Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most
common subtype of non-Hodgkin lymphoma (NHL) and constitutes a
heterogeneous category of aggressive lymphomas (1). Numerous studies have shown that genes
and microRNAs (miRNAs) exert various roles in DLBCL (2–5).
Furthermore, differentially expressed genes and miRNAs are
important in the pathogenesis of DLBCL; for example, TRX1 is
key in cell growth and survival, as well as in the chemoresistance
of relapsed/refractory DLBCL (6).
In addition, the misregulation of hsa-miR-155 and hsa-miR-146a acts
as a diagnostic and prognostic marker (7). Genes and miRNAs that are not
differentially expressed, but are associated with DLBCL also exert
particular roles in DLBCL. The B cell receptor isotype is a
reliable indicator for the GCB and ABC subtypes in
DLBCL (8), and hsa-miR-135b
contributes to tumorigenesis through modulation of the tumor
immune-phenotype and microenvironment (9).
Gene regulatory factors are predominantly comprised
of transcription factors (TFs) and miRNAs, and these control the
expression of genomic information in multicellular genomes
(10). TFs are proteins that bind
to specific DNA sequences, controlling the transfer of genetic
information between DNA and messenger RNA (11). TFs may regulate (activate or
repress) gene expression alone or in conjunction with other
proteins. miRNAs are small non-coding RNA molecules (~22
nucleotides in length) that function in the transcriptional and
post-transcriptional regulation of gene expression (12). miRNAs regulate gene expression by
silencing genes or targeting genes for degradation, and influence
various cancer processes, including proliferation, differentiation
and apoptosis.
miRNAs target thousands of human genes, usually
known as target genes (targets), which are important in analyzing
the biological functions of miRNAs. Currently, a number of
arithmetic methods (13) and
experimentally validated databases (14,15)
have provided sufficient data to investigate the associations among
different miRNAs.
In DNA sequences, miRNAs are encoded by certain
genes; these are usually termed miRNA host genes. In the
transcription process, miRNAs and the corresponding host gene are
transcribed simultaneously (16).
The host gene and the intronic miRNA are coordinately expressed in
certain biological processes (17),
and together accomplish certain functions and are involved in
signaling pathways (18).
The occurrence of DLBCL cannot be attributed to a
single gene, miRNA or signaling pathway; rather, DLBCL is the
result of various biological functions acting together. A number of
genes and miRNAs associated with DLBCL have been identified;
however, the underlying mechanism of miRNA and gene involvement in
DLBCL remains largely unknown. The present study focused on the
networks of TFs, miRNAs, miRNA targets and miRNA host genes to
examine the key regulatory associations in DLBCL and partially
reveal the underlying control mechanisms. Experimentally validated
associations (among TFs and miRNAs, miRNAs and the respective
target genes, and miRNAs and the corresponding host genes) were
collected from TarBase, miRTarBase, TransmiR and miRBase databases
(14,15,19,20).
Differentially expressed genes and miRNAs in DLBCL, and
DLBCL-associated genes and miRNAs were collected from databases and
the relevant literature. To further investigate the DLBCL
transcriptional network, TFs were obtained by the P-match method
and were considered as DLBCL-associated genes. The associated genes
and miRNAs included those that were differentially and
non-differentially expressed. Three networks were constructed to
gradually understand the mechanism of DLBCL. The first is an
experimentally validated network of miRNAs and genes constructed
from all data. The second is a differential expression network;
differentially expressed genes and miRNAs were mapped onto the
first network, then extracted to construct the second network. The
third is a DLBCL-associated network (similar methods were used to
construct this third network). The regulatory associations between
differentially expressed genes, differentially expressed miRNAs and
predicted TFs were separately extracted from the three networks,
and the similarities and differences were compared to distinguish
the key regulatory associations in DLBCL.
Materials and methods
Data collection and processing
An experimentally validated dataset of miRNAs and
the corresponding targets were extracted from TarBase 5.0 (Diana
Lab, Philadelphia, PA, USA) (14)
and miRTarBase (15). The National
Center for Biotechnology Information (NCBI) gene database
(http://www.ncbi.nlm.nih.gov/gene/)
was used to unify the official symbols of miRNAs and genes; this
dataset was designated set U1.
An experimentally validated dataset of TFs and
miRNAs was extracted from the TransmiR database (http://www.cuilab.cn/transmir) (19), and this dataset was termed set
U2.
The dataset of host genes and the respective miRNAs
was extracted from the miRBase (http://www.mirbase.org/)(20) and NCBI gene databases. This dataset
was set U3.
In the present study, the differentially expressed
genes included genetically mutated genes, abnormally expressed
protein genes, single nucleotide polymorphisms (SNPs) and
overexpressed, downregulated, upregulated genes. The dataset of
differentially expressed genes was retrieved from Cancer Genetics
Web (http://www.cancerindex.org/geneweb/index.html) and the
relevant literature was obtained from the NCBI SNP database
(http://www.ncbi.nlm.nih.gov/snp/).
Numerous DLBCL-associated genes have been detected,
such as those involved in the development and metastasis of human
DLBCL, as well as those used therapeutically in DLBCL prevention,
diagnosis and radial therapy. The DLBCL-associated genes also
include the differentially expressed genes. The dataset of
DLBCL-associated genes was collected from the GeneCards
database(http://www.genecards.org/) (21) and the relevant studies found using
PubMed. To gain an improved understanding of the transcriptional
network of TFs, miRNAs and targets, popular TFs were extracted
using the P-match method (Biobase, Wolfenbüttel, Germany) (22). These TFs are termed the
DLBCL-associated genes. The present study focused on the TFs that
appear on the TransmiR database. Since miRNAs regulate gene
expression together with TFs, 1,000-nt promoter region sequences of
targets that are targeted by differentially expressed miRNAs were
downloaded from the University of California, Santa Cruz Genome
Browser (23). The P-match method
was used to identify TF binding sites (TFBSs) in the 1,000-nt
promoter region sequences. These TFBSs were mapped onto the
promoter regions of targets, then the corresponding TFs of these
TFBSs were obtained. In the P-match method, the vertebrate matrix
was selected with a high-quality criterion for the extracted TFs.
The dataset of differentially expressed and associated genes was
set U4.
Differentially expressed miRNAs include
overexpressed, downregulated and upregulated miRNAs. The associated
miRNAs involved in various DLBCL processes include differentially
expressed and non-differentially expressed miRNAs. The dataset of
differentially expressed miRNAs was retrieved from the mir2Disease
database (http://www.miR2Disease.org (24). The dataset of DLBCL-associated
miRNAs was collected from the differentially expressed miRNA data
and information from previous relevant studies; this dataset was
termed set U5.
Construction of the three networks
To construct the experimentally validated,
differentially expressed and associated networks, regulatory
associations among TFs, miRNAs, targets and host gene were
extracted from U1, U2 and
U3, and the associations were combined to
construct the experimentally validated network. Differentially
expressed genes and miRNAs were extracted from U4
and U5, then mapped onto the experimentally
validated network. The differential expression network was
constructed by extracting these associations and combining them.
Certain differentially expressed genes and miRNAs were not present
in U1, U2 and
U3; these were presented as single nodes in the
differential expression network. Similar methods were used to
construct the DLBCL-associated network.
Results
DLBCL differential expression
network
A number of important miRNAs, genes and the
corresponding regulatory associations may result in the occurrence
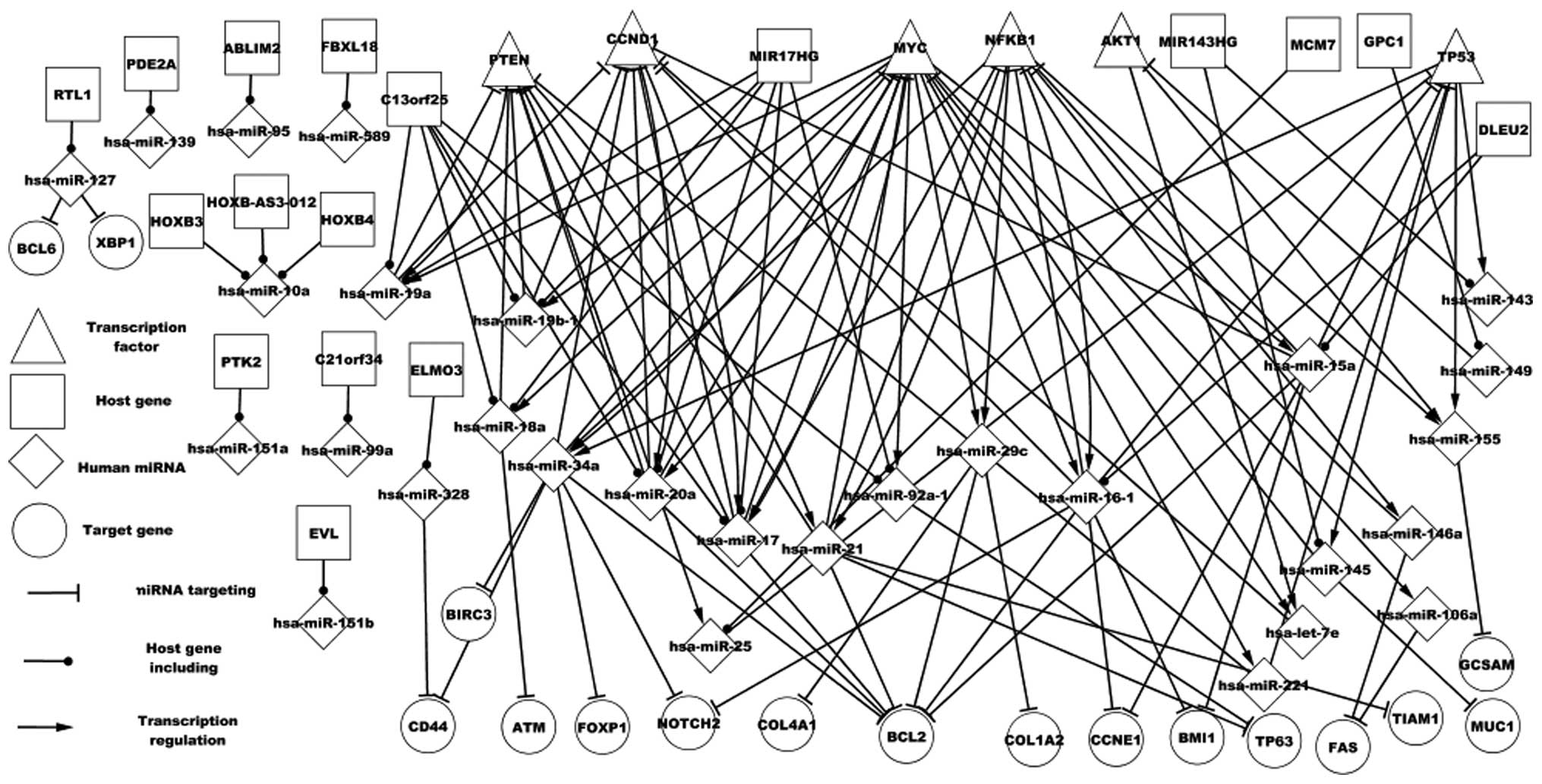
of DLBCL (Fig. 1). All single nodes
that do not have a regulatory association with miRNAs, such as
BCL10 and CASP10, were omitted, although these nodes also exert key
roles in DLBCL. A total of six TFs, 23 miRNA targets, and 21 miRNAs
and the respective host genes are presented in Fig. 1. With the exception of certain host
genes, the other nodes all indicate differentially expressed miRNAs
and proteins in DLBCL. Fig. 1
demonstrates various types of regulatory association between miRNAs
and genes. One miRNA may target one gene or numerous genes, one TF
may regulate one miRNA or numerous miRNAs, numerous TFs may
regulate one miRNA or numerous miRNAs, and numerous miRNAs may
target one gene or numerous genes. Particular features of host
genes and the respective miRNAs are revealed in Fig. 1. A host gene may encode one or
numerous miRNAs that target other genes; for example GPC1 encodes
hsa-miR-149, which targets AKT1. An miRNA may be located in various
host genes; for example, three host genes (HOXB3, HOXB4 and
HOXB-AS3-012) encode hsa-miR-10a. Thus, the differential expression
network partially revealed the regulatory mechanism of DLBCL.
DLBCL-associated network
Numerous regulatory associations among genes and
miRNAs were identified in the DLBCL-associated network. Compared
with the differential expression network, the DLBCL-associated
network includes additional TFs, miRNAs and mass targets. The
DLBCL-associated network also reveals further regulatory
associations between genes and miRNAs; for example, EGFR was
found to regulate hsa-miR-21, which targets MYC; hsa-miR-21 was
identified to target E2F1, which regulates hsa-miR-17; and TP53 was
revealed to regulate hsa-miR-125b (hsa-miR-125b-1 and -2), which
targets AKT1. These nodes are all associated with DLBCL, however,
certain nodes do not demonstrate differentially expressed data. The
DLBCL-associated network expands on the differential expression
network and these newly identified regulatory associations may
contribute to tumor growth, migration, prevention, diagnosis,
development and other processes in DLBCL.
Host genes and the corresponding miRNAs
in DLBCL
Two differentially expressed genes (TP63 and FOXP1)
were identified as host genes in the present study, although the
respective miRNAs were not differentially expressed in DLBCL.
hsa-miR-127 was found to be encoded in RTL1, and to target two
differentially expressed genes, XBP1 and BCL6. Although certain
host genes are not differentially expressed in DLBCL, the genes may
be involved in certain DLBCL processes when the miRNAs are
differentially expressed. In the differential expression network,
certain host genes and the corresponding miRNAs exhibit the feature
where a host gene encodes numerous miRNAs that alone or together
target specific genes.
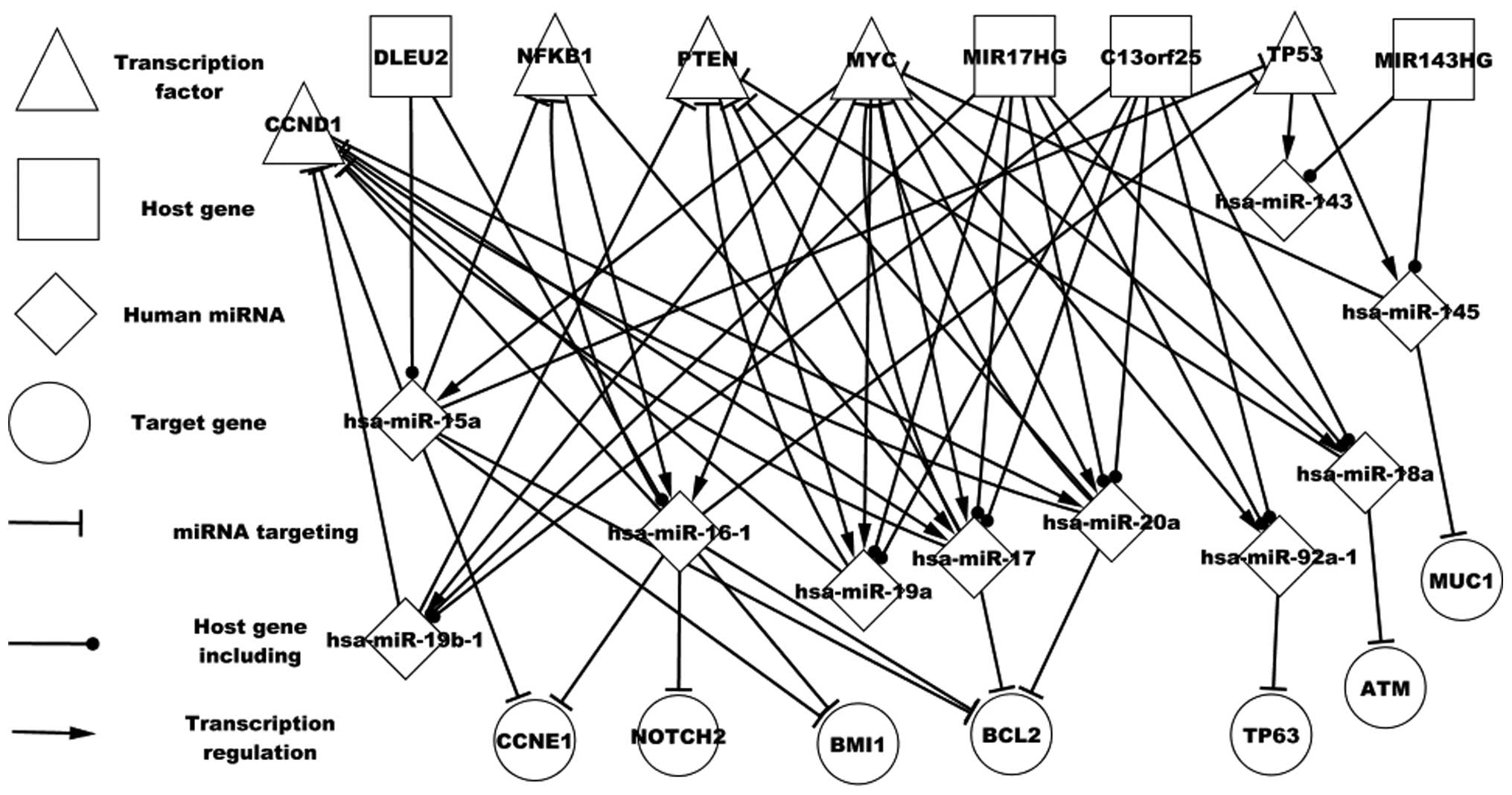
Fig. 2 shows certain
particular host genes and the regulatory associations between these
genes and TFs, and miRNAs and the corresponding targets. For
example, MIR17HG encodes six miRNAs, and four of these miRNAs
(hsa-miR-17, hsa-miR-20a, hsa-miR-19a and hsa-miR-19b-1) target
CCND1. hsa-miR-19a is regulated by MYC and PTEN and hsa-miR-19a and
PTEN form a self-adaptation association. hsa-miR-17 is regulated by
three TFs (MYC, NFKB1 and CCND1) and MIR143HG encodes two miRNAs
(hsa-miR-143 and hsa-miR-145) that are regulated by TP53.
hsa-miR-145 targets MUC1, however, hsa-miR-143 does not target any
differentially expressed genes. hsa-miR-17 and hsa-miR-16-1 are
regulated by NFKB1, which is targeted by hsa-miR-15a and
hsa-miR-16-1. In conclusion, host genes and the respective miRNAs
may aid in understanding the pathogenesis of DLBCL.
Transcriptional network of predicted
TFs
A total of 16 differentially expressed miRNAs
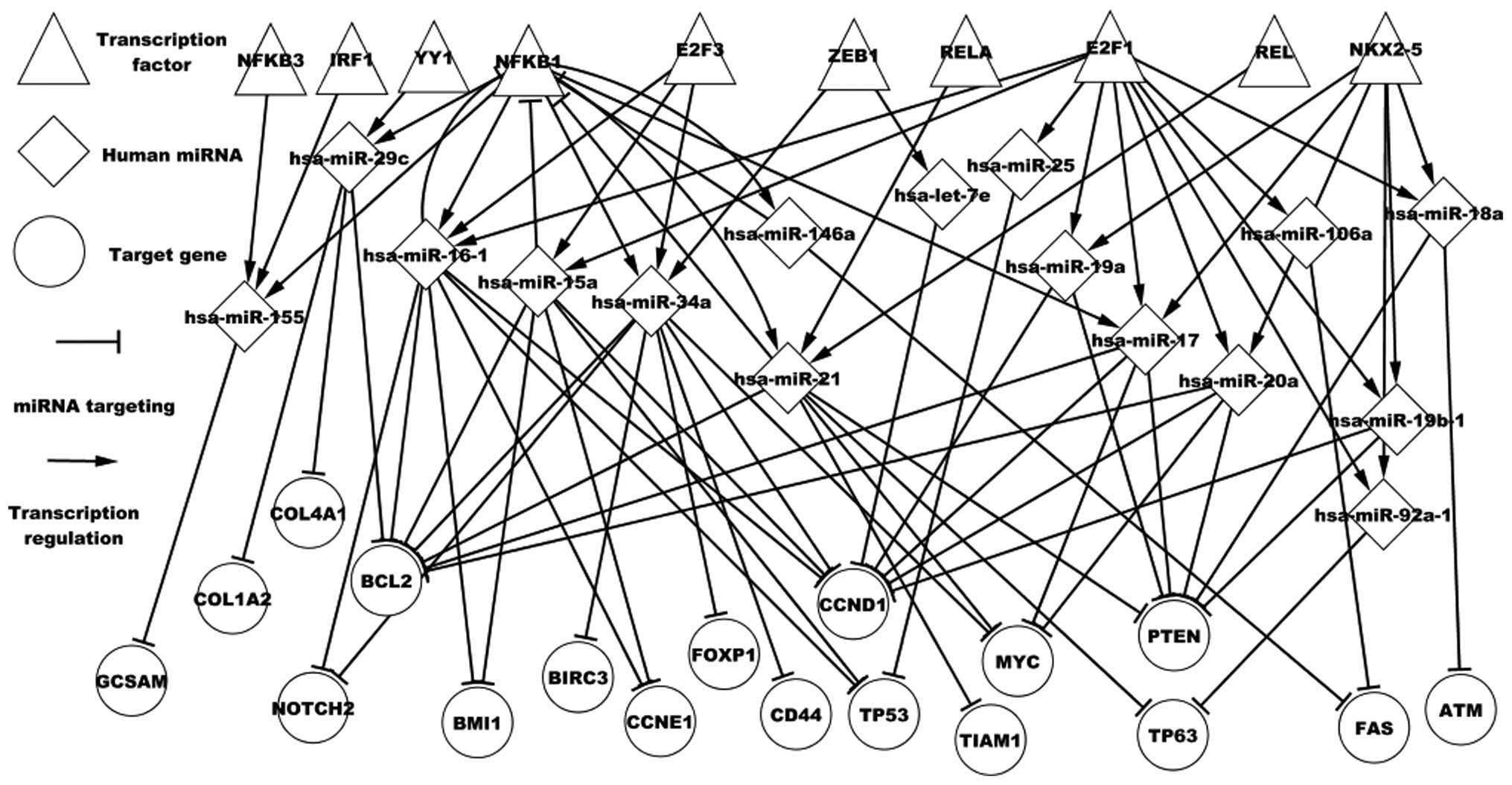
regulated by the predicted TFs were further analyzed. Fig. 3 shows the regulatory associations
among predicted TFs, differentially expressed miRNAs and
differentially expressed targets in DLBCL. These TFs and miRNAs in
turn influence the respective successors. For example, the E2F1,
PAX5, REL, RELA, STAT1 and YY1 TFs have been experimentally
validated in DLBCL. NFKB1 is a differentially expressed TF gene in
DLBCL. Fig. 3 shows that NFKB1
regulates seven miRNAs and is targeted by four miRNAs.
hsa-miR-146a, hsa-miR-21, hsa-miR-16-1 and NFKB1 form three
self-adaptation associations. NFKB1 regulates hsa-miR-17, which
targets MYC, CCND1, PTEN and BCL2. Fig.
3 also shows that a differentially expressed miRNA may be
regulated by various TFs, a target may be targeted by numerous
differentially expressed miRNAs, an miRNA may indirectly influence
other miRNAs through particular TFs and a TF may indirectly
influence other genes through various differentially expressed
miRNAs. For example, hsa-miR-29c is regulated by YY1 and NFKB1;
hsa-miR-17, hsa-miR-21, hsa-miR-20a and hsa-miR-34a target MYC: YY1
regulates hsa-miR-29c, which targets BCL2; and hsa-miR-21 targets
NFKB1, which regulates hsa-miR-155. This transcription network may
contribute to the further understanding of DLBCL pathogenesis.
Regulatory associations among
differentially expressed genes
To understand the regulatory network more clearly,
the regulatory associations of each node (differentially expressed
genes, differentially expressed miRNAs and predicted TFs) were
extracted and compared according to the predecessors and successors
of the gene, the nodes preceeding the current one on the path or
the node following the current one on the path, respectively. Among
these genes, MYB, MYC and CCND1 and four miRNAs (hsa-miR-17,
hsa-miR-155, hsa-miR-15a and hsa-miR-34a) formed five
self-adaptation associations.
For the differentially expressed genes, PTEN may be
used as an example. Table I shows
PTEN, and the respective predecessors and successors in the three
networks. Seven miRNAs target PTEN-mediated regulation of three
miRNAs in the differential expression network, eight miRNAs target
PTEN-mediated regulation of three miRNAs in the DLBCL-associated
network and 21 miRNAs target PTEN-mediated regulation of nine
miRNAs in the experimentally validated network. In the human body,
the regulatory associations between PTEN and DLBCL-associated
miRNAs influence multiple DLBCL processes. However, the regulatory
associations between PTEN and the non-associated miRNAs may not
influence DLBCL. Predecessors may indirectly influence successors
by regulation of PTEN. Two miRNAs (hsa-miR-19a and hsa-miR-21) were
found to target PTEN (Table I) and
form two self-adaptation associations. Furthermore, hsa-miR-19a and
hsa-miR-21 are also regulated by PTEN. The expression of another
miRNA may subsequently be affected when either of these miRNAs are
differentially expressed.
 | Table IRegulatory associations between miRNAs
and PTEN. |
Table I
Regulatory associations between miRNAs
and PTEN.
| miRNAs that target
PTEN | miRNAs regulated by
PTEN |
|---|
|
|
|---|
| Differential
expression network | Associated
network | Experimentally
validated network | Differential
expression network | Associated
network | Experimentally
validated network |
|---|
| | miR-106b | | | |
| | miR-141 | | | |
| | miR-17 | | | |
| | miR-18a | | | |
| | miR-19a | | | |
| | miR-19b-1 | | | |
| miR-17 | miR-17 | miR-19b-2 | miR-19a | miR-19a | miR-19a |
| miR-18a | miR-18a | miR-20a | miR-21 | miR-21 | miR-21 |
| miR-19a | miR-19a | miR-21 | miR-25 | miR-25 | miR-22 |
| miR-19b-1 | miR-19b-1 | miR-214 | | | miR-25 |
| miR-20a | miR-20a | miR-216a | | | miR-302a |
| miR-21 | miR-21 | miR-221 | | | miR-302b |
| miR-221 | miR-221 | miR-222 | | | miR-302c |
| miR-222 | miR-26a-1 | | | miR-302d |
| | miR-26a-2 | | | miR-302f |
| | miR-29b | | | |
| | miR-217 | | | |
| | miR-494 | | | |
| | miR-519a | | | |
| | miR-519d | | | |
| | miR-93 | | | |
Regulatory associations among
differentially expressed miRNAs
As with the differentially expressed genes, the
regulatory associations among differentially expressed miRNAs were
extracted and compared according to the predecessors and successors
of miRNA.
The analysis only focused on hsa-miR-20a to
illustrate the regulatory associations regarding differentially
expressed miRNAs in three networks (differentially expressed,
related and experimentally validated networks). Table II shows hsa-miR-20a, and its
predecessors and successors in the three networks. CCND1 and MYC
regulate hsa-miR-20a, which targets four genes in the differential
expression network. In the DLBCL-associated network, four genes
regulate hsa-miR-20a, which targets 14 genes. In the experimentally
validated network, 10 genes regulate hsa-miR-20a, which targets 26
genes. MYC, CCND1 and hsa-miR-20a were found to form two
self-adaptation associations (Table
II). As with PTEN, regulatory associations between hsa-miR-20a
and DLBCL-associated genes influence multiple DLBCL processes;
however, other regulatory associations between hsa-miR-20a and
non-associated genes may not influence DLBCL.
 | Table IIRegulatory associations between
hsa-miR-20a and various genes. |
Table II
Regulatory associations between
hsa-miR-20a and various genes.
| Genes that regulate
hsa-miR-20a | Genes targeted by
hsa-miR-20a |
|---|
|
|
|---|
| Differential
expression network | Associated
network | Experimentally
validated network | Differential
expression network | Associated
network | Experimentally
validated network |
|---|
| CCND1 | CCND1 | CCND1 | BCL2 | CCND1 | APP, CCND1 |
| MYC | E2F1 | E2F1 | CCND1 | BCL2 | BCL2, BMPR2 |
| MYC | MYC | MYC | RUNX1 | BNIP2, RUNX1 |
| NKX2-5 | MYCN | PTEN | CCND2 | CCND2, CDKN1A |
| | NKX2-5 | | CDKN1A | E2F1, E2F3 |
| | TLX1 | | E2F1 | HIF1A, IRF2 |
| | TLX3 | | E2F3 | KIT, SMAD4 |
| | ESR1 | | HIF1A | MEF2D, MYC |
| | STAT5B | | IRF2 | NRAS, MAPK9 |
| | SPI1 | | KIT | PTEN, RB1 |
| | | | MYC | RBL1, RBL2 |
| | | | PTEN | TGFBR2, THBS1 |
| | | | RB1 | VEGFA, WEE1 |
| | | | TGFBR2 | MAP3K12, EGLN3 |
| | | | THBS1 | MUC17 |
| | | | VEGFA | |
Regulatory associations among predicted
TFs
The predecessors and successors of the predicted TFs
were used to extract and compare the regulatory associations of
each predicted TF. Three TFs (E2F1, E2F3 and NFKB1) and five
differentially expressed miRNAs form five self-adaptation
associations. Notably, NFKB1 is a differentially expressed gene and
E2F1 is associated with DLBCL.
The present study only focused on E2F3 to illustrate
regulatory associations regarding differentially expressed miRNAs
in three networks (differentially expressed, related and
experimentally validated networks). Table III shows E2F3, and its
predecessors and successors in the three networks. Four
differentially expressed miRNAs target E2F3, which regulates three
differentially expressed miRNAs. In the DLBCL-associated network,
six miRNAs target E2F3, which regulates three miRNAs in the same
network. In the experimentally validated network, 12 miRNAs target
E2F3, which regulates 11 miRNAs. hsa-miR-34a and E2F3 were found to
form a self-adaptation association in the differential expression
network. In this self-adaptation association, E2F3 is not
differentially expressed in DLBCL, although hsa-miR-34a is
differentially expressed. Therefore, hsa-miR-34a may indirectly
cause other miRNAs to be erroneously expressed by E2F3.
 | Table IIIRegulatory associations between
miRNAs and the transcription factor, E2F3. |
Table III
Regulatory associations between
miRNAs and the transcription factor, E2F3.
| miRNAs that target
E2F3 | miRNAs regulated by
E2F3 |
|---|
|
|
|---|
| Differential
expression network | Associated
network | Experimentally
validated network | Differential
expression network | Associated
network | Experimentally
validated network |
|---|
| miR-17 | miR-125b-1 | miR-106b | miR-15a | miR-15a | let-7a-1 |
| miR-20a | miR-125b-2 | miR-125b-1 | miR-16-1 | miR-16-1 | let-7a-2 |
| miR-210 | miR-17 | miR-125b-2 | miR-34a | miR-34a | let-7a-3 |
| miR-34a | miR-20a | miR-128b | | | miR-15b |
| miR-210 | miR-17 | | | miR-15a |
| miR-34a | miR-195 | | | miR-16-1 |
| | miR-20a | | | miR-16-2 |
| | miR-200b | | | miR-195 |
| | miR-203a | | | miR-106b |
| | miR-210 | | | miR-34a |
| | miR-34a | | | let-7i |
| | miR-34c | | | |
Discussion
Certain important regulatory associations among
differentially expressed genes, differentially expressed miRNAs and
predicted TFs were identified in the present study. For the
differentially expressed genes and miRNAs, the results indicated
numerous important regulatory associations, including the fact that
hsa-miR-149 targets AKT1, which regulates hsa-let-7e. The signaling
pathways identified may exert key biological functions in DLBCL or
affect normal physiological processes, thus resulting in the
occurrence of DLBCL. Certain signaling pathways have been detected
that influence particular processes in DLBCL; for example
hsa-miR-155 regulates the PI3K-AKT signaling pathway in DLBCL
(25). Other signaling pathways
have not been identified in DLBCL, however, may influence numerous
processes in other types of cancer; for example,
hsa-miR-15a/hsa-miR-16-1 targets BCL2 and exerts an etiological and
therapeutic role in keratocystic odontogenic tumors (26), and hsa-miR-16-1 targets CCND1 in
mantle cell lymphoma (27).
Poliseno et al (28)
observed that PTEN regulates hsa-miR-25 and Kumar et al
(29) indicated that TP53 is
targeted by hsa-miR-25. hsa-miR-25 is differentially expressed in
DLBCL, and forms an association between PTEN and TP53; PTEN may
influence TP53 expression via hsa-miR-25. The present study may
expand the understanding of the associations among these genes.
Although certain signaling pathways were not found to influence
DLBCL processes in the present study, the biological functions of
these signaling pathways in other types of cancer may contribute to
DLBCL. The remaining signaling pathways, which have not been
associated with any type of cancer, may exert potential functions
in DLBCL; for example, in the present study, hsa-miR-92a-1 was
found to target TP63. For the predicted TF signaling pathways,
certain signaling pathways have been detected in other types of
cancer; for instance, ZEB1 regulates hsa-miR-34a in lung cancer
(30) and hsa-miR-21 targets E2F2
in breast cancer (31).
In conclusion, in the present study, numerous
important regulatory associations in DLBCL were identified.
Furthermore, certain genes and miRNAs exhibited self-adaptation
associations. The differential expression network partially
revealed the pathogenesis of DLBCL and the DLBCL-associated network
supplied comprehensive data with regard to the genes and miRNAs
associated with DLBCL processes, including prevention, diagnosis,
development and therapy. Therefore, the present study contributes
to the understanding of the underlying molecular mechanisms and
potential treatment of DLBCL.
Acknowledgements
This study was supported by the grants from the
National Natural Science Foundation of China (grant no. 60973091)
and the Science and Technology Development Plan of Jilin Province
(grant no. 20130101166JC).
Abbreviations:
|
miRNA
|
microRNA
|
|
TFs
|
transcription factors
|
|
targets
|
target genes
|
|
DLBCL
|
diffuse large B-cell lymphoma
|
|
NCBI
|
National Center for Biotechnology
Information
|
|
TFBSs
|
transcription factor binding sites
|
|
NHL
|
non-Hodgkin lymphoma
|
References
|
1
|
No authors listed. A clinical evaluation
of the International Lymphoma Study Group classification of
non-Hodgkin’s lymphoma. The Non-Hodgkin’s Lymphoma Classification
Project. Blood. 89:3909–3918. 1997.
|
|
2
|
Nishiu M, Yanagawa R, Nakatsuka S, et al:
Microarray analysis of gene-expression profiles in diffuse large
B-cell lymphoma: identification of genes related to disease
progression. Jpn J Cancer Res. 93:894–901. 2002.
|
|
3
|
Bavi P, Uddin S, Bu R, et al: The
biological and clinical impact of inhibition of NF-κB-initiated
apoptosis in diffuse large B cell lymphoma (DLBCL). J Pathol.
224:355–366. 2011.
|
|
4
|
Malumbres R, Sarosiek KA, Cubedo E, et al:
Differentiation stage-specific expression of microRNAs in B
lymphocytes and diffuse large B-cell lymphomas. Blood.
113:3754–3764. 2009.
|
|
5
|
Alencar AJ, Malumbres R, Kozloski GA, et
al: MicroRNAs are independent predictors of outcome in diffuse
large B-cell lymphoma patients treated with R-CHOP. Clin Cancer
Res. 17:4125–4135. 2011.
|
|
6
|
Li C, Thompson MA, Tamayo AT, et al:
Over-expression of Thioredoxin-1 mediates growth, survival, and
chemoresistance and is a druggable target in diffuse large B-cell
lymphoma. Oncotarget. 3:314–326. 2012.
|
|
7
|
Zhong H, Xu L, Zhong JH, et al: Clinical
and prognostic significance of miR-155 and miR-146a expression
levels in formalin-fixed/paraffin-embedded tissue of patients with
diffuse large B-cell lymphoma. Exp Ther Med. 3:763–770. 2012.
|
|
8
|
Ruminy P, Etancelin P, Couronné L, et al:
The isotype of the BCR as a surrogate for the GCB and ABC molecular
subtypes in diffuse large B-cell lymphoma. Leukemia. 25:681–688.
2011.
|
|
9
|
Matsuyama H, Suzuki HI, Nishimori H, et
al: miR-135b mediates NPM-ALK-driven oncogenicity and renders
IL-17-producing immunophenotype to anaplastic large cell lymphoma.
Blood. 118:6881–6892. 2011.
|
|
10
|
Hobert O: Gene regulation by transcription
factors and microRNAs. Science. 319:1785–1786. 2008.
|
|
11
|
Latchman DS: Transcription factors: an
overview. Int J Biochem Cell Biol. 29:1305–1312. 1997.
|
|
12
|
Chen K and Rajewsky N: The evolution of
gene regulation by transcription factors and microRNAs. Nat Rev
Genet. 8:93–103. 2007.
|
|
13
|
Betel D, Wilson M, Gabow A, et al: The
microRNA.org resource: targets and expression. Nucleic Acids Res.
36:D149–D153. 2008.
|
|
14
|
Papadopoulos GL, Reczko M, Simossis VA, et
al: The database of experimentally supported targets: a functional
update of TarBase. Nucleic Acids Res. 37:D155–D158. 2009.
|
|
15
|
Hsu SD, Lin FM, Wu WY, et al: miRTarBase:
a database curates experimentally validated microRNA-target
interactions. Nucleic Acids Res. 39:D163–D169. 2011.
|
|
16
|
Rodriguez A, Griffiths-Jones S, Ashurst JL
and Bradley A: Identification of mammalian microRNA host genes and
transcription units. Genome Res. 14:1902–1910. 2004.
|
|
17
|
Baskerville S and Bartel DP: Microarray
profiling of microRNAs reveals frequent coexpression with
neighboring miRNAs and host genes. RNA. 11:241–247. 2005.
|
|
18
|
Cao G, Huang B, Liu Z, et al: Intronic
miR-301 feedback regulates its host gene, ska2, in A549 cells by
targeting MEOX2 to affect ERK/CREB pathways. Biochem Biophys Res
Comm. 396:978–982. 2010.
|
|
19
|
Wang J, Lu M, Qiu C and Cui Q: TransmiR: a
transcription factor-microRNA regulation database. Nucleic Acids
Res. 38:D119–D122. 2010.
|
|
20
|
Kozomara A and Griffiths-Jones S: miRBase
integrating microRNA annotation and deep-sequencing data. Nucleic
Acids Res. 39:D152–D157. 2011.
|
|
21
|
Safran M, Dalah I, Alexander J, et al:
GeneCards Version 3: the human gene integrator. Database (Oxford).
baq0202010.
|
|
22
|
Chekmenev DS, Haid C and Kel AE: P-Match:
transcription factor binding site search by combining patterns and
weight matrices. Nucleic Acids Res. 33:W432–W437. 2005.
|
|
23
|
Fujita PA, Rhead B, Zweig AS, et al: The
UCSC Genome Browser database: update 2011. Nucleic Acids Res.
39:D876–D882. 2011.
|
|
24
|
Jiang Q, Wang Y, Hao Y, et al:
miR2Disease: a manually curated database for microRNA deregulation
in human disease. Nucleic Acids Res. 37:D98–D104. 2009.
|
|
25
|
Huang X, Shen Y, Liu M, et al:
Quantitative proteomics reveals that miR-155 regulates the PI3K-AKT
pathway in diffuse large B-Cell lymphoma. Am J Pathol. 181:26–33.
2012.
|
|
26
|
Diniz MG, Gomes CC, de Castro WH, et al:
miR-15a/16–1 influences BCL2 expression in keratocystic odontogenic
tumors. Cellular Oncol. 35:285–291. 2012.
|
|
27
|
Chen RW, Bemis LT, Amato CM, et al:
Truncation in CCND1 mRNA alters miR-16–1 regulation in mantle cell
lymphoma. Blood. 112:822–829. 2008.
|
|
28
|
Poliseno L, Salmena L, Riccardi L, et al:
Identification of the miR-106b~25 microRNA cluster as a
proto-oncogenic PTEN-targeting intron that cooperates with its host
gene MCM7 in transformation. Sci Signal. 3:ra292010.
|
|
29
|
Kumar M, Lu Z, Takwi AA, et al: Negative
regulation of the tumor suppressor p53 gene by microRNAs. Oncogene.
30:843–853. 2011.
|
|
30
|
Ahn YH, Gibbons DL, Chakravarti D, et al:
ZEB1 drives prometastatic actin cytoskeletal remodeling by
downregulating miR-34a expression. J Clin Invest. 122:3170–3183.
2012.
|
|
31
|
Bhat-Nakshatri P, Wang G, Collins NR, et
al: Estradiol-regulated microRNAs control estradiol response in
breast cancer cells. Nucleic Acids Res. 37:4850–4861. 2009.
|