Introduction
Periodontitis is one of the most common chronic
inflammatory diseases worldwide and a leading cause of tooth loss
in adults (1). Its prevalence is
steadily increasing among aging populations, with significant
effects not only on oral health but also on overall health and
quality of life. Periodontitis is also associated with systemic
chronic inflammatory diseases, including inflammatory bowel disease
(IBD), cardiovascular disease, autoimmune conditions, and
Alzheimer's disease (2).
Furthermore, the increasing incidence of periodontitis has led to
an increase in healthcare costs (3). Periodontitis is caused by oral
bacteria, whereby complex interactions between the host immune
response and bacterial toxins lead to chronic inflammation
(4). This results in alveolar bone
destruction and the loss of tooth-supporting tissues, ultimately
leading to tooth loss (5).
Therefore, early periodontitis prevention and delaying progression
are crucial for maintaining oral health and improving patient
quality of life in later life.
The periodontal ligament (PDL) plays a key role in
periodontitis pathophysiology. The PDL is a connective tissue
attaching teeth to the alveolar bone. In addition to its roles in
alleviating mechanical stress and transmitting sensations, the PDL
also helps regulate immune responses and inflammation. In
particular, in early-stage periodontitis, bacterial stimulation
activates PDL cells to secrete various inflammatory cytokines
[interleukin (IL)-1β, IL-6, tumor necrosis factor (TNF)-α, etc.]
and matrix-degrading enzymes (MMPs) to induce an inflammatory
response and promote alveolar bone destruction (6,7).
Therefore, inflammation-inducing models using PDL cells are
important experimental tools in periodontitis research. Among these
models, lipopolysaccharide (LPS) is a toxin derived from the cell
wall of gram-negative bacteria, such as Porphyromonas
gingivalis, and is a major causative agent of periodontitis. It
induces a strong inflammatory response by activating inflammatory
signaling pathways [mitogen-activated protein kinase (MAPK) and
nuclear factor-κB (NF-κB), etc.] through the innate immune receptor
Toll-like receptor 4 (TLR4) (8).
Therefore, LPS is the most widely used stimulant in in vitro
inflammatory models mimicking periodontitis, producing an
inflammatory response similar to that of periodontitis (9).
Recent studies have increasingly focused on the
regulatory effects of extracellular vehicles (EVs) on inflammatory
diseases. EVs, including small vesicles often referred to as
exosomes (30–150 nm), are secreted by various cell types and carry
bioactive molecules such as mRNA, miRNA, and proteins, thereby
mediating intercellular communication and regulating target cell
function (10). Stem cell-derived
EVs have been shown to exert anti-inflammatory and tissue
regeneration-promoting effects in several disease models (11). Among these, EVs derived from
tonsil-derived stem cells (T-MSC-EVs) modulate immune cell
function, suppress pro-inflammatory cytokine production, and
promote tissue repair (12,13).
Given that periodontitis is characterized by both
excessive inflammation and impaired tissue regeneration,
therapeutic strategies that simultaneously suppress inflammation
and support osteogenic repair are of particular interest.
Therefore, this study investigated the dual anti-inflammatory and
osteogenic effects of T-MSC-EVs in an LPS-stimulated human
periodontal ligament fibroblast (hPDLF) model of periodontitis.
Materials and methods
Cell culture
The hPDLFs were commercially obtained as primary
cells from company (hPDLF, #2630, ScienCell Research Laboratories).
This cell type was classified as a primary human cell tissue and
was not immortalized cell lines. Detailed product information is
available at the supplier's website: https://sciencellonline.com/en/human-periodontal-ligament-fibroblasts/.
Although the cells were commercially sourced, the study was
conducted in accordance with the Declaration of Helsinki and the
research design was officially approved by the Institutional Review
Board of Kyung Hee University (IRB No. KHSIRB-25-417). The hPDLF
cells were cultured in Dulbecco's Modified Eagle's Medium (DMEM)
supplemented with 10% fetal bovine serum (FBS), 100 U/ml
penicillin, and 100 µg/ml streptomycin in a humidified atmosphere
of 5% CO2 at 37°C. Osteogenic medium (OM) was prepared
by supplementing culture media with 50 µg/ml L-ascorbic acid
(Sigma-Aldrich; Merck KGaA) and 10 mM β-glycerophosphate
(Sigma-Aldrich; Merck KGaA) (14).
The hPDLF cells were sub-cultured at 80–90% confluence, and
experiments were conducted using passages 4–7.
EV preparation from T-MSCs and
characterization by nanoparticle tracking analysis (NTA) and
transmission electron microscopy (TEM)
T-MSCs were obtained from human tonsil tissues,
following approval by the Institutional Review Board (Department of
Otorhinolaryngology, Yonsei University Wonju College of Medicine,
IRB-CR320104), with written informed consent obtained from all
donors. This cell type was classified as a primary human cell
tissue and was not immortalized cell lines. T-MSCs were cultured in
medium supplemented with EV-depleted FBS prepared by
ultracentrifugation, as previously described. At 70–80% confluency,
the cells were washed, switched to fresh EV-depleted medium, and
conditioned for 24–48 h. Conditioned medium (CM) was collected and
cleared of cells, apoptotic bodies, and debris by sequential low-
to medium-speed centrifugation and 0.22 µm filtration. Small EVs
were then isolated from the clarified CM by high-speed
ultracentrifugation, washed by a second ultracentrifugation step,
and resuspended in Dulbecco's PBS (DPBS). The resulting EVs were
then aliquoted and stored at −80°C while minimizing freeze-thaw
cycles. The T-MSC-EVs used in the experiments in the present study
were manufactured and purified by Hyundai Meditech Co., Ltd. using
internal standard operating procedures (SOPs). EV identity,
particle size distribution, and concentration were confirmed by
nanoparticle tracking analysis (NTA). Transmission electron
microscopy (TEM) was performed using a Talos L120C microscope (FEI,
USA) operated at 120 kV with a LaB6 electron gun. A
carbon-film-coated copper grid (400 mesh, Ted Pella) was rendered
hydrophilic using a PELCO easiGlow™ glow discharge
system (Ted Pella) under negative polarity for 20 sec before
mounting the samples. For negative staining, a 2% uranyl acetate
(UA) solution was freshly prepared. One drop of EV suspension was
placed on a glow-discharged grid and incubated for approximately 1
min. Excess liquid was gently removed using filter paper. The grid
was then floated on a 20 µl droplet of 2% UA solution for 20 sec
for staining, and excess stain was blotted off. Finally, the grid
was air-dried for 10 min at room temperature before imaging.
EV dosing
T-MSC-EVs were quantified by NTA and administered to
hPDLFs at final concentrations of 1×108 or 5×108 particles/ml,
depending on the experimental condition. Particle number-based
dosing was used because the total EV protein content was
insufficient for reliable quantification and did not necessarily
reflect the number of vesicles or functional EV cargo. The
selection of these specific doses (1×108 or 5×108 particles/ml) was
based on previous studies demonstrating that MSC-derived EVs exert
significant immunomodulatory effects within this concentration
range in in-vitro human cell models (15).
RT-qPCR
Total RNA was isolated from cells using TRIzol
reagent (Invitrogen, Carlsbad, CA, USA) according to the
manufacturer's instructions. Reverse transcription was performed
using AccuPower RT PreMix (Bioneer). qPCR was performed on the cDNA
samples using AxenTM qPCR Master Mix (Macrogen) on a
7500 Real-Time PCR System (Thermo Fisher Scientific, Inc.). The
relative mRNA levels of target genes were normalized to β-actin
mRNA levels and analyzed using the comparative Ct method (ΔΔCt)
(16). The primer sequences used
in this study are listed in Table
I.
 | Table I.PCR primers. |
Table I.
PCR primers.
| Gene | Sequences
(5′-3′) | Ta, °C | Size, bp |
|---|
| GAPDH | Forward:
CTCTTCACCACCATGGAGAAG | 60 | 201 |
|
| Reverse:
GTTGTCATGGATGACCTTGGC |
|
|
| IFN-γ | Forward:
TGTCGCCAGCAGCTAAAACA | 60 | 91 |
|
| Reverse:
TGCAGGCAGGACAACCATTA |
|
|
| IL-6 | Forward:
CACCGGGAACGAAAGAGAAGC | 61 | 75 |
|
| Reverse:
CGAAGGCGCTTGTGGAGA |
|
|
| IL-1β | Forward:
AGCTACGAATCTCCGACCAC | 59 | 186 |
|
| Reverse:
CGTTATCCCATGTGTCGAAGAA |
|
|
| ALP | Forward:
CTATCCTGGCTCCGTGCTCC | 62 | 133 |
|
| Reverse:
AGAGATGCAATCGACGTGGG |
|
|
| BSP | Forward:
GACACCACAGAGACCGGAAG | 60 | 232 |
|
| Reverse:
AATTGTCCCCACGAGGTTCC |
|
|
| OCN | Forward:
ATGAGAGCCCTCACACTCCT | 59 | 117 |
|
| Reverse:
CTTGGACACAAAGGCTGCAC |
|
|
| OPN | Forward:
ACAAATACCCAGATGCTGTGGC | 61 | 86 |
|
| Reverse:
ACTTGGAAGGGTCTGTGGGG |
|
|
| SOST | Forward:
ACACAGCCTTCCCGTGTAGTG | 62 | 187 |
|
| Reverse:
CGGACACGTCTTTGGTCTCA |
|
|
Western blotting
Total cells were lysed in radioimmunoprecipitation
assay (RIPA) buffer supplemented with protease and phosphatase
inhibitor cocktails. Whole-cell lysates and nuclear extracts were
prepared according to standard protocols. Protein concentrations
were determined using a protein assay. Equal amounts of protein (25
µg per lane) were separated by sodium dodecyl
sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and
transferred onto polyvinylidene fluoride (PVDF) membranes. After
transfer, the membranes were rinsed three times with Tris-buffered
saline containing 0.1% Tween-20 (TBST) and blocked with a blocking
solution for 1 h at room temperature. The membranes were then
incubated overnight at 4°C with primary antibodies, followed by
incubation with horseradish peroxidase-conjugated secondary
antibodies for 1 h at 37°C. Primary antibodies against β-actin,
p65, c-Jun, and c-Fos (Santa Cruz Biotechnology, Inc.), as well as
phosphorylated or total extracellular signal-regulated kinase (ERK)
and c-Jun N-terminal kinase (JNK) (Cell Signaling Technology) were
used at a dilution of 1:1,000.
For characterization, EVs were lysed in RIPA buffer
containing protease inhibitors, and protein concentrations were
determined. Equal amounts of vesicle proteins were subjected to
SDS-PAGE and transferred onto PVDF membranes, as described above.
The membranes were probed with primary antibodies against the
exosome markers cluster of differentiation (CD)63 (rabbit
monoclonal, Cell Signaling Technology, #52090, 1:1,000) and CD9
(rabbit monoclonal, #13174, 1:1,000). Albumin (rabbit polyclonal,
#4929; Cell Signaling Technology, 4929, 1:1,000) was used as a
negative marker to evaluate contamination by soluble serum
proteins. Protein bands were visualized using an enhanced
chemiluminescence (ECL) detection system (Amersham), and images
were captured using an Amersham™ ImageQuant™
500 imaging system (Cytiva).
Cell proliferation
Cell proliferation was evaluated using EZ-Cytox
(DoGenBio), according to the manufacturer's instructions. Briefly,
cells were seeded in 96-well plates at a density of
5×103 cells/well and cultured under the indicated
conditions. At the designated time points, 100 µl of solution,
diluted 1:10, was added to the culture was added to each well and
incubated for 1 h at 37°C. The absorbance at 450 nm was measured
using a microplate reader (Thermo Fisher Scientific, Inc.). Cell
proliferation was expressed as a percentage relative to the control
group. All experiments were performed in triplicate.
Alkaline phosphatase (ALP) activity
assay and staining
ALP activity was measured using a
sensoLyte® pNpp alkaline phosphatase assay kit
(Anaspec). Cells were cultured in 96-well plates, and osteogenic
differentiation was induced under the specified conditions. After
the designated incubation period, the culture medium was removed,
and the cells were gently washed twice with PBS to eliminate serum
components that could interfere with the assay. Subsequently, 100
µl of pNPP substrate solution was directly added to each well. The
plate was then incubated at room temperature for 1 h, protected
from light, and quantified by measuring absorbance at 405 nm using
a microplate reader. ALP staining was performed using a TRACP &
ALP double staining kit (Takara) according to the manufacturer's
instructions. The cells were fixed in a fixation solution for 20
min, washed twice with PBS, and stained with ALP staining solution.
After incubation in the dark at room temperature for 1 h, the
stained cells were photographed under a light microscope (ECLIPSE
TS100; Nikon).
Alizarin Red staining
Calcium deposition was assessed by Alizarin Red S
staining. Cells were cultured in 24-well plates, and osteogenic
differentiation was induced under the specified conditions. The
cells were then washed twice with PBS and fixed in 70% ethanol for
20 min at room temperature. After fixation, the cells were rinsed
with distilled water and incubated with Alizarin Red S solution
(Samchun Pure Chemical) for 2 h in the dark at room temperature.
The stained cells were washed thoroughly with distilled water to
remove excess dye and then air-dried. The stained calcium deposits
were imaged using an ECLIPSE TS100 system (Nikon).
Statistical analysis
Statistical analyses were performed using GraphPad
Prism, version 8.4.3 (GraphPad Software). For the mRNA expression
of inflammatory markers (PCR data), ordinary one-way analysis of
variance (ANOVA) was performed, followed by Bonferroni's post-hoc
test. For all other experiments, two-way ANOVA with Tukey's
multiple comparison test was used. All values of *P<0.05,
**P<0.01, ***P<0.001, ****P<0.0001 were regarded as
indicative of statistical significance.
Results
NTA characterization of marker
validation and T-MSC-derived EVs
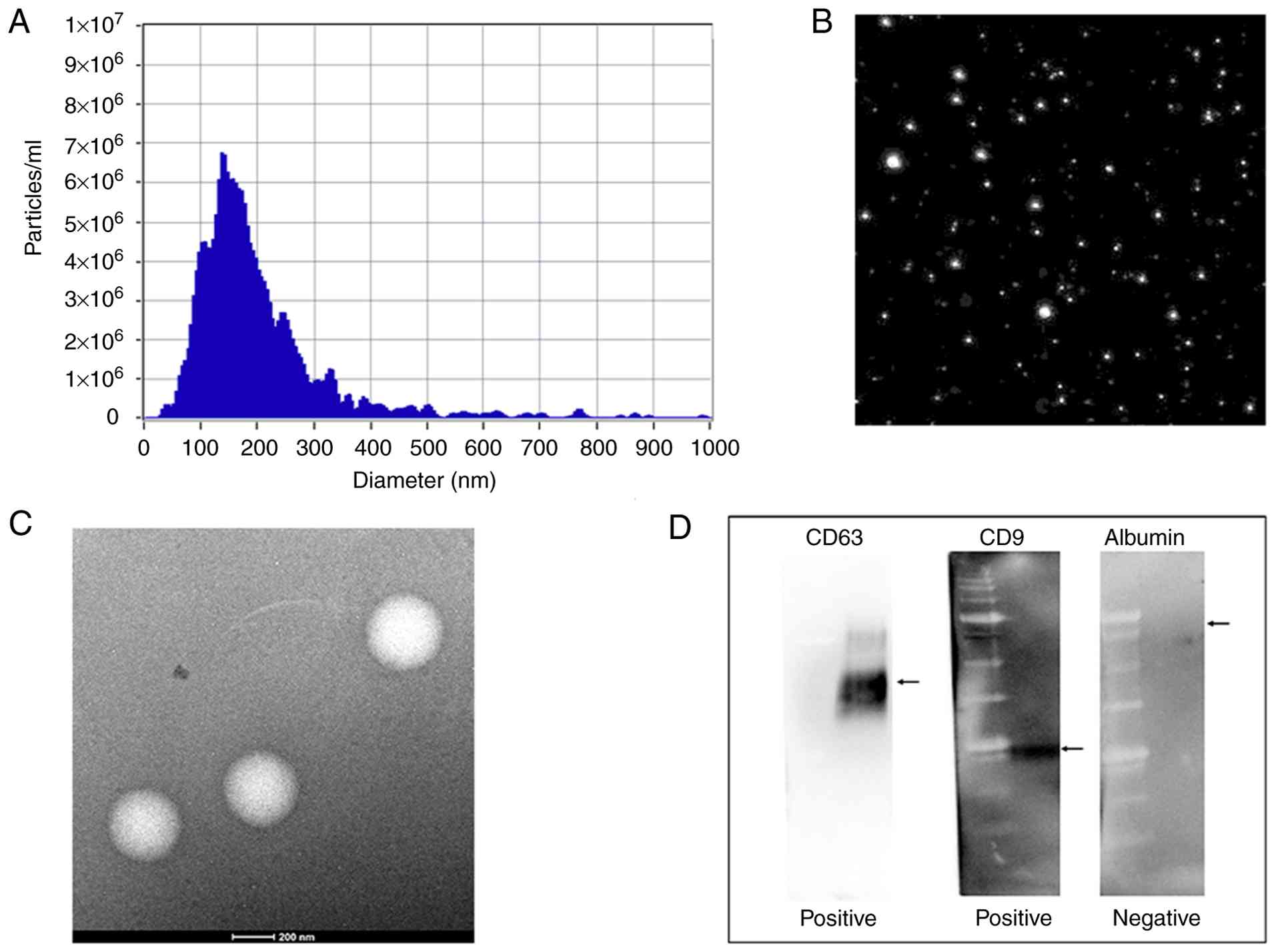
NTA, TEM, and western blotting were performed to
confirm the physical characteristics and molecular identity of
T-MSC-derived EVs (Fig. 1). The
NTA analysis considered only particles ≤200 nm as EV-sized
populations; additionally, minor high-diameter shoulders (>250
nm) were excluded as potential aggregates. The size-concentration
curve showed a unimodal distribution with a peak at approximately
140–150 nm. The particle concentration was approximately 1.5×108
particles/ml. Consistent with these measurements, TEM revealed
round vesicle-like structures within the expected exosomal size
range (50–150 nm). The isolated T-MSC-derived EVs showed a typical
exosomal morphology without apparent aggregation. Western blotting
to assess the expression of EV-associated markers and to further
validate the exosomal nature and purity of the isolated vesicles
revealed tetraspanins CD63 and CD9 in the T-MSC-derived EVs,
confirming the enrichment of exosome-associated proteins. In
contrast, albumin, which was used as a negative marker for soluble
protein contamination, was not detected in the EV preparations.
Collectively, these results demonstrated that the
isolated T-MSC-derived EVs exhibited the size distribution,
morphology, and molecular marker profile characteristic of
exosomes, indicating successful isolation and high purity (Fig. 1).
LPS inhibition of cell proliferation
and the protective effect of EVs
Fig. 2 shows the
effects of LPS and EVs on the PDL. LPS inhibited cell proliferation
in a time- and dose-dependent manner. Although no significant
differences were observed on day 2, marked differences were
observed among the groups on day 3. T-MSC-EVs effectively prevented
the LPS-induced inhibition of cell proliferation at both
concentrations, and higher concentrations promoted cell growth to a
level exceeding that of the control.
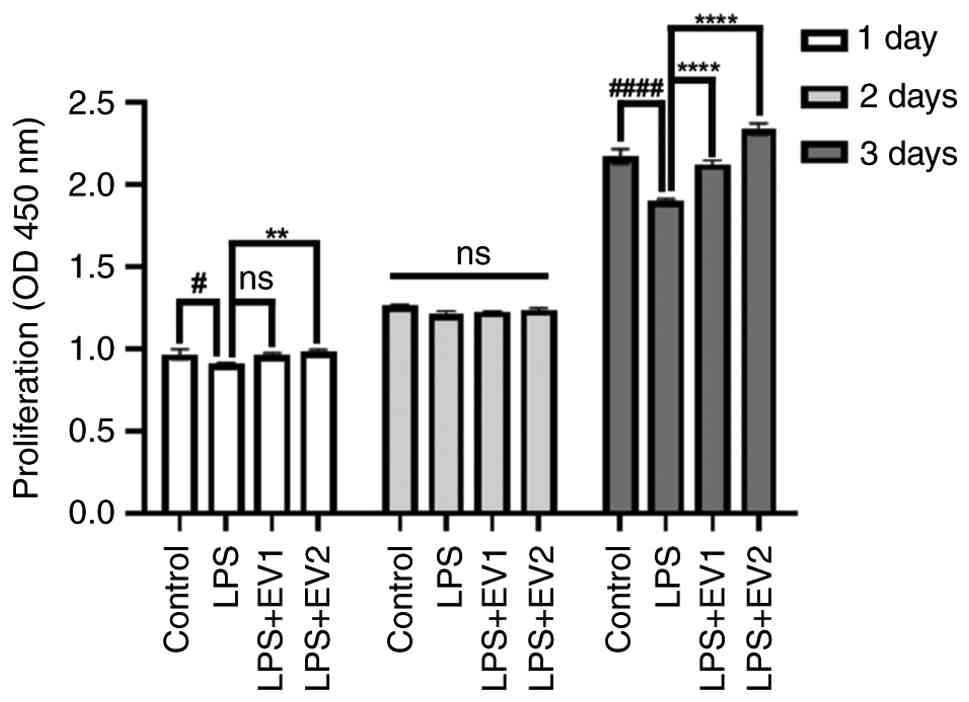 | Figure 2.Cell proliferation was evaluated by
measuring the OD at 450 nm on days 1, 2 and 3 following treatment
with LPS (1 µg/ml) and T-MSC-EVs. EVs were used at concentrations
of 1×108 particles/ml or 5×108 particles/ml.
LPS treatment suppressed proliferation in a time-dependent manner,
with a pronounced reduction observed on day 3 (P<0.0001 vs.
control). By contrast, EV treatment effectively prevented
LPS-induced proliferation inhibition, and notably, treatment with
T-MSC-EVs at 5×108 particles/ml resulted in a
significant increase in cell proliferation beyond the control level
on day 3. No significant differences were observed among the groups
on day 2. All experiments were performed in triplicate.
#P<0.05 and ####P<0.0001, control group
vs. LPS group. **P<0.01 and ****P<0.0001, LPS group vs.
EV-treated groups. EV1, T-MSC-derived EVs (1×108
particles/ml); EV2, T-MSC-derived EVs (5×108
particles/ml); EV, extracellular vesicle; LPS, lipopolysaccharide;
ns, not significant; OD, optical density; T-MSC, tonsil-derived
mesenchymal stem cell. |
Changes in inflammatory cytokine
expression
The expression levels of inflammatory cytokines,
including IL-1β, IL-6, IL-8, and IFN-γ, were markedly
elevated following LPS stimulation in hPDLFs (Fig. 3). In the T-MSC-EVs-treated group,
IL-6 and IL-1β levels were stable or slightly
increased, with no statistically significant change in
IL-1β, while T-MSC-EV treatment significantly reduced
IL-8 and IFN-γ levels. These results demonstrated a
cytokine response pattern characterized by selective reduction of
IL-8 and IFN-γ in the experiment.
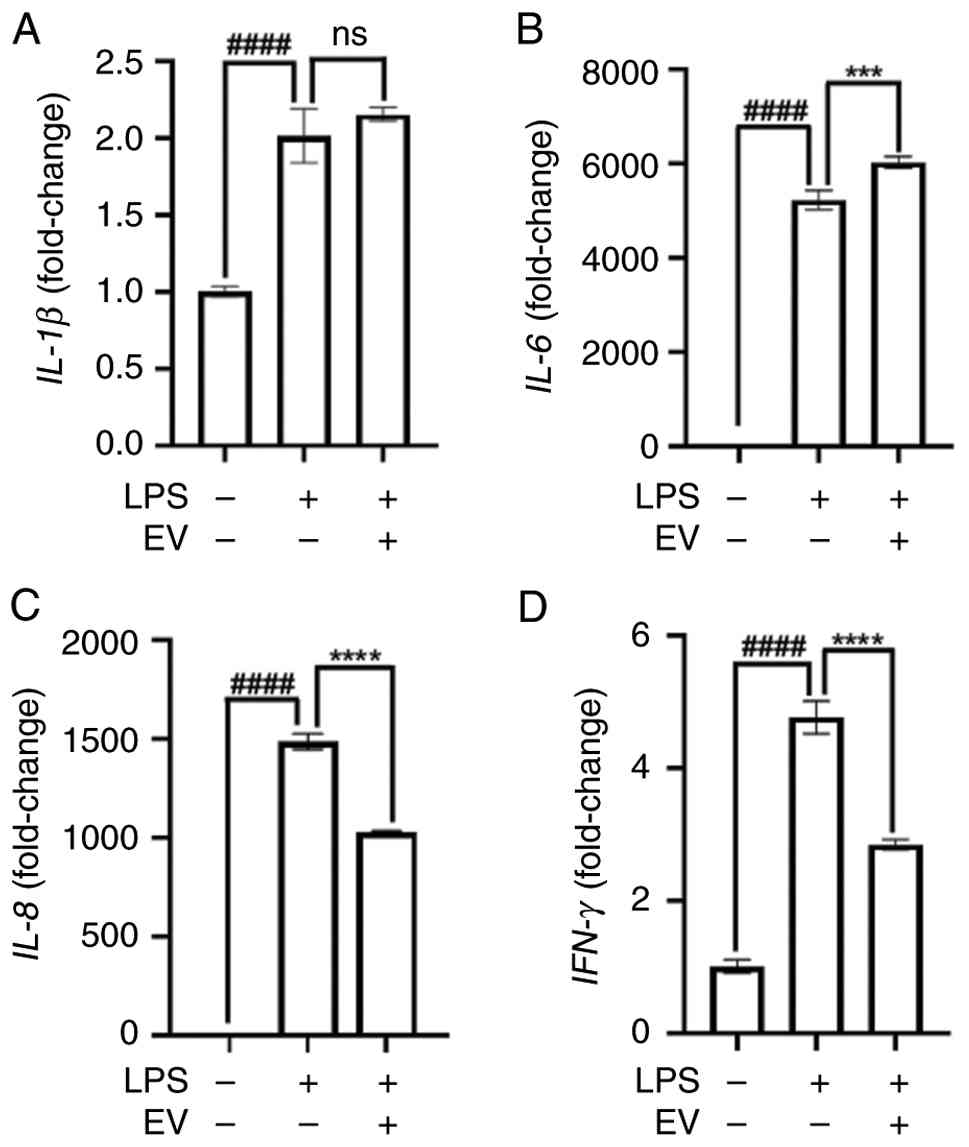 | Figure 3.Quantitative PCR analysis of
proinflammatory cytokines (IL-1β, IL-6, IL-8 and
IFN-γ) in cells stimulated with LPS (1 µg/ml), with or
without tonsil-derived mesenchymal stem cell-derived EVs
(5×108 particles/ml) treatment. (A) IL-1β mRNA
expression. (B) IL-6 mRNA expression. (C) IL-8 mRNA
expression. (D) IFN-γ mRNA expression. LPS significantly
induced the expression of all four cytokines. EV treatment further
increased IL-6 expression but had no significant effect on
IL-1β. By contrast, IL-8 and IFN-γ levels were
significantly decreased following EV treatment, suggesting partial
attenuation of the inflammatory response. All experiments were
performed in triplicate. ####P<0.0001, control group
vs. LPS group. ***P<0.001 and ****P<0.0001, LPS group vs.
EV-treated group. EV, extracellular vesicle; LPS,
lipopolysaccharide; ns, not significant. |
Modulation of inflammatory signaling
pathways
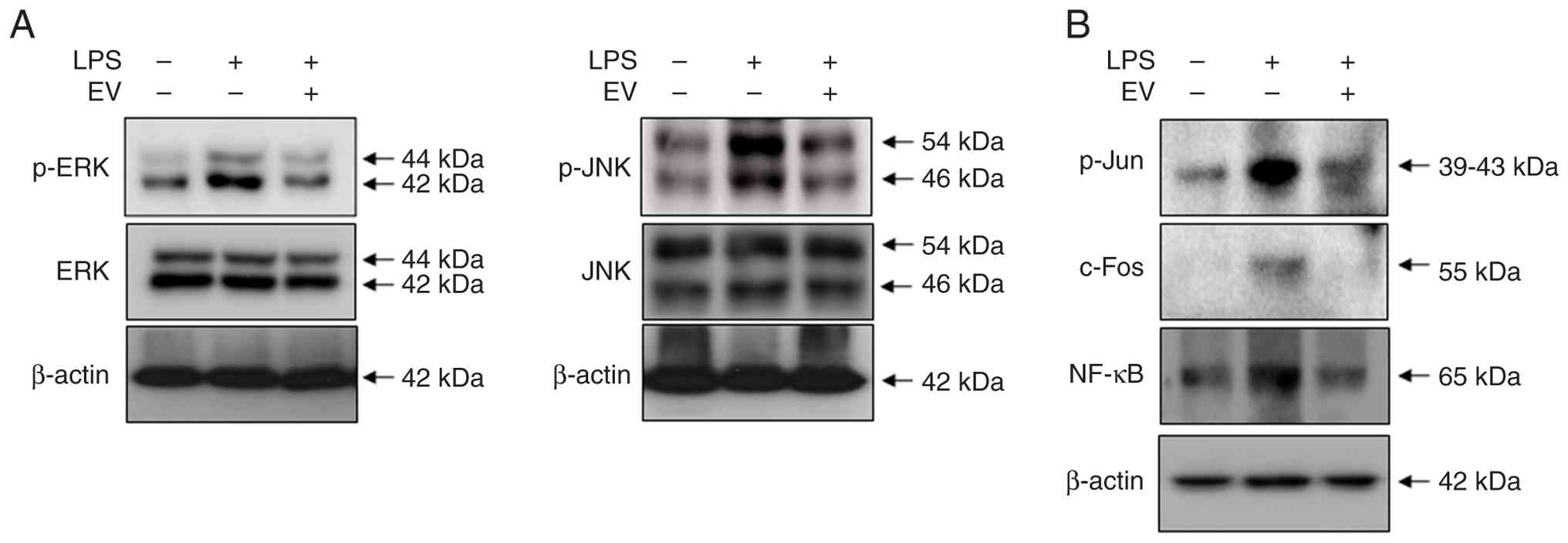
LPS stimulation activated classical MAPK pathways,
including JNK and ERK, which mediate inflammatory responses and
lead to the activation of transcription factors, including AP-1
(c-Jun, c-Fos) and NF-κB (Fig. 4).
The EV-treated group showed markedly decreased phosphorylation of
MAPK components (ERK and JNK), suggesting that T-MSC-EVs suppressed
the upstream inflammatory signaling cascade (Fig. 4A). Furthermore, downstream
transcription factors c-Jun, c-Fos, and NF-κB were also inhibited,
supporting the anti-inflammatory effects of T-MSC-EVs (Fig. 4B).
Restoration of osteogenic activity by
EV treatment under inflammatory conditions
The effects of T-MSC-EVs on bone formation were
evaluated using ALP activity assay, ALP staining, and Alizarin Red
staining over 21 days. LPS treatment markedly reduced ALP activity
and mineralization capacity (Fig.
5). However, T-MSC-EV treatment restored or even enhanced ALP
activity and mineralized nodule formation, especially at later time
points (days 14 and 21), indicating a potential osteogenic effect
of T-MSC-EVs despite inflammatory conditions.
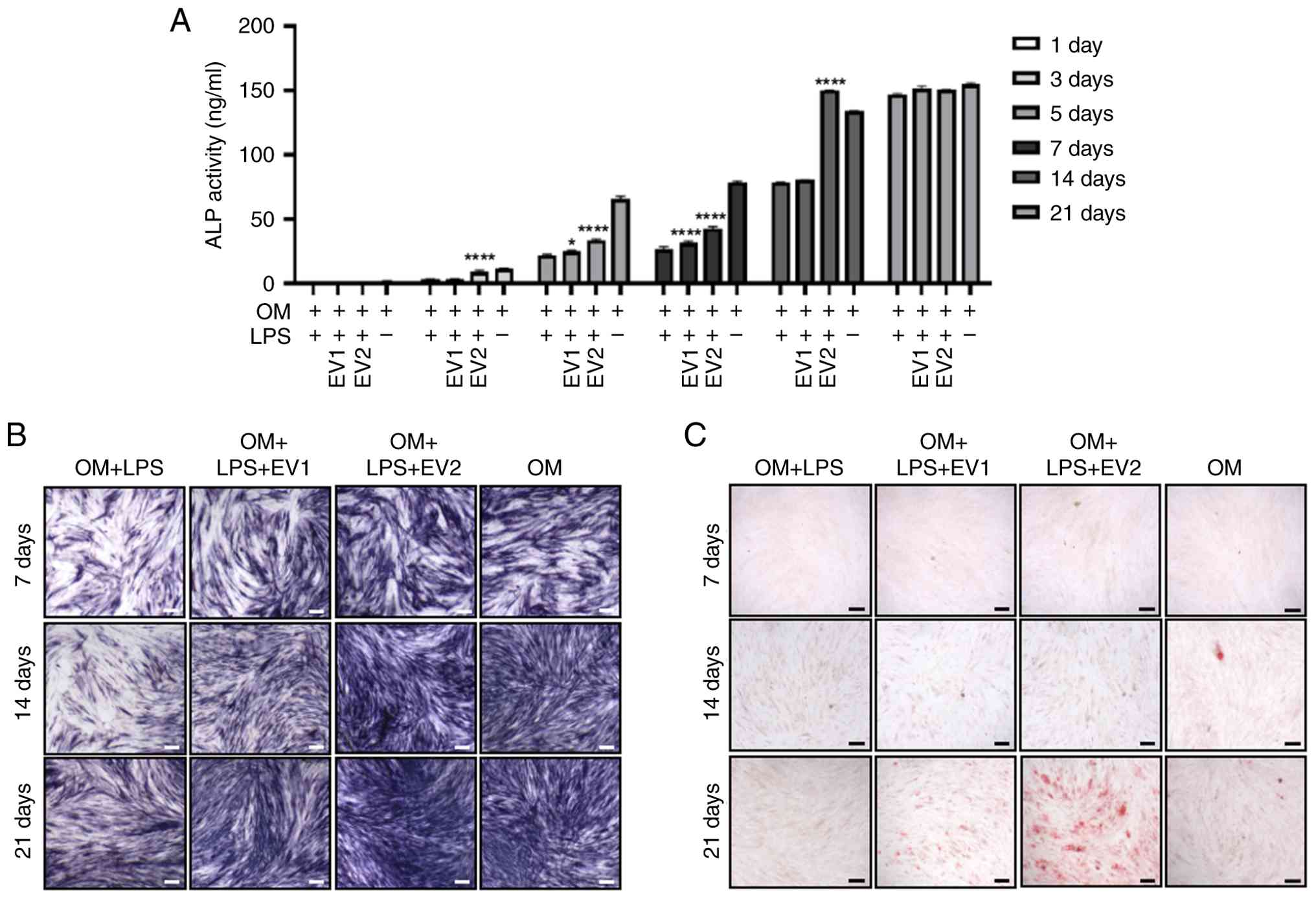 | Figure 5.T-MSC-derived EVs restore osteogenic
activity under LPS stimulation. (A) An ALP activity assay was
performed at 1, 3, 5, 7, 14 and 21 days. ALP activity remained
suppressed in the LPS (1 µg/ml)-treated group compared with the OM
group up to day 14. By contrast, compared with the LPS-treated
group, in the EV1-treated group, ALP activity was significantly
increased at day 5. In the EV2-treated group, LPS-suppressed ALP
activity was significantly restored at day 3. (B) The ALP staining
intensity was reduced in the LPS group, which was reversed by EV
treatment. Scale bar, 200 µm. (C) Alizarin red staining revealed
decreased mineralization in the LPS group at 21 days, while EV
treatment restored calcium deposition, suggesting enhanced bone
formation capacity. Scale bar, 200 µm. All experiments were
performed in triplicate. *P<0.05, ****P<0.0001, OM + LPS
group vs. OM + LPS + EV-treated groups. ALP, alkaline phosphatase;
EV1, T-MSC-derived EVs (1×108 particles/ml); EV2,
T-MSC-derived EVs (5×108 particles/ml); EV,
extracellular vesicle; LPS, lipopolysaccharide; OM, osteogenic
medium; T-MSC, tonsil-derived mesenchymal stem cell. |
EVs restore osteogenic gene expression
suppressed by LPS treatment
Fig. 6 shows
changes in gene expression induced by LPS and T-MSC-EVs. LPS
treatment resulted in an overall decrease in the mRNA expression of
key osteogenic genes, including ALP, BSP, OPN, and
OCN, with BSP and OPN exhibiting a more
pronounced suppression trend over time. In contrast, treatment with
EVs resulted in recovered or increased gene expression, with a
significant level of recovery observed for some indicators. In
particular, OPN and OCN, which are mid- and
late-stage osteogenic differentiation markers (17,18),
showed a tendency for expression to increase rapidly after 14 days
due to T-MSC-EVs. The levels of SOST, an osteogenic
inhibitor (19), increased
following LPS treatment but were suppressed again upon T-MSC-EV
treatment, suggesting the positive effect of EVs on the normal
regulation of osteogenic gene expression, even under inflammatory
conditions.
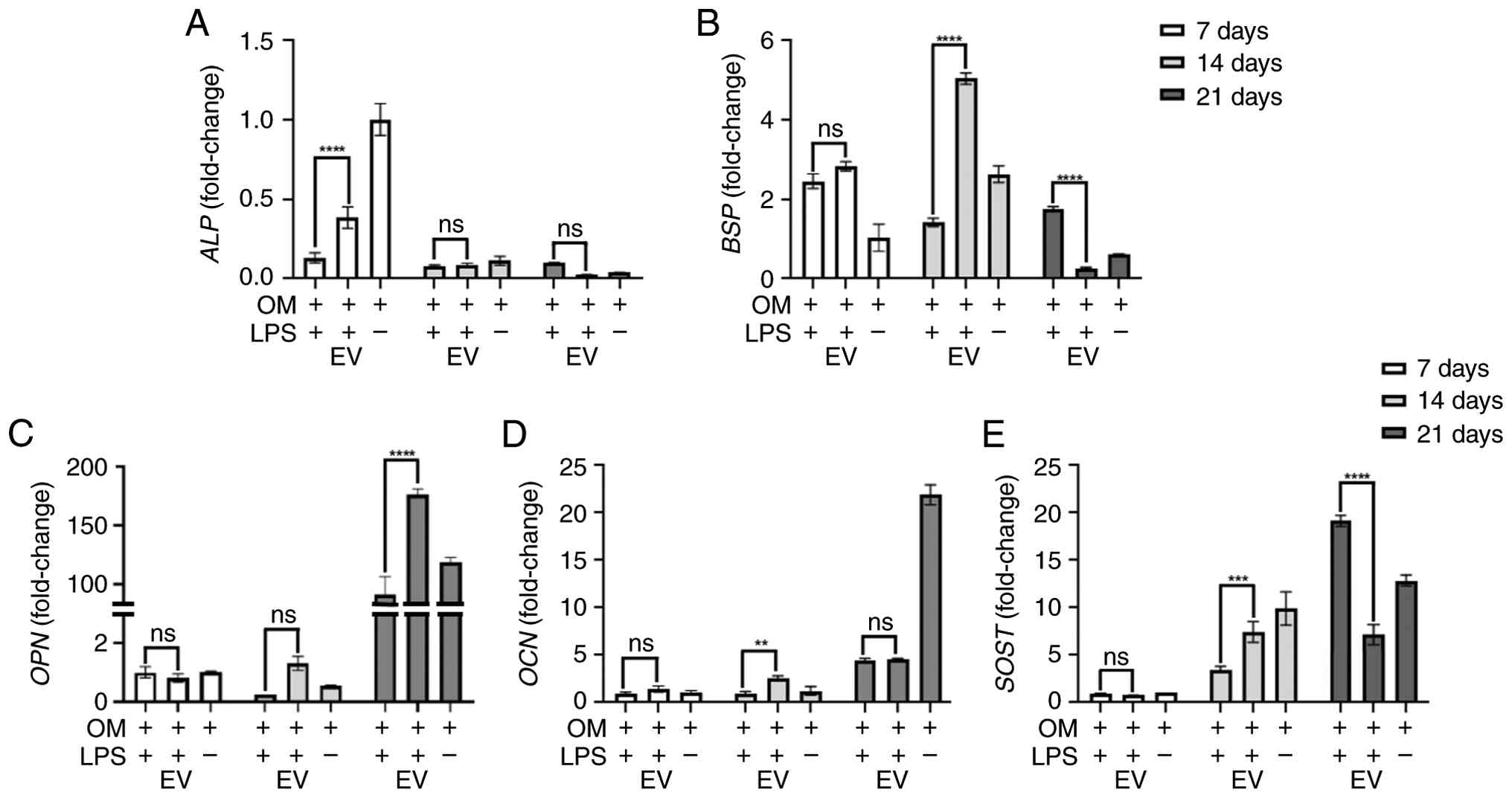 | Figure 6.Reverse transcription-quantitative
PCR analysis of osteogenic markers was conducted at 7, 14 and 21
days after treatment. (A) ALP mRNA expression. (B)
BSP mRNA expression. (C) OPN mRNA expression. (D)
OCN mRNA expression. (E) SOST mRNA expression.
ALP gene expression, which was reduced by LPS (1 µg/ml)
treatment, was enhanced by tonsil-derived mesenchymal stem
cell-derived EVs (5×108 particles/ml) at day 7.
BSP and OCN gene expression was increased following
LPS + EV treatment at day 14; however, BSP expression was
lower in the EV-treated group than in the LPS-treated group at day
21. SOST expression was markedly higher in the EV-treated
group than in the LPS-treated group at day 14, whereas at day 21,
SOST expression was significantly higher in the LPS-treated
group. All experiments were performed in triplicate. **P<0.01,
***P<0.001, ****P<0.0001, OM + LPS group vs. OM + LPS +
EV-treated group. ALP, alkaline phosphatase; BSP,
bone sialoprotein; EV, extracellular vesicle; LPS,
lipopolysaccharide; ns, not significant; OCN, osteocalcin;
OM, osteogenic medium; OPN, osteopontin; SOST,
sclerostin. |
Discussion
In present study, we evaluated the regulatory
effects and underlying mechanisms of EVs derived from T-MSC on
LPS-induced inflammation in hPDLFs. T-MSCs represent an attractive
source of stem cell because they can be obtained as surgical waste
after tonsillectomy, providing an easily accessible and ethically
favorable material. Additionally, T-MSCs are characterized by a
strong proliferative capacity and an immune-privileged phenotype,
which endows their EVs with enhanced immunomodulatory and
regenerative potential.
LPS activates innate immunity through TLR4, leading
to NF-κB and MAPK (p38, ERK, and JNK) signaling, which promotes
inflammatory cytokine production, inhibits cell proliferation, and
induces cell damage (20). Our
findings showed that T-MSC-EVs restored the LPS-induced suppression
of hPDLF proliferation. This protective effect was more evident at
higher particle doses.
T-MSC-EVs selectively modulated inflammatory
cytokines by suppressing IL-8 and IFN-γ, key
mediators of neutrophil recruitment and Th1 signaling (21,22),
while generally maintaining IL-6 and IL-1β levels,
which are essential for immune activation and tissue repair
(23). This selective cytokine
regulation, combined with EV-mediated inhibition of MAPK-AP-1
components (p-ERK, p-JNK, c-Fos, c-Jun), suggests that EVs
attenuate excessive inflammation while preserving pro-repair
signals (24).
In addition to controlling inflammation, T-MSC-EVs
also restored osteogenic activity under inflammatory conditions.
ALP activity and mineralized nodule formation were significantly
enhanced at later stages, and key osteogenic genes (ALP, BSP,
OPN, and OCN) showed marked recovery. Particularly,
OPN and OCN, which are markers of mid-to-late
osteogenic differentiation, were robustly reactivated (17,18).
Moreover, SOST, a negative regulator of bone formation, was
suppressed following T-MSC-EV treatment (19). These findings indicate that
T-MSC-EVs not only protect cells from inflammatory insults but also
actively promote bone regeneration by reinstating osteogenic gene
expression profiles.
Although the present study did not analyze the EV
cargo, the same T-MSC-EVs were previously reported to contain
multiple highly expressed miRNAs (25) including miR-199a-3p, miR-145-5p,
miR-24-3p, miR-214-3p, and let-7 family members. These enriched
miRNAs may target components of the MAPK-AP-1 and TLR4-associated
NF-κB pathways in hPDLFs, contributing to the selective reduction
of IL-8 and IFN-γ and to the reversal of LPS-induced proliferation
and osteogenic suppression. However, whether the miRNA and protein
cargo profiles of T-MSC-EVs in the present study, and their
functional relevance, are fully consistent with those reported
previously remains to be directly validated.
Several studies have reported the anti-inflammatory
and regenerative effects of EVs derived from various stem cell
sources (26–29). In the present study, T-MSC-EVs
exhibited significant immunomodulatory and regenerative potential,
likely attributable to the immune-privileged nature of
tonsil-derived cells and the enriched expression of
immune-regulatory genes (30,31).
These findings highlight the importance of selecting an appropriate
source of EVs. Nevertheless, challenges remain, including
heterogeneous EV composition, inconsistent therapeutic efficacy,
lack of standardized quantification protocols, and limitations in
large-scale production and storage stability (32). Addressing these issues is critical
for a successful clinical translation.
In summary, EVs derived from T-MSCs restored cell
viability, attenuated tissue-destructive cytokines, and preserved
regeneration-associated immune responses. They also inhibited the
MAPK-AP-1 signaling pathway and promoted osteogenic
differentiation, even under inflammatory conditions. These
multifaceted actions highlight T-MSC-EVs as intelligent biological
modulators that can fine-tune the immune response while also
supporting tissue regeneration. Higher concentrations of T-MSC-EVs
enhanced cell proliferation and differentiation. Future studies
should aim to elucidate active cargo components (e.g., miRNAs and
proteins), standardize production and quality control processes,
and validate the therapeutic potential of these EVs in vivo
and in clinical settings. Collectively, the results of this study
provide a robust scientific basis for the development of precise
regenerative therapies using stem cell-derived EVs.
Acknowledgements
Not applicable.
Funding
The present study was supported by the National Research
Foundation of Korea grant funded by the Korea government (grant no.
RS-2024-00341119).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
WJB obtained and analyzed the data and prepared the
graphical outputs. SKK contributed to the conception and
experimental design and critically revised the manuscript for
important intellectual content. HSK provided expert consultation on
the design of extracellular vesicle-related experiments and
contributed to data interpretation. SWK contributed to data
analysis and interpretation and critically revised the manuscript.
JYB conceived and supervised the study, contributed to experimental
design and data interpretation, critically revised the manuscript,
and finalized the manuscript. SWK and JYB confirm the authenticity
of all the raw data. All authors have read and approved the final
version of the manuscript.
Ethics approval and consent to
participate
The present study was conducted in accordance with
The Declaration of Helsinki and the research design was officially
approved by the Institutional Review Board of Kyung Hee University
(IRB No. KHSIRB-25-417; Seoul, South Korea).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ray RR: Periodontitis: An oral disease
with severe consequences. Appl Biochem Biotechnol. 195:17–32. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Sedghi LM, Bacino M and Kapila YL:
Periodontal disease: The good, the bad, and the unknown. Front Cell
Infect Microbiol. 11:7669442021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Peres MA, Macpherson LMD, Weyant RJ, Daly
B, Venturelli R, Mathur MR, Listl S, Celeste RK, Guarnizo-Herreño
CC, Kearns C, et al: Oral diseases: A global public health
challenge. Lancet. 394:249–260. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Socransky SS, Haffajee AD, Cugini MA,
Smith C and Kent RL Jr: Microbial complexes in subgingival plaque.
J Clin Periodontol. 25:134–144. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kwon T, Lamster IB and Levin L: Current
concepts in the management of periodontitis. Int Dent J.
71:462–476. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lekic P and McCulloch CA: Periodontal
ligament cell population: The central role of fibroblasts in
creating a unique tissue. Anat Rec. 245:327–341. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Proff P, Reicheneder C, Faltermeier A,
Kubein-Meesenburg D and Römer P: Effects of mechanical and
bacterial stressors on cytokine and growth-factor expression in
periodontal ligament cells. J Orofac Orthop. 75:191–202. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Maldonado RF, Sá-Correia I and Valvano MA:
Lipopolysaccharide modification in Gram-negative bacteria during
chronic infection. FEMS Microbiol Rev. 40:480–493. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Meng D, Wang Y and Liu T: Protective
effects of silibinin on LPS-induced inflammation in human
periodontal ligament cells. Front Chem. 10:10196632022. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yanuar A, Agustina H, Budhiparama NC and
Atik N: Prospect of exosome in ligament healing: A systematical
review. Stem Cells Cloning. 16:91–101. 2023.PubMed/NCBI
|
|
11
|
Qiao X, Tang J, Dou L, Yang S, Sun Y, Mao
H and Yang D: Dental pulp stem cell-derived exosomes regulate
anti-inflammatory and osteogenesis in periodontal ligament stem
cells and promote the repair of experimental periodontitis in rats.
Int J Nanomedicine. 18:4683–4703. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Tao Y, Liu T, Jing F, Tan X, Zhao X,
Bernaerts KV, Jia R, Zhao J, Yin Y and Zhang T: Adipose-derived
stem-cell-derived exosomes encapsulated patch for modulating
inflammation and promoting tissue regeneration. ACS Nano.
19:21271–21289. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Cho KA, Cha JE, Kim J, Kim YH, Ryu KH and
Woo SY: Mesenchymal stem cell-derived exosomes attenuate
TLR7-mediated mast cell activation. Tissue Eng Regen Med.
19:117–129. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bae WJ, Auh QS, Kim GT, Moon JH and Kim
EC: Effects of sodium tri- and hexameta-phosphate in vitro
osteoblastic differentiation in Periodontal Ligament and
osteoblasts, and in vivo bone regeneration. Differentiation.
92:257–269. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang J, Moosavizadeh S, Jammes M, Tabasi
A, Bach T, Ryan AE and Ritter T: In-vitro immunomodulatory efficacy
of extracellular vesicles derived from TGF-β1/IFN-γ dual licensed
human bone marrow mesenchymal stromal cells. Stem Cell Res Ther.
16:3572025. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhu S, Chen W, Masson A and Li YP: Cell
signaling and transcriptional regulation of osteoblast lineage
commitment, differentiation, bone formation, and homeostasis. Cell
Discov. 10:712024. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Huang W, Yang S, Shao J and Li YP:
Signaling and transcriptional regulation in osteoblast commitment
and differentiation. Front Biosci. 12:3068–3092. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
van Bezooijen RL, ten Dijke P, Papapoulos
SE and Löwik CW: SOST/sclerostin, an osteocyte-derived negative
regulator of bone formation. Cytokine Growth Factor Rev.
16:319–327. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li Z, Scott MJ, Fan EK, Li Y, Liu J, Xiao
G, Li S, Billiar TR, Wilson MA, Jiang Y and Fan J: Tissue damage
negatively regulates LPS-induced macrophage necroptosis. Cell Death
Differ. 23:1428–1447. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bosco MC, Gusella GL, Espinoza-Delgado I,
Longo DL and Varesio L: Interferon-gamma upregulates interleukin-8
gene expression in human monocytic cells by a posttranscriptional
mechanism. Blood. 83:537–542. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Rodrigues DR, Fernandes RK, Balderramas
HDA, Penitenti M, Bachiega TF, Calvi SA, Dias-Melicio LA, Ikoma MR
and Soares ÂM: Interferon-gamma production by human neutrophils
upon stimulation by IL-12, IL-15 and IL-18 and challenge with
Paracoccidioides brasiliensis. Cytokine. 69:102–109. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hirano T: IL-6 in inflammation,
autoimmunity and cancer. Int Immunol. 33:127–148. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Yan L, Wang J, Cai X, Liou YC, Shen HM,
Hao J, Huang C, Luo G and He W: Macrophage plasticity: Signaling
pathways, tissue repair, and regeneration. MedComm (2020).
5:e6582024. View
Article : Google Scholar : PubMed/NCBI
|
|
25
|
Choi DW, Cho KA, Kim J, Kim J, Lee HJ, Kim
YH, Park JW and Woo SY: Extracellular vesicles from tonsil-derived
mesenchymal stromal cells show anti-tumor effect via miR-199a-3p.
Int J Mol Med. 48:2212021. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ahmad P, Estrin N, Farshidfar N, Zhang Y
and Miron RJ: Mechanistic insights into periodontal ligament stem
cell-derived exosomes in tissue regeneration. Clin Oral Investig.
29:3572025. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Feng H, Gong S, Liu J, Aghayants S, Liu Y,
Wu M, Wu Y and Song J: Adipose-derived stem cell exosomes:
Mechanisms and therapeutic potentials in wound healing. Biomark
Res. 13:882025. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Tan F, Li X, Wang Z, Li J, Shahzad K and
Zheng J: Clinical applications of stem cell-derived exosomes.
Signal Transduct Target Ther. 9:172024. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ning X, Liu R, Huang Y, Huang Z, Li H, Li
Q, Sheng Z and Wu J: Dental stem cell-derived exosomes: A review of
their isolation, classification, functions, and mechanisms. Stem
Cells Int. 2024:21873922024. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kim DK, Lee HJ, Lee IH and Lee JJ:
Immunomodulatory effects of primed tonsil-derived mesenchymal stem
cells on atopic dermatitis via B cell regulation. Cells. 13:802023.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Shin SC, Seo Y, Park HY, Jung DW, Shin TH,
Son H, Kim YK, Lee JC, Sung ES, Jang JY, et al: Regenerative
potential of tonsil mesenchymal stem cells on surgical cutaneous
defect. Cell Death Dis. 9:1832018. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Palakurthi SS, Shah B, Kapre S, Charbe N,
Immanuel S, Pasham S, Thalla M, Jain A and Palakurthi S: A
comprehensive review of challenges and advances in exosome-based
drug delivery systems. Nanoscale Adv. Oct 29–2024.(Epub ahead of
print). doi: 10.1039/d4na00501e. View Article : Google Scholar : PubMed/NCBI
|