Introduction
Previous studies have demonstrated that the
overexpression of basic fibroblast growth factor (bFGF) and
epithelial growth factor (EGF), and their receptors, epithelial
growth factor receptor (EGFR) and bFGF receptor (bFGFR), occurs
frequently in high-grade gliomas and glioblastomas (1–3). These
studies indicated the existence of autocrine and/or paracrine loops
of EGF-EGFR and FGF-FGFR in the glioblastoma microenvironment.
Glioblastomas are the most malignant neuroepithelial tumors of the
brain, and typically express glial fibrillary acidic protein
(GFAP).
An increasing number of studies have indicated that
glioblastomas develop from multipotent cancer stem cells, which are
a small subset of tumor cells possessing the capacity of
self-renewal and the ability to proliferate into heterogeneous
lineages of cancer cells under specific environmental stimuli
(4–6). It has been observed that EGF and bFGF
are critical components of the culture medium for cancer stem cells
and human embryonic stem cells, and are necessary for maintaining
these stem cells in an undifferentiated state (7–9). These
findings indicate that EGF/EGFR and FGF/FGFR may be responsible for
a microenvironment that maintains glioblastomas and
glioblastoma-derived stem cells in an undifferentiated state.
In the present study, the functions of EGF and bFGF
were inhibited using small interfering RNA (siRNA) to interfere
with their receptors, and the expression of the markers for stem
cells [cluster of differentiation (CD)133], astrocytes (GFAP),
neurons [tubulin-β3 (TUBB4)] and oligodendrocytes [myelin basic
protein (MBP)] were analyzed for the first time in U251
glioblastoma cells. The results provide valuable data on the
effects of EGF and bFGF in glioma-derived cancer stem cells.
Materials and methods
Cell culture
The human glioblastoma U251 cell line and the human
embryonic kidney 293 cell line (HEK 293) (ATCC, Manassas, VA, USA)
were used in all experiments and were maintained in Dulbecco’s
modified Eagle’s medium (DMEM; Hyclone) supplemented with 10% fetal
bovine serum (Hyclone), 1% penicillin-streptomycin solution and 1%
GlutaMAX (Sigma, St. Louis, MO, USA). The cells were cultured under
standard conditions (humidified atmosphere of 5% CO2 in
air at 37°C).
ELISA
Conditioned medium from the U251 cells was collected
for the ELISA following incubation for 1, 3 or 5 days.
Concentrations of EGF and bFGF in the cell culture supernatant were
determined using a Quantikine Human EGF Immunoassay kit (Shanghai
Weiao Biotech Ltd., Shanghai, China) and a bFGF Immunoassay kit
(Shanghai Weiao Biotech Ltd.), respectively, according to the
manufacturer’s instructions.
Preparation of siRNA and construction and
identification of enhanced green fluorescent protein (EGFP)-labeled
lentiviral expression vector
Four siRNA duplexes targeting human EGFR (GenBank
accession no. NM_201283.1) and bFGFR (GenBank accession no.
NM_023107.2) were used. All siRNA duplexes were designed and
synthesized by Shanghai R&S Biotechnology Co. Ltd. (Shanghai,
China), using 2′-bis(acetoxyethoxy)-methyl ether (ACE) protection
chemistry.
The siRNA duplexes were prepared by annealing the
complementary oligos and were ligated into the expression vector
pcDNA6.2-EGFP (Invitrogen, Carlsbad, CA, USA) to form
pcDNA6.2-EGFP-EGFR and pcDNA6.2-EGFP-bFGFR, respectively. The
products were used to transform Escherichia coli DH5α cells.
Following overnight culture of the cells with 50 mg/l
spectinomycin, pcDNA6.2-EGFP-EGFR and pcDNA6.2-EGFP-bFGFR were
extracted and the sequences of the EGFR and bFGFR siRNA were
confirmed by sequencing. The siRNA sequences were screened based on
their interference efficiency in HEK 293 cells, which was evaluated
by fluorescence microscopy and reverse transcription polymerase
chain reaction (RT-PCR). The most efficient duplex was selected for
subsequent studies. The target sequences of the siRNAs were
5′-AAGTGTGTAACGGAATAGGTA-3′ for EGFR and
5′-GCAACGTGGAGTTCATGTGTA-3′ for bFGFR.
To construct the lentiviral expression vector,
pcDNA6.2-EGFP-EGFR and pcDNA6.2-EGFP-bFGFR were amplified by PCR
using a primer pair with Asc1 and Pme1 sequences
(5′-TACTGGCGCGCCGCCACCATGGTGAG CAAGGGCGAGGA-3′ and
5′-ACTAGTTTAAACTGCGGC CAGATCTGGGC-3′). Subsequent to restriction
endonuclease digestion with Asc1 and Pme1, the
product was ligated into the lentiviral expression vector,
pLenti6.3-MCS/V5 DEST (Invitrogen), which was used to transform
E. coli DH5α cells. Positive recombinant colonies (named
pLenti6.3-EGFP-EGFR-miR and pLenti6.3-EGFP-bFGFR-miR) were screened
and identified by PCR, Asc1 and Pme1 digestion and
DNA sequencing. To produce lentiviral particles, the positive
recombinant plasmid DNA was extracted from the E. coli DH5α
cells using a plasmid extraction kit (Axygen Biotechnology Ltd.,
Hangzhou, China) and mixed with 6 μg pLenti6.3-MCS/V5 DEST and 6 μg
plasmid packaging mix for co-transfection of the HEK 293 cells
using POLOdeliverer 3000 Transfection Reagent (Shanghai R&S),
according to the manufacturer’s instructions. Following incubation
for 48–72 h, the lentiviral particle supernatant was collected for
the next experiment. Packaging efficiency was assessed using
fluorescence microscopy.
Transfection of siRNAs
Exponentially growing U251 cells were plated at a
density of 1×105 cells/dish and incubated in 5%
CO2 at 37°C. At 80% confluence, the cells were divided
into five groups [RNA interference (RNAi)-EGFR, RNAi-bFGFR,
RNAi-EGFR + bFGFR, RNAi-negative control (NC) and control] and
transfected with Lenti-EGFP-EGFR-microRNA (miR),
Lenti-EGFP-bFGFR-miR, Lenti-EGFP-EGFR-miR plus
Lenti-EGFP-bFGFR-miR, Lenti-GFP and blank DMEM, respectively
(multiplicity of infection, MOI = 10). To intensify infection,
polybrene (4–8 μg/ml final concentration) was also used. Subsequent
to 24 h of incubation, the medium was replaced with fresh DMEM and
the U251 cells were incubated for another 48 h. Infection
efficiency was assessed by fluorescence microscopy based on the
fluorescence of the reporter gene, GFP.
RNA isolation and cDNA synthesis
Total RNA was extracted from the cultured cells with
TRIzol reagent (Invitrogen). cDNA was synthesized using Moloney’s
murine leukemia virus (M-MLV; Fermentas, Waltham, MA, USA) in a
20-μl reaction volume containing 1 μg total RNA, 1 μl random primer
(0.5 μg/μl), 1 μl oligo(dT) primer (0.5 μg/μl) and RNase-free
water. The reaction was incubated at 65°C for 10 min and quickly
chilled on ice. Next, 4 μl 5X reaction buffer, 0.5 μl RiboLock
ribonuclease inhibitor (20 U/μl), 2 μl 10 mM dNTP mix (Majorbio
Bio-Pharm Technology Co., Ltd., Shanghai, China) and 1 μl of
ReverTra Ace (Fermentas) was added. The mixture was incubated at
37°C for 60 min. The process was stopped by heating the solution to
70°C for 10 min and then quickly chilling it on ice.
Quantitative (q)PCR
EGFR, bFGFR, CD133, GFAP, TUBB4 and MBP mRNA was
detected by qPCR using the primers listed in Table I. PCR amplification was performed in
a 20-μl volume that contained 1 μl reverse transcriptase
(Fermentas), 10 μl 2X SYBR Green mix (Invitrogen), 8 μl 10X buffer
and 1 μl primers. PCR was performed using an Eppendorf Realplex
(Eppendorff, Hamberg, Germany) with initial denaturation at 95°C
for 2 min, followed by 40 cycles of 95°C for 15 sec, 59°C for 15
sec and 72°C for 20 sec. The dissociation of the reaction products
was conducted from 60 to 95°C for 30 sec with 39 cycles.
Fluorescence data were converted into cycle threshold (CT)
measurements. The CT of each sample was averaged and analyzed by a
comparative CT method. The expression level of a specific gene was
determined quantitatively by calculating the ratio relative to an
internal standard [human β-actin (hACTB)] as 2−ΔΔT.
 | Table ISequences of PCR primers used in the
study. |
Table I
Sequences of PCR primers used in the
study.
| Gene name | Forward primer | Reverse primer | Size, bp |
|---|
| EGFR |
TGACTGAGGACAGCATAGACGA |
GGGCTGGACAGTGTTGAGATAC | 203 |
| FGFR |
GGCTGCCAAGACAGTGAAGTT |
GGTTGATGCTGCCGTACTCAT | 215 |
| GFAP |
GCACGCAGTATGAGGCAATG |
ACTCCAGGTCGCAGGTCAAG | 177 |
| TUBB3 |
CCAAGGGTCACTACACGGAG |
ATGATGCGGTCGGGATACTC | 187 |
| CD133 |
CCGCAGGAGTGAATCTTTTATC |
CCATTCCCTGTGCGTTGA | 199 |
| MBP2 |
CCGAGAAGGCCAGTACGA |
GTGAAAGTTCACCCAGGTTTCT | 74 |
| hACTB |
TCCTTCCTGGGCATGGAGT |
CAGGAGGAGCAATGATCTTGAT | 208 |
Statistical analysis
Statistical analyses were performed using SPSS
software (version 11.5.0 for Windows; SPSS, Inc., Chicago, IL,
USA). Statistical significance was evaluated with Student’s t-test,
using data from at least three independent experiments. Values of
P<0.05 were considered to indicate a statistically significant
difference.
Results
Confirmation of recombinant lentiviral
expression vectors
Following transformation with the lentiviral
expression plasmids, pLenti6.3-EGFP-EGFR-miR and
pLenti6.3-EGFP-bFGFR-miR, the E. coli DH5α cells were grown
in oscillating culture and spread onto ampicillin-lysogeny broth
plates. Single clones were selected for PCR amplification. Agarose
gel electrophoresis of the PCR products showed the expected 911-bp
insert, and digestion with Asc1 and Pme1 yielded the
appropriate fragments (Fig. 1).
Thus, the recombinant lentiviral expression vectors were
constructed successfully.
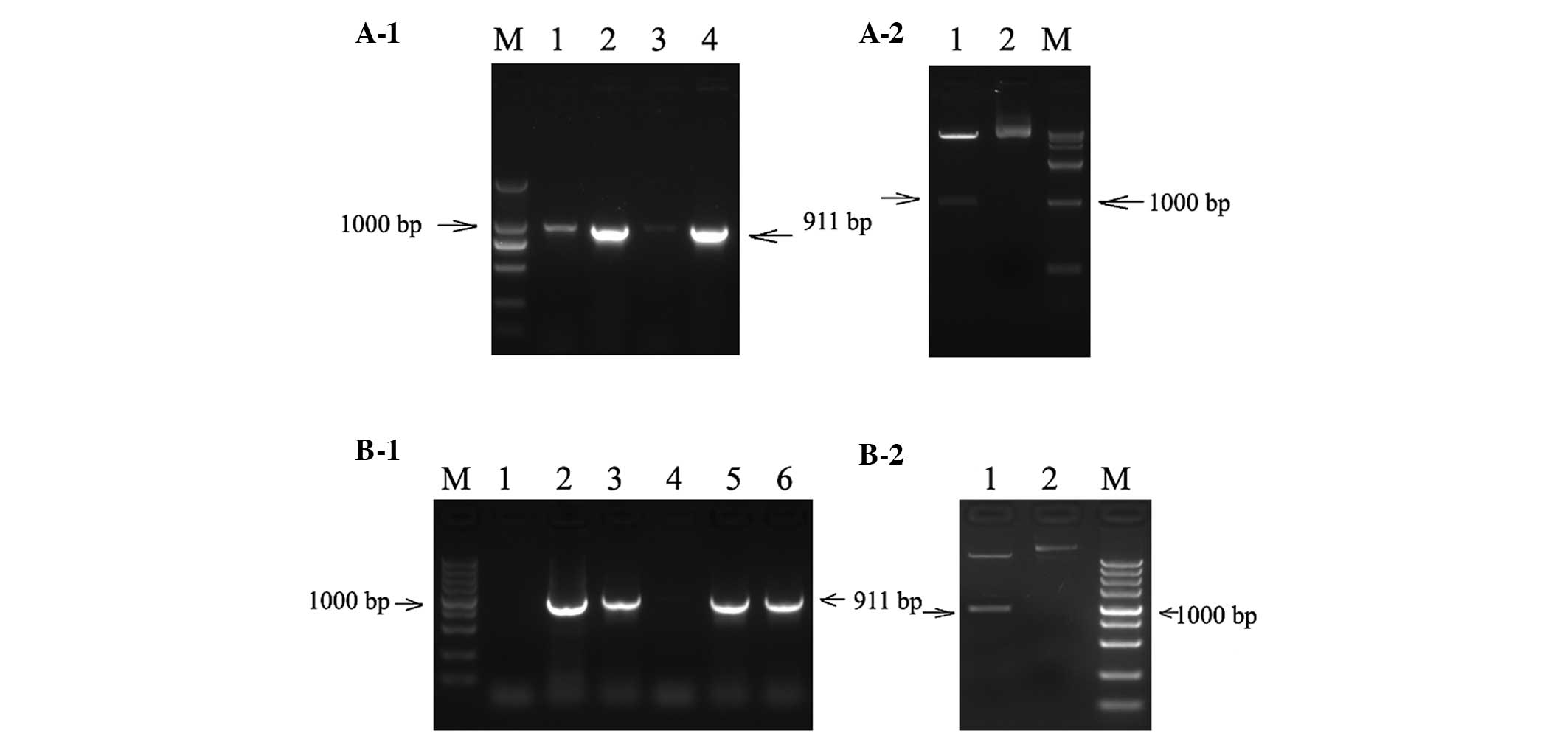 | Figure 1Identification of EGFP-labeled
lentiviral expression vector. (A-1) Gel electrophoresis of the PCR
product of pLenti6.3-EGFP-EGFR-miR. M, marker DL2000; lane 1,
negative control; lanes 2–4, PCR product of recombinant vector (911
bp). (A-2) Restriction map of the PCR positive vector of
pLenti6.3-EGFP-EGFR-miR. M, marker DL15000; lane 1, digestion
product of double restriction endonuclease of AscI and
PmeI; lane 2, the recombinant plasmid
pLenti6.3-EGFR-EGFR-miR. (B-1) Gel electrophoresis of the PCR
product of pLenti6.3-EGFP-FGFR-miR. M, marker DL5000; lane 1,
negative control; lanes 2–6, PCR product of recombinant vector (911
bp). (B-2) Restriction map of the PCR positive vector of
pLenti6.3-EGFP-FGFR-miR. M, marker DL5000; lane 1, digestion
product of double restriction endonuclease of AscI and PmeI; lane
2, the recombinant plasmid pLenti6.3-EGFP-FGFR-miR. EGFP, enhanced
green fluorescent protein; EGFR, epithelial growth factor receptor;
miR, microRNA; PCR, polymerase chain reaction; FGFR, fibroblast
growth factor receptor. |
Selected EGFR and FGFR siRNA sequences markedly
decreased target expression in the U251 cells. Target mRNA
expression in the U251 cells was assayed 72 h after transfection
with the lentiviral particles carrying the siRNA sequences
(Lenti-EGFP-EGFR-miR and/or Lenti-EGFP-bFGFR-miR). Transfection was
confirmed by the expression of EGFP in the U251 cells, as assessed
by fluorescence microscopy (Fig.
2). No green fluorescence was observed in the non-transfected
U251 control cells. The expression levels of EGFR and bFGFR mRNA
was analyzed by qPCR. The mRNA expression of EGFR and bFGFR was
significantly reduced compared with the expression in the control
cells (P<0.01). The siRNA sequences targeting EGFR and bFGFR
effectively knocked down the mRNA levels of these receptors by at
least 80% (Fig. 3).
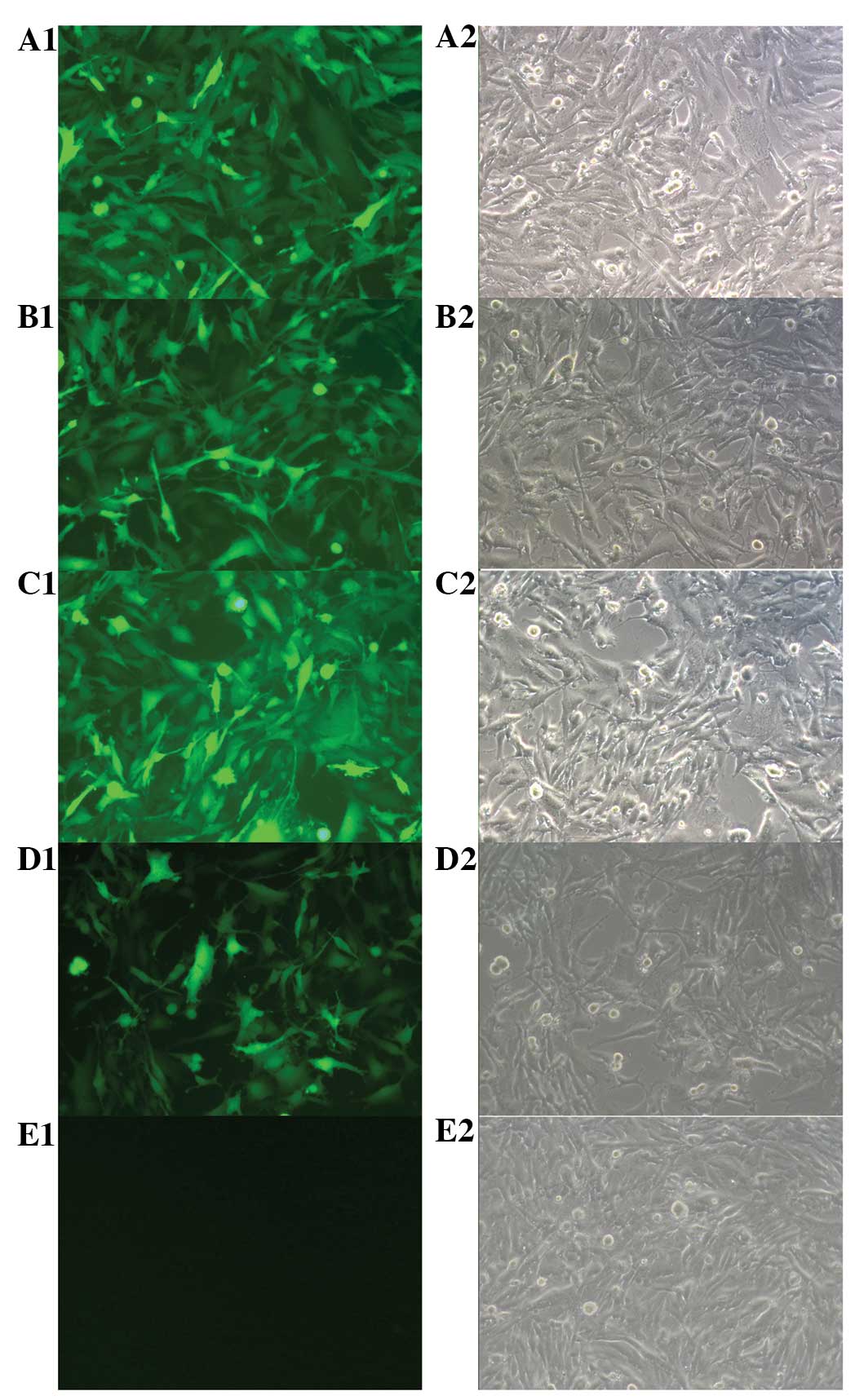 | Figure 2Fluorescence microscope map of U251
cells. U251 cells display green fluorescence after transfection
with (A1) Lenti-EGFP-EGFR-miR, (B1) Lenti-EGFP-bFGFR-miR, (C1)
Lenti-EGFP-EGFR-miR+Lenti-EGFP-bFGFR-miR, or (D1) Lenti-EGFP at 60
h. (E1) Blank control (A2, B2, C2, D2 and E2): phase contrast
microscope (Original magnification, ×400). EGFP, enhanced green
fluorescent protein; EGFR, epithelial growth factor receptor; miR,
microRNA; bFGFR, basic fibroblast growth factor receptor. |
U251 cells express EGF/EGFR and
bFGF/bFGFR
The expression of EGF/bFGF and EGFR/bFGFR in the
U251 cells was examined. qPCR showed that the EGFR and bFGFR mRNA
was expressed in the U251 cells (Fig.
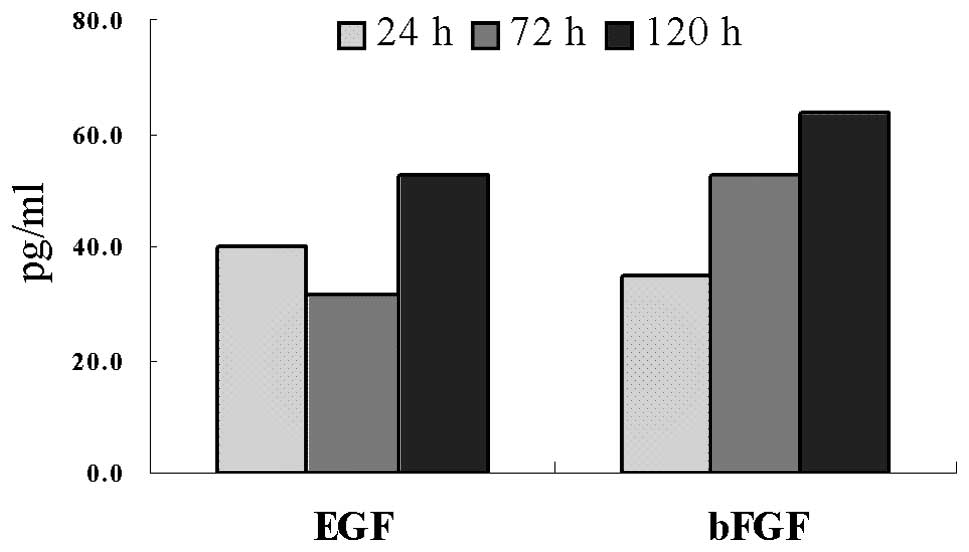
3). To determine the expression of EGF and bFGF, ELISAs of the
U251 cell culture supernatants were performed. The levels of EGF
and bFGF in the supernatants increased with the culture time. At 5
days, the concentrations of EGF and bFGF were 52.2 and 63.4 pg/ml,
respectively (Fig. 4).
CD133, GFAP, MBP and TUBB3 mRNA
expression in U251 cells with EGFR and bFGFR gene knockdown
To evaluate mRNA expression levels of the genes of
interest, qPCR was performed once the U251 cells had been
transfected with Lenti-EGFP-EGFR-miR, Lenti-EGFP-FGFR-miR or both.
The results are shown in Fig. 5.
Non-transfected U251 control cells expressed CD133, GFAP, MBP2 and
TUBB3 mRNA. After silencing EGFR and/or bFGFR expression, the mRNA
expression levels of CD133, GFAP, MBP and TUBB3 were altered. CD133
mRNA expression was significantly reduced in the RNAi-EGFR,
RNAi-bFGFR and RNAi-EGFR + RNAi-FGFR groups compared with that in
the control (RNAi-NC) group (P<0.05), with particularly marked
downregulation in the RNAi-EGFR + RNAi-FGFR group. The mRNA
expression levels of TUBB3 and GFAP in the U251 cells were
significantly increased in the RNAi-EGFR, RNAi-bFGFR and RNAi-EGFR
+ RNAi-FGFR groups compared with those in the RNAi-NC group
(P<0.05). However, the mRNA expression level of CD133, GFAP and
TUBB3 did not differ significantly among the RNAi-EGFR, RNAi-bFGFR
and RNAi-EGFR + RNAi-FGFR groups. The expression of the MBP mRNA
was decreased in the RNAi-EGFR and RNAi-bFGFR groups, but was
increased in the RNAi-EGFR + RNAi-FGFR group; however, the MBP mRNA
level in the RNAi-EGFR + RNAi-FGFR group was not significantly
different from that in the RNAi-NC group.
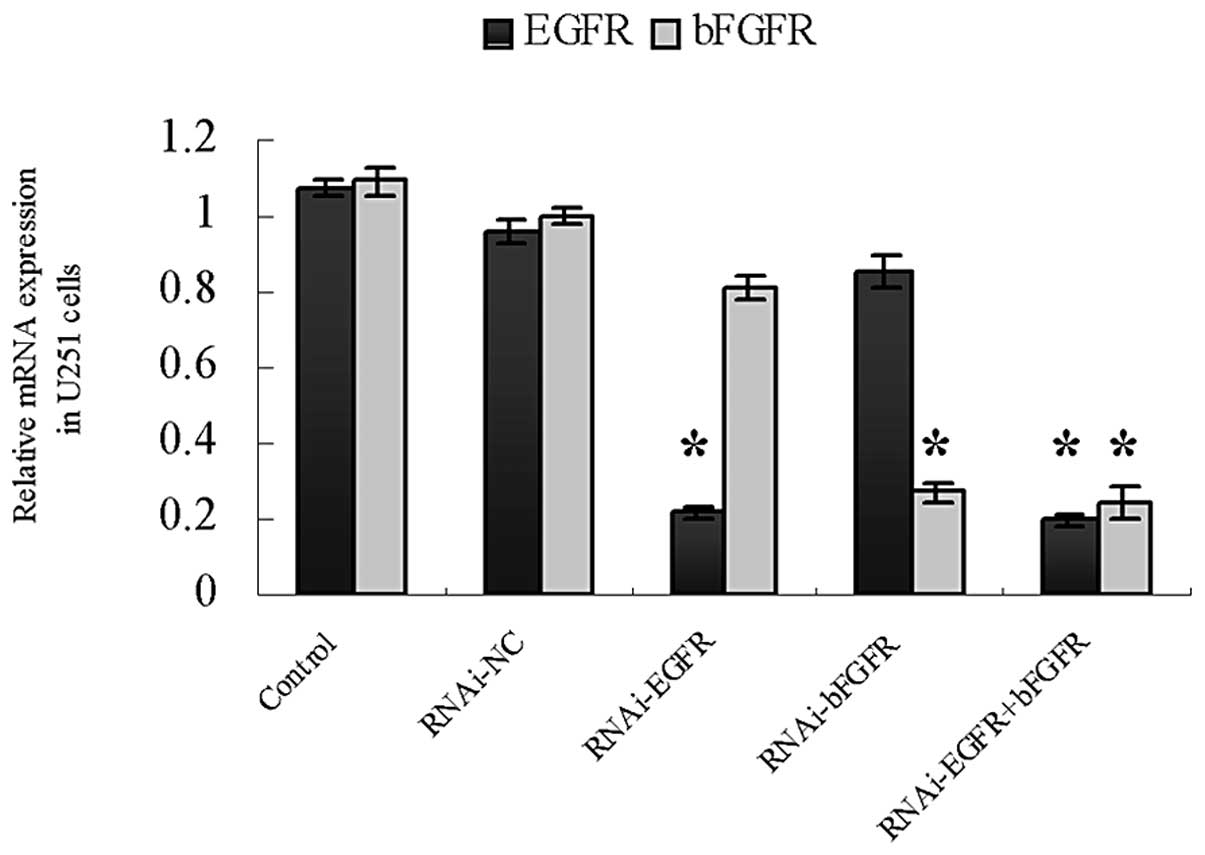
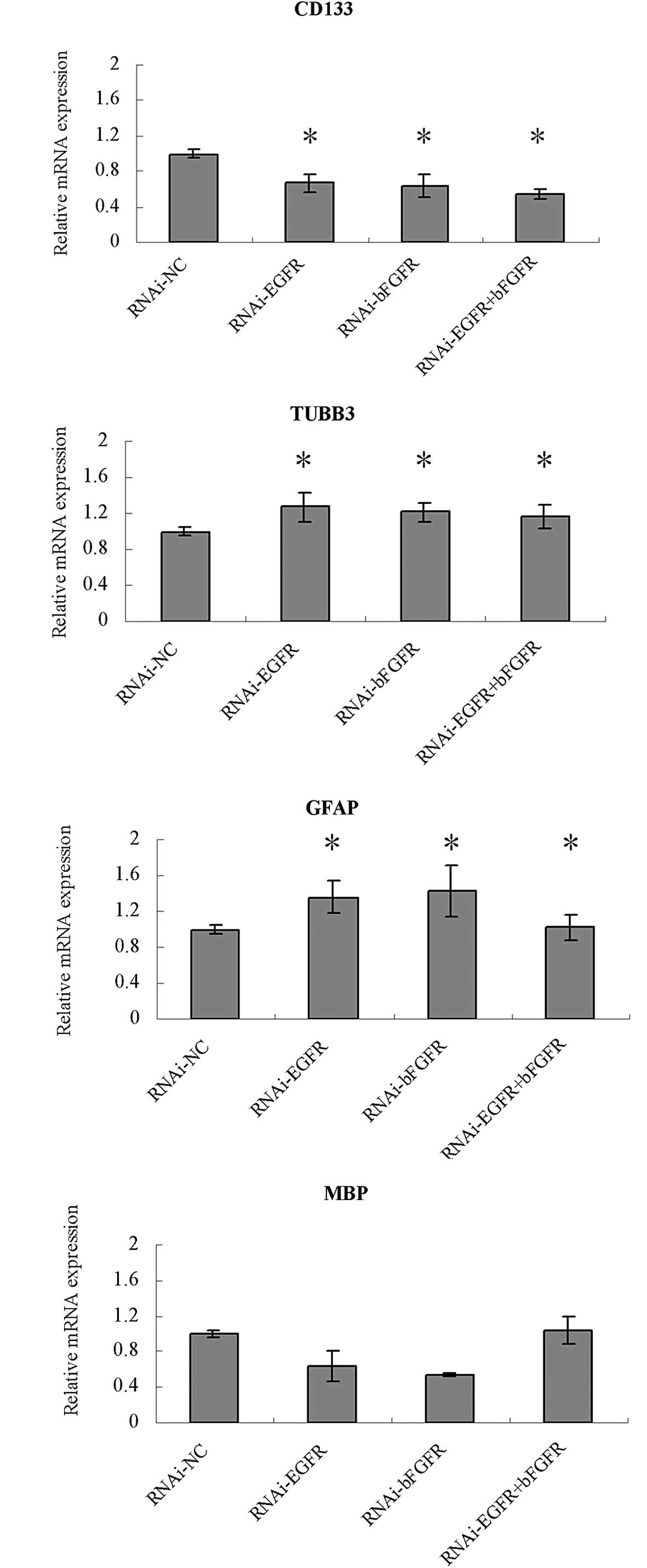 | Figure 5Relative mRNA expression of CD133,
TUBB3, GFAP and MBP in U251 cells measured by qPCR. U251 cells were
transfected with Lenti-EGFP-EGFR-miR or Lenti-EGFP-FGFR-miR, or
both Lenti-EGFP-EGFR-miR and Lenti-EGFP-FGFR-miR. U251 cells
transfected with Lenti-EGFP were set up as control groups.
*P<0.05. CD133, cluster of differentiation 133;
TUBB3, tubulin-β3; GFAP, glial fibrillary acidic protein; MBP,
myeline basic protein; EGFP, enhanced green fluorescent protein;
EGFR, epithelial growth factor receptor; miR, microRNA; bFGFR,
basic fibroblast growth factor receptor; RNAi, RNA interference;
NC, negative control; qPCR, quantitative polymerase chain
reaction. |
Discussion
Glioblastoma, the most common and malignant
astrocytoma, is believed to have a glial cell origin. GFAP, a
specific marker for immature or neoplastic astrocytes, is usually
expressed in glioblastoma cells (10). The present study assessed the
expression of a stem cell marker (CD133), an astrocyte marker
(GFAP), a neuronal marker (TUBB4) and an oligodendrocyte marker
(MBP) using qPCR. The U251 cells expressed CD133 mRNA, which is
consistent with our previous observations using
immunohistochemistry and PCR in different WHO-grade astrocytomas
(11). Co-expression of the
neuronal (TUBB4) and oligodendrocytic (MBP) markers was also
observed in the U251 cells. These results indicate that
multilineage phenotypes exist in U251 cells. Stem cell markers and
multilineage phenotypes have also been described in glioblastomas
(12). The expression of neuronal,
astrocytic and oligodendroglial lineage markers indicates that U251
glioblastoma cells are multipotent or plastic cells. This
multilineage phenotype may be a consequence of the differentiation
of glioblastoma cells under environmental regulation (13).
Determining the differentiation mechanism(s) of
glioblastoma cells may help in developing opportunities for
interfering with the glioblastoma phenotype, which may be useful in
treating glioblastomas (14).
Various autocrine factors found in adult tissue and malignant
tumors provide growth signals. Previous studies have demonstrated
the overexpression of EGFR and FGFR, and their co-expression with
their ligands occurs frequently in gliomas (1–3). These
studies indicate that EGFR and FGFR are candidate receptors for
autocrine growth factors in gliomas.
In the present study, mRNA expression of EGFR and
bFGFR was detected in U251 glioblastoma cells, and EGF and FGF were
detected in U251 cell supernatants. These results indicate that
active EGF-EGFR and FGF-FGFR autocrine pathways exist in U251
cells. It has been reported that EGF and bFGF are critical
components of human embryonic stem cell and cancer stem cell
culture media, and that they are crucial for maintaining glioma
stem cells in an undifferentiated state (15–17).
The EGF-EGFR and FGF-FGFR autocrine pathways may be involved in the
dedifferentiation mechanism(s) of glioblastoma cells.
siRNA mediates sequence-specific mRNA degradation
and has become a useful tool for silencing specific target genes.
In the present study, siRNA duplexes targeting human EGFR and bFGFR
were designed, synthesized and used to knock down EGFR and/or FGFR
gene expression, in order to aid in the elucidation of the roles of
EGF-EGFR and FGF-FGFR autocrine pathways in the differentiation of
U251 glioblastoma cells. EGFP-labeled recombinant lentiviral
expression vectors for the siRNAs were constructed and confirmed by
PCR, restriction enzyme digestion and DNA sequencing. The siRNAs
targeting EGFR and FGFR effectively knocked down the receptor mRNA
levels in the U251 cells.
EGF and bFGF are important autocrine factors in
glioblastomas and enable cancer stem-like cells to persist in
established glioblastoma cell lines without the addition of
exogenous growth factors (18,19).
Activated EGFR and bFGFR signaling increases cell proliferation,
survival and migration, and blocks glial and neuronal
differentiation in glioblastomas. In the present study, knockdown
of the EGFR and/or bFGFR genes significantly downregulated CD133
mRNA expression and facilitated U251 cell glial and neuronal
differentiation. Knockdown of both EGFR and FGFR appeared to act
synergistically to downregulate CD133 expression. This demonstrates
the importance of EGF and bFGF in maintaining the undifferentiated
state of cancer stem cells and stem cells. However, this is not the
case for promoting glial and neuronal differentiation of U251
cells. In fact, the facilitation of U251 cell glial and neuronal
differentiation was almost the same whether the EGFR pathway, the
FGFR pathway or both pathways were blocked. Notably, knocking down
EGFR and/or FGFR did not facilitate oligodendrocyte differentiation
of the U251 cells, as others have reported in stem cells (20). By contrast, MBP mRNA expression was
downregulated subsequent to blocking either the EGFR or FGFR
pathway. These findings indicate that EGFR and/or FGFR contribute
to the regulation of stem cell marker gene expression and
differentiation.
Stem cell markers and multilineage phenotypes have
been described in glioblastomas. The present study showed that U251
glioblastoma cells express stem cell markers and neuronal,
astrocytic and oligodendroglial lineage markers. EGFR and bFGFR
mRNA and EGF and bFGF proteins were detected in U251 cells and
culture supernatant, respectively. This indicated that autocrine
and/or paracrine EGF and/or bFGF mechanisms exist in U251 cells.
The knockdown of the EGFR and/or FGFR genes downregulated CD133
mRNA expression and facilitated glial and neuronal differentiation
in the U251 cells.
Acknowledgements
This study was supported by grants from the Science
and Technology Department of Zhejiang Province and the Health
Bureau of Zhejiang Provincial.
Abbreviations:
|
EGF
|
epithelial growth factor
|
|
bFGF
|
basic fibroblast growth factor
|
|
EGFR
|
epithelial growth factor receptor
|
|
bFGFR
|
basic fibroblast growth factor
|
|
GFAP
|
glial fibrillary acidic protein
|
|
HEK293
|
human embryonic kidney cell line
293
|
|
DMEM
|
Dulbecco’s modified Eagle’s medium
|
|
FBS
|
fetal bovine serum
|
References
|
1
|
Ekstrand AJ, James CD, Cavenee WK, Seliger
B, Pettersson RF and Collins VP: Genes for epidermal growth factor
receptor, transforming growth factor alpha, and epidermal growth
factor and their expression in human gliomas in vivo. Cancer Res.
51:2164–2172. 1991.
|
|
2
|
Bredel M, Pollack IF, Hamilton RL and
James CD: Epidermal growth factor receptor expression and gene
amplification in high-grade non-brainstem gliomas of childhood.
Clin Cancer Res. 5:1786–1792. 1999.PubMed/NCBI
|
|
3
|
Morrison RS, Gross JL, Herblin WF, Reilly
TM, LaSala PA, Alterman RL, Moskal JR, Kornblith PL and Dexter DL:
Basic fibroblast growth factor-like activity and receptors are
expressed in a human glioma cell line. Cancer Res. 50:2524–2529.
1990.PubMed/NCBI
|
|
4
|
Yuan X, Curtin J, Xiong Y, Liu G,
Waschsmann-Hogiu S, Farkas DL, Black KL and Yu JS: Isolation of
cancer stem cells from adult glioblastoma multiforme. Oncogene.
23:9392–9400. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Singh SK, Clarke ID, Terasaki M, Bonn VE,
Hawkins C, Squire J and Dirks PB: Identification of a cancer stem
cell in human brain tumors. Cancer Res. 63:5821–5828.
2003.PubMed/NCBI
|
|
6
|
Singh SK, Hawkins C, Clarke ID, Squire JA,
Bayani J, Hide T, Henkelman RM, Cusimano MD and Dirks PB:
Identification of human brain tumour initiating cells. Nature.
432:396–401. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Eiselleova L, Matulka K, Kriz V, Kunova M,
Schmidtova Z, Neradil J, Tichy B, Dvorakova D, Pospisilova S, Hampl
A and Dvorak P: A complex role for FGF-2 in self-renewal, survival,
and adhesion of human embryonic stem cells. Stem Cells.
27:1847–1857. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
8
|
Howard BM, Gursel DB, Bleau AM, Beyene RT,
Holland EC and Boockvar JA: EGFR signaling is differentially
activated in patient-derived glioblastoma stem cells. J Exp Ther
Oncol. 8:247–260. 2010.PubMed/NCBI
|
|
9
|
Dvorak P and Hampl A: Basic fibroblast
growth factor and its receptors in human embryonic stem cells.
Folia Histochem Cytobiol. 43:203–208. 2005.PubMed/NCBI
|
|
10
|
McComb RD and Burger PC: Pathologic
analysis of primary brain tumors. Neurol Clin. 3:711–728.
1985.PubMed/NCBI
|
|
11
|
Ma YH, Mentlein R, Knerlich F, Kruse ML,
Mehdorn HM and Held-Feindt J: Expression of stem cell markers in
human astrocytomas of different WHO grades. J Neurooncol. 86:31–45.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rieske P, Golanska E, Zakrzewska M,
Piaskowski S, Hulas-Bigoszewska K, Wolańczyk M, Szybka M,
Witusik-Perkowska M, Jaskolski DJ, Zakrzewski K, et al: Arrested
neural and advanced mesenchymal differentiation of glioblastoma
cells-comparative study with neural progenitors. BMC Cancer.
9:542009. View Article : Google Scholar
|
|
13
|
Ricci-Vitiani L, Pallini R, Larocca LM,
Lombardi DG, Signore M, Pierconti F, Petrucci G, Montano N, Maira G
and De Maria R: Mesenchymal differentiation of glioblastoma stem
cells. Cell Death Differ. 15:1491–1498. 2008. View Article : Google Scholar
|
|
14
|
Abhold EL, Kiang A, Rahimy E, et al: EGFR
kinase promotes acquisition of stem cell-like properties: a
potential therapeutic target in head and neck squamous cell
carcinoma stem cells. PLoS One. 7:e324592012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Soeda A, Inagaki A, Oka N, Ikegame Y, Aoki
H, Yoshimura S, Nakashima S, Kunisada T and Iwama T: Epidermal
growth factor plays a crucial role in mitogenic regulation of human
brain tumor stem cells. J Biol Chem. 283:10958–10966. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Kitchens DL, Snyder EY and Gottlieb DI:
FGF and EGF are mitogens for immortalized neural progenitors. J
Neurobiol. 25:797–807. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kondo T, Setoguchi T and Taga T:
Persistence of a small subpopulation of cancer stem-like cells in
the C6 glioma cell line. Proc Natl Acad Sci USA. 101:781–786. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Kelly JJ, Stechishin O, Chojnacki A, Lun
X, Sun B, Senger DL, Forsyth P, Auer RN, Dunn JF, Cairncross JG, et
al: Proliferation of human glioblastoma stem cells occurs
independently of exogenous mitogens. Stem Cells. 27:1722–1733.
2009. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li G, Chen Z, Hu YD, Wei H, Li D, Ji H and
Wang DL: Autocrine factors sustain glioblastoma stem cell
self-renewal. Oncol Rep. 21:419–424. 2009.PubMed/NCBI
|
|
20
|
Ju P, Zhang S, Yeap Y and Feng Z:
Induction of neuronal phenotypes from NG2+ glial progenitors by
inhibiting epidermal growth factor receptor in mouse spinal cord
injury. Glia. 60:1801–1814. 2012.
|