|
1
|
Dumitrescu RG: Epigenetic targets in
cancer epidemiology. Methods Mol Biol. 471:457–467. 2009.
|
|
2
|
Jaenisch R and Bird A: Epigenetic
regulation of gene expression: how the genome integrates intrinsic
and environmental signals. Nat Genet. 33(Suppl): S245–S254.
2003.
|
|
3
|
Liu WR, Shi YH, Peng YF and Fan J:
Epigenetics of hepatocellular carcinoma, a new horizon. Chin Med J
(Engl). 125:2349–2360. 2012.
|
|
4
|
Logan CY and Nusse R: The Wnt signaling
pathway in development and disease. Annu Rev Cell Dev Biol.
20:781–810. 2004.
|
|
5
|
Kim M, Lee HC, Tsedensodnom O, et al:
Functional interaction between Wnt3 and Frizzled-7 leads to
activation of the Wnt/betacatenin signaling pathway in
hepatocellular carcinoma cells. J Hepatol. 48:780–791. 2008.
|
|
6
|
Parkin DM, Bray F, Ferlay J and Pisani P:
Global cancer statistics 2002. CA Cancer J Clin. 55:74–108.
2005.
|
|
7
|
Calvisi DF, Ladu S, Gorden A, et al:
Mechanistic and prognostic significance of aberrant methylation in
the molecular pathogenesis of human hepatocellular carcinoma. J
Clin Invest. 117:2713–2722. 2007.
|
|
8
|
Nishida N, Nagasaka T, Nishimura T, Ikai
I, Boland CR and Goel A: Aberrant methylation of multiple tumor
suppressor genes in aging liver, chronic hepatitis, and
hepatocellular carcinoma. Hepatology. 47:908–918. 2008.
|
|
9
|
Shin SH, Kim BH, Jang JJ, Suh KS and Kang
GH: Identification of novel methylation markers in hepatocellular
carcinoma using a methylation array. J Korean Med Sci.
25:1152–1159. 2010.
|
|
10
|
Jung N, Won JK, Kim BH, et al:
Pharmacological unmasking microarray approach-based discovery of
novel DNA methylation markers for hepatocellular carcinoma. J
Korean Med Sci. 27:594–604. 2012.
|
|
11
|
Huang J, Zhang YL, Teng XM, et al:
Down-regulation of SFRP1 as a putative tumor suppressor gene can
contribute to human hepatocellular carcinoma. BMC Cancer.
7:1262007.
|
|
12
|
Zhang C, Li H, Zhou G, et al:
Transcriptional silencing of the TMS1/ASC tumour suppressor gene by
an epigenetic mechanism in hepatocellular carcinoma cells. J
Pathol. 212:134–142. 2007.
|
|
13
|
Park HJ, Yu E and Shim YH: DNA
methyltransferase expression and DNA hypermethylation in human
hepatocellular carcinoma. Cancer Lett. 233:271–278. 2006.
|
|
14
|
Kanai Y, Ushijima S, Tsuda H, Sakamoto M
and Hirohashi S: Aberrant DNA methylation precedes loss of
heterozygosity on chromosome 16 in chronic hepatitis and liver
cirrhosis. Cancer Lett. 148:73–80. 2000.
|
|
15
|
Kanai Y, Hui AM, Sun L, et al: DNA
hypermethylation at the D17S5 locus and reduced HIC-1 mRNA
expression are associated with hepatocarcinogenesis. Hepatology.
29:703–709. 1999.
|
|
16
|
Zhang X, Li HM, Liu Z, et al: Loss of
heterozygosity and methylation of multiple tumor suppressor genes
on chromosome 3 in hepatocellular carcinoma. J Gastroenterol.
48:132–143. 2012.
|
|
17
|
Saxonov S, Berg P and Brutlag DL: A
genome-wide analysis of CpG dinucleotides in the human genome
distinguishes two distinct classes of promoters. Proc Natl Acad Sci
USA. 103:1412–1417. 2006.
|
|
18
|
Weber M, Hellmann I, Stadler MB, et al:
Distribution, silencing potential and evolutionary impact of
promoter DNA methylation in the human genome. Nat Genet.
39:457–466. 2007.
|
|
19
|
Baylin SB and Chen WY: Aberrant gene
silencing in tumor progression, implications for control of cancer.
Cold Spring Harb Symp Quant Biol. 70:427–433. 2005.
|
|
20
|
Pogribny IP, Muskhelishvili L, Tryndyak VP
and Beland FA: The role of epigenetic events in genotoxic
hepatocarcinogenesis induced by 2-acetylaminofluorene. Mutat Res.
722:106–113. 2011.
|
|
21
|
Matsuda Y, Ichida T, Matsuzawa J, Sugimura
K and Asakura H: p16 (INK4) is inactivated by extensive CpG
methylation in human hepatocellular carcinoma. Gastroenterology.
116:394–400. 1999.
|
|
22
|
Lin YW, Chen CH, Huang GT, et al:
Infrequent mutations and no methylation of CDKN2A (P16/MTS1) and
CDKN2B (p15/MTS2) in hepatocellular carcinoma in Taiwan. Eur J
Cancer. 34:1789–1795. 1998.
|
|
23
|
Shen L, Ahuja N, Shen Y, et al: DNA
methylation and environmental exposures in human hepatocellular
carcinoma. J Natl Cancer Inst. 94:755–761. 2002.
|
|
24
|
Marotta F, Vangieri B, Cecere A and
Gattoni A: The pathogenesis of hepatocellular carcinoma is
multifactorial event. Novel immunological treatment in prospect.
Clin Ter. 155:187–199. 2004.
|
|
25
|
Cougot D, Neuveut C and Buendia MA: HBV
induced carcinogenesis. J Clin Virol. 34(Suppl 1): S75–S78.
2005.
|
|
26
|
Anzola M: Hepatocellular carcinoma, role
of hepatitis B and hepatitis C viruses proteins in
hepatocarcinogenesis. J Viral Hepat. 11:383–393. 2004.
|
|
27
|
Barazani Y, Hiatt JR, Tong MJ and Busuttil
RW: Chronic viral hepatitis and hepatocellular carcinoma. World J
Surg. 31:1243–1248. 2007.
|
|
28
|
Kiran M, Chawla YK and Kaur J: Methylation
profiling of tumor suppressor genes and oncogenes in hepatitis
virus-related hepatocellular carcinoma in northern India. Cancer
Genet Cytogenet. 195:112–119. 2009.
|
|
29
|
Zhong S, Yeo W, Tang MW, et al: Intensive
hypermethylation of the CpG island of Ras association domain family
1A in hepatitis B virus-associated hepatocellular carcinomas. Clin
Cancer Res. 9:3376–3382. 2003.
|
|
30
|
Schagdarsurengin U, Wilkens L, Steinemann
D, Flemming P, et al: Frequent epigenetic inactivation of the
RASSF1A gene in hepatocellular carcinoma. Oncogene. 22:1866–1871.
2003.
|
|
31
|
Zheng D, Liu BB, Liu YK, et al: Screening
for differential methylation status by CpG island microarray in the
hepatocellular carcinoma cell lines. Zhonghua Zhong Liu Za Zhi.
30:891–896. 2008.(In Chinese).
|
|
32
|
Liu BB, Zheng D, Liu YK, et al:
Array-based profiling of the differential methylation status of CpG
islands in hepatocellular carcinoma cell lines. Oncol Lett.
1:815–820. 2010.
|
|
33
|
Jurkowska RZ and Jeltsch A: Silencing of
gene expression by targeted DNA methylation: concepts and
approaches. Methods Mol Biol. 649:149–161. 2010.
|
|
34
|
Kim JK, Samaranayake M and Pradhan S:
Epigenetic mechanisms in mammals. Cell Mol Life Sci. 66:596–612.
2009.
|
|
35
|
Oh BK, Kim H, Park HJ, et al: DNA
methyltransferase expression and DNA methylation in human
hepatocellular carcinoma and their clinicopathological correlation.
Int J Mol Med. 20:65–73. 2007.
|
|
36
|
Liao YJ, Liu SP, Lee CM, et al:
Characterization of a glycine N-methyltransferase gene knockout
mouse model for hepatocellular carcinoma: Implications of the
gender disparity in liver cancer susceptibility. Int J Cancer.
124:816–826. 2009.
|
|
37
|
Zhao Z, Wu Q, Cheng J, et al: Depletion of
DNMT3A suppressed cell proliferation and restored PTEN in
hepatocellular carcinoma cell. J Biomed Biotechnol.
2010:7375352010.
|
|
38
|
Fan H, Chen L, Zhang F, et al: MTSS1, a
novel target of DNA methyltransferase 3B, functions as a tumor
suppressor in hepatocellular carcinoma. Oncogene. 31:2298–2308.
2012.
|
|
39
|
Park IY, Sohn BH, Yu E, et al: Aberrant
epigenetic modifications in hepatocarcinogenesis induced by
hepatitis B virus X protein. Gastroenterology. 132:1476–1494.
2007.
|
|
40
|
Bartel DP: MicroRNAs, genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
|
|
41
|
Zhang X, Liu S, Hu T, Liu S, He Y and Sun
S: Up-regulated microRNA-143 transcribed by nuclear factor kappa B
enhances hepatocarcinoma metastasis by repressing fibronectin
expression. Hepatology. 50:490–499. 2009.
|
|
42
|
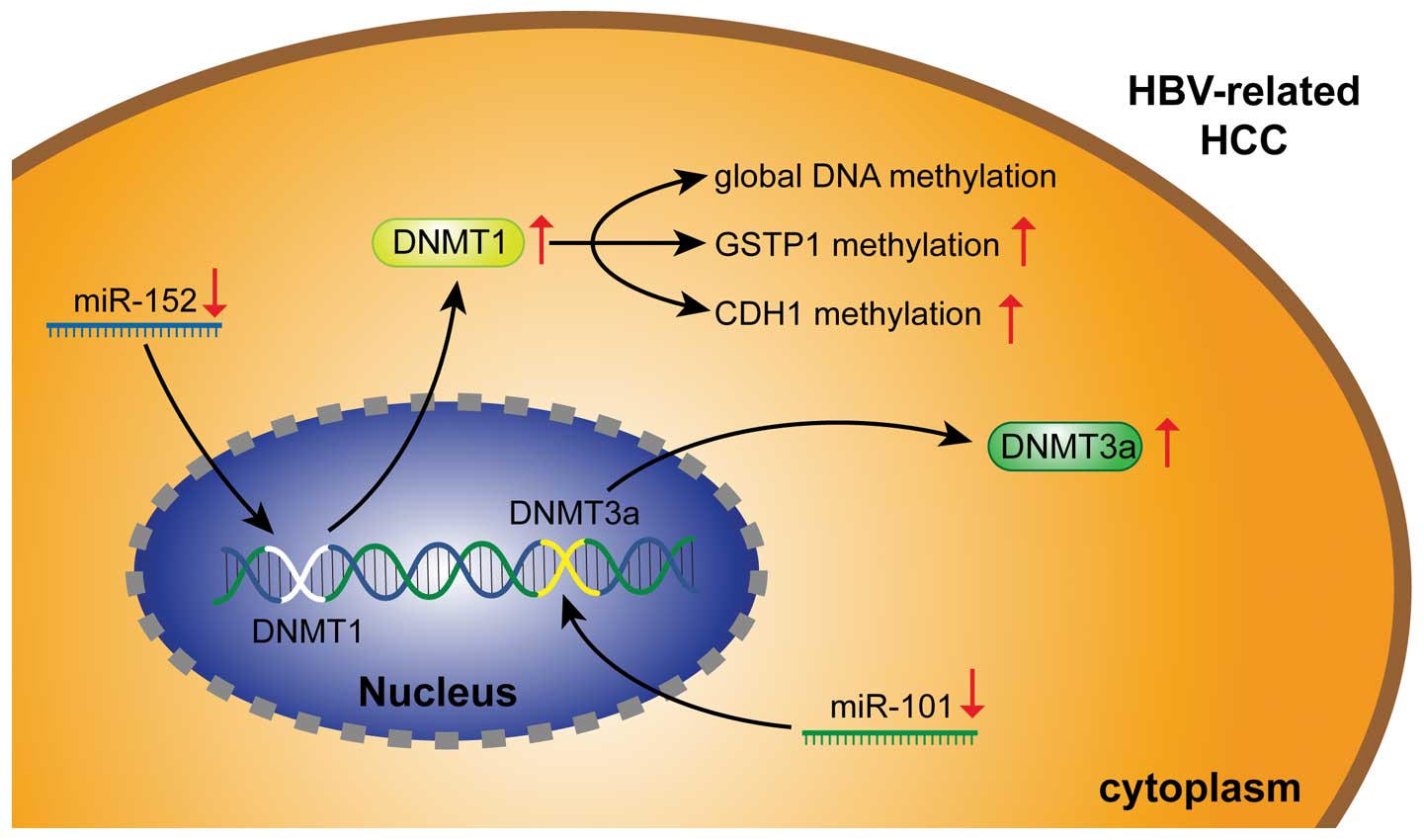
Huang J, Wang Y, Guo Y and Sun S:
Down-regulated microRNA-152 induces aberrant DNA methylation in
hepatitis B virus-related hepatocellular carcinoma by targeting DNA
methyltransferase 1. Hepatology. 52:60–70. 2010.
|
|
43
|
Wei X, Xiang T, Ren G, et al: miR-101 is
down-regulated by the hepatitis B virus x protein and induces
aberrant DNA methylation by targeting DNA methyltransferase 3A.
Cell Signal. 25:439–446. 2013.
|
|
44
|
Lim SO, Gu JM, Kim MS, et al: Epigenetic
changes induced by reactive oxygen species in hepatocellular
carcinoma: methylation of the E-cadherin promoter.
Gastroenterology. 135:2128–2140. 2008.
|
|
45
|
Blackburn RV, Spitz DR, Liu X, et al:
Metabolic oxidative stress activates signal transduction and gene
expression during glucose deprivation in human tumor cells. Free
Radic Biol Med. 26:419–430. 1999.
|
|
46
|
Lee YJ, Galoforo SS, Berns CM, et al:
Glucose deprivation-induced cytotoxicity and alterations in
mitogen-activated protein kinase activation are mediated by
oxidative stress in multidrug-resistant human breast carcinoma
cells. J Biol Chem. 273:5294–5299. 1998.
|
|
47
|
Rahman I, Marwick J and Kirkham P: Redox
modulation of chromatin remodeling: impact on histone acetylation
and deacetylation, NF-kappaB and pro-inflammatory gene expression.
Biochem Pharmacol. 68:1255–1267. 2004.
|
|
48
|
Adenuga D, Yao H, March TH, Seagrave J and
Rahman I: Histone deacetylase 2 is phosphorylated, ubiquitinated,
and degraded by cigarette smoke. Am J Respir Cell Mol Biol.
40:464–473. 2009.
|
|
49
|
Ito K, Lim S, Caramori G, Chung KF, Barnes
PJ and Adcock IM: Cigarette smoking reduces histone deacetylase 2
expression, enhances cytokine expression, and inhibits
glucocorticoid actions in alveolar macrophages. FASEB J.
15:1110–1112. 2001.
|
|
50
|
Quint K, Agaimy A, Di Fazio P, et al:
Clinical significance of histone deacetylases 1, 2, 3, and 7, HDAC2
is an independent predictor of survival in HCC. Virchows Arch.
459:129–139. 2011.
|
|
51
|
Zopf S, Ocker M, Neureiter D, et al:
Inhibition of DNA methyltransferase activity and expression by
treatment with the pan-deacetylase inhibitor panobinostat in
hepatocellular carcinoma cell lines. BMC Cancer. 12:3862012.
|
|
52
|
Jia Y, Yang Y, Liu S, Herman JG, Lu F and
Guo M: SOX17 antagonizes WNT/β-catenin signaling pathway in
hepatocellular carcinoma. Epigenetics. 5:743–749. 2010.
|
|
53
|
Grigoryan T, Wend P, Klaus A and
Birchmeier W: Deciphering the function of canonical Wnt signals in
development and disease: conditional loss- and gain-of-function
mutations of beta-catenin in mice. Genes Dev. 22:2308–2341.
2008.
|
|
54
|
Xu X, Liu RF, Wan BB, et al: Expression of
a novel gene FAM43B repressing cell proliferation is regulated by
DNA methylation in hepatocellular carcinoma cell lines. Mol Cell
Biochem. 354:11–20. 2011.
|
|
55
|
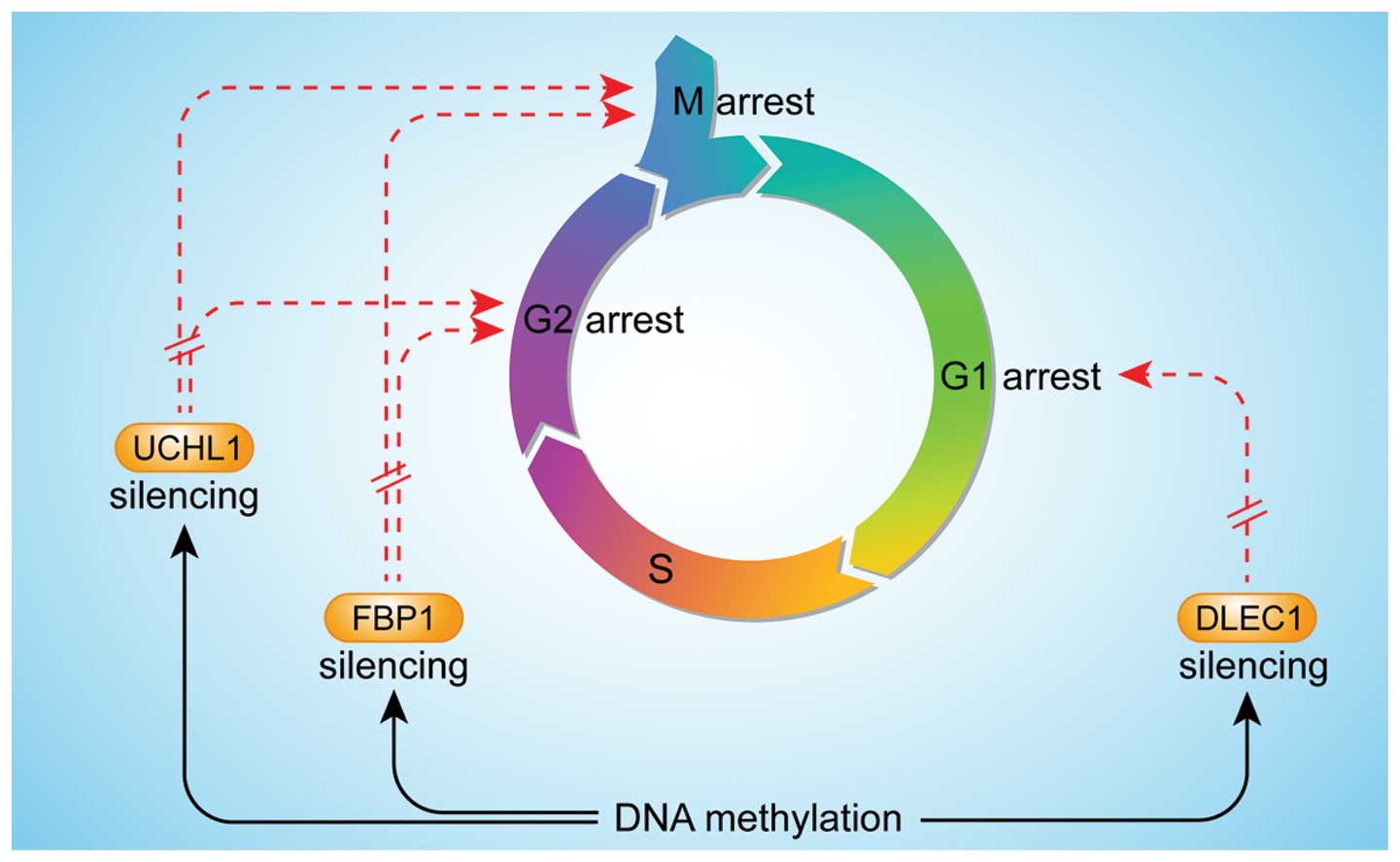
Qiu GH, Salto-Tellez M, Ross JA, et al:
The tumor suppressor gene DLEC1 is frequently silenced by DNA
methylation in hepatocellular carcinoma and induces G1 arrest in
cell cycle. J Hepatol. 48:433–441. 2008.
|
|
56
|
Yu J, Tao Q, Cheung KF, et al: Epigenetic
identification of ubiquitin carboxyl-terminal hydrolase L1 as a
functional tumor suppressor and biomarker for hepatocellular
carcinoma and other digestive tumors. Hepatology. 48:508–518.
2008.
|
|
57
|
Chen M, Zhang J, Li N, Qian Z, Zhu M, Li
Q, Zheng J, Wang X and Shi G: Promoter hypermethylation mediated
downregulation of FBP1 in human hepatocellular carcinoma and colon
cancer. PLoS One. 6:e255642011.
|
|
58
|
Komatsu S, Okazaki Y, Tateno M, et al:
Methylation and downregulated expression of mac25/insulin-like
growth factor binding protein-7 is associated with liver
tumorigenesis in SV40T/t antigen transgenic mice, screened by
restriction landmark genomic scanning for methylation (RLGS-M).
Biochem Biophys Res Commun. 267:109–117. 2000.
|
|
59
|
Berasain C, Hevia H, Fernández-Irigoyen J,
Larrea E, Caballería J, Mato JM, Prieto J, Corrales FJ,
García-Trevijano ER and Avila MA: Methylthioadenosine phosphorylase
gene expression is impaired in human liver cirrhosis and
hepatocarcinoma. Biochim Biophys Acta. 1690:276–284. 2004.
|
|
60
|
Zhou X, Popescu NC, Klein G and Imreh S:
The interferon-alpha responsive gene TMEM7 suppresses cell
proliferation and is downregulated in human hepatocellular
carcinoma. Cancer Genet Cytogenet. 177:6–15. 2007.
|
|
61
|
Nishida N, Kudo M, Nagasaka T, Ikai I and
Goel A: Characteristic patterns of altered DNA methylation predict
emergence of human hepatocellular carcinoma. Hepatology.
56:994–1003. 2012.
|
|
62
|
Xie B, Zhou J, Yuan L, Ren F, Liu DC, Li Q
and Shu G: Epigenetic silencing of Klotho expression correlates
with poor prognosis of human hepatocellular carcinoma. Hum Pathol.
44:795–801. 2012.
|
|
63
|
Lou C, Yang B, Gao YT, Wang YJ, Nie FH,
Yuan Q, Zhang CL and Du Z: Aberrant methylation of multiple genes
and its clinical implication in hepatocellular carcinoma. Zhonghua
Zhong Liu Za Zhi. 30:831–836. 2008.(In Chinese).
|
|
64
|
Nishida N, Nagasaka T, Nishimura T, Ikai
I, Boland CR and Goel A: Aberrant methylation of multiple tumor
suppressor genes in aging liver, chronic hepatitis, and
hepatocellular carcinoma. Hepatology. 47:908–918. 2008.
|
|
65
|
Piao GH, Piao WH, He Y, Zhang HH, Wang GQ
and Piao Z: Hyper-methylation of RIZ1 tumor suppressor gene is
involved in the early tumorigenesis of hepatocellular carcinoma.
Histol Histopathol. 23:1171–1175. 2008.
|
|
66
|
Lal G, Padmanabha L, Smith BJ, Nicholson
RM, Howe JR, O’Dorisio MS and Domann FE Jr: RIZ1 is epigenetically
inactivated by promoter hypermethylation in thyroid carcinoma.
Cancer. 107:2752–2759. 2006.
|
|
67
|
Chadwick RB, Jiang GL, Bennington GA, Yuan
B, Johnson CK, Stevens MW, Niemann TH, Peltomaki P, Huang S and de
la Chapelle A: Candidate tumor suppressor RIZ is frequently
involved in colorectal carcinogenesis. Proc Natl Acad Sci USA.
97:2662–2667. 2000.
|
|
68
|
Piao GH, Piao WH, He Y, et al:
Hyper-methylation of RIZ1 tumor suppressor gene is involved in the
early tumorigenesis of hepatocellular carcinoma. Histol
Histopathol. 23:1171–1175. 2008.
|
|
69
|
Zhang C, Li H, Wang Y, et al: Epigenetic
inactivation of the tumor suppressor gene RIZ1 in hepatocellular
carcinoma involves both DNA methylation and histone modifications.
J Hepatol. 53:889–895. 2010.
|
|
70
|
Formeister EJ, Tsuchiya M, Fujii H, et al:
Comparative analysis of promoter methylation and gene expression
endpoints between tumorous and non-tumorous tissues from
HCV-positive patients with hepatocellular carcinoma. Mutat Res.
692:26–33. 2010.
|
|
71
|
Yang JD, Seol SY, Leem SH, Kim YH, Sun Z,
Lee JS, Thorgeirsson SS, Chu IS, Roberts LR and Kang KJ: Genes
associated with recurrence of hepatocellular carcinoma: integrated
analysis by gene expression and methylation profiling. J Korean Med
Sci. 26:1428–1438. 2011.
|
|
72
|
Iyer P, Zekri AR, Hung CW, Schiefelbein E,
Ismail K, Hablas A, Seifeldin IA and Soliman AS: Concordance of DNA
methylation pattern in plasma and tumor DNA of Egyptian
hepatocellular carcinoma patients. Exp Mol Pathol. 88:107–111.
2010.
|
|
73
|
Kimura N, Moribe T, Iizuka N, Miura T,
Tamatsukuri S, Ishitsuka H, Hamamoto Y and Oka M: Rapid and
quantitative detection of CpG-methylation status using TaqMan PCR
combined with methyl-binding-domain polypeptide. Clin Biochem.
42:1113–1122. 2009.
|