Introduction
Bladder cancer is among the most common malignancies
of the urinary system worldwide, with an estimated 614,000 new
cases and nearly 220,000 related deaths globally in 2022 (1). Bladder cancer is divided into
non-muscle-invasive (NMIBC) and muscle-invasive (MIBC) subtypes.
Although NMIBC has an improved prognosis, it necessitates costly
long-term monitoring, and ~20% of cases progress to MIBC. The high
incidence and recurrence rates of bladder cancer make it one of the
most challenging diseases in the field of urology. While early
detection and treatment can markedly improve patient outcomes, the
prognosis for those with invasive or metastatic disease remains
poor. Although traditional therapeutic approaches, such as surgery,
radiation therapy, and chemotherapy, are effective to some extent
in delaying disease progression, they are often limited by
significant side effects, suboptimal efficacy, and the development
of drug resistance (2,3). As a result, the development of novel
therapeutic agents to improve clinical outcomes for bladder cancer
patients, reduce disease recurrence, and mitigate treatment-related
toxicity is urgently needed.
Ferroptosis is an iron-dependent cell death
mechanism driven by lipid peroxidation and shows promise for
treating cancers resistant to conventional therapies. Nevertheless,
the therapeutic potential of ferroptosis induction in bladder
cancer remains unclear (4). The
identification of ferroptosis-inducing factors and the exploration
of their underlying mechanisms have emerged as promising research
directions. For instance, cirSIRT5 has been shown to promote
ferroptosis in bladder cancer (5)
and abietic acid was also reported to induce ferroptotic cell death
in this malignancy (6).
Reactive oxygen species (ROS) play a central role in
inducing ferroptosis by promoting lipid peroxidation, leading to
cellular damage and death (7). In
this context, the glutathione peroxidase 4 (GPX4) enzyme is a key
regulator, as it protects cells from ROS-induced lipid peroxidation
(8). The dysregulation of GPX4 has
been associated with increased susceptibility to ferroptosis,
making it a critical target in cancer therapy (9). Targeting the ferroptosis pathway,
particularly through the modulation of ROS levels and GPX4
activity, represents a promising therapeutic strategy for the
treatment of bladder cancer, as it offers a novel approach for
overcoming the limitations of conventional therapies and addressing
drug resistance (10).
Traditional Chinese medicine (TCM) and its bioactive
components have garnered significant attention because of their
role in modulating ferroptosis, offering new strategies for cancer
treatment (11). Among these
compounds, Paris polyphylla and its polyphyllin constituents
have been reported to exert antitumor effects by promoting
ferroptosis through various mechanisms. For example, research has
shown that polyphyllin I (PPI) can induce ferroptosis by
downregulating GPX4 expression and accumulating ROS, which triggers
inhibitory effects on the proliferation, invasion and metastasis of
hepatocellular carcinoma cells (12). Additionally, polyphyllin III has
been shown to deplete GPX4 levels, accumulate ROS and induce
ferroptosis in breast cancer cells (13). Despite these findings, the specific
role of polyphyllin II (PPII) in regulating ferroptosis in bladder
cancer remains unexplored. The sequencing results of the present
study suggested that PPII may regulate the ROS levels and
ferroptosis in bladder cancer. In addition, GPX4 is a key
regulatory factor for ferroptosis. Therefore, it was hypothesized
that PPII is likely to promote ROS accumulation, cause
Fe2+ overload by downregulating GPX4 levels, and
ultimately induce ferroptosis.
Materials and methods
Cell culture
MIBC (T24 and 5637) provided by Professor Shengtian
Zhao (First Clinical Medical College, Qilu Hospital of Shandong
University, Jinan, Shandong) were used in the present study. The
T24 and 5637 cell lines are both well-established models of MIBC
and share a common TP53 mutation background. Using two distinct
cell lines helps demonstrate that the observed pro-ferroptotic
effect of PPII is not an isolated phenomenon limited to a single
genetic context but can be reproduced across different models of
MIBC. The cells were cultured in RPMI-1640 medium (cat. no.
CM10041, Macgene Technology Ltd.) supplemented with 10% fetal
bovine serum (cat. no. 10270–106; Gibco; Thermo Fisher Scientific,
Inc.). The two cell lines were maintained at 37°C in a humidified
incubator with 5% CO2.
Cell viability assay
Cell viability was assessed using a CCK-8 assay kit
(E-CK-A362, Elabscience Bionovation Inc.). T24 (3×103)
and 5637 (6×103) cells were seeded into 96-well plates
and incubated at 37°C with 5% CO2 overnight, after which
the cells were treated with cisplatin or PPII. Following treatment,
10% CCK-8 solution was added to each well and gently mixed, and the
plates were then incubated for an additional 2 h. The absorbance
was measured at 450 nm using a spectrophotometer (Type 1510; Thermo
Fisher Scientific, Inc.).
Colony formation assay
To assess the proliferative capacity of the cells, a
colony formation assay was performed. Cells (800 cells/well)
treated with gradient concentrations of PPII (0–0.8 µM) were seeded
into six-well plates and cultured in medium supplemented with 10%
fetal bovine serum, after which they were allowed to grow at 37°C
with 5% CO2 for 10 days until visible colonies formed.
The cells were then gently washed with PBS and fixed with 4%
paraformaldehyde for 15 min at room temperature (RT). After
fixation, the colonies were stained with 0.1% crystal violet for 30
min at RT, followed by rinsing with water to remove excess dye. The
number of stained colonies was counted to evaluate the clonogenic
potential of the cells.
RNA-sequencing and data
pre-processing
T24 cells were treated with gradient concentrations
of PPII [0 (NC group), 0.2 (L group) and 0.4 (H group) µM] for 48 h
and total RNA were extracted with a MJzol animal RNA Extraction Kit
(cat. no. Majorbio) following by the manufacturer's protocol. RNA
quality and integrity were analyzed using a NanoPhotometer
spectrophotometer (Implen GmbH) and an Agilent 2100 Bioanalyzer
(Agilent Technologies, Inc.). To construct RNA-seq libraries,
ribosomal RNA (rRNA) was removed from total RNA, and the remaining
mRNA was then randomly fragmented. The RNA-seq libraries were
constructed using an Illumina Truseq RNA Sample Prep Kit (Illumina,
Inc.) and sequenced on a NovaSeq 6000 system (Illumina, Inc.).
Prior to bioinformatics analysis, FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/)
was used to assess the quality of the raw data and the raw data
were preprocessed to obtain high-quality clean read data. Cleaned
reads were then mapped to the human reference genome GRCh38/hg38
using the spliced-read aligner HISAT2 (14) and StringTie (15) to obtain raw read counts and
transcripts per million (TPM).
Gene expression and dynamic expression
model analyses
Raw read count data were used for gene expression
analyses. Genes with low counts might represent a sequencing bias
and contribute less to further analysis; thus, genes with zero
expression values were excluded. After data filtering, differential
expression analysis was performed with the Bioconductor package
DESeq2 (16). Any gene with a
P-value <0.05 and a fold change >1.25 was regarded as a
significantly differentially expressed gene (DEG).
TPM of DEGs were used for c-means clustering with
the R package Mfuzz to characterize dynamic changes in expression
patterns (17). Fuzzy c-means
(FCM) clustering is a soft clustering method performed with the
Mfuzz algorithm with two key parameters (c=number of clusters and
m=fuzzification parameter). The algorithm iteratively assigns the
profile to the cluster with the shortest Euclidean distance while
minimizing any objective function. In the present study, the data
were clustered with the parameters c=10 and m=2.
Pathway enrichment and Gene Ontology
(GO) analysis
GO functional enrichment and Kyoto Encyclopedia of
Genes and Genomes (KEGG) pathway analyses were performed using the
R package clusterProfiler (18).
GO terms were divided into three separate subgroups: Molecular
functions (MFs), cellular components (CCs) and biological processes
(BPs). Enriched GO terms and KEGG pathways were identified
according to the cutoff criterion of P-values <0.05.
Immunoblotting
Western blotting was carried out following described
protocols (19). Briefly, total
cellular proteins were extracted using RIPA lysis buffer (cat. no.
P0013C; Beyotime Biotechnology), and their concentrations were
quantified with a BCA assay kit (cat. no. P0010S; Beyotime
Biotechnology). An equal amount of protein (20 µg per lane) was
separated by 10% SDS-PAGE gel and subsequently electrophoretically
transferred onto a PVDF membrane (cat. no. IPVH00010;
MilliporeSigma). The membrane was blocked with 5% skimmed milk
powder in TBST buffer (0.1% Tween-20) at RT for 1 h and then probed
with the primary antibody at 4°C overnight. After washing, it was
incubated with an HRP-conjugated goat anti-mouse secondary antibody
(1:3,000 for GPX4 and 1:10,000 for ACTB; cat. no. A0216; Beyotime
Biotechnology) at RT for 1 h. Finally, the target protein bands
were visualized using an enhanced chemiluminescence (ECL)
substrate. The primary antibodies used in the present study
included GPX4 (mouse-sourced antibody, 1:1,000; cat. no.
67763-1-Ig) (20) purchased from
Proteintech Group, Inc. and ACTB (mouse-sourced antibody, 1:10,000;
cat. no. A5441) (21), which was
obtained from Sigma-Aldrich (Merck KGaA). The intensity of the
protein bands was quantified using ImageJ (version 1.53q; National
Institutes of Health).
ROS detection analysis
Intracellular ROS levels were determined using a
Reactive Oxygen Species Assay Kit (cat. no. S0033S; Beyotime
Biotechnology), with DCFH-DA as the primary reagent. In brief, T24
and 5637 cells were plated in 6-well plates and treated with
gradient concentrations of PPII (0, 0.4 and 0.8 µM for T24 cells
and 0, 0.85 and 1.70 µM for 5637 cells) for 48 h. After treatment,
the cells were incubated with DCFH-DA for 30 min at 37°C and the
nuclei were stained with Hoechst 33342 (cat. no. C1029; Beyotime
Biotechnology) for 10 min. The fluorescence was then visualized
using a fluorescence microscope (Vert.A1; Zeiss GmbH) and the mean
fluorescence intensity was quantified using ImageJ (version 1.53q;
National Institutes of Health).
Fe2+ detection
analysis
Fe2+ detection was performed according to
the FerroOrange manufacturer's instructions (cat. no. F374; Dojindo
Laboratories, Inc.). After 48 h of PPII treatment with various
concentrations of PPII (as described in the ROS detection analysis
section), the culture medium was removed and the BC cells were
washed three times with HBSS. The cells were then incubated with
FerroOrange working solution (1 µM) at 37°C in a 5% CO2 atmosphere
for 30 min. Changes in fluorescence intensity were observed using a
fluorescence microscope (Vert.A1; Zeiss GmbH).
Analysis of oxidative stress-related
markers
T24 and 5637 cells were treated as aforementioned,
followed by washing with PBS and lysis with lysis buffer. After
centrifugation (19,480 × g), the supernatant was collected for the
detection of malondialdehyde (MDA) levels using a MDA detection kit
(cat. no. S0131S; Beyotime Biotechnology). The absorbance at 532 nm
was measured with a spectrophotometer (Type 1510; Thermo Fisher
Scientific, Inc.). Similarly, superoxide dismutase (SOD) levels
were measured following the instructions of the corresponding
reagent kit (cat. no. S0101M; Beyotime Biotechnology), with the
absorbance recorded at 450 nm.
Detection of oxidized and non-oxidized
lipids
T24 and 5637 cells were treated with gradient
concentrations of PPII for 48 h as aforementioned. The medium was
then replaced with complete medium containing BODIPY 581/591 C11
(10 µM; cat. no. D3861; Invitrogen; Thermo Fisher Scientific, Inc.)
and the cells incubated at 37°C with 5% CO2 for an additional 30
min. The cells were then washed three times with PBS (5 min per
wash), followed by staining with Hoechst 33342 for 10 min at 37°C.
After the samples were washed, the fluorescence intensities of
oxidized and non-oxidized lipids were observed under a fluorescence
microscope (Vert.A1; Zeiss GmbH), and the mean fluorescence
intensity was quantified using ImageJ (version 1.53q; National
Institutes of Health).
Statistical analysis
All the statistical analyses were performed using
GraphPad Prism (version 5.0; Dotmatics). Data are presented as the
mean ± standard deviation (SD) from at least three independent
experiments. For comparisons between two groups, an unpaired
two-tailed Student's t-test was used. For comparisons among more
than two groups, one-way analysis of variance (ANOVA) was
performed, followed by Dunnett's test for multiple comparisons when
all groups were compared against a single control group. DEGs were
identified using an adjusted P-value (False Discovery Rate; FDR)
threshold. Furthermore, Pearson correlation analysis was conducted
using R software (version 4.3.0; http://cran.r-project.org/bin/windows/base/old/4.3.0/)
to assess the relationships between oxidative stress indicators.
P<0.05 was considered to indicate a statistically significant
difference.
Results
PPII inhibits the cell viability and
colony formation of BC cells
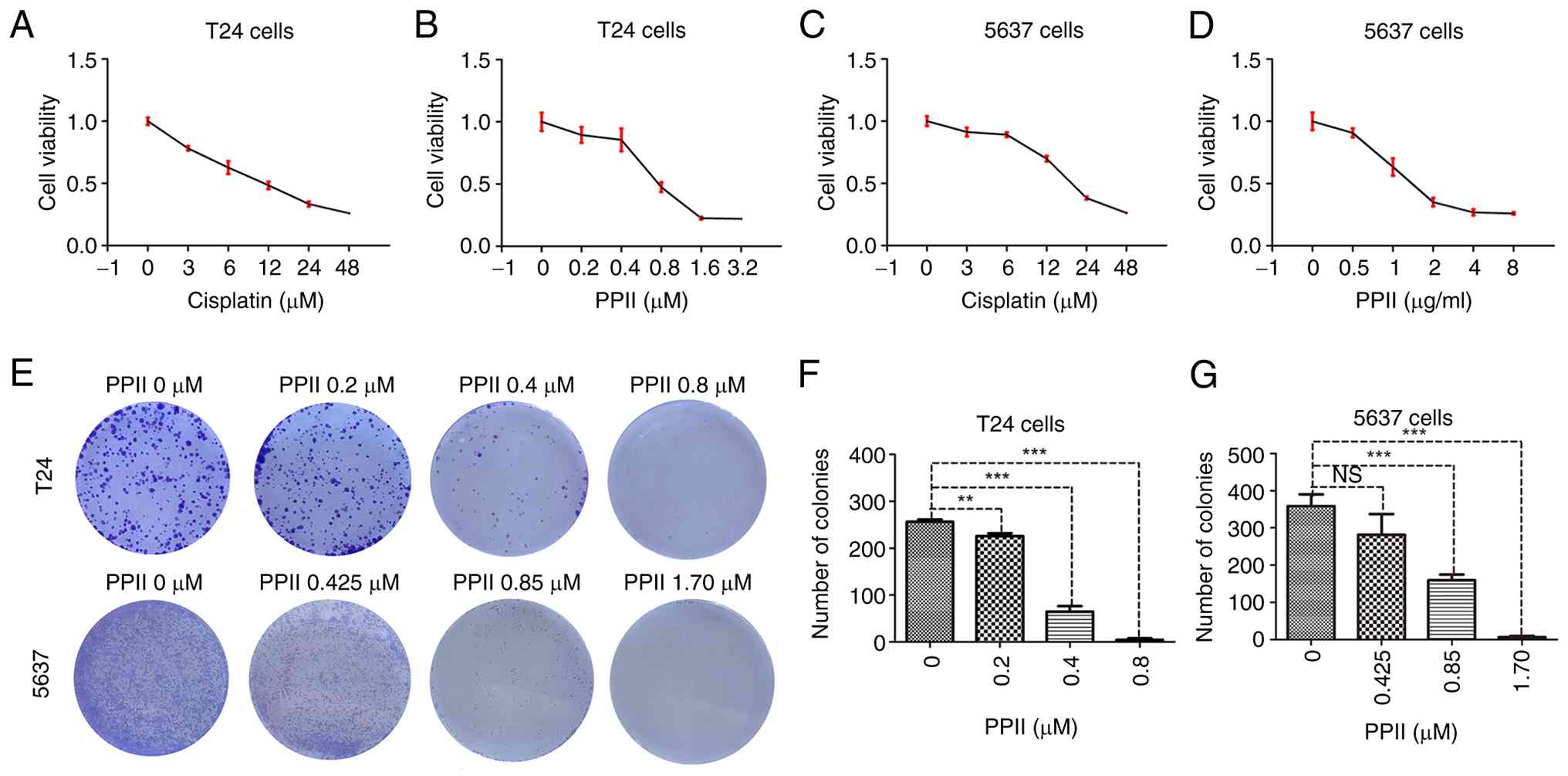
The effect of PPII on bladder cancer cell viability
was assessed using a CCK-8 assay. T24 and 5637 cells were treated
with increasing concentrations of cisplatin (used as a positive
control, Fig. 1A and C) or PPII
(Fig. 1B and D) for 48 h. The
CCK-8 results revealed that PPII could inhibited the viability of
bladder cancer cells and the calculated IC50 values of
cisplatin were 11.70±1.31 µM for T24 cells and 20.17±0.82 µM for
5637 cells, whereas the IC50 values of PPII were
0.86±0.09 µM for T24 cells and 1.72±0.21 µM for 5637 cells.
The effect of the PPII on the proliferative capacity
of bladder cancer cells was evaluated through colony formation
assays. The results suggested that PPII effectively reduced the
number of colonies formed by T24 and 5637 cells in a dose-dependent
manner (Fig. 1E). Statistical
analysis revealed that 0.2, 0.4, and 0.8 µM PPII significantly
decreased the number of colonies formed by T24 cells (P<0.01,
Fig. 1F). Similarly, PPII at
concentrations of 0.85 and 1.70 µM PPII markedly inhibited colony
formation in 5637 cells (P<0.001, Fig. 1G).
RNA sequencing analysis of BC cells
treated with PPII
Previous experiments had indicated that PPII
effectively inhibits the proliferation of BC cells. To minimize the
confusing effects of cytotoxicity from high concentrations of
drugs, the present study chose PPII drug concentrations (0, 0.2,
and 0.4 µM) that did not affect T24 cell viability for RNA
transcriptome sequencing (Fig.
S1A), aiming to elucidate the potential mechanisms involved in
this process.
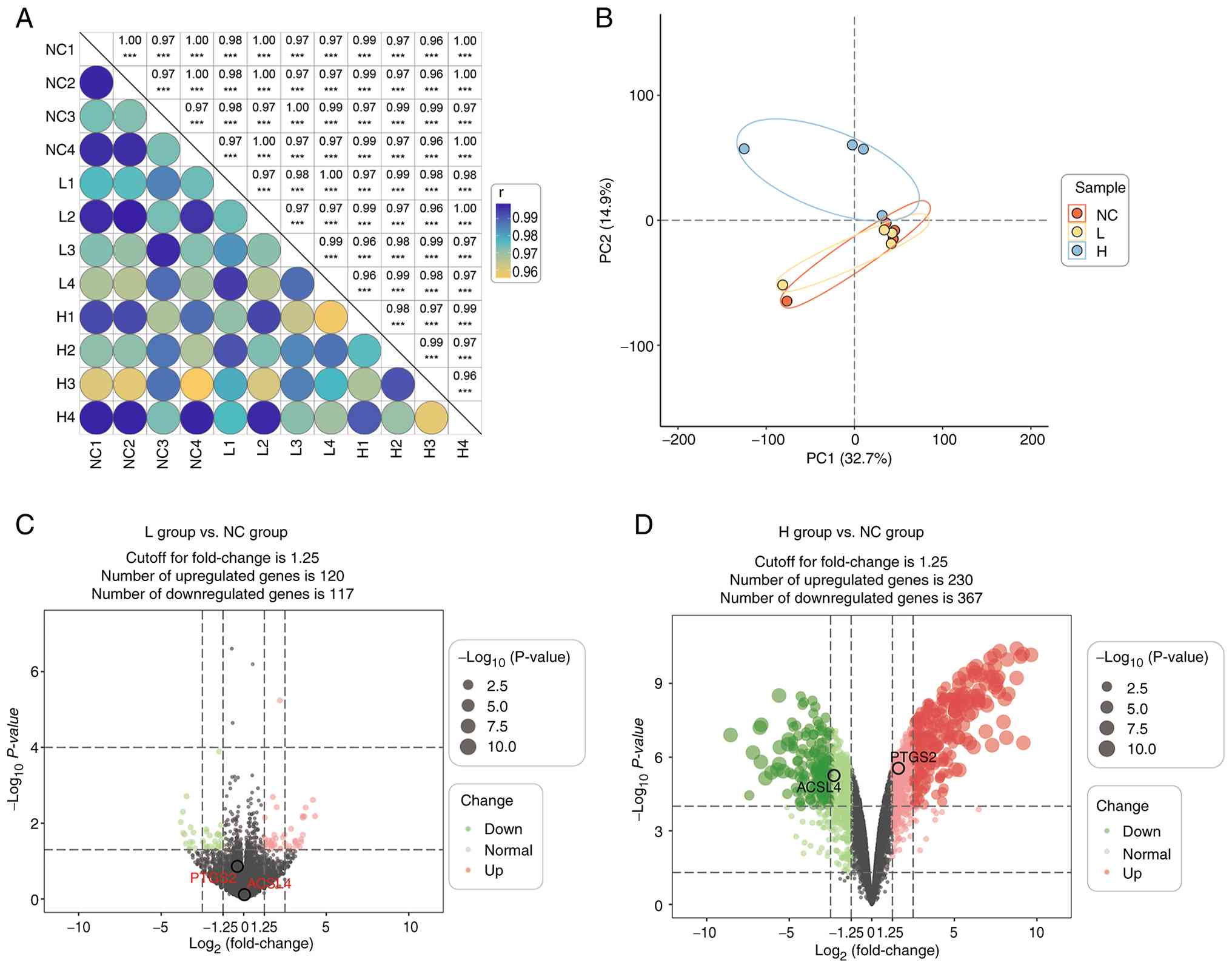
Overview of the transcriptomic
analysis
T24 cells were treated with high-concentration and
low-concentration PPII for 48 h, after which the RNA was harvested
in quadruplicate. A total of 12 samples were divided into three
groups, namely, the NC, L and H groups, according to the
concentrations of PPII (Fig. 2A).
DEGs between each group were identified using a P-value <0.05
and a fold change greater than the cutoff. A total of 834 DEGs were
identified between the NC, L and H groups. Principal components
analysis revealed that the between-group samples were clearly
separated from each other and revealed good clustering of samples
within the same group, indicating distinct gene expression profiles
upon PPII treatment (Fig. 2B).
Volcano plots also revealed the DEGs that differed from each other
in the groups (Fig. 2C and D). All
the DEGs were included in the subsequent analyses.
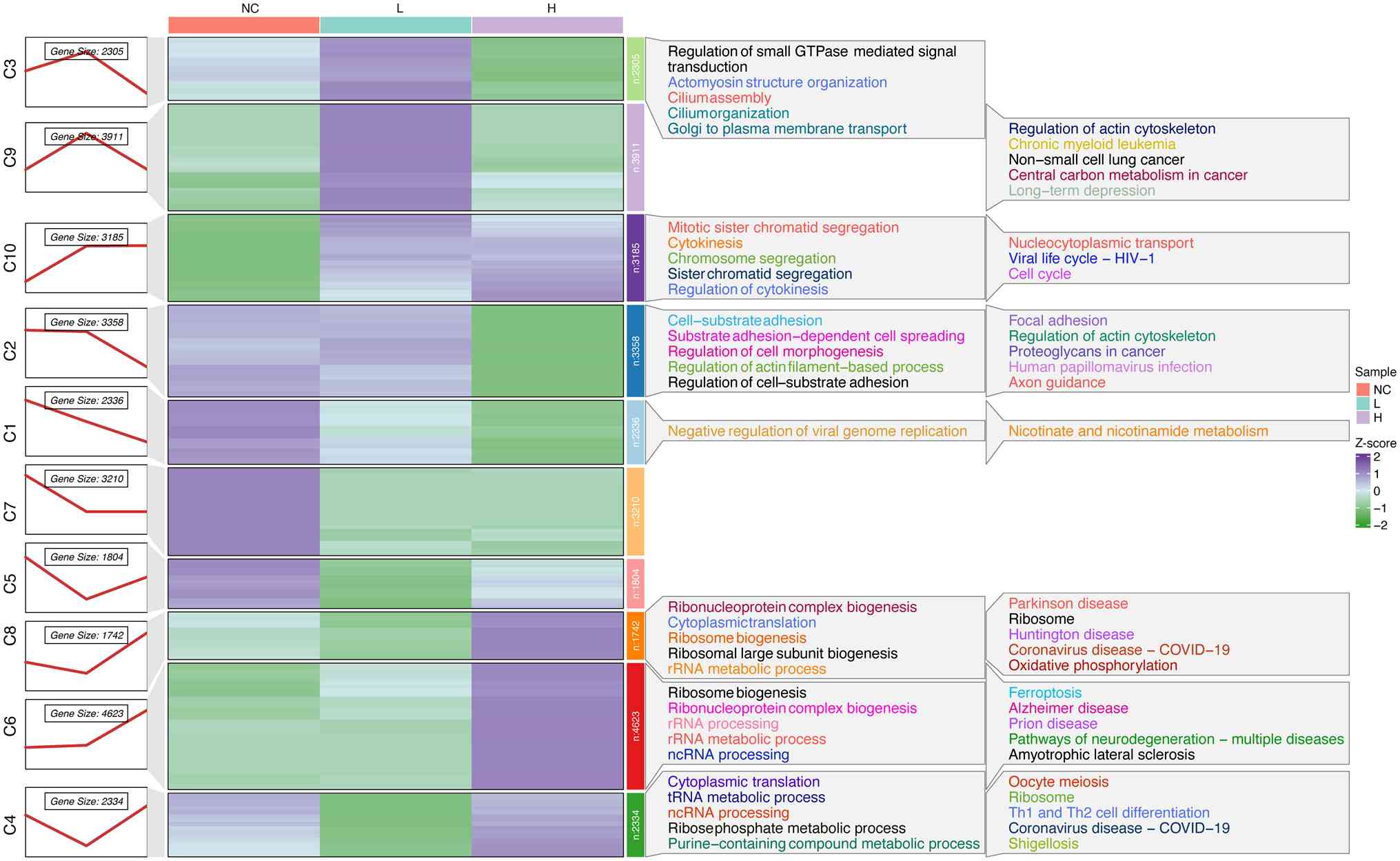
Identification of dynamic expression
patterns
To understand the dynamic alteration expression
profiles in T24 cells during the progression of PPII treatment, 834
DEGs were subjected to soft clustering to determine their
expression trends in the three series groups. The DEGs were divided
into 10 clusters according to their trend similarity over time
using a soft Mfuzz clustering algorithm and not all the clusters
exhibited consistent expression trends with a distinct peak. A
heatmap revealed that both Cluster 6 and Cluster 8 showed
transcriptional upregulation response to PPII treatment, Cluster 6
demonstrated a more consistent concentration-dependent pattern,
showing gradual upregulation with increasing PPII concentrations.
By contrast, Cluster 8 exhibited a biphasic response with initial
downregulation at lower concentrations followed by upregulation at
higher doses (Fig. 3). On the
basis of this clear concentration-dependent response, Cluster 6 was
selected for further investigation.
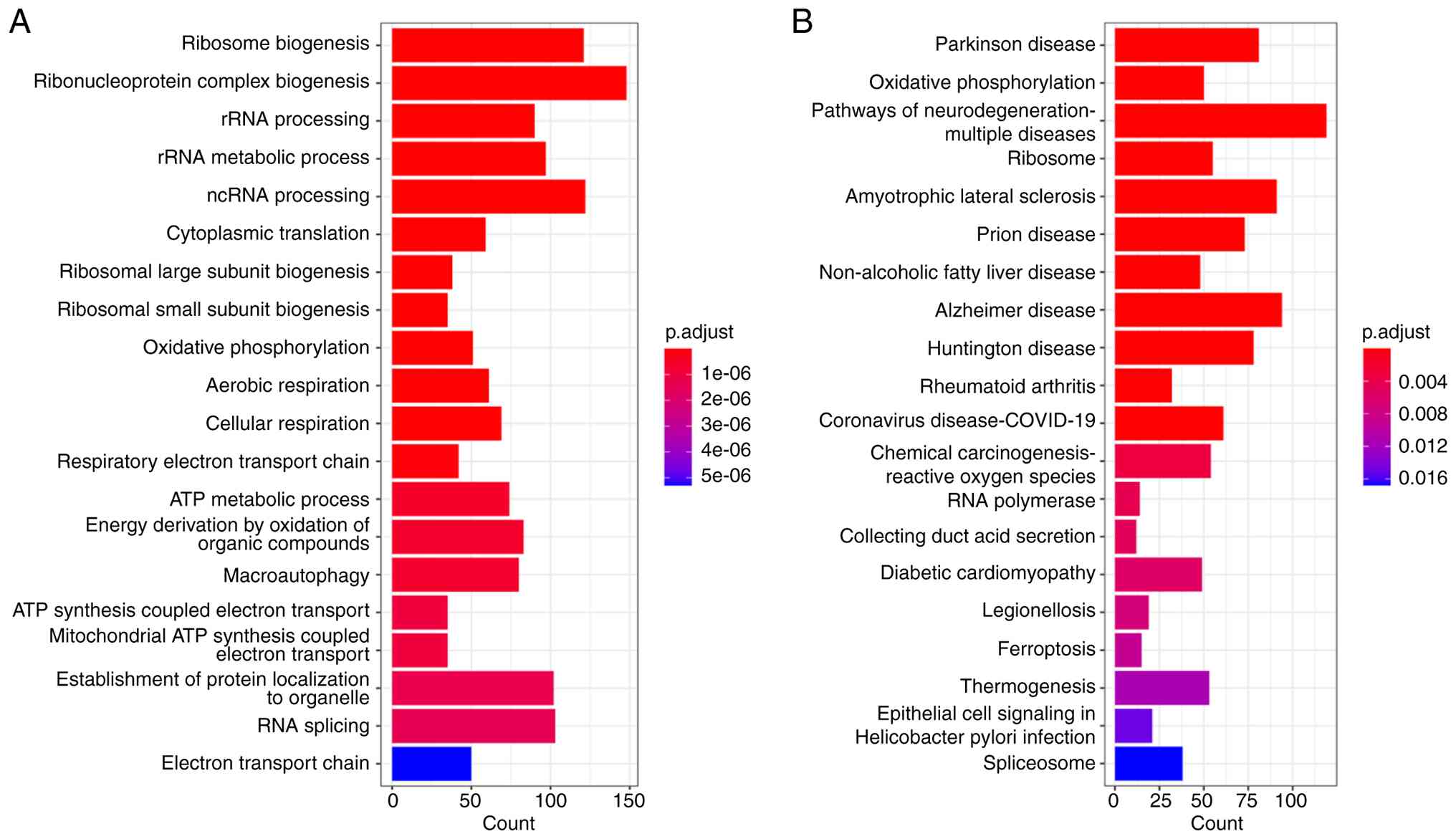
GO and KEGG analyses of Cluster 6
GO and KEGG pathway analyses of the DEGs in Cluster
6 revealed enrichment of ribosome-related and RNA
processing-related categories in the GO analysis (Fig. 4A). KEGG analysis revealed pathways
related to ferroptosis and ROS (Fig.
4B), and ferroptosis-related factors, including PTGS2 and
ACSL4, showed varying degrees of changes with dose-dependent manner
in volcano maps (Fig. 2C and D).
Given that ferroptosis is closely associated with the regulation of
cell proliferation and that ROS play a crucial role in the
ferroptosis process, subsequent experiments focused on validating
the regulatory effects of PPII on ROS and ferroptosis.
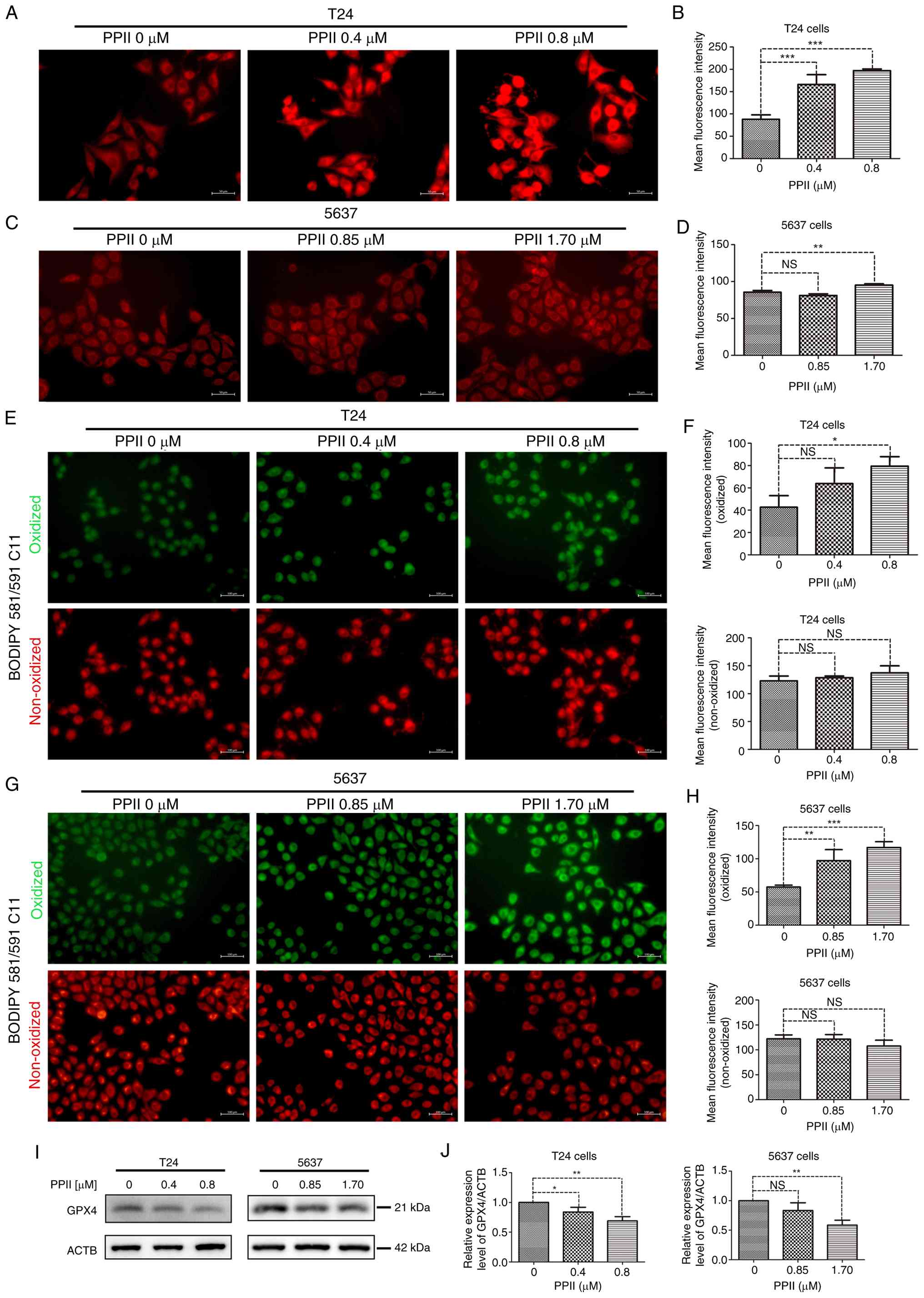
PPII promotes ROS in BC cells
The effect of PPII on ROS levels in bladder cancer
cells was assessed using a ROS detection kit, as detailed in the
methods section. The fluorescence intensity was used to quantify
the ROS levels, revealing that T24 cells exhibited a significant
increase in fluorescence intensity following PPII treatment.
Notably, the fluorescence intensity in the 0.8 µM PPII treatment
group was comparable to that in the positive control group (CRTL +
Rosup; Fig. 5A). Statistical
analysis demonstrated that as the concentration of PPII increased,
the average fluorescence intensity significantly increased
(Fig. 5C; P<0.001) in a
dose-dependent manner. Similar results were observed in 5637 cells
(Fig. 5B and C). Since MDA and SOD
are critical regulators of ROS levels, their concentrations were
measured. High concentrations of PPII (0.8 µM for T24 cells and
1.70 µM for 5637 cells) significantly increased MDA levels
(Fig. 5D, P<0.05) but inhibited
SOD activity (Fig. 5E; P<0.05).
Furthermore, Pearson analysis revealed a negative correlation
between ROS and SOD, whereas ROS and MDA showed a positive
correlation (Fig. S1B). These
findings suggested that PPII effectively increases ROS levels in
bladder cancer cells.
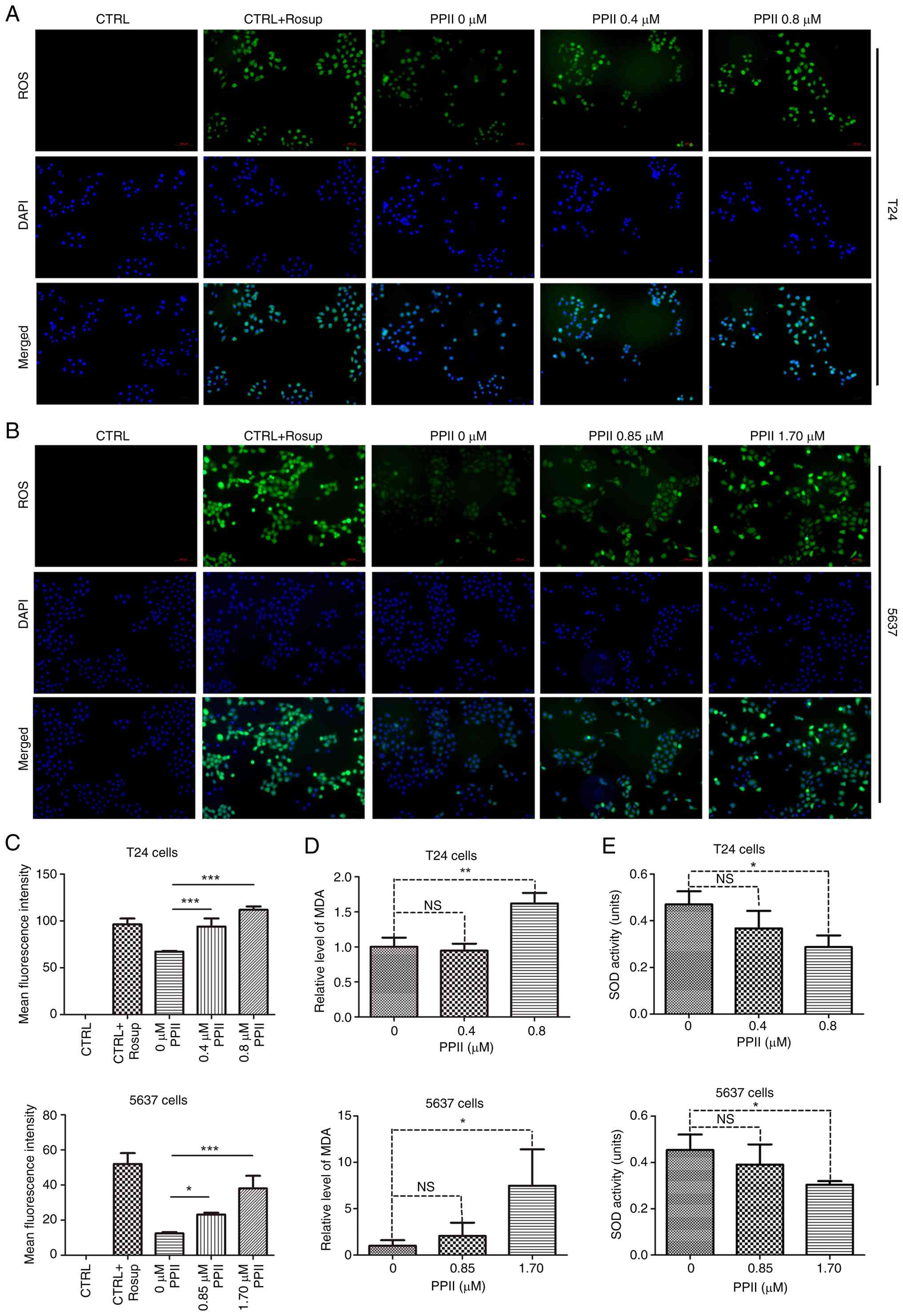 | Figure 5.ROS level detection. (A) T24 and (B)
5637 cells (magnification, ×100) were treated with gradient
concentrations of PPII and (C) their ROS levels were measured using
a reagent kit, with the average fluorescence intensity analyzed
statistically. The levels of (D) MDA and (E) SOD in BC cells were
determined using the corresponding assay kits as described in the
methods section. NS, no significant difference, *P<0.05,
**P<0.01 and ***P<0.001. ROS, reactive oxygen species; PPII,
polyphyllin II; MDA, malondialdehyde; SOD, superoxide dismutase;
BC, bladder cancer. |
PPII promotes ferroptosis in BC
cells
Fe2+ accumulation is a key indicator of
ferroptosis. To assess changes in Fe2+ levels after PPII
treatment in BC cells, the FerroOrange kit was used. The results
revealed that the Fe2+ staining intensity increased with
increasing PPII concentration in T24 cells (Fig. 6A). Statistical analysis revealed
that the average fluorescence intensity significantly increased in
a dose-dependent manner (Fig. 6B;
P<0.001). Consistent findings were also observed in 5637 cells
(Fig. 6C and D).
The accumulation of oxidized lipids serves as a key
marker in the process of ferroptosis. Using specific probes, the
levels of both oxidized and non-oxidized lipids was assessed.
Treatment with 0.8 µM PPII significantly elevated the levels of
oxidized lipids in T24 cells (P<0.05), but did not markedly
affect the levels of non-oxidized lipids (Fig. 6E and F). Similarly, in 5637 cells,
treatment with 0.85 µM and 1.70 µM PPII markedly increased oxidized
lipid levels (Fig. 6G and H).
Molecular docking suggests that PPII and GPX4, a key
regulatory factor of Ferroptosis, have potential good binding
ability (Fig. S1C and D; binding
energy, −8.7 kcal/mol). Further analysis of the expression of GPX4
was conducted using western blotting. The results indicated that
treatment with high concentrations of PPII markedly decreased GPX4
levels in both T24 and 5637 cells (Fig. 6I and J). Furthermore, PPII reduces
the increase in GPX4 levels caused by the ferroptosis inhibitor
Fer-1 (Fig. S1E). These findings
suggested that the regulatory effect of PPII on ferroptosis is
mediated by GPX4.
Discussion
ROS play pivotal roles in ferroptosis, a form of
programmed cell death characterized by iron-dependent lipid
peroxidation (22).
Fe2+ is crucial in this process, serving as a catalyst
in the Fenton reaction, which converts hydrogen peroxide into
highly active hydroxyl radicals. These radicals exacerbate
oxidative stress by promoting the peroxidation of polyunsaturated
fatty acids within cellular membranes (23). The resulting accumulation of
oxidized lipids leads to cellular damage and ultimately triggers
ferroptosis (24). This
relationship between Fe2+ and ROS underscores the
importance of iron metabolism in the regulation of ferroptosis,
particularly in cancer cells in which these pathways are often
dysregulated (10,25).
The antioxidant enzyme GPX4 plays a critical role in
counteracting oxidative stress. By reducing lipid hydroperoxides to
their corresponding alcohols, GPX4 prevents the propagation of
lipid peroxidation and protects cells from ferroptosis (26). However, when GPX4 activity is
compromised, either through genetic downregulation or
pharmacological inhibition, cells become more susceptible to
ferroptosis because of the unchecked accumulation of ROS and
oxidized lipids (27).
According to the Chinese Pharmacopoeia, the
Polyphyllins, including PPI, PPII, Polyphyllin VI (PPVI), and
Polyphyllin VII (PPVII), are the major active compounds used for
the authentication of Paris polyphylla. Our previous
experimental data indicated that, among these compounds, compared
with PPI, PPVI, and PPVII, PPII has superior efficacy in inhibiting
the viability of both T24 and 5637 cells (28). In the present study, PPII treatment
markedly downregulated GPX4 expression, leading to elevated ROS
levels and increased lipid peroxidation in bladder cancer cells.
These findings suggest that PPII may promote ferroptosis by
disrupting the delicate balance between ROS production and
antioxidant defense mechanisms mediated by GPX4.
In addition to GPX4, other oxidative stress-related
markers, such as MDA and SOD, play significant roles in the
regulation of ferroptosis. MDA is a byproduct of lipid peroxidation
and serves as a biomarker for oxidative stress (29). The present study revealed that PPII
treatment leads to increased MDA levels in the bladder cancer
cells, indicating increased lipid peroxidation. Conversely, the
activity of SOD, an enzyme responsible for detoxifying superoxide
radicals (30), was inhibited by
PPII, contributing to the accumulation of ROS. These findings
suggested that PPII modulates multiple components of the oxidative
stress pathway, promoting ferroptosis through both the upregulation
of lipid peroxidation and the inhibition of antioxidant
defenses.
The interplay between Fe2+, ROS and lipid
peroxidation is further complicated by the involvement of various
signaling pathways. For instance, ROS can activate signaling
cascades that promote the expression of pro-ferroptotic genes,
whereas Fe2+-mediated lipid peroxidation can amplify
these signals, creating a feed-forward loop that drives ferroptosis
(31,32). Additionally, the downregulation of
GPX4 disrupts redox homeostasis, tipping the balance in favor of
oxidative stress and ferroptosis (33). The present study highlighted the
potential of targeting this axis in bladder cancer therapy. While
the initial transcriptomic profiling was conducted in T24 cells as
a resource-efficient hypothesis-generating step, all key
mechanistic findings, particularly regarding ROS accumulation and
ferroptosis activation, were rigorously validated in both T24 and
5637 cell lines. This dual-cell-line validation approach ensures
that the conclusions of the present study about PPII's
pro-ferroptotic effects are generalizable across different MIBC
models, as PPII-induced GPX4 downregulation, combined with
increased Fe2+ accumulation and ROS production, leads to
a potent ferroptotic response in bladder cancer cells. Although
PPII affects the ferroptosis process of bladder cancer cells
through GPX4, GPX4 may not be the direct target of PPII. Fe ions
not only participate in the induction of ferroptosis but also cause
pyroptosis (34). Does the PPII
regulate the accumulation of Fe ions to induce other forms of cell
death? It is hoped to clarify the upstream regulatory factors
through which PPII affects ferroptosis through GPX4, such as
Nrf2/HO-1 pathway, in future studies and explore other forms of
cell death induced by PPII.
In conclusion, the promotion of ferroptosis through
the modulation of Fe2+ levels, ROS production, and lipid
peroxidation represents a promising therapeutic strategy in bladder
cancer. The downregulation of GPX4 by PPII and the associated
increase in oxidative stress suggest that targeting the above
processes could be an effective approach for increasing
ferroptosis. PPII is a potential small molecule drug for the
treatment of bladder cancer and future studies should further
investigate the molecular mechanisms underlying the effects of PPII
and explore its potential in combination with other
ferroptosis-inducing agents. Furthermore, in vivo validation
and pharmacokinetic analysis should also be conducted.
Supplementary Material
Supporting Data
Acknowledgements
The authors would like to thank Professor Shengtian
Zhao (First Clinical Medical College, Qilu Hospital of Shandong
University, Jinan, Shandong) for providing the T24 and 5637 cell
lines.
Funding
The present study was funded by the Natural Science Foundation
of Shandong Province, China (grant no. ZR2020QH337).
Availability of data and materials
The data generated in the present study are included
in the figures of this article. The data generated in the present
study may be found in the Genome Sequence Archive (Genomics,
Proteomics & Bioinformatics, 2021) at the National Genomics
Data Center (Nucleic Acids Research, 2022), China National Center
for Bioinformation/Beijing Institute of Genomics, Chinese Academy
of Sciences (GSA-Human: HRA005259; http://ngdc.cncb.ac.cn/gsa-human). The original images
and analytical data may be found in the figshare repository and can
be accessed via https://figshare.com/articles/dataset/Polyphyllin_II_regulates_ROS_levels_and_promotes_ferroptosis_in_bladder_cancer_cells/30448508.
Authors' contributions
ZL and YX conceived and designed the study. QQ and
YS conducted the experiments. Data analysis and display (writing
the R language code for the bioinformatics analysis and for
generating the figures in the article) were performed by ZS and RG.
RG and ZS confirm the authenticity of all the raw data. All authors
participated in the manuscript writing which was ultimately edited
by YX and ZL. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The T24 and 5637 cell lines used in this experiment
were approved by the Ethics Committee of the Affiliated Hospital of
Shandong University of Traditional Chinese Medicine.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI
|
|
2
|
Li F, Zhang H, Huang Y, Li D, Zheng Z, Xie
K, Cao C, Wang Q, Zhao X, Huang Z, et al: Single-cell transcriptome
analysis reveals the association between histone lactylation and
cisplatin resistance in bladder cancer. Drug Resist Updat.
73:1010592024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Li F, Zheng Z, Chen W, Li D, Zhang H, Zhu
Y, Mo Q, Zhao X, Fan Q, Deng F, et al: Regulation of cisplatin
resistance in bladder cancer by epigenetic mechanisms. Drug Resist
Updat. 68:1009382023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Liu T, Xu X, Li J, Bai M, Zhu W, Liu Y,
Liu S, Zhao Z, Li T, Jiang N, et al: ALOX5 deficiency contributes
to bladder cancer progression by mediating ferroptosis escape. Cell
Death Dis. 14:8002023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Li W, Ou Y, Ye F, Cheng Z, Chen Z, Zhou Q,
Yan X and Jiang H: cirSIRT5 induces ferroptosis in bladder cancer
by forming a ternary complex with SYVN1/PHGDH. Cell Death Discov.
10:3912024. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Xu Y, Tong Y, Lei Z, Zhu J and Wan L:
Abietic acid induces ferroptosis via the activation of the HO-1
pathway in bladder cancer cells. Biomed Pharmacother.
158:1141542023. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wang B, Wang Y, Zhang J, Hu C, Jiang J, Li
Y and Peng Z: ROS-induced lipid peroxidation modulates cell death
outcome: Mechanisms behind apoptosis, autophagy, and ferroptosis.
Arch Toxicol. 97:1439–1451. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wu S, Guo N, Xu H, Li Y, Sun T, Jiang X,
Fu D, You T, Diao S, Huang Y and Hu C: Caveolin-1 ameliorates
hepatic injury in non-alcoholic fatty liver disease by inhibiting
ferroptosis via the NOX4/ROS/GPX4 pathway. Biochem Pharmacol.
230:1165942024. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Liu H, Schreiber SL and Stockwell BR:
Targeting dependency on the GPX4 lipid peroxide repair pathway for
cancer therapy. Biochemistry. 57:2059–2060. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhang C, Liu X, Jin S, Chen Y and Guo R:
Ferroptosis in cancer therapy: A novel approach to reversing drug
resistance. Mol Cancer. 21:472022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Qin L, Zhong Y, Li Y and Yang Y: TCM
targets ferroptosis: Potential treatments for cancer. Front
Pharmacol. 15:13600302024. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yang R, Gao W, Wang Z, Jian H, Peng L, Yu
X, Xue R, Peng W, Li K and Zeng P: Polyphyllin I induced
ferroptosis to suppress the progression of hepatocellular carcinoma
through activation of the mitochondrial dysfunction via
Nrf2/HO-1/GPX4 axis. Phytomedicine. 122:1551352024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhou Y, Yang J, Chen C, Li Z, Chen Y,
Zhang X, Wang L and Zhou J: Polyphyllin III–Induced Ferroptosis in
MDA-MB-231 Triple-negative breast cancer cells can be protected
against by KLF4-Mediated upregulation of xCT. Front Pharmacol.
12:6702242021. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kim D, Paggi JM, Park C, Bennett C and
Salzberg SL: Graph-based genome alignment and genotyping with
HISAT2 and HISAT-genotype. Nat Biotechnol. 37:907–915. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Pertea M, Pertea GM, Antonescu CM, Chang
TC, Mendell JT and Salzberg SL: StringTie enables improved
reconstruction of a transcriptome from RNA-seq reads. Nat
Biotechnol. 33:290–295. 2015. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Love MI, Huber W and Anders S: Moderated
estimation of fold change and dispersion for RNA-seq data with
DESeq2. Genome Biol. 15:5502014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kumar L and Futschik M: Mfuzz: A software
package for soft clustering of microarray data. Bioinformation.
2:5–7. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z,
Feng T, Zhou L, Tang W, Zhan L, et al: clusterProfiler 4.0: A
universal enrichment tool for interpreting omics data. Innovation
(Camb. 2:1001412021.PubMed/NCBI
|
|
19
|
Shi YX, Xu L, Wang X, Zhang KK, Zhang CY,
Liu HY, Ding P, Shi W and Liu Z: Paris polyphylla ethanol extract
and polyphyllin I ameliorate adenomyosis by inhibiting
epithelial-mesenchymal transition. Phytomedicine. 127:1554612024.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zeng Y, Wu R, Wang F, Li S, Li L, Li Y,
Qin P, Wei M, Yang J, Wu J, et al: Liberation of daidzein by gut
microbial β-galactosidase suppresses acetaminophen-induced
hepatotoxicity in mice. Cell Host Microbe. 31:766–780.e7. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bayona C, Alza L, Ranđelović T, Sallán MC,
Visa A, Cantí C, Ochoa I, Oliván S and Herreros J: Tetralol
derivative NNC-55-0396 targets hypoxic cells in the glioblastoma
microenvironment: An organ-on-chip approach. Cell Death Dis.
15:1272024. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zi Y, Wang X, Zi Y, Yu H, Lan Y, Fan Y,
Ren C, Liao K and Chen H: Cigarette smoke induces the ROS
accumulation and iNOS activation through deactivation of
Nrf-2/SIRT3 axis to mediate the human bronchial epithelium
ferroptosis. Free Radic Biol Med. 200:73–86. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Henning Y, Blind US, Larafa S, Matschke J
and Fandrey J: Hypoxia aggravates ferroptosis in RPE cells by
promoting the Fenton reaction. Cell Death Dis. 13:6622022.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Pope LE and Dixon SJ: Regulation of
ferroptosis by lipid metabolism. Trends Cell Biol. 33:1077–1087.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhao L, Zhou X, Xie F and Zhang L, Yan H,
Huang J, Zhang C, Zhou F, Chen J and Zhang L: Ferroptosis in cancer
and cancer immunotherapy. Cancer Commun (Lond). 42:88–116. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Xie Y, Kang R, Klionsky DJ and Tang D:
GPX4 in cell death, autophagy, and disease. Autophagy.
19:2621–2638. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wang H, Cheng Y, Mao C, Liu S, Xiao D,
Huang J and Tao Y: Emerging mechanisms and targeted therapy of
ferroptosis in cancer. Mol Ther. 29:2185–2208. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Niu W, Xu L, Li J, Zhai Y, Sun Z, Shi W,
Jiang Y, Ma C, Lin H, Guo Y and Liu Z: Polyphyllin II inhibits
human bladder cancer migration and invasion by regulating
EMT-associated factors and MMPs. Oncol Lett. 20:2928–2936. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ye T, Yang W, Gao T, Yu X, Chen T, Yang Y,
Guo J, Li Q, Li H and Yang L: Trastuzumab-induced cardiomyopathy
via ferroptosis-mediated mitochondrial dysfunction. Free Radic Biol
Med. 206:143–161. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ji HS, Bang SG, Ahn MA, Kim G, Kim E, Eom
SH and Hyun TK: Molecular cloning and functional characterization
of heat Stress-responsive superoxide dismutases in garlic (Allium
sativum L.). Antioxidants (Basel). 10:8152021. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Liu J, Kang R and Tang D: Signaling
pathways and defense mechanisms of ferroptosis. FEBS J.
289:7038–7050. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wu ZF, Liu XY, Deng NH, Ren Z and Jiang
ZS: Outlook of Ferroptosis-targeted lipid peroxidation in
cardiovascular disease. Curr Med Chem. 30:3550–3561. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang W, Liu Y, Liao Y, Zhu C and Zou Z:
GPX4, ferroptosis, and diseases. Biomed Pharmacother.
174:1165122024. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhou B, Zhang JY, Liu XS, Chen HZ, Ai YL,
Cheng K, Sun RY, Zhou D, Han J and Wu Q: Tom20 senses
iron-activated ROS signaling to promote melanoma cell pyroptosis.
Cell Res. 28:1171–1185. 2018. View Article : Google Scholar : PubMed/NCBI
|