Introduction
Periodontitis is a chronic inflammatory disease of
the oral cavity that arises from untreated gingivitis,
progressively destroying the tooth-supporting structures and
leading to tooth loss. The condition is typically associated with
immune dysregulation, bacterial infection and systemic disease such
as diabetes (1). Although the
prevalence of severe periodontitis increases with age, it can
affect individuals across all age groups. In 2011–2020, the
estimated rate of dentate periodontitis in adults was ~62% and the
rate of severe periodontitis was 23.6% (2). Diagnosis typically involves clinical
and radiographic evaluation of periodontal tissue, with hallmark
features including alveolar bone resorption and deepened
periodontal pockets. Treatment for mild periodontitis typically
involves mechanical debridement and antibiotic therapy. However,
severe periodontitis results in irreversible destruction of
periodontal tissue and therapeutic efforts typically focus on
promoting alveolar bone regeneration (3–5). The
periodontal ligament, a specialized connective tissue layer,
anchors the tooth to the alveolar bone, supplies nutrients and
contributes to tissue repair through the regulation of cell
proliferation, growth factors and calcification homeostasis
(6–8). Periodontal ligament stem cells
(PDLSCs) possess multipotent differentiation capacity and can give
rise to osteoblasts, adipocytes and periodontal neuronal cells
(9). Owing to their regenerative
potential and compatibility with biomaterials, PDLSCs are regarded
as good seed cells for periodontal tissue regeneration (10,11).
The MAPK signaling pathway is a key intracellular
cascade regulating cell proliferation, differentiation, apoptosis
and stress responses (12,13). This pathway is markedly involved in
the pathogenesis of chronic periodontitis, influencing inflammatory
responses and alveolar bone remodeling (14). Furthermore, the osteogenic
differentiation potential of PDLSCs is associated with MAPK
signaling (15), underscoring its
dual importance in both disease progression and tissue
regeneration. The MAPK pathway operates through a phosphorylation
cascade in which MAP kinase kinase kinases (MAPKKKs), such as RAF
family members, activate MAP kinase kinases, which phosphorylate
MAPKs that translocate to the nucleus to directly phosphorylate
transcription factors or activate downstream kinases to modulate
gene expression (16). C-Raf
proto-oncogene serine/threonine-protein kinase (CRAF), a
representative member of the RAF family, contains three conserved
regions (CR1, CR2 and CR3) and participates in cellular processes
through downstream MAPK signaling by dimerizing and phosphorylating
MEK to activate the MEK-ERK pathway (17). RuvB-like AAA ATPase-1 (RUVBL1), a
multifunctional ATPase, is implicated in SC maintenance,
differentiation and migration by functioning as an essential
component of chromatin remodeling complexes thereby regulating
transcription, DNA repair and protein assembly (18). RUVBL1 is a CRAF-binding protein
that activates the RAF/MEK/ERK pathway by preventing
phosphorylation at serine 259 within the CR2 domain of CRAF
(19). RUVBL1 may interact with
CRAF to promote or inhibit MAPK pathway activation in PDLSCs,
potentially impacting periodontal regeneration. However, this
hypothesis remains to be experimentally validated. In addition,
oligodeoxynucleotide (ODN)-MT01, an inhibitory ODN designed based
on human mitochondrial DNA, attenuates alveolar bone resorption and
enhances osteogenic differentiation of bone marrow mesenchymal SCs
(MSCs) in periodontitis models via ERK- and p38-mediated MAPK
signaling (20,21). However, whether ODN MT01 enhances
PDLSC-mediated periodontal regeneration remains unclear.
The present study aimed to elucidate the roles of
RUVBL1 and CRAF in PDLSCs and their involvement in the MAPK
signaling pathway in the treatment of periodontitis. The present
findings may provide mechanistic insight into the osteogenic
potential of PDLSCs and their application in periodontal tissue
regeneration.
Materials and methods
Cell culture
Primary human PDLSCs (cat. no. CP-H234; passage
three) were obtained from Procell Life Science & Technology
Co., Ltd. and cultured in DMEM/F12 basal medium (88%; cat. no.
12400-024; Gibco; Thermo Fisher Scientific, Inc.) containing FBS
(10%; cat. no. 10099-141; Gibco; Thermo Fisher Scientific, Inc.),
penicillin-streptomycin (P/S, 1%; cat. no. 1902417; Gibco; Thermo
Fisher Scientific, Inc.) and glutamine (1%; cat. no. 1894153;
Gibco; Thermo Fisher Scientific, Inc.). The cells were incubated at
37°C with 5% CO2 and 95% humidity.
Flow cytometry
Cell suspension was placed in a flow tube at the
1×106 cells/ml and 5 µl anti-human CD90 FITC (cat. no.
555595; dilution 1:20; BD Biosciences), CD105 APC (cat. no. 562408;
dilution 1:20; BD Biosciences) and CD45 PE-Cy7 (cat. no. 557748;
dilution 1:20; BD Biosciences) fluorescent-labeled antibodies was
added. Cells were incubated at 4°C in the dark for 30 min.
Following incubation, the cells were washed with PBS containing 2%
FBS and centrifuged at 4°C at 300 × g for 5 min. The cell pellet
was resuspended in 500 µl PBS and analyzed immediately. Flow
cytometry was performed using a BD FACSCanto II flow cytometer (BD
Biosciences) equipped with 488 and 633 nm lasers. Data acquisition
and analysis were performed using BD FACSDiva software (version
8.0; BD Biosciences).
Cell induction
PDLSCs were cultured for 14 days at 37°C with 5%
CO2 in a DMEM/F12 medium (cat. no. 12400-024; Gibco;
Thermo Fisher Scientific, Inc.) containing osteogenic or lipogenic
inducers: Osteogenic inducers were as follows: 10−8
mol/l dexamethasone (cat. no. D4902; MilliporeSigma), 50 µg/ml
vitamin C (cat. no. A8960; MilliporeSigma) and 10 mmol/l sodium
β-glycerophosphate (cat. no. G9422; MilliporeSigma). Lipogenic
inducers were as follows: 200 µmol/ml indomethacin (cat. no. I7378;
MilliporeSigma), 10 µg/ml insulin (cat. no. I3536; MilliporeSigma),
0.5 mmol/ml IBMX (cat. no. I5879; MilliporeSigma) and 1 µmol/ml
dexamethasone (cat. no. D4902; MilliporeSigma). To explore the
potential of ODN MT01, a synthetic ODN known to promote osteoblast
maturation (20), in PDLSC
osteogenesis, cells were treated with ODN MT01 (0.5–4.0 µg/ml)
following osteogenic induction.
Construction of overexpression and
interference plasmids
Overexpression or knockdown plasmids and negative
controls (NCs) were constructed by Genomeditech (Shanghai) Co.,
Ltd. The CRAF overexpression vector was
PGMLV-CMV-MCS-3×Flag-PGK-Puro, the RUVBL1 overexpression vector was
PGMLV-CMV-MCS1-3×FIag-PGK-PuroxFIag-PGK-Puro and the interference
vector was pGMLV-SC5 RNA inteference. For the overexpression NC,
insert-free PGMLV-CMV-MCS-3×Flag-PGK-Puro (for CRAF) and
PGMLV-CMV-MCS1-3×FIag-PGK-PuroxFIag-PGK-Puro (for RUVBL1) were
used. NC and short hairpin (sh)RNA targets were as follows: sh-NC,
5′-TTCTCCGAACGTGTCACGT-3′; sh-CRAF-1, 5′-GGAGTAACATCAGACAACTCT-3′;
sh-CRAF-2, 5′-GGATTTCGATGTCAGACTTGT-3′; sh-CRAF-3,
5′-GAAGACGTTCCTGAAGCTTGC-3′; sh-RUVBL1-1,
5′-GGGAGTGAAGTTTACTCAACT-3′; sh-RUVBL1-2,
5′-GCCACAGAATTCGACCTTGAA-3′; and sh-RUVBL1-3,
5′-GTCCATGATGGGCCAGCTAAT-3′. Lentiviral particles were produced
using a third-generation packaging system in 293T cells (ATCC
CRL-3216). For packaging, 20 µg lentiviral plasmid (pGMLV-CRAF,
pGMLV-RUVBL1 or pGMLV-shRNA) was co-transfected with 10 µg psPAX2
(cat. no. 12260; Addgene) and 5 µg pMD2.G envelope plasmid (cat.
no. 12259; Addgene) at a 4:2:1 mass ratio using
Lipofectamine® 3000 (cat. no. L3000015; Thermo Fisher
Scientific, Inc.). The transfection complex was incubated with 293T
cells for 6–8 h at 37°C, and viral supernatants were collected at
48 h and 72 h, filtered (0.45 µm; cat. no. SLHV033RS;
MilliporeSigma) and concentrated using Lenti-X Concentrator (cat.
no. 631232; Takara Bio, Inc.). PDLSCs were infected at an MOI of
10–20 with 8 µg/ml polybrene (cat. no. H9268; MilliporeSigma) for
24 h at 37°C. Puromycin (cat. no. P8833; MilliporeSigma; 2 µg/ml
for selection and 1 µg/ml for maintenance) was added 48 h
post-infection. Reverse transcription-quantitative (RT-q)PCR and
western blotting were used to perform expression validation and
shRNA lentiviral screening.
Cell Counting Kit-8 (CCK-8) assay
Cells were inoculated into 96-well plates at a
density of 2.5×103 cells/well. The medium was replaced
with complete medium [DMEM/F12 (cat. no. 11330032; Gibco; Thermo
Fisher Scientific, Inc.) without P/S when the cell fusion reached
40–50%. CCK-8 assay was performed according to the manufacturer's
instructions (cat,. no. CK-04; Dojindo Laboratories). Briefly, 10
µl CCK-8 reagent was added to each well and incubated for 2 h at
37°C with 5% CO2. Absorbance was measured at 450 nm.
RT-qPCR
RNA was extracted using using Trizol®
solution (Thermo Fisher Scientific, Inc.) in an ice bath. cDNA was
obtained according to the instructions of the FastKing cDNA First
Strand Synthesis kit (cat. no. KR116; Tiangen Biotech Co., Ltd.)
and amplified. The color developer was the Taq Pro Universal SYBR
qPCR Master Mix (cat. no. Q712-03; Vazyme Biotech, Co., Ltd.).
Thermocycling conditions were as follows: 95°C for 30 sec; followed
by 40 cycles of 95°C for 10 sec and 60°C for 30 sec. GAPDH was used
as an internal reference gene. Experimental results were quantified
using the 2−ΔΔCq method (22). Primer information is listed in
Table I.
 | Table I.Primer information. |
Table I.
Primer information.
| Gene | Forward
(5′-3′) | Reverse
(5′-3′) |
|---|
| GAPDH |
AAAGGGTCATCATCTCTG |
GCTGTTGTCATACTTCTC |
| CRAF |
ACTCCTATGGCATCGTATT |
TCTGATCTCGGTTGTTGA |
| RUVBL1 |
GATACCTATGCCACAGAAT |
ATTAGCCACATCCAAGTC |
| ARAF |
TTCTGTGACTTCTGCCTTA |
ATGCTGGTGGAACTTGTA |
| BRAF |
CTTGTATCACCATCTCCATA |
GGCGTGTAAGTAATCCAT |
| MEK1 |
CAGTGGAGTGTTCAGTCT |
ACTTCCTCAGCATCAGAT |
| MEK2 |
CTGGACTATATTGTGAAC |
TTGATGAGGCATTTATTG |
| ERK1 |
GATGGAGACTGACCTGTA |
CTGGTAGAGGAAGTAGCA |
| ERK2 |
TGGTGTGCTCTGCTTATG |
AGTAGGTCTGGTGCTCAA |
Western blotting
Protein extraction from cells was performed with
RIPA (Beyotime Biotechnology) containing protease inhibitors [Roche
Diagnostics (Shanghai) Co., Ltd]. Protein concentration was
determined using the BCA Protein Assay Kit (cat. no. P0012;
Beyotime Biotechnology). Sample proteins (60 µg per lane) were
subjected to SDS-PAGE (10% gel) electrophoresis, PVDF membrane
transfer and incubated with skimmed milk (5%) for 2 h at room
temperature. Primary antibodies (5 ml) were added overnight at 4°C
in a refrigerator. Enzyme-labeled secondary antibody was added and
incubated for 2 h at room temperature. Color development was
performed using an ECL kit (cat. no. P0018; Beyotime
Biotechnology). Chemiluminescent signals were detected using
ChemiDoc XRS+ Imaging System (Bio-Rad Laboratories) and analyzed
using Image Lab software (version 6.1; Bio-Rad Laboratories). The
antibodies were as follows: GAPDH (1:1,000; cat. no. P30008M;
Abmart Pharmaceutical Technology Co., Ltd.), A-Raf proto-oncogene
serine/threonine-protein kinase (ARAF) (cat. no. bs-2251R), CRAF
(cat. no. bs-23170R), MEK1/2 (cat. no. bs-1041R), phosphorylated
(p)-MEK1/2 (cat. no. bs-3270R), ERK1/2 (cat. no. bsm-33232M),
p-ERK1/2 (cat. no. bs-3016R; all BIOSS), RUVBL1 (cat. no. 74775;
Cell Signaling Technology, Inc.), ERK1/2 (all 1:500; cat. no.
bsm-33232M; BIOSS), B-Raf proto-oncogene serine/threonine-protein
kinase (BRAF) (1:2,000; cat. no. ab33899; Abcam) and HRP-conjugated
universal secondary antibody [cat. nos. 7074 (anti-rabbit) and 7076
(anti-mouse); Cell Signaling Technology, Inc.].
Oil Red O staining
Cell slides were fixed in paraformaldehyde (4%) for
10 min at room temperature. The slides were washed in 60%
isopropanol for 20 sec and stained for 10 min with Oil Red O at
room temperature. Slides were differentiated with 60% isopropanol
and the excess dye was removed. Finally, the slides were restained
in hematoxylin (0.5%) for 2 min at room temperature and rinsed in
distilled water before being dried and sealed for microscopic
observation. The stained sections (5-µm thick) were observed under
a light microscope (Eclipse E100; Nikon Corporation).
Alkaline phosphatase (ALP)
staining
ALP staining was performed using ALP Staining
Solution (cat. no. G1480; Solarbio Science & Technology Co.,
Ltd.) according to the manufacturer's instructions. Cells were
fixed with ALP fixative (4% paraformaldehyde; 0.5 ml per well) for
3 min at room temperature and incubated with ALP incubation
solution (BCIP/NBT working solution; 0.5 ml per well) for 30 min at
room temperature, with PBS washing after each step. Finally,
microscopic examination was performed using a light microscope
(Eclipse E100; Nikon Corporation).
Alizarin red staining
Cells were fixed using a paraformaldehyde solution
(4%) for 10 min at room temperature. Staining was performed with
0.1% alizarin red-Tris-HCl (pH 8.3) solution for 30 min at room
temperature. Rinsing was performed with distilled water and drying
were performed for sealing. The degree of cell mineralization was
observed under the light microscope (Eclipse E100; Nikon
Corporation).
Statistical analysis
Data are presented as the mean ± SD from three
independent experiments (n=3). One-way ANOVA followed by Tukey's
HSD post hoc test was performed using SPSS (version 23.0; IBM
Corp.) under a two-tailed significance level. Visualization was
conducted using Origin® 2021 software (OriginLab
Corporation). P<0.05 was considered to indicate a statistically
significant difference.
Results
PDLSCs exhibit clonogenic capacity and
multilineage differentiation potential
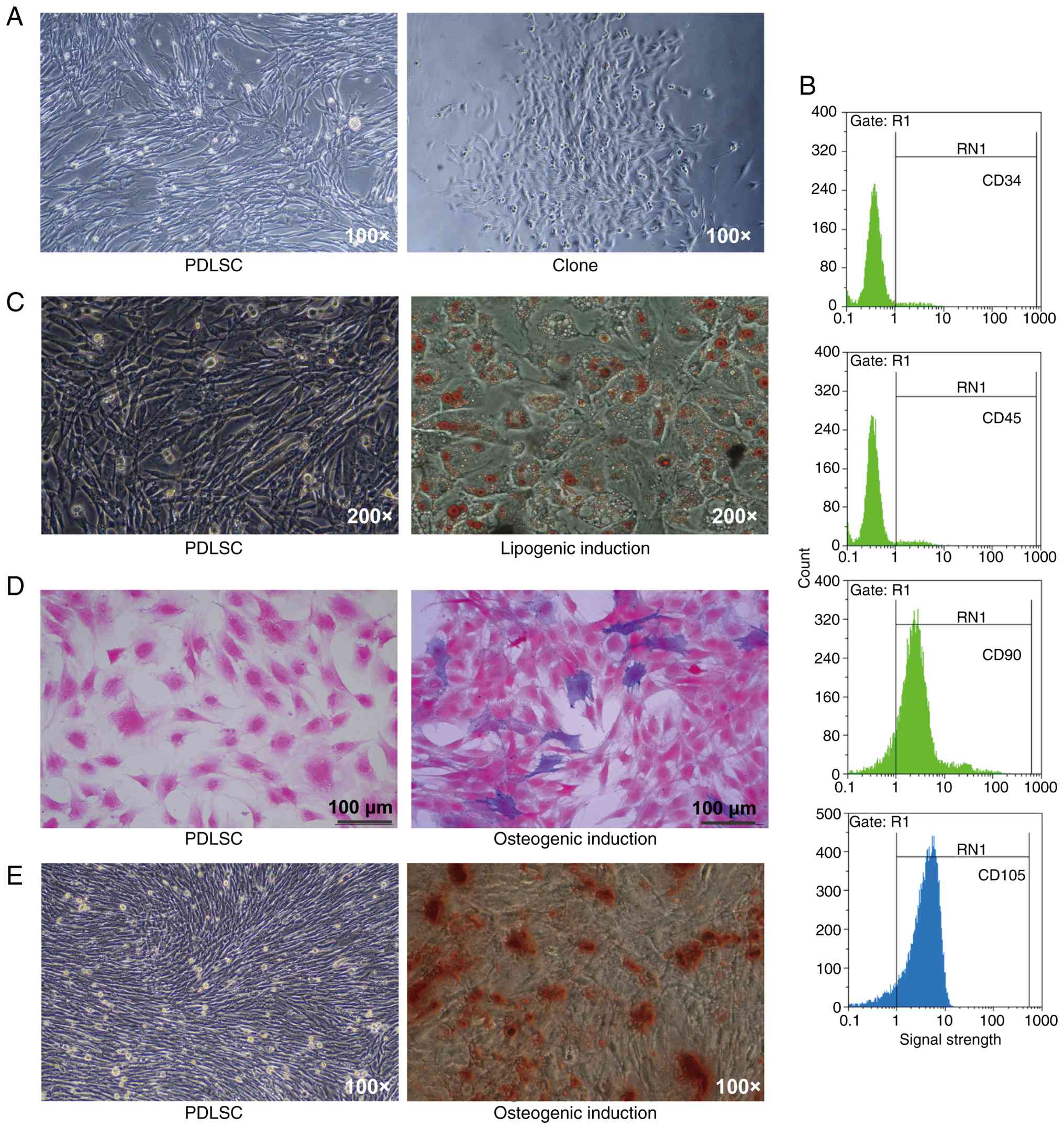
Following successful cloning (Fig. 1A), the expression of MSC surface
markers in PDLSCs was analyzed by flow cytometry. The cells were
positive for CD90 and CD105 but negative for CD34 and CD45
(Fig. 1B), demonstrating their MSC
phenotype. PDLSCs possessed both adipogenic and osteogenic
differentiation potential (Fig. 1C and
D). Alizarin red staining revealed a marked increase in
mineralized nodule formation following osteogenic induction
(Fig. 1E). Collectively, these
results indicated that the isolated PDLSCs exhibited robust
self-renewal and multipotent differentiation capabilities.
PDLSC viability is positively
associated with CRAF and negatively associated with RUVBL1
To investigate the roles of CRAF and RUVBL1 in PDLSC
viability, lentiviral transfection was performed. qPCR and western
blotting analyses determined the successful overexpression of CRAF
(Fig. 2A and C) and RUVBL1
(Fig. 2B, D), as well as effective
knockdown using sh-CRAF-3 and sh-RUVBL1-1 (Fig. 2E-H). CCK-8 assay demonstrated that
CRAF overexpression markedly enhanced PDLSC viability, whereas CRAF
knockdown suppressed it (Fig. 2I).
Conversely, RUVBL1 overexpression decreased PDLSC viability,
whereas RUVBL1 knockdown restored it (Fig. 2J). These findings indicated that
PDLSC viability was positively regulated by CRAF and negatively
regulated by RUVBL1.
RUVBL1 and CRAF enhance osteogenic
differentiation, whereas CRAF promotes while RUVBL1 inhibits lipid
accumulation
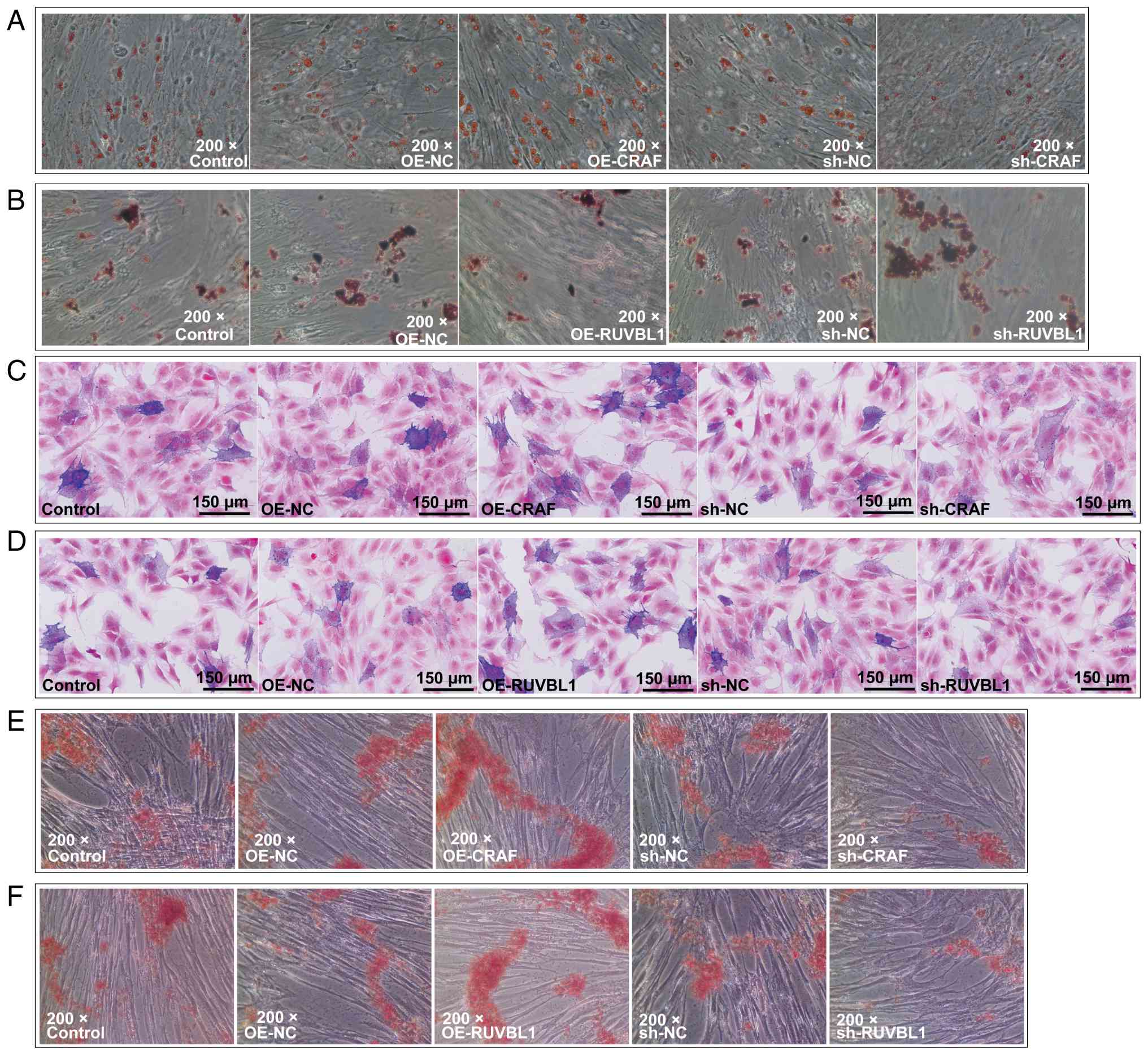
To examine the effects of RUVBL1 and CRAF on PDLSC
differentiation, Oil Red O, ALP and alizarin red staining were
performed. CRAF overexpression enhanced adipogenic differentiation,
whereas CRAF knockdown suppressed it (Fig. 3A). By contrast, RUVBL1
overexpression inhibited adipogenic differentiation, while its
knockdown enhanced lipid accumulation (Fig. 3B). Both CRAF and RUVBL1
overexpression increased ALP activity and mineralized nodule
formation, indicating enhanced osteogenic differentiation (Fig. 3C-F). Knockdown of either gene
reversed these effects, leading to decreased ALP activity and
mineralization. Collectively, these data demonstrated that both
RUVBL1 and CRAF promoted osteogenic differentiation of PDLSCs,
while exhibiting divergent effects on adipogenesis.
RUVBL1 does not regulate CRAF, but
both activate the MEK/ERK signaling pathway
As the MAPK pathway serves a key role in PDLSC
differentiation (12), the
association between RUVBL1, CRAF and this pathway was assessed.
Overexpression of CRAF did not significantly affect ARAF or BRAF
expression at either the mRNA or protein level (Fig. 4A, B, H and I). Similarly, RUVBL1
overexpression did not significantly alter the expression of ARAF,
BRAF or CRAF (Fig. 5A-C and H-J),
suggesting that RUVBL1 did not regulate CRAF. However, CRAF
overexpression significantly increased MEK1/2 and ERK1/2 expression
and phosphorylation compared with NCs, while CRAF knockdown
resulted in the opposite effect (Fig.
4C-G and J-P). Similarly, RUVBL1 overexpression significantly
elevated MEK1/2 and ERK1/2 mRNA and protein levels, as well as
their phosphorylation, whereas RUVBL1 knockdown significantly
suppressed them compared with NCs (Fig. 5D-G and K-P). These results indicate
that although RUVBL1 did not directly regulate CRAF, both proteins
may have independently activated the MEK/ERK signaling pathway in
PDLSCs.
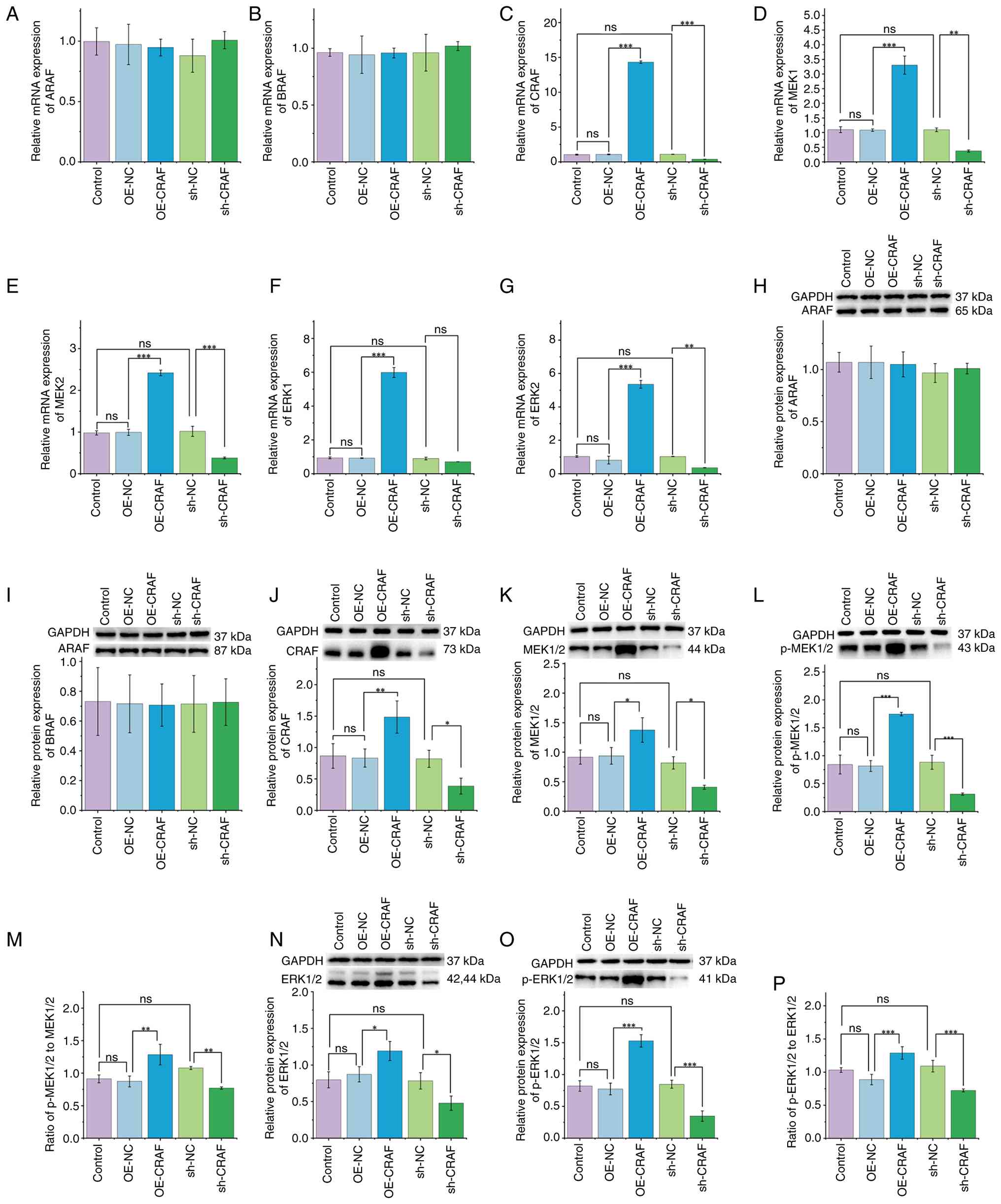 | Figure 4.Overexpression of CRAF activates the
RAF/MEK/ERK signaling pathway. Reverse transcription-quantitative
PCR was used to analyze the mRNA expression of (A) ARAF, (B) BRAF,
(C) CRAF, (D) MEK1, (E) MEK2, (F) ERK1and (G) ERK2. Western
blotting was used to analyze the protein levels of (H) ARAF, (I)
BRAF, (J) CRAF, (K) MEK1/2, (L) p-MEK1/2, (M) p-MEK1/2 to MEK1/2,
(N) ERK1/2, (O) p-ERK1/2 and (P) p-ERK1/2 to ERK1/2. *P<0.05,
**P<0.01 and ***P<0.001. p-, phosphorylated; OE,
overexpression; NC, negative control; sh, short hairpin; ns, not
significant; ARAF, A-Raf proto-oncogene serine/threonine-protein
kinase; BRAF, B-Raf proto-oncogene serine/threonine-protein kinase;
CRAF, C-Raf proto-oncogene serine/threonine-protein kinase. |
 | Figure 5.Overexpression of RUVBL1 activates the
MEK/ERK signaling pathway. Reverse transcription-quantitative PCR
was used to analyze the mRNA expression of (A) ARAF, (B) BRAF, (C)
CRAF, (D) MEK1, (E) MEK2, (F) ERK1 and (G) ERK2. Western blotting
was used to analyze protein levels of (H) ARAF, (I) BRAF, (J) CRAF,
(K) MEK1/2, (L) p-MEK1/2, (M) p-MEK1/2 to MEK1/2, (N) ERK1/2, (O)
p-ERK1/2 and (P) p-ERK1/2 to ERK1/2. *P<0.05, **P<0.01 and
***P<0.001. RUVBL1, RuvB-like AAA ATPase-1; p-, phosphorylated;
OE, overexpression; NC, negative control; sh, short hairpin; ns,
not significant; ARAF, A-Raf proto-oncogene
serine/threonine-protein kinase; BRAF, B-Raf proto-oncogene
serine/threonine-protein kinase; CRAF, C-Raf proto-oncogene
serine/threonine-protein kinase. |
ODN MT01 enhances RUVBL1- and
CRAF-mediated osteogenic differentiation
ALP activity increased in a dose-dependent manner
and 4.0 µg/ml was used for subsequent experiments (Fig. 6A and B). ODN MT01 markedly enhanced
osteogenic differentiation of PDLSCs (Fig. 6C and D). In addition, combined
treatment with ODN MT01 and overexpression of CRAF or RUVBL1
further augmented osteogenic differentiation (Fig. 6E and F). Conversely, knockdown of
CRAF or RUVBL1 decreased osteogenesis, however this inhibitory
effect was partially rescued by ODN MT01 treatment (Fig. 6G and H). These findings suggested
that ODN MT01 promoted PDLSC osteogenic differentiation and acted
additively with with RUVBL1 and CRAF activation to potentially
enhance bone-forming potential.
 | Figure 6.ODN MT01 promotes osteogenic
differentiation of PDLSCs. (A) PDLSC proliferation and (B) ALP
staining following treatment with varying concentrations of ODN
MT01. (C) Osteogenic differentiation and (D) mineralization after
the addition of ODN MT01 and osteogenic inducers. ALP staining was
used to assess the effect of (E) CRAF and (F) RUVBL1 on osteogenic
differentiation of PDLSCs following the addition of ODN MT01 and
osteogenic inducers. Alizarin Red staining was used to assess the
effect of (G) CRAF and (H) RUVBL1 on the degree of mineralization
of PDLSCs following the addition of ODN MT01 and osteogenic
inducers. *P<0.05 and ***P<0.001. ODN, oligodeoxynucleotide;
PDLSC, periodontal ligament stem cell; ALP, alkaline phosphatase,
RUVBL1, RuvB-like AAA ATPase-1; NC, negative control; OE,
overexpression; sh, short hairpin; ns, not significant; CRAF, C-Raf
proto-oncogene serine/threonine-protein kinase; OI, osteogenic
induction; OM, ODN MT01. |
Discussion
As pluripotent SCs with multilineage differentiation
potential, PDLSCs serve a key role in the regeneration of
periodontal tissue. Their osteogenic and adipogenic differentiation
capacities are important for periodontitis therapy. RUVBL1, an
ATP-binding protein belonging to the AAA+ ATPase family,
participates in cellular processes, including DNA damage repair,
transcriptional regulation and chromatin remodeling (23). A previous study has suggested that
RUVBL1 serves as a negative regulator of cell differentiation
(24). However, its specific role
in PDLSC differentiation has not been fully elucidated. The present
study demonstrated that RUVBL1 overexpression enhanced the
osteogenic potential of PDLSCs while suppressing adipogenic
differentiation. Notably, although RUVBL1 positively regulates cell
proliferation (25), the present
results indicated RUVBL1 overexpression reduced PDLSC viability.
Excessive RUVBL1 expression may induce replication stress, leading
to cell cycle arrest and a shift from proliferation toward
osteogenic differentiation (26).
CRAF, a member of the RAF kinase family, regulates SC function
through both MAPK-dependent and -independent mechanisms (27), though to the best of our knowledge,
its role in PDLSCs has not been previously reported. In the present
study, CRAF overexpression enhanced PDLSC viability and increased
both osteogenic and adipogenic differentiation. By contrast, RUVBL1
promoted osteogenesis but suppressed viability and adipogenesis.
RUVBL1 promotes a shift from mTOR-driven lipogenesis towards
AMPK-induced fatty acid catabolism in a hepatocellular carcinoma
model (28). Although this
differed from the present research model and may reflect
cell-specificity, it offers a potential explanation of the present
findings, suggesting that the divergent effects of RUVBL1 and CRAF
on lipid synthesis may stem from distinct signaling pathways.
Collectively, these data suggested that both RUVBL1 and CRAF
enhance osteogenic differentiation through distinct regulatory
mechanisms.
The MAPK signaling pathway is implicated in PDLSC
osteogenic differentiation (12,15).
Extracellular stimuli are important in activation of the MAPK
pathway (29). ERK, a traditional
MAPK, primarily regulates cell proliferation and differentiation
(30). As a MAPKKK, CRAF activates
MEK1/2, which phosphorylates ERK1/2 to regulate downstream
transcriptional responses associated with proliferation,
differentiation and apoptosis (31). Consistent with these canonical
functions, the present study demonstrated that CRAF overexpression
increased MEK and ERK phosphorylation, thereby activating the
MEK/ERK signaling pathway to enhance PDLSC proliferation and
differentiation. RUVBL1 is a CRAF-binding protein capable of
activating the RAF/MEK/ERK pathway by preventing phosphorylation at
serine 259 within the CR2 domain of CRAF (19). This suggests RUVBL1 may affect
PDLSC function by binding to CRAF and promoting its kinase activity
through dephosphorylation of the inhibitory S259 site, thereby
facilitating CRAF-mediated MEK/ERK activation.. However, the
present results did not reveal a regulatory association between
RUVBL1 and CRAF in PDLSCs. Despite this, RUVBL1 overexpression
independently enhanced MEK/ERK phosphorylation, indicating that it
may activate this pathway via a CRAF-independent mechanism.
Specifically, RUVBL1 has been demonstrated to exert its effects
through numerous pathways, including regulating chromatin
remodeling via its AAA+ ATPase activity to alter the
transcriptional activity of oncogenes such as CTNNB1 (25), inducing transcription-dependent
replication stress and DNA damage when expression levels are
deregulated (26), promoting cell
cycle progression through direct interaction with AHNAK2 (32). These findings indicate that RUVBL
function does not necessarily require RAF activation. Collectively,
these results indicate that RUVBL1 and CRAF function independently
yet converge on the MEK/ERK pathway to promote osteogenic
differentiation in PDLSCs. In addition, ODN MT01, a synthetic ODN
derived from human mitochondrial DNA, upregulates osteoblast
differentiation markers such as RUNX family transcription factor 2
and osteocalcin (21) and exerts
anti-inflammatory effects (33).
In the present study, ODN MT01 enhanced PDLSC osteogenesis and
partially rescued the inhibitory effects of RUVBL1 or CRAF
knockdown. These results suggested that ODN MT01 may have
potentiated RUVBL1- and CRAF-mediated activation of the MEK/ERK
pathway, thereby amplifying PDLSC osteogenic differentiation.
The present study demonstrated that the
overexpression of RUVBL1 or CRAF in PDLSCs activated the MEK/ERK
signaling pathway and promoted osteogenic differentiation, an
effect augmented by ODN MT01 (Fig.
S1). This has marked implications for the therapeutic
management of periodontal tissue. The present results warrant
further investigation, including additional replication studies and
in vivo research to assess their clinical translational
potential. Furthermore, the delivery potential of RUVBL1/CRAF
modulators based on biomaterials requires development. In addition,
studies investigating the upstream pathways regulating RUVBL1 and
CRAF as well as whether the osteogenic differentiation-promoting
effect of ODN MT01 occurs through direct interaction with PDLSCs or
other cytokine-mediated pathways are required.
In PDLSCs, RUVBL1 and CRAF independently activated
the MEK/ERK signaling pathway to promote osteogenic
differentiation, despite the absence of a direct regulatory
association between them. The present study identified RUVBL1 and
CRAF as potential molecular targets for enhancing periodontal
tissue regeneration. ODN MT01 further augmented this osteogenic
effect, suggesting its potential as an adjunctive therapeutic agent
for periodontitis. Collectively, the present study provided novel
mechanistic insight and a basis for future translational research
into PDLSC-based periodontal regeneration.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by the National Natural Science
Foundation of China (grant no. 82460460), the High-End Talent Grant
of Yunnan Province (grant no. YNWR-QNBJ-2020-272), the Special
Funding for the Cultivation of High-Level Health and Medical
Technology Talents in Yunnan Province (grant no. H-2024015) and the
Yunnan Revitalization Talent Support Program (grant no.
XDYC-QNRC-2024-449).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
XYZ conceived and designed the study, performed the
experiments, analyzed the data and wrote the manuscript. PH
performed the experiments and analyzed the data. MYQ performed the
experiments, analyzed the data and wrote the manuscript. HG
conceived and designed the experiments, performed the experiments,
analyzed the data and edited the manuscript. HG and XYZ confirm the
authenticity of all the raw data. All authors have read and
approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Caton JG, Armitage G, Berglundh T, Chapple
ILC, Jepsen S, Kornman KS, Mealey BL, Papapanou PN, Sanz M and
Tonetti MS: A new classification scheme for periodontal and
peri-implant diseases and conditions - Introduction and key changes
from the 1999 classification. J Periodontol. 89 (Suppl 1):S1–S8.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Trindade D, Carvalho R, Machado V,
Chambrone L, Mendes JJ and Botelho J: Prevalence of periodontitis
in dentate people between 2011 and 2020: A systematic review and
meta-analysis of epidemiological studies. J Clin Periodontol.
50:604–626. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ramanauskaite A, Becker K, Cafferata EA
and Schwarz F: Clinical efficacy of guided bone regeneration in
peri-implantitis defects. A network meta-analysis. Periodontol
2000. 93:236–253. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wang C, Zhao Q, Chen C, Li J, Zhang J, Qu
S, Tang H, Zeng H and Zhang Y: CD301b(+) macrophage: the new
booster for activating bone regeneration in periodontitis
treatment. Int J Oral Sci. 15:192023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wei J, Wang K, Li Y, Huang J, Deng P, Xia
X, Yang C, Xu L and Xu J: Metformin carbon dots-based osteogenic
and protein delivery system to promote bone regeneration in
periodontitis. Bioact Mater. 53:459–479. 2025.PubMed/NCBI
|
|
6
|
Zhang JJ, Li X, Tian Y, Zou JK, Gan D,
Deng DK, Jiao C, Yin Y, Tian BM, Wu RX, et al: Harnessing
mechanical stress with viscoelastic biomaterials for periodontal
ligament regeneration. Adv Sci (Weinh). 11:e23095622024. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhang S, Liu J, Feng F, Jia Y, Xu F, Wei Z
and Zhang M: Rational design of viscoelastic hydrogels for
periodontal ligament remodeling and repair. Acta Biomater.
174:69–90. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Shimono M, Ishikawa T, Ishikawa H,
Matsuzaki H, Hashimoto S, Muramatsu T, Shima K, Matsuzaka K and
Inoue T: Regulatory mechanisms of periodontal regeneration. Microsc
Res Tech. 60:491–502. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Trubiani O, Pizzicannella J, Caputi S,
Marchisio M, Mazzon E, Paganelli R, Paganelli A and Diomede F:
Periodontal ligament stem cells: current knowledge and future
perspectives. Stem Cells Dev. 28:995–1003. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang Y, Papagerakis S, Faulk D, Badylak
SF, Zhao Y, Ge L, Qin M and Papagerakis P: Extracellular matrix
membrane induces cementoblastic/osteogenic properties of human
periodontal ligament stem cells. Front Physiol. 9:9422018.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhu Y, Wang W, Chen Q, Ren T, Yang J, Li
G, Qi Y, Yuan C and Wang P: Bioprinted PDLSCs with
high-concentration GelMA hydrogels exhibit enhanced osteogenic
differentiation in vitro and promote bone regeneration in vivo.
Clin Oral Investig. 27:5153–5170. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Liu X, Zhao W, Peng Y, Liu N and Liu Q:
The relationship between MAPK signaling pathways and osteogenic
differentiation of periodontal ligament stem cells: A literature
review. PeerJ. 13:e191932025. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kim HJ and Bar-Sagi D: Modulation of
signalling by Sprouty: A developing story. Nat Rev Mol Cell Biol.
5:441–450. 2004. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhao N, Fang H, Tang YP and Liu Q:
Advancement on Correction of MAPK Signal Pathway and Chronic
Periodontitis. J Oral Sci Res. 33:1012–1015. 2017.(In Chinese).
|
|
15
|
Wu Y, Yang Y, Yang P, Gu Y, Zhao Z, Tan L,
Zhao L, Tang T and Li Y: The osteogenic differentiation of PDLSCs
is mediated through MEK/ERK and p38 MAPK signalling under hypoxia.
Arch Oral Biol. 58:1357–1368. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yang SH, Sharrocks AD and Whitmarsh AJ:
MAP kinase signalling cascades and transcriptional regulation.
Gene. 513:1–13. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Riaud M, Maxwell J, Soria-Bretones I,
Dankner M, Li M and Rose AAN: The role of CRAF in cancer
progression: from molecular mechanisms to precision therapies. Nat
Rev Cancer. 24:105–122. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jha S and Dutta A: RVB1/RVB2: Running
rings around molecular biology. Mol Cell. 34:521–533. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Guo H, Zhang XY, Peng J, Huang Y, Yang Y,
Liu Y, Guo XX, Hao Q, An S and Xu TR: RUVBL1, a novel C-RAF-binding
protein, activates the RAF/MEK/ERK pathway to promote lung cancer
tumorigenesis. Biochem Biophys Res Commun. 498:932–939. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhang Q, Qu X, Liang C, Li H, Du S, Wang
C, Xie Y, Zheng Y and Wang L: Effect of oligonucleotide MT01
delivered by N-isopropylacrylamide modified polyethyleneimine for
bone regeneration. Front Bioeng Biotechnol. 11:12045712023.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hou X, Shen Y, Zhang C, Zhang L, Qin Y, Yu
Y, Wang L and Sun X: A specific oligodeoxynucleotide promotes the
differentiation of osteoblasts via ERK and p38 MAPK pathways. Int J
Mol Sci. 13:7902–7914. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yenerall P, Das AK, Wang S, Kollipara RK,
Li LS, Villalobos P, Flaming J, Lin YF, Huffman K, Timmons BC, et
al: RUVBL1/RUVBL2 ATPase activity drives PAQosome maturation, DNA
replication and radioresistance in lung cancer. Cell Chem Biol.
27:105–121.e14. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lin D, Lin B, Bhanot H, Riou R, Abt NB,
Rajagopal J and Saladi SV: RUVBL1 is an amplified epigenetic factor
promoting proliferation and inhibiting differentiation program in
head and neck squamous cancers. Oral Oncol. 111:1049302020.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang C and Wu S: RUVBL1-modulated
chromatin remodeling alters the transcriptional activity of
oncogenic CTNNB1 in uveal melanoma. Cell Death Discov. 9:1322023.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hristova RH, Stoynov SS, Tsaneva IR and
Gospodinov AG: Deregulated levels of RUVBL1 induce
transcription-dependent replication stress. Int J Biochem Cell
Biol. 128:1058392020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Khacho M, Clark A, Svoboda DS, Azzi J,
MacLaurin JG, Meghaizel C, Sesaki H, Lagace DC, Germain M, Harper
ME, et al: Mitochondrial dynamics impacts stem cell identity and
fate decisions by regulating a nuclear transcriptional program.
Cell Stem Cell. 19:232–247. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Guida A, Simeone I, Papini D, Polvani S,
Dragoni G, Ceni E, Picariello L, Galli A and Mello T: Targeting
RuvBL1 reduces mTOR-driven NASH-HCC progression in conditional
PTEN-KO mice. Dig Liver Dis. 56:S962024. View Article : Google Scholar
|
|
29
|
Xie B, He X, Guo Y, Shen J, Yang B, Cai R,
Chen J and He Y: Cyclic tensile stress promotes osteogenic
differentiation via upregulation of Piezo1 in human dental follicle
stem cells. Hum Cell. 37:1649–1662. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Lavoie H, Gagnon J and Therrien M: ERK
signalling: A master regulator of cell behaviour, life and fate.
Nat Rev Mol Cell Biol. 21:607–632. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Bahar ME, Kim HJ and Kim DR: Targeting the
RAS/RAF/MAPK pathway for cancer therapy: From mechanism to clinical
studies. Signal Transduct Target Ther. 8:4552023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li X, Li H, Shao MM, Miao J, Fu Y and Hu
B: Downregulation of AHNAK2 inhibits cell cycle of lung
adenocarcinoma cells by interacting with RUVBL1. Thorac Cancer.
14:2093–2104. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Yu H, Shen Y, Liu Y, Gao H, Zhou Y, Hu T
and Lin C: Effect of specific sequence oligodeoxynucleotide MT01 on
the proliferation, apoptosis, and cell cycle of osteoblasts invaded
by Porphyromonas gingivalis. West China Journal of Stomatology.
33:617–621. 2015.(In Chinese). PubMed/NCBI
|