Introduction
MicroRNAs (miRs) are small, non-coding RNAs that
regulate gene expression at the transcriptional,
post-transcriptional and/or translational level. Approximately 30%
of human genes are estimated to be the targets of miRs, which play
a role in diverse biological processes such as development,
proliferation, differentiation and apoptosis (1,2).
Altered levels of the expression of miRs resulting from their
deregulation occur in a wide range of human diseases, including
cancer, where miRs act as either tumor suppressor genes or
oncogenes and are involved in the initiation, progression and
dissemination of cancer (1,2). Aberrant regulation of miRs is caused
by either genetic or epigenetic alterations, impairment in miR
processing or both (1,2). The genetic alteration of miRs is
caused mainly by chromosomal abnormalities, since more than 50% of
miR genes are located on fragile sites, although mutations occur
(1). The epigenetic cause of miR
deregulation involves silencing miR expression by CpG island
methylation and repressive histone modifications (1). Defects in the miR processing machinery
include altered levels of processing enzymes and defective
activation of pathways that facilitate the processing (1,2).
In colorectal carcinogenesis, epigenetic changes are
common phenomena in pre-cancerous and cancerous lesions, in
addition to the genomic changes that frequently occur in these
lesions (3–7). Epigenetic changes may be found in the
apparently normal colonic mucosa adjacent to cancer exhibiting
similar changes, indicating a field effect (8,9).
Recently, an altered level of the expression of miRs has been
reported in colorectal cancer (CRC) and certain miRs are expressed
at low levels in CRC due to the hypermethylation of miR genes
(10). To identify miRs with
epigenetically silenced expression in CRC, colon cancer cells, with
or without genetic deficiency for the DNA methyltransferase
enzymes, were used in combination with miR expression profiling
(11,12). Among the epigenetically silenced
miRs identified using these screening methods were miR-124a and
miR-34b/c, which showed frequent epigenetic silencing in CRC
tissues, but not in the adjacent normal mucosa, miR-124a (42/56,
75%) and miR-34b/c (101/111, 90%) (11,12).
The present study aimed to compare the frequency of
the epigenetic silencing of miRNA-124a and miRNA-34b/c in mucinous
and non-mucinous CRCs, various histological types of colorectal
polyps, uninvolved mucosa adjacent to CRC, normal mucosa from
non-cancer bearing subjects and ulcerative colitis, as well as to
compare the frequencies of miR expression in cell lines and cancer
tissues from different organs.
Materials and methods
Cell lines and primary tumors
Colorectal cancer cell lines C, C1a, Caco2, Colo201,
Colo320, H498, HCT8, HCT1116, HRT18, HT29, Lovo, LS123, LS174T,
RKO, RW2982, SW48, SW620, SW1116, SW1463, T74, VACO5, VACO6,
VACO10P, VACO411 and VACO703 were grown in DMEM, supplemented with
10% fetal bovine serum at 37°C in a 5% CO2 atmosphere.
Formalin-fixed, paraffin-embedded tissue blocks of primary CRCs,
colorectal polyps, ulcerative colitis, normal mucosa and primary
cancers from the pancreas, stomach, liver, lung, breast, kidney,
prostate and melanomas were obtained from the Department of
Pathology, University of California at San Francisco and the San
Francisco Veterans Affairs Medical Center, USA. This study was
approved by the institutional review boards of the
institutions.
5-Aza-2′-deoxycytidine (5-aza-dC)
treatment
Colo320, RKO, SW48 and VACO5 cells were grown in
6-well plates on day 0 and treated with 5 μM 5-aza-dC (Sigma, St.
Louis, MO, USA) on day 1. The medium was replaced on day 2. The
cells were grown for 2 days and harvested on day 4.
Measurement of the expression of miR-124a
and miR-34b/c
Total small RNA was isolated from the cultured cells
using a miRNeasy Mini kit (Qiagen, Valencia, CA, USA), according to
the manufacturer's instructions. cDNAs were synthesized using
specific miRNA primers (Applied Biosystems, Foster City, CA, USA),
or using hexanucleotides mix and SuperScript first-strand cDNA
synthesis kit (Invitrogen, Carlsbad, CA, USA). Gene expression was
measured by quantitative real-time PCR with Applied Biosystem 7500
Fast Sequence Detection and Taq Man assay kits (Applied
Biosystems), or by regular PCR using the primers:
5′-TGAGGGCCCCTCTGCGTGT (miR-124a, forward) and
5′-AGGCGCCTCTCTTGGCATT (miR-124a, reverse); or
5′-GTGCTCGGTTTGTAGGCAGT (miR-34b/c, forward) and
5′-GTGCCTTGTTTTGATGGCAG (miR-34b/c, reverse).
Methylation analysis
DNA obtained from the cell lines or tissues were
modified with NaHSO3 (3). The DNA methylation status of miR-124a
and miR-34b/c was determined by methylation-specific PCR (MSP). For
miR-124a, the NaHSO3-modified DNA was amplified
separately by PCR with either methylation- specific primers, i.e.,
5′-GCGAGGATTTTACGTAAGTTC (forward) and 5′-CAAAAAAACCCTCAAAACTAAAAC
GAACG (reverse), or unmethylation-specific primers, i.e.,
5′-TGGGTGAGGATTTTATGTAAGTTT (forward) and
5′-CACAAAAAAACCCTCAAAACTAAAACAAACA (reverse). For miR-34b/c, the
modified DNAs were amplified by PCR with either
methylation-specific primers, i.e, 5′-CGGTGAAATGGGGTTCGAGGC
(forward) and 5′-CCGAACACCGAACACCCGCG (reverse), or unmethylation-
specific primers, i.e., 5′-TGTTTTTTGGTGAAATGGGG TTTGAGGT (forward)
and 5′-CCTACAAACCAAACAC CAAACACCCACA (reverse).
Analysis of microsatellite instability
(MSI), CpG island methylation phenotype (CIMP), loss of
heterozygosity (LOH) and BRAF, KRAS and p53 mutations
DNA was isolated from the microdissected samples and
the analysis of MSI, CIMP, LOH and BRAF, KRAS and p53 gene
mutations was performed as previously described (13).
Statistical analysis
A comparison of the categorical variables was
performed using the Chi-square and Fisher's exact tests. The
Student's t-test was used for comparison of the mean age of the
patients. P<0.05 was considered to be statistically
significant.
Results
Correlation of the expression levels of
miR-124a and miR-34b/c with the methylation status in colon cancer
cell lines
The expression levels of miR-124a and miR-34b/c,
quantified by real-time PCR using 124a- and miR-34b/c-specific
primers, showed a positive correlation with their methylation
status (Table I). MiR-124a was
expressed at significantly lower levels in HCT116, RKO and VACO5
cells compared to Caco2 and Colo320 cells, concomitant with
methylation in HCT116, RKO and VACO5 cells, but not in Caco2 and
Colo320 cells. The low expression level of miR-34b/c also
correlated with methylation (Table
I). The cell lines with a significantly lower expression of
miR-34b/c (RKO, Colo201, Colo320 and VACO5) showed methylation,
while LS174 cells which showed higher levels of miR-34b/c
expression did not show methylation of miR-34b/c. Methylation of
these miR genes is a common event in colon cancer cells; 88%
(22/25) of colon cancer cell lines showed methylation of miR-124a
and 89% (17/19) showed methylated miR-34b/c.
 | Table IComparison of miR-124a and miR-34b/c
expression levels with methylation status. |
Table I
Comparison of miR-124a and miR-34b/c
expression levels with methylation status.
| miR-124a | miR-34b/c |
|---|
|
|
|
|---|
| Expression | Methylation | Expression | Methylation |
|---|
| Caco2 | 330.39 | − | nd | nd |
| HCT116 | 1.65 | + | nd | nd |
| RKO | 1.00 | + | 1.74 | + |
| Colo201 | nd | nd | 1.00 | + |
| Colo320 | 85.72 | − | 1.08 | + |
| LS174T | nd | nd | 15.25 | − |
| VACO5 | 1.58 | + | 1.50 | + |
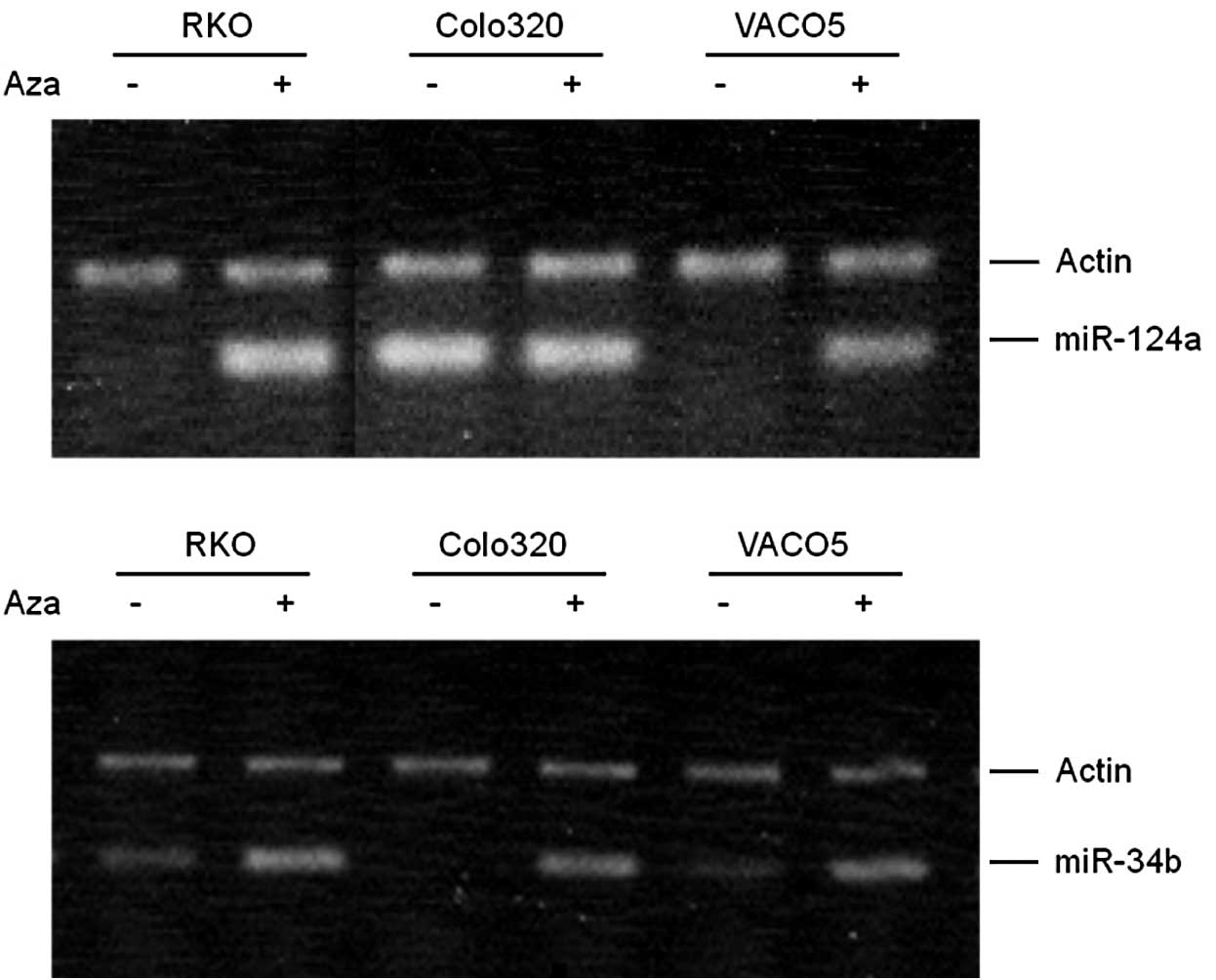
Expression of miR-124a and miR-34b was
induced by the 5-aza-dC treatment
RKO, Colo320 and VACO5 cells were treated with
5-aza-dC and the expression of miR-124a and miR-34b/c was measured
by RT-PCR (Fig. 1). RKO and VACO5
cells, which had no detectable miR-124a, showed a significant
induction of miR-124a following treatment, indicating that miR-124a
was methylated, while Colo320, which had a high-level expression of
miR-124a, did not show induction, indicating that miR-124a was
unmethylated. After the 5-aza-dC treatment, the three cell lines
with either low (RKO and VACO5) or undetectable (Colo320) levels of
miR-34b/c expression showed significant induction of miR-34b/c,
indicating that miR-34b/c is methylated at a moderate to high level
in these cells.
Methylation of miR-124a and miR-34b is
frequently present in colorectal cancers and colonic polyps
The methylation status of miR-124a and miR-34b was
determined in the primary tumors of the colorectum, stomach,
pancreas, liver, lung, breast, kidney, prostate and melanomas
(Table II). miR-124a was most
frequently methylated in CRC (80/81, 99%) and prostate cancer (4/4,
100%) compared to other cancer types whose frequency ranged from 17
to 67%. Hepatocellular carcinoma showed no methylation of miR-124a
(0/9, 0%). Methylation of miR-34b/c was also most frequently
observed in CRC (63/68, 93%), compared to other cancer types which
showed methylation frequencies ranging from 17 to 69% (Table II). Mucinous and non-mucinous CRCs
showed similar methylation frequencies of miR-124a and miR-34b/c
(Table III). The methylation of
both miR-124a and miR-34b/c were observed frequently in all
histological types of colonic polyps (121/131, 92% for miR-124a;
145/152, 95% for miR-34b/c) (Table
III). This result indicates that the methylation of miR-124a
and miR-34b/c is an early event in colorectal carcinogenesis.
 | Table IIMethylation of miR-124a and miR-34b/c
in primary tumors. |
Table II
Methylation of miR-124a and miR-34b/c
in primary tumors.
| miR-124a | miR-34b/c |
|---|
|
|
|
|---|
| Total | Methylated | % | Total | Methylated | % |
|---|
| Colorectum | 81 | 80 | 99 | 68 | 63 | 93 |
| Pancreas | 9 | 5 | 56 | 10 | 5 | 50 |
| Stomach | 15 | 10 | 67 | 41 | 31 | 69 |
| Liver | 9 | 0 | 0 | 9 | 2 | 22 |
| Lung | 9 | 3 | 33 | 7 | 2 | 29 |
| Breast | 24 | 4 | 17 | 40 | 22 | 55 |
| Kidney | 8 | 2 | 25 | 10 | 4 | 40 |
| Prostate | 4 | 4 | 100 | 5 | 3 | 60 |
| Melanoma | 5 | 3 | 60 | 6 | 1 | 17 |
 | Table IIIMethylation of miR-124a and miR-34b/c
in colorectal tissues. |
Table III
Methylation of miR-124a and miR-34b/c
in colorectal tissues.
| Colorectal
tissue | miR-124a | miR-34b/c |
|---|
|
|
|
|---|
| Total | Methylated | % | Total | Methylated | % |
|---|
| Normal mucosa | 12 | 0 | 0 | 19 | 0 | 0 |
| Ulcerative
colitis | 20 | 0 | 0 | 24 | 0 | 0 |
| Colonic polyps | 131 | 121 | 92 | 152 | 145 | 95 |
| HP | 38 | 35 | 92 | 42 | 42 | 100 |
| SSA | 25 | 24 | 96 | 27 | 26 | 96 |
| SA | 23 | 22 | 96 | 24 | 23 | 96 |
| TVA | 22 | 20 | 90 | 31 | 30 | 97 |
| TA | 23 | 20 | 87 | 28 | 24 | 86 |
| Mucosa adjacent to
cancer | 49 | 29 | 59 | 46 | 12 | 26 |
| Cancer | 81 | 80 | 99 | 68 | 63 | 93 |
| MC | 10 | 10 | 100 | 14 | 12 | 86 |
| NMC | 71 | 70 | 99 | 54 | 51 | 94 |
Methylation of miR-124a and miR-34b was
present in normal mucosa adjacent to colorectal cancers
Normal mucosa adjacent to CRC was microdissected and
the methylation status of miR-124a and miR-34b was analyzed
(Table III). Methylation of
miR-124a was observed in the adjacent mucosa in 59% of cases
(29/49), compared to 26% for miR-34b/c (12/46) (Table III). By contrast, none of the
colonic mucosa from the subjects without CRC showed methylation of
miR-124a (0/12) or of miR-34b/c (0/19). In addition, none of the
ulcerative colitis specimens showed methylation of miR-124a (0/20)
or miR-34b/c (0/24) (Table
III).
Correlation of the methylation status of
miR-124a and miR-34b/c in CRC and adjacent normal mucosa with
clinical and molecular charateristics of CRC
When the presence of miR-124a methylation in the
adjacent apparently normal colonic mucosa was compared to the
clinical and molecular characteristics of CRC, a significant
correlation was observed between the methylation of miR-124a in the
adjacent mucosa and CIMP (p<0.001) (Table IV). A borderline correlation was
observed between miR-124a methylation and proximal tumor location
(p<0.081), MSI (p<0.070) and BRAF mutations (p<0.069). By
contrast, the methylation of miR-34b/c in the adjacent colonic
mucosa was significantly correlated with patient age (p<0.004),
but no correlation was detected with other clinical and molecular
characteristics of CRC (Table
IV).
 | Table IVCorrelation of methylation status of
miR-124a and miR-34b/c in colorectal cancer and adjacent normal
mucosa with clinical and molecular characteristics in cancer. |
Table IV
Correlation of methylation status of
miR-124a and miR-34b/c in colorectal cancer and adjacent normal
mucosa with clinical and molecular characteristics in cancer.
| miR-124a
(adjacent/cancer) | miR-34b/c
(adjacent/cancer) |
|---|
|
|
|
|---|
| +/+ | −/+ | p-value | +/+ | −/+ | p-value |
|---|
| Total | 29 | 20 | | 12 | 34 | |
| Age | | | ns | | | 0.004 |
| Mean | 69.0 | 68.2 | | 76.8 | 66.4 | |
| SD | 12.1 | 15.2 | | 7.7 | 14.6 | |
| Gender | | | ns | | | ns |
| Male | 14 (54%) | 12 (46%) | | 7 (29%) | 17 (71%) | |
| Female | 14 (64%) | 8 (36%) | | 5 (24%) | 16 (76%) | |
| Stage | | | ns | | | ns |
| A/B | 17 (57%) | 13 (43%) | | 6 (21%) | 23 (79%) | |
| C/D | 12 (63%) | 7 (37%) | | 6 (35%) | 11 (65%) | |
| Tumor side | | | 0.081 | | | ns |
| Proximal | 15 (75%) | 5 (25%) | | 5 (26%) | 14 (74%) | |
| Distal | 14 (48%) | 15 (52%) | | 7(26%) | 20 (74%) | |
| Survival (5
years) | | | ns | | | ns |
| − | 16 (70%) | 7 (30%) | | 7 (32%) | 15 (68%) | |
| + | 13 (50%) | 13 (50%) | | 5 (21%) | 19 (79%) | |
| MSI | | | 0.070 | | | ns |
| + | 5 (100%) | 0 (0%) | | 0 (0%) | 5 (100%) | |
| − | 24 (55%) | 20 (45%) | | 12 (29%) | 29 (71%) | |
| CIMP (≥3/6) | | | <0.001 | | ns | |
| + | 13 (100%) | 0 (0%) | | 3 (23%) | 10 (77%) | |
| − | 16 (44%) | 20 (56%) | | 9 (27%) | 24 (73%) | |
| LOH (≥1/4) | | | ns | | | ns |
| + | 21 (55%) | 17(45%) | | 11 (30%) | 26 (70%) | |
| − | 8 (73%) | 3(27%) | | 1 (11%) | 8 (89%) | |
| BRAF mutations | | | 0.069 | | | ns |
| + | 6 (100%) | 0 (0%) | | 1 (14%) | 6 (86%) | |
| − | 23 (53%) | 20 (47%) | | 11 (28%) | 28 (72%) | |
| KRAS mutations | | | ns | | | ns |
| + | 7 (54%) | 6 (46%) | | 3 (21%) | 11 (79%) | |
| − | 22 (61%) | 14 (39%) | | 9 (28%) | 23 (72%) | |
| p53 mutations | | | ns | | | ns |
| + | 16 (57%) | 12 (43%) | | 6 (24%) | 19 (76%) | |
| − | 13 (62%) | 8 (38%) | | 6 (29%) | 15 (71%) | |
Discussion
The present study showed that miR-124a and miR-34b/c
are most frequently methylated in CRC compared to eight other tumor
types, indicating a relative colon tissue type-specific expression
of these miRs. We observed that the two miRs are frequently
methylated in colon cancer cell lines (miR-124a, 22/25, 88% and
miR-34b/c, 17/19, 89%) and that methylation is correlated with the
low-level expression of these miRs. Lujambio et al have
reported that HCT116 colon cancer cells that express low levels of
miR-124a, showed a 2- to 3-fold up-regulation of miR-124a
expression, following treatment with the demethylating agent
5-aza-dC, or when made genetically deficient for the DNA
methylatransferase enzymes (11).
The data indicate that DNA methylation-associated repression of
miR-124a occurs frequently in colon cancer cells. A similar result
was observed with miR-34b/c in HCT116 cells (12). Epigenetic loss of miR-124a
expression was found to be associated with the activation of an
oncogene, cyclin D kinase 6, and the phosphorylation of the
retinoblastoma tumor suppressor gene, and that miR-34b/c are
targets of p53, indicating the functional significance of the
epigenetic regulation of these genes (11,12,14).
We observed that RKO and VACO5 colon cancer cells exhibited no
detectable levels of miR-124a, and that RKO, VACO5 and Colo320
colon cancer cells exhibited low or undetectable levels of
miR-34b/c, which showed a significant induction of the respective
miRs, following 5-aza-dC treatment, indicating that the epigenetic
silencing of these miRs occurs in colon cancer cells. These
observations are consistent with previous reports (10–12).
No detectable methylation of miR-124a or miR-34b/c
was noted in the normal colonic tissues from the subjects without
cancer, or in the colonic mucosa of patients with ulcerative
colitis. However, frequent methylation of miR-124a (59%) and
miR-34b/c (26%) was noted in the apparently normal mucosa adjacent
to the CRCs with methylation of these genes, indicating a field
effect or field cancerization (8,15,16).
We have recently reported that epigenetic silencing of oncostatin M
receptor (OSMR) occurs in 55% of apparently normal mucosa adjacent
to CRCs (9). Together, these data
indicate that epigenetic regulation of OSMR, miR-124a and miR-34b/c
may function as mediators of the field effect or field
cancerization. We previously reported that different histological
types of colonic polyps (serrated vs. tubular adenoma) and colon
cancer (mucinous vs. non-mucinous) show different frequencies and
patterns of CpG island methylation of various genes (17,18).
However, in the present study, high frequencies of methylation of
miR-124a and miR-34b/c were observed in all histological types of
colonic polyps and CRC. Together with the data regarding adjacent
normal mucosa, our data on colonic polyps indicate that methylation
of miR-124a and miR-34b/c occurs early in multi-step colorectal
carcinogenesis.
The genetic instability pathways, chromosomal
instability (CIN) and MSI, as well as additional pathways involving
epigenetic abnormalities, are known to be involved in colorectal
carcinogenesis (4–6,19).
CRCs with CIN are characterized by frequent LOH, KRAS and p53
mutations, whereas CRCs with MSI frequently show a high degree of
concurrent promoter methylation in multiple genes, i.e., CIMP, and
BRAF mutations (5–7,19).
CRCs with MSI show different clinicopathological characteristics
from CIN tumors, such as a younger age at diagnosis, proximal
location, presence of tumor-infiltrating lymphocytes and mucinous
histology (5–7,19). The
potential relationship of the methylation status of miR-124a and
miR-34b/c of CRCs and apparently normal adjacent mucosa to these
molecular and clinicopathological characteristics was examined. We
noted a significant correlation of the methylation of miR-124a in
the apparently adjacent normal mucosa to CIMP, and a borderline
correlation with proximal tumor location, MSI and BRAF mutations,
molecular characteristics frequently found in CRCs with MSI. By
contrast, the methylation of miR-34b/c showed a significant
correlation only with older age at diagnosis. These findings are
significant since promoter methylation in CRCs was broadly
classified into cancer- and age-associated types (5,6,20).
In conclusion, we confirmed the frequent occurrence
of epigenetic silencing of miR-124a and miR-34b/c in CRCs. We also
noted that the methylation of the two miRs occurs frequently in
apparently normal adjacent colonic mucosa, indicating a field
effect or field cancerization; that mucinous and non-mucinuous CRCs
and all histological types of colonic polyps show frequent
methylation of the two miRs, indicating that methylation of these
miRs is an early event in all pathways of colorectal
carcinogenesis; and finally, that the methylation of miR-124a in
the apparently adjacent normal colonic mucosa is associated with
MSI of CRCs, while the methylation of miR-34b/c is associated with
older age at diagnosis, underscoring the importance of the
relationship between promoter methylation, cancer and aging.
Acknowledgements
This study was supported in part by the Department
of Veterans Affairs Medical Research Service, the Oberkotter
Foundation grant and the Theodora Betz Foundation grant.
References
|
1
|
Croce CM: Causes and consequences of
microRNA dysregulation in cancer. Nat Rev Genet. 10:704–714. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Davalos V and Esteller M: MicroRNAs and
cancer epigenetics: a macrorevolution. Curr Opin Oncol. 22:35–45.
2010. View Article : Google Scholar
|
|
3
|
Vogelstein B and Kinzler KW: Cancer genes
and the pathways they control. Nat Med. 10:789–799. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Jones PA and Baylin SB: The epigenomics of
cancer. Cell. 128:683–692. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Issa JP: CpG island methylator phenotype
in cancer. Nat Rev Cancer. 4:988–993. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kim YS and Deng G: Epigenetic changes
(aberrant DNA methylation) in colorectal neoplasia. Gut Liver.
1:12–21. 2007.PubMed/NCBI
|
|
7
|
Deng G, Nguyen A, Tanaka H, et al:
Regional hypermethylation and global hypomethylation are associated
with altered chromatin conformation and histone acetylation in
colorectal cancer. Int J Cancer. 118:2999–3005. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Shen L, Kondo Y, Rosner GL, et al: MGMT
promoter methylation and field defect in sporadic colorectal
cancer. J Natl Cancer Inst. 97:1330–1338. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Deng G, Karkar S, Okudaira K, Choi E,
Sleisenger MH and Kim YS: Unique methylation pattern of oncostatin
M receptor gene in cancer of colorectum and other digestive organs.
Clin Cancer Res. 15:1519–1526. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Faber C, Kirchner T and Hlubeck F: The
impact of microRNAs in colorectal cancer. Virchows Arch.
454:359–367. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lujambio A, Ropero S, Ballestar E, et al:
Genetic unmasking of an epigenetically silenced microRNA in human
cancer cells. Cancer Res. 67:1424–1429. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Toyota M, Suzuki H, Sasakin Y, et al:
Epigenetic silencing of microRNA-34b/c and B-cell translocation
gene 4 is associated with CpG island methylation in colorectal
cancer. Cancer Res. 68:4123–4132. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Deng G, Kakar S, Tanaka H, Matsuzaki K,
Miura S, Sleisenger MH and Kim YS: Proximal and distal colorectal
cancers show distinct gene-specific methylation profile and
clinical and molecular characteristics. Eur J Cancer. 44:1290–1301.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
He L, He X, Lowe SW and Hannon GJ:
microRNAs join the p53 network – another piece in the tumor
suppressor puzzle. Nat Rev Cancer. 7:819–822. 2007.
|
|
15
|
Braakhuis BJ, Tabor MP, Kummer JA, Leemans
CR and Brakenhoff RH: A genetic explanation of Slaughter's concept
of field cancerization: evidence of clinical implication. Cancer
Res. 63:1727–1730. 2003.
|
|
16
|
Leedham SJ, Graham TA, Oukuf D, et al:
Clonality, founder mutations, and field cancerization in human
ulcerative colitis-associated neoplasia. Gastroenterology.
136:542–550. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kim YH, Kakar S, Cun L, Deng G and Kim YS:
Distinct CpG island methylation profile and BRAF mutation status in
serrated and adenomatous colorectal polyps. Int J Cancer.
123:2587–2593. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tanaka H, Deng G, Matsuzaki K, et al: BRAF
mutation, CpG island methylator phenotype and microsatellite
instability occur more frequently and concordantly in mucinous than
non-mucinous colorectal cancer. Int J Cancer. 118:2765–2771. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wong JJL, Hawkins NJ and Ward RL:
Colorectal cancer: a model for epigenetic tumorigenesis. Gut.
56:140–148. 2007. View Article : Google Scholar
|
|
20
|
Ahuja N, Li Q, Mohan AL, Baylin SB and
Issa JP: Aging and DNA methylation in colorectal mucosa and cancer.
Cancer Res. 58:5489–5494. 1998.PubMed/NCBI
|