
|
64
|
Castillo RL, Sidhu I, Dolgalev I, Chu T,
Prystupa A, Subudhi I, Yan D, Konieczny P, Hsieh B, Haberman RH, et
al: Spatial transcriptomics stratifies psoriatic disease severity
by emergent cellular ecosystems. Sci Immunol. 8:eabq79912023.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Francis L, McCluskey D, Ganier C, Jiang T,
Du-Harpur X, Gabriel J, Dhami P, Kamra Y, Visvanathan S, Barker JN,
et al: Single-cell analysis of psoriasis resolution demonstrates an
inflammatory fibroblast state targeted by IL-23 blockade. Nat
Commun. 15:9132024. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Kim J, Lee J, Lee J, Kim K, Li X, Zhou W,
Cao J and Krueger JG: Psoriasis harbors multiple pathogenic type 17
T-cell subsets: Selective modulation by risankizumab. J Allergy
Clin Immunol. 155:1898–1912. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Johansen C, Rittig AH, Mose M, Bertelsen
T, Weimar I, Nielsen J, Andersen T, Rasmussen TK, Deleuran B and
Iversen L: STAT2 is involved in the pathogenesis of psoriasis by
promoting CXCL11 and CCL5 production by keratinocytes. PLoS One.
12:e01769942017. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Hald A, Andrés RM, Salskov-Iversen ML,
Kjellerup RB, Iversen L and Johansen C: STAT1 expression and
activation is increased in lesional psoriatic skin. Br J Dermatol.
168:302–310. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Lizzul PF, Aphale A, Malaviya R, Sun Y,
Masud S, Dombrovskiy V and Gottlieb AB: Differential expression of
phosphorylated NF-kappaB/RelA in normal and psoriatic epidermis and
downregulation of NF-kappaB in response to treatment with
etanercept. J Invest Dermatol. 124:1275–1283. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Bai L, Fang H, Xia S, Zhang R, Li L,
Ochando J, Xu J and Ding Y: STAT1 activation represses IL-22 gene
expression and psoriasis pathogenesis. Biochem Biophys Res Commun.
501:563–569. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Liang Y, Han D, Zhang S and Sun L: FOSL1
regulates hyperproliferation and NLRP3-mediated inflammation of
psoriatic keratinocytes through the NF-kB signaling via
transcriptionally activating TRAF3. Biochim Biophys Acta Mol Cell
Res. 1871:1196892024. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Zeng F, Liu H, Lu D, Liu Q, Chen H and
Zheng F: Integrated analysis of gene expression profiles identifies
transcription factors potentially involved in psoriasis
pathogenesis. J Cell Biochem. 120:12582–12594. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Xue X, Soroosh P, De Leon-Tabaldo A,
Luna-Roman R, Sablad M, Rozenkrants N, Yu J, Castro G, Banie H,
Fung-Leung WP, et al: Pharmacologic modulation of RORγt translates
to efficacy in preclinical and translational models of psoriasis
and inflammatory arthritis. Sci Rep. 6:379772016. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Skepner J, Ramesh R, Trocha M, Schmidt D,
Baloglu E, Lobera M, Carlson T, Hill J, Orband-Miller LA, Barnes A,
et al: Pharmacologic inhibition of RORγt regulates Th17 signature
gene expression and suppresses cutaneous inflammation in vivo. J
Immunol. 192:2564–2575. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Leppkes M, Becker C, Ivanov II, Hirth S,
Wirtz S, Neufert C, Pouly S, Murphy AJ, Valenzuela DM, Yancopoulos
GD, et al: RORγ-Expressing Th17 cells induce murine chronic
intestinal inflammation via redundant effects of IL-17A and IL-17F.
Gastroenterology. 136:257–267. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Tang L, Yang X, Liang Y, Xie H, Dai Z and
Zheng G: Transcription factor retinoid-related orphan receptor γt:
A promising target for the treatment of psoriasis. Front Immunol.
9:12102018. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Vanaki E, Ataei M, Sanati M, Mansouri P,
Mahmoudi M, Zarei F and Jadali Z: Expression patterns of Th1/Th2
transcription factors in patients with guttate psoriasis. Acta
Microbiol Immunol Hung. 60:163–174. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Endo H, Momota Y, Oikawa A and Shinkai H:
Psoriatic skin expresses the transcription factor Gli1: Possible
contribution of decreased neurofibromin expression. Br J Dermatol.
154:619–623. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Wang Z, Sun Y, Lou F, Bai J, Zhou H, Cai
X, Sun L, Yin Q, Tang S, Wu Y, et al: Targeting the transcription
factor HES1 by L-menthol restores protein phosphatase 6 in
keratinocytes in models of psoriasis. Nat Commun. 13:78152022.
View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Dragan M, Sun P, Chen Z, Ma X, Vu R, Shi
Y, Villalta SA and Dai X: Epidermis-intrinsic transcription factor
ovol1 coordinately regulates barrier maintenance and neutrophil
accumulation in psoriasis-like inflammation. J Invest Dermatol.
142:583–593.e5. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Goldminz AM, Au SC, Kim N, Gottlieb AB and
Lizzul PF: NF-κB: An essential transcription factor in psoriasis. J
Dermatol Sci. 69:89–94. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Sobolev VV, Tchepourina E, Korsunskaya IM,
Geppe NA, Chebysheva SN, Soboleva AG and Mezentsev A: The role of
transcription factor PPAR-γ in the pathogenesis of psoriasis, skin
cells, and immune cells. Int J Mol Sci. 23:97082022. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
He C, Song TC, Qi RQ and Gao XH:
Integrated single-cell and spatial transcriptomics reveals
heterogeneity of fibroblast and pivotal genes in psoriasis. Sci
Rep. 13:171342023. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Yuan X, Xin T, Yu H, Huang J, Xu Y, Ou C
and Chen Y: Transcription Factor IRF7 is involved in psoriasis
development and response to guselkumab treatment. J Inflamm Res.
17:1039–1055. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Morizane S and Gallo RL: Antimicrobial
peptides in the pathogenesis of psoriasis. J Dermatol. 39:225–230.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Pfaff CM, Marquardt Y, Fietkau K, Baron JM
and Lüscher B: The psoriasis-associated IL-17A induces and
cooperates with IL-36 cytokines to control keratinocyte
differentiation and function. Sci Reps. 7:156312017. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Nestle FO, Kaplan DH and Barker J:
Psoriasis. N Engl J Med. 361:496–509. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Kromann B, Olsson A, Zhang YM, Løvendorf
MB, Skov L and Dyring-Andersen B: Proteins in the skin and blood in
patients with psoriasis: A systematic review of proteomic studies.
Dermatology. 240:317–328. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Wang H, Wang C, Qin R, He J, Zhang X, Ma
C, Li S, Fan L, Wang L and Cao L: Integrative analysis of plasma
proteomics and transcriptomics reveals potential therapeutic
targets for psoriasis. Biomedicines. 13:13802025. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Schonthaler HB, Guinea-Viniegra J, Wculek
SK, Ruppen I, Ximénez-Embún P, Guío-Carrión A, Navarro R, Hogg N,
Ashman K and Wagner EF: S100A8-S100A9 protein complex mediates
psoriasis by regulating the expression of complement factor C3.
Immunity. 39:1171–1181. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Zhang B and Wu F: Proteomic identification
of exosomes derived from psoriasis cells using data-independent
acquisition mass spectrometry. Arch Dermatol Res. 316:2242024.
View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Møller LBP, Kromann B, Kabatnik S,
Hjortlund JH, Haulrig MB, Sølberg JBK, Bzorek M, Clark RA, Skov L,
Mann M, et al: Spatial proteomic profiling reveals increased levels
of cholesterol synthesis proteins in psoriasis vulgaris. J Invest
Dermatol. 145:3051–3063.e2. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Lu C, Yang F, He S, Yu H, Wang Q, Li M,
Zeng X and Leng X: Serum proteome analysis identifies a potential
biomarker for axial psoriatic arthritis. Eur J Med Res. 29:1462024.
View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Cai YX, Zheng DS, Chen XL, Bai ZP, Zhang
J, Deng W and Huang XF: An integrated multi-omics analysis
identifies protein biomarkers and potential drug targets for
psoriatic arthritis. Commun Biol. 8:2402025. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Liu S, Li L, Liang Y, Tan Y, Wang X, Feng
Y, Chen N and Lei X: Novel genetic insight for psoriasis:
Integrative genome-wide analyses in 863 080 individuals and
proteome-wide Mendelian randomization. Brief Bioinform.
26:bbaf0322024. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Qiao P, Zhang C, Yu J, Shao S, Zhang J,
Fang H, Chen J, Luo Y, Zhi D, Li Q, et al: Quinolinic acid, a
tryptophan metabolite of the skin microbiota, negatively regulates
NLRP3 inflammasome through AhR in psoriasis. J Invest Dermatol.
142:2184–2193.e6. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Kang H, Li X, Zhou Q, Quan C, Xue F, Zheng
J and Yu Y: Exploration of candidate biomarkers for human psoriasis
based on gas chromatography-mass spectrometry serum metabolomics.
Br J Dermatol. 176:713–722. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Li SS, Liu Y, Li H, Wang LP, Xue LF, Yin
GS and Wu XS: Identification of psoriasis vulgaris biomarkers in
human plasma by non-targeted metabolomics based on UPLC-Q-TOF/MS.
Eur Rev Med Pharmacol Sci. 23:3940–3950. 2019.PubMed/NCBI
|
|
99
|
Madsen RK, Lundstedt T, Gabrielsson J,
Sennbro CJ, Alenius GM, Moritz T, Rantapää-Dahlqvist S and Trygg J:
Diagnostic properties of metabolic perturbations in rheumatoid
arthritis. Arthritis Res Ther. 13:R192011. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Li L, Chuan-Jian L, Ling H, Jing-Wen D,
Ze-Hui H, Yu-Hong Y and Zhong-Zhao Z: Untargeted serum metabonomics
study of psoriasis vulgaris based on ultra-performance liquid
chromatography coupled to mass spectrometry. Oncotarget.
8:95931–95944. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Coras R, Kavanaugh A, Boyd T, Huynh D,
Lagerborg KA, Xu YJ, Rosenthal SB, Jain M and Guma M: Choline
metabolite, trimethylamine N-oxide (TMAO), is associated with
inflammation in psoriatic arthritis. Clin Exp Rheumatol.
37:481–484. 2019.PubMed/NCBI
|
|
102
|
Kamleh MA, Snowden SG, Grapov D, Blackburn
GJ, Watson DG, Xu N, Ståhle M and Wheelock CE: LC-MS metabolomics
of psoriasis patients reveals disease severity-dependent increases
in circulating amino acids that are ameliorated by anti-TNFα
treatment. J Proteome Res. 14:557–566. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Dutkiewicz EP, Hsieh KT, Wang YS, Chiu HY
and Urban PL: Hydrogel micropatch and mass spectrometry-assisted
screening for Psoriasis-Related skin metabolites. Clin Chem.
62:1120–1128. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Xiong Q, Zhong D, Li Q, Yu Y, Zhang S,
Liang J and Zhang X: LC-MS metabolomics reveal skin metabolic
signature of psoriasis vulgaris. Exp Dermatol. 32:889–899. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Takeichi T, Kinoshita F, Tanaka H, Fujita
S, Kobayashi Y, Nakatochi M, Sugiura K and Akiyama M: The
lipoxygenase-hepoxilin pathway is activated in cutaneous plaque
lesions of psoriasis. J Cutaneous Immunol Allergy. 2:15–24. 2019.
View Article : Google Scholar
|
|
106
|
Łuczaj W, Wroński A, Domingues P, Rosário
Domingues M and Skrzydlewska E: Lipidomic analysis reveals specific
differences between fibroblast and keratinocyte ceramide profile of
patients with psoriasis vulgaris. Molecules. 25:6302020. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Setkowicz M, Mastalerz L, Gielicz A,
Wojas-Pelc A and Sanak M: Lack of association of ALOX12 and ALOX15B
polymorphisms with psoriasis despite altered urinary excretion of
12(S)-hydroxyeicosatetraenoic acid. Br J Dermatol. 172:337–344.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Alonso A, Julià A, Vinaixa M, Domènech E,
Fernández-Nebro A, Cañete JD, Ferrándiz C, Tornero J, Gisbert JP,
Nos P, et al: Urine metabolome profiling of immune-mediated
inflammatory diseases. BMC Med. 14:1332016. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Lunjani N, Ahearn-Ford S, Dube FS, Hlela C
and O'Mahony L: Mechanisms of microbe-immune system dialogue within
the skin. Genes Immun. 22:276–288. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Hurabielle C, Link VM, Bouladoux N, Han
SJ, Merrill ED, Lightfoot YL, Seto N, Bleck CKE, Smelkinson M,
Harrison OJ, et al: Immunity to commensal skin fungi promotes
psoriasiform skin inflammation. Proc Natl Acad Sci USA.
117:16465–16474. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Fyhrquist N, Muirhead G, Prast-Nielsen S,
Jeanmougin M, Olah P, Skoog T, Jules-Clement G, Feld M,
Barrientos-Somarribas M, Sinkko H, et al: Microbe-host interplay in
atopic dermatitis and psoriasis. Nat Commun. 10:47032019.
View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Grice EA, Kong HH, Conlan S, Deming CB,
Davis J, Young AC; NISC Comparative Sequencing Program, ; Bouffard
GG, Blakesley RW, Murray PR, et al: Topographical and temporal
diversity of the human skin microbiome. Science. 324:1190–1192.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Quan C, Chen XY, Li X, Li X, Xue F, Chen
LH, Liu N, Wang B, Wang LQ, Wang XP, et al: Psoriatic lesions are
characterized by higher bacterial load and imbalance between
Cutibacterium and Corynebacterium. J Am Acad Dermatol. 82:955–961.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Olejniczak-Staruch I, Ciążyńska M,
Sobolewska-Sztychny D, Narbutt J, Skibińska M and Lesiak A:
Alterations of the skin and gut microbiome in psoriasis and
psoriatic arthritis. Int J Mol Sci. 22:39982021. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Gao Z, Tseng CH, Strober BE, Pei Z and
Blaser MJ: Substantial alterations of the cutaneous bacterial biota
in psoriatic lesions. PLoS One. 3:e27192008. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Paulino LC, Tseng CH, Strober BE and
Blaser MJ: Molecular analysis of fungal microbiota in samples from
healthy human skin and psoriatic lesions. J Clin Microbiol.
44:2933–2941. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Gao T, Wang X, Li Y and Ren F: The role of
probiotics in skin health and related gut-skin axis: A review.
Nutrients. 15:31232023. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Salem I, Ramser A, Isham N and Ghannoum
MA: The gut microbiome as a major regulator of the gut-skin axis.
Front Microbiol. 9:3826982018. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Sikora M, Stec A, Chrabaszcz M,
Waskiel-Burnat A, Zaremba M, Olszewska M and Rudnicka L: Intestinal
fatty acid binding protein, a biomarker of intestinal barrier, is
associated with severity of psoriasis. J Clin Med. 8:10212019.
View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Sikora M, Chrabąszcz M, Waśkiel-Burnat A,
Rakowska A, Olszewska M and Rudnicka L: Claudin-3-a new intestinal
integrity marker in patients with psoriasis: Association with
disease severity. J Eur Acad Dermatol Venereol. 33:1907–1912. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Sikora M, Stec A, Chrabaszcz M, Knot A,
Waskiel-Burnat A, Rakowska A, Olszewska M and Rudnicka L: Gut
microbiome in psoriasis: An updated review. Pathogens. 9:4632020.
View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Hidalgo-Cantabrana C, Gómez J, Delgado S,
Requena-López S, Queiro-Silva R, Margolles A, Coto E, Sánchez B and
Coto-Segura P: Gut microbiota dysbiosis in a cohort of patients
with psoriasis. Br J Dermatol. 181:1287–1295. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Chen YJ, Ho HJ, Tseng CH, Lai ZL, Shieh JJ
and Wu CY: Intestinal microbiota profiling and predicted metabolic
dysregulation in psoriasis patients. Exp Dermatol. 27:1336–1343.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Tan LR, Zhao S, Zhu W, Wu L, Li J, Shen M,
Lei L, Chen X and Peng C: The Akkermansia muciniphila is a gut
microbiota signature in psoriasis. Exp Dermatol. 27:144–149. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Yerushalmi M, Elalouf O, Anderson M and
Chandran V: The skin microbiome in psoriatic disease: A systematic
review and critical appraisal. J Transl Autoimmun. 2:1000092019.
View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Guo H, Gao J, Gong L and Wang Y:
Multi-omics analysis reveals novel causal pathways in psoriasis
pathogenesis. J Transl Med. 23:1002025. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Xing L, Wu T, Yu L, Zhou N, Zhang Z, Pu Y,
Wu J and Shu H: Exploration of biomarkers of psoriasis through
combined multiomics analysis. Mediators Inflamm. 2022:77310822022.
View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Deng J, Leijten E, Zhu Y, Nordkamp MO, Ye
S, Pouw J, Tao W, Balak D, Zheng G, Radstake T, et al: Multi-omics
approach identifies PI3 as a biomarker for disease severity and
hyper-keratinization in psoriasis. J Dermatol Sci. 111:101–108.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Glasebach H, Rupp S and Burger-Kentischer
A: A standardizable human-based psoriasis skin model for drug
development. Front Med (Lausanne). 12:15394842025. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Tamari M, Saeki H, Hayashi M, Umezawa Y,
Ito T, Fukuchi O, Nobeyama Y, Yanaba K, Nakagawa H, Tsunemi Y, et
al: An association study of 36 psoriasis susceptibility loci for
psoriasis vulgaris and atopic dermatitis in a Japanese population.
J Dermatol Sci. 76:156–157. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Nasser A El and Hamid N: Unraveling the
genetic landscape of psoriasis: A genome-wide association study in
Egypt. Scientific J Dermatol Venereol. 3:24–38. 2025.
|
|
132
|
Yang JS, Liu TY, Lu HF, Tsai SC, Liao WL,
Chiu YJ, Wang YW and Tsai FJ: Genome-wide association study and
polygenic risk scores predict psoriasis and its shared phenotypes
in Taiwan. Mol Med Rep. 30:1–22. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Elinghaus E, Elinghaus D, Stuart PE, Nair
RP, Debrus S, Raelson JV, Belouchi M, Fournier H, Reinhard C, Ding
J, et al: Genome-wide association study identifies a psoriasis
susceptibility locus at TRAF3IP2. Nat Genet. 42:991–995. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Zhou X, Zhou H, Luo X and Wu RF: Discovery
of biomarkers in the psoriasis through machine learning and dynamic
immune infiltration in three types of skin lesions. Front Immunol.
15:13886902024. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Yang Y, Xie S, Jiang W, Tang S and Shi Y:
Discovering novel biomarkers associated with the pathogenesis of
psoriasis: Evidence from bioinformatic analysis. Int J Gen Med.
15:2817–2833. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Mao R, Zhang T, Yang Z and Li J: Unveiling
novel protein biomarkers for psoriasis through integrated analysis
of human plasma proteomics and mendelian randomization. Psoriasis
(Auckl). 14:179–193. 2024.PubMed/NCBI
|
|
137
|
Zhou Y, Wang P, Yan BX, Chen XY, Landeck
L, Wang ZY, Li XX, Zhang J, Zheng M and Man XY: Quantitative
proteomic profile of psoriatic epidermis identifies OAS2 as a novel
biomarker for disease activity. Front Immunol. 11:14322020.
View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Song Q, Chen Y, Ma J, Zhou W, Song J, Wu C
and Liu J: Metabolomics reveals molecular signatures for psoriasis
biomarkers and drug targets discovery. Clin Cosmet Investig
Dermatol. 16:3181–3191. 2023. View Article : Google Scholar : PubMed/NCBI
|